Abstract
Candida parapsilosis is an opportunistic yeast that was recently deemed a high- importance fungal pathogen by the World Health Organization. In fact, C. parapsilosis poses an escalating threat in healthcare settings due to its ability to adapt to diverse environments, propensity for human-to-human transmission, and capacity to develop antifungal resistance. Recent studies emphasize its rising clinical importance, particularly with the increasing resistance to antifungals and the emergence of clonal outbreaks, making it a serious threat to public health. This review provides an up-to-date synthesis of our current knowledge on this yeast, addressing its epidemiology, environmental adaptability, and the molecular mechanisms driving resistance to azoles and echinocandins. In particular, it provides a comprehensive overview of the resistome of C. parapsilosis, offering insights into the genetic determinants associated with antifungal resistance. We also identify key unresolved questions and emphasize the need for further research to mitigate its impact on healthcare systems.
Similar content being viewed by others
Candida parapsilosis is a diploid ascomycetous yeast and a member of the Debaryomycetaceae. Like other medically relevant yeasts such as Candida albicans, Candidozyma auris or Candida tropicalis, C. parapsilosis belongs to the CUG-Ser1 clade in which the CUG codon has been reassigned to code for serine rather than leucine1. Notably, C. parapsilosis is part of a complex of closely related species (C. parapsilosis sensu stricto, Candida orthopsilosis, Candida metapsilosis, and the two recently described Candida theae and Candida margitis), some of which emerged through hybridization events2,3. However, C parapsilosis sensu stricto (from here on indicated as C. parapsilosis) is by far the predominant species of the complex in human clinical samples. Behaving as a natural commensal of the skin and digestive tract, C. parapsilosis may also act as an opportunistic pathogen and cause bloodstream infections (i.e. candidemia), endocarditis, peritonitis, and other invasive diseases in susceptible hosts4.
Candida parapsilosis has a global distribution and is particularly prevalent in healthcare environments4. It is recognized as the second or third most common cause of invasive candidiasis worldwide, following C. albicans and, in some regions, Nakaseomyces glabratus (formerly Candida glabrata)5. The incidence of C. parapsilosis infections is notably high in neonatal intensive care units (NICUs) and among immunocompromised patients. Geographically, higher incidence rates have been reported in Southern Europe, South Africa and Latin America, reflecting regional variations in species distribution and possibly infection control practices6. Global mortality rates for C. parapsilosis bloodstream infections range from 17% to 47%, which—while lower than those associated with C. albicans and N. glabratus—still represent a substantial clinical burden7,8. According to the recent multicentre study Candida III, the attributable mortality rate for C. parapsilosis candidemia was ~8%, underscoring the serious nature of these infections9.
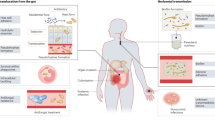
Several risk factors contribute to the susceptibility to invasive C. parapsilosis infections, including the use of central venous catheters, total parenteral nutrition, broad-spectrum antibiotics, corticosteroids, neutropenia, and prolonged hospital stays. It was recently demonstrated that prophylactic treatment with broad-spectrum antibiotics induces gut dysbiosis in hematopoietic stem transplant recipients (10). The resulting reduction in bacterial load promotes yeast overgrowth in the gut, thus facilitating the translocation of C. parapsilosis into the bloodstream to cause candidemia10. In neonates, particularly premature infants, additional risk factors include low birth weight and immature immune systems, making C. parapsilosis a leading cause of fungal sepsis in this vulnerable population11. Moreover, C. parapsilosis ability to form biofilms on medical devices (e.g. intravenous catheters) often results in infections that are persistent and difficult-to-treat (though this trait is highly strain-dependent)12. It is interesting to note that candidalysin, firstly discovered in C. albicans and further unveiled as a major virulence factor of C. albicans, C. dubliniensis and C. tropicalis, does not have an ortholog in the genome of C. parapsilosis13,14.
Until not long ago, the vast majority of C. parapsilosis isolates were susceptible to azoles. However, since 2018, the frequency of isolation of azole-resistant C. parapsilosis strains has been alarmingly increasing, parallelled by a higher number of reports of fluconazole-resistant C. parapsilosis causing clonal outbreaks in the clinics4,15,16. Although there are some conflicting data regarding whether or not fluconazole resistance in C. parapsilosis results in a higher mortality17,18,19, the delay in effective therapy – as sometimes fluconazole is started empirically - together with the ability of these strains to spread across different hospitals, cities, and potentially continents20,21, make for an explosive mix. It is therefore not surprising that this species has been included in the high-priority group of the recently published WHO Fungal Priority Pathogen List (FPPL)22.
Candida parapsilosis can adapt to and thrive in various environments
C. parapsilosis has long been known in the medical setting as a skin colonizer, which helps explaining its potential for inter-human transmission via horizontal transfer23. Besides humans, it has also been isolated from a large variety of animals, including wild birds24,25,26. Of note, animal isolates of C. parapsilosis - and other opportunistic yeasts - displaying acquired azole resistance to medical antifungals have been reported, suggesting that selective pressure may occur outside the human host, raising additional concerns27,28.
An environmental study conducted in the US identified several Candida species (including C. parapsilosis) in soil samples, suggesting that they may evolve and possibly thrive outside human hosts, which challenge the dogma of these yeasts being strict commensals29,30. C. parapsilosis has also been isolated from different environmental niches, such as agricultural fields, compost, fruits and recreational water31,32,33,34,35. It is however still unclear whether C. parapsilosis is merely leaking into the environment passively from animals, waste or human activities, or if it actively evolves and persists independently from human/animal presence. Notably, C. parapsilosis has been frequently isolated from environmental and water samples within hospital settings36. As an example, 42% of environmental samplings taken from an intensive care unit in Italy grew C. parapsilosis37. Like C. auris, C. parapsilosis has a remarkable ability to survive for several days to weeks under harsh conditions, such as dry surfaces and nutrient limitation38,39; this species is even frequently isolated from laundry machines40. In addition, quaternary ammonium disinfectants are poorly effective in killing C. parapsilosis, as is the case for other Candida species41,42. Moreover, many strains of C. parapsilosis are robust biofilm producers, a trait that enhances their ability to colonize and persist on medical devices43.
C. parapsilosis response to stressful stimuli has been investigated in several studies: this species has demonstrated tolerance to high concentrations of potassium chloride (KCl), sodium chloride (NaCl), and lithium chloride (LiCl), as well as the ability to grow across a wide range of pH levels39,44,45. Compared to the other common opportunistic yeast species, it shows greater tolerance to oxidative stress induced by hydrogen peroxide and cell wall stress induced by calcofluor white45. However, it is less thermotolerant than other species, although some isolates may grow at temperatures as high as 47 °C45. A large gene disruption library recently revealed a unique regulation of sulphur metabolism in this species, which may be involved in scavenging essential sulphur from its environment46.
Overall, C. parapsilosis ability to persist in harsh environments suggests that contaminated surfaces/devices may act as reservoirs in healthcare settings, thus contributing to its propensity to spread and cause outbreaks.
In the last years, acquired antifungal resistance has been emerging dramatically
C. parapsilosis intrinsically shows diminished susceptibility to echinocandins as a consequence of the naturally occurring P660A amino acid substitution in the hot spot region 1 (HS1) of the FKS1 gene47. Nonetheless, this species has historically been reported as mostly susceptible to azoles48,49. Unfortunately, the landscape is evolving as C. parapsilosis isolates showing echinocandin resistance, azole resistance, or both, are looming in many parts of the globe since the turn of the XXI century. To date, true echinocandin resistance is still anecdotic, but has been reported in Greece, Iran, and China50,51,52. Some isolates may show echinocandin tolerance rather than true resistance, but the clinical impact of this observation is still unknown53.
