Abstract
Understanding how microorganisms adapt to novel physical and chemical environments requires integrating evolutionary, regulatory, and phenotypic perspectives. Here, we examined Streptococcus mutans populations previously evolved for 100 days under simulated microgravity (sMG) or combined microgravity and silver nitrate (sMGAg), generating new transcriptomic and phenotypic datasets and integrating them with prior whole-genome sequencing. These environments model key pressures encountered in enclosed spaceflight habitats, including altered fluid shear, oxidative challenges, and exposure to disinfectants. Populations maintained under normal gravity (NG) largely preserved ancestral metabolic and redox characteristics. In contrast, sMG populations exhibited divergent physiological and transcriptional outcomes that were not predictable from genomic variants alone, including multiple ROS response patterns, broad reductions in carbohydrate metabolism, and consistent retention of trehalose utilization. Populations evolved under sMGAg showed more convergent patterns, characterized by broad activation of oxidoreductase and metal-handling pathways, elevated basal ROS relative to the ancestral strain with reduced inducibility, and a consistent gain in nitrate-reduction capability. These outcomes reflect condition-associated physiological states resolved only through combined genomic, transcriptomic, and phenotype-level data, as no single data type was sufficient to capture the full structure of adaptive responses. Together, these findings illustrate how distinct physical and chemical stress regimes reshape the landscape of accessible evolutionary responses, with microgravity alone permitting a wider range of adaptive trajectories and microgravity combined with silver favoring more uniform physiological states. More broadly, this work demonstrates that integrated multi-level datasets are essential for accurately characterizing adaptive outcomes in extreme or non-terrestrial environments.
Similar content being viewed by others
Introduction
Life on Earth has evolved within a narrow band of biophysical constraints, including gravity, oxygen-rich atmospheres, and terrestrial chemistry1,2. When organisms encounter environments that violate these conditions, such as in outer space, it is essential to understand how they adapt. Space imposes a suite of selective pressures on terrestrial microorganisms, including microgravity, elevated radiation, limited nutrients, and altered atmospheric composition3. These conditions can dramatically affect microbial physiology, triggering changes in stress responses, virulence, and gene regulation4,5. Due to logistical constraints of spaceflight, analog environments provide valuable platforms to explore microbial responses under space-like conditions6,7,8,9.
The human oral microbiome is a complex and dynamic ecosystem that plays a critical role in maintaining systemic health10. Among its members, Streptococcus mutans is of particular importance due to its central role in the development of dental caries. This is primarily due to its acidogenic metabolism, its ability to form robust biofilms, and its capacity to tolerate acidic environments10,11,12,13. While S. mutans has been well studied in terrestrial contexts such as pH fluctuation, dietary sugar exposure, and microbial competition10,13, far less is known about how it adapts to environmental extremes that differ from its ancestral environment. Studies using S. mutans in simulated microgravity have shown links to increased acid tolerance, modified gene expression, and alterations in the cell wall14,15,16,17. These adaptations may have clinical relevance, as astronauts have reported increased susceptibility to dental caries during spaceflight, likely due to altered salivary flow, biofilm dynamics, and reduced oral hygiene18,19,20.
A wide range of molecular pathways have been implicated in microbial survival under spaceflight or simulated microgravity, revealing diverse transcriptional strategies that support physiological resilience. Short-term studies show that space-relevant environments rapidly induce oxidative stress responses, including the upregulation of peroxidases and redox-maintaining enzymes such as katA, sodA, and thioredoxins, which help mitigate intracellular damage from reactive oxygen species8,21. Heat shock proteins and general stress regulators including dnaK, hrcA, htrA, ciaR/H, and rpoE are also commonly elevated, supporting protein folding, surface adhesion, and tolerance to thermal and oxidative challenges8,22. In parallel, space-adapted bacteria often downregulate glycolytic and phosphotransferase systems, shifting toward alternative carbohydrate metabolism involving trehalose, galactose, and glycogen utilization21,23,24. Collectively, these transcriptional responses reflect broad physiological plasticity that facilitates short-term acclimation to environmental extremes. Yet while these regulatory changes can be induced quickly, they are often reversible and non-heritable. In contrast, long-term exposure to spaceflight or simulated microgravity conditions can result in stable genetic mutations that permanently fix adaptive traits.
Longitudinal whole-genome sequencing across multiple evolved lineages has revealed convergent mutations in stress-related networks, particularly those governing cell envelope integrity, redox balance, and carbohydrate transport. These findings suggest that extended selection in low-shear environments drives consistent shifts in stress-response and metabolic pathways16,17,25,26,27.
Among the chemical pressures encountered in spacecraft systems, silver-based disinfection represents a particularly potent and relevant selective force. Silver-based disinfection is currently employed in Russian spaceflight systems and is under consideration by NASA as a replacement for iodine due to its broad-spectrum antimicrobial activity and compatibility with closed-loop environments28,29,30. Silver ions act by disrupting membrane integrity, generating reactive oxygen species, and interfering with core biosynthetic pathways31,32,33. However, long-term silver exposure has been shown to drive adaptive resistance via efflux systems, redox remodeling, and cross-resistance to antibiotics34,35,36. Our recent work on Streptococcus mutans evolving under silver and simulated microgravity revealed stable phenotypes such as reduced antibiotic susceptibility and mutations in redox, transport, and membrane-associated genes17. Here, we aim to link reversible gene regulation with irreversible genomic change in the context of long-term silver and microgravity exposure. This challenge is well suited to long-term experimental evolution frameworks, which allow us to track how phenotypic plasticity becomes genetically encoded over time. This process is central to understanding how biological complexity emerges from sustained selection in non-terrestrial environments.
The evolution of microbial resistance in spaceflight settings has consequences that extend beyond molecular biology, influencing both astronaut health and the integrity of built spacecraft environments. The closed nature of these habitats amplifies such risks as microbial populations introduced by crew members can persist, adapt, and colonize interior surfaces, especially when exposed to repeated selective pressures like silver-based disinfection33,37. Biofilm-forming organisms such as S. mutans are particularly well suited for survival in these conditions, combining stress resilience with adhesive capacity14,15,16. Studies of enclosed life support systems, including Lunar Palace 1 and the International Space Station, have shown that the human oral microbiome significantly contributes to the airborne and surface-associated microbial load in space habitats38,39. These findings underscore the need to understand how human-associated bacteria respond to extreme and chemically selective environments, as their adaptation may undermine sterilization efforts and pose long-term risks to crew safety and infrastructure stability.
Despite growing interest in microbial resilience in space, most studies have not addressed the long-term adaptive response of bacteria under persistent spaceflight-like conditions. Most of the existing studies have examined short-term acclimatory responses over hours to a few days. In contrast, long-term evolutionary adaptation under persistent spaceflight-like conditions remains far less explored. Our previous work16,17 used long-term serial passage to study the adaptive evolution of S. mutans under simulated microgravity and under combined silver nitrate exposure across 100 days. This approach enables the characterization of stable, evolved traits rather than transient responses, providing a rare window into bacterial adaptation under sustained spaceflight-like conditions. To investigate how S. mutans respond to the compounded pressures of microgravity and silver, we further analyzed our experimentally evolved populations derived from long-term selection under these conditions. Using RNA sequencing and targeted phenotypic assays, we explored transcriptional, metabolic, and structural adaptations that contribute to microbial persistence in closed, resource-limited spaceflight environments. This approach allows us to dissect both shared and condition-specific molecular responses and to clarify how extreme environmental factors shape bacterial evolution over time.
This work enhances our understanding of microbial evolution in non-terrestrial settings. It also informs approaches in synthetic biology, space medicine, and biocontrol strategies for long-term missions where environmental conditions differ drastically from those on Earth.
