Abstract
Background
Healthcare Artificial Intelligence (AI) offers transformative potential but often inherits biases from training data, worsening disparities. While bias mitigation has focused on structured data, mental health relies on unstructured clinical notes, where linguistic differences and data sparsity pose challenges. This study aims to detect and reduce non-biological textual bias in AI models supporting pediatric mental health screening.
Methods
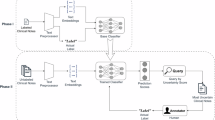
We analyzed ~20,000 pediatric anxiety cases and matched controls (ages 5-15) from Cincinnati Children’s Hospital records, where gender prevalence transitions from male-dominant in early childhood to female-dominant in adolescence. Anxiety prediction models were fine-tuned using a Transformer architecture optimized for computational efficiency. Classification parity across sex subgroups was evaluated, and we also verified that the model relied on clinically relevant words (using the LIME tool). Bias was mitigated through informative term filtering and systematic gender-biased text replacement.
Results
Here, we show systematic under-diagnosis of female adolescents, with 4% lower accuracy and 9% higher false-negative rates compared to male patients. Notes for male patients are on average 500 words longer, and linguistic similarity metrics reveal distinct word distributions between sexes. Applying our de-biasing framework reduces diagnostic bias by up to 27%, improving equity in model performance.
Conclusions
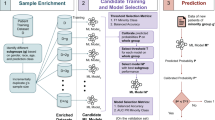
We develop and evaluate a data-centric de-biasing framework to address gender-based disparities in clinical text arising from non-biological differences, such as reporting practices and documentation styles. Our method selectively de-biases data by neutralizing biased language and normalizing information density while preserving clinically relevant content. Further validation across different models is essential before clinical deployment.
Plain language summary
Artificial Intelligence (AI) is increasingly used in healthcare, but it can unintentionally reflect biases found in medical records. These biases may lead to unfair predictions, especially in mental health, where information comes from written notes rather than tabular data. Our study looks at anxiety in children and teenagers and explores whether differences in how doctors write notes for boys and girls affect AI predictions. We analyzed thousands of records and found that girls were more likely to be underdiagnosed. To address this, we developed a method that removes biased language and balances information without losing important clinical details. This approach improves fairness in AI predictions, but more testing is needed before it can be used in real-world healthcare.
Similar content being viewed by others
Data availability
The textual notes used in this study are derived from sensitive clinical sources and cannot be shared publicly due to patient confidentiality and institutional data-sharing agreements. Access to the data may be granted through collaboration, subject to appropriate governance and ethical approvals. Researchers interested in accessing the data should contact the corresponding author.
Code availability
The original version of our code is tightly linked to confidential data pipelines and cannot be shared in its raw form. To ensure reproducibility without compromising patient privacy, we provide a publicly available version that has been carefully sanitized to remove sensitive components while closely replicating the functionality used in this study. The repository is accessible at: https://github.com/julia-ive/bias-pediatric-anxiety50. The code is compatible with Python 3.12 and associated libraries. To ensure reproducibility without compromising patient confidentiality, the repository includes synthetic data that mimics the structure and characteristics of the original dataset.
References
COVID-19 Mental Disorders Collaborators Global prevalence and burden of depressive and anxiety disorders in 204 countries and territories in 2020 due to the COVID-19 pandemic. Lancet 398, 1700–1712 (2025).
Racine, N. et al. Global prevalence of depressive and anxiety symptoms in children and adolescents during COVID-19: a meta-analysis. JAMA Pediatrics 175, 1142–1150 (2021).
Timmons, A. C. A call to action on assessing and mitigating bias in artificial intelligence applications for mental health. Perspect. Psychol. Sci. 18, 1062 (2022).
Behrens, B., Swetlitz, C., Pine, D. S. & Pagliaccio, D. The screen for child anxiety related emotional disorders (SCARED): informant discrepancy, measurement invariance, and test-retest reliability. Child psychiatry Hum. Dev. 50, 473–482 (2019).
Strawn, J. R., Lu, L., Peris, T. S., Levine, A. & Walkup, J. T. Research review: Pediatric anxiety disorders - what have we learnt in the last 10 years?. J. child Psychol. psychiatry, allied Discip. 62, 114–139 (2021).
