Abstract
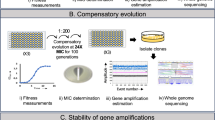
Antibiotic resistance is surging, demanding approaches that detect resistance before genetic fixation, since MIC, sequencing, and culture assays detect resistance late. We present Metabolomics-Driven Intervention Antibiotic Design (MDAD), which stages resistance evolution from metabolic compensation to sub-lethal adaptation to genetic fixation. A brief pulse challenge with a targeted metabolomics panel yields a pre-genetic risk index, interpreted with time-kill assays to situate early survival behaviours and guide mechanism-aware treatment and surveillance.
Similar content being viewed by others
Introduction: the temporal blind spot in resistance detection
Antibiotic resistance remains a major global health threat1. Surveillance systems and therapeutic decisions continue to rely heavily on genetic markers or culture-based susceptibility tests, which detect resistance only after it has become stable and often irreversible2,3. However, recent insights indicate that bacteria undergo metabolic reprogramming in response to antibiotic stress well before acquiring stable genetic resistance mutations4. These early metabolic adaptations may serve as critical biomarkers for impending resistance. Yet, current diagnostic approaches fail to capture this temporal window, representing a significant blind spot in both research and clinical practice4,5. Herein, we focus on articulating the MDAD framework, using representative exemplars to derive cross-cutting metabolic axes and testable directions, rather than cataloguing the literature exhaustively.
MDAD: a conceptual model for resistance evolution
The MDAD model redefines antibiotic resistance not as a binary state, but as a dynamic and metabolically driven continuum, with multiple intervention points prior to irreversible genetic fixation. It delineates the progression of resistance as follows:
Stage 1: Metabolic compensation
Upon antibiotic exposure, bacteria initiate stress response pathways and rewire central metabolism to maintain cellular homoeostasis. Operationally, this unfolds over minutes (~5–30 min) and is marked by quick shifts in energy/redox indices (ATP/ADP, NADH/NAD⁺), transient rerouting of central carbon flow, and induction of stress regulators. These changes are short-lived and typically recede on washout; durable envelope remodelling is not expected at this timescale and generally arises under sustained sub-MIC exposure (Stage 2)6,7,8.
Stage 2: Sub-lethal adaptation
Prolonged sub-inhibitory antibiotic exposure drives changes in membrane composition (e.g., phosphatidylglycerol and cardiolipin enrichment), upregulation of efflux systems, and altered cell envelope architecture. These adaptations improve bacterial survival under stress, yet remain pre-genetic (i.e., not genetically fixed)9,10,11. Typically, these shifts unfold over tens of minutes to a few hours (~30–240 min) under continued sub-MIC stress. Phenotypically, killing becomes slower or biphasic at unchanged MIC (tolerance/persistence), and the state is largely reversible with recovery12,13.
Conceptually, the early (Stages 1–2) changes reflect a bidirectional coupling: the pre-exposure metabolic state can influence uptake and early survival, while class-specific mechanisms of action place predictable demands on central metabolism and the envelope (for example, energy/redox balance, envelope precursor/headgroup pools, and efflux capacity). A class-aware reading helps avoid conflating exposure confirmation with tolerance physiology. We therefore distinguish the mechanism of action (MoA) imprints from metabolic compensation. MoA imprints are immediate, mechanism-aligned metabolic or lipid shifts right after exposure. Metabolic compensation is downstream homoeostatic reprogramming that supports survival over tens of minutes to hours. Tolerance is quantified at the population level using the minimum duration for killing (MDK) metrics derived from time-kill kinetics, which capture the time required to achieve a defined reduction (e.g., 99–99.9%) in bacterial population under sustained antibiotic exposure12,13,14,15.
Stage 3: Genetic fixation
Eventually, stable resistance-conferring mutations accumulate across regulatory/structural loci and, importantly, in direct antibiotic targets and efflux systems. In Staphylococcus aureus, acquisition of mecA and alterations in PBP2 underpin β-lactam resistance10,16; activation of the VraSR cell-wall stress response contributes to reduced glycopeptide susceptibility17. Mutations in target-site genes such as rpoB (RNA polymerase β-subunit) confer rifampicin resistance, and changes in efflux pumps decrease intracellular exposure18. Collectively, these heritable alterations fix the resistant phenotype and markedly limit therapeutic flexibility. At this stage, bacteria rely less on acute metabolic adaptation to survive antibiotic exposure, and intervention becomes substantially less effective16,19,20. Evidence across pathogens and antibiotic classes indicates that pre-genetic metabolic states can bias and accelerate the evolutionary path to genetic resistance. For example, tolerance/persistence increases survival and mutation rates, accelerating resistance evolution21, and metabolic mutations can directly confer resistance phenotypes7,22.