Results
Our previous experimental evolution studies established that Streptococcus mutans adapt to simulated microgravity and dual silver stress through combinations of fixed genomic mutations and phenotype-level shifts in adhesion, acid tolerance, antimicrobial susceptibility, and other metabolic traits. These studies identified key genetic targets but did not resolve how these adaptations manifest at the level of global transcription or how genotype and expression patterns relate to downstream phenotypes. Here, we used RNA sequencing to characterize transcriptomic remodeling in 14 independently evolved populations after 100 days under normal gravity (NG), simulated microgravity (sMG), or combined sMG and silver nitrate (sMGAg); all RNA-seq quality metrics are reported in Supplemental Table 1. Differential expression relative to the ancestral strain was quantified for each population (Supplemental Tables 2a–2n). When integrated with prior genomic data and phenotype-level measurements (biochemical assays and ROS profiles), these transcriptomic analyses enabled us to define how evolutionary pressures shape global expression programs, how replicate populations converge or diverge within conditions, and how expression changes align with both pathway-level remodeling and lineage-specific physiological outcomes.
Global transcriptomic shifts differ across conditions
We first compared the total number of differentially expressed genes in each condition relative to the ancestor (Fig. 1a). A one-way ANOVA indicated significant differences among conditions (F₂,₁₁ = 5.26, p = 0.025). Post hoc Tukey tests showed that NG populations had significantly more DEGs than both sMG and sMGAg (p < 0.05 for each comparison), whereas sMG and sMGAg did not differ. Indicating that, microgravity-based evolution produced a more streamlined transcriptional response compared to the broader remodeling observed in NG.
a Total numbers of differentially expressed genes (DEGs) for each condition, classified as upregulated or downregulated relative to the ancestral strain. Statistical contrasts for each condition are provided in Supplemental Table 1, with lineage-specific DEG lists in Supplemental Table 2a–n. b Principal component analysis (PCA) of log₂ fold-change values shows distinct transcriptional profiles across conditions. Ellipses indicate the dispersion of sMG and sMGAg populations; NG contains two replicates and is shown for reference. c Replicate dispersion, calculated as each sample’s distance to its group centroid in PC1–PC2 space. NG is displayed without statistical testing due to n = 2; no significant difference was detected between sMG and sMGAg. d Pairwise Jaccard similarity of DEG sets across replicates within each condition. Significance levels: ns = not significant; *p < 0.05; **p < 0.01; ***p < 0.001.
Principal Component Analysis of all expressed genes (Fig. 1b) showed a distinct separation between NG and both microgravity-derived conditions. NG replicates occupied a distinct region of PC space, whereas sMG and sMGAg clusters partially overlapped, indicating similar but non-identical transcriptional trajectories under microgravity with or without silver. Because NG included only two populations, a 95% confidence ellipse could not be calculated.
Replication-level variability was quantified using each population’s Euclidean distance to its condition centroid in PC1–PC2 space (Fig. 1c). NG populations each had identical distances (9.92). Because only two replicate populations were available, NG was excluded from statistical comparisons. sMG populations showed the widest within-condition dispersion (median = 4.923, n = 8), while sMGAg populations showed a numerically lower dispersion (median = 3.660, n = 4). However, this difference was not statistically significant. An exact two-tailed Mann–Whitney test yielded U = 9 and p = 0.2828, indicating that the variability among independently evolved lineages was comparable under microgravity alone and microgravity plus silver.
Pairwise transcriptional similarity was also assessed by computing Jaccard similarity indices for DEGs between replicate populations (Fig. 1d). sMG showed a broad range of Jaccard values (0.19–0.68, median = 0.309, n = 28), reflecting the diversity observed in PCA. sMGAg populations showed moderately higher similarity (0.34–0.70, median = 0.360, n = 6). A two-group Mann–Whitney comparison again showed no significant difference between the conditions (p = 0.283). Together, Fig. 1a–d show that NG produces a broader and more distinct transcriptional program, whereas sMG and sMGAg share partially convergent trajectories with comparable replicate-level variability. Because NG, sMG, and sMGAg include different numbers of independently evolved populations, any apparent differences in transcriptional dispersion should be interpreted cautiously and may partly reflect sampling depth in addition to biological variability.
Carbohydrate uptake and metabolism, secondary metabolism, lipid biosynthesis, and ion homeostasis represent the dominant axes of transcriptomic remodeling
To identify functional processes underlying these transcriptional changes, DEGs were mapped to Clusters of Orthologous Groups (COGs) and enrichment was tested using Fisher’s exact tests with Benjamini–Hochberg correction (Supplemental Table 3). Carbohydrate transport and metabolism (COG G) and secondary metabolite biosynthesis, transport, and catabolism (COG Q) were significantly enriched in all 14 populations. Lipid metabolism (COG I) was enriched in sMG1, sMG4, sMG7, sMG8, sMGAg2, and sMGAg4. Inorganic ion transport (COG P) was enriched in NG2, NG4, and sMG4. Heatmaps of all COG-assigned DEGs are shown in Fig. 2a–d.
Heatmaps show log₂FC in gene expression relative to the ancestral strain for all differentially expressed genes assigned to major Clusters of Orthologous Groups (COGs). Each column represents an independently evolved population under normal gravity (NG), simulated microgravity (sMG), or combined microgravity with silver nitrate (sMGAg), and each row corresponds to a single COG-annotated gene. The color scale is centered at log₂FC = 0, rendered in white, with upregulated genes shown in pink and downregulated genes shown in purple. a COG G: carbohydrate transport and metabolism. b COG Q: secondary metabolite biosynthesis, transport and catabolism. c COG I: lipid transport and metabolism. d COG P: inorganic ion transport and metabolism. The heatmaps summarize condition-associated transcriptional remodeling without implying direct statistical contrasts between environments. Statistical tests for enrichment are reported in Supplemental Table 3 and populations showing statistically significant enrichment are denoted by *p < 0.05; **p < 0.01; ***p < 0.001 in the population name on the x-axis.
COG G genes showed consistent remodeling across all lineages (Fig. 2a). Core phosphotransferase system components including levA–D, manN, manM, manL, fruA, fruD, and ptsG were downregulated in nearly all populations (typical log₂FC ≈ −1.3 to −3.6). Glycogen-associated genes (glgP, glgB, glgC, glgD, glgA) showed similarly broad repression. In contrast, the trehalose utilization genes treC and treB were upregulated in 13 of 14 lineages, with log₂FC values ranging from approximately +1.3 to +4.8. NG populations additionally showed induction of the cellobiose operon (celA–D), especially in NG4. Most sMG and sMGAg lineages strongly upregulated lactose catabolic genes (lacA, lacB, lacD, lacE, lacG, lacX), with the exception of sMG3 and sMG4 where they were repressed. Thus, although all populations exhibited broad PTS downregulation and trehalose induction, each condition displayed additional characteristic features within carbohydrate pathways.
COG Q genes, dominated by the mutanobactin biosynthetic cluster, were uniformly downregulated in all 14 populations (Fig. 2b). mubA, mubB, mubC, mubD, mubH, mubI, mubJ, mubM, mubP, mubZ, and adjacent loci such as pgap_annot_000793 and pgap_annot_000804 showed negative log₂FC values across all lineages (approximately −1.5 to −5.7). No population showed activation nor directional variability within this cluster.
Although COG I was enriched only in six lineages, lipid metabolism genes were differentially expressed across all populations (Fig. 2c). The fatty-acid biosynthesis enzyme fabD was strongly downregulated in nearly every lineage ( − 1.6 to −5.5), with particularly high magnitude reductions in NG4, sMG5, sMG7, and sMGAg2. Additional biosynthetic genes (accA, accB, accC, accD, fabF) were significantly reduced in sMGAg2. The dehydrogenase hgdC was broadly repressed across all populations. A few genes showed lineage-restricted upregulation, such as pgap_annot_001143 in NG2, NG4, sMG3, sMG4, sMG6, and sMG7, and bcrC in sMGAg4.
