Abstract
Mutations in mitochondrial-related genes underlie numerous neurodegenerative diseases, yet the significance of most variants remains uncertain concerning disease phenotypes. Several thousand genes have been shown to regulate mitochondria in eukaryotic cells, but which of these genes are necessary for proper mitochondrial function and dynamics? We investigated the degree of morphological disruptions in mitochondrial gene-silenced cells to understand the genetic contribution to the expected mitochondrial phenotype and to identify potentially pathogenic variants like pathogenic mutations in MFN2. We analyzed 5835 gRNAs in a high dimensional phenotypic dataset produced by the image-based pooled analysis platform Raft-Seq. Using the MFN2-mutant cell phenotype, we identified several genes, including TMEM11, TIMM8A, NDUFAF4, NDUFAF7, and NDUFS5 (NADH ubiquinone oxidoreductase-related genes), as crucial for normal mitochondrial dynamics in human U2OS cells. Additionally, we found several missense and UTR variants within the genes SLC25A19 and ATAD3A as drivers of mitochondrial aggregation. By examining multiple features instead of a single readout, this analysis was powered to detect genes which had morphological ‘signatures’ aligned with MFN2-mutant phenotypes. Reanalysis with anomaly detection revealed other critical genes, including APOOL, MCEE, NIT, PHB, and SLC16A7, which perturb mitochondrial network morphology in a manner divergent from MFN2. These studies show causal links between gene knockouts and gene-specific variants into the assembly or maintenance of mitochondrial dynamics and can hopefully lead to a better understanding of mitochondrial related diseases.
Similar content being viewed by others
Introduction
Mitochondria are complex organelles that play a crucial role in energy production and various cellular processes including cellular growth, differentiation, and cell death1,2,3. Neurons are particularly sensitive to mitochondrial disruptions, due to their elongated, complex shape and metabolic burden4. The processes of fission, fusion, and mitophagy induce morphological changes in these organelles, broadly defined as mitochondrial dynamics5,6,7,8. When these dynamics are disrupted, mitochondrial morphology can change, as seen in pathogenic mutations of the gene Mitofusin2 (MFN2) in Charcot-Marie-Tooth Disease Type 2 A (CMT2A) which often produce aggregated mitochondrial morphology9,10,11,12,13. Although numerous studies have been conducted in the realm of mitochondrial dynamics in relation to human disease, it is difficult to establish a connection between organellar phenotype and genotype14,15. Several thousand genes have been shown to regulate mitochondria in eukaryotic cells, but it is unknown which are required for proper mitochondrial dynamics. Also, the degree of cellular disruption when these genes are knocked out is unknown. Recognizing the necessity and magnitude of genetic contribution to properly functioning mitochondria could help define pathogenic variants that cause human disease.
While the well-studied MFN2 gene is primarily recognized for its role in human disease and disruptions in mitochondrial morphology, solute carrier family 25 member 19 (SLC25A19) and ATPase family AAA domain containing 3 A (ATAD3A) are two genes also associated with human disease. Although less is known regarding their impact on mitochondrial dynamics, both genes code for proteins located in the inner mitochondrial membrane. SLC25A19 is a thiamine pyrophosphate carrier and mutations that disrupt this gene, and related transport pathways, are linked to several conditions such as microcephaly, central nervous system malformations, anemias, neuropathy, aciduria, carnitine deficiency, HHH syndrome, early infantile epileptic encephalopathy type 3, aspartate/glutamate deficiency, Fontaine progeroid syndrome, and citrullinemia type II16,17,18. ATAD3A encodes a mitochondrial inner membrane protein essential for mitochondrial dynamics, nucleoid organization, protein translation, and cholesterol metabolism19,20,21,22. Both genes are expected to have a different impact on mitochondria than MFN2. Understanding the impact of SLC25A19 and ATAD3A at the variant level could provide mechanistic insights and set the stage for more targeted therapeutics given their relevance in human disease.
