Abstract
SARS-CoV-2 infection disrupts the host’s immune system, altering autoimmune responses. This study investigated host autoreactivities in SARS-CoV-2 infections, their association with severe COVID-19, and the neutralizing antibody response. Circulating autoantibodies were detected in convalescent serum samples from unvaccinated SARS-CoV-2-infected patients. Clustering, correlation analysis, principal component analysis, and neural network modeling were used to explore the relationship between autoantibodies, hospitalization, and SARS-CoV-2 neutralization. The presence of one autoantibody correlated with the detection of multiple others. Anti-IFNα antibodies were associated with elevated levels of anti-ENAs (extractable-nuclear antigen) but not with clinical outcome. COVID-19 hospitalization was significantly associated with the collective expression of autoantibodies targeting three ENAs: SSA/Ro52, Jo-1, and RNP. In contrast, autoantibodies targeting RNP/Sm, PCNA, Scl-70, and PL-12 were strongly associated with SARS-CoV-2 neutralization. In summary, this study has identified self-antigens targeted in hospitalized COVID-19 patients and highlights a novel association between the autoantibody response and the antiviral humoral response.
Similar content being viewed by others
Introduction
Autoantibodies that target various cellular components are linked to several autoimmune diseases (ADs). Over 100 ADs have been described and numerous autoepitopes identified, since the discovery of anti-DNA antibodies in systemic lupus erythematosus (SLE) patients in the 1940s1,2,3. Common targets of autoantibodies include antinuclear antibodies (ANA) targeting DNA and histones, extractable nuclear antigens (ENA) present in the nucleus and cytoplasm, phospholipids, and antineutrophil cytoplasmic antibodies (ANCA) directed against the cytoplasmic components of neutrophils. Preexisting autoantibodies can influence the course of infectious diseases, with viruses potentially triggering autoimmune conditions2,4,5,6,7. Elevated autoantibody levels are observed in severe SARS-CoV-2 infected patients, although it remains unclear if autoantibodies are the cause or consequence of severe COVID-198,9,10,11. During SARS-CoV-2 infection, autoantibodies are likely triggered by virus-induced cytokine storm12,13, molecular mimicry between viral and host proteins14,15, virus-induced T cell exhaustion16,17, and dysfunctional regulatory T cells18,19. While ANAs can be elevated in long COVID-19 patients20,21, ANA positivity has also been shown to be substantially more prevalent than in the general population22. Furthermore, COVID-19 outpatients had an increased risk of developing inflammatory arthritis, connective tissue diseases, and intestinal-related AD in compared to inpatients, highlighting the complex relationship between autoantibodies and COVID-1923.
Type I interferon (IFN) induced innate immune responses are crucial for protective immunity as they reduce virus replication. Inborn defects in IFN pathway genes could impair critical antiviral interferon responses, increasing susceptibility to severe viral infections24,25,26,27,28,29,30,31,32,33 Individuals with severe COVID-19 often harbor defects in TLR3, IRF3, and IRF7, critical for IFN responses34,35,36,37. Consistent with inborn genetic disorders, anti-IFN autoantibodies, which block IFN function, contribute to poorer clinical outcomes. These anti-IFN autoantibodies were first detected in patients treated with IFNβ for nasopharyngeal carcinoma in the early 1980s and are prevalent in autoimmune conditions like SLE and autoimmune polyendocrine syndrome type 1 (APS-1)38,39,40. Neutralizing anti-IFN autoantibodies are linked to a subset of critically ill COVID-19 patients with low levels of IFN41,42. Anti-IFN autoantibodies are found in ~10% of patients with critical COVID-19, are linked to multi-organ failure, and compromise antiviral defenses in the nasal mucosa, enabling virus dissemination43,44,45,46,47.
Here, we detected 20 common autoantibody signatures, along with anti-IFNα autoantibodies, using convalescent serum samples from unvaccinated, SARS-CoV-2-infected patients, and examined their relationship to disease severity. We further explored the correlation between circulating autoantibodies in the context of SARS-CoV-2 infection and identified a subset of anti-ENAs associated with hospitalization. Finally, we establish a new association between autoantibodies and the humoral immune response targeting the spike region of SARS-CoV-2 variants, mediating its neutralization.
Results
Study population
The clinical summary is outlined in Table 1. The study cohort included 38 unvaccinated individuals infected with SARS-CoV-2. Serum samples were collected during the first and third Canadian pandemic waves in 2020 and 2021. Of the 38 patients, 63% (24/38) were managed as outpatients, while 37% (14/38) required hospitalization (Fig. 1a). The median age was 56 and 66.5 years for non-hospitalized and hospitalized groups, respectively, with a higher proportion of females in the non-hospitalized group. Serum samples were collected at a median of 37 and 82 days following the first positive COVID-19 test for non-hospitalized and hospitalized groups, respectively. For the hospitalized group, the median time from onset of symptoms to hospital admission was 6 days. Common symptoms included fever, cough, and shortness of breath, with shortness of breath being more prevalent in hospitalized group. The hospitalized group had a higher prevalence of co-morbidities, including cardiac, vascular, pulmonary, and renal illnesses, as well as cancer and mental health conditions. The ICU group had four patients, two of whom required intubation.
a Overview of cohort used in this study. b Fold increase in the levels of 20 clinically relevant autoantibodies and total IgG levels. Anti-ENA (black), anti-ANCA (red), anti-phospholipid (blue), anti-aminoacyl tRNA synthetase (green), anti-complement (brown), and total IgG (purple). c Clustered heatmap showing the clustering of autoantibodies. Clusters are horizontally separated using dashed lines. d Principal component analysis in ICU, hospitalized, and non-hospitalized groups. (red, ICU; green, hospitalized; blue, non-hospitalized) e–h Heatmap analysis on non-hospitalized, hospitalized, non-ICU hospitalized, and ICU groups.
