Abstract
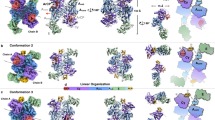
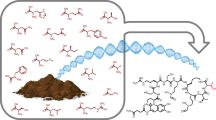
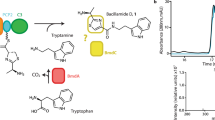
Non-ribosomal peptide synthetases (NRPSs) produce diverse bioactive peptides, including antibiotics and antitumour agents, by integrating proteinogenic and non-proteinogenic amino acids alongside tailoring modifications, such as epimerization (E) and cyclization. Here we demonstrate that the tetrahydropyrimidine ring of pyoverdine is synthesized by the condensation (C) domain PvdL-C3, revealing cyclization capability of a canonical NRPS domain. Analyses of PvdL-E2C3A3 cross-module structures reveal a stable E-C didomain catalytic platform maintained by the donor communication-mediating domain. Through mutagenesis and evolutionary analyses, we identified key active site residues shaping the catalytic channel, controlling both efficiency and specificity. We further engineered PvdL-C3 to accept l-epimers, unveiling latent substrate flexibility. Ultimately, our work provides a unifying model where domain architecture and catalytic channel features collectively govern NRPS function, elucidating a cyclization mechanism and providing a structural blueprint for future enzyme engineering.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data that support the findings of this study are available within the tables and figures in Supplementary Information. The 3D cryo-EM maps generated in this study have been deposited in the Electron Microscopy Data Bank (https://www.emdataresource.org/) under accession numbers EMD-64387, EMD-62988, EMD-62989 and EMD-62990 (Supplementary Table 4). The atomic coordinates have been deposited in the Protein Data Bank (https://www.rcsb.org) under accession numbers 9UP2, 9LCO, 9LCP and 9LCQ. Cryo-EM maps are available via figshare at https://doi.org/10.6084/m9.figshare.30618215 (ref. 72). Source data are provided with this paper.
References
Walsh, C. T. Polyketide and nonribosomal peptide antibiotics: modularity and versatility. Science 303, 1805–1810 (2004).
Felnagle, E. A. et al. Nonribosomal peptide synthetases involved in the production of medically relevant natural products. Mol. Pharm. 5, 191–211 (2008).
Walsh, C. T., O’Brien, R. V. & Khosla, C. Nonproteinogenic amino acid building blocks for nonribosomal peptide and hybrid polyketide scaffolds. Angew. Chem. Int. Ed. Engl. 52, 7098–7124 (2013).
Hur, G. H., Vickery, C. R. & Burkart, M. D. Explorations of catalytic domains in non-ribosomal peptide synthetase enzymology. Nat. Prod. Rep. 29, 1074–1098 (2012).
Patel, K. D., MacDonald, M. R., Ahmed, S. F., Singh, J. & Gulick, A. M. Structural advances toward understanding the catalytic activity and conformational dynamics of modular nonribosomal peptide synthetases. Nat. Prod. Rep. 40, 1550–1582 (2023).
Baltz, R. H. Function of MbtH homologs in nonribosomal peptide biosynthesis and applications in secondary metabolite discovery. J. Ind. Microbiol. Biotechnol. 38, 1747–1760 (2011).
Miller, B. R., Drake, E. J., Shi, C., Aldrich, C. C. & Gulick, A. M. Structures of a nonribosomal peptide synthetase module bound to MbtH-like proteins support a highly dynamic domain architecture. J. Biol. Chem. 291, 22559–22571 (2016).
Bloudoff, K. & Schmeing, T. M. Structural and functional aspects of the nonribosomal peptide synthetase condensation domain superfamily: discovery, dissection and diversity. Biochim. Biophys. Acta Proteins Proteom. 1865, 1587–1604 (2017).
Walsh, C. T. & Nolan, E. M. Morphing peptide backbones into heterocycles. Proc. Natl Acad. Sci. USA 105, 5655–5656 (2008).