Still, MFN2, SLC25A19, and ATAD3A are only three of hundreds of genes that have been identified to have mitochondrial implications in disease. A renewed effort in functional genomics to enhance understanding of the intricate biology behind sequencing data has led to the development of genetic CRISPR screens, including stand-alone and combined formats like survival, FACS-based, arrayed, and single-cell, for greater insight23. Image-based screening techniques linking phenotype to genotype by microscopy allow the interrogation of a comprehensive array of perturbations in live cell populations24,25,26, as do screening techniques designed to investigate the morphologies of specific organelles and their links to human disease13,27. Deep cellular phenotyping techniques have also emerged exploring molecular characterization, cellular morphology, functional single cell analysis, and even cellular interactions28,29,30,31,32,33. Such techniques signify a critical shift beyond cell survival and toward a deeper exploration of organelle structure and function34.
Here, we utilize the existing Raft-Seq pipeline to examine the morphology and activity (depolarization) of mitochondria in single human cells. We used the U2OS osteosarcoma cell line for its mitochondrial network and extensive characterization with MFN2 variants13. By knocking out the endogenous locus for 1,410 known mitochondrial-related genes, we aim to determine those which produce a morphological signature akin to mutations in MFN2, which are known to cause neurodegenerative disease. Although many of these mitochondrial-related genes are well-studied, there continue to be surprises regarding their impact on mitochondrial dynamics35. Our findings are carefully validated in independent assays where we assess Cas9 cutting efficiency and cellular morphology across thousands of cells. Not only can we measure the relative disruption of mitochondrial morphology in different knockouts, but we can also determine this for specific indel variants along a particular gene. In this study, we leverage Raft-Seq to ascertain which mitochondrial-related genes are required for normal mitochondrial morphology.
Results
Mitochondrial phenotypes in 1410 mitochondrial-related genes using Raft-seq
Normally, mitochondria are dispersed densely around the perinuclear region and sparsely networked throughout the cytoplasm of human U2OS cells. These cells display an aggregated mitochondrial phenotype when overexpressing a dominant mutant of MFN2 (L76P or R94Q, for example), or when edited to contain an endogenous MFN2 variant13. To identify additional genes that perturb mitochondrial morphology and possibly cause neurodegenerative disorders if mutated, we screened 5,835 gRNAs for 1,410 mitochondrial-related genes (Table S1) using Raft-Seq, a pooled high content imaging pipeline (Fig. 1a). These guides were organized into several pools, packaged into lentivirus, and delivered into U2OS doxycycline-inducible (Dox) Cas9 cells at a low multiplicity-of-infection (MOI, 0.1) to ensure one gRNA per cell before raft plating. In the primary screen, 77 raft plates were seeded, stained, and imaged. Cells were stained to both visualize the extent of all mitochondria (using MitoTracker) and their polarization state (using TMRM). Single-cell data points were acquired then filtered for debris, health, and focus (Fig. S1), resulting in 455,006 healthy and viable cells, each assessed for its mitochondrial signature.