On the contrary, the sharp increase in the number of azole-resistant isolates has been a matter of concern in hospitals located in a growing number of countries during the last 10 years15. A study by Pfaller et al. based on data collected under the radar of the SENTRY Program in 2019 reported an overall fluconazole resistance rate of up to 4.6% in Europe — the region where the highest rates were observed54. However, rates of azole resistance are quite variable from one centre to another; whereas many hospitals remain unaffected, other ones report extremely high rates of azole resistance, as reported in Spain, South Africa, Greece, and Italy49,55,56,57,58,59. Of note, high resistance rates should be interpreted with caution, as they may reflect extensive horizontal transfer of resistant strains between patients rather than increased selection of resistant strains. Notably, whereas a study on candidemia conducted in 29 Spanish hospitals in 2010 and 2011 reported fluconazole resistance in up to 3% of C. parapsilosis isolates48, two recent studies conducted in Madrid between 2019 and 2022 revealed that some hospitals experienced a rising number of fluconazole-resistant C. parapsilosis isolates throughout the study period, while other hospitals in the same city did not report the presence of such isolates49,58. If the centre-to-centre variation requires performing antifungal susceptibility testing in clinical isolates to obtain a close monitoring of the local epidemiology, national data are also needed to establish a framework of the situation on a wider level. A prospective study on antifungal resistance in isolates causing fungemia among patients admitted to 85 hospitals located across Spain is currently ongoing in 2025. Data from this study will be useful to monitor epidemiological changes in the country in the last 15 years. Recently, a US reference laboratory reported an increase in fluconazole resistance in C. parapsilosis, rising from 8.2% in 2015 to 20.3% in 202460. Altogether, fluconazole-resistant C. parapsilosis is an increasing problem that can affect hospitals in a disparate manner, even in hospitals settled in the same city, highlighting the need for prospective surveillance programs. It is worth noting that a recent multicentre retrospective study from Italy identified fluconazole resistance as a risk factor for recurrent candidemia within one year18.
Molecular mechanisms of antifungal resistance in C. parapsilosis are highly diverse
The publication of the draft genome of the C. parapsilosis CDC 317 strain in 2009 was a significant milestone in the study of this major human fungal pathogen, which among other impacts, facilitated research into the molecular mechanisms underlying antifungal resistance61. This genome assembly is regularly updated and accessible via the Candida Genome Database62. Last year, the first chromosome-level genome assembly of C. parapsilosis was published63. C. parapsilosis has a diploid genome of 13 Mbp, containing ~5800 genes, spread across eight chromosomes. Compared to other Candida species, its genome exhibits an exceptionally low level of heterozygosity, with only one heterozygous single nucleotide polymorphism (SNP) per 15,553 bases44,61.
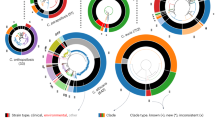
Molecular studies of clinical isolates exhibiting acquired antifungal resistance, particularly to azoles, have uncovered a broad array of mechanisms that enable C. parapsilosis to evade antifungal treatments15. Specifically, mutations leading to amino acid substitutions in the ERG11 gene, which encodes lanosterol demethylase—the target enzyme for azole drugs—have been documented in several studies55,56,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82. To date, about 20 distinct Erg11 amino acid substitutions have been identified in clinical isolates. Importantly, the advent of genome manipulation, and more recently the introduction of CRISPR-Cas9, have streamlined dissecting the potential contribution of these variants to azole resistance67,83,84. A map of the ERG11 variants described so far, along with their documented impact on fluconazole susceptibility is provided in Fig. 1. The effect of an amino acid substitution on susceptibility to fluconazole depends on its location within the enzyme 3D structure, allelic dosage (heterozygous or homozygous), and the genetic background of the clinical isolates84. For example, while the R398I mutation is a neutral polymorphism (i.e. no impact on azole resistance), the Y132F mutation—the most commonly reported worldwide and also present in the type strain CDC317—primarily affects short-tailed azoles (fluconazole and voriconazole), long-tailed azoles (posaconazole) being mostly unaffected84. Other mechanisms include the upregulation of ERG11 and active efflux resulting from the overexpression of membrane transporters belonging to the ATP-binding cassette (Cdr1) or Major Facilitator (Mdr1) super families. Both are driven by gain-of-function mutations which produce hyperactive alleles in genes encoding transcription factors, namely UPC2, MRR1, and TAC183,85,86. The presence of both ERG11 mutations and hyperactive transcription factors is likely to contribute synergistically to resistance. Although Tac1, Mrr1 and Upc2 orchestrate the regulation of CDR1, MDR1 and ERG11, respectively, in C. albicans, Mrr1 has been shown to regulate MDR1, MDR1B (CPAR2_603010) but also CDR1B (CPAR2_304370) in C. parapsilosis87. Doorley et al., further demonstrated that Tac1 in C. parapsilosis directly modulates fluconazole susceptibility by interacting with multiple CDR transporter genes, namely CDR1 (CPAR2_405290), CDR1B (CPAR2_307370) and CDR1C (CPAR2_300010)83. Multiple amino acid variants have been reported from clinical isolates in Tac120,53,65,68,71,77,83,88,89 and Mrr120,56,64,65,68,74,76,79,86,87,88,89,90,91 (Fig. 1). Yet, only a few of them have been demonstrated to result in hyperactive alleles, thereby contributing to azole resistance. One of these is G583R in Mrr1, which also reduces the susceptibility of C. parapsilosis to flucytosine84. A similar cross-resistance pattern has been reported in C. lusitaniae, and related to MDR1 upregulation92,93. Compared to Tac1 and Mrr1, fewer than 20 variants have been reported in Upc2, and none have been shown to alter fluconazole susceptibility68,71,88,94. However, the current picture of amino acid variants occurring in these transcription factors is possibly incomplete as most studies focus only on Erg11 amino acid polymorphism.
Amino acid changes associated with fluconazole resistance by functional validation are highlighted in red. Substitutions only reported in fluconazole resistant/susceptible dose-dependent isolates are depicted in black; note that these do not necessarily confer fluconazole resistance, as the resistant phenotype may be due to their combination with other mutations/other mechanisms completely. Substitutions either identified in fluconazole-susceptible strains or not associated with resistance according to functional validation are depicted in green. *Mutation leading to a frameshift/stop codon. Note: although Erg11G458S direct impact on resistance was not experimentally validated, this amino acid change has already been associated with fluconazole resistance in C. orthopsilosis140 and is related to G464S in C. albicans, which is also known to confer fluconazole resistance141.
A few isolates harbouring mutations (D14Y, G111R and R135I) that impair C-5 sterol desaturase activity - encoded by the ERG3 gene - have also been reported85,95,96. Although exposure to azoles in wild-type Erg3 yeasts results in the accumulation of toxic sterol intermediates, when Erg3 is non-functional due to missense mutations, these intermediates are no longer converted into the toxic products, thereby reducing cellular toxicity and allowing yeast cells to persist even in the presence of azole drugs. In addition, as a consequence of Erg3 impairment, these mutants also display an altered membrane sterol profile characterized by a marked decrease in ergosterol and the accumulation of intermediate sterols (mainly ergosta-7,22-dienol), leading to a reduced susceptibility to amphotericin B. Surprisingly, this uncommon phenotype is also associated with increased tolerance to echinocandins in C. parapsilosis84,95.
Few clinical isolates showing evidence for acquired echinocandin resistance have been reported so far, and all had FKS1 mutations50,51,52. Although still uncommon, the simultaneous emergence of echinocandin and fluconazole resistance poses the risk of multidrug resistance, as already observed in Turkey50. Interestingly, in vitro evolution experiments exposing C. parapsilosis isolates to micafungin have unveiled additional substitutions in HS1 (amino acid 652–660) and HS2 (1369–1376)97,98. Substitutions located outside these hotspots were also reported in clinical isolates, although they do not seem to confer resistance53,99,100. Instead, they have been associated with echinocandin tolerance (which can facilitate the acquisition of resistance) and enhanced growth, while reducing fitness costs in response to oxidative stress99. Table 1 lists all Fks1 amino acid substitutions described to date.