Although category-level enrichment for COG P occurred only in NG2, NG4, and sMG4, nearly all populations exhibited remodeling of ion transport and homeostasis genes (Fig. 2d). Multiple metal- and ion-handling genes including sitB which was only differentially expressed in sMG and sMGAg, mtsC, trkA, gltJ, gltK, yxeN, and ytmL were upregulated in many lineages, while the msm ABC transporter system (msmX, msmG, msmF) was strongly downregulated ( − 1.7 to −4.9) in all lineages but sMG5. Copper efflux genes copA and copZ were induced in several populations, including NG2, NG4, sMG2, and the sMGAg lineages.
Taken together, the COG analysis shows that all populations converged upon broad suppression of PTS-mediated carbohydrate uptake, uniform repression of secondary metabolite biosynthesis, widespread downregulation of fatty-acid biosynthesis, and remodeling of metal and ion transport pathways, with NG, sMG, and sMGAg each exhibiting additional condition-specific transcriptional signatures. Because the total number of DEGs varied across lineages, condition-specific enrichment patterns should be interpreted cautiously, as differences in replicate number (NG = 2 vs. sMG = 8 vs. sMGAg = 4) can influence apparent enrichment.
GO analysis reinforces major COG-level themes and adds pathway resolution
Gene Ontology Biological Process enrichment was performed using the DAVID Bioinformatics Resources, which evaluates over-representation of functional categories using modified Fisher’s exact tests with Benjamini–Hochberg correction. Significant and near-significant terms for all populations are summarized in Supplemental Table 4. GO enrichment reinforced the same major axes of transcriptomic remodeling detected in the COG analysis, with added pathway-level granularity.
The most consistently enriched GO term across lineages was “phosphoenolpyruvate-dependent sugar phosphotransferase system” (GO:0009401), which was significant in 12 populations and near significant in the remaining two. This pattern directly reflected the broadly shared downregulation of ptsG, fruC, fruI, ptnA, and bglP observed in Fig. 3. Similarly, “secondary metabolite biosynthetic process” (GO:0044550) and “amino acid activation for nonribosomal peptide biosynthesis” (GO:0043101) were enriched in nearly all lineages and both were driven by the same mutanobactin-associated gene cluster that dominated COG Q.
Heatmaps show log₂ fold-change (log₂FC) in gene expression relative to the ancestral strain across populations evolved under NG, sMG, and sMGAg. The color scale is centered at log₂FC = 0 (white), with upregulation in pink and downregulation in purple. Genes are grouped by enriched GO terms. Rows represent individual genes and columns represent evolved populations. Heatmaps summarize transcriptional patterns and do not imply statistical contrasts between environments. Significant and near-significant terms are summarized in Supplemental Table 4.
GO analysis also resolved specific carbohydrate-processing pathways, including “lactose catabolic process via tagatose-6-phosphate” and “D-tagatose-6-phosphate catabolic process”, which corresponded directly to the induction or repression of lacC, lacD, and lacG in Fig. 3. These pathway-specific enrichments showed that although all evolved populations remodeled carbohydrate metabolism broadly, each lineage utilized different combinations of sugar pathways.
Thus, GO enrichment confirmed the robustness of the major functional shifts observed in COG analysis and clarified how specific carbohydrate and secondary-metabolite gene sets contributed to multiple GO categories. GO analysis did not reveal additional axes of adaptation beyond those already detected by COGs.
Shared differentially expressed genes reveal a conserved core and condition-specific modules
We next evaluated convergence within and across conditions by identifying DEGs shared among replicate lineages. NG populations shared 173 DEGs (86 downregulated, 87 upregulated), sMG populations shared 46 DEGs (35 down, 11 up), and sMGAg populations shared 62 DEGs (50 down, 12 up) as shown in Fig. 4a. These differences partly reflect the number of replicate populations in each condition. Chi-square tests demonstrated significant differences in the ratio of downregulated to upregulated shared DEGs when comparing NG to sMG (χ² = 10.23, p = 0.0014) and NG to sMGAg (χ² = 17.83, p < 0.0001), whereas sMG and sMGAg did not differ (χ² = 0.324, p = 0.569). As with general DE pattersn, these results indicate that both microgravity conditions shared a more strongly downregulated common expression signature relative to NG.
a Total number of DEGs shared among populations within each condition (NG, sMG, sMGAg), separated into upregulated (orange) and downregulated (yellow) genes relative to the ancestral strain. Bars show the number of genes consistently differentially expressed across all populations within each environment. b Venn diagram showing the number of DEGs unique to or shared among NG, sMG, and sMGAg populations. Counts indicate genes consistently differentially expressed within each condition. These summaries describe overlap patterns but do not imply statistical contrasts between conditions. Gene identities are located in Supplemental Table 5.
A core set of 38 DEGs was shared among all three conditions (Fig. 4b, Supplemental Table 5). This set consisted largely of carbohydrate-associated genes such as dexB, galT, ugpC, treC, and treP, and nearly the entire mutanobactin/ABC transporter locus including mub family genes and msmE. These genes represent the most consistently remodeled transcriptional module across the evolutionary environments shown in the COG and GO analysis.
Each condition also had its own distinct set of repeatedly remodeled genes (Supplemental Table 5). NG-specific shared DEGs included carbohydrate-active and protein-quality control genes such as pgap_annot_000087 (α-amylase) and chaperone-associated loci (dnaK, grpE). sMG-specific DEGs included lacD and multiple mutanobactin and ABC transporter genes, consistent with widespread changes to carbohydrate uptake and secondary metabolism pathways. sMGAg-specific shared DEGs included pgap_annot_000162 (MerR-like regulator), pgap_annot_001649, and multiple membrane-associated transporters.
These patterns show that although a small, conserved core was shared across all conditions, each environment repeatedly remodeled additional pathway modules specific to its selective pressures. Again, differences in the number of shared DEGs partly reflect differing numbers of replicate populations (NG = 2, sMG = 8, sMGAg = 4), which influence how often DEGs recur across lineages.
DNA-seq integration identifies lineage-specific genomic variants that align with local expression changes
To contextualize the transcriptomic patterns, we compared RNA-seq results with previously published DNA-seq datasets for the same 14 evolved populations16,17. Comparison showed that only a limited subset of genomic variants occurred in pathways that also displayed expression changes (Supplemental Table 2), and these relationships were lineage-specific rather than condition-wide. Most major transcriptional programs: such as widespread PTS downregulation, uniform suppression of the mutanobactin locus, reduced fatty-acid biosynthesis, and broad remodeling of ion-homeostasis genes, occurred in populations regardless of whether they carried mutations in the corresponding pathways.
A small number of redox-associated variants occurred across the evolved populations. rex mutations were present only in NG4 and sMG2, whereas gorA mutations appeared in sMG2, sMG5, sMG7, sMG8, sMGAg2, and sMGAg3, and sMGAg1 carried a variant upstream of ahpC. Several of these same lineages also exhibited differential expression of antioxidant genes: NG4 showed induction of ahpF, ahpC, and sodA; sMG2 showed sodA induction; sMG3 and sMG5 showed ahpC induction (with sMG5 additionally inducing sodA); sMGAg1 showed increased ahpF and ahpC expression; and sMGAg2 showed increased sodA expression. Although these RNA-level shifts were not restricted to mutated backgrounds, the co-occurrence of rex, gorA, and ahpC-adjacent mutations with altered expression of antioxidant and thioredoxin-related genes provides lineage-specific genomic context for interpreting subsequent ROS phenotypes in the subsequent section.
Ion-transport variants were also found across multiple backgrounds. Mutations in trk potassium-uptake genes were present in NG2, sMG3, sMG5, sMG6, sMGAg1, sMGAg2, and sMGAg3. Many of the same lineages also showed differential expression of trxA, including NG2, sMG2, sMG3, sMG4, and sMGAg2. trxA is involved in maintaining the reduced thiol state of proteins, and potassium gradients influence redox homeostasis and membrane energetics, therefore the co-occurrence of trk mutations and trxA induction in several lineages may indicate overlapping pressures on ion-handling and oxidative-stress balance, although again these patterns were not consistent across all populations with trk variants.