Tulisiak, A. K. et al. Antidepressant prescribing by pediatricians: A mixed-methods analysis. Curr. Probl. Pediatr. Adolesc. Health Care 47, 15–24 (2017).
Golden, G. et al. Applying artificial intelligence to clinical decision support in mental health: What have we learned? Health Policy Technol. 13, 100844 (2024).
Perlman, K. et al. Development of a differential treatment selection model for depression on consolidated and transformed clinical trial datasets. Transl. Psychiatry 14, 263 (2024).
Hou, J. & Wang, L. L. Explainable AI for clinical outcome prediction: A survey of clinician perceptions and preferences. AMIA Summits Transl. Sci. Proc. 2025, 215–224 (2025).
Zhang, T., Schoene, A. M., Ji, S. & Ananiadou, S. Natural language processing applied to mental illness detection: a narrative review. npj Digital Med. 5, 1–13 (2022).
Ji, Y. et al. Mitigating the risk of health inequity exacerbated by large language models. npj Digit. Med. 8, 246 (2025).
Coley, R. Y., Johnson, E., Simon, G. E., Cruz, M. & Shortreed, S. M. Racial/ethnic disparities in the performance of prediction models for death by suicide after mental health visits. JAMA Psychiatry 78, 726–734 (2021).
Angwin, L. J. M. S., J. & Kirchner, L. Machine bias: There’s software used across the country to predict future criminals. and it’s biased against blacks. ∣ benton institute for broadband & society https://www.benton.org/headlines/machine-bias-theres-software-used-across-country-predict-future-criminals-and-its-biased (2016).
Chouldechova, A. & Roth, A. A snapshot of the frontiers of fairness in machine learning. Commun. ACM 63, 82–89 (2020).
Corbett-Davies S., Gaebler J., D., Nilforoshan H., Shroff R. & Goel S. The measure and mismeasure of fairness. J. Mach. Learn. Res. 24, 14730–14846 (2023).
Caton, S. & Haas, C. Fairness in machine learning: a survey. ACM Comput. Surv. 56, 1–38 (2024).
Bolukbasi, T., Chang, K.-W., Zou, J., Saligrama, V. & Kalai, A. Man is to computer programmer as woman is to homemaker? debiasing word embeddings. In Proc. of the 30th International Conference on Neural Information Processing Systems (NIPS). 4356–4364 (Red Hook, NY, USA, 2016).
Tokpo, E. K., Delobelle, P., Berendt, B. & Calders, T. How far can it go? On intrinsic gender bias mitigation for textclassification. In: Proc. of the 17th Conference of the European Chapter of the Association for Computational Linguistics (EACL) 3418–3433 (Stroudsburg, PA, USA,, 2023).
Raza, S., Garg, M., Reji, D. J., Bashir, S. R. & Ding, C. Nbias: a natural language processing framework for bias identification in text. Expert Syst. Appl. 237, 121542 (2024).
Fang, T., Lu, N., Niu, G. & Sugiyama, M. Rethinking importance weighting for deep learning under distribution shift. In Proc. of the 34th International Conference on Neural Information Processing Systems (NIPS), 11996–12007 (Red Hook, NY, USA, 2020).
Clark, C., Yatskar, M. & Zettlemoyer, L. Don’t take the easy way out: Ensemble based methods for avoiding knowndataset biases. In Proc. of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9thInternational Joint Conference on Natural Language Processing (EMNLP-IJCNLP), 4069–4082 (Stroudsburg, PA, USA, 2019).
Blagus, R. & Lusa, L. Smote for high-dimensional class-imbalanced data. BMC Bioinformatics 14, 1–16 (2013).
He, H., Bai, Y., Garcia, E. A. & Li, S. Adasyn: adaptive synthetic sampling approach for imbalanced learning. In Proc. International Joint Conference on Neural Networks 1322–1328 (IEEE, 2008).
Liang, P. P. et al. Towards Debiasing Sentence Representations. In Proceedings of the 58th Annual Meeting of theAssociation for Computational Linguistics (ACL), 5502–5515 (Stroudsburg, PA, USA, 2020).
Beutel, A., Chen, J., Zhao, Z. & Chi, E. H. Data decisions and theoretical implications when adversarially learning fair representations. https://doi.org/10.48550/arXiv.1707.00075 (2017).