Detection of autoimmune antibodies in convalescent sera
We examined the relationship between autoimmune antibodies and hospitalization in SARS-CoV-2 infection by measuring autoantibody levels against 20 self-antigens associated with various clinical phenotypes, utilizing a magnetic bead-based multiplex approach. Of these, 75% (15 of 20) were ANAs targeting various nuclear components, including DNA and RNA. All ANAs targeted ENAs, a subgroup comprising non-chromatin nuclear proteins. The panel also included two ANCAs, one anti-aminoacyl tRNA synthetase (ARS) antibody, one antiphospholipid antibody, and one C1q antibody targeting the complement C1 complex. Sham control beads without any conjugated antigen were used for background fluorescence intensities. We observed a substantial variation in the prevalence of autoantibodies in our cohort with a 2.8- to 465.7-fold increase in autoantibody levels compared to sham-control beads (Fig. 1b). To determine whether variations in autoantibody levels were related to differences in IgG, we measured the total polyclonal IgG concentration (Fig. 1b). The mean IgG level was 1337 mg/dL (range, 1279–1397). There was no significant association between IgG levels and autoantibodies, except for CENP-B (p = 0.0357), suggesting that the variation in autoantibodies is largely independent of total IgG levels.
Correlation matrix and clustering of autoantibodies
We then evaluated the association between each of the 20 autoantibodies and their clustering patterns. Correlation matrix grouped the autoantibodies into four clusters (Fig. 1c). The first cluster (from the top) included five ANAs targeting various nuclear components (Ku, CENP-A, RNP/Sm, PCNA, and Scl-70) and an anti-ARS antibody, PL-12. Each of the six autoantibodies in this cluster strongly correlated with each other—intracluster correlation showed meaningful grouping of autoantibodies, providing further insight into those autoantibodies that are frequently co-expressed in this cohort. The second cluster contained two ANAs, Sm and PM/Scl-100, which significantly correlated with each other. The third cluster contained four ANAs (Ribosomal P, SSA/Ro52, RNP, and Jo-1) and an anti-phospholipid antibody, β2-Glycoprotein, all of which significantly correlated with each other. The final cluster had seven autoantibodies: four ANAs (SSA/Ro60, SSB/La, Mi-2, and CENP-B), two ACNAs (Myeloperoxidase and Proteinase 3), and one anti-C1q antibody targeting the complement C1 component. The intra-cluster correlation was weakest for this cluster. Overall, the intracluster correlation analysis provided additional insight into patterns of autoantibody co-expression.
In addition to the intra-cluster correlation, we observed significant inter-cluster correlations (Fig. 1c). While intra-cluster correlation helped identify autoantibodies that are closely related within specific groups, inter-cluster correlation allowed us to explore how autoantibodies in different clusters are connected. Autoantibodies from cluster 1 were strongly associated (p < 0.001) with those in cluster 2 and weakly with cluster 4; however, no association was found between cluster 1 and cluster 3. The two ANAs in cluster 2 significantly correlated with the four ANAs in cluster 3, while eliciting a weak correlation with cluster 4. Finally, there was significant association between autoantibodies in cluster 3 and cluster 4. Our clustering data reveal that the presence of one autoantibody correlates with the detection of multiple other autoantibodies, suggesting a complex and heterogeneous autoimmune response in this cohort.
Association of autoantibodies with COVID-19 progression
Next, we performed principal component analysis (PCA) to reduce the dimensionality of the dataset and identify autoantibodies that are associated with COVID-19 hospitalization (Fig. 1d). Dimension 1 (PC1) 1 and PC2 accounted for 40.2% and 21.2% of the total variance, respectively. However, PCA did not reveal any distinct separation between non-hospitalized, hospitalized, and ICU groups, indicating that autoantibodies by themselves are not associated with hospitalization in this cohort.
Next, we focused on identifying signature patterns associated with disease progression using heatmap analysis (Fig. 1e–h). This enabled visualization of fold-changes in autoantibody levels across samples and identification of clustering patterns that might mediate hospitalization. Heatmap analyses were performed on four groups: non-hospitalized, hospitalized, non-ICU hospitalized, and ICU. While the non-hospitalized and non-ICU hospitalized groups showed interspersed clustering, there was a distinct separation of autoantibody clusters in the hospitalized and ICU groups. Notably, three ENAs from the third cluster (Fig. 1c), SSA/Ro52, Jo-1, and RNP, clustered in the hospitalized and ICU groups (Fig. 1f, h). We noticed 3.8-, 3.5- and 1.9-fold increases in SSA/Ro52, Jo-1 and RNP levels, respectively, in hospitalized versus non-hospitalized groups (Table 2), suggesting that these antigens could be targets in COVID-19-associated hospitalized patients and may be associated to ICU admission. Next, we investigated whether the systemic autoantibody response was associated with hospitalization (Fig. 2). The mean fold increase in autoantibody levels compared to sham-conjugated controls for the hospitalized and non-hospitalized groups is shown for each of the 20 autoantibodies (Table 2). There was no significant difference in the fold-change of autoantibody levels while comparing the non-hospitalized group to the hospitalized group. However, the collective presence of SSA/Ro52, Jo-1, and RNP autoantibodies predicted hospitalization status with 62.5% accuracy using a neural network model (Fig. 2b). These findings indicate that this subset of autoantibodies could be synergistically associated with hospitalization, and future studies are needed to validate our findings.
a Mean autoantibody levels for each of the 20 autoantibodies between hospitalized and non-hospitalized groups. Statistical significance calculated using unpaired t-test (ns - not significant). b SSA/Ro52, Jo-1, and RNP autoantibodies collectively predicted hospitalization status in a neural network model.