Mutations in carbohydrate-associated genes were more limited. NG2 carried a high-frequency 30-bp deletion in a PTS IIB component, and sMGAg1 possessed an intergenic mutation immediately upstream of lacA. These variants occurred in lineages that also showed altered PTS or lactose-operon transcription, though similar expression shifts were observed in populations without such mutations, indicating that genomic variation was not the primary driver of carbohydrate remodeling.
Several microgravity-evolved lineages also carried envelope-associated variants, including mutations in vicK (sMG2, sMG3), mltG (sMG3), mreD (sMG6), and a PBP1A-like gene (sMG7). These variants were found in populations exhibiting changes in envelope remodeling and cell-wall biosynthesis gene expression, though again, these patterns also occurred in lineages lacking mutations in these loci.
Overall, the comparison between DNA-seq and RNA-seq shows that the major transcriptomic signatures of adaptation were not explained by parallel genomic changes. These lineage-specific variants cannot be interpreted as causal drivers of the observed expression changes, as similar transcriptional patterns occurred in lineages without mutations in these pathways. The handful of lineage-specific variants that occurred in redox, ion-transport, carbohydrate, or envelope-associated pathways provide additional mechanistic context for those specific lineages, but do not account for the broad, condition-scale regulatory programs revealed by RNA-seq. This reinforces that transcriptomic data were essential for resolving the dominant adaptive responses to microgravity and under silver stress.
Biochemical profiling reveals environment-specific metabolic shifts not captured by RNA-seq or DNA-seq alone
To evaluate metabolic changes among lineages adapted NG, sMG and sMGAg, we used the API 20 NE and API 20 STREP biochemical identification systems from bioMérieux®. These standardized panels assess carbohydrate fermentation, enzymatic activities, and other metabolic traits widely used for bacterial characterization. Although API 20 NE is designed for non-enteric Gram-negative bacteria, several assays report broadly conserved biochemical functions that provide a useful survey of metabolic capabilities across taxa. The API 20 STREP system is optimized for catalase-negative Gram-positive cocci, enabling confirmation of species identity and assessment of whether evolved lineages retained characteristic Streptococcus mutans biochemical signatures. Together, these assays provided a consistent and comparable set of measurements for assessing phenotypic outcomes across all evolved populations, biochemical profile data from both tests are summarized in Table 1a and b.
The ancestral strain fermented glucose, mannose, sorbitol, lactose, raffinose, and inulin, hydrolyzed esculin, produced β-galactosidase activity and a positive Voges–Proskauer reaction, and was negative for nitrate reduction, urease, indole production, and gelatin hydrolysis. All NG populations retained this complete ancestral profile, consistent with phenotypic stability in this condition.
In contrast, all sMG populations lost the ability to ferment glucose, mannose, sorbitol, lactose, raffinose, and inulin and were negative for esculin hydrolysis, β-galactosidase activity, and the Voges–Proskauer test, while retaining trehalose fermentation. Although several carbohydrate uptake and processing genes were downregulated in sMG, many of these genes were also differentially expressed in NG without associated phenotype changes, indicating that differential expression alone did not predict API-measured activities. Therefore, phenotype-level gains or losses of these activities occurred even when transcriptional profiles at the gene and pathway level appeared similar.
sMGAg populations shared most sMG-associated carbohydrate losses but differed in several traits. All four retained β-galactosidase activity, and three of four partially recovered esculin hydrolysis, whereas this activity was fully absent in sMG. Genes commonly associated with esculin processing, such as dexB and bglP, were reduced in expression across NG, sMG, and sMGAg, including in lineages that regained activity, indicating that esculin restoration in sMGAg does not map directly onto expression patterns of these loci. Lactose fermentation, lost in sMG, was restored only in sMGAg1. Although lactose operon genes were upregulated in several sMGAg lineages, again, these transcriptional patterns alone cannot explain the phenotype. Across all sMG and sMGAg populations, the trehalose operon (treR, treP, treC) was consistently upregulated, and trehalose was the only fermentative carbohydrate phenotype maintained, indicating that trehalose utilization was uniquely preserved under simulated microgravity.
A distinct phenotype emerged exclusively in sMGAg: all four populations reduced nitrate to nitrogen. Because nitrate was present during evolution in the form of silver nitrate and available during the assay, this likely reflects reduction of the environmental nitrate supplied during evolution and assay conditions, rather than a silver-specific adaptation. Several oxidoreductase-associated genes were induced in sMGAg relative to the ancestor, but no mutations were detected in these loci.
Across conditions, these biochemical profiles demonstrate clear and reproducible environment-specific metabolic outcomes. The results also show that RNA-seq and DNA-seq alone cannot fully capture the functional consequences of long-term evolution; direct phenotypic assays are essential for resolving the integrated metabolic outputs shaped by regulatory and genomic change.
Distinct ROS profiles emerge from lineage-specific redox and ion-homeostasis changes and from silver exposure
We then quantified intracellular ROS using CM-H₂DCFDA fluorescence under untreated and H₂O₂-treated conditions. CM-H₂DCFDA passively diffuses into cells, is deacetylated by intracellular esterases, and subsequently forms thiol-conjugated adducts; oxidation of these intracellular adducts generates a stable fluorescent signal retained within the cytoplasm. This enhanced retention makes CM-H₂DCFDA well suited for quantifying ROS in long-term or dynamically changing conditions such as those examined here. Statistical contrasts were performed relative to the ancestral control for NG (Supplemental Table 6), sMG (Supplemental Table 7), and sMGAg (Supplemental Table 8), and for +/-AgNO₃ in the growth and assay conditions within sMGAg (Supplemental Table 9).
When ancestral S. mutans is grown overnight in HARVs we see a small decrease in basal ROS and a loss of induction upon H2O2 treatment (Fig. 5a). Basal ROS remained largely unchanged among NG lineages compared to the ancestral grown in normal conditions except for NG2 which showed only a slight elevation. After H₂O₂ challenge, NG1 showed a significant increase in induction, NG2 was uninducible and NG3 and NG4 exhibited induction to levels seen in the ancestor. These patterns suggest that NG populations maintained ancestral-like basal redox physiology and differed modestly in inducibility, with only one population showing increased ROS upon oxidative challenge.
a Basal and oxidant-treated reactive oxygen species (ROS) levels in NG and sMG populations. Bars show averaged fluorescence (RFU) normalized to OD₆₀₀ for untreated (black) and H₂O₂-treated (pink) samples. Letters indicate significant groupings within each treatment based on multiple-comparison procedures, and asterisks denote significance between untreated and treated samples within the same lineage. All lineage-specific statistical contrasts relative to the ancestral strain are reported in Supplemental Table 6 (NG) and Supplemental Table 7 (sMG). b Basal and oxidant-treated ROS levels in sMGAg populations, measured first after growth without silver nitrate and again after growth in the presence of AgNO₃ ( + AgNO₃). Bars follow the same normalization and statistical notation used in panel (a). Error bars represent the standard error of the mean (SEM). Lineage-level contrasts relative to the ancestor are reported in Supplemental Table 8, and all within-lineage ±AgNO₃ comparisons are reported in Supplemental Table 9.
sMG produced a more heterogeneous outcome. sMG populations either showed basal ROS comparable to the ancestor (sMG1 and sMG6), while the remainder showed basal ROS levels significantly lower than that of the ancestral. After H₂O₂ exposure, sMG1, sMG2, sMG4, showed ROS levels higher than that of the ancestor, sMG5 and sMG8 showed ROS levels lower than ancestral while sMG3, sMG5 and sMG7 had ROS levels similar to the treated ancestor. sMG5 and 6 were uninducible upon H₂O₂ challenge. These divergent inducibility patterns are consistent with the genomic heterogeneity present in these populations: rex mutations were found only in NG4 and sMG2, gorA mutations occurred in sMG2, sMG5, sMG7, and sMG8, and sMGAg1 carried a variant upstream of ahpC. Several of these same lineages also showed differential expression of antioxidant genes in RNA-seq (sMG2 induction of sodA; sMG3 and sMG5 induction of ahpC; sMG5 induction of sodA), although these genomic and transcriptional differences do not map cleanly onto the ROS phenotypes. These associations are correlative and do not demonstrate that rex, gorA, or ahpC variants directly caused the observed ROS phenotypes.