This model highlights the predictive value of metabolomics in detecting stages 1 and 2, offering a critical window for intervention before resistance becomes genetically entrenched (Fig. 1).
Early metabolic adaptation and sub-lethal shifts precede the genetic fixation of resistance. The window for pre-emptive intervention is highlighted along the resistance trajectory. Created with BioRender.com.
In framing pre-genetic adaptation, we place tolerance and persistence within the metabolically plastic window (see Box 1). Stage 1 reflects rapid, reversible metabolic compensation without durable survival gains; Stage 2 captures sustained physiological shifts that manifest as tolerance/persistence in time-kill curves (i.e., right-shifted killing or biphasic survival) without a change in MIC. These states prolong survival, expand opportunity for mutation, and are therefore critical targets for early intervention. Consistent with the stochastic, single-cell origin of resistance, we use bulk metabolomics as a probabilistic, population-level risk index, not a cell-level predictor, with signal strengthened by a brief pulse challenge before extraction.
Evidence across pathogens and drug classes: pre-genetic metabolic signatures
We present representative exemplars and then distil cross-cutting metabolic axes that recur across pathogens and antibiotic classes. As outlined in Box 1, these early adaptive states correspond to tolerance (slower kill at unchanged MIC) and persistence (biphasic killing), so the exemplars below illustrate metabolic underpinnings of these phenotypes rather than isolated anecdotes.
Metabolic adaptations in Staphylococcus aureus: tolerance and pre-resistance remodelling
Recent studies demonstrate that S. aureus undergoes coordinated metabolic rewiring in response to glycopeptide exposure, forming a continuum between antibiotic tolerance and resistance. In a multi-omics investigation of MRSA under vancomycin pressure, Castro et al.23 identified early metabolic shifts including enhanced peptidoglycan biosynthesis, redox adaptation, and alterations in alanine, glutamine, and serine pathways. These changes emerged before phenotypic resistance was detectable, supporting their role as early biomarkers23. Parallel findings by Freiberg et al. revealed that arginine restriction modulates protein synthesis and stress responses, promoting tolerance phenotypes that enhance S. aureus survival during vancomycin treatment in vivo24. Together, these studies highlight metabolic reprogramming, particularly in amino acid and cell wall pathways, as an early, reversible adaptation that anticipates or enables resistance emergence.
Metabolic context shapes β-Lactam resistance in Escherichia coli
Zampieri et al.7 demonstrated that sub-lethal exposure to β-lactam antibiotics under different carbon sources induces distinct metabolic adaptations in E. coli prior to the emergence of classical resistance mechanisms. These early shifts, identified through metabolomics and flux balance analysis, influenced both the pace and direction of resistance evolution, including β-lactamase gene expression. The study highlights how environmental metabolic conditions actively shape adaptive trajectories, supporting the concept that resistance can be anticipated through metabolic profiling before genetic fixation occurs7.
Membrane lipid remodelling detected by metabolomics and lipidomics prior to resistance
Using integrated metabolomics and lipidomics, Jiang et al.25 demonstrated that S. aureus initiates membrane lipid remodelling as a pre-genetic strategy to evade daptomycin and innate immune attack. Prior to acquiring canonical resistance mutations, the bacteria increased lysyl-phosphatidylglycerol (L-PG) levels via upregulation of mprF, reducing daptomycin binding through altered membrane charge. This adaptation also impaired recognition by cationic antimicrobial peptides, enhancing immune evasion. These metabolic and lipidomic changes occurred before resistance fixation25. Genetic disruption of mprF reversed both daptomycin tolerance and immune escape, underscoring the metabolic basis of this transient adaptive phase25. Functionally, the same lipid headgroup and acyl-chain shifts tune surface charge and membrane fluidity, thereby altering cell-envelope permeability and the energetic capacity for efflux, which together reduce effective intracellular daptomycin exposure despite an unchanged MIC.
