Abstract
Staphylococcus argenteus is implicated in various superficial and invasive diseases in humans, and multidrug resistance has been increasingly reported. Here, we aim to elucidate the bacterial features shaping the global spread of antimicrobial resistant S. argenteus. The global population contained a heterogeneous collection of antimicrobial resistance (AMR) genes, virulence genes and plasmid replicon types. Of the 13 sequence types (ST) present, ST2250 made up 56.2% of the dataset. We estimate its most recent common ancestor to have emerged in 1967. Two sublineages of ST2250 with distinct AMR gene content appeared separately in Asia and Europe in 1989 and 1992, respectively. The effective population size of ST2250 has soared in the past five decades, although there is indication of a recent decline. Overall, our results show that the AMR pool of S. argenteus is driven in part by the clonal expansion and subsequent diversification of ST2250 worldwide.
Similar content being viewed by others
Introduction
The coagulase-positive Staphylococcus argenteus is a close relative of the more well-known opportunistic pathogen Staphylococcus aureus. It was initially identified as a divergent lineage of S. aureus and was formerly known as clonal complex 75 (CC75)1. In 2015, S. argenteus was officially designated as a distinct species2. It has been implicated in a variety of superficial and invasive diseases, including toxin-mediated food poisoning3, skin and soft tissue infections4,5, bloodstream infections6,7, bone and joint infections8, and purulent lymphadenitis (infection of the lymph nodes)9. S. argenteus has also been documented in food products10 and diseased animals11,12. Reports of methicillin resistant and multidrug resistant strains of S. argenteus from human disease cases13,14,15 pose a grave concern, and instances of their transmission have been documented16,17. While the prevalence of S. argenteus in diseases relative to S. aureus remains low, misidentification with S. aureus and the lack of a systematic surveillance effort may have underestimated its prevalence.
Population genomic studies of S. argenteus are few, but those few reports have uncovered notable genomic features that underlie its growing importance as a pathogen. A genomic analysis of 153 globally distributed genomes strains from different geographical origins indicated that geographical distribution may have propelled the divergence of S. argenteus and its horizontal acquisition of mobile genetic elements18. In the Dutch national surveillance of methicillin-resistant S. aureus (MRSA), a total of 54 isolates were identified as S. argenteus, with the earliest methicillin resistant S. argenteus documented in 200815. In a retroactive assessment of Taiwanese staphylococcal bacteremia between 2010 and 2012, 47 cases that were previously attributed to S. aureus were determined to be due to S. argenteus infection6. In Canada and the United States, 22 S. argenteus isolates displayed evidence of considerable diversity in terms of sequence types (ST), with many genomic features related to virulence in common with those of S. aureus19. Analysis of 132 S. argenteus genomes from fourteen countries spanning over 13 years revealed the growing prevalence of sequence type (ST) 2250, which was notable for the presence of a CRISPR/Cas system that was absent in most other STs20. In Thailand, the rapid spread of the dominant ST2250 in the past 20 years is attributed to the acquisition of multiple exotoxin and antimicrobial resistance (AMR) genes that have been linked previously to livestock-associated S. aureus21. The global distribution of S. argenteus suggests international dissemination and the presence of geographic hotspots of the species in Southeast Asia, Australia, and the Amazon22.
Despite growing evidence of the clinical importance of S. argenteus, knowledge about the underlying causes of the growing threat of AMR emerging in the species remains limited. Here, we analyzed 379 genomes of S. argenteus to describe its global population structure and evolutionary history. Overall, our results show that the AMR pool of S. argenteus is driven in part by the global clonal expansion of ST2250 in the last five decades and subsequent divergence into sublineages with distinct AMR gene content. Continued surveillance of S. argenteus will be critical in developing effective strategies to reduce the risk for multidrug-resistant strains to emerge and further disseminate.
Results
The global S. argenteus population is phylogenetically diverse but dominated by ST2250
We retrieved a total of 379 high-quality genome sequences of S. argenteus derived from 28 countries in six continents, four ecological sources, and collection years of 2005–2023 (Fig. 1 and Supplementary Data 1). The median number of genes per genome was 2540 (range: 2333–2719) (Supplementary Fig. 1A). The genome fluidity23 was 0.109 (Supplementary Fig. 1B), indicating that on average, 10.9% of genes were unique (i.e., not shared) between genome pairs while 89.1% of genes were common between pairs. This low genome fluidity suggests that S. argenteus exhibits limited gene content variability. To further evaluate gene expansion dynamics, pan-genome openness was assessed using Heaps’ law24. The γ coefficient obtained for ST2250 was 0.064, while the value for the complete dataset was 0.084 (Supplementary Fig. 1C). Although the pangenome is considered open, the low γ values observed are consistent with the genome fluidity results and indicate restricted gene content variability in S. argenteus.
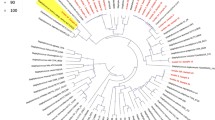
A Midpoint-rooted maximum likelihood tree showing the phylogenetic relationships of globally distributed S. argenteus genomes. The tree was built using 59,124 single nucleotide polymorphisms (SNPs) derived from aligned and concatenated sequences of 2180 core genes (i.e., genes present 99% of genomes). Tree scale represents number of nucleotide substitutions per site. The colored lines emanating from the branches of the phylogeny are colored by sequence type (ST) with only the most common six STs shown in color while less common STs are grouped as others (i.e., present in fewer than 10 genomes or without ST assignment). The four outer rings (from inner to outer) represent the SCCmec type, source, continent and time period. For visual clarity, only the six major STs are shown. B World map showing the major geographical regions from where the 379 S. argenteus were derived. The pie charts show the distribution of the six major ST per continent. The colors in the pie charts represent the STs following the color scheme in (A). The number next to each pie chart shows the total number of genomes per region. C Proportion of STs and number of genomes per year. The stacked colors in each bar represent the STs following the color scheme in (A). The numbers above each bar represent the total number of genomes per year. D Number of genomes per ST. The stacked colors in each bar represent the SCCmec types, following the color scheme in (A). The numbers above each bar indicate the total number of genomes of each ST. Detailed information of the features and associated metadata of each genome is presented in Supplementary Data 1.
Multi-locus sequence typing (MLST) analysis revealed the presence of 13 previously recognized STs (Supplementary Data 1). Six sequence types (ST) comprised 88.39% (335/379) of the genomes in the population: ST2250 (n = 213 genomes), ST1223 (n = 44), ST2198 (n = 24), ST2793 (n = 23), ST5961 (n = 17), and ST2854 (n = 14) (Fig. 1A). ST2250 alone accounted for 56.2% of all genomes. The remaining seven STs were each found in eight or fewer genomes and included both known and unassigned STs. Six other STs were represented by a single genome.