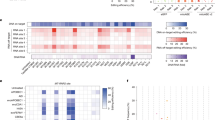
a Overview of pooled gRNA screening process using Raft-Seq. Mitochondrial-related gRNAs were packaged into lentivirus and infected into U2OS cells with Dox-inducible Cas9. Cells were imaged and analyzed on a subset of phenotypic mitochondrial features before being individually selected for isolation and genotyping. b Fluorescent 20x confocal images of U2OS cells on microrafts of cells from no Cas9 (no Dox, the WT-like control), MFN2R94Q (mutant-like control), Non-Target gRNA (WT-like), and spiked in MFN2V459fs/WT mutants (two examples each). Only the first two (blue line) were ‘labeled’, and used to train the multi-feature classifications, whereas the cells under the purple line were isolated from the mixed screening population and their gRNAs identified through Raft-Seq. The edge of a single raft is demarcated by dotted lines. The cell is shown again to the right of each full-raft image with adjusted look-up-tables to visualize what is sometimes a bright perinuclear cluster of mitochondria, especially in the MFN2V459fs/WT mutant. Linear adjustments were made to the images for presentation, but only full color-depth images were used in the analysis. c Line plot showing the six different multi-feature models comprising Mito Path ScoreR, used to distinguish MFN2 mutant cells from controls. Each line represents the model score for a particular gRNA in the primary screen. Six models are shown as positions along the X-axis, and the connecting points of each line represent the score for a particular gRNA by a particular model. The lines are colored by the average of all the models (Mito Path ScoreR), with red indicating more MFN2-mutant like phenotypes, green control-like, and white intermediate. d Volcano plot listing compiled gRNA results and their position along the X-axis showing WT-like phenotypes (left) or mutant-like phenotypes (right). Blue dots indicate Non-Target gRNAs while green dots represent selections taken into upcoming experiments. Yellow dots are two gRNAs selected as anti-hits. The spiked MFN2V459fs/WT positive control is indicated with a red marker. −Log P-values of T-tests were capped at 5. Dashed line indicates p-value threshold where FDR is 0.2. e Plot showing gRNA results in relation to Nuclei anti-health (vertical axis), Mito Count (left axis), and Mito Path ScoreR (right axis). Nuclei anti-health is calculated as nuclei intensity/nuclei area. Only markers with FDR < = 0.2 are shown.
In the Raft-Seq experiments, the WT-like control consisted of U2OS cells containing the Mito gRNA library where no Cas9 cutting (no Dox) was induced. In contrast, the genetically perturbed control was comprised of an isogenic line with a mutant endogenous MFN2 (frameshift at V459). These control cell lines were placed in specific wells to serve as anchors, facilitating the distinction between healthy and perturbed in the screen (Fig. 1b). These controls were used to train ‘vanguard’ logistic regression models to distinguish the MFN2-mutant from the WT-like line. Those model scores were subsequently utilized to select Cas9-active (Dox induced) cells displaying both normal and disrupted phenotypes (Fig. S2) for Raft-based capture. In total, 29,260 cells were picked into 304 96-well plates and single-cell genotyped to identify the corresponding gRNA. This resulted in 10,730 cells that revealed a single distinct gRNA from the library. In addition, the MFN2V459fs/WT cell line was also introduced to the Dox-induced cells at 0.2% of the total cell count before being plated. Recovery of this “spiked-in” control verified that the Raft-Seq pipeline was successfully able to detect and extract cells with MFN2 mutant-like phenotypes. The result was a highly enriched spiked-in MFN2 control, achieving an AUC (Area Under the ROC Curve) of 0.848 (Fig. S3). Importantly, the gRNA identities were not known until after genotyping, and the admixture additionally minimized the impact of batch effects on the score of pathogenicity. Thus, this AUC and enrichment indicate a well-tuned assay.
We trained six multi-feature models consisting of an even mix of binary classifiers (3 Logistic Regressions and 3 Random Forests, 12 random features each, 6 to 11 of which were mitochondrial features. Table S2) on the normalized data across all plates (Fig. 1c). Models only saw wild type-like no Cas9 (no Dox) and mutant-like (MFN2V459fs/WT) labeled wells for training. Each model scored the differences between the wild type and mutant phenotypes. To prevent bias towards a particular model, we made a soft-voting ensemble called the Mito Path ScoreR (Path is short for Pathogenicity, “R” indicates raft-based, Table S3) by mean aggregating the prediction probabilities across the six models. Using this Mito Path ScoreR, we assembled the gRNAs across multiple observations over numerous raft replicates and experiment days. We compared each gRNA’s Mito Path ScoreR to the median score of all Non-Target gRNAs under a lenient false discovery rate (FDR), befitting an exploratory analysis (Fig. 1d). Unlike other types of pooled screens, Raft-Seq can link the gRNA to specific single cells (Fig. S4). Recovered MFN2V459fs/WT cells were consistently among the most significant results, and a subset of the highest scoring gRNAs were selected for further validation.