Detection of binding and neutralizing anti-IFNα antibodies and their association with hospitalization
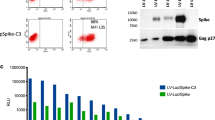
Next, we investigated the role of circulating anti-IFNα autoantibodies on COVID-19-related hospitalization. A549 cells were pre-treated with serial ten-fold dilutions of IFNα and assayed for VSV-GFP replication to determine the minimal concentration required to maintain an antiviral state (Fig. 3a). IFNα at concentrations of 0.01 ng/mL or higher resulted in a partial to complete antiviral state, with an IC50 of 0.0125 ng/mL (95% CI of 0.01009 to 0.01580; Fig. 3b). We next incubated A549 cells with 0.1 and 1 ng/mL IFNα in the presence of serially diluted control anti-human mouse IgG1 monoclonal antibody (MMHA-2) that binds and neutralizes human IFNα followed by VSV-GFP infection (Fig. 3c). Treatment of A549 cells with IFNα in the absence of MMHA-2 reduced VSV-GFP infection compared to untreated control (Fig. 3c lower panel). Incubation of MMHA-2 with IFNα resulted in a concentration-dependent increase in VSV-GFP infection for both IFNα treatments (Fig. 3c upper and middle panels), with an IC50 of 16.6 and 119.7 ng/mL of MMHA-2 for neutralizing 0.1 and 1.0 ng/mL of IFNα, respectively (Fig. 3d). We next evaluated if IFNα autoantibodies in serum samples could dampen IFNα-induced antiviral activity, with ID50 values, referred to as a “neutralization score”, derived for each serum sample. Two patient samples had a substantial neutralization score with autoantibodies functionally neutralizing IFNα. We further detected IFNα binding antibodies using ELISA in 11% (4/38) of the cohort. Of these, three were in the non-hospitalized category, while one was admitted to the ICU. We did not notice an association between the presence of anti-IFNα neutralizing or binding antibodies with hospitalization status (Fig. 3e–G). Corroborating these findings, PCA analysis showed no distinct clustering between serum samples with and without anti-IFNα binding autoantibodies (Fig. 3h). Our data provide evidence that a subset of SARS-CoV-2 patients have binding IFNα autoantibodies that are also capable of restricting an IFNα-induced antiviral state.
a Activity of IFNα on VSV-GFP replication in human A549 lung epithelial cells. b IC50 determination of IFNα activity from (a). c Anti-IFNα neutralization assay using a control anti-IFNα monoclonal neutralizing antibody (nAb), MMHA-2. d IC50 determination of neutralization activity from (c). e, f Association of anti-IFNα neutralizing antibodies to COVID-19 hospitalization. g Association of anti-IFNα binding antibodies to COVID-19 hospitalization. h Principal component analysis in anti-IFNα binding antibody positive and negative groups. (red, anti-IFNα pos; blue, anti-IFNα neg) i Heatmap analysis showing association between 20 autoantibodies and anti-IFNα binding antibody. Statistical significance was calculated using an unpaired t-test (e–g) or Pearson coefficient (i).
Anti-IFNα binding autoantibodies are strongly associated with ENA autoantibodies
Next, we examined associations between binding anti-IFNα autoantibodies and the other 20 autoantibodies (Fig. 3i). Notably, a positive correlation was noticed between binding anti-IFNα autoantibodies and seven other autoantibodies. Among them, four (SSA/Ro52, RNP, PM/Scl-100, Jo-1) showed a strong correlation, indicating that the presence of anti-IFNα autoantibodies is associated with a higher frequency of other autoantibodies.
Circulating autoantibodies are associated with SARS-CoV-2 neutralization
Last, we tested whether autoantibodies were associated with an increased humoral immune response against SARS-CoV-2. Five live SARS-CoV-2 isolates were used to determine neutralization activity: SB3 (ancestral), B.1.351 (beta), R.1 645 (R.1), B.1.617.2 (delta), and BA.5 (omicron)48. SB3, R.1 645 and B.1.617.2 were neutralized by 63% (24/38), 76% (29/38) and 61% (23/38) of samples respectively at an ID50 > 25 (Fig. 4a). In contrast, B.1.351 and BA.5 were neutralized by only 16% (6/38) and 13% (5/38) of samples respectively (Fig. 4a). B.1.351 was significantly more resistant to neutralization than SB3 (p = 0.0012), R.1 645 (p < 0.0001), and B.1.617.2 (p = 0.0199). Similarly, BA.5 was significantly more resistant to neutralization than SB3 (p = 0.0006), R.1 645 (p < 0.0001), and B.1.617.2 (p = 0.0181).
a Sensitivity of SB3, B.1.351, R.1 645, B.1.617.2 and BA.5 variants to neutralizing antibodies (brown, SB3; green, B.1.351; blue, R.1 645; red, B.1.617.2; purple, BA.5). Statistical significance was calculated using an unpaired t-test (*p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001). b Circa plots to visualize association of the level of autoantibodies to SARS-CoV-2 neutralization (brown, SB3; green, B.1.351; blue, R.1 645; red, B.1.617.2; purple, BA.5). Statistical significance was calculated using Pearson coefficient (*p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001) c–g Principal component analysis in neutralizer and non-neutralizer groups (red, neutralizer; blue, non-neutralizer).
