Abstract
Background
Medulloblastoma (MB), the most common malignant pediatric brain tumor, lacks prognostic tools integrating clinical, molecular, and treatment-related characteristics for individualized management.
Methods
We developed machine learning models using multicenter data from 729 Chinese patients (2001-2023), of whom 509 were assigned to the training set and 220 to the testing set, and further validated the models on 201 patients from international MB consortia. To accommodate patients and researchers with varying datatypes, four application scenarios were established, including clinical-molecular-radiotherapy (CMR), clinical-molecular (CM), clinical-radiotherapy (CR), and clinical-only (CO).
Results
We construct four model scenarios and assess their predictive performance in the testing set: an XGBoost-based CMR model (incorporating 11 features, including molecular subgroup, radiotherapy dose, and key gene expression) with a C-index of 0.612; an XGBoost-based CM (C-index = 0.609); a GBM-based CR (C-index = 0.637); and a GBM-based CO (C-index = 0.635). External validation demonstrates robust performance, with radiotherapy and molecular data contributing significantly to enhanced efficacy. In addition, interactive web-based Shiny applications have been launched to facilitate dynamic risk assessment and treatment optimization.
Conclusions
By integrating multidimensional data, our framework enables the tailored prognostication and clinical decision to meet the multidimensional requirements of research and medicine.
Plain language summary
Medulloblastoma is a fast-growing brain tumor that is the most common brain tumor in children. Current clinical treatments are not designed for individualized patient management. In this study, we analyzed data from 729 people in China and 201 people internationally with medulloblastoma to develop four computational models for different scenarios. The models were able to predict the likely outcome for people with medulloblastoma. To facilitate clinical implementation, we have developed web-based applications that enable their use for risk assessment. This work could enable more personalized treatment decision-making and potentially improve the management of people with medulloblastoma clinically.
Similar content being viewed by others
Introduction
Medulloblastoma (MB) is the most common malignant pediatric tumor originating from the cerebellum, accounting for approximately 20% of pediatric brain tumor cases1. The 5-year survival rate varies by risk stratification: average-risk patients exhibit a 5-year survival rate of approximately 80%2,3,4, whereas high-risk patients have a 5-year survival rate of 60~65%5,6,7,8,9. Radiotherapy is a crucial element in the comprehensive treatment strategies for MB, enhancing survival rates in both standard-risk and high-risk patients4,6.
Predicting postoperative survival probabilities is critical for informing treatment strategies and patient counselling. Large clinical and molecular data actually make the traditional hypothesis-driven traditional statistical method less effective10,11. Machine learning (ML) algorithms have been employed to develop prognostic models for predicting mortality and endpoints, contributing to capturing complex relationships, and thereby improving predictive accuracy. ML applications in cancer survival analysis have significantly emerged, particularly in MB. Most of these studies rely on data from the Surveillance, Epidemiology, and End Results (SEER) database10,12,13, which offers comprehensive and regularly updated clinical information, including clinical and demographic variables. However, the SEER database has some drawbacks such as relatively limited included variables, limited representativeness of the population, and lack of long-term follow-up information, diagnostic and genetic profiles. In addition, some models may produce inaccurate predictions due to small sample sizes, thereby limiting their clinical applicability. Although some studies have developed models to predict the survival of MB patients, the previous tools often fail to fully account for radiation dose and critical genetic events. Therefore, it is imperative to integrate genetic events with radiotherapy strategy to establish more effective survival prediction methods and develop individualized radiotherapy regimens and follow-up strategies.
Irrespective of molecular subgroups, craniospinal irradiation (CSI) and posterior fossa tumor bed (PFTB) boost for primary MB are indispensable for potential cures14,15,16. However, long-term follow-up has revealed that intensive treatment was associated with an increased risk of late toxicity, resulting in treatment sequelae that severely affected patients’ quality of life17,18. Additionally, the NCCN guidelines also highlight that the molecular classification and associated risk stratification function ascrucial considerations during clinical practices19, playing a critical role in prognosis. Integrating molecular subgroups, gene signatures, and radiation dosages into a unified modeling framework is essential to yield clinically relevant and realistic prognostic predictions. However, non-specialists such as patients may not always have an accurate understanding of radiotherapy dosing, highlighting the need for inclusivity. Developing diversified applications tailored to different audiences is essential to fully realize the practical value of the study.
In the current study, we aim to introduce a predictive model for postoperative survival probabilities of MB patients in international multi-center longitudinal cohorts with six algorithms. Four scenario-specific models are developed to predict survival rates at 1, 3, 5 and 10 years. In addition to standard clinical variables, we incorporate transcriptomic prognostic signatures, such as MYC and OTX2, to enhance predictive power. The optimal model is translated as an interactive online calculator to facilitate utility for clinicians and patients. The final models of four scenarios effectively predict survival rates at 1, 3, 5, and 10 years, providing reliable prognostic information for clinical decision-making.
Methods
Study cohort
We retrospectively collected data on patients with MB who underwent surgery followed by radiotherapy and/or chemotherapy from September 2001 to April 2023. This multicenter cohort was led by Beijing Tiantan Hospital, Capital Medical University (Beijing, China). Written informed consent was obtained from all subjects, and the study was approved by the Institutional Review Board (IRB) of Beijing Tiantan Hospital (KY2022-133-03 and KY2023-130-01).
A total of 729 patients met the inclusion criteria: (1) a pathological diagnosis of MB, (2) availability of complete clinicopathological data, and (3) complete follow-up information. Patients were excluded if: (1) outcome or treatment information was absent (n = 277), (2) no treatment or only chemotherapy was administered (n = 18), or (3) treatment data were insufficient (n = 19).