To further explore the global population structure, we constructed a maximum likelihood phylogenetic tree based on the alignment of 59,124 single nucleotide polymorphisms (SNPs) derived from 2180 core genes (Fig. 1A and Supplementary Data 2). Each of the six major STs was located on a long branch of the phylogenetic tree and formed a distinct monophyletic cluster, indicating deep time divergence from other STs. The very short branches on each of the phylogenetic clusters of the six major ST indicate strong genetic homogeneity within each ST. Genomes from disparate continents and ecological sources were intermingled throughout the tree. Although no strict segregation by geography or source was observed, the phylogeny revealed trends of partial clustering. For instance, ST5961 comprised predominantly of genomes from food and the environment and was mainly found in Asia. In contrast, ST2793 included a large number of European genomes, while ST2198 was primarily associated with North American genomes (Fig. 1B). The majority of genomes used in this study were of human origin (n = 312), representing 82.3% of the total dataset. Within the predominant lineage ST2250, humans were the main source (n = 205). Among these human-associated ST2250 genomes, most genomes originated from Asia (112/205) and Europe (74/205). Beginning in the year 2008 onwards, ST2250 was consistently present every year (Fig. 1C).
The mobile genetic element Staphylococcal Cassette Chromosome mec (SCCmec) is a unique feature of methicillin-resistant Staphylococcus species25. It carries the mecA gene encoding a low-affinity penicillin-binding protein 2a (PBP2a) that confers resistance to beta-lactam antibiotics26. In S. aureus, 15 structurally distinct types of SCCmec have been identified26,27. In our S. argenteus dataset, we found SCCmec present in 106 genomes (Fig. 1D and Supplementary Data 1). We identified five types of SCCmec: Ia (n = 2 genomes), IVa (n = 13), IVb (n = 1), IVc (n = 89), and IVd (n = 1). SCCmec type IVc elements were frequently detected in ST1223, ST2250, and ST2793, in 13.64%, 30.51%, and 95.65% of the genomes of each ST, respectively.
A variety of virulence genes is present in ST2250 and in less common STs
We screened all genomes for the presence and distribution of genes associated with virulence. We identified a total of 86 unique virulence genes, with a median of 60 genes per genome (range: 51–71) (Supplementary Data 3). A total of 45 virulence genes were present in 99–100% of the genomes. We found a small but significant difference in the number of virulence genes per genome between ST2250 and non-ST2250 genomes, with a median of 60 and 64, respectively (Fig. 2A). When comparing specific functional categories of virulence genes per genome between ST2250 and non-ST2250 genomes, we found significant differences for those genes associated with adherence, immune modulation, exoenzyme, exotoxin, and effector delivery (all with p < 0.05, Mann–Whitney test), but not for biofilm (Fig. 2B–G, Supplementary Data 4). Except for the effector delivery systems, the set of specific virulence genes did not vary tremendously between the two groups in terms of their median, with differences in dispersion or the presence of outliers leading to a statistically significant difference.
A Comparison of the total number of virulence genes per genome in ST2250 versus non-ST2250. Mean values are 63.90 vs 59.23. Comparison of the total number of virulence genes of specific class B adherence [mean values are 5.0 vs 4.5], C biofilm [mean values are 5.0 vs 5.0], D modulation [mean values are 18.78 vs 18.34], E exoenzyme [mean values are 6.78 vs 6.42], F exotoxin [mean values are 12.50 vs 12.09], and G effector delivery system [mean values are 6.0 vs 9.0]. For all panels, red and gray dots represent individual genomes of ST2250 and non-ST2250, respectively. The median is represented by the large blue dot, the box represents the interquartile range, the horizontal line in the middle of the box represents the median, and the lower and upper ends of the violin jitter plots represent the lowest data point without the outliers and the highest data point without outliers, respectively. Median was used as the central measure due to the non-normality of the data. Mann–Whitney test was used for all comparisons. Detailed information about the presence and absence of virulence genes in each genome is presented in Supplementary Data 3 and the list of virulence genes in each functional category in Supplementary Data 4.
When considering only human-derived genomes, we found similar results. For the total number of virulence genes per genome, the difference between ST2250 (n = 205) and non-ST2250 (n = 107) genomes remained highly significant (p = 1.87e − 37, Mann–Whitney test). Broken down into functional categories, we found significant differences between ST2250 and non-ST2250 in the number of genes per genome related to adherence (p = 3.39e − 07), exoenzyme (p = 5.26e − 10), exotoxin (p = 2.04e − 05), and effector delivery system (p = 3.38e − 44), but not for biofilm (p = 0.47) and modulation (p = 0.92).
Altogether, these results show that both ST2250 and non-ST2250 lineages generally have similar potential to cause disease.
AMR genes are varied, numerous and co-occurring
We identified 29 distinct genes associated with AMR, representing 12 antimicrobial classes and disinfectants (Supplementary Data 3). When comparing ST2250 and non-ST2250, we observed variation in the composition of AMR genes present in individual genomes (Fig. 3A) and in the total number of AMR genes per genome, with median values of 13 and 10 for ST2250 and non-ST2250 genomes, respectively (p = 3.29e-24, Mann–Whitney test; Fig. 3B). Across the entire dataset, there were a total of 45 distinct AMR gene profiles, with the number of genes per profile varying between 9 and 16 (Fig. 3C). Nine AMR genes were present in ≥99% of the genomes: lmrS, norA, mepAR and mgrA (multiple drugs), tet(38) (tetracycline), blaZ and mecC-type blaZ (beta-lactam), and arlRS (fluoroquinolone and disinfectant) (Fig. 3C, D). In contrast, some AMR genes were only detected in single genomes, such as fusB (fusidic acid), lnuA (lincosamide), TEM-181 (beta-lactam), and mupA (mupirocin).
A Histogram showing the number of genomes containing a specific AMR profile based on unique combinations of AMR genes shown in (C). Pink portions of a bar represent the ST2250 genomes, while the gray portion represents non-ST2250 genomes. The number above each bar shows the total number of genomes. B Comparison of the total number of AMR genes per genome in ST2250 versus non-ST2250 genomes. The median is represented by the large blue dot, the box represents the interquartile range, the horizontal line in the middle of the box represents the median, and the lower and upper ends of the violin jitter plots represent the lowest data point without the outliers and the highest data point without outliers, respectively. Median was used as the central measure due to the non-normality of the data. Mann–Whitney test was used to compare the two groups. C AMR profiles identified based on specific combination of AMR genes, shown by the connected dots. Gene names are shown on the left. D Number of genomes that carry each AMR gene listed in (C). For (C, D), colors of the dots and bars represent different antimicrobial classes. E Heatmap displaying the conditional probabilities of co-occurrence of AMR genes, i.e., gene A in relation to gene B. Gray squares represent gene pairs excluded from the analysis to avoid self-query. F Association network illustrating the directionality of co-occurrence among AMR genes based on conditional probability, relative risk, bidirectional probability score, and fold change. Nodes of the network represent AMR genes and are colored according to their respective antimicrobial classes. Edge width and color are scaled according to the magnitude of relative risk. In the co-occurrence network, arrows indicate the directionality of co-occurrence. Detailed information about the presence and absence of AMR genes in each genome is presented in Supplementary Data 3.