In light of the increase of azole resistance, new antifungal drugs with original modes of action are now more than even required. Among them, fosmanogepix (formerly APX001)—the prodrug of manogepix—is currently in phase III clinical trials101. This new antifungal targets Gwt1, a fungal glycosylphosphatidylinositol acyltransferase. With the exception of Pichia kudriavzevii, manogepix displays strong activity against various yeasts, including C. parapsilosis102. Although resistance to this new drug is still underexplored, in vitro evolution experiments of C. albicans, N. glabratus and C. parapsilosis isolates have led to the selection of resistant isolates showing a 4–32 fold increase in manogepix MIC103. Sequencing of the resistant isolates identified point mutations in the GWT1 gene (leading to amino acid substitutions V162A and V163A in C. albicans and N. glabratus, respectively), although not in C. parapsilosis, suggesting the existence of both GWT1-dependent and -independent mechanisms of manogepix resistance. In 2020, Arendrup et al., demonstrated a correlation between manogepix and fluconazole MICs and suggested a possible role for efflux pumps104. Recently, it was demonstrated that G650E, a gain-of-function mutation in Tac1 conferring fluconazole resistance, also reduces the susceptibility of C. parapsilosis to manogepix through both CDR-dependent and independent mechanisms105.
When addressing antifungal resistance, genomic plasticity—an important trait in fungal pathogens—should be considered, as it enables fungal cells to adapt rapidly and withstand stressful conditions. As in other yeast species, aneuploidy has also been reported in C. parapsilosis106. As an example, chromosome 5 trisomy has recently been shown to facilitate in vitro adaptation to caspofungin—likely due to CHS7 gene amplification—and cross tolerance to flucytosine107. Genomic plasticity can also translate into segmental aneuploidy or shorter-range amplifications, triggering copy number variations (CNV). CNVs have recently been reported for both ERG11 and CDR1B, and may explain azole resistance in some clinical isolates79,108.
Finally, heteroresistance, which has been elegantly demonstrated in C. parapsilosis, could explain breakthrough infections in haematology patients receiving echinocandin prophylaxis109. Heteroresistance refers to a phenomenon in which a small subpopulation of cells—typically around 1 in 10,000—displays resistance to a given antifungal agent, while the majority of the population remains susceptible. Notably, these resistant cells can proliferate under antifungal pressure, but the phenotype is reversible upon removal of the selective agent. Yet, although documented in other opportunistic yeasts, heteroresistance to azoles has not been observed in C. parapsilosis110,111. Importantly, heteroresistance detection poses a significant challenge for clinical laboratories, as standard antifungal susceptibility tests often fail to identify it. Instead, it requires a dedicated approach such as population analysis profiling112.
Overall, the emergence and selection of allelic variants in C. parapsilosis, combined with its genome plasticity, allow this species to cope with antifungal drugs.
Azole-resistant C. parapsilosis can cause clonal outbreaks in clinical settings
Resistant isolates have been detected as part of the colonising microbiota of patients and in the surrounding environment where the patients are hosted55. In the presence of the specific risk factors described above, the isolates can then cause endogenous infections or spread to other patients via patient-to-patient transmission or environment-to-patient transmission. Genotyping of isolates is therefore crucial for understanding the molecular epidemiology of circulating clones. This approach provides better insight into the dynamics of their spread between patients, which may explain the unexpectedly high rates of fluconazole resistance observed at some centres. This route of infection acquisition is now well supported by reports of clonal outbreaks involving C. parapsilosis isolates harbouring the Y132F substitution, which is by far the most common Erg11 substitution in fluconazole-resistant isolates15. For yet unknown reasons, isolates harbouring this substitution seems more prone to cause clonal local outbreaks and potentially become widespread among many hospitals worldwide20,21,68,69,71,75,113. The presence of Y132F isolates was first reported in 2004 in a Turkish hospital74. Since then, they have been detected in a growing number of countries. Table 2 summarizes studies reporting outbreaks of invasive candidiasis caused by C. parapsilosis isolates harbouring the Y132F substitution over the last 20 years. Most outbreaks have revealed the presence of one dominating genotype, often accompanied by additional, less frequent genotypes that are genetically closely related. Some studies reported the presence of identical genotypes in different hospitals114, sometimes located at different cities115, which outlines the capacity of isolates harbouring the Y132F substitution to spread. The fact that the vast majority of patients infected by fluconazole-resistant isolates had not received any azole by the time they became infected strongly points towards the patient-to-patient transmission as the main driver of these outbreaks116. There are yet only a few studies comparing the genetic relationship between azole-resistant isolates collected from different countries. A recent regional study conducted in Spain showed that the genotypes found were city-specific, except for some genotypes found simultaneously at relatively close cities, which may suggest transmission across hospitals after patient transfers115. Another study recently demonstrated that a 2018–22 outbreak that occurred at multiple hospitals in Berlin, Germany, was caused by isolates that were closely related to those responsible for outbreaks in Turkey, Kuwait and South Korea20. The fact that unrelated genotypes (or lineages) harbouring the Y132F substitution have been emerging at different geographic areas for the last 20 years is an enigma20,115.
Fast detection of fluconazole-resistant C. parapsilosis isolates
The emergence of fluconazole-resistant strains represents both a therapeutic and epidemiological challenge. Empirical treatment may be ineffective in such cases, potentially delaying appropriate antifungal therapy and thereby increasing patient mortality. Traditional antifungal susceptibility testing methods, such as broth microdilution (CLSI or EUCAST), although accurate, require 24–48 h after fungal growth is achieved. This slow turnaround time compromises early antifungal decision-making. Therefore, the availability of rapid detection tools would be a game-changer allowing antifungal treatment to be adjusted in hours rather than days, improving clinical outcomes and facilitating the control of nosocomial outbreaks.
Among phenotypic approaches, direct gradient strips directly performed on positive blood cultures may speed antifungal susceptibility results117. While this method has proven useful for fast detection of antifungal resistance in N. glabratus and C. albicans118,119, it has not yet been validated for fluconazole-resistant C. parapsilosis. Fluconazole-supplemented agar plates are a low-cost alternative used as screening tool for identifying colonizing or invasive resistant strains, including C. auris120,121. This approach has been shown to be effective in detecting fluconazole-resistant C. parapsilosis strains during hospital outbreaks122. The MALDI-TOF MS platform, now commonly used for yeast identification in clinical labs, has expanded toward antifungal susceptibility testing. New methods such as Composite Correlation Index (CCI) and MBT ASTRA (Antifungal Susceptibility Test Rapid Assay) combine mass spectrometry with fungal growth assessment in the presence of antifungals123,124,125. These systems provide results in 3–8 h, with high concordance compared to reference standard methods (CLSI/EUCAST), enabling phenotypic classification of susceptible or resistant isolates without extended incubation. One study showed that MALDI-TOF MS could detect echinocandin resistance in the C. parapsilosis complex after just 3 h of incubation, with 95% concordance with CLSI results (for anidulafungin and other drugs), supporting its potential use for fluconazole resistance detection as long as drug concentrations are optimized126. Although most studies have focused on C. albicans and N. glabratus, their principles are applicable to C. parapsilosis.
An alternative approach is molecular testing: point mutations in the ERG11 gene—particularly leading to the Y132F and G458S substitutions—have enabled the development of PCR-based protocols for rapid detection of resistant strains, either directly from fungal DNA or from cultured isolates80,127,128,129. A recent study developed conventional and real-time PCR assays to detect the Y132F substitution in pure cultures, skipping DNA extraction130. Both approaches showed 100% sensitivity and specificity; the conventional PCR delivered results in ~3 h, while real-time PCR required only 1.5 h130. A rapid method based on real-time PCR and high-resolution melting curve analysis (HRM) was developed to simultaneously detect SNPs in ERG11 leading to both Y132F and G458S substitutions131. This approach differentiated between homozygous and heterozygous strains, showing high specificity, and optimal sensitivity at a threshold of ~5 × 10⁵ cells per reaction. However, strict PCR conditions are required to avoid false positives.