Schofield, C. J. et al. Proteins of the penicillin biosynthesis pathway. Curr. Opin. Struct. Biol. 7, 857–864 (1997).
Haslinger, K., Peschke, M., Brieke, C., Maximowitsch, E. & Cryle, M. J. X-domain of peptide synthetases recruits oxygenases crucial for glycopeptide biosynthesis. Nature 521, 105–109 (2015).
Trauger, J. W., Kohli, R. M., Mootz, H. D., Marahiel, M. A. & Walsh, C. T. Peptide cyclization catalysed by the thioesterase domain of tyrocidine synthetase. Nature 407, 215–218 (2000).
Gao, X. et al. Cyclization of fungal nonribosomal peptides by a terminal condensation-like domain. Nat. Chem. Biol. 8, 823–830 (2012).
Quadri, L. E. N., Keating, T. A., Patel, H. M. & Walsh, C. T. Assembly of the Pseudomonas aeruginosa nonribosomal peptide siderophore pyochelin: in vitro reconstitution of aryl-4,2-bisthiazoline synthetase activity from PchD, PchE, and PchF. Biochemistry 38, 14941–14954 (1999).
Gaudelli, N. M., Long, D. H. & Townsend, C. A. β-Lactam formation by a non-ribosomal peptide synthetase during antibiotic biosynthesis. Nature 520, 383–387 (2015).
Ringel, M. T. & Brüser, T. The biosynthesis of pyoverdines. Microb. Cell 5, 424–437 (2018).
Wilson, B. R., Bogdan, A. R., Miyazawa, M., Hashimoto, K. & Tsuji, Y. Siderophores in iron metabolism: from mechanism to therapy potential. Trends Mol. Med. 22, 1077–1090 (2016).
Gulick, A. M. Nonribosomal peptide synthetase biosynthetic clusters of ESKAPE pathogens. Nat. Prod. Rep. 34, 981–1009 (2017).
Clugston, S. L., Sieber, S. A., Marahiel, M. A. & Walsh, C. T. Chirality of peptide bond-forming condensation domains in nonribosomal peptide synthetases: the C5 domain of tyrocidine synthetase is a dCl catalyst. Biochemistry 42, 12095–12104 (2003).
Peng, H., Schmiederer, J., Chen, X., Panagiotou, G. & Kries, H. Controlling substrate and stereospecificity of condensation domains in nonribosomal peptide synthetases. ACS Chem. Biol. 19, 599–606 (2024).
Rausch, C., Hoof, I., Weber, T., Wohlleben, W. & Huson, D. H. Phylogenetic analysis of condensation domains in NRPS sheds light on their functional evolution. BMC Evol. Biol. 7, 78 (2007).
Wheadon, M. J. & Townsend, C. A. Evolutionary and functional analysis of an NRPS condensation domain integrates β-lactam, D-amino acid, and dehydroamino acid synthesis. Proc. Natl Acad. Sci. USA 118, e2026017118 (2021).
Heberlig, G. W., La Clair, J. J. & Burkart, M. D. Crosslinking intermodular condensation in non-ribosomal peptide biosynthesis. Nature 638, 261–269 (2025).
Diecker, J. et al. Structural basis for a scaffolding role of the COM domain in nonribosomal peptide synthetases. Angew. Chem. Int. Ed. Engl. https://doi.org/10.1002/anie.202506621 (2025).
Cao, W. et al. In-situ purification of non-ribosomal peptide synthetases assembly line for structural and biochemical studies. Int. J. Mol. Sci. 26, 1750 (2025).
Owen, J. G., Calcott, M. J., Robins, K. J. & Ackerley, D. F. Generating functional recombinant NRPS enzymes in the laboratory setting via peptidyl carrier protein engineering. Cell Chem. Biol. 23, 1395–1406 (2016).