A total of 289 genomes carried fosB (fosfomycin resistance) (Supplementary Fig. 2) and includes genomes from ST2250 (n = 210 genomes have fosB), ST1223 (n = 44), ST1850 (n = 8), one from each of ST5964, ST5057, ST5978, and ST3262, and 23 from undesignated STs. Within the ST2250 clade in the phylogeny are five recently diverged non-ST2250 genomes that also harbor fosB. The five genomes share at least six of the seven MLST loci with ST2250, with four genomes containing novel alleles and one genome lacking one locus.
We next sought to determine if there were AMR genes that tend to co-occur more often than is expected by chance. Here, we analyzed only those AMR genes that were present in 5 – 95% of the genomes. These include aph(3’)-IIIa (aminoglycoside), PC1-type blaZ (beta-lactam), fosB (fosfomycin), dfrG (trimethoprim), mecA (beta-lactam), and tetL (tetracycline). For every pair of the five AMR genes, we calculated the conditional probability (CP) and relative risk (RR) of their co-occurrence28 (Fig. 3E and Supplementary Data 5). Results showed certain AMR gene pairs that have conditional probabilities of at least 0.75 (i.e., there is at least 75% probability of observing gene A given the presence of gene B). These include (written in the order gene A – gene B): aph(3’)-IIIa, dfrG, mecA, and tetL (gene A) that showed significant CP of occurrence in the presence of PC1-type blaZ (gene B), as well as aph(3’)-IIIa, PC1-type blaZ, mecA, and tetL (gene A) that were frequently observed in association with fosB (gene B). We also identified 10 gene pairs with RR values > 1, indicating that the presence of one gene is associated with a higher likelihood of occurrence of another28 (Fig. 3F). The pair with the highest RR is tetL and aph(3’)-IIIa. It should be noted that although high values indicate a strong relative association, they do not necessarily reflect high absolute probabilities28.
Among the AMR gene pairs analyzed, some resistance determinants stood out for both high CP and RR values, suggesting consistent co-occurrence patterns. The gene tetL showed a strong association with aph(3’)-IIIa (CP = 0.997; RR = 23.7), while dfrG was frequently observed in the presence of mecA (CP = 0.997; RR = 12.75). Other significant associations included aph (3’)-IIIa in the presence of PC1-type blaZ (CP = 0.967; RR = 1.75), fosB in the presence of aph (3’)-IIIa (CP = 0.978; RR = 1.43), tetL in the presence of fosB (CP = 0.997; RR = 1.44), dfrG in the presence of PC1-type blaZ (CP = 0.972; RR = 1.67), and mecA in the presence of PC1-type blaZ (CP = 0.942; RR = 1.69).
Altogether, the AMR results show that S. argenteus, mainly driven by ST2250, harbor a diverse array of AMR genes. Co-occurring AMR genes in different combinations indicate co-inheritance, whether vertically or horizontally, and high potential for multidrug-resistant lineages to emerge when multiple genetic determinants of AMR converge in the same genetic background.
Heterogeneous distribution of plasmid replicons and co-occurrence with AMR genes
Across the 379 genomes evaluated, we identified 43 distinct plasmid replicon types, consisting of 41 rep-type and two Col-type (Supplementary Data 6). Among the STs with more than 10 genomes, ST2793 showed the highest proportion of genomes carrying plasmids (22/23 genomes, equivalent to 95.7%), followed by ST2250 (178/213 genomes, 83.5%) (Supplementary Fig. 3A). All genomes from the five most frequent STs carried rep types. Plasmid type ColRNAI_1 was detected in an ST1850 genome (Accession no. ERR3501086), while ColpVC_1 was present in an ST2250 genome (Accession no. GCA_010570875.1_ASM1057087v1). No plasmid replicon was detected in ST5961 (0/17) genomes. Overall, these results reveal heterogeneity in plasmid element distribution among S. argenteus lineages.
Because many of the genomes in our dataset consisted of short-read sequences, we were unable to verify the gene content of plasmids. Nonetheless, it is still possible to investigate whether any of the plasmid replicon types co-occur with specific AMR genes more often than expected by chance. We identified three plasmid replicon types and six AMR genes that exhibited significant co-occurrence (Supplementary Fig. 3B and Data 7). Plasmid types rep16_1_CDS8 (pSAS) and rep5a_1_repSAP001 (pN315) both co-occur with the same AMR genes PC1-type blaZ, aph(3’)-IIIa, fosB and tet(L) (all with p-values ranging from 1.10 × 10⁻⁴⁷ to 6.56 × 10⁻²⁸, Fisher’s exact test), and all pairs were detected between 89 and 209 genomes. Among these pairwise relationships, plasmid pairings with tetL and aph(3’)-IIIa stood out due to their high odds ratio (OR) values, reflecting the high probability of their presence in genomes carrying these plasmid replicon types. We also found significant co-occurrence of AMR genes PC1-type blaZ and fosB with the three plasmid replicon types, although co-occurrence with rep20_3_rep (pTW20) were relatively fewer compared to rep16_1_CDS8 (pSAS) and rep5a_1_repSAP001 (pN315). Nonetheless, it is notable that dfrG (p = 1.18 × 10⁻¹⁰; OR = 165.2; in 21 genomes) and mecA (p = 3.17 × 10⁻⁸; OR = 109.1; in 21 genomes) showed statistically significant though less frequent co-occurrence with this replicon rep20_3_rep (pTW20), but not the other two plasmid types.
Molecular dating of ST2250
ST2250 comprised more than half of our dataset. We therefore sought to determine the evolutionary history of ST2250 using a time-calibrated phylogenetic tree. We estimated that the most recent common ancestor of ST2250 emerged in 1967 (95% highest posterior density (HPD) interval = 1950–1978) (Fig. 4A). Nearly all of ST2250 (n = 200/213 genomes or 93.90%) contain the fosB gene; in contrast, of the 166 non-ST2250 genomes, 79 carry the fosB gene (corresponding to 47.60%) which included ST1850, a small cluster of eight genomes that is the most closely related clade to ST2250 (Supplementary Fig. 2).
A Bayesian maximum clade credibility time-calibrated phylogeny of ST2250 based on non-recombining regions of the core genome alignment. The colored lines emanating from the branches of the tree represent the major geographical regions. genomes were also annotated based on geographical origin and antimicrobial classes, both represented by colored dots next to the branches. Different SCCmec types are indicated by distinct colors of colored bars. Antimicrobial resistance elements present in each genome are indicated by colored dots and their colors represent the antimicrobial classes. The divergence date (median estimate with 95% highest posterior density dates) is indicated as a horizontal blue line on the tree. For visual clarity, only the relevant divergence dates related to the clonal origin and SCCmec acquisition are shown. B Bayesian skygrowth plot showing changes in effective population size (Ne) over time. Median is represented by a solid line, and 95% confidence intervals are represented by the shaded blue area around the median. Details of the molecular dating analyses are presented in Supplementary Fig. 4, Data 10 and Data 11.