Silver-adapted populations assayed without silver exhibited generally higher basal ROS than ancestral (Fig. 5b). After H₂O₂ treatment, sMGAg2, sMGAg3, and sMGAg4 showed significant induction, whereas sMGAg1 remained non-inducible. When the sMGAg populations were assayed in the presence of AgNO₃, a clear pattern emerged. Specifically, the ancestral strain exposed to AgNO₃ showed the highest ROS values observed in any condition. ROS levels rose within each lineage relative to its own no-silver baseline, demonstrating that silver exposure continued to impose oxidative load (Supplemental Table 9). However, the magnitude of this increase was markedly lower than that seen in the ancestral strain exposed to AgNO₃. Across all sMGAg lineages, ROS levels in AgNO₃ remained significantly below the ancestral +AgNO₃ benchmark (Supplemental Table 9), indicating that silver adaptation reduced the severity of silver-triggered oxidative stress relative to the naïve ancestor. At the same time, every sMGAg lineage still responded to AgNO₃ with an increase over its own untreated state, showing that silver continues to elicit oxidative pressure but that silver-evolved populations are preconditioned to tolerate it. These responses may have occurred in the context of several lineage-specific mutations affecting redox-relevant pathways, including gorA (sMGAg2 and sMGAg3), the upstream ahpC variant in sMGAg1, and multiple trk potassium-transport mutations (sMGAg1–3). Some of these same lineages also exhibited expression changes in ahpF, ahpC, sodA, or trxA, but again these transcriptional signatures were not consistently aligned with ROS outcomes.
Together, these results indicate that NG, sMG and sMGAg populations each exhibited distinct ROS phenotypes relative to the ancestral strain. These findings again underscore that phenotype-level assays are essential for interpreting adaptive outcomes that could not be inferred from genomic or transcriptomic data alone.
Discussion
Microbial persistence in extraterrestrial environments depends on the balance between regulatory plasticity, lineage-specific genetic variation, and phenotype-level expression. Streptococcus mutans, an opportunistic oral pathogen with strong metabolic and redox sensitivity, provides a tractable model for examining how microorganisms adapt to space-relevant stressors. Across NG, sMG, and sMGAg, single-gene mutations and differential expression did not reliably predict metabolic or redox phenotypes, and populations with similar DEGs often occupied distinct regions of PCA space. These patterns indicate that phenotype-level outcomes emerge from coordinated network-level transcriptional remodeling rather than changes in individual genes, underscoring the need to integrate genomic, transcriptomic, and biochemical data to resolve adaptive states.
DNAseq, ROS and biochemical assays were performed across all four NG, eight sMG, and four sMGAg populations, so interpretations are supported by greater population-level replication. In contrast, RNA-seq was available for only two NG populations, limiting interpretation of NG transcriptional variance. Carbohydrate pathways represented a dominant axis of adaptation both at the phenotypic and transcriptomic level. NG lineages retained broad sugar fermentation, consistent with reversible regulatory plasticity rather than genetic fixation. In contrast, sMG produced systematic contraction of carbohydrate use; glucose, raffinose, mannose, sorbitol, and lactose fermentation were lost, despite many associated PTS genes being downregulated in both NG and sMG. This disconnect reinforces that transcriptional shifts alone did not predict carbohydrate phenotypes, consistent with broader reports that microgravity disrupts central metabolism in S. mutans15 and other bacteria6,40. Trehalose metabolism was the notable exception: tre operon upregulation occurred in nearly every lineage, and trehalose was the only fermentative trait consistently maintained, aligning with its role as a stabilizing sugar across diverse stresses41,42. Under sMGAg, lactose operons were transcriptionally induced in several lineages, yet phenotypic recovery occurred only in the population carrying an upstream lacA mutation, echoing prior observations of stress-enabled metabolic reactivation43,44.
Redox management showed divergent phenotypes between lineages under microgravity but converged under dual stress. Some sMG lineages carried variants in rex or gorA, yet these did not map cleanly onto basal or induced ROS phenotypes, consistent with rex functioning as a redox sensor45 and prior evidence of destabilized redox homeostasis in low-shear systems46. Repeated silver exposure drove all sMGAg lineages toward elevated basal ROS combined with markedly attenuated inducibility when exposed to silver, reflecting physiological preconditioning rather than mutation-driven convergence. This pattern is consistent with well-established silver-induced oxidative cycling and envelope-associated stress responses47,48.
Envelope regulation also shifted in environment-specific ways. Under sMG, LPXTG-anchored wall proteins were broadly altered and several lineages acquired variants in vicK or mltG, mirroring known microgravity effects on envelope-associated signaling49. Under sMGAg, populations instead upregulated mreD and multiple LPXTG/LysM-associated wall genes, often without corresponding genomic mutations. This potential strengthening of the cell envelope under dual stress parallels responses to redox-active antimicrobials, including nanosilver, which target envelope integrity and stimulate efflux-associated defenses50. These patterns hint that the cell wall is not simply collateral damage but potentially a recurrent target of adaptive remodeling, underscoring the need for further investigation.
One of the most distinctive outcomes was the emergence of nitrate reduction capabilities in all sMGAg lineages, absent in both NG and sMG. Although S. mutans lacks the canonical membrane-bound nitrate reductase complex (NarG/NarH) found in respiratory nitrate-reducing bacteria51, RNAseq showed that sMGAg lineages induced multiple oxidoreductase-associated genes. This novel phenotype likely reflects expanded use of environmental nitrate, present throughout evolution as silver nitrate, rather than a silver-specific response.
Synthesizing these outcomes, we propose a general model of stress adaptation: NG populations rely on reversible plasticity; sMG destabilizes metabolism, generating divergent lineage-specific trajectories; and dual stress channels this diversity into convergence, with shared redox and envelope remodeling (Fig. 6) consistent with other bacterial outcomes in low-shear stress7. As replicate numbers vary across datasets (RNA-seq: NG = 2 vs. sMG = 8 and sMGAg = 4; DNA-seq/ROS/API: NG = 4), we interpret convergence and divergence as qualitative patterns supported by full replication for phenotypes and more limited replication for NG transcriptomic variance. For spaceflight microbiology, these results suggest that although microgravity alone may destabilize metabolic capacity, layered environmental pressures, such as concurrent antimicrobial exposure, can paradoxically stabilize physiological outcomes, selecting for narrower but more predictable functional repertoires.
Schematic overview of the dominant patterns documented in genomic, transcriptomic, and phenotypic datasets from each evolutionary environment. (Top) under normal gravity (NG), (Middle) under simulated microgravity (sMG), and (Bottom) under simulated microgravity co-adapted with silver nitrate (sMGAg). Generated in BioRender.
More broadly, our findings underscore that microbial adaptation in engineered habitats is not defined by uniform genetic solutions but by flexible regulatory systems that navigate multiple evolutionary routes. Critically, genotype and transcriptional data alone were insufficient to predict adaptive outcomes: carbohydrate losses, nitrate reduction, and redox phenotypes each required direct phenotypic assays for correct interpretation. Integrating DNA-seq, RNA-seq, and phenotype-level measurements therefore provides the most complete view of microbial adaptation under microgravity and dual stressors, offering a framework for anticipating microbial behavior in long-duration spaceflight systems and for designing targeted antimicrobial strategies in clinical and built environments.