Regarding treatment protocols, the extent of resection was assessed by contrast-enhanced MRI within 72 hours post-surgery, with a gross-total resection rate of 91.2% in this cohort. All patients received X-ray photon radiotherapy delivered with conventional fractionation (1.8 Gy per fraction) in accordance with NCCN guidelines. Intensity-modulated radiotherapy (IMRT) was predominantly used before 2012, whereas volumetric-modulated arc therapy (VMAT) was gradually adopted thereafter. The median interval between surgery and the initiation of radiotherapy was 38 days, and the median radiotherapy duration was 42 days. The median craniospinal irradiation (CSI) dose was 30.6 Gy, and the median posterior fossa/tumor bed (PFTB) boost dose was 55.8 Gy. No concurrent chemotherapy was administered during radiotherapy. Adjuvant chemotherapy was initiated 4 – 6 weeks after the completion of radiotherapy, with regimens determined according to risk stratification. Before 2010, radiotherapy was the primary treatment approach. Following 2010, the standard of care shifted predominantly to a combination of radiotherapy and chemotherapy. Given the potential time-related effects of this treatment transition, we incorporated treatment strategy as a factor when evaluating batch effects via principal component analysis (PCA). The PCA results showed samples clustered tightly, with no distinct separation related to assay platforms, protocols, or sequencing dates—confirming no obvious batch effects (Supplementary Fig. 1). This ensures no batch-related confounding in the molecular predictors. Therefore, the treatment strategies for RT and RT + CT were accounted for in the two clinical scenarios analyzed in our modeling.
Additionally, data from 201 patients enrolled in international MB consortia—comprising 32 medical centers from 16 countries led by Texas Children’s Hospital and The Hospital for Sick Children—were utilized for external validation20. Data variables in the external cohort were harmonized to ensure consistency with the training cohort.
Data collection
Data for variables evaluated in the current study were obtained from medical records and follow-up information, including demographics, histological and molecular pathology, molecular events, metastatic (M) stage, operation, treatment strategies, radiation dose, and survival follow-up. Demographic variables included age at diagnosis (infant: 0–3 years, child: 4–10 years, adolescent: 11–17 years, adult: Greater than or equal to 18 years) and sex. Histological and molecular pathology included molecular subgroup [WNT, SHH, Group_3 (Gr.3), and Group_4 (Gr.4)] and histological subgroup [classic (CMB), desmoplastic nodular (DNMB), MB extensive nodularity (MBEN), and large cell/anaplastic (LC/AMB)]. Operation resection extent included gross or nearly total resection (GTR/NTR; residual tumor ≤ 1.5 cm²) and subtotal resection (STR; residual disease > 1.5 cm²). Treatment strategies included radiotherapy only and concurrent radiotherapy and chemotherapy. M stage was divided into M0 and M+ stages. Radiotherapeutic strategies were subdivided into craniospinal irradiation (<36.0 Gy, 30.6 ~ < 36.0 Gy, and 36.0 Gy) and posterior fossa tumor bed boost (<55.8 Gy and ≥ 55.8 Gy). Molecular events included MYC, MYCN, OTX2, and GFI1 expression levels. The interval, measured in months from the surgery to the recorded date of death or the last follow-up time point if patients were alive, was defined as overall survival (OS). This study is strictly compliant with the “Guidance of the Ministry of Science and Technology (MOST) for the Review and Approval of Human Genetic Resources”. The multi-omics data used in this study have been formally filed for the external provision of human genetic resource information (Filing No. *BF2022062811508). To protect patient identities, all samples were coded, de-identified, and desensitized. The raw genomic, clinical, and imaging data are securely archived at the China National GeneBank DataBase (CNGBdb, https://db.cngb.org). Due to national regulations on human genetic resources and privacy protection, these data are available to qualified researchers for academic purposes upon reasonable request, subject to approval by the institutional Data Access Committee (DAC).
471 patients from an international cohort of 57 medical centers worldwide to externally validate the extrapolation of the predictive models20. Information on operative resection and radio-/chemotherapy was provided by the collaborative team at Texas Children’s Hospital. The data information and variables in the external database were harmonized and consistent with the training cohort. All studies within the consortia received approval from their respective IRBs, and all participants provided informed consent. The cohort flow chart of Chinese cohort and international cohort are shown in Fig. 1a, b, which demonstrates the study subject screening process. The same inclusion and exclusion criteria were applied to both databases. The final included study subjects had complete clinical data and comprehensive follow-up records.
a Cohort flow diagram of Chinese cohort. b Cohort flow diagram of international cohort.
During surgery, MB samples were procured from 303 chemo-/radiotherapy-naïve patients. Portions of tissues were embedded in paraffin, whereas the remaining tissues were preserved in liquid nitrogen. In total, 279 MB samples were classified according to DNA methylation using whole-genome bisulfite sequencing (WGBS). Next-generation sequencing (NGS) was conducted on 224 samples, and bulk RNA sequencing was performed on 252 samples. The samples were coded with a distinct patient identifier to uphold patient confidentiality. Multi-omics approaches were employed to refine molecular subgroup classification.
WGBS
For the Chinese cohort, all molecular profiling (DNA methylation arrays, NGS, and RNA-seq) underwent standardized library construction and sequencing at Beijing Genomics Institute (BGI Research, Chongqing), ensuring consistent technical quality. DNA was extracted, with 0.5% unmethylated λ phage DNA incorporated to evaluate bisulfite conversion efficiency, and the efficiency was confirmed to exceed 99.5% in all samples. DNA was fragmented to an average length of 250 bp using a Covaris M220 (RRID: SCR_019818) focused-ultrasonicator. Bisulfite conversion was carried out with the EZ DNA Methylation-Gold Kit, followed by library preparation using the Accel-NGS Methyl-Seq Kit. Libraries underwent quality verification on a 2100 Bioanalyzer, and sequencing was performed on the MGISEQ-2000 system to generate 151 bp paired-end reads.