Three SCCmec types were detected in ST2250 genomes. SCCmec type IVc was identified in 59 genomes that form a single cluster within the phylogeny and are mainly found in Europe. We estimated the origin of this cluster to be 1992 (HPD = 1984–1999). Nearly all members of this cluster (n = 55/59 genomes) also harbor the AMR gene dfrG. SCCmec type IVa was detected in three genomes, with its origins estimated in 1998 (HPD = 1991–2006) and 2011 (HPD = 2009–2012). SCCmec type Ia was identified in two genomes, with an estimated origin of 2017 (HPD = 2012–2017). These events indicate multiple independent acquisitions of distinct SCCmec elements within ST2250. Another phylogenetic cluster within ST2250 (n = 112 genomes) that is mainly found in Asia emerged in 1989 (HPD = 1979–1995). This Asian cluster is characterized by the high prevalence of aph(3’)-IIIa (n = 92 genomes, equivalent to 82.14%) and tetL (n = 84 genomes; 75%), but only two genomes carry SCCmec (type IVa). Both Asian and European sublineages of ST2250 exhibited multiple instances of intercontinental dissemination based on the presence of some genomes from one continent intermingled within the phylogenetic cluster of genomes from the other continent.
Demographic reconstruction revealed a rapid increase in the effective population size of ST2250 in the past 56 years, peaking in 2012 then rapidly declining afterwards (Fig. 4B). The molecular dating results show that the evolutionary history of ST2250 is punctuated by sublineages that have diversified and independently acquired various combinations of AMR genes; however, there is some indication that the ST2250 population is experiencing a decline in recent years.
Discussion
The growing number of reports of human diseases attributed to S. argenteus worldwide, as well as its presence in non-clinical environments, call for a deeper understanding of this emerging pathogen. The recent identification of this species in foodborne outbreaks further highlights this concern and underscores the importance of understanding its biology as an opportunistic pathogen3,29,30. Our study contributes to addressing this gap in knowledge by elucidating the population genomic features of S. argenteus, especially with respect to AMR dissemination. Overall, our results show that the AMR pool of S. argenteus is driven in part by the global clonal expansion of ST2250 in the last five decades and its subsequent diversification into sublineages with distinct AMR gene content.
The demographic pattern of S. argenteus up until 2012 (shown in Fig. 4B) mirrors that of antimicrobial resistant S. aureus31, and we can only speculate as to the underlying causes of this surge in population growth in S. argenteus. The expansion of ST2250 has certainly contributed to shaping the global population dynamics of S. argenteus. In S. aureus, demographical characteristics point to a series of epidemic waves of antimicrobial resistant strains instigated by one or a few successful clones that have caused a large proportion of infections in hospitals and communities worldwide31. At least four waves of resistance are known and are attributed to a penicillin-resistant clone in the 1940s and four distinct MRSA clones (MRSA I, II, III, IV) between the 1960s and 2000s31. These resistant clones appeared immediately after antimicrobial agents were widely introduced in clinics, thereby creating a strong selective pressure on nosocomial strains31. Moreover, the dominance of ST2250 and its rapid expansion in recent evolutionary time may partly explain the low values of genome fluidity and Heaps’ γ coefficient, indicating limited gene content variation.
The high prevalence of fosB in ST2250 genomes (93.90%) compared to non-ST2250 genomes (47.60%) may suggest that the gene’s persistence in this lineage played an important role in the evolutionary expansion of ST2250. Discovered in 1969, fosfomycin inhibits bacterial cell wall synthesis by impeding the formation of the peptidoglycan precursor32. It is used primarily as an oral treatment for urinary tract infections33. While considered an “old” antibiotic, fosfomycin has seen a resurgence in interest and use due to the rise of multidrug-resistant bacteria, including MRSA strains34, which are known to also exhibit multidrug resistance35. The origin of fosB in ST2250 is likely linked to plasmid acquisition, since this gene has been reported widely in S. aureus36 as well as in other Staphylococcus species37. The contribution of mobile genetic elements in ST2250 evolution has previously been highlighted; for example, in Thailand, AMR and exotoxin genes linked to livestock-associated S. aureus were mobilized via plasmids and phages to ST225021. In-depth investigations of mobile genetic elements and their gene cargo using long-read sequencing of S. argenteus strains will be paramount to understanding how ST2250 has acquired its AMR genes. The closest relative of ST2250 is ST1850, and while it is not frequently detected in our dataset (eight genomes), it also harbors fosB, which may suggest the presence of fosB in the common ancestor of ST2250 genomes.
Two sublineages of ST2250 emerged independently at nearly the same time (Asian sublineage in 1989 and European sublineage in 1992), and their distinct AMR profiles likely explain the co-occurrence patterns of AMR genes shown in Fig. 3E, F. The tight co-occurrence of aph(3’)-IIIa, blaZ, and tetL in the Asian sublineage, as well as that of mecA, blaZ, and dfrG in the European sublineage, may be indicative of an initial horizontal co-acquisition of these genes in the common ancestor of each sublineage, followed by vertical inheritance in descendants as each sublineage diversified. The date of origin of the Asian lineage that we report here is consistent with that reported by Moradigaravand et al.21. Their study on 68 Thai S. argenteus genomes sampled between 2006 and 2013 inferred an introduction of ST2250 from Malaysia to Thailand 20–30 years ago (therefore 1983–1993)21. Previous studies suggest that this divergence may have been accompanied by ecological adaptation events, facilitated by plasmid-mediated acquisition of AMR genes18,21. A similar scenario has occurred in S. aureus ST59, which experienced concurrent but independent evolution of distinct sublineages in North America and East Asia, driven by mobile genetic elements38.
The population decline of ST2250 after 2012 is certainly intriguing. However, it is important to note that this observed decline is in part influenced by uneven temporal sampling, particularly from 2021 to 2023, and should be interpreted with caution. Skyline-based demographic reconstructions are known to be sensitive to temporal sampling density39. Nonetheless, such a decline is not inconceivable. In S. aureus, fluctuations in the effective population size of antimicrobial-resistant lineages have been documented and reflect a scenario of lineage replacement. In S. aureus, a gradual replacement of ST5 by ST8 in Colombia40, ST764 in Japan41, and ST72 in South Korea42 has been reported. Another example is methicillin-resistant ST239 being superseded by ST5943. ST5 and ST8 from bloodstream infections in the United States have alternatively fluctuated in their effective population sizes44. In S. argenteus, the second most common ST is ST1223, with some genomes carrying any of the four SCCmec types. All ST1223 genomes analyzed here carry the fosB gene as well as differential distribution of aph(3’)-IIIa, PC1_Type blaZ, and mecA. On the other hand, nearly all ST2793 genomes have SCCmec type IVc. Both STs have been detected in multiple years. If lineage replacement is indeed occurring, these two STs are possible contenders to replace ST2250. It is certainly interesting that all six major STs exhibit similar topological structure in the phylogeny – each one is found on a very long branch and from which very short branches emanate, indicating a rapid diversification in more recent time (Fig. 1A). Thus, continuous monitoring of these less common STs is needed to identify those with surging population sizes. Importantly, such replacement does not necessarily involve only lineages of the same species and may also reflect the emergence of other staphylococcal lineages or species occupying overlapping ecological niches. While the precise range of ecological niches inhabited by S. argenteus remains obscure, co-occurrence of different Staphylococcus species in the same niches have been previously reported45,46,47.