To explore potential association between autoantibody levels and neutralization capacity, a correlation analysis was performed (Fig. 4b). Strikingly, 12 of the 20 autoantibodies were significantly correlated with B.1.351 neutralization, suggesting that autoantibodies may enhance the neutralization activity against resistant SARS-CoV-2 isolates. Six ENAs (PCNA, Scl-70, RNP/Sm, CENP-A, Sm, and Ku), along with an aminoacyl-tRNA synthetase (PL-12), showed the strongest correlation (p < 0.0001), suggesting that targeting of specific autoantigens is associated with significantly greater B.1.351 neutralization. Similarly, 10 autoantibodies were significantly correlated with B.1.617.2 neutralization, with RNP/Sm, PCNA, Scl-70, and PL-12 showing the strongest correlations (p < 0.0001). Interestingly, the four autoantibodies strongly associated with both B.1.351 and B.1.617.2 neutralization were grouped in cluster 1 (Fig. 1c). Only two autoantibodies, proteinase 3 and CENP-B, were associated with BA.5 neutralization. There was no association observed for the other two sensitive SARS-CoV-2 isolates, SB3 and R.1 645. To further explore the relationship, we conducted PCA by categorizing samples as “neutralizers” (ID50 > 25) or “non-neutralizers” (ID50 < 25). There was no clustering between the two groups, indicating that autoantibodies collectively do not influence SARS-CoV-2 neutralization (Fig. 4c–g). These findings imply that autoantibodies targeting RNP/Sm, PCNA, Scl-70, and PL-12 correlates with SARS-CoV-2 neutralization activity.
Discussion
We detected levels of 20 clinically relevant, circulating autoantibodies frequently found in ADs using serum samples from unvaccinated, SARS-CoV-2-infected, recovered individuals and determined if autoreactivity was associated with severe COVID-19 using hospitalization as the primary criterion (Figs. 1 and 2). Although we observed the presence of a diverse repertoire of autoantibodies in acute SARS-CoV-2, there was no association between individual autoantibody frequency and hospitalization. However, the presence of three autoantibodies targeting SSA/Ro52, Jo-1, and RNP was collectively associated with hospitalization (Fig. 2). Our data indicate that the disruption of immunological tolerance is associated with hospitalization in SARS-CoV-2 acute infections and highlights the need for prospective studies to further investigate the association between infectious diseases and autoimmunity2,4,6. Furthermore, 11% and 5% of our cohort had binding and neutralizing anti-IFNα autoantibodies, respectively, with anti-IFNα antibodies significantly associated with six ENAs and β2-glycoprotein (Fig. 3). Finally, the presence of a subset of autoantibodies was found to be associated with enhanced neutralization of B.1.351, B.1.617.2, and BA.5 (Fig. 4).
Transcriptomic and proteomic analyses from our group and others have highlighted the enrichment of antiviral responses and type I IFN signaling pathways leading to the expression of ISGs during SARS-CoV-2 infection48,49,50. Here, we demonstrate the presence of anti-IFNα neutralizing antibodies in acute SARS-CoV-2 infection. The prevalence of these antibodies is slightly higher than the 1.5% observed in a blood bank study51 and lower than the 10% observed in individuals with life-threatening COVID-19 pneumonia, of whom 94% were men43. While systemic anti-IFN autoantibodies have been linked to higher viral loads and severe COVID-19, they serve to reduce excessive inflammation in APS-1 patients, leading to milder COVID-19 symptoms43,44,52. The homeostatic function of anti-IFNα neutralizing autoantibodies was demonstrated in the nasal mucosa of SARS-CoV-2 infected patients, where these autoantibodies appeared within weeks of infection, reduced excessive nasal IFNα secretion and inflammation, thereby contributing to an efficient recovery53. Our data show that neutralizing anti-IFNα antibodies are not driving hospitalization and may, in turn, be balancing the IFN activity to prevent excessive inflammation.
Finally, we establish a correlation between SARS-CoV-2 neutralization and autoantibody response (Fig. 4). Although previous studies show an association between autoantibodies and anti-SARS-CoV-2 IgG targeting the receptor-binding domain (RBD), non-structural protein 1 (NSP1), and nucleocapsid protein8,54, it remained unclear if autoantibodies in COVID-19 patients associate with SARS-CoV-2 neutralization. Of the 20 autoantibodies examined, 12 were significantly associated with the neutralization of B.1.351, and 10 were significantly associated with B.1.617.2 neutralization, suggesting that the autoimmune response might modulate the anti-viral humoral immune response. A higher frequency of autoantibodies correlates with the ability to neutralize multiple human immunodeficiency virus-1 (HIV-1) subtypes, thereby broadening the neutralization spectrum in some HIV-1-infected individuals55,56. Similarly, anti-influenza antibodies from SLE patients have a higher avidity and neutralization capacity57. The development of neutralizing antibodies may be controlled by immune tolerance mechanisms, as observed in HIV-1 and influenza virus infections, where antibodies develop long complementarity-determining region 3 (CDR3) loops and undergo extensive somatic hypermutation58,59. While factors such as infecting strain, timing of infection, viral load, and disease severity can influence the development of SARS-CoV-2 neutralization response60,61,62, future studies are needed to clarify our observed relationship between autoantibodies and neutralization.