Methods
Bacterial Strains
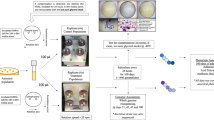
The ancestral Streptococcus mutans strain Clarke NCTC 10449 (ATCC 25175) was used as the baseline reference strain, consistent with our previous long-term adaptation experiments16. For this study, we analyzed bacterial populations that had undergone 100-days of evolution under three environmental conditions: normal gravity (NG), simulated microgravity (sMG), and sMG with silver nitrate exposure (sMGAg). This included four NG strains; however, only two were used for RNA-seq due to contamination (observed via microscopy and plating on BHI agar) and loss of the other two glycerol stocks during experimentation, eight sMG strains, and four sMGAg strains. All cultures were grown in Brain Heart Infusion (BHI) broth or on BHI agar at 37 °C in a 5% CO₂ atmosphere unless otherwise noted. In all sMGAg culture conditions, silver nitrate was maintained at a final concentration of 10 µg/mL (the original selection concentration).
Simulated microgravity and HARV preparation
Low-shear modeled microgravity (LSMMG) was simulated using High Aspect Rotating Vessels (HARVs) manufactured by Synthecon Inc. (Houston, TX). The vessels were operated with their rotation axis oriented perpendicular to the gravity vector, a configuration that randomizes sedimentation and shear forces, effectively mimicking the microgravity conditions encountered during spaceflight6,7. A rotation speed of 25 rpm was selected based on previous studies that identified this setting as optimal for achieving LSMMG in bacterial systems6,7,8. This speed maintains uniform suspension and nutrient diffusion, minimizing gradients and ensuring consistent physiological stress7,52. The resulting shear environment also approximates fluid forces in the oral cavity, making it relevant for modeling S. mutans physiology14. Prior to inoculation, HARV components were cleaned with a mild detergent, soaked in a 25% bleach solution for 15 min, and rinsed thoroughly with distilled water. The vessels were autoclaved at 121 °C for 20 min and cooled under sterile conditions. To confirm sterility, HARVs containing sterile BHI were incubated at 25 rpm and 37 °C for 24 h prior to inoculation.
RNA extraction and culture conditions
Frozen glycerol stock cultures were revived and inoculated into sterile HARVs in triplicate for each population. NG1 and NG3 showed contamination upon revival. When plated on BHI agar, these cultures produced larger yellow colonies within 24 h, whereas S. mutans typically forms small white colonies after 48 h under the same conditions. Microscopy confirmed cocci clusters rather than characteristic S. mutans chains. Based on these observations NG1 and NG3 were excluded from RNA sequencing. Cultures were incubated under their respective conditions, NG, sMG and sMGAg until mid-exponential phase (OD₆₀₀ = 0.5), followed by an additional 1-h incubation to stabilize gene expression. After this incubation, the three biological replicates from each condition were pooled into a single composite sample. The pooled cultures were centrifuged at 5000 rpm for 20 min to pellet the cells. Pooling of biological replicates was performed in accordance with established protocols in bacterial transcriptomics, where such pooling has been validated as a reliable approach for representing population-level gene expression and reducing technical noise, especially when condition-level trends are the primary focus53,54. Pellets were immediately preserved using RNAprotect Bacteria Reagent (Qiagen) and stored at −80 °C until RNA extraction. Total RNA was extracted using the Qiagen RNeasy PowerBiofilm Kit® according to the manufacturer’s protocol.
Library preparation and RNA sequencing
Ribosomal RNA was depleted using the Illumina Ribo-Zero Plus rRNA Removal Kit. Strand-specific RNA-seq libraries were prepared with the Illumina Stranded Total RNA Prep with Ribo-Zero Plus Kit, which includes cDNA synthesis, fragmentation, end repair, adapter ligation, and library amplification. Libraries were then sequenced on the Illumina NextSeq 2000 platform at North Carolina A&T State University to generate paired-end reads at high depth for bacterial transcriptomic analysis.
Bioinformatics analysis
Sequencing data were processed using the onboard bcl-convert pipeline to generate FASTQ files, and read quality was assessed using FastQC55. All samples exhibited high sequencing quality (mean Phred >32; ≥88% bases ≥Q30; 1.2–1.7% adapter content). Total base yields ranged from 288–799 million per read direction. Adapter contamination was minimal (1.2%–1.7%), and demultiplexing accuracy was high, with >93% of reads perfectly matching their sample barcodes.
High-quality reads were aligned to the Streptococcus mutans ATCC 25175 reference genome using HISAT2 (v2.2.1) with default parameters56. A custom genome index was built from the reference FASTA using hisat2-build, and this index was used for all alignments. Gene-level quantification was performed using HTSeq-count (v0.13.5) with position-sorted BAM files57. The following parameters were used: -s no (unstranded protocol), -t CDS (to count only coding sequences), -i gene_id (to match gene identifiers), and a minimum alignment quality of 5 (-a 5). Annotations were provided using a curated GTF file. Output files were manually standardized by adding gene ID and count headers for downstream integration. Alignment quality was robust, with 98.2%–99.8% of total reads successfully mapped and 95.5%–99.3% identified as primary alignments. No samples showed cross-chromosomal mate pairing, confirming alignment specificity and integrity. Coverage was deep and uniform (median ~2200 reads/gene; ~5500–11,200 reads/gene per sample).
Differential gene expression analysis was conducted using the edgeR58 package in RStudio59. Read counts were normalized using the Trimmed Mean of M-values (TMM) method. A custom design matrix contrasted each evolved condition against the ancestral reference. Statistical testing was performed using the quasi-likelihood F-test (glmQLFit and glmQLFTest), and genes with an FDR < 0.05 were considered significantly differentially expressed. Differential expression results were further annotated using a custom Python script that parsed GenBank annotations from Streptococcus mutans ATCC 25175 (.gbk file) to extract locus tags, gene names, and product descriptions. These annotations were mapped to each differentially expressed gene, and the resulting files were exported in tsv format for downstream interpretation. All RNAseq QC data is reported in supplemental Table 1. The RNA-seq datasets generated and analyzed in this study are available in the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/) under BioProject ID PRJNA1278699, with associated BioSample accession numbers SAMN49234291–SAMN49234305.
Replicate heterogeneity analysis
To quantify transcriptomic heterogeneity among replicate populations, we performed two complementary analyses using base functions in RStudio59. First, we conducted principal component analysis (PCA) on variance-stabilized logFC values from all differentially expressed genes. For each condition (NG, sMG, sMGAg), we then calculated the Euclidean distance of each replicate to its group centroid in PC1–PC2 space, providing a measure of within-group dispersion. Group differences were tested by Kruskal–Wallis with post hoc Mann–Whitney U tests and Bonferroni correction. Second, we quantified replicate overlap at the gene level by calculating pairwise Jaccard similarity indices of differentially expressed gene sets (FDR < 0.05). Jaccard indices were computed as the size of the intersection divided by the union of DEGs between replicates. Comparisons between sMG and sMGAg were tested by Mann–Whitney U, while NG (n = 2) was summarized without statistical testing.
Custom mapping and annotation integration
Differentially expressed (DE) genes from RNA-seq analyses of Streptococcus mutans populations evolved under normal gravity (NG), simulated microgravity (sMG), and silver-adapted simulated microgravity (sMGAg) conditions were first annotated using a custom multi-source strategy to maximize functional coverage. Since S. mutans, ATCC 25175 (the experimental strain), lacks full annotation in standard pipelines, we constructed a cross-reference table that integrated multiple data sources. Predicted proteins from the S. mutans UA159 reference genome (NC_004350.2; UniProt proteome UP000002512) served as the annotation backbone. UniProtKB mapping files (including gene names, EC numbers, GO terms, and functional descriptions) were linked to PGAP-formatted locus tags (originating from the ATCC 25175 sequence file) via sequence-based loose matching, ensuring accurate correspondence between experimental DE gene nomenclature used in our original alignment and reference annotations using eggNOG-mapper 6.0.560. To further enrich functional mapping, we integrated files from the Gene Ontology Consortium: interpro2go, pfam2go, ec2go, and kegg_reaction2go. These enabled indirect assignment of GO Biological Process terms to genes with InterPro, Pfam, EC, or KEGG annotations even in the absence of direct GO mappings. This framework improved annotation coverage for under-annotated genes.