Bulk RNA sequencing
Gene expression profiling was conducted on the BGISEQ-50 platform. Reads underwent alignment to the hg38 human reference genome, followed by quantification through Expectation-Maximization software. Mann–Whitney tests were employed for global integration analyses, contrasting the expression levels of amplified or deleted genes against those in copy-number-balanced regions. To control statistical errors, false discovery rate correction was independently implemented at each locus, with adjusted thresholds applied to minimize false-positive.
Model development and performance comparison
The data from the Chinese cohort were divided, with 70% utilized for training and 30% for testing, in order to avoid overfitting. Additionally, an external international set was used for external validation. We constructed four scenarios for predictor variables, each including different combinations of clinical, molecular, and radiotherapeutic information: Scenario 1 includes clinical, molecular, and radiotherapeutic information (CMR); Scenario 2 includes clinical and molecular information (CM); Scenario 3 includes clinical and radiotherapeutic information (CR); and Scenario 4 includes only clinical information (CO). These scenarios represent predictor variables with or without radiotherapy and molecular information, respectively. Six algorithms were applied to construct models to predict prognosis in each of four scenarios, including Cox Proportional Hazards (CoxPH), Random Survival Forests (RSF), Extreme Gradient Boosting (XGBoost), Elastic Net (ENET), DeepSurv, and Gradient Boosting Machine (GBM). These methodologies have previously been adapted into survival models proficiently in managing right-censored survival data. For non-WNT/non-SHH (Gr.3/4) MB patients in molecular scenarios, features were selected with reference to molecular risk stratification and incorporation of key molecular events based on previous studies21,22. Briefly, manual feature selection of molecular scenarios was performed based on clinical expertise, previous studies, and accessibility, referring to univariate COX analysis to select candidate predictors (p < 0.1). Model hyperparameters were optimized through grid search, utilizing a repeated 5-fold cross-validation method to maximize the concordance index (C-index) (Supplementary Table 5).
Several common evaluation metrics, such as the C-index, Brier score, and Integrated Brier Score (IBS), were involved to assess the performance of these models. Furthermore, the predictive accuracy for 5- and 10-years survival was assessed using the area under the receiver operating characteristic (ROC) curve (AUC) and calibration curves. Decision curve analysis (DCA) curves for 5- and 10-years periods were presented to evaluate the net benefits of different models. The generalizability of the final model was validated in international MB consortia.
Model explanation
For model explanations, we utilized SHAP (SHapley Additive exPlanations) values to explain how the final model works23. This interpretable approach provided two distinct forms of explanations: an overarching explanation of the model at the feature level and a personalized explanation at the individual level. The overarching explanation described the overall functionality of the model.
Statistics and reproducibility
The outcome indicators included the survival status and the overall survival time information. Data for categorical variables were processed using the one-hot encoding method. Continuous variables were presented as means ± standard deviation (SD), and categorical data as frequencies (percentages). For categorical variables, group comparisons were performed using the Chi-square test or Fisher’s exact test; for continuous variables, the Wilcoxon rank-sum test was utilized. We calculated the AUC to determine the model’s discriminative ability and select the optimal model. We used the DCA to evaluate the clinical utility of the prognostic model. The DeLong test was used for the comparison between AUC values. After selecting the best model, we developed a browser-based interactive application for computing individualized survival probabilities. Statistical analyses were carried out using R software (Version_4.4.0). A two-sided p < 0.05 was considered statistically significant.
Results
Baseline clinical information
This retrospective study involved 729 patients with MB in the Chinese cohort for the identification of the prediction model. These 729 patients were allocated into separate training and testing sets (Fig. 1a). The international cohort consisting of 201 patients was used as the external validation set (Fig. 1b). Supplementary Tables 1 and 2 summarize the comparison of clinicopathological and molecular characteristics of patients who received postoperative radiotherapy and/or chemotherapy among the training, testing, and external validation sets. The details of study design are displayed in Fig. 2. The design consists of five parts: data preparation, model development, optimal model selection, model interpretation, and an online calculator construction.
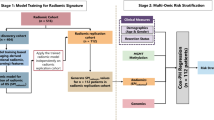
This figure illustrates the five core phases of the research, ranging from data preparation to the construction of clinical application tools. a Data preparation: flowcharts detailing the screening process for the Chinese discovery cohort (n = 1043) and the international consortium external validation cohort (n = 471). Based on exclusion criteria, 729 Chinese patients and 201 international patients with complete clinical data were ultimately included. The Chinese cohort was further partitioned into training and testing sets at a 7:3 ratio. b Model development: integration of clinical features, molecular features, and radiotherapy parameters. Six algorithms were employed to construct predictive models across four distinct application scenarios: CMR, CM, CR, and CO. c Selecting the best model: evaluation of model performance for 5- and 10-year survival predictions via ROC curves, AUC, calibration curves, and DCA. d Model interpretation: utilization of the SHAP method to rank feature contributions and provide individualized explanations for the final selected model. e Online calculator: translation of the optimized models into interactive web-based Shiny applications that support the input of clinical and molecular features to dynamically assess patient survival probabilities at various time points.