The fast and accurate detection of resistant clones is key for outbreak control. Combining mutation-targeted PCRs with genotyping methods will enable early detection of transmissible resistant clones, particularly in healthcare settings. Since 2019, nosocomial outbreaks of fluconazole-resistant C. parapsilosis have been increasingly reported15. These outbreaks were tagged by using molecular typing techniques such as microsatellite markers130, multilocus sequence typing (MLST), or whole genome sequencing (WGS)20. As an alternative to PCR-based genotyping methods, MALDI-TOF combined with deep learning132 and Fourier-transform infrared spectroscopy (FTIR) using systems like IR Biotyper® has shown promising results for outbreak tracking. While it does not directly detect resistance, FTIR can generate biochemical fingerprints that distinguish clonally related resistant strains. For example, two recent studies demonstrated that FTIR correctly clustered fluconazole-resistant C. parapsilosis strains harbouring the Y132F substitution, with 74% concordance compared to microsatellite genotyping133,134. Both genotyping and FTIR allow the detection of clonal patterns of transmission and guide infection control measurement; these can be useful tools in epidemiological surveillance and hospital outbreak control. Rapid detection and molecular surveillance should be integral elements of outbreak handling in hospital settings. This multidimensional approach not only improves early therapeutic decisions but also strengthens infection prevention and control strategies in healthcare institutions.
While targeted PCR and accurate detection of ERG11 gene mutations provide mutation-based prediction of fluconazole resistance in a matter of hours, they do not replace phenotypic antifungal susceptibility testing. However, they are highly valuable in epidemiological surveillance and in settings where prevalent mutations are known. Notably, current PCR methods are still limited to the detection of a few mutations leading to amino acid changes (e.g., Y132F, G458S). In the future, commercial multiplex PCR panels (e.g., FilmArray BCID, T2Candida) should incorporate resistance markers detection, although current platforms primarily focus on species identification rather than antifungal resistance. Since all these methods have limitations (Table 3), future research should focus on specific validations on the fluconazole resistance detection in C. parapsilosis, improving standardisation, expanding the mutations detectable by PCR, and demonstrating their direct impact on drug susceptibility and clinical outcomes.
Unanswered questions
Despite significant advances, several critical questions about C. parapsilosis remain unanswered, limiting our understanding of its epidemiology, transmission dynamics, and resistance mechanisms. As discussed above, C. parapsilosis is commonly associated with nosocomial (hospital-acquired) infections, but the exact transmission dynamics remain unclear although efficient tools are now available (whole-genome sequencing, microsatellite genotyping) for detailed molecular epidemiological investigations, enabling the identification of clonal clusters, distinguishing epidemiologically related strains, and tracing possible routes of transmission such as patient-to-patient spread mediated by healthcare workers’ hands or contaminated hospital environments. What are the underlying factors driving the rapid and aggressive emergence of antifungal resistance in this species? Why are there such differences in the prevalence of fluconazole resistance between hospitals? Is person-to-person transmission more common than initially believed, and what role do environmental reservoirs and hospital equipment play in sustaining outbreaks? Although one explanation for the rapid increase and spread of fluconazole-resistant strains may be related to high fluconazole exposure and poor hand hygiene, the absence (or low) fitness costs of ERG11 substitutions, also supported by recent findings in C. albicans, may play a role84,135. Interestingly, biofilm production, which might account for the persistence of these strains, could facilitate their persistence in the environment and potentially their spread. However, there are some contradictory results on whether azole-resistant isolates produce less or more biofilm76,136. More research is needed to determine whether azole-resistant strains involved in clinical outbreaks may exhibit increased resistance to disinfection protocols.
What should we do now? Roadmap for the future
The emergence of azole-resistant C. parapsilosis capable of spreading and causing hospital outbreaks is a major concern that requires action to prevent this phenomenon from following the same trajectory as C. auris137. When isolation in pure culture confirms C. parapsilosis as cause of invasive infections, antifungal susceptibility testing should be prioritized (along with molecular genotyping when an outbreak is suspected). In fact, the results can not only inform clinical decisions regarding treatment options, but also facilitate the early detection of resistant isolates with the potential to cause uncontrolled outbreaks, thereby helping to prevent patients from acquiring new infections. Nipping a potential outbreak in the bud also translates to better outcomes for patients and less need to use more expensive drugs alternative to fluconazole. In this scenario, detecting antifungal resistance through phenotypic testing and simultaneously scan the mechanism of resistance would be the most helpful.
An ideal diagnostic workflow would combine multiple methods including rapid identification of C. parapsilosis by MALDI-TOF, simultaneous PCR targeting mutations known to be associated with resistance, rapid antifungal susceptibility testing (AFST) using MALDI-TOF with MBT ASTRA or CCI-performed alongside regular AFST to phenotypically assess antifungal resistance- and DNA-based molecular genotyping—or FTIR—to assess the relatedness between isolates and rule out potential nosocomial outbreaks. This integrated approach would optimize turnaround time and provide valuable epidemiological insights, thereby reducing therapeutic errors, limiting hospital transmission, and strengthening prevention strategies. Further research should include validation of existing methods specifically for fluconazole resistance detection in C. parapsilosis, expansion of the mutation coverage of PCR-based tools, the standardization of protocols across laboratories, and the demonstration of the impact of rapid resistance detection on patient outcomes. Lastly, the wider adoption of WGS in clinical laboratories—enabled by decreasing costs and the availability of robust, standardized pipelines—will enhance our understanding of its resistome, facilitate the detection of acquired resistance, and strengthen epidemiological surveillance of resistant C. parapsilosis clones.
Overall, establishing consensual guidelines for the management of C. parapsilosis in hospital settings—including environmental cleaning and disinfection procedures—would be highly valuable. Existing guidelines for C. auris infection control, such as those issued by the CDC138 and the UK139, provide a useful reference, as both opportunistic yeasts share important features, notably their ability to spread between humans and persist in the environment137.
Finally, the concerning spread of C. parapsilosis resistant clones poses a difficult challenge that only a holistic approach combining phenotypic, molecular, and epidemiological tools can win, by improving early therapeutic decisions and strengthening hospital infection control strategies.
Data availability
No datasets were generated or analysed during the current study.
References
Turner, S. A. & Butler, G. The Candida pathogenic species complex. Cold Spring Harb. Perspect. Med. 4, 1–18 (2014).
Mixão, V. et al. Genome analysis of five recently described species of the CUG-Ser clade uncovers Candida theae as a new hybrid lineage with pathogenic potential in the Candida parapsilosis species complex. DNA Res. 29, dsac010 (2022).
Del Olmo, V. & Gabaldón, T. Hybrids unleashed: exploring the emergence and genomic insights of pathogenic yeast hybrids. Curr. Opin. Microbiol 80, 102491 (2024).
Govrins, M. & Lass-Flörl, C. Candida parapsilosis complex in the clinical setting. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-023-00961-8 (2023).
Arendrup, M. C. et al. European candidaemia is characterised by notable differential epidemiology and susceptibility pattern: results from the ECMM Candida III study. J. Infect. 87, 428–437 (2023).
Lass-Flörl, C. et al. Invasive candidiasis. Nat. Rev. Dis. Prim. 10, 20 (2024).
Asogan, M. et al. Candida parapsilosis: a systematic review to inform the World Health Organization fungal priority pathogens list. Med. Mycol. 62, myad131 (2024).
Horn, D. L. et al. Epidemiology and outcomes of candidemia in 2019 patients: data from the prospective antifungal therapy alliance registry. Clin. Infect. Dis. 48, 1695–1703 (2009).
Salmanton-García, J. et al. Attributable mortality of candidemia - Results from the ECMM Candida III multinational European Observational Cohort Study. J. Infect. 89, 106229 (2024).
Zhai, B. et al. High-resolution mycobiota analysis reveals dynamic intestinal translocation preceding invasive candidiasis. Nat. Med. 26, 59–64 (2020).