COG enrichment and directionality analysis
To characterize functional shifts in orthologous gene groups, DE genes were mapped to COG (Clusters of Orthologous Groups)61 categories using EggNOG-mapper 6.0.260. Each population’s DE gene list was tested for enrichment of specific COG categories using Fisher’s exact test, comparing the number of DE genes in each COG to the total number of genes in that COG in the background genome. Bonferroni-adjusted p-values were used to control for multiple testing, and COG categories with FDR < 0.05 were considered significantly enriched. For enriched COGs, regulation direction was assessed from log₂FC values within each COG that showed significant enrichment and generated heatmaps in GraphPad Prism ® to visualize changes in regulation associated with each COG category. Significance’ corresponds specifically to the Bonferroni-adjusted p-value from Fisher’s exact enrichment test for that COG category.
Gene Ontology (GO) and DAVID Analysis
GO enrichment was performed using the DAVID Bioinformatics Resources v6.862, with each population’s significant DE genes (FDR < 0.05) used as foreground lists. The full S. mutans UA159 UniProt proteome (UP000002512) was supplied as a custom background for consistent cross-sample comparison. Enrichment of GO Biological Process terms was assessed using modified Fisher’s exact testing, and significance was defined using the Benjamini–Hochberg method (FDR < 0.05).
KEGG pathway enrichment
KEGG pathway enrichment was also performed in DAVID62 using the same DE gene lists and the S. mutans UA159 UniProt background. No KEGG pathways were significantly enriched (Benjamini–Hochberg FDR < 0.05) in any of the NG, sMG, or sMGAg populations. As a result, KEGG data were not included in the results.
Biochemical profiling
To evaluate metabolic activity, Analytical Profile Index (API) 20 NE and API Strep test strips (bioMérieux Inc.) were used on ancestral, NG, sMG, and sMGAg populations in triplicate. Cultures were grown in HARVs overnight and standardized to a 0.5 McFarland turbidity in 0.85% NaCl solution. Each test strip was inoculated with 100 µL of bacterial suspension per well, and mineral oil was added to anaerobic wells as per protocol. Strips were incubated at 37 °C for 24–48 h in a humidified chamber. Reactions were scored using the manufacturer’s chart. Nitrate reduction was confirmed using NIT1 and NIT2 reagents, and comparative metabolic profiles in comparison to the ancestral were recorded across all conditions.
Reactive Oxygen Species (ROS) detection
Intracellular ROS accumulation was quantified using CM-H₂DCFDA fluorescence which detects total ROS in the cell. Cultures were grown in HARVs (37 °C, 25 rpm, 5% CO₂) for 48 h. Two 1.5 mL aliquots were harvested from each population, centrifuged at 13,000 rpm for 3 min, and washed in Hank’s Balanced Salt Solution (HBSS). Cells were resuspended in a 5 µM CM-H₂DCFDA solution and incubated at 37 °C for 60 min to allow dye loading. Samples were washed, resuspended in HBSS, and split into untreated and 5 mM H₂O₂-treated groups. A total of 200 µL per condition was plated in triplicate in a 96-well clear-bottom plate. Fluorescence was measured using excitation/emission wavelengths of 492–495 nm and 517–527 nm. RFU values were normalized to OD₆₀₀. ROS fluorescence values (RFU/OD₆₀₀) were compared between untreated and H₂O₂-treated samples within each lineage using unpaired two-tailed Student’s t-tests in GraphPad Prism ®. Each lineage was also compared directly against the ancestral strain under both conditions to determine whether evolved populations exhibited altered basal ROS levels or inducibility. Technical replicates (n = 3) were used for each condition, and significance was assigned at p < 0.05. Significance levels are indicated in figures as ns (not significant), *p < 0.05, **p < 0.01, ***p < 0.001, and ***p < 0.0001. To assess broader trends across conditions, one-way ANOVA was also performed separately for NG vs sMG populations, sMG vs sMGAg populations, and sMGAg populations ± AgNO₃. Post-hoc Tukey’s multiple comparisons were applied to test all pairwise contrasts within each grouping. This analysis allowed direct evaluation of differences between groups of evolved populations rather than within-lineage changes alone. Mean differences, 95% confidence intervals, adjusted p-values, replicates (n = 3) were included for each lineage, and significance was assigned at adjusted p < 0.05.
Data availability
The RNA-seq datasets generated and analyzed in this study are available in the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/) under BioProject ID PRJNA1278699, with associated BioSample accession numbers SAMN49234291–SAMN49234305. These data sets will be publicly available at time of publication.
References
Cockell, C. S. Habitable worlds with no signs of life. Philos. Trans. R. Soc. A 372, 20130082 (2014).
Ward P. D., Brownlee D. Messengers from the Stars. Rare Earth: Why Complex Life is Uncommon in the Universe. 2000:277-287.
Horneck, G., Klaus, D. M. & Mancinelli, R. L. Space Microbiology. Microbiol. Mol. Biol. Rev. 74, 121–156 (2010).
Rosenzweig, J. A. et al. Spaceflight and modeled microgravity effects on microbial growth and virulence. Applied microbiology and biotechnology; 2010. 885-891.
Singh, N. K., Wood, J. M., Karouia, F. & Venkateswaran, K. Succession and persistence of microbial communities and antimicrobial resistance genes associated with International Space Station environmental surfaces. Microbiome 6, 1–23 (2018).
Crabbé, A. et al. Use of the rotating wall vessel technology to study the effect of shear stress on growth behaviour of Pseudomonas aeruginosa PA01. Environ. Microbiol. 10, 2098–2110 (2008).
Nickerson, C. A., Ott, C. M., Wilson, J. W., Ramamurthy, R. & Pierson, D. L. Microbial responses to microgravity and other low-shear environments. Microbiol Mol. Biol. Rev. 68, 345–361 (2004).
Wilson, J. W. et al. Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq. Proc. Natl. Acad. Sci. Usa. 104, 16299–16304 (2007).
Zea, L. et al. A Molecular Genetic Basis Explaining Altered Bacterial Behavior in Space. PLoS One 11, e0164359 (2016).
Wade, W. G. The oral microbiome in health and disease. Pharmacol. Res. 69, 137–143 (2013).
Ajdić, D. et al. Genome sequence of Streptococcus mutans UA159, a cariogenic dental pathogen. Proc. Natl. Acad. Sci. 99, 14434–14439 (2002).
Kilian, M. et al. The oral microbiome–an update for oral healthcare professionals. Br. Dent. J. 221, 657–666 (2016).
Lemos, J. A. et al. The biology of Streptococcus mutans. Microbiol. Spectr. 7, gpp3–gpp2018 (2019).
Cheng, X. et al. Effects of simulated microgravity on Streptococcus mutans physiology and biofilm structure. FEMS Microbiol Lett. 359, 94–101 (2014).
Orsini, S. S., April, M. L. & Kelly, C. R. Investigation of simulated microgravity effects on Streptococcus mutans physiology and global gene expression. npj Microgravity 3, 4 (2017).
Fernander, M. C. et al. Adaptation to simulated microgravity in Streptococcus mutans. NPJ microgravity 8, 17 (2022).
Fernander, M. C. et al. Co-adaptation of Streptococcus mutans to simulated microgravity and silver nitrate. Front. Astron. Space Sci. 10, 1183867 (2023).
Del Fabbro, M. et al. Oral Health of Astronauts in Short-and Long-Term Missions in Space. Aerosp. Med. Hum. Perform. 96, 168–179 (2025).
Harika, P. D. et al. Oral health in zero gravity: a comprehensive review of orofacial effects and countermeasures in spaceflights. Cureus. 2023;15.
Lloro, V., Giovannoni, L. M., Lozano-de Luaces, V., Lloro, I. & Manzanares, M. C. Is oral health affected in long period space missions only by microgravity? A systematic review. Acta Astronautica. 167, 343–350 (2020).
Kim, W. et al. Effect of spaceflight on Pseudomonas aeruginosa final cell density is modulated by nutrient and oxygen availability. BMC Microbiol. 13, 1–10 (2013).