Among the 729 patients in the derivation cohort, molecular subgroups were identified in 424 (58.2%) patients, including 40 WNT-MB, 84 SHH-MB, 84 Gr.3-MB, and 216 Gr.4-MB. Patients were followed up for a median of 6.9 years (95% CI, 6.6 to 7.4). The 5-year and 10-year cumulative OS rates were 82.7% and 67.5%, respectively. The median age at diagnosis was 8 years (IQR, 6 to 11). Based on the presence of dissemination on cytology or gadolinium-enhanced craniospinal MRI, 110 patients (15.1%) had metastases (M + ). Histopathological classification was performed for 525 (72.0%) patients. Of those, the proportion of CMB (41.8%) was the highest, followed by DNMB (20.6%) and LC/AMB (5.6%), and the lowest percentage was found in MBEN (4.0%). GTR/NTR was achieved in 665 (91.2%) patients and STR in 64 (8.8%). A total of 533 cases (73.1%) received postoperative radiotherapy combined with chemotherapy, while the other 196 cases (26.9%) had only radiotherapy after the operation. The median CSI dose was 30.6 Gy (IQR, 28.8 to 36.0) with PFTB boosted to a median dose of 55.8 Gy (IQR, 54.0 to 55.8). According to the classification of the observed outcomes, 729 cases were divided into alive and deceased groups, of which 542 cases were survivors. The clinicopathological and molecular characteristics of the training and testing sets are listed in Supplementary Table 1 and 2. There was no statistically significant difference between the two sets for all the analyzed characteristics (all p > 0.05).
Supplementary Tables 1 and 2 provide the clinicopathological and molecular characteristics of 201 patients in the international cohort, which had a median follow-up of 5.8 years (Q1-Q3: 5.3–6.7). The median age was 8 (5.0, 12.6) years, and 135 (67.2%) were male. Molecular subgroups were identified in 200 patients (99.5%), including WNT-MB (n = 4), SHH-MB (n = 61), Gr.3-MB (n = 45), and Gr.4-MB (n = 90). The M+ stage was observed in 60 (29.9%) patients and the M0 stage in 141 (70.2%). Histological subgroups were identified in 179 patients (89.1%), including CMB (n = 121), DNMB (n = 27), MBEN (n = 6) and LC/AMB (n = 25). GTR/NTR was achieved in 181 (90.0%) patients and STR in 20 (10.0%). A total of 154 cases (76.6%) received postoperative radiotherapy in combination with chemotherapy, while the other 47 cases (23.4%) received postoperative radiotherapy alone. The median CSI dose was 24.0 Gy (IQR, 23.4 to 36.0) with PFTB boosted to a median dose of 54.0 Gy (IQR, 54.0 to 55.8).
Feature selection
In the CMR and CM scenarios, candidate predictors were selected using univariate Cox analysis (Fig. 3a), based on NCCN guidelines, previous studies, clinical expertise, and data accessibility. The expression levels of molecular markers MYC, MYCN, OTX2, and GFI1 (p = 0.026) were ultimately included in the analysis. Specific data for each variable from the training set were utilized to develop six algorithms—CoxPH, RSF, XGBoost, ENET, DeepSurv, and GBM—to predict 5- and 10-year prognoses in the four scenarios.
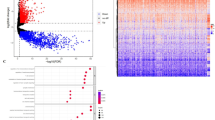
a Univariate COX regression feature selection for model construction. b Comparison of Area Under the Receiver Operating Characteristic curves (AUROCs) for predicting medulloblastoma prognosis using six algorithms within different time spans in testing set (left) and external validation set (right). c Receiver operating characteristic (ROC) curves with different algorithms for predicting 5- (left) and 10-year (right) survival predictions in medulloblastoma patients in testing set (n = 59). d ROC curves with Extreme Gradient Boosting algorithm (XGBoost) for predicting 5- (left) and 10-year (right) survival predictions in medulloblastoma patients in external validation set (n = 73). e Calibration plots of overall survival predictions for XGBoost algorithm in testing (left) (n = 59) and external validation set (right) (n = 73). f Decision curves for evaluating the clinical utility (net benefit) of XGBoost algorithm for 5- (left) and 10-year (right) survival predictions (n = 73).
Model performance comparison
Given the significant differences between WNT-MB and other molecular subgroups, we included interaction terms in the multivariate Cox analysis. As shown in Supplementary Tables 3 and 4, no significant interactions were found when PFTB boost dose and treatment strategies were analyzed separately with molecular subgroups. Therefore, we combined WNT with other subgroups to construct the predictive model.
In four scenarios (including CMR, CM, CR, and CO), we utilized six algorithms to predict patient prognosis. Their comparative performances are summarized in Supplementary Table 6, demonstrating that the XGBoost and GBM algorithms exhibited moderate yet consistent discrimination and favorable calibration across different scenarios. In the CMR scenario for Gr.3/4-MB in testing set, XGBoost algorithm (IBS = 0.122, C-index = 0.612) exhibited exceptional predictive performance for the prognosis of MB patients, achieving an AUC of 0.601 at 5 years and 0.734 at 10 years (Figs. 3b, 3c), followed by CoxPH algorithm (C-index = 0.578), RSF algorithm (C-index = 0.531), GBM algorithm (C-index = 0.510), ENET algorithm (C-index = 0.495), and DeepSurv algorithm (AUC = 0.465). The calibration plots in testing set presented in Fig. 3e illustrated that XGBoost algorithm maintained good consistency between its predictions and the observations for the 5- and 10-year OS rates (IBS = 0.122). Moving on to the CM scenario, XGBoost algorithm continued to show its superiority. It also boasted the best predictive performance (IBS = 0.131, C-index = 0.609), achieving an AUC of 0.618 at 5 years and 0.737 at 10 years, and the Time-dependent AUC values plot and ROC curves and for ML algorithms are presented in Supplementary Fig. 2a and 2b respectively. The calibration curves in Supplementary Fig. 2d show that the predicted probabilities and observed outcomes for the XGBoost algorithm were similar for the 5- and 10-year overall survival rates (IBS = 0.131). Moreover, in the CR scenario for the testing set, the GBM model demonstrated the best predictive performance in predicting the prognosis of MB patients (IBS = 0.114, C-index = 0.637), with 5-year AUCs of 0.662 and 10-year of 0.736 (Supplementary Fig. 3a, b). As shown in Supplementary Fig. 4a, b, of the abovementioned six algorithms, the GBM model fared best in terms of predicting the prognosis of MB patients in the CO scenario (IBS = 0.112, C-index = 0.635). GBM algorithm also showed good agreement between predicted and observed 5-year and 10-year OS rates in the CR and CO scenarios (Supplementary Figs. 3d, 4d). Overall, the superiority of XGBoost and GBM method was quantitatively validated through C-index, IBS metrics, and calibration plots across testing cohorts, demonstrating their potential for clinical practice.