SARS-CoV-2 infections are associated with the development of new-onset IgG autoantibodies targeting numerous self-protein targets across a wide range of tissues8,9,11. Autoimmunity is a key feature of post-acute sequelae of COVID-19 (PASC), observed in approximately 10% of SARS-CoV-2 infections with persistent symptoms over 3 months after the initial infection63. Polyautoimmunity occurs in 62% of patients with PASC, characterized by the presence of ANAs detected over 12 months post-COVID-19 and correlating with neurological disorder intensity20,54,64. Although this observation suggests that autoantibodies could serve as biomarkers of PASC, limited evidence supports that targeting specific autoantibodies could reverse PASC. Interestingly, health records suggest that vaccination decreases the likelihood of developing AD post SARS-CoV-2 infection23,65. The autoantibody levels were found to be stable in vaccinated individuals compared to acute COVID-19 patients; however, increased anti-SNURF antibodies have been reported following vaccination66,67. Finally, it is noteworthy that the immunological mechanisms that drive virus-induced ADs are not exclusive to SARS-CoV-2; as infections with human cytomegalovirus, influenza virus, Epstein-Barr virus, and chikungunya virus have also been linked to autoimmunity68,69,70,71.
A limitation of our study is the absence of longitudinal samples that would have been valuable to explore whether autoantibody levels were transient. Additionally, we did not have a direct comparison between our SARS-CoV-2 infection to an uninfected control group. However, seroprevalence data suggest that 76% of the Canadian population had infection-acquired antibodies from prior SARS-CoV-2 infections72, making the establishment of a clear control group for comparison difficult. Some of our interpretations on hospitalization may be influenced by confounding factors—higher age, higher proportion of males, higher prevalence of vascular illnesses, and differences in sampling time. Another limitation is that we focused exclusively on the IFNα2 subtype. Finally, limited serum sample availability precluded us from performing an IgG depletion in IFN-neutralizing patients.
In conclusion, our findings suggest that the presence of autoantibodies in an acute setting may not independently lead to a pathological clinical phenotype; however, the presence of multiple, distinct autoantibodies could contribute to severe COVID-19. Identifying specific autoantibodies associated with severe COVID-19 provides valuable insights into biomarkers, and prospective studies are needed to explore this further. Notably, we observed that the generation of anti-IFNα antibodies is correlated with the presence of several other autoantibodies, indicating a common feature governing the disruption of immune tolerance mechanism. Finally, our data showing a correlation between autoantibodies and SARS-CoV-2 neutralization in COVID-19 infections emphasize the need of further research to explore cross-reactivity of SARS-CoV-2 proteins with other autoepitopes and to determine if SARS-CoV-2-infected patients with AD develop a broader neutralizing antibody response.
Methods
Human donors
Informed consent was obtained for the collection of convalescent serum from 38 patients with laboratory-confirmed SARS-CoV-2 infection. This study was approved by Sunnybrook Research Institute (REB#2218) and Sinai Health System (REB# 02-0118-U and 05-0016-C) ethics boards.
Cells and viruses
Human A549 lung and monkey Vero E6 epithelial cells were cultured as described previously48. Vesicular stomatitis virus expressing green fluorescent protein (VSV-GFP) was used for IFNα neutralization assays at MOI 1.0. SB3, an ancestral SARS-CoV-2 variant, and R.1 645, a variant under monitoring (VuM), were isolated and purified as described in refs. 48,73. The B.1.351 (beta), B.1.617.2 (delta), and BA.5 (omicron) were obtained from BEI Resources (Manassas, VA, United States). Experiments with SARS-CoV-2 were performed in a biosafety containment level 3 facility as approved by the institutional biosafety committee at McMaster University.
Detection of autoantibodies
Autoantibodies were detected using a multiplex panel (MilliporeSigma, Catalog # HAIAB-10K) following manufacturer’s guidelines. Briefly, to measure autoantibodies, beads were sonicated, vortexed, and thoroughly mixed. Assay plates were washed with 200 µL of wash buffer for 10 min. After decanting the buffer, 25 μL of assay buffer was added to each well, followed by 25 μL of serum samples diluted 1:100 in assay buffer. Subsequently, 25 μL of the mixed bead suspension was added to each well. Plates were incubated overnight at 4 °C on a plate shaker. The following day, plates were washed three times with wash buffer. Then, 50 μL of PE-conjugated IgG detection reagent was added to each well, and the plates were incubated at room temperature for 90 min. After incubation, plates were washed three times with wash buffer and the beads were resuspended in 150 μL of drive fluid. The plate was shaken for 5 min to ensure uniform bead suspension. Fluorescence from labeled magnetic beads was measured with a MAGPIX instrument and data analyzed using xPONENT software. The specificity of the 20-plex panel ranged from 92% to 100% (manufacturer’s data), indicating minimal cross-reactivity. Sham control beads were used to measure background fluorescence intensities. These negative control beads (without any antigen) went through the same process of conjugation as the panel beads. Sham control beads exhibited low background fluorescence, with a median fluorescence intensity (MFI) of 36.6 (interquartile range: 23.7–82.0). Fold increases in MFI were calculated by dividing the MFI of each autoantibody with the MFI of sham-conjugated controls.
Interferon treatment
A549 cells were left untreated or treated with ten-fold serially diluted IFNα (Catalog # I4276, Sigma-Aldrich). Cells were pre-treated for 6 h before infection with VSV-GFP at MOI 1.0. GFP was measured 24 h post-infection to determine initiation of virus replication.