Taken together, these exemplars indicate recurring, pre-genetic metabolic axes across species and antibiotic classes, supporting generalizable metabolomic readouts and motivating the diagnostic workflow outlined next.
Diagnostic and therapeutic implications
Metabolomics offers a unique lens to detect antibiotic resistance at its earliest metabolic inflection points, before genetic fixation occurs23,26,27. Unlike conventional diagnostics that rely on genetic markers, metabolomics captures dynamic shifts, such as those in glycolysis, TCA cycle activity, and membrane lipid remodelling, that mark the transition toward intermediate adaptive states preceding stable resistance7,13,15,23. These early fingerprints enable more timely classification and intervention, with the explicit understanding that markers quantify population-level risk rather than identify individual mutant cells. In routine interpretation, we co-report simple context variables (e.g., growth phase, media, oxygen, exposure design) and apply class-aware rules so that a given signature is read in the correct physiological frame rather than as a context-free label. In routine use, the panel would be interpreted alongside MDK/time-kill metrics to contextualize population-level tolerance/persistence risk. While tolerance and persistence are useful survival phenotypes for interpretation, MDAD is not limited to these; the panel detects broader, class-aware metabolic liabilities (e.g., envelope charge/fluidity shifts, redox load, efflux-linked exposure loss) that inform regimen adjustment before genetic fixation. A two-stage discovery to clinical reading workflow summarizing this diagnostic framework is shown in Fig. 2. For already resistant isolates, the same panel can guide selection of partner agents or adjuvants and flag cases at high risk of failure despite “susceptible” MICs to alternative drugs.
Stage I (Discovery/Training): Source isolates are profiled under condition-rich designs (media, growth phase, oxygen, inoculum, exposure) using untargeted LC–MS/MS to nominate candidate features. Models are trained with harmonized QC (internal standards, pooled QCs, reference materials) and full context metadata, then stress-tested for generalizability (cross-site, temporal holdouts, untargeted to targeted transfer). This stage outputs a locked, targeted biomarker panel with fixed thresholds, SOP/QC package, and context-aware interpretation rules. Stage II (Clinical Reading): A patient isolate is measured on the locked panel, optionally after a brief, class-aware pulse-challenge, to reveal actionable liabilities. The resulting profile is compared to the fixed biomarkers for risk classification (read in the context of MDK metrics derived from time-kill kinetics), with off-protocol contexts flagged for confirmatory re-measurement under the standard pulse. The report couples the risk call with mechanism-aware treatment adjustments (e.g., agent selection less sensitive to charge/fluidity, efflux inhibition, or partner-drug choice) and co-reports key context variables to ensure physiologically grounded interpretation. Created with BioRender.com.
In practice, a compact panel combining headgroup ratios (e.g., Lys-PG: PG), energy/redox indices (ATP/ADP, NADH/NAD⁺), and a single permeability/efflux check is sufficient to classify samples as envelope-remodelled, a state that predicts reduced intracellular antibiotic exposure at unchanged MIC. To stabilize readouts across contexts, we use a short, standardized pulse-challenge before extraction and a small set of internal controls. When context differs materially from the training conditions, the panel flags the result for confirmatory testing or re-measurement under the standard pulse. When this classification is positive, treatment should prioritize envelope-directed adjustments (e.g., switch to agents less sensitive to charge/fluidity, add an efflux inhibitor, or pair with a membrane-active adjuvant) while confirming the call on the next sample. This keeps the readout actionable within 24–48 h and links metabolomics directly to a regimen change in the pre-genetic window.
To translate this panel into practice, a feasible sample to result pathway is as follows: (i) obtain a colony or positive blood-culture pellet, (ii) apply a brief pulse-challenge with the index antibiotic for minutes to enrich risk bearing states, (iii) quench and extract, and (iv) run a targeted LC–MS/MS panel (10–30 features with isotope-labelled internal standards) covering energy/redox indices (ATP/ADP, NADH/NAD⁺), envelope precursors/headgroups (for example Lys-PG: PG), and selected central-carbon markers. Results are expressed as a population-level risk index and reported alongside the MIC within 24–48 hours. Analytical validation follows MSI-style reporting with shared reference materials and pooled-QC to monitor drift and precision28,29, with inter-instrument and inter-site reproducibility assessed under harmonized QC schedules. Clinical validation then compares the risk index with orthogonal standards (MIC per CLSI/EUCAST; tolerance/MDK from time-kill assays12,14; and clinical endpoints such as time to clearance or treatment failure), adopts TRIPOD + AI for model reporting30, and uses AUROC and, for imbalanced outcomes, AUPRC as evaluation metrics31. This anchors the panel in existing diagnostic practice while enabling prospective assessment of decision impact.