Tanovic, A., Samel, S. A., Essen, L.-O. & Marahiel, M. A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008).
Hahn, M. & Stachelhaus, T. Selective interaction between nonribosomal peptide synthetases is facilitated by short communication-mediating domains. Proc. Natl Acad. Sci. USA 101, 15585–15590 (2004).
Dehling, E. et al. Mapping of the communication-mediating interface in nonribosomal peptide synthetases using a genetically encoded photocrosslinker supports an upside-down helix-hand motif. J. Mol. Biol. 428, 4345–4360 (2016).
Fage, C. D. et al. Communication breakdown: dissecting the COM interfaces between the subunits of nonribosomal peptide synthetases. ACS Catal. 11, 10802–10813 (2021).
Reimer, J. M. et al. Structures of a dimodular nonribosomal peptide synthetase reveal conformational flexibility. Science 366, eaaw4388 (2019).
Klau, L. J. et al. The natural product domain Seeker version 2 (NaPDoS2) webtool relates ketosynthase phylogeny to biosynthetic function. J. Biol. Chem. 298, 102480 (2022).
Marchler-Bauer, A. et al. CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res. 39, D225–D229 (2011).
Bloudoff, K., Fage, C. D., Marahiel, M. A. & Schmeing, T. M. Structural and mutational analysis of the nonribosomal peptide synthetase heterocyclization domain provides insight into catalysis. Proc. Natl Acad. Sci. USA 114, 95–100 (2017).
Wang, J. et al. Catalytic trajectory of a dimeric nonribosomal peptide synthetase subunit with an inserted epimerase domain. Nat. Commun. 13, 592 (2022).
Bloudoff, K., Rodionov, D. & Schmeing, T. M. Crystal structures of the first condensation domain of CDA synthetase suggest conformational changes during the synthetic cycle of nonribosomal peptide synthetases. J. Mol. Biol. 425, 3137–3150 (2013).
Zhong, L. et al. Engineering and elucidation of the lipoinitiation process in nonribosomal peptide biosynthesis. Nat. Commun. 12, 296 (2021).
Samel, S. A., Schoenafinger, G., Knappe, T. A., Marahiel, M. A. & Essen, L.-O. Structural and functional insights into a peptide bond-forming bidomain from a nonribosomal peptide synthetase. Structure 15, 781–792 (2007).
Keating, T. A., Marshall, C. G., Walsh, C. T. & Keating, A. E. The structure of VibH represents nonribosomal peptide synthetase condensation, cyclization and epimerization domains. Nat. Struct. Mol. Biol. 9, 522–526 (2002).
Izoré, T. et al. Structures of a non-ribosomal peptide synthetase condensation domain suggest the basis of substrate selectivity. Nat. Commun. 12, 2511 (2021).
Pistofidis, A. et al. Structures and mechanism of condensation in non-ribosomal peptide synthesis. Nature 638, 270–278 (2025).
Pfeifer, B. A. Biosynthesis of complex polyketides in a metabolically engineered strain of E. coli. Science 291, 1790–1792 (2001).
Yin, J., Lin, A. J., Golan, D. E. & Walsh, C. T. Site-specific protein labeling by Sfp phosphopantetheinyl transferase. Nat. Protoc. 1, 280–285 (2006).
Thompson, R. F., Iadanza, M. G., Hesketh, E. L., Rawson, S. & Ranson, N. A. Collection, pre-processing and on-the-fly analysis of data for high-resolution, single-particle cryo-electron microscopy. Nat. Protoc. 14, 100–118 (2019).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Kimanius, D. et al. Data-driven regularization lowers the size barrier of cryo-EM structure determination. Nat. Methods 21, 1216–1221 (2024).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
DiMaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D 74, 519–530 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Terlouw, B. R. et al. MIBiG 3.0: a community-driven effort to annotate experimentally validated biosynthetic gene clusters. Nucleic Acids Res. 51, D603–D610 (2023).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51, D1373–D1380 (2023).