Anti-IFNα neutralization assay
The anti-IFNα neutralizing monoclonal antibody MMHA-2 (Invitrogen, Catalog # 211002) or convalescent serum samples were serially diluted and incubated with 0.1 and 1.0 ng/mL IFNα (Catalog # I4276, Sigma-Aldrich) for 2 h at 37 °C. A549 cells were pre-treated with the antibody:IFNα mixture for 6 h before infection with VSV-GFP at MOI 1.0. GFP was measured 24 h post-infection to determine initiation of virus replication.
Enzyme-linked immunosorbent assay (ELISA)
Anti-IFNα binding antibodies (catalog # 5018034, Invitrogen) and total IgG antibodies (catalog # BMS2091, Invitrogen) were determined using ELISA following manufacturer’s guidelines.
SARS-CoV-2 neutralization assay
Serially diluted serum samples were incubated with SARS-CoV-2 (150 PFU/well) at 37 °C for 1 h before adding to pre-plated Vero E6 cells. Five days post-infection, luminescence was quantified with CellTiter-Glo 2.0 Reagent (catalog # G9243, Promega) using a BioTek Synergy H1 microplate reader. The luminescent signal, which was proportional to the amount of ATP present, was measured in Relative Light Units (RLU).
Bioinformatics and statistical analysis
Statistical power analysis was carried out in R using “pwr” package. This study is sufficiently powered, with most analyses having a power greater than 0.8 and a few marginal ones above 0.7. Correlation was executed in R using “rcorr” function from the “Hmisc” package. Heatmaps were generated using “pheatmap” package. “symnum,” an in-built function in R, was used to replace correlation coefficients with symbols based on the degree of relation. Principal component analysis (PCA) was executed in R using “prcomp” function from the “stats” package and visualized using “ggfortify” package. Neural network was executed in R using “neuralnet” package using 80% of the data for training and 20% for testing. Circa plots (https://omgenomics.com/circa) were used to visualize association of the level of autoantibodies with SARS-CoV-2 neutralization.
Unpaired t-test with Welch’s correction was used to calculate differences between hospitalized and non-hospitalized groups. The correlation between autoantibody levels and anti-IFNα antibody or SARS-CoV-2 neutralization was determined using the Pearson r product. Neutralization score was determined using a non-linear regression curve fit model. An unpaired t-test was used to compare the neutralization of the SARS-CoV-2 isolates. GraphPad Prism 10 was used for the above statistical tests.
Data availability
All data used are available upon reasonable request to the corresponding author.
References
Robbins, W. C., Holman, H. R., Deicher, H. & Kunkel, H. G. Complement fixation with cell nuclei and DNA in lupus erythematosus. Proc. Soc. Exp. Biol. Med. 96, 575–579 (1957).
Davidson, A. & Diamond, B. Autoimmune diseases. N. Engl. J. Med. 345, 340–350 (2001).
Mackay, I. R. & Rowley, M. J. Autoimmune epitopes: autoepitopes. Autoimmun. Rev. 3, 487–492 (2004).
Mobasheri, L. et al. SARS-CoV-2 triggering autoimmune diseases. Cytokine 154, 155873 (2022).
Smatti, M. K. et al. Viruses and autoimmunity: a review on the potential interaction and molecular mechanisms. Viruses 11 (2019).
Puel, A., Bastard, P., Bustamante, J. & Casanova, J. L. Human autoantibodies underlying infectious diseases. J. Exp. Med. 219, https://doi.org/10.1084/jem.20211387 (2022).
Abdel-Wahab, N., Talathi, S., Lopez-Olivo, M. A. & Suarez-Almazor, M. E. Risk of developing antiphospholipid antibodies following viral infection: a systematic review and meta-analysis. Lupus 27, 572–583 (2018).
Chang, S. E. et al. New-onset IgG autoantibodies in hospitalized patients with COVID-19. Nat. Commun. 12, 5417 (2021).
Zuo, Y. et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci. Transl. Med. 12, https://doi.org/10.1126/scitranslmed.abd3876 (2020).
Woodruff, M. C. et al. Dysregulated naive B cells and de novo autoreactivity in severe COVID-19. Nature 611, 139–147 (2022).
Wang, E. Y. et al. Diverse functional autoantibodies in patients with COVID-19. Nature 595, 283–288 (2021).
Hu, B., Huang, S. & Yin, L. The cytokine storm and COVID-19. J. Med Virol. 93, 250–256 (2021).
Song, P., Li, W., Xie, J., Hou, Y. & You, C. Cytokine storm induced by SARS-CoV-2. Clin. Chim. Acta 509, 280–287 (2020).
Adiguzel, Y. Molecular mimicry between SARS-CoV-2 and human proteins. Autoimmun. Rev. 20, 102791 (2021).
Arevalo-Cortes, A., Rodriguez-Pinto, D. & Aguilar-Ayala, L. Evidence for molecular mimicry between SARS-CoV-2 and human antigens: implications for autoimmunity in COVID-19. Autoimmune Dis. 2024, 8359683 (2024).
Diao, B. et al. Reduction and functional exhaustion of t cells in patients with coronavirus disease 2019 (COVID-19). Front. Immunol. 11, 827 (2020).
Kusnadi, A. et al. Severely ill COVID-19 patients display impaired exhaustion features in SARS-CoV-2-reactive CD8(+) T cells. Sci. Immunol. 6, https://doi.org/10.1126/sciimmunol.abe4782 (2021).