Therapeutically, metabolomics highlights actionable vulnerabilities in bacterial physiology, including altered energy metabolism4,15, membrane restructuring in M. tuberculosis and S. aureus10,18, and upregulated cell wall biosynthesis under β-lactams or glycopeptides10,22. These findings provide a rationale for combination therapies that incorporate metabolic adjuvants, efflux pump inhibitors, or inhibitors targeting specific metabolic pathways26,32,33. Clinically, the use of real-time metabolic profiling offers potential to refine diagnostics and tailor treatments, particularly in critical care settings13,27,34. In parallel, metabolomics continues to drive the identification of novel antibiotic targets in priority pathogens26,32,35,36.
Challenges and technological needs
Despite the promise of metabolomics in forecasting resistance, several hurdles must be overcome before clinical integration. A lack of standardized protocols for sample preparation, data acquisition, and biomarker validation limits reproducibility across studies37,38,39. Condition-dependence is intrinsic to metabolomics and, in MDAD, is treated as design input rather than noise. Panels are trained and validated under condition-rich designs with explicit recording of media chemistry, growth rate, inoculum, temperature, pH, atmosphere, and exposure regimen, so that models learn context-conditioned decision rules. Integration with genomics, proteomics, and transcriptomics can deepen mechanistic insight, but multi-omics data fusion remains computationally intensive7,37,40.
Interpreting complex datasets generated by LC-MS/MS and NMR still requires advanced tools38,41. Before machine learning can guide decisions, data quality and comparability must be established41,42. To that end, we pair standard operating conditions with explicit context metadata and class-aware interpretation. This preserves physiological relevance while improving portability and auditability across sites. For MDAD-style biomarkers, models must (i) train on condition-rich, standardized datasets (strains, media, stressors, exposure times, instruments)28,29, (ii) pre-register features/algorithms and prevent leakage30, (iii) prove transportability via temporal holdouts, cross-site validation, and assay-shift tests (untargeted to targeted)30, and (iv) prioritize feature stability and decision-impact over maximizing AUROC (area under the receiver operating characteristics (ROC) curve; a threshold-free measure of ranking performance). For imbalanced problems, prefer AUPRC (area under the precision–recall curve), which is more sensitive to rare positives31. Practically, this requires shared reference materials and internal-standard mixes, harmonized QC (including pooled-QC frequency and acceptance criteria), explicit metadata standards (growth rate, inoculum, media chemistry, temperature, pH, atmosphere/gas composition [e.g., aerobic, anaerobic, microaerophilic; O₂/CO₂ %, headspace/aeration], exposure design)28, and prospective validation in which the metabolomics result alters a therapeutic decision within 24–48 h and improves outcomes (e.g., faster clearance, fewer failures)30. This moves the discussion from feasibility to clinical reliability. The principal bottleneck is standardized, multi-site, condition-rich data and metadata; with that foundation, AI/ML becomes a tractable implementation problem. Even with these modelling and standardization steps in place, deployment still depends on practical constraints, per-sample cost, instrument access, and staff expertise, which vary widely across diagnostic settings38,43.
Finally, reported metabolomic signatures often lack causal validation and can vary across strains and growth conditions7,8,26. Addressing these limitations, through time-resolved designs, interventional perturbations (genetic or chemical), multi-strain panels, and integration with surveillance networks, will be essential for translating discovery into meaningful clinical impact39,44.