Guo, Y. et al. Comparative genomic analysis of Klebsiella pneumonia (LCT-KP214) and a mutant strain (LCT-KP289) obtained after spaceflight. BMC genomics. 15, 1–11 (2014).
Iturriaga, G., Suárez, R. & Nova-Franco, B. Trehalose metabolism: from osmoprotection to signaling. Int. J. Mol. Sci. 10, 3793–3810 (2009).
Novikova, N. et al. Survey of environmental biocontamination on board the International Space Station. Res Microbiol. 157, 5–12 (2006).
Fajardo-Cavazos, P. et al. Evolution of Bacillus subtilis to enhanced growth at low pressure: up-regulated transcription of des-desKR, encoding the fatty acid desaturase system. Astrobiology 12, 258–270 (2012).
Tirumalai, M. R. et al. The adaptation of Escherichia coli cells grown in simulated microgravity for an extended period is both phenotypic and genomic. npjMicrogravity 3, 15 (2017).
Tirumalai, M. R. et al. Evaluation of acquired antibiotic resistance in Escherichia coli exposed to long-term low-shear modeled microgravity and background antibiotic exposure. Mbio; 2019. 2637.
Hicks P., Nelson J., Callahan M. R., editors. Initial Trade Study for In-line Silver Sensor for Spacecraft Potable Water Systems. International Conference on Environmental Systems; (2019).
Ley, S. et al., editors. Fate of Silver Biocide on the International Space Station Living Environment2021: 50th International Conference on Environmental Systems.
Perry, S. Silver-based Microbial Check Valve for Spacecraft Potable Water Systems. 2019.
Espinosa-Cristóbal, L. F. et al. Adherence inhibition of Streptococcus mutans on dental enamel surface using silver nanoparticles. Mater. Sci. Eng.: C. 33, 2197–2202 (2013).
Liau, S. Y. R. Interaction of silver nitrate with readily identifieable groups: relationship to the antibacterial action of silver ions. Lett. Appl. Microbiol. 25, 279–283 (1997).
Mijnendonckx, K., Leys, N., Mahillon, J., Silver, S. & Van Houdt, R. Antimicrobial silver: uses, toxicity and potential for resistance. Biometals 26, 609–621 (2013).
Graves, J. L. Jr. et al. Rapid evolution of silver nanoparticle resistance in Escherichia coli. Front Genet. 6, 42 (2015).
Seiler, C. & Berendonk, T. U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol. 3, 399 (2012).
Tajkarimi, M. et al. Selection for Ionic-silver Confers Silver Nanoparticle Resistance in Escherichia coli. JSM Nanotechnol. Nanomed. 5, 1047 (2017).
Paton, S., Moore, G., Campagnolo, L. & Pottage, T. Antimicrobial surfaces for use on inhabited space craft: A review. Life Sci. Space Res. 26, 125–131 (2020).
Checinska Sielaff, A. et al. Characterization of the total and viable bacterial and fungal communities associated with the International Space Station surfaces. Microbiome 7, 1–21 (2019).
Yang, J., Fu, Y. & Liu, H. Microbiomes of air dust collected during the ground-based closed bioregenerative life support experiment” Lunar Palace 365. Environ. Microbiome. 17, 1–20 (2022).
Sharma, G. & Curtis, P. D. The impacts of microgravity on bacterial metabolism. Life 12, 774 (2022).
Elbein, A. D., Pan, Y. T., Pastuszak, I. & Carroll, D. New insights on trehalose: a multifunctional molecule. Glycobiology 13, 17R–27R (2003).
Tapia, H. & Koshland, D. E. Trehalose is a versatile and long-lived chaperone for desiccation tolerance. Curr. Biol. 24, 2758–2766 (2014).
Notley-McRobb, L. & Ferenci, T. The generation of multiple co-existing mal-regulatory mutations through polygenic evolution in glucose-limited populations of Escherichia coli. Environ. Microbiol. 1, 45–52 (1999).
Manché, K., Notley-McRobb, L. & Ferenci, T. Mutational adaptation of Escherichia coli to glucose limitation involves distinct evolutionary pathways in aerobic and oxygen-limited environments. Genetics 153, 5–12 (1999).
Bitoun, J. P., Nguyen, A. H., Fan, Y., Burne, R. A. & Wen, Z. T. Transcriptional repressor Rex is involved in regulation of oxidative stress response and biofilm formation by Streptococcus mutans. FEMS Microbiol. Lett. 320, 110–117 (2011).
Abshire, C. F. et al. Exposure of Mycobacterium marinum to low-shear modeled microgravity: effect on growth, the transcriptome and survival under stress. npjMicrogravity 2, 1–14 (2016).
Dakal, T. C., Kumar, A., Majumdar, R. S. & Yadav, V. Mechanistic basis of antimicrobial actions of silver nanoparticles. Front. Microbiol. 7, 1831 (2016).
Kamat, S. & Kumari, M. Emergence of microbial resistance against nanoparticles: Mechanisms and strategies. Front. Microbiol. 14, 1102615 (2023).
Foster, J. S., Wheeler, R. M. & Pamphile, R. Host-microbe interactions in microgravity: assessment and implications. Life 4, 250–266 (2014).
Gunawan, C. et al. Nanosilver targets the bacterial cell envelope: the link with generation of reactive oxygen radicals. ACS Appl. Mater. interfaces 12, 5557–5568 (2020).
Moreno-Vivián, C., Cabello, P., Martínez-Luque, M., Blasco, R. & Castillo, F. Prokaryotic nitrate reduction: molecular properties and functional distinction among bacterial nitrate reductases. J. Bacteriol. 181, 6573–6584 (1999).
Klaus, D. M. Microgravity and its implications for fermentation biotechnology. Trends Biotechnol. 16, 369–373 (1998).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Todd, E. V., Black, M. A. & Gemmell, N. J. The power and promise of RNA-seq in ecology and evolution. Mol. Ecol. 25, 1224–1241 (2016).
Andrews S. FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (referenced 02/25/2018).
Zhang, Y., Park, C., Bennett, C., Thornton, M. & Kim, D. Rapid and accurate alignment of nucleotide conversion sequencing reads with HISAT-3N. Genome Res. 31, 1290–1295 (2021).
Anders, S. P. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. bioinformatics 26, 139–140 (2010).
Team RC. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-projectorg/ (2016).
Hernández-Plaza, A. et al. eggNOG 6.0: enabling comparative genomics across 12 535 organisms. Nucleic acids Res. 51, D389–D394 (2023).
Tatusov, R. L., Koonin, E. V. & Lipman, D. J. A genomic perspective on protein families. Science 278, 631–637 (1997).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic acids Res. 50, W216–W221 (2022).
Acknowledgements
This work, along with support for M.C.F., was primarily funded by the Engineering Research Centers Program of the National Science Foundation under NSF Cooperative Agreement No. EEC-2133504 awarded to M.D.T. and J.L.G. K.R.M. was supported by the National Institutes of Health (NIH), National Institute of General Medical Sciences (NIGMS) MARC Undergraduate NRSA Institutional Grant (Award No. 5T34GM083980).
Author information
Authors and Affiliations
Contributions
M.C.F. oversaw and conducted the RNA sequencing experiments. K.R.M. developed and performed the reactive oxygen species (ROS) and biochemical assays, with J.E.S. providing additional support on ROS measurements. B.T.S. conducted functional data analysis of the RNA-seq results and created the associated visualizations. M.D.T. performed library preparation, sequencing, and initial data processing, drafted the first version of the manuscript, and contributed to funding acquisition. J.L.G. contributed to study conception and design, as well as the analysis and interpretation of all data. All authors reviewed and edited the manuscript and approved the final submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fernander, M.C., McClure, K.R., Sanders, B.T. et al. Adaptive transcriptional remodeling of Streptococcus mutans under simulated microgravity and silver stress reveals evolutionary innovation in artificial environments. npj Complex 3, 6 (2026). https://doi.org/10.1038/s44260-025-00068-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44260-025-00068-6