Pammi, M., Holland, L., Butler, G., Gacser, A. & Bliss, J. M. Candida parapsilosis is a significant neonatal pathogen: a systematic review and meta-analysis. Pediatr. Infect. Dis. J. 32, e206–e216 (2013).
Neji, S. et al. Virulence factors, antifungal susceptibility and molecular mechanisms of azole resistance among Candida parapsilosis complex isolates recovered from clinical specimens. J. Biomed. Sci. 24, 67 (2017).
Moyes, D. L. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016).
Richardson, J. P. et al. Candidalysins Are a New Family of Cytolytic Fungal Peptide Toxins. mBio 13, e0351021 (2022).
Daneshnia, F. et al. Worldwide emergence of fluconazole-resistant Candida parapsilosis: current framework and future research roadmap. Lancet Microbe 4, e470–e480 (2023).
Gabaldón, T. Threats from the Candida parapsilosis complex: the surge of multidrug resistance and a hotbed for new emerging pathogens. Microbiol. Mol. Biol. Rev. 88, e0002923 (2024).
Bartalucci, C., Vena, A., Giacobbe, D. R. & Bassetti, M. Clinical impact of fluconazole-resistant Candida parapsilosis: a narrative review. Fut. Microbiol. 1–10, https://doi.org/10.1080/17460913.2025.2544443 (2025).
Vena, A. et al. Impact of fluconazole resistance on the outcomes of patients with Candida parapsilosis bloodstream infections: a retrospective multicenter study. Clin. Infect. Dis. 80, 540–550 (2025).
Semet, C. et al. Risk factors and outcome for bloodstream infections due to fluconazole-resistant Candida parapsilosis: a 22-year single-center retrospective study. Antimicrob. Resist. Infect. Control 14, 49 (2025).
Brassington, P. J. T. et al. Genomic reconstruction of an azole-resistant Candida parapsilosis outbreak and the creation of a multi-locus sequence typing scheme: a retrospective observational and genomic epidemiology study. Lancet Microbe 6, 100949 (2025).
Díaz-García, J. et al. Evidence of fluconazole-resistant Candida parapsilosis genotypes spreading across hospitals located in Madrid, Spain and Harboring the Y132F ERG11p Substitution. Antimicrobial Agents Chemother. 66, e0071022 (2022).
Fisher, M. C. & Denning, D. W. The WHO fungal priority pathogens list as a game-changer. Nat. Rev. Microbiol. 21, 211–212 (2023).
Bonassoli, L. A., Bertoli, M. & Svidzinski, T. I. E. High frequency of Candida parapsilosis on the hands of healthy hosts. J. Hospital Infect. 59, 159–162 (2005).
Cordeiro, R. deA. et al. Candida parapsilosis complex in veterinary practice: a historical overview, biology, virulence attributes and antifungal susceptibility traits. Vet. Microbiol. 212, 22–30 (2017).
Dornelles, G. et al. Harris’ hawk (Parabuteo unicinctus) as a source of pathogenic human yeasts: a potential risk to human health. Fut. Microbiol. 17, 169–175 (2022).
Glushakova, A. M., Rodionova, E. N. & Kachalkin, A. V. Yeasts in Feces of Pigeons (Columba livia) in the City of Moscow. Curr. Microbiol. https://doi.org/10.1007/s00284-020-02251-5 (2020).
Castelo-Branco, D. et al. Collateral consequences of agricultural fungicides on pathogenic yeasts: a one health perspective to tackle azole resistance. Mycoses 65, 303–311 (2022).
Castelo-Branco, D. de S. C. M. et al. Azole resistance in Candida from animals calls for the One Health approach to tackle the emergence of antimicrobial resistance. Med. Mycol. 1–10, https://doi.org/10.1093/mmy/myz135 (2020).
Opulente, D. A. et al. Pathogenic budding yeasts isolated outside of clinical settings. FEMS Yeast Res. 19, 1–6 (2019).
Morio, F. Dear medical mycologists, it is time to look outside the box. FEMS Yeast Res. 20, foz080 (2020).
Vadkertiova, R., Dudasova, H. & Balascakova, M. Yeasts in agricultural and managed soils. In Yeasts in Natural Ecosystems: Diversity 1–499, https://doi.org/10.1007/978-3-319-62683-3 (2017).
Lo, H.-J. et al. Fruits as the vehicle of drug resistant pathogenic yeasts. J. Infect. 75, 254–262 (2017).
Glushakova, АМ & Kachalkin, АV. Yeast community succession in cow dung composting process. Fungal Biol. 127, 1075–1083 (2023).
Ramos, L. S. et al. Candida spp. isolated from recreational coastal waters of Rio de Janeiro - Brazil: focus on antifungal resistance and virulence attributes. Sci. Total Environ. 947, 174662 (2024).
Liu, Z., Rossi, J. M. & Myers, L. C. Candida albicans Zn cluster transcription factors Tac1 and Znc1 are activated by farnesol to upregulate a transcriptional program including the multidrug efflux pump CDR1. Antimicrobial Agents Chemother. 62, e00968–18 (2018).
Arroyo, M. G. et al. Broad diversity of fungi in hospital water. Sci. World J. 2020, 1–6 (2020).
Prigitano, A. et al. ICU environmental surfaces are a reservoir of fungi: species distribution in northern Italy. J. Hosp. Infect. 123, 74–79 (2022).
Piedrahita, C. T. et al. Environmental surfaces in healthcare facilities are a potential source for transmission of Candida auris and Other Candida Species. Infect. Control Hosp. Epidemiol. 38, 1107–1109 (2017).
Allert, S. et al. From environmental adaptation to host survival: Attributes that mediate pathogenicity of Candida auris. Virulence 13, 191–214 (2022).
Dögen, A. et al. Candida parapsilosis in domestic laundry machines. Med. Mycol. 55, 813–819 (2017).
Fu, L. et al. Different efficacies of common disinfection methods against Candida auris and other Candida species. J. Infect. Public Health 13, 730–736 (2020).
Cadnum, J. L. et al. Effectiveness of disinfectants against Candida auris and other Candida Species. Infect. Control Hosp. Epidemiol. 38, 1240–1243 (2017).
Cavalheiro, M. & Teixeira, M. C. Candida biofilms: threats, challenges, and promising strategies. Front. Med. 5, 28 (2018).
Tóth, R. et al. Candida parapsilosis: from genes to the bedside. Clin. Microbiol. Rev. 32, e00111–18 (2019).
Heaney, H. et al. The environmental stress sensitivities of pathogenic Candida species, including candida auris, and implications for their spread in the hospital setting. Med. Mycol. 58, 744–755 (2020).
Lombardi, L. et al. Alternative sulphur metabolism in the fungal pathogen Candida parapsilosis. Nat. Commun. 15, 9190 (2024).
Garcia-Effron, G., Katiyar, S. K., Park, S., Edlind, T. D. & Perlin, D. S. A naturally occurring proline-to-alanine amino acid change in Fks1p in Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis accounts for reduced echinocandin susceptibility. Antimicrob. Agents Chemother. 52, 2305–2312 (2008).
Guinea, J. et al. Molecular identification and antifungal susceptibility of yeast isolates causing fungemia collected in a population-based study in Spain in 2010 and 2011. Antimicrob. Agents Chemother. 58, 1529–1537 (2014).
Díaz-García, J. et al. Yeasts from blood cultures in the wake of the COVID-19 pandemic in a tertiary care hospital: shift in species epidemiology, steady low antifungal resistance and full in vitro ibrexafungerp activity. Med. Mycol. 61, myad072 (2023).
Arastehfar, A. et al. Genetically related micafungin-resistant Candida parapsilosis blood isolates harbouring novel mutation R658G in hotspot 1 of Fks1p: a new challenge?. J. Antimicrobial. Chemother. 76, 418–422 (2021).
Ning, Y. et al. Decreased echinocandin susceptibility in Candida parapsilosis causing candidemia and emergence of a pan-echinocandin resistant case in China. Emerg. Microbes Infect. 12, 2153086 (2023).