Outlook: toward a pre-resistance paradigm
By detecting resistance trajectories before they become genetically entrenched, metabolomics offers a paradigm shift in how antibiotic resistance is diagnosed, monitored, and managed7,26,40. Rather than reacting to fixed mutations, clinicians could intervene during earlier, metabolically plastic stages, potentially reversing or halting resistance before it becomes heritable26,40,41. In this view, condition-dependence becomes actionable: by reading metabolic state in its context, clinicians can adjust therapy or microenvironment to reduce pre-genetic survival and narrow the window to mutation. MDAD is not limited to pre-genetic states. In isolates with established resistance, targeted metabolomic profiling maps compensatory costs and fragile pathways18,40. Brief probe challenges with intended partner drugs or adjuvants expose actionable liabilities (for example, efflux-dependent exposure loss or envelope stress)15,25, supporting mechanism-aware rescue therapy and surveillance for second-step evolution32.
In clinical diagnostics, real-time metabolic fingerprints could guide tailored therapies based on bacterial adaptation profiles, improving outcomes in time-critical settings such as intensive care units38,39,45. At the population level, metabolic surveillance may support early detection of emerging resistance patterns, enriching global initiatives with functional data beyond genomics26,46.
Looking ahead, metabolomics could drive the discovery of new drug targets that exploit transient vulnerabilities in central metabolism, membrane remodelling, or redox regulation, which are key pillars of the MDAD framework47,48,49. Integration with AI and machine learning will be critical to decoding complex metabolic datasets and predicting resistance phenotypes in diverse pathogens41,50.
Translational validation is essential to establish whether early metabolic shifts reliably precede resistance and predict treatment failure. Key approaches include time-kill metabolomics, animal infection models, and longitudinal clinical cohort studies7,33,48,51,52,53. Demonstrating reproducibility across pathogens and settings will be pivotal for embedding metabolomics into next-generation antimicrobial stewardship strategies39,41,46.
Data availability
No datasets were generated or analysed during the current study.
References
WHO. WHO bacterial priority pathogens list, 2024: Bacterial pathogens of public health importance to guide research, development and strategies to prevent and control antimicrobial resistance 2024 (2024).
Yamin, D. et al. Current and Future Technologies for the Detection of Antibiotic-Resistant Bacteria. Diagnostics 13. https://doi.org/10.3390/diagnostics13203246 (2023).
Muntean, M. M. et al. Phenotypic and genotypic detection methods for antimicrobial resistance in ESKAPE pathogens (Review). Exp. Ther. Med. 24, 508 (2022).
Dunphy, L. J., Yen, P. & Papin, J. A. Integrated Experimental and Computational Analyses Reveal Differential Metabolic Functionality in Antibiotic-Resistant Pseudomonas aeruginosa. Cell Syst. 8, 3–14.e13 (2019).
Santi, I., Manfredi, P., Maffei, E., Egli, A. & Jenal, U. Evolution of antibiotic tolerance shapes resistance development in chronic Pseudomonas aeruginosa infections. mBio 12, e03482–20 (2021).
Wong, F. et al. Reactive metabolic byproducts contribute to antibiotic lethality under anaerobic conditions. Mol. Cell 82, 3499–3512.e3410 (2022).
Zampieri, M. et al. Metabolic constraints on the evolution of antibiotic resistance. Mol. Syst. Biol. 13, 917 (2017).
Zampieri, M., Zimmermann, M., Claassen, M. & Sauer, U. Nontargeted metabolomics reveals the multilevel response to antibiotic perturbations. Cell Rep. 19, 1214–1228 (2017).
MacDermott-Opeskin, H. I., Gupta, V. & O’Mara, M. L. Lipid-mediated antimicrobial resistance: a phantom menace or a new hope?. Biophys. Rev. 14, 145–162 (2022).
Nikolic, P. & Mudgil, P. The cell wall, cell membrane and virulence factors of Staphylococcus aureus and their role in antibiotic resistance. Microorganisms 11. https://doi.org/10.3390/microorganisms11020259 (2023).
Wee, G. N. et al. Phenotypic convergence of bacterial adaption to sub-lethal antibiotic treatment. Front. Cell. Infect. Microbiol. 12, 913415 (2022).
Brauner, A., Fridman, O., Gefen, O. & Balaban, N. Q. Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nat. Rev. Microbiol. 14, 320–330 (2016).
Lopatkin, A. J. et al. Bacterial metabolic state more accurately predicts antibiotic lethality than growth rate. Nat. Microbiol. 4, 2109–2117 (2019).
Brauner, A., Shoresh, N., Fridman, O. & Balaban, N. Q. An Experimental Framework for Quantifying Bacterial Tolerance. Biophys. J. 112, 2664–2671 (2017).