Predictive performance of disease-free survival (DFS) models
To further address the clinical significance of tumor recurrence, we analyzed DFS, defined as the interval from treatment initiation to the first recurrence or death. In the testing set, the predictive performance across the four scenarios was modest, with C-indices of 0.657 for the CoxPH-based CMR model, 0.632 for the GBM-based CM model, 0.659 for the RSF-based CR model, and 0.643 for the RSF-based CO model. Furthermore, external validation was unfeasible due to the absence of recurrence data in the international cohort. Considering the limited predictive efficacy and the current lack of external generalizability, these DFS models do not yet meet the requirements for robust clinical application. Consequently, we did not develop an interactive web-based calculator for DFS at this stage (Supplementary Fig. 6).
External validation
To evaluate the generalizability of the model, we performed external validation using the XGBoost and GBM algorithms in the international MB cohort. When combining molecular information and radiotherapy strategy in the CMR scenario, as depicted in Fig. 3d, the externally validated ROC curve attained a 5-year AUC of 0.807 (95% CI: 0.685–0.930) and a 10-year AUC of 0.787 (95% CI: 0.610–0.963), which was comparable to that of the testing set (p = 0.12). For the external validation the CM scenario (Supplementary Fig. 2c), the XGBoost algorithm achieved a 5-year AUC of 0.692 (95% CI: 0.584–0.801) and a 10-year AUC of 0.729 (95% CI: 0.581–0.877), which was similar to the AUC obtained in the testing set (p = 0.29). Additionally, for the CR scenario, as illustrated in Supplementary Fig. 3c, the externally validated ROC curve demonstrated a 5-year AUC of 0.722 (95% CI: 0.608–0.836) and a 10-year AUC of 0.727 (95% CI: 0.547–0.906), comparable to that observed in the testing set (p = 0.96). For the CO scenario (Supplementary Fig. 4c), the GBM algorithm achieved a 5-year AUC of 0.730 (95% CI: 0.638–0.822) and a 10-year AUC of 0.712 (95% CI: 0.591–0.833), which was not statistically distinguishable from the AUC obtained in testing set (p = 0.92).
Model calibration is depicted in the external validation set (Fig. 3e, Supplementary Figs. 2d, 3d, and 4d), which show favorable consistency between the predictions and the observed outcomes of the four scenarios. Collectively, the XGBoost and GBM algorithms in the four scenarios showed favorable calibration and consistent performance in external validations.
The DCA further demonstrated the predictive and clinical application potential for XGBoost and GBM algorithms in the four scenarios. In the CMR and CO scenarios, the DCA presented in Fig. 3f and Supplementary Fig. 4e revealed the XGBoost and GBM algorithms each perform optimally across the wide threshold range for predicting the 5- and 10-year OS rates in their corresponding scenarios. Furthermore, regarding the clinical applicability of the CM and CR scenarios, the XGBoost and GBM algorithms achieved a robust net benefit only within a narrow range of threshold probabilities (Supplementary Figs. 2e and 3e).
Model explanation
Given that the SHAP method interprets the final model output by calculating each variable’s contribution to prediction, we employed this method to analyze the results of the XGBoost algorithm. We evaluated the feature-importance rankings based on SHAP values for the CMR scenario(Fig. 4a). In this plot, the contributions of each indicator to the prediction model were assessed using the average SHAP values and presented in descending order as the five most essential features: GFI1 expression level, M stage, Subgroup, MYCN expression level, and MYC expression level. We performed dot plot analysis to uncover the direction and strength of the influence of each feature on model prediction. Features, such as M+ stage and high MYC and GFI1 expression, significantly resulted in the poor prognosis, further underscoring the significance of molecular events in predictive modeling (Fig. 4b). The top 5 most influential variables in the summary plot of the CM scenario were roughly the same as the CMR scenario (Supplementary Fig. 5a). In CR and CO scenarios, histological and molecular subgroups ranked as the top two most important variables (Supplementary Fig. 5b, c). Notably, under these two clinical scenarios, the M stage significantly increases the risk of poor prognosis, while a higher PFTB dose (≥ 55.8 Gy) and RT + CT can dramatically reduce this risk (Supplementary Fig. 5b, c).
a Visualization of SHAP summary bar plot depicting the contribution ranking of the XGBoost algorithm’s features in CMR scenario. The bars represent the importance of the variables and their overall contribution to the model prediction. b SHAP summary dot plot. Each patient gets one dot per feature in the model, with the dot color (dark blue = high, light blue = low) showing the actual feature value. Dots stack vertically to show density.
Implementation of the web calculator
The XGBoost-based survival predictor was integrated into a web application for utilization in clinical scenarios. To improve clinical practicality, two interactive web-based Shiny apps by the CMR and CM scenarios were created: https://prognosticmodel.shinyapps.io/Scenario1_CMR/ for cases where the radiotherapeutic dose information was available and https://prognosticmodel.shinyapps.io/Scenario2_CM/ for cases where it was not. The web applications for the CR and CO scenarios are accessible online at the following links: https://prognosticmodel.shinyapps.io/Scenario3_CR/ and https://prognosticmodel.shinyapps.io/Scenario4_CO/. Practical demonstration using a representative case in CMR scenario. By inputting the actual values of the features required for the scenarios, the application can automatically predict the survival rates and clinical risk groups of individual patients with MB. In this case, users input complete data entry by responding to 11 queries and the calculator could predict survival rates at different time spans and the importance of variables (Fig. 5a, b). The results in Fig. 5c showed that LC/AMB, Gr.3, and MYC high level were associated with poorer prognosis, while CSI dose contributed positively to the favorable prognosis.
a Practical demonstration using a representative case in CMR scenario. Users input complete data entry by responding to 11 queries. b Survival line chart illustrating predicted survival probabilities at multiple key future time points (12, 36, 60, 80, 100, and 120 months) from the current moment. The y-axis indicates survival probability (%), while the x-axis represents time points. Specific survival values are displayed at the top of the graph. c Feature contribution plot based on SHAP values. Positive SHAP values indicate an increased risk of death, whereas negative values suggest a protective effect.