However, our dataset does not include a sufficient number of non-ST2250 genomes to carry out a time-calibrated phylogenetic analysis and elucidate the demography of ST1223, ST2793, and other less common STs. This gap in knowledge should therefore be an impetus to conduct a broad, long-term sampling scheme and robust genomic surveillance of S. argenteus to discover sources and genetic diversity of non-ST2250 lineages, as well as determine whether the decline of ST2250 will continue. Moreover, fitness and competition experiments of different S. argenteus STs will be particularly informative in understanding the genetic factors determining the success or failure of individual clones.
We acknowledge the limitations of our study. First, the reliance on publicly available genomes led to a disproportionate representation of genomes from a few geographical and ecological sources. We recognize that our findings reflect more of what exists in clinical human sources, which comprise the majority of our dataset. It is also highly likely that numerous S. argenteus strains remain unidentified in S. aureus genomic surveillance studies. We also do not have phenotypic data on AMR and virulence; hence, we cannot ascertain the antimicrobial susceptibility features of individual strains. Plasmid analysis is also limited because many of the genomes in our dataset were short-read sequenced. Second, existing databases for ST identification and for querying AMR genes, virulence-associated genes, and SCCmec types are limited to those built for the more thoroughly studied S. aureus. Hence, genetic variants specific to S. argenteus are likely to exist but remain obscured from current in silico detection methods and databases. Third, the co-occurrence analysis of AMR genes did not treat genomes as independent observations nor adjust for population structure or phylogenetic non-independence. Consequently, particularly given the overrepresentation of ST2250 in the dataset, the observed association patterns may in part due to clonal inheritance rather than independent horizontal transfer events. Regardless, it does not minimize our results showing the importance of ST2250 in the global population. Lastly, any investigation of ancestral lineages is limited by the diversity of contemporary lineages that are being analyzed. This means that missing lineages are likely to influence our inference of the time to the most recent common ancestor and dates of clonal origins. Notwithstanding these limitations, our study should be considered as a baseline census of the standing genomic and lineage diversity of S. argenteus worldwide. We anticipate that the results presented here will form the basis for outstanding questions and open multiple avenues for future research and surveillance efforts. For example, the range of disease presentations and the mechanisms of disease transmission remain to be elucidated in S. argenteus, as well as the prevalence of S. argenteus in asymptomatic carriage in the human population.
In summary, our results reveal that the global spread of resistance genes related to various antimicrobial classes is driven in part by the rapid lineage expansion and AMR acquisition of ST2250 in the past five decades. Our study will help inform public health efforts to developing effective strategies to reduce the risk for further dissemination of S. argenteus lineages with multidrug-resistant and virulent characteristics.
Methods
Genome collection and annotation
A total of 437 genome sequences were retrieved from the National Center for Biotechnology Information (NCBI) Sequence Read Archive in September 2023. Paired-end reads were assembled de novo using the Shovill v.1.1.0 pipeline (https://github.com/tseemann/shovill), implementing the k-mer-based assembly algorithm SKESA v.2.4.048, and followed by pre-and post-processing methods in Shovill. We employed the—trim flag for trimming of adapter sequences. To assess genome quality, we used QUAST v.5.0.249 and CheckM v.1.1.350. Genomes with <90% completeness and >5% contamination were excluded. We also excluded assemblies with >200 contigs and N50 < 40,000 bp. In total, our dataset included 379 genomes, which were used in all subsequent analyses. The number of contigs ranged from 1 to 128 (median = 18), and N50 values ranged from 47,737 to 2,793,527 bp (median = 447,539 bp) (Supplementary Data 1). We obtained genome completeness (mean = 99.73%, range: 96.6 – 99.81%) and contamination (mean = 0.12%, range: 0.08–2.33%) (Supplementary Data 1). All 379 genomes were within the quality thresholds recommended by CheckM. To confirm that the genomes belonged to the same species, we calculated the average nucleotide identity (ANI) for all possible genome pairs using FastANI v.1.3251 (Supplementary Data 8). We applied the recommended threshold of ≥95% ANI to define species boundaries. Additionally, the 379 genomes were compared to the reference genome S. argenteus GCA_000236925.1 (ASM23692v1) to verify species identity. The genomes were annotated using Prokka v.1.14.652.
Pan-genome analysis and phylogenetic tree reconstruction
We used Panaroo v.1.3.453 to characterize the collective set of genes present in all genomes in our dataset, i.e., pan-genome54. Panaroo was run in strict mode with the—remove-invalid-genes option to ignore annotations that do not conform to the expected gene structure (i.e., with a premature stop codon or genes of an invalid length). We defined core genes as those present in ≥99% of the genomes (i.e., at least 375 genomes), whereas the accessory genes were genes present in <99% of the dataset (i.e., less than 375 genomes) (Supplementary Data 2). To estimate gene content variability across the population, genome fluidity was calculated using a custom R script based on the method described by Kislyuk et al.23, where the number of unique genes between each pair of genomes is divided by the total number of genes present in either genome, and the mean of all pairwise comparisons is reported as the fluidity value. To complement the evaluation of gene content variability across genomes, pan-genome openness was also estimated using a custom R script based on Heaps’ law as proposed by Tettelin et al.24. Pan-genome size (P) was modeled as a function of the number of genomes included (N) using the power-law model P = k·N^γ. The γ parameter was derived from the mean rarefaction curve (with 100 permutations) and used to classify the pan-genome as open or closed according to the original Tettelin criterion (0 < γ < 1 indicates an open pan-genome, γ ≤ 0 or γ ≥ 1 indicates a closed pan-genome). This analysis was performed both for the complete genome dataset (n = 379) and separately for ST2250 (n = 213 genomes) to evaluate potential differences in gene expansion dynamics at the population and lineage levels.
Nucleotide sequences were aligned using MAFFT v.7.47155. Sequence alignments of 2180 core genes were concatenated, and SNPs were extracted from the core gene alignment using SNP-sites v.2.5.156. A total of 59,124 SNPs were identified. The aligned core SNPs were used to construct a maximum likelihood phylogenetic tree with IQ-TREE v.2.1.4-Beta57, using the general time-reversible (GTR) model of nucleotide substitution58 with empirical base frequencies, ascertainment bias correction, and Gamma-distributed rate variation with three categories (SYM + ASC + R3), as selected by ModelFinder59. Node support was assessed with 1000 bootstrap replicates. Phylogenetic trees were visualized and annotated using the Interactive Tree of Life (iTOL)60.