Stokes, J. M., Lopatkin, A. J., Lobritz, M. A. & Collins, J. J. Bacterial Metabolism and Antibiotic Efficacy. Cell Metab. 30, 251–259 (2019).
Munita, J. M. & Arias, C. A. Mechanisms of Antibiotic Resistance. Microbiol Spectr. 4. https://doi.org/10.1128/microbiolspec.VMBF-0016-2015 (2016)
Howden, B. P., Davies, J. K., Johnson, P. D. R., Stinear, T. P. & Grayson, M. L. Reduced Vancomycin Susceptibility in Staphylococcus aureus, Including Vancomycin-Intermediate and Heterogeneous Vancomycin-Intermediate Strains: Resistance Mechanisms, Laboratory Detection, and Clinical Implications. Clin. Microbiol. Rev. 23, 99–139 (2010).
Gygli, S. M., Borrell, S., Trauner, A. & Gagneux, S. Antimicrobial resistance in Mycobacterium tuberculosis: mechanistic and evolutionary perspectives. FEMS Microbiol. Rev. 41, 354–373 (2017).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Vivekanandan, K. E., Kumar, P. V., Jaysree, R. C. & Rajeshwari, T. Exploring molecular mechanisms of drug resistance in bacteria and progressions in CRISPR/Cas9-based genome expurgation solutions. Glob. Med. Genet. 12, 100042 (2025).
Windels, E. M. et al. Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME J. 13, 1239–1251 (2019).
Schrader, S. M. et al. Multiform antimicrobial resistance from a metabolic mutation. Sci. Adv. 7, eabh2037 (2021).
Castro, B. E. et al. Multiomics characterization of methicillin-resistant Staphylococcus aureus (MRSA) isolates with heterogeneous intermediate resistance to vancomycin (hVISA) in Latin America. J. Antimicrob. Chemother. 78, 122–132 (2022).
Freiberg, J. A. et al. Restriction of arginine induces antibiotic tolerance in Staphylococcus aureus. Nat. Commun. 15, 6734 (2024).
Jiang, J.-H. et al. Antibiotic resistance and host immune evasion in Staphylococcus aureus mediated by a metabolic adaptation. Proc. Natl. Acad. Sci. 116, 3722–3727 (2019).
Kok, M., Maton, L., van der Peet, M., Hankemeier, T. & van Hasselt, J. G. C. Unraveling antimicrobial resistance using metabolomics. Drug Discov. Today 27, 1774–1783 (2022).
Venkataraman, R., Nakedi, K., Akhade, A. S. & Soni, V. In Antimicrobial Resistance: Factors to Findings: Omics and Systems Biology Approaches 151–177 (Springer, 2024).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Broadhurst, D. et al. Guidelines and considerations for the use of system suitability and quality control samples in mass spectrometry assays applied in untargeted clinical metabolomic studies. Metabolomics 14, 72 (2018).
Collins, G. S. et al. TRIPOD+ AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ 385 (2024).
Saito, T. & Rehmsmeier, M. The precision-recall plot is more informative than the ROC plot when evaluating binary classifiers on imbalanced datasets. PloS One 10, e0118432 (2015).
Campos, A. I. & Zampieri, M. Metabolomics-DRIVEN EXPLORATION OF THE CHEMICAL DRUG SPACE TO PREDICT COMBINATION ANTIMICROBIAL THERAPies. Mol. Cell 74, 1291–1303.e1296 (2019).
Hussein, M. et al. Metabolomics STUDY OF THE SYNERGISTIC KILling of Polymyxin B in combination with Amikacin against Polymyxin-SUSCEPTIBLE AND -RESIstant Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 64, https://doi.org/10.1128/aac.01587-01519 (2019).
Vo, D. K. & Trinh, K. T. L. Emerging BIOMARKERS IN METABOLOMICS: ADVANCEMENTS IN PRECISION HEALTH AND DISEASE DIAGNOSis. Int. J. Mol. Sci. 25. https://doi.org/10.3390/ijms252313190 (2024).
Hussein, M. et al. Metabolic profiling unveils enhanced antibacterial synergy of polymyxin B and teixobactin against multi-drug resistant Acinetobacter. baumannii. Sci. Rep. 14, 27145 (2024).