The Mito Path ScoreR reduces the dimensionality of the large set of perturbations to a single axis, specifically to detect MFN2 mutant-like morphological signatures. This low-dimensional representation can possibly mask other interesting phenotypes (expanded below in the section Underlying Details of the Multi-feature Mitochondrial Pathogenicity Phenotype in Fig. 4), as some may relate more strongly to health or another cellular state (Fig. 1e). For our downstream analyses, we focused on gRNAs which strongly perturbed the cellular phenotype towards the MFN2-mutant control while not adversely affecting nuclear health as measured by Nuclei Anti-Health (nuclei intensity/nuclei area)36. Such gRNAs result in more clustered mitochondria without detrimental effects on the overall cell health.
Knockout of TMEM11, TIMM8A, and three NADH ubiquinone oxidoreductase-related genes phenocopy MFN2 variants in U2OS cells
To validate the knockout-induced phenotypes elucidated from the screen, we selected thirty gRNAs against twenty-eight genes based on a high Mito Path ScoreR, low nuclei anti-health, and low mitochondrial count (Fig. 1e). We transfected, into U2OS cells, synthetic versions of these gRNAs (syngRNAs) (Table S4) complexed with Cas9 as ribonucleoprotein complexes. These “founder” cell populations were grown out and subsequently split for imaging and cutting analysis (Cas9/syngRNA cutting efficiency was above 95% for most of the syngRNAs Fig. S5). In analysis of cell morphologies, Non-Target, ATP5G3, and PRSS35 gRNA-nucleofected U2OS cells served as the negative controls (the latter two had WT-like results in the primary screen), and an MFN2 gRNA was used as the positive control. Rather than imaging on rafts, validation cells were arrayed in 96-well optical plates for confocal imaging (plate-based assay). Using a plate-based assay instead of a raft-based assay increased cost-effectiveness and enhanced the number of single-cell instances per condition, thereby improving analytical power. We trained another set of binary classifiers to discriminate between Non-Target and MFN2 mutant cell morphology and mean aggregated their prediction scores to create a soft-voting ensemble called Mito Path ScoreP (“P” indicates plate-based, Table S5). The results highlighted a group of syngRNAs which yielded strong MFN2 mutant-like phenotypes (Fig. 2a, Kruskal-Wallis amongst plate replicates p = 3.51 × 10−11). Out of all the validated gRNAs, TMEM11 yielded the highest Mito Path ScoreP, followed by NDUFAF4, TIMM8A, MRPS18C, NDUFAF7, NDUFS5, and RARS2 (Table S5). The specific phenotypic signature and penetrance (fraction of cells expressing the mitochondrial disruption) are expected to be different for each of these gRNAs. We observed a highly variable penetrance in the different gRNAs (Fig. S6) and a consistent aggregated phenotype (Fig. 2b). We also confirmed the robustness (COD r2 = 0.773) of this phenotype amongst technical replicates performed on different days with different physical experimental layouts (Fig. 2c). All of the constituent classifiers making up the Mito Path ScoreP ensemble generally agreed on the top gRNAs perturbing mitochondrial phenotypes (Fig. 2d).
a U2OS cells nucleofected with syngRNAs selected from the primary screen and their mitochondrial pathogenicity scores. Significant gRNAs with score >0.2 are listed with asterisks denoting P values: * P ≤ 0.05, ** P ≤ 0.01, *** P ≤ 0.001 (Wilcoxon rank sum test on plate replicates), overall Kruskal-Wallis p = 3.51 × 10−11. Orange bars indicate NADH ubiquinone oxidoreductase-related gene hits, blue bars were purposely selected non-hits