Siopi, M. et al. Pan-echinocandin resistant C. parapsilosis harboring an F652S Fks1 alteration in a patient with prolonged echinocandin therapy. J. Fungi 8, 931 (2022).
Daneshnia, F. et al. Determinants of fluconazole resistance and echinocandin tolerance in C. parapsilosis isolates causing a large clonal candidemia outbreak among COVID-19 patients in a Brazilian ICU. Emerg. Microbes Infect. 11, 2264–2274 (2022).
Pfaller, M. A., Diekema, D. J., Turnidge, J. D., Castanheira, M. & Jones, R. N. Twenty years of the SENTRY antifungal surveillance program: results for Candida species from 1997-2016. Open Forum Infect. Dis. 6, S79–S94 (2019).
Thomaz, D. Y. et al. Environmental clonal spread of azole-resistant Candida parapsilosis with Erg11-Y132F mutation causing a large candidemia outbreak in a Brazilian cancer referral center. J. Fungi 7, 259 (2021).
Magobo, R. E., Lockhart, S. R. & Govender, N. P. Fluconazole-resistant Candida parapsilosis strains with a Y132F substitution in the ERG11 gene causing invasive infections in a neonatal unit, South Africa. Mycoses 63, 471–477 (2020).
Siopi, M. et al. Increase in candidemia cases and emergence of fluconazole-resistant Candida parapsilosis and C. auris isolates in a tertiary care academic hospital during the COVID-19 pandemic, Greece, 2020 to 2023. Eur. Surveill. 29, 2300661 (2024).
Díaz-García, J. et al. Monitoring the epidemiology and antifungal resistance of yeasts causing fungemia in a tertiary care hospital in madrid, spain: any relevant changes in the last 13 years?. Antimicrob. Agents Chemother. 65, e01827–20 (2021).
Caggiano, G. et al. Genotypic and phenotypic characteristics of Candida parapsilosis bloodstream isolates: health care associated infections in a teaching Hospital in Italy. J. Infect. Public Health 17, 967–974 (2024).
McHugh, J., Chesdachai, S., Dunsirn, M., Wengenack, N. & Vergidis, P. Increasing fluconazole resistance in Candida parapsilosis: a 10-Year Analysis of Blood Culture Isolates at a US Reference Laboratory (2015-2024). J. Infect. Dis. jiaf249 https://doi.org/10.1093/infdis/jiaf249 (2025).
Butler, G. et al. Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature 459, 657–662 (2009).
Skrzypek, M. S. et al. The Candida Genome Database (CGD): incorporation of Assembly 22, systematic identifiers and visualization of high throughput sequencing data. Nucleic Acids Res. 45, D592–D596 (2017).
Mutalová, S. et al. Chromosome-level genome assembly of an auxotrophic strain of the pathogenic yeast Candida parapsilosis. Microbiol. Resour. Announc. 13, e0034724 (2024).
Grossman, N. T., Pham, C. D., Cleveland, A. A. & Lockhart, S. R. Molecular mechanisms of fluconazole resistance in Candida parapsilosis isolates from a U.S. surveillance system. Antimicrob. Agents Chemother. 59, 1030–1037 (2015).
Berkow, E. L. et al. Multidrug transporters and alterations in sterol biosynthesis contribute to azole antifungal resistance in Candida parapsilosis. Antimicrob. Agents Chemother. 59, 5942–5950 (2015).
Souza, A. C. R. et al. Candida parapsilosis resistance to fluconazole: molecular mechanisms and in vivo impact in infected Galleria mellonella larvae. Antimicrob. Agents Chemother. 59, 6581–6587 (2015).
Singh, A. et al. Emergence of clonal fluconazole-resistant Candida parapsilosis clinical isolates in a multicentre laboratory-based surveillance study in India. J. Antimicrob. Chemother. 74, 1260–1268 (2019).
Choi, Y. J. et al. Fluconazole-resistant Candida parapsilosis bloodstream isolates with Y132F mutation in ERG11 gene, South Korea. Emerg. Infect. Dis. 24, 1768–1770 (2018).
Castanheira, M., Deshpande, L. M., Messer, S. A., Rhomberg, P. R. & Pfaller, M. A. Analysis of global antifungal surveillance results reveals predominance of Erg11 Y132F alteration among azole-resistant Candida parapsilosis and Candida tropicalis and country-specific isolate dissemination. Int. J. Antimicrob. Agents 55, 105799 (2020).
Martini, C. et al. Prevalence and clonal distribution of azole-resistant Candida parapsilosis isolates causing bloodstream infections in a large Italian hospital. Front. Cell. Infect. Microbiol. 10, 232 (2020).
Ning, Y.-T. et al. Emergence of multiple fluconazole-resistant Candida parapsilosis sensu stricto clones with persistence and transmission in China. J. Antimicrob. Chemother. 79, 128–133 (2024).
Thomaz, D. Y. et al. An azole-resistant Candida parapsilosis outbreak: clonal persistence in the intensive care unit of a Brazilian Teaching Hospital. Front. Microbiol. 9, 2997 (2018).
Hare, R. K. et al. Candidemia among hospitalized pediatric patients caused by several clonal lineages of Candida parapsilosis. J. Fungi 8, 183 (2022).
Demirci-Duarte, S., Arikan-Akdagli, S. & Gülmez, D. Species distribution, azole resistance and related molecular mechanisms in invasive Candida parapsilosis complex isolates: Increase in fluconazole resistance in 21 years. Mycoses 64, 823–830 (2021).
Corzo-Leon, D. E., Peacock, M., Rodriguez-Zulueta, P., Salazar-Tamayo, G. J. & MacCallum, D. M. General hospital outbreak of invasive candidiasis due to azole-resistant Candida parapsilosis associated with an Erg11 Y132F mutation. Med. Mycol. https://doi.org/10.1093/mmy/myaa098 (2020).
Arastehfar, A. et al. First report of candidemia clonal outbreak caused by emerging fluconazole-resistant Candida parapsilosis isolates harboring Y132F and/or Y132F+K143R in Turkey. Antimicrob. Agents Chemother. 64, e01001–e01020 (2020).
Arastehfar, A. et al. Evaluation of molecular epidemiology, clinical characteristics, antifungal susceptibility profiles, and molecular mechanisms of antifungal resistance of Iranian Candida parapsilosis species complex blood isolates. Front. Cell Infect. Microbiol. 10, 206 (2020).
Arastehfar, A. et al. Clonal candidemia outbreak by Candida parapsilosis carrying Y132F in Turkey: evolution of a persisting challenge. Front. Cell Infect. Microbiol. 11, 676177 (2021).
Bergin, S. et al. Analysis of clinical Candida parapsilosis isolates reveals copy number variation in key fluconazole resistance genes. Antimicrob. Agents Chemotherapy e0161923, https://doi.org/10.1128/aac.01619-23 (2024).
Asadzadeh, M., Ahmad, S., Al-Sweih, N. & Khan, Z. Epidemiology and Molecular Basis of Resistance to Fluconazole Among Clinical Candida parapsilosis Isolates in Kuwait. Microb. Drug Resist. 23, 966–972 (2017).
Asadzadeh, M., Dashti, M., Ahmad, S., Alfouzan, W. & Alameer, A. Whole-genome and targeted-amplicon sequencing of fluconazole-susceptible and -resistant Candida parapsilosis isolates from Kuwait reveals a previously undescribed N1132D polymorphism in CDR1. Antimicrob. Agents Chemother. 65, 1–3 (2021).
Sharma, V. et al. Genotypic diversity and molecular basis of fluconazole resistance in Candida parapsilosis clinical isolates collected over 7 years in a tertiary-care hospital in North India. Mycoses 68, e70062 (2025).
Doorley, L. A., Barker, K. S., Zhang, Q., Rybak, J. M. & Rogers, P. D. Mutations in TAC1 and ERG11 are major drivers of triazole antifungal resistance in clinical isolates of Candida parapsilosis. Clin. Microbiol. Infect. 29, 1602.e1–1602.e7 (2023).