Discussion
Although researchers have proposed numerous prediction models, the current study presents using an ML-based prognostic model that integrates radiotherapy dose and molecular events to predict survival outcomes in MB patients, and this model was based on the XGBoost algorithm using a multi-center follow-up database. Our study revealed that XGBoost outperformed other models in accurately predicting the survival outcomes of MB patients in four scenarios. Among the four scenarios, scenarios involving radiotherapy information were superior to those without radiotherapy. In the CMR and CM scenarios testing sets, the C-indexes were 0.612 (with radiotherapy) and 0.609 (without), respectively. In the CR and CO scenarios, they were 0.637 (with radiotherapy) and 0.635 (without), respectively. These findings confirm the model’s high predictive performance and emphasize the significance of radiotherapy dose as a key variable in prognostic assessment. Moreover, the model offers better recommendations for radiotherapy doses tailored to MB cases and provides an intuitive understanding of how various treatment strategies influence patient survival.
In this study, incorporating radiotherapy dose into the prognostic prediction model for MB represented a key innovation that addresses a critical gap in molecular identification and therapeutic planning. Following the introduction of molecular subgroups, dose reduction in radiotherapy for low-risk groups has become a prevailing trend24. Current MB prognostic models are primarily based on radiomics, clinical information, or transcriptomic data10,12,25,26. However, these models generally fail to account for the impact of radiotherapy dose, a fundamental therapeutic variable. The novelty of incorporating radiotherapy dose lies in its ability to capture not only the direct effects of treatment intensity on prognosis but also to surpass traditional single-variable prediction methods, providing clinicians with more precise guidance for radiotherapy dose optimization and demonstrating its breakthrough potential in personalized treatment of MB.
In some countries, medical centers still lack the ability to determine molecular subgroups through sequencing-based approaches. In order to facilitate the use of the model calculator, undefined molecular or histology subgroup data were not excluded in the modeling process, which was convenient for patients and their doctors. Additionally, since patients may not easily understand or obtain radiotherapy dose indicators, we designed four scenarios with/without radiotherapy and molecular information to improve usability. Since GTR or NTR is the standard for clinical risk group classification in the NCCN guidelines, the excision variable was added to the clinical scenario. Consistent with previous studies27, our cohort analysis found that the extent of tumor resection had no significant impact on prognosis, which led to its exclusion as a predictive variable in CMR and CM scenario construction.
The evaluation of feature significance revealed the contribution to prediction accuracy of the model. Our findings were in close alignment with prior studies, which had consistently identified molecular markers, radiation, and chemotherapy as critical determinants of prognosis. Since the advent of genetic examination and therapy, molecular characteristics have become the most important prognostic factors, second only to treatment strategies, in various malignancies, including gliomas28, breast cancer29, and hematological cancers30. For instance, Pfister et al., in their analysis of MB patients, emphasized the significance of genomic amplifications of MYC or MYCN as the important prognostic factor31. As Northcott PA et al. suggested, GFI1 is abnormally overexpressed via the mechanism of enhancer hijacking in Gr.3 and Gr.4 MB driving tumor progression and influencing prognosis. However, GFI1 has rarely been incorporated into prognostic models for brain tumors32. In our study, the influence of high GFI1 expression levels on prognosis was evident in the CMR and CM scenarios. Additionally, Kuo et al.’s study on MB patients from SEER database identified radiotherapy and chemotherapy as significant factors in predicting OS10. Similarly, Qaddoumi et al. highlighted the radiotherapy, tumor grade, and surgical methods as the independent adverse prognostic factors in pediatric glioma using SEER database33. Although these studies included radiotherapy variables, they did not explore the effect of detailed radiation doses on the prognosis of patients with brain tumors. Notably, a higher PFTB dose (≥ 55.8 Gy) significantly reduced the risk of poor prognosis in our study, highlighting the predictive value of increasing the PFTB boost dose in CMR and CM scenarios. Extended follow-up analysis demonstrated that intensive treatments substantially affected the quality of life in pediatric patients17,18, with children under the age of 7 being particularly vulnerable to neurological impairments due to radiotherapy34. Overall, our model provides a comprehensive framework that enables clinicians and patients to systematically assess prognosis during the development of treatment plans.
As an integrating multiple weak-classifier learning method, XGBoost algorithm isoptimized based on gradient-boosting decision trees, making it especially suitable for large datasets and complex features. Recently, prediction models based on the XGBoost algorithm have been widely used in the medical field, demonstrating excellent performance in multiple areas, such as breast cancer prognosis35 and in-hospital mortality prediction36,37. Compared with the traditional Cox regression algorithm, the XGBoost model can effectively capture the nonlinear relationship and improve generalization ability. Furthermore, the XGBoost model is robust to high-dimensional and highly variable datasets, which can further reduce the noise within the datasets. Similarly, XGBoost and GBM algorithms were utilized under four scenarios to develop the well-calibrated and explainable prediction model for prognosis assessment of MB patients.