In silico identification of STs, AMR genes, virulence genes, and SCCmec
The ST was determined using mlst v.2.19.0 (https://github.com/tseemann/mlst), which extracts seven housekeeping genes and compares allelic profiles against previously characterized STs in the PubMLST database61. Because an MLST scheme specific for S. argenteus has not been developed yet, we used the MLST scheme for S. aureus62, as has been done by previous studies of S. argenteus19,20 (Supplementary Data 1). STs were determined based on sequence variation in seven single-copy housekeeping genes (arc, aroE, glpF, gmk, pta, tpi, yqiL). We used ABRicate v.1.0.1 (https://github.com/tseemann/abricate) to identify the presence of acquired AMR and virulence genes (Supplementary Data 3). We used the minimum thresholds of > 80% for sequence coverage and >80% sequence identity for comparing the query sequence with the curated Comprehensive Antimicrobial Resistance Database (CARD)63 and the Virulence Factor Database (VFDB)64. We screened for the presence and types of SCCmec using the stand-alone tool staphopia-sccmec v.165.
We employed ReGAIN v.1.0.528 to investigate probabilistic patterns of co-occurrence among AMR genes. To minimize noise and exclude genes that are either rare or nearly ubiquitous, we focused on resistance genes present in 5–95% of the genomes for the construction of the Bayesian network. For each pair of AMR genes, we computed different metrics to estimate co-occurrence: conditional probability (CP), relative risk (RR), bidirectional probability score (BDPS), and fold change28 (Supplementary Data 5). CP refers to the probability of observing gene A given the presence of gene B, expressed on a scale of 0–128. RR is defined as how much more likely it is to observe gene A in the presence of gene B, compared to its absence, and is calculated as the ratio between the CP of occurrence of A given B and the CP of occurrence of A in the absence of B28. BDPS refers to bidirectional strength of co-occurrence and is calculated as the ratio of CP of gene A given gene B to the CP of gene B given gene A28. Fold change refers to the ratio of the RR in both directions. The analysis was carried out with the bnS option in module 2 (optimized for small and moderately sized datasets), 500 bootstrap replicates, and 100 resamples, as recommended by the developers. A significance threshold of 0.5 was applied to exclude gene pairs with weak statistical support. To build the network, we selected gene A - gene B relationships that were statistically validated through CP, RR, BDPS, and fold change (Supplementary Data 5).
Plasmid replicon detection and co-occurrence analysis with AMR genes
We used ABRicate (version 1.0.1) with the PlasmidFinder database66 to detect and classify plasmid replicon types. Minimum thresholds of 80% sequence identity and 80% coverage were applied for plasmid sequence identification (Supplementary Data 6). We selected only those AMR genes and plasmid replicons that were present in 5–95% of the genomes to reduce noise due to both ubiquitously occurring elements and those that are extremely rare elements. This selection allowed us to increase the robustness of our results. Hence, our analysis only included PC1_Type blaZ, aph(3’)-IIIa, fosB, tet(L), dfrG and mecA and the plasmid replicon types rep16_1_CDS8(pSAS), rep5a_1_repSAP001(pN315) and rep20_3_rep(pTW20). Only significantly supported pairwise associations between plasmid replicon and AMR genes were included in the co-occurrence network (Fig. 3F). The network was visualized in RStudio 2024.09.167.
Estimating the date of clonal origin and effective population size
In creating the time-calibrated phylogeny of ST2250, we needed to use a reference genome of the same ST. We opted to use a genome from our dataset based on simple distance estimation using Mash v.2.368. Using the tidyverse v.2.0.0 package69 in RStudio67, we identified the genome with the lowest average distance to all others. The selected reference genome was SAMEA3449138 that was sampled in 2015 (Supplementary Data 9). SNPs in the core genome alignment were identified using Snippy v.4.6.0 (https://github.com/tseemann/snippy) and mapped to the reference genome using the snippy-multi script. Multiple sequence alignments were then screened for evidence of recombination using Gubbins v.3.2.170, generating recombination-masked full-genome length and SNP-only alignments. Maximum likelihood phylogenies were inferred from the SNP-only alignments using IQ-TREE v.2.1.457, applying the Transversion model with empirical base frequencies and invariable sites (TVM + F + I) substitution model selected according to the Bayesian Information Criterion71. For evolutionary rate estimation using BactDating, we used the recombination-masked full-genome alignment, which retains invariant sites and avoids ascertainment bias associated with SNP-only alignments. Using the recombination-free phylogeny generated by Gubbins for ST2250, we used TempEst v.1.5.372 to assess the temporal signal and identify genomes with potential issues, such as aberrant sequence or mutation rates, mislabeled dates, or unique evolutionary events that intensify genetic divergence. Based on this analysis, one outlying genome was identified and excluded from further analyses (Supplementary Fig. 4A and Data 1,0).
We used BactDating v.1.1.2 to carry out a root-to-tip linear regression analysis of ST225073. We calculated the coefficient of determination (R2) to assess the significance of the temporal signal based on random permutations of sampling dates (Supplementary Fig. 4B). We then used BactDating to estimate the dates of the most recent common ancestor73. We carried out 107 iterations, removed the first half as burn-in, and subsequently sampled every 100 iterations. To test the robustness of the molecular clock signal, we performed a random permutation of the sample collection dates, maintaining the original interval from 2008 to 2023 but breaking the temporal association between genome and collection date (Supplementary Data 10). The dates were shuffled using the dplyr v.1.1.474 and lubridate v.1.9.475 packages in R v.4.4.376. The convergence and mixing of the Markov Chain Monte Carlo (MCMC) chains were evaluated with the coda package v.0.19-4.177 (Supplementary Data 10, Fig. 4C, D). Subsequently, BactDating’s model compare function was used to calculate the Deviance Information Criterion, a metric for comparing the fit and complexity of Bayesian models78 and results confirmed that our selected model was better than the model with the shuffled dates (Supplementary Data 10). To guarantee the quality of the samples of the inferred parameters (coalescent time unit α, substitution rate μ, per-branch substitution rate σ) in BactDating, we calculated the effective sample size, ensuring that the values exceeded the recommended limit of 200 (Supplementary Data 9). We also performed an additional BactDating analysis using the same sampling dates and settings as the original run. We compared this second run with the initial analysis to check that the Gelman-Rubin multivariate statistic was lower than the recommended threshold of 1.179 (Supplementary Data 11). This statistic is used to monitor the convergence of the simulations by evaluating the variances within and between the MCMC chains80. We used the same approach to carry out a time-calibrated phylogeny for ST1223; however, even after 108 iterations, the data failed to converge.
To determine changes in the effective population size of ST2250 over time, we ran the R package Skygrowth v. 0.3.181 on the recombination-free phylogeny of ST2250. Skygrowth employs a Bayesian Gibbs-within-Metropolis MCMC and fast maximum a posteriori algorithm to estimate the effective population size through time and growth rates of the effective population size. We ran Skygrowth with the default mode of 100,000 iterations and a smoothing parameter of 10 exponential for the prior.