Hartuis, S. et al. Precise genome editing underlines the distinct contributions of mutations in ERG11, ERG3, MRR1, and TAC1 genes to antifungal resistance in Candida parapsilosis. Antimicrob. Agents Chemother. 68, e0002224 (2024).
Branco, J. et al. Impact of ERG3 mutations and expression of ergosterol genes controlled by UPC2 and NDT80 in Candida parapsilosis azole resistance. Clin. Microbiol. Infect. 23, 575.e1–575.e8 (2017).
Branco, J. et al. Fluconazole and voriconazole resistance in Candida parapsilosis is conferred by gain-of-function mutations in MRR1 transcription factor gene. Antimicrob. Agents Chemother. 59, 6629–6633 (2015).
Doorley, L. A. et al. Candida parapsilosis Mdr1B and Cdr1B are drivers of Mrr1-mediated clinical fluconazole resistance. Antimicrob. Agents Chemotherapy e0028922, https://doi.org/10.1128/aac.00289-22 (2022).
McTaggart, L. R., Cabrera, A., Cronin, K. & Kus, J. V. Antifungal susceptibility of clinical yeast isolates from a large canadian reference laboratory and application of whole-genome sequence analysis to elucidate mechanisms of acquired resistance. Antimicrob. Agents Chemother. 64, e00402–e00420 (2020).
Kim, T. Y. et al. Evolution of fluconazole resistance mechanisms and clonal types of Candida parapsilosis isolates from a tertiary care hospital in South Korea. Antimicrob. Agents Chemother. 66, e0088922 (2022).
Branco, J. et al. Clinical azole cross-resistance in Candida parapsilosis is related to a novel MRR1 gain-of-function mutation. Clin. Microbiol. Infect. 28, 1655.e5–1655.e8 (2022).
Franconi, I. et al. The S862C amino acid change in CpMrr1 confers fluconazole resistance in Candida parapsilosis. JAC Antimicrob. Resist 7, dlaf051 (2025).
Kannan, A. et al. Comparative Genomics for the Elucidation of Multidrug Resistance in Candida lusitaniae. mBio 10, S1–S159 (2019).
Rajesh-Khanna, D.-S. et al. Coordinated regulation of Mdr1- and Cdr1-mediated protection from antifungals by the Mrr1 transcription factor in emerging Candida spp. mBio. e0132325, https://doi.org/10.1128/mbio.01323-25 (2025).
Arastehfar, A. et al. The quiet and underappreciated rise of drug-resistant invasive fungal pathogens. J. Fungi 6, 138 (2020).
Rybak, J. M. et al. Loss of C-5 sterol desaturase activity results in increased resistance to azole and echinocandin antifungals in a clinical isolate of Candida parapsilosis. Antimicrob. Agents Chemother. 61, 1–13 (2017).
Papp, C. et al. Triazole evolution of Candida parapsilosis results in cross-resistance to other antifungal drugs, influences stress responses, and alters virulence in an antifungal drug-dependent manner. mSphere 5, 1–15 (2020).
Khalifa, H. O., Watanabe, A. & Kamei, K. Genetic Mutations in FKS1 gene associated with acquired echinocandin resistance in Candida parapsilosis complex. Mycopathologia 189, 40 (2024).
Papp, C. et al. Echinocandin-induced microevolution of Candida parapsilosis influences virulence and abiotic stress tolerance. mSphere 3, e00547–18 (2018).
Daneshnia, F. et al. Candida parapsilosis isolates carrying mutations outside of FKS1 hotspot regions confer high echinocandin tolerance and facilitate the development of echinocandin resistance. Int. J. Antimicrob. Agents 106831, https://doi.org/10.1016/j.ijantimicag.2023.106831 (2023).
Martí-Carrizosa, M., Sánchez-Reus, F., March, F., Cantón, E. & Coll, P. Implication of Candida parapsilosis FKS1 and FKS2 mutations in reduced echinocandin susceptibility. Antimicrob. Agents Chemother. 59, 3570–3573 (2015).
Hoenigl, M. et al. Novel antifungals and treatment approaches to tackle resistance and improve outcomes of invasive fungal disease. Clin. Microbiol. Rev. 37, e00074–23 (2024).
Pfaller, M. A., Huband, M. D., Rhomberg, P. R., Bien, P. A. & Castanheira, M. Activities of manogepix and comparators against 1435 recent fungal isolates collected during an International Surveillance Program (2020). Antimicrob. Agents Chemother. 66, e0102822 (2022).
Kapoor, M., Moloney, M., Soltow, Q. A., Pillar, C. M. & Shaw, K. J. Evaluation of resistance development to the Gwt1 inhibitor manogepix (APX001A) in Candida species. Antimicrob. Agents Chemother. 64, 1–12 (2019).
Arendrup, M. C. & Jørgensen, K. M. Manogepix (APX001A) displays potent in vitro activity against human pathogenic yeast, but with an unexpected correlation to fluconazole MICs. Antimicrob. Agents Chemother. 64, e00429–20 (2020).
Barker, K. S. et al. Mutations in transcription factors that confer fluconazole resistance also confer reduced susceptibility to manogepix in Candida auris (Candidozyma auris), Candida albicans, Candida parapsilosis, and Candida glabrata (Nakaseomyces glabratus). Antimicrob. Agents Chemother. e00680-25, https://doi.org/10.1128/aac.00680-25 (2025).
Yang, F. et al. Aneuploidy underlies tolerance and cross-tolerance to drugs in Candida parapsilosis. Microbiol. Spectr. 9, e0050821 (2021).
Sun, L. L. et al. Aneuploidy enables cross-tolerance to unrelated antifungal drugs in Candida parapsilosis. Front. Microbiol. 14, 1137083 (2023).
Ning, Y., Dai, R., Zhang, L., Xu, Y. & Xiao, M. Copy number variants of ERG11: mechanism of azole resistance in Candida parapsilosis. Lancet Microbe 5, e10 (2024).
Zhai, B. et al. Antifungal heteroresistance causes prophylaxis failure and facilitates breakthrough Candida parapsilosis infections. Nat. Med. 30, 3163–3172 (2024).
Gautier, C., Maciel, E. I. & Ene, I. V. Approaches for identifying and measuring heteroresistance in azole-susceptible Candida isolates. Microbiol. Spectr. 12, e0404123 (2024).
Schutz, K. S. et al. Polygenetic determinants of azole resistance, tolerance, and heteroresistance in Candida albicans. bioRxiv, https://doi.org/10.1101/2025.06.29.662174 (2025).
Hérivaux, A., Papon, N. & Morio, F. Heteroresistance: a hidden cause behind antifungal prophylaxis failure. Trends Microbiol. 32, 1042–1044 (2024).
Fekkar, A. et al. Hospital outbreak of fluconazole-resistant Candida parapsilosis: arguments for clonal transmission and long-term persistence. Antimicrob. Agents Chemotherapy https://doi.org/10.1128/AAC.02036-20 (2021).
Díaz-García, J. et al. Candida genotyping of blood culture isolates from patients admitted to 16 hospitals in Madrid: genotype spreading during the COVID-19 pandemic driven by fluconazole-resistant C. parapsilosis. J. Fungi 8, 1228 (2022).
Mesquida, A. et al. Fluconazole-resistant Candida parapsilosis genotypes from hospitals located in five Spanish cities and one in Italy: description of azole-resistance profiles associated with the Y132F ERG11p substitution. Mycoses 67, e13706 (2024).
Alcoceba, E. et al. Fluconazole-resistant Candida parapsilosis clonally related genotypes: first report proving the presence of endemic isolates harbouring the Y132F ERG11 gene substitution in Spain. Clin. Microbiol. Infect. https://doi.org/10.1016/j.cmi.2022.02.025 (2022).