Li, Y., Baldwin, T. & Cohn, T. Towards Robust and Privacy-preserving Text Representations. In Proc. of the 56th Annual Meeting of the Association for Computational Linguistics (ACL), (Melbourne, Australia, 2018).
Woodworth, B., Gunasekar, S., Ohannessian, M. I. & Srebro, N. Learning Non-Discriminatory Predictors. In Proc. of the 2017 Conference on Learning Theory (PMLR) Vol. 65, 1920–1953 (2017).
Berk, R. et al. A convex framework for fair regression. arXiv.org https://doi.org/10.48550/arXiv.1706.02409 (2017).
Vaidya, A. et al. Demographic bias in misdiagnosis by computational pathology models. Nat. Med. 30, 1174–1190 (2024).
Guo, C., Pleiss, G., Sun, Y. & Weinberger, K. Q. On calibration of modern neural networks. In Proc. of the 34th International Conference on Machine Learning (PMLR), 1321–1330 (2017).
Kessler, R. C. Epidemiology of women and depression. J. Affect. Disord. 74, 5–13 (2003).
Bird, S., Klein, E. & Loper, E. Natural Language Processing With Python: Analyzing Text With The Natural Language Toolkit (O’Reilly Media, Inc., 2009).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Li, Y., Wehbe, R. M., Ahmad, F. S., Wang, H. & Luo, Y. A comparative study of pretrained language models for long clinical text. J. Am. Med. Inform. Assoc. 30, 340–347 (2023).
Wolf, T. et al. Transformers: State-of-the-art natural language processing. In Proc. of the 2020 Conference on EmpiricalMethods in Natural Language Processing: System Demonstrations (EMNLP), 38–45 (Stroudsburg, PA, USA, 2020).
Johnson, A. E. W. et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci. Data 10, 1 (2023).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. In Proc. of the 2019 NAACL-HLT, 4171–4186 (New Orleans, Louisiana, USA, 2019).
Chopra, S., Agarwal, P., Ahmed, J., Biswas, S. S. & Obaid, A. J. Roberta and BERT: revolutionizing mental healthcare through natural language. SN Comput. Sci. 5, 1–12 (2024).
Hossain, M. M., Hossain, M. S., Mridha, M. F., Safran, M. & Alfarhood, S. Multi task opinion enhanced hybrid BERT model for mental health analysis. Sci. Rep. 15, 3332 (2025).
Chen, Q. et al. Benchmarking large language models for biomedical natural language processing applications and recommendations. Nat. Commun. 16, 3280 (2025).
Loshchilov, I. & Hutter, F. Decoupled weight decay regularization. In Proc. 7th International Conference on Learning Representations (2017).
Feldman, M., Friedler, S. A., Moeller, J., Scheidegger, C. & Venkatasubramanian, S. Certifying and Removing DisparateImpact. In Proc. of the 21th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, (New York, NY, USA 2015).
Chouldechova, A. Fair prediction with disparate impact: a study of bias in recidivism prediction instruments. Big Data 5, 153–163 (2017).
Kurita, K., Vyas, N., Pareek, A., Black, A. W. & Tsvetkov, Y. Measuring bias in contextualized word representations. In Proc.of the First Workshop on Gender Bias in Natural Language Processing ACL, 166–172 (Florence, Italy, 2019).
Ribeiro, M. T., Singh, S. & Guestrin, C. Why should I trust you?: Explaining the predictions of any classifier. In Proc. of the22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, (New York, NY, USA, 2016).
Shapley, L. S. A value for n-person games. Contribution to the Theory of Games (Princeton University Press, 1953).
Rousseau, A. XenC: an open-source tool for data selection in natural language processing. Prague Bull. Math. Linguist. 100, 73–82 (2013).
Kiritchenko, S. & Mohammad, S. Examining gender and race bias in two hundred sentiment analysis systems. In Proc. of the Seventh Joint Conference on Lexical and Computational Semantics, 43–53 (New Orleans, Louisiana, USA, 2018).