By analyzing the involved variables, certain limitations of the current mode include the lack of key imaging features such as tumor location and size, MRI contrast enhancement intensity, as well as the degree of brainstem invasion. Additionally, critical pathological features, including the Ki-67 index or mitotic count, as well as clinical indicators reflecting postoperative functional status, such as assessments of speech, balance, mutism, respiration, and swallowing function were also omitted. The Ki-67 index has emerged as a critical prognostic indicator for various types of cancer38,39. We attempted to incorporate this metric into the current model; however, due to the limited sample size and the absence of this indicator in the external validation set, it was ultimately excluded from the final analysis. Future studies incorporating higher-resolution datasets encompassing these variables will be essential for refining predictive accuracy and guiding longitudinal data follow-up.
Although DFS is critical for informing surveillance strategies in MB, the predictive performance of our DFS models was slightly lower than that of the OS models. This discrepancy may stem from the inherent complexity of accurately recording recurrence events and the limited number of such events in retrospective cohorts. Furthermore, the lack of recurrence data in the international consortia precluded external validation of the DFS models, limiting their current generalizability. Consequently, OS remains the primary endpoint of this study, while future efforts will focus on aggregating multicenter recurrence data to refine and validate DFS-based prognostic tools.
In conclusion, we developed XGBoost and GBM models with better predictive performance to predict the prognosis of MB patients and validated their potential as clinically reliable tools. Based on the interactive applications, this study addresses a gap in previous models that overlooked treatment-specific details and provides additional assistance to radiation oncologists for strategy determination.
Data availability
The WGBS dataset is deposited in the China National Genomics Bank (CNGB) with the accession code (CNP0006196). The NGS data are deposited in CNGB with the accession code (CNP0006197). The total bulk RNA sequencing data are deposited in CNGB with the accession code (CNP0006198). The data will be made available upon reasonable request. The source data supporting the findings of this study are available in Figshare40. Access to the international cohort data can be requested from the original data custodians at Texas Children’s Hospital20.
References
Leece, R. et al. Global incidence of malignant brain and other central nervous system tumors by histology, 2003-2007. Neuro Oncol. 19, 1553–1564 (2017).
Ramaswamy, V. & Taylor, M. D. Medulloblastoma: from myth to molecular. J. Clin. Oncol. 35, 2355–2363 (2017).
Taylor, R. E. et al. Results of a randomized study of preradiation chemotherapy versus radiotherapy alone for nonmetastatic medulloblastoma: The International Society of Paediatric Oncology/United Kingdom Children’s Cancer Study Group PNET-3 Study. J. Clin. Oncol. 21, 1581–1591 (2003).
Packer, R. J. et al. Phase III study of craniospinal radiation therapy followed by adjuvant chemotherapy for newly diagnosed average-risk medulloblastoma. J. Clin. Oncol. 24, 4202–4208 (2006).
Gajjar, A. et al. Risk-adapted craniospinal radiotherapy followed by high-dose chemotherapy and stem-cell rescue in children with newly diagnosed medulloblastoma (St Jude Medulloblastoma-96): long-term results from a prospective, multicentre trial. Lancet Oncol. 7, 813–820 (2006).
Tarbell, N. J. et al. High-risk medulloblastoma: a pediatric oncology group randomized trial of chemotherapy before or after radiation therapy (POG 9031). J. Clin. Oncol. 31, 2936–2941 (2013).
Gandola, L. et al. Hyperfractionated accelerated radiotherapy in the Milan strategy for metastatic medulloblastoma. J. Clin. Oncol. 27, 566–571 (2009).
von Bueren, A. O. et al. Treatment of children and adolescents with metastatic medulloblastoma and prognostic relevance of clinical and biologic parameters. J. Clin. Oncol. 34, 4151–4160 (2016).
Esbenshade A. J. et al. A Phase II feasibility study of oral etoposide given concurrently with radiotherapy followed by dose intensive adjuvant chemotherapy for children with newly diagnosed high-risk medulloblastoma (protocol POG 9631): A report from the Children’s Oncology Group. Pediatr Blood Cancer, 2017, 64.
Kuo, C. C. et al. An online calculator using machine learning for predicting survival in pediatric patients with medulloblastoma. J. Neurosurg. Pediatr. 33, 85–94 (2024).
Chen, K. et al. Two machine learning methods identify a metastasis-related prognostic model that predicts overall survival in medulloblastoma patients. Aging 12, 21481–21503 (2020).
Sun, M., Sun, J. & Li, M. Deep learning models for predicting the survival of patients with medulloblastoma based on a surveillance, epidemiology, and end results analysis. Sci. Rep. 14, 14490 (2024).
Li, T. et al. A novel survival nomogram for stage I-III acral lentiginous melanoma patients, based on the SEER database and a multi-center external validation cohort. Holist. Integr. Oncol. 3, 23 (2024).
Mynarek, M. et al. MEDB-04. Young children with metastatic medulloblastoma: frequent requirement for radiotherapy in children with non-WNT/non-SHH medulloblastoma despite highly intensified chemotherapy–results of the MET-HIT2000-BIS4 trial [M]. Neuro Oncol. 24, i104 (2022).
Cohen, K. J. et al. A pilot study omitting radiation in the treatment of children with newly diagnosed Wnt-activated medulloblastoma. Clin. Cancer Res 29, 5031–5037 (2023).
Hovestadt, V. et al. Medulloblastomics revisited: biological and clinical insights from thousands of patients. Nat. Rev. Cancer 20, 42–56 (2020).
Saraf, A. et al. Long-term outcomes and late toxicity of adult medulloblastoma treated with combined modality therapy: a contemporary single-institution experience. Neuro Oncol. 24, 2180–2189 (2022).
Salloum, R. et al. Late morbidity and mortality among medulloblastoma survivors diagnosed across three decades: a report from the childhood cancer survivor study. J. Clin. Oncol. 37, 731–740 (2019).