Goncalves-Pereira, M. H. et al. Dysfunctional phenotype of systemic and pulmonary regulatory T cells associate with lethal COVID-19 cases. Immunology 168, 684–696 (2023).
Meckiff, B. J. et al. Imbalance of regulatory and cytotoxic SARS-CoV-2-reactive CD4(+) T Cells in COVID-19. Cell 183, 1340–1353.e1316 (2020).
Son, K. et al. Circulating anti-nuclear autoantibodies in COVID-19 survivors predict long COVID symptoms. Eur. Respir J. 61, https://doi.org/10.1183/13993003.00970-2022 (2023).
Seessle, J. et al. Persistent symptoms in adult patients 1 year after coronavirus disease 2019 (COVID-19): a prospective cohort study. Clin. Infect. Dis. 74, 1191–1198 (2022).
Peluso, M. J., Thomas, I. J., Munter, S. E., Deeks, S. G. & Henrich, T. J. Lack of antinuclear antibodies in convalescent coronavirus disease 2019 patients with persistent symptoms. Clin. Infect. Dis. 74, 2083–2084 (2022).
Chang, R. et al. Risk of autoimmune diseases in patients with COVID-19: a retrospective cohort study. EClinicalMedicine 56, 101783 (2023).
Lim, H. K. et al. Severe influenza pneumonitis in children with inherited TLR3 deficiency. J. Exp. Med. 216, 2038–2056 (2019).
Bucciol, G. & Desmet, L. Leuven laboratory of inborn errors of, I., Corveleyn, A. & Meyts, I. Pathogenic P554S Variant in TLR3 in a patient with severe influenza pneumonia. J. Clin. Immunol. 42, 430–432 (2022).
Zhang, S. Y. Herpes simplex virus encephalitis of childhood: inborn errors of central nervous system cell-intrinsic immunity. Hum. Genet. 139, 911–918 (2020).
Lim, H. K. et al. TLR3 deficiency in herpes simplex encephalitis: high allelic heterogeneity and recurrence risk. Neurology 83, 1888–1897 (2014).
Guo, Y. et al. Herpes simplex virus encephalitis in a patient with complete TLR3 deficiency: TLR3 is otherwise redundant in protective immunity. J. Exp. Med. 208, 2083–2098 (2011).
Chen, J. et al. Inborn errors of TLR3- or MDA5-dependent type I IFN immunity in children with enterovirus rhombencephalitis. J. Exp. Med. 218, https://doi.org/10.1084/jem.20211349 (2021).
Lamborn, I. T. et al. Recurrent rhinovirus infections in a child with inherited MDA5 deficiency. J. Exp. Med. 214, 1949–1972 (2017).
Andersen, L. L. et al. Functional IRF3 deficiency in a patient with herpes simplex encephalitis. J. Exp. Med. 212, 1371–1379 (2015).
Hernandez, N. et al. Life-threatening influenza pneumonitis in a child with inherited IRF9 deficiency. J. Exp. Med. 215, 2567–2585 (2018).
Ciancanelli, M. J. et al. Infectious disease. Life-threatening influenza and impaired interferon amplification in human IRF7 deficiency. Science 348, 448–453 (2015).
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370, https://doi.org/10.1126/science.abd4570 (2020).
Matuozzo, D. et al. Rare predicted loss-of-function variants of type I IFN immunity genes are associated with life-threatening COVID-19. Genome Med. 15, 22 (2023).
Saheb Sharif-Askari, N. et al. Multiple inborn errors of type I IFN immunity in a 33-year-old male with a fatal case of COVID-19. Heliyon 10, e29338 (2024).
Kholaiq, H. et al. Human Genetic and immunological determinants of SARS-CoV-2 infection and multisystem inflammatory syndrome in children. Clin. Exp. Immunol. https://doi.org/10.1093/cei/uxae062 (2024).
Panem, S., Check, I. J., Henriksen, D. & Vilcek, J. Antibodies to alpha-interferon in a patient with systemic lupus erythematosus. J. Immunol. 129, 1–3 (1982).
Levin, M. Anti-interferon auto-antibodies in autoimmune polyendocrinopathy syndrome type 1. PLoS Med. 3, e292 (2006).
Vallbracht, A., Treuner, J., Flehmig, B., Joester, K. E. & Niethammer, D. Interferon-neutralizing antibodies in a patient treated with human fibroblast interferon. Nature 289, 496–497 (1981).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Trouillet-Assant, S. et al. Type I IFN immunoprofiling in COVID-19 patients. J. Allergy Clin. Immunol. 146, 206–208 e202 (2020).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Bastard, P. et al. Autoantibodies neutralizing type I IFNs are present in ~4% of uninfected individuals over 70 years old and account for ~20% of COVID-19 deaths. Sci. Immunol. 6, https://doi.org/10.1126/sciimmunol.abl4340 (2021).
Koning, R. et al. Autoantibodies against type I interferons are associated with multi-organ failure in COVID-19 patients. Intens. Care Med. 47, 704–706 (2021).
Goncalves, D. et al. Antibodies against type I interferon: detection and association with severe clinical outcome in COVID-19 patients. Clin. Transl. Immunol. 10, e1327 (2021).
Lopez, J. et al. Early nasal type I IFN immunity against SARS-CoV-2 is compromised in patients with autoantibodies against type I IFNs. J. Exp. Med. 218, https://doi.org/10.1084/jem.20211211 (2021).
Jacob, R. A. et al. Sensitivity to neutralizing antibodies and resistance to type I interferons in SARS-CoV-2 R.1 lineage variants, Canada. Emerg. Infect. Dis. 29, 1386–1396 (2023).