Guinea, J. et al. Rapid antifungal susceptibility determination for yeast isolates by use of Etest performed directly on blood samples from patients with fungemia. J. Clin. Microbiol. 48, 2205–2212 (2010).
Bordallo-Cardona, M. Á et al. Resistance to echinocandins in Candida can be detected by performing the etest directly on blood culture samples. Antimicrob. Agents Chemother. 62, e00162–18 (2018).
Escribano, P. et al. The Etest performed directly on blood culture bottles is a reliable tool for detection of fluconazole-resistant Candida albicans isolates. Antimicrob. Agents Chemother. 61, e00400–e00417 (2017).
Linares, M. J., Charriel, G., Solís, F. & Casal, M. CHROMAgar Candida with fluconazole: comparison with microdilution techniques]. Enferm. Infecc. Microbiol Clin. 21, 493–497 (2003).
Ruiz-Gaitán, A. et al. Usefulness of chromogenic media with fluconazole supplementation for presumptive identification of Candida auris. Diagnostics 13, 231 (2023).
Martin-Gomez, M. et al. Usefulness of Fluconazole-Supplemented Chromogenic Agar (FSCA) for the detection of fluconazole non-susceptible Candida parapsilosis (FNSCP) in clinical samples. In 32nd European Society of Clinical Microbiology and Infectious Diseases meeting. Lisbon (Portugal) from April 23 to 26 (2022).
De Carolis, E. et al. Are we ready for nosocomial candida auris infections? Rapid identification and antifungal resistance detection using MALDI-TOF mass spectrometry may be the answer. Front. Cell Infect. Microbiol. 11, 645049 (2021).
Vatanshenassan, M. et al. Proof of concept for MBT ASTRA, a rapid matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS)-based method to detect caspofungin resistance in Candida albicans and Candida glabrata. J. Clin. Microbiol. 56, e00420–18 (2018).
Knoll, M. A., Ulmer, H. & Lass-Flörl, C. Rapid antifungal susceptibility testing of yeasts and molds by MALDI-TOF MS: a systematic review and meta-analysis. J. Fungi 7, 63 (2021).
Roberto, A. E. M. et al. Rapid detection of echinocandins resistance by MALDI-TOF MS in Candida parapsilosis complex. Microorganisms 8, 109 (2020).
Hubler, C. M., Carvalhaes, C. G. & Castanheira, M. Recent increase in fluconazole-nonsusceptible Candida parapsilosis in a global surveillance with the expansion of the Erg11 Y132F genotype and a rapid detection method for this alteration. Diagn. Microbiol. Infect. Dis. 106, 115957 (2023).
Karthika, K., Premamalini, T., Vijayakishore, T. & Kindo, A. J. Standardization of polymerase chain reaction for detection of fluconazole resistance targeting Y132F mutation in ERG11 gene in Candida parapsilosis. Curr. Med. Mycol. 10, e2024.345209.1517 (2024).
Öztürk, S. et al. Rapid tetra-primer amplification refractory mutation system-polymerase chain reaction protocol for detection of Y132F mutation in fluconazole resistant Candida parapsilosis. Micro. Drug Resist. 30, 210–213 (2024).
Guinea, J. et al. Fluconazole-resistant Candida parapsilosis: fast detection of the Y132F ERG11p substitution, and a proposed microsatellite genotyping scheme. Clin. Microbiol. Infect. 30, 1447–1452 (2024).
Trevijano-Contador, N. et al. High-resolution melting assay to detect the mutations that cause the Y132F and G458S substitutions at the ERG11 gene involved in azole resistance in Candida parapsilosis. Mycoses 67, e13811 (2024).
Mohammad, N. et al. Improving the detection of epidemic clones in Candida parapsilosis outbreaks by combining MALDI-TOF mass spectrometry and deep learning approaches. Microorganisms 11, 1071 (2023).
De Carolis, E. et al. The Fourier-transform infrared spectroscopy-based method as a new typing tool for Candida parapsilosis clinical isolates. Microbiol. Spectr. 11, e0238823 (2023).
De Carolis, E. et al. Follow the path: unveiling an azole resistant Candida parapsilosis outbreak by FTIR spectroscopy and STR analysis. J. Fungi 10, 753 (2024).
Bédard, C. et al. Most azole resistance mutations in the Candida albicans drug target confer cross-resistance without intrinsic fitness cost. Nat. Microbiol. 9, 3025–3040 (2024).
El Hady, R. et al. Phenotypic and genotypic characterization of Candida parapsilosis complex isolates from a Lebanese hospital. Sci. Rep. 15, 4853 (2025).
Steixner, S., Bauer, A., Grubwieser, P. & Lass-Flörl, C. Can we prevent azole-resistant Candida parapsilosis from becoming the next Candidozyma auris? Expert Rev. Anti Infect. Ther. 1–3, https://doi.org/10.1080/14787210.2025.2544694 (2025).
Infection Control Guidance: Candida auris. https://www.cdc.gov/candida-auris/hcp/infection-control/index.html (2024).
Jesudason, T. UKHSA publishes updated guidance for C auris. Lancet Microbe. 101151 (2025).
Morio, F. et al. Precise genome editing using a CRISPR-Cas9 method highlights the role of CoERG11 amino acid substitutions in azole resistance in Candida orthopsilosis. J. Antimicrob. Chemother. 74, 2230–2238 (2019).
Morio, F., Loge, C., Besse, B., Hennequin, C. & Le Pape, P. Screening for amino acid substitutions in the Candida albicans Erg11 protein of azole-susceptible and azole-resistant clinical isolates: new substitutions and a review of the literature. Diagn. Microbiol. Infect. Dis. 66, 373–384 (2010).
Daneshnia, F. et al. Whole-genome sequencing confirms a persistent candidaemia clonal outbreak due to multidrug-resistant Candida parapsilosis. J. Antimicrob. Chemother. 78, 1488–1494 (2023).
Misas, E. et al. Molecular and Epidemiological Investigation of Fluconazole-resistant Candida parapsilosis-Georgia, United States, 2021. Open Forum Infect. Dis. 11, ofae264 (2024).
Thomaz, D. Y. et al. A Brazilian inter-hospital candidemia outbreak caused by fluconazole-resistant Candida parapsilosis in the COVID-19 Era. J. Fungi 8, 100 (2022).
Ceballos-Garzon, A. et al. Emergence and circulation of azole-resistant C. albicans, C. auris and C. parapsilosis bloodstream isolates carrying Y132F, K143R or T220L Erg11p substitutions in Colombia. Front. Cell Infect. Microbiol. 13, 1136217 (2023).
Zeeshan, M. et al. Fluconazole-resistant Candida parapsilosis complex candidemia and analysis of mutations in the ERG11 gene from Pakistan. Mycoses 67, e13677 (2024).
Abdolrasouli, A. et al. Cluster of clonal fluconazole-resistant Candida parapsilosis harbouring Y132F mutation in ERG11 gene in a neonatal ICU in the UK. J. Antimicrob. Chemother. 80, 887–890 (2025).
Acknowledgements
J.G. has received funds for participating in educational activities organized on behalf of Gilead, Pfizer, and Mundipharma; he has also received research funds from FIS, Gilead, F2G, Shionogi, Scynexis, Mundipharma, outside the submitted work. F.M. has received funds for participating in educational activities organized on behalf of Gilead, and Mundipharma; he has also received research funds from Pfizer, Gilead, Mundipharma, Agence Nationale de la Recherche and CAPES-COFECUB, outside the submitted work. F.M. acknowledges funding from Mundipharma to investigate antifungal resistance in yeasts.
Author information
Authors and Affiliations
Contributions
J.G., P.E., and F.M. conceived and designed the manuscript. All authors (J.G., P.E., M.C., L.L. and F.M.) participated in writing, critically reviewed the content, have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guinea, J., Escribano, P., Cadeau, M. et al. Emerging antifungal resistance in Candida parapsilosis: the end of the innocence. npj Antimicrob Resist 3, 99 (2025). https://doi.org/10.1038/s44259-025-00173-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00173-5