Gajjar, A. et al. Pediatric central nervous system cancers, version 2.2025, NCCN Clinical Practice Guidelines In Oncology. J. Natl. Compr. Canc Netw. 23, 113–130 (2025).
Cavalli, F. M. G. et al. Intertumoral heterogeneity within medulloblastoma subgroups. Cancer Cell 31, 737–54.e6 (2017).
Gajjar, A. et al. Outcomes by clinical and molecular features in children with medulloblastoma treated with risk-adapted therapy: results of an International Phase III Trial (SJMB03). J. Clin. Oncol. 39, 822–835 (2021).
Sharma, T. et al. Second-generation molecular subgrouping of medulloblastoma: an international meta-analysis of Group 3 and Group 4 subtypes. Acta Neuropathol. 138, 309–326 (2019).
Lundberg S. M., Lee S.-I. A unified approach to interpreting model predictions. Advances in neural information processing systems, 30 (2017).
Seidel C. et al. Radiotherapy in medulloblastoma-evolution of treatment, current concepts and future perspectives. Cancers, 13 (2021).
Yan, J. et al. Incremental prognostic value and underlying biological pathways of radiomics patterns in medulloblastoma. EBioMedicine 61, 103093 (2020).
Li, C. et al. Construction and Validation of a 13-gene signature for prognosis prediction in medulloblastoma. Front. Genet. 11, 429 (2020).
Thompson, E. M. et al. Prognostic value of medulloblastoma extent of resection after accounting for molecular subgroup: a retrospective integrated clinical and molecular analysis. Lancet Oncol. 17, 484–495 (2016).
Eckel-Passow, J. E. et al. Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N. Engl. J. Med. 372, 2499–2508 (2015).
Jiao, Y. et al. Lactylation-related gene signature for prognostic prediction and immune infiltration analysis in breast cancer. Heliyon 10, e24777 (2024).
Eckardt, J. N. et al. Prediction of complete remission and survival in acute myeloid leukemia using supervised machine learning. Haematologica 108, 690–704 (2023).
Pfister, S. et al. Outcome prediction in pediatric medulloblastoma based on DNA copy-number aberrations of chromosomes 6q and 17q and the MYC and MYCN loci. J. Clin. Oncol. 27, 1627–1636 (2009).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Qaddoumi, I., Sultan, I. & Gajjar, A. Outcome and prognostic features in pediatric gliomas: a review of 6212 cases from the Surveillance, Epidemiology, and End Results database. Cancer 115, 5761–5770 (2009).
Ris, M. D. et al. Intellectual outcome after reduced-dose radiation therapy plus adjuvant chemotherapy for medulloblastoma: a Children’s Cancer Group study. J. Clin. Oncol. 19, 3470–3476 (2001).
Fang, Y. et al. Mitochondrial-related genes as prognostic and metastatic markers in breast cancer: insights from comprehensive analysis and clinical models. Front. Immunol. 15, 1461489 (2024).
Mao, B. et al. Machine learning for the prediction of in-hospital mortality in patients with spontaneous intracerebral hemorrhage in intensive care unit. Sci. Rep. 14, 14195 (2024).
Yang, J. et al. A machine learning model for predicting in-hospital mortality in Chinese patients with ST-segment elevation myocardial infarction: findings from the China Myocardial Infarction Registry. J. Med Internet Res 26, e50067 (2024).
Zhao, F. et al. Prognostic value of Ki-67 index in adult medulloblastoma after accounting for molecular subgroup: a retrospective clinical and molecular analysis. J. Neurooncol. 139, 333–340 (2018).
Petrelli, F. et al. Prognostic value of different cut-off levels of Ki-67 in breast cancer: a systematic review and meta-analysis of 64,196 patients. Breast Cancer Res. Treat. 153, 477–491 (2015).
Data from: An interpretable machine learning model for predicting prognosis of medulloblastoma integrating genetic and clinical features. Figshare. https://doi.org/10.6084/m9.figshare.31016692.
Acknowledgements
This work was supported by the Natural Science Foundation of Beijing Joint Project (L2510021 and L232079), Natural Science Foundation of Beijing for Distinguished Young Scholars (JQ24040), National Natural Science Foundation of China (82273343, 82204127, and 82573715), Beijing Medical and Health Science and Technology Promotion “UPlan” Project (YC202501QX0950), Capital Medical University Startup (3100/12100208), National Key Research and Development Program of China (2022ZD0210100), Capital Medical Foundation for Excellent Young Scholars (KCB2304), and Capital Medical University Beijing Tiantan Hospital Clinical Key Research Project (2025-B23).
Author information
Authors and Affiliations
Contributions
Yu Tian, Hailong Liu, and Jiao Zhang for study concept and design, obtainment of funding, and study supervision. Yu Su and Kaiwen Deng for study concept and design, collation of data, statistical analysis, interpretation of data, and drafting of the manuscript. Xuan Chen for DNA methylation and RNA expression analysis. Yu Tian, Hailong Liu, and Jiao Zhang for manuscript edition. Xiaoguang Qiu, Hailong Liu, Tao Jiang, Chunde Li, Kaiwen Deng, Fei Liu, Zhaoyang Feng, and Dongyang Wang for management of patients, acquisition of data, and collation of data. Craig Daniels, Hyun Yong Koh, Ricardo Daniel Gonzalez, Hiromichi Suzuki, Tsubasa Miyauchi, and Michael D. Taylor for the acquisition of data. Rui Chen, Shuaicheng Li, Jiankang Li, and Wei Wang for improvements in methodology and technical support. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Anahita Fathi Kazerooni and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Su, Y., Deng, K., Chen, X. et al. An interpretable machine learning model for predicting prognosis of medulloblastoma integrating genetic and clinical features. Commun Med 6, 134 (2026). https://doi.org/10.1038/s43856-026-01454-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-026-01454-4