Statistical tests
All statistical analyses were carried out using the ggstatsplot v.0.13.082 package in R v.4.4.376. We checked for normality using the Shapiro.test() function implemented in R. Because the data were not normally distributed, Mann–Whitney test was used to compare the number of virulence and AMR genes per genome between ST2250 and non-ST2250 (Figs. 2 and 3B). Results were considered significant when p < 0.05. Co-occurrence was assessed using Fisher’s exact test, calculating for each gene–plasmid pair the p-value (with p < 0.005 considered statistically significant), the odds ratio (interpreted as a positive association when >1), and n_overlap values corresponding to the number of genomes carrying both elements (Supplementary Data 11). Only associations with statistical support were included in the co-occurrence networks (Supplementary Fig. 3B).
Data availability
The dataset supporting the conclusions of this article is included within the article and its supplementary files. Genome sequence data used in this study are publicly available in the NCBI Sequence Read Archive, and BioSample accession numbers for each genome are listed in Supplementary Data 1.
References
Tong, S. Y. C. et al. Virulence of endemic nonpigmented northern Australian Staphylococcus aureus clone (clonal complex 75, S. argenteus) is not augmented by staphyloxanthin. J. Infect. Dis. 208, 520–527 (2013).
Tong, S. Y. C. et al. Novel staphylococcal species that form part of a Staphylococcus aureus-related complex: the non-pigmented Staphylococcus argenteus sp. nov. and the non-human primate-associated Staphylococcus schweitzeri sp. nov. Int J. Syst. Evol. Microbiol. 65, 15–22 (2015).
Wakabayashi, Y. et al. Staphylococcal food poisoning caused by Staphylococcus argenteus harboring staphylococcal enterotoxin genes. Int J. Food Microbiol. 265, 23–29 (2018).
Monecke, S. et al. Characterisation of PVL-Positive Staphylococcus argenteus from the United Arab Emirates. Antibiotics 13, 401 (2024).
Supriadi, I. R. et al. Identification and characterization of Staphylococcus argenteus from Indonesia. Int. J. Med. Microbiol. 316, 151629 (2024).
Chen, S.-Y. et al. High mortality impact of Staphylococcus argenteus on patients with community-onset staphylococcal bacteraemia. Int J. Antimicrob. Agents 52, 747–753 (2018).
Lee, M. et al. Staphylococcus argenteus bacteremia in the Republic of Korea. Microbiol. Spectr. 12, e0279823 (2024).
Rigaill, J. et al. Community-acquired Staphylococcus argenteus sequence type 2250 bone and joint infection, France, 2017. Emerg. Infect. Dis. 24, 1958–1961 (2018).
Ohnishi, T. et al. Purulent lymphadenitis caused by Staphylococcus argenteus, representing the first Japanese case of Staphylococcus argenteus (multilocus sequence type 2250) infection in a 12-year-old boy. J. Infect. Chemother. 24, 925–927 (2018).
Rong, D. et al. Prevalence and characterization of Staphylococcus aureus and Staphylococcus argenteus isolated from rice and flour products in Guangdong, China. Int. J. Food Microbiol. 406, 110348 (2023).
Indrawattana, N. et al. Staphylococcus argenteus from rabbits in Thailand. Microbiologyopen 8, e00665 (2019).
Meijer, E. F. J. et al. Canine Staphylococcus argenteus: case report from The Netherlands. Pathogens 11, 153 (2022).
Hao, S. et al. Genomic profiling of evolving daptomycin resistance in a patient with recurrent Staphylococcus argenteus sepsis. Antimicrob. Agents Chemother. 64, e00961-20 (2020).
Shittu, A. O. et al. First report of a methicillin-resistant, high-level mupirocin-resistant Staphylococcus argenteus. Front. Cell Infect. Microbiol. 12, 860163 (2022).
Witteveen, S. et al. Genetic characteristics of methicillin-resistant Staphylococcus argenteus isolates collected in the Dutch National MRSA Surveillance from 2008 to 2021. Microbiol. Spectr. 10, e0103522 (2022).
Giske, C. G. et al. Transmission events and antimicrobial susceptibilities of methicillin-resistant Staphylococcus argenteus in Stockholm. Clin. Microbiol. Infect. 25, 1289.e5–1289.e8 (2019).
Jauneikaite, E. et al. Staphylococcus argenteus transmission among healthy Royal Marines: a molecular epidemiology case-study. J. Infect. 83, 550–553 (2021).
Wu, S. et al. Evolutionary divergence of the novel Staphylococcal species Staphylococcus argenteus. Front. Microbiol. 12, 769642 (2021).
Eshaghi, A. et al. Phenotypic and genomic profiling of Staphylococcus argenteus in Canada and the United States and recommendations for clinical result reporting. J. Clin. Microbiol. 59, e02470-20 (2021).
Goswami, C., Fox, S., Holden, M., Leanord, A. & Evans, T. J. Genomic analysis of global Staphylococcus argenteus strains reveals distinct lineages with differing virulence and antibiotic resistance gene content. Front. Microbiol. 12, 795173 (2021).
Moradigaravand, D. et al. Evolution of the Staphylococcus argenteus ST2250 clone in northeastern Thailand is linked with the acquisition of livestock-associated Staphylococcal genes. mBio 8, e00802–17 (2017).
Yan, A., Kus, J. V. & Sant, N. The Staphylococcus aureus complex: implications for the clinical microbiology laboratory. J. Clin. Microbiol. 63, e0127624 (2025).
Kislyuk, A. O., Haegeman, B., Bergman, N. H. & Weitz, J. S. Genomic fluidity: an integrative view of gene diversity within microbial populations. BMC Genom. 12, 32 (2011).
Tettelin, H., Riley, D., Cattuto, C. & Medini, D. Comparative genomics: the bacterial pan-genome. Curr. Opin. Microbiol. 11, 472–477 (2008).
Liu, J. et al. Staphylococcal chromosomal cassettes mec (SCCmec): a mobile genetic element in methicillin-resistant Staphylococcus aureus. Micro. Pathog. 101, 56–67 (2016).
International Working Group on the Classification of Staphylococcal Cassette Chromosome Elements (IWG-SCC) Classification of staphylococcal cassette chromosome mec (SCCmec): guidelines for reporting novel SCCmec elements. Antimicrob. Agents Chemother. 53, 4961–4967 (2009).
Uehara, Y. Current status of Staphylococcal Cassette Chromosome mec (SCCmec). Antibiotics 11, 86 (2022).
Horvath, E. R. B., Stein, M. G., Mulvey, M. A., Hernandez, E. J. & Winter, J. M. Resistance Gene Association and Inference Network (ReGAIN): a bioinformatics pipeline for assessing probabilistic co-occurrence between resistance genes in bacterial pathogens. Preprint at https://doi.org/10.1101/2024.02.26.582197 (2024).
Suzuki, Y. et al. Food poisoning outbreak in Tokyo, Japan caused by Staphylococcus argenteus. Int. J. Food Microbiol. 262, 31–37 (2017).
Cavaiuolo, M. et al. First report of enterotoxigenic Staphylococcus argenteus as a foodborne pathogen. Int. J. Food Microbiol. 394, 110182 (2023).