Qi, P., Zhang, Y., Zhang, Y., Bolton, J. & Manning, C. D. Stanza: A Python natural language processing toolkit for many human languages. In Proc. of the 58th Annual Meeting of the Association for Computational Linguistics: System Demonstrations (ACL), 101–108 (Stroudsburg, PA, USA, 2020).
julia ive. julia-ive/bias-pediatric-anxiety: Code for detecting and mitigating demographic bias in pediatric mental health text https://doi.org/10.5281/zenodo.18359989 (2026).
Zhang, Y., Zhang, Y., Qi, P., Manning, C. D. & Langlotz, C. P. Biomedical and clinical English model packages for the Stanza Python NLP library. J. Am. Med. Inform. Associat. 28, 1892–1899. (2021).
Obermeyer, Z., Powers, B., Vogeli, C. & Mullainathan, S. Dissecting racial bias in an algorithm used to manage the health of populations. Science 366, 447–453 (2019).
Guevara, M. Large language models to identify social determinants of health in electronic health records. npj Digit. Med. 7, 6 (2024).
Seedat, N., Imrie, F. & Schaar, M. V. D. Navigating data-centric artificial intelligence with dc-check: Advances, challenges, and opportunities. IEEE Trans. Artif. Intell. 5, 2589–2603 (2024).
Li, N., Goel, N. & Ash, E. Data-Centric Factors in Algorithmic Fairness. In Proc. of the 2022 AAAI/ACM Conference on AI, Ethics, and Society (AIES 2022), 396–410 (2022).
Warner, E. N. et al. Developmental epidemiology of pediatric anxiety disorders. Child Adolesc. Psychiatr. Clin. N. Am. 32, 511–530 (2023).
Dalsgaard, S. et al. Incidence rates and cumulative incidences of the full spectrum of diagnosed mental disorders in childhood and adolescence. JAMA Psychiatry 77, 155–164 (2020).
Sellergren, A. et al. MedGemma technical report. arXiv [cs.AI] https://doi.org/10.48550/arXiv.2507.05201 (2025).
Singhal, K. et al. Toward expert-level medical question answering with large language models. Nat. Med. 31, 943–950 (2025).
Blanzeisky, W. & Cunningham, P. Algorithmic factors influencing bias in machine learning. In Proc. of Joint European Conference on Machine Learning and Knowledge Discovery in Databases, 559–574 (2021).
Ding, X., Xi, R. & Akoglu, L. Outlier detection bias busted: understanding sources of algorithmic bias through data-centric factors. In Proc. AAAI/ACM Conf. AI Ethics, Soc. 7, 384–395 (2024).
Shah, D. S., Schwartz, H. A. & Hovy, D. Predictive biases in natural language processing models: A conceptual framework and overview. In Proc. of the 58th Annual Meeting of the Association for Computational Linguistics (ACL), 5248–5264 (Stroudsburg, PA, USA, 2020).
Acknowledgements
This work was funded by Cincinnati Children’s Hospital Medical Center’s Mental Health Trajectory program. The views expressed are those of the authors and not necessarily those of the Cincinnati Children’s Hospital Medical Center’s Decode program. This work was authored in part by UT-Battelle, LLC, under Contract No. DE-AC05-00OR22725 with the U.S. Department of Energy. The United States Government retains, and the publisher, by accepting the article for publication, acknowledges that the United States Government retains a non-exclusive, paid-up, irrevocable, world-wide license to publish or reproduce the published form of this manuscript or allow others to do so, for United States Government purposes. The Department of Energy will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan).
Author information
Authors and Affiliations
Contributions
J.I.: Conceptualisation, methodology, software, validation, formal analysis, investigation, writing—final draft preparation, writing—reviewing and editing. P.B.: Early draft preparation, writing—reviewing and editing. V.Y. and D.S.: Resources, data curation. J.P. and T.G.: Conceptualisation, methodology, formal analysis, writing—reviewing and editing. J.R.S., G.A., J.T., S.C., M.C., and A.J.K.: Conceptualisation, writing—reviewing and editing. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.I. is an Editorial Board Member for Communications Medicine but was not involved in the editorial review or peer review, nor in the decision to publish this article. All other authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ive, J., Bondaronek, P., Yadav, V. et al. A data-centric approach to detecting and mitigating demographic bias in pediatric mental health text. Commun Med (2026). https://doi.org/10.1038/s43856-026-01480-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43856-026-01480-2