Sayers, E. W. et al. Database resources of the national center for biotechnology information. Nucleic Acids Res. 50, D20–D26 (2022).
Blin, K. et al. antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucleic Acids Res. 51, W46–W50 (2023).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Hoang, D. T., Chernomor, O., Von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Cao, W. & Chen, S. L. Cryo-EM maps for PvdL-E2-C3-A3-PCP3 and related constructs. figshare https://doi.org/10.6084/m9.figshare.30618215 (2025).
Acknowledgements
We thank the National Facility for Protein Science in Shanghai (NFPS), Zhangjiang Laboratory, for collecting the cryo-EM images used in this study. The computations in this paper were run on the π 2.0 cluster supported by the Center for High Performance Computing at Shanghai Jiao Tong University. This work was financially supported by the National Key R&D Program of China (grants 2021YFA0910500 to J.L.), the National Science Foundation of China (grants 32270033 and 22577073 to Z.W., 32271302 to J.L., 32201035 to J.W., 32171252 to Z.D.), the China Postdoctoral Science Foundation (grants 2022T150412 and 2022M720085 to J.W.) and the Shanghai ‘Super Postdoctoral’ Incentive Program (grant 202112 to J.W.), Shanghai Pilot Program for Basic Research-Shanghai Jiao Tong University. We declare that the English grammar and flow of the text were polished by using Google’s Gemini large language models.
Author information
Authors and Affiliations
Contributions
Z.W., W.C., J.W. and J.L. conceived of the study. W.C. and S.L.C. performed the biochemical reconstitution and mutagenesis experiments. W.C., J.W. and D.L. conducted the protein structural analysis and W.C. and X.W. performed the compound synthesis and analysis. Data curation was performed by W.C. and J.W. and formal analysis was conducted by Z.W., W.C., J.W. and J.L. The original draft of the paper was written by W.C. and Z.W. and all authors contributed to the review and editing. The project was supervised by J.L. (biochemical experiments) and Z.W. (biochemical and structural analysis experiments). Overall project administration was handled by Z.W. and Z.D., who also led the funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Synthesis thanks Max Cryle, Andrew Gulick and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Peter Seavill, in collaboration with the Nature Synthesis team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Tables 1–6 and Figs. 1–33.
Supplementary Video 1 (download MP4 )
3D variability analysis (3DVA) of the A conformation dataset revealing an oscillation of approximately 20° of the E2 domain relative to the C3 structural domain.
Supplementary Video 2 (download MP4 )
3D variability analysis (3DVA) of the D-conformation dataset showing the oscillation of the E2 domain relative to the C3 domain, and the movement of PCP2 from the E2 active site to the C3 donor site.
Supplementary Video 3 (download MP4 )
3D variability analysis (3DVA) illustrating the conformational switch of the C3 domain between open and closed states, synchronized with the binding and dissociation of the PCP2 domain.
Supplementary Data 1 (download XLSX )
Source data for the chiral analysis bar chart presented in Supplementary Fig. 28.
Source data
Source Data Fig. 2 (download TXT )
Source data for the COMD mutation analysis bar chart (Fig. 2h) and the sequence conservation analysis (Fig. 2i).
Source Data Fig. 4 (download XLSX )
Source data for the relative production bar charts presented in Fig. 4b,d.
Source Data Fig. 5 (download TXT )
Source data for the phylogenetic tree analysis (Fig. 5a) and the enzymatic activity bar chart (Fig. 5c).
Source Data Fig. 6 (download TXT )
Source data for the multiple sequence alignment (Fig. 6a) and relative abundance bar chart (Fig. 6c).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cao, W., Wang, J., Chen, S.L. et al. Analysis of heterocycle formation and stereochemical control by a non-ribosomal peptide synthetase condensation domain. Nat. Synth (2026). https://doi.org/10.1038/s44160-026-01014-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44160-026-01014-7