Banerjee, A. et al. Experimental and natural evidence of SARS-CoV-2-infection-induced activation of type I interferon responses. iScience 24, 102477 (2021).
Hatton, C. F. et al. Delayed induction of type I and III interferons mediates nasal epithelial cell permissiveness to SARS-CoV-2. Nat. Commun. 12, 7092 (2021).
Vazquez, S. E. et al. Neutralizing autoantibodies to type I interferons in COVID-19 convalescent donor plasma. J. Clin. Immunol. 41, 1169–1171 (2021).
Meisel, C. et al. Mild COVID-19 despite autoantibodies against type I IFNs in autoimmune polyendocrine syndrome type 1. J. Clin. Investig. 131, https://doi.org/10.1172/JCI150867 (2021).
Babcock, B. R. et al. Transient anti-interferon autoantibodies in the airways are associated with recovery from COVID-19. Sci. Transl. Med. 16, eadq1789 (2024).
Rojas, M. et al. Autoimmunity is a hallmark of post-COVID syndrome. J. Transl. Med. 20, 129 (2022).
Borrow, P. & Moody, M. A. Immunologic characteristics of HIV-infected individuals who make broadly neutralizing antibodies. Immunol. Rev. 275, 62–78 (2017).
Haynes, B. F. et al. Strategies for HIV-1 vaccines that induce broadly neutralizing antibodies. Nat. Rev. Immunol. 23, 142–158 (2023).
Kaur, K. et al. High affinity antibodies against influenza characterize the plasmablast response in SLE patients after vaccination. PLoS ONE 10, e0125618 (2015).
Krammer, F. The human antibody response to influenza A virus infection and vaccination. Nat. Rev. Immunol. 19, 383–397 (2019).
Landais, E. & Moore, P. L. Development of broadly neutralizing antibodies in HIV-1 infected elite neutralizers. Retrovirology 15, 61 (2018).
Qi, H., Liu, B., Wang, X. & Zhang, L. The humoral response and antibodies against SARS-CoV-2 infection. Nat. Immunol. 23, 1008–1020 (2022).
Trinite, B. et al. SARS-CoV-2 infection elicits a rapid neutralizing antibody response that correlates with disease severity. Sci. Rep. 11, 2608 (2021).
Peluso, M. J. et al. SARS-CoV-2 antibody magnitude and detectability are driven by disease severity, timing, and assay. Sci. Adv. 7, eabh3409 (2021).
Altmann, D. M., Whettlock, E. M., Liu, S., Arachchillage, D. J. & Boyton, R. J. The immunology of long COVID. Nat. Rev. Immunol. 23, 618–634 (2023).
Seibert, F. S. et al. Severity of neurological Long-COVID symptoms correlates with increased level of autoantibodies targeting vasoregulatory and autonomic nervous system receptors. Autoimmun. Rev. 22, 103445 (2023).
Tesch, F. et al. Incident autoimmune diseases in association with SARS-CoV-2 infection: a matched cohort study. Clin. Rheumatol. 42, 2905–2914 (2023).
Jernbom, A. F. et al. Prevalent and persistent new-onset autoantibodies in mild to severe COVID-19. Nat. Commun. 15, 8941 (2024).
Jaycox, J. R. et al. SARS-CoV-2 mRNA vaccines decouple anti-viral immunity from humoral autoimmunity. Nat. Commun. 14, 1299 (2023).
Maek, A. N. W. & Silachamroon, U. Presence of autoimmune antibody in chikungunya infection. Case Rep. Med. 2009, 840183 (2009).
Loza-Tulimowska, M., Semkow, R., Michalak, T. & Nowoslawski, A. Autoantibodies in sera of influenza patients. Acta Virol. 20, 202–207 (1976).
Houen, G. & Trier, N. H. Epstein-barr virus and systemic autoimmune diseases. Front. Immunol. 11, 587380 (2020).
Gugliesi, F. et al. Human cytomegalovirus and autoimmune diseases: Where are we? Viruses 13, https://doi.org/10.3390/v13020260 (2021).
Murphy, T. J. et al. The evolution of SARS-CoV-2 seroprevalence in Canada: a time-series study, 2020-2023. CMAJ 195, E1030–E1037 (2023).
Banerjee, A. et al. Isolation, sequence, infectivity, and replication kinetics of severe acute respiratory syndrome coronavirus 2. Emerg. Infect. Dis. 26, 2054–2063 (2020).
Acknowledgements
This work was supported by a COVID-19 response grant to KM from the Canadian Institutes of Health Research (CIHR #OV4-170645). MSM is supported by a CIHR COVID-19 rapid response grant, a CIHR new investigator award, and an Ontario early researcher award. Convalescent serum sample collection was supported by a COVID-19 response grant to AM and SM from the CIHR (#439999).
Author information
Authors and Affiliations
Contributions
R.A.J. and K.M. conceptualized and designed the study. R.A.J., A.S., A.J.M. and S.M. collected the data. M.R.D., A.B., M.S.M., A.J.M. and S.M. contributed resources. R.A.J. and H.O.A. analyzed and interpreted the data. R.A.J., H.O.A., and K.M. drafted the manuscript. All authors revised and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jacob, R.A., Ajoge, H.O., D’Agostino, M.R. et al. Distinct circulating autoantibodies are associated with COVID-19 hospitalization and SARS-CoV-2 neutralization activity. npj Viruses 3, 64 (2025). https://doi.org/10.1038/s44298-025-00149-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44298-025-00149-2