Hussein, M. et al. Providing insight into the mechanism of action of cationic lipidated oligomers using metabolomics. mSystems 9, e0009324 (2024).
Mohr, A. E., Ortega-Santos, C. P., Whisner, C. M., Klein-Seetharaman, J. & Jasbi, P. Navigating Challenges and Opportunities in Multi-Omics Integration for Personalized Healthcare. Biomedicines 12, 1496 (2024).
Pinu, F. R., Goldansaz, S. A. & Jaine, J. Translational METABOLOMICS: CURRENT CHALLENGES AND FUTURE OPPORTUNITies. Metabolites 9. https://doi.org/10.3390/metabo9060108 (2019).
Rahman, M. & Schellhorn, H. E. Metabolomics of infectious diseases in the era of personalized medicine. Front. Mol. Biosci. 10, 1120376 (2023).
Wareth, G., Neubauer, H. & Sprague, L. D. A silent network’s resounding success: how mutations of core metabolic genes confer antibiotic resistance. Signal Transduct. Target Ther. 6, 301 (2021).
Chi, J. et al. Artificial intelligence in metabolomics: a current review. TrAC Trends Anal. Chem. 178, 117852 (2024).
Zhao, A. P. et al. AI for science: predicting infectious diseases. J. Saf. Sci. Resil. 5, 130–146 (2024).
Emwas, A. H. et al. NMR Spectroscopy for metabolomics research. Metabolites 9. https://doi.org/10.3390/metabo9070123 (2019)
Hassall, J., Coxon, C., Patel, V. C., Goldenberg, S. D. & Sergaki, C. Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects. npj Antimicrob. Resist. 2, 16 (2024).
Ashrafian, H. et al. Metabolomics: The Stethoscope for the Twenty-First Century. Med Princ. Pr. 30, 301–310 (2021).
Elbehiry, A. et al. Detection of antimicrobial resistance via state-of-the-art technologies versus conventional methods. Front. Microbiol. 16, 1549044 (2025).
Zhu, S. et al. Metabolomics revealed mechanism for the synergistic effect of sulbactam, polymyxin-B and amikacin combination against Acinetobacter baumannii. Front. Microbiol. 14, 1217270 (2023).
Hussein, M. et al. Mechanisms Underlying Synergistic Killing of Polymyxin B in Combination with Cannabidiol against Acinetobacter baumannii: A Metabolomic Study. Pharmaceutics 14, 786 (2022).
Ye, D. et al. β-lactam antibiotics induce metabolic perturbations linked to ROS generation leads to bacterial impairment. Front. Microbiol. 15, 1514825 (2024).
Chang, A. & Chen, J. H. BSAC Vanguard Series: Artificial intelligence and antibiotic stewardship. J. Antimicrob. Chemother. 77, 1216–1217 (2022).
Pacheco-Navarro, A. E. & Rogers, A. J. The metabolomics of critical illness. Handb. Exp. Pharm. 277, 367–384 (2023).
Oliveira, L. B. et al. Metabolomic profiling of plasma reveals differential disease severity markers in COVID-19 patients. Front. Microbiol. 13, 844283 (2022).
Dutta, N. K. et al. Integration of metabolomics and transcriptomics reveals novel biomarkers in the blood for tuberculosis diagnosis in children. Sci. Rep. 10, 19527 (2020).
Malachowa, N. et al. Insights into the molecular basis of reduced vancomycin susceptibility among three prominent Staphylococcus aureus clonal complexes. Microbiol. Spectr. 12, e00486–00424 (2024).
Author information
Authors and Affiliations
Contributions
M.H. conceived the central idea and developed the MDAD conceptual model. M.H. and T.V. outlined the manuscript structure and drafted the initial text. M.H. wrote the full manuscript, conducted literature synthesis, and prepared the figures. D.J.C., M.B., G.G.R., J.L., and T.V. contributed critical insights into metabolomics, clinical translation, and antimicrobial pharmacology. All authors reviewed, edited, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hussein, M., Creek, D.J., Baker, M. et al. Metabolomics-driven prediction of antibiotic resistance: a perspective on pre-genetic intervention. npj Antimicrob Resist 3, 101 (2025). https://doi.org/10.1038/s44259-025-00168-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00168-2