Chambers, H. F. & Deleo, F. R. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol. 7, 629–641 (2009).
Hendlin, D. et al. Phosphonomycin, a new antibiotic produced by strains of Streptomyces. Science 166, 122–123 (1969).
Matthews, P. C. et al. Oral fosfomycin for treatment of urinary tract infection: a retrospective cohort study. BMC Infect. Dis. 16, 556 (2016).
Omori, K. et al. Fosfomycin as salvage therapy for persistent methicillin-resistant Staphylococcus aureus bacteremia: a case series and review of the literature. J. Infect. Chemother. 30, 352–356 (2024).
Turner, N. A. et al. Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat. Rev. Microbiol. 17, 203–218 (2019).
Monte, D. F. M. & de Oliveira, C. J. B. Global trends in the increasing prevalence of the fosfomycin resistance gene in Staphylococcus aureus. Lancet Microbe 5, e104 (2024).
Etienne, J., Gerbaud, G., Courvalin, P. & Fleurette, J. Plasmid-mediated resistance to fosfomycin in Staphylococcus epidermidis. FEMS Microbiol. Lett. 52, 133–137 (1989).
McClure, J.-A. et al. Staphylococcus aureus ST59: concurrent but separate evolution of North American and East Asian lineages. Front. Microbiol. 12, 631845 (2021).
Hall, M. D., Woolhouse, M. E. J. & Rambaut, A. The effects of sampling strategy on the quality of reconstruction of viral population dynamics using Bayesian skyline family coalescent methods: a simulation study. Virus Evol. 2, vew003 (2016).
Arias, C. A. et al. A prospective cohort multicenter study of molecular epidemiology and phylogenomics of Staphylococcus aureus bacteremia in nine Latin American countries. Antimicrob. Agents Chemother. 61, e00816-17 (2017).
Hisatsune, J. et al. Staphylococcus aureus ST764-SCCmecII high-risk clone in bloodstream infections revealed through national genomic surveillance integrating clinical data. Nat. Commun. 16, 2698 (2025).
Choi, S. H. et al. A longitudinal study of adult patients with Staphylococcus aureus bacteremia over 11 years in Korea. J. Korean Med. Sci. 36, e104 (2021).
Chen, H. et al. Drivers of methicillin-resistant Staphylococcus aureus (MRSA) lineage replacement in China. Genome Med. 13, 171 (2021).
Souza, S. S. R. et al. Demographic fluctuations in bloodstream Staphylococcus aureus lineages configure the mobile gene pool and antimicrobial resistance. NPJ Antimicrob. Resist. 2, 14 (2024).
Méric, G. et al. Ecological overlap and horizontal gene transfer in Staphylococcus aureus and Staphylococcus epidermidis. Genome Biol. Evol. 7, 1313–1328 (2015).
Kastman, E. K. et al. Biotic interactions shape the ecological distributions of Staphylococcus species. mBio 7, e01157-16 (2016).
Ikhimiukor, O. O. et al. Leaky barriers to gene sharing between locally co-existing coagulase-negative Staphylococcus species. Commun. Biol. 6, 482 (2023).
Souvorov, A., Agarwala, R. & Lipman, D. J. SKESA: strategic k-mer extension for scrupulous assemblies. Genome Biol. 19, 153 (2018).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Medini, D., Donati, C., Tettelin, H., Masignani, V. & Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet Dev. 15, 589–594 (2005).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Page, A. J. et al. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Micro. Genom. 2, e000056 (2016).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Tavaré, S. Some probabilistic and statistical problems in the analysis of DNA sequences. Lect. Math. Life Sci. 17, 57–86 (1986).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 3, 124 (2018).
Enright, M. C., Day, N. P., Davies, C. E., Peacock, S. J. & Spratt, B. G. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 38, 1008–1015 (2000).
Alcock, B. P. et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the comprehensive antibiotic resistance database. Nucleic Acids Res. 51, D690–D699 (2023).
Liu, B., Zheng, D., Zhou, S., Chen, L. & Yang, J. VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res. 50, D912–D917 (2022).
Petit, R. A. & Read, T. D. Staphylococcus aureus viewed from the perspective of 40,000+ genomes. PeerJ 6, e5261 (2018).
Carattoli, A. et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
RStudio Team. RStudio: Integrated Development for R. http://www.rstudio.com/ (2020).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Schwarz, G. Estimating the dimension of a model. Ann. Stat. 6, 461–464 (1978).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Didelot, X., Croucher, N. J., Bentley, S. D., Harris, S. R. & Wilson, D. J. Bayesian inference of ancestral dates on bacterial phylogenetic trees. Nucleic Acids Res. 46, e134 (2018).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: a grammar of data manipulation. R package version 1.1.4 https://dplyr.tidyverse.org (2025).
Grolemund, G. & Wickham, H. Dates and times made easy with lubridate. J. Stat. Softw. 40, 1–25 (2011).
R Core Team. R: A language and environment for statistical computing https://www.R-project.org/ (2025).
Plummer, M., Best, N., Cowles, K. & Vines, K. CODA: convergence diagnosis and output analysis for MCMC. R. N. 6, 7–11 (2006).
Spiegelhalter, D. J., Best, N. G., Carlin, B. P. & Van Der Linde, A. Bayesian measures of model complexity and fit. J. R. Stat. Soc. Ser. B Stat. Methodol. 64, 583–639 (2002).
Gelman, A. & Rubin, D. B. Inference from iterative simulation using multiple sequences. Stat. Sci. 7, 457–472 (1992).
Brooks, S. P. & Gelman, A. General methods for monitoring convergence of iterative simulations. J. Comput. Graph. Stat. 7, 434–455 (1998).
Volz, E. M. & Didelot, X. Modeling the growth and decline of pathogen effective population size provides insight into epidemic dynamics and drivers of antimicrobial resistance. Syst. Biol. 67, 719–728 (2018).
Patil, I. Visualizations with statistical details: the ‘ggstatsplot’ approach. J. Open Source Softw. 6, 3167 (2021).
Acknowledgements
The authors thank the SUNY University at Albany Information Technology Services where all bioinformatics analyses were carried out. The study was supported by the National Institutes of Health Award no. R35GM142924 to C.P.A. The funders had no role in study design, data collection and analysis, decision to publish, and preparation of the manuscript and the findings do not necessarily reflect views and policies of the authors’ institutions and funders.
Author information
Authors and Affiliations
Contributions
C.P.A. designed and guided the work. L.R.M.C., M.M., and A.B.G.B. carried out all bioinformatics analyses. L.R.M.C. and M.M. wrote the initial manuscript. L.R.M.C. and C.P.A. wrote the final manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Costa, L.R.M., Marmion, M., Buiatte, A.B.G. et al. Five-decade global expansion of Staphylococcus argenteus ST2250 shapes antimicrobial resistance dynamics. npj Antimicrob Resist 4, 14 (2026). https://doi.org/10.1038/s44259-026-00188-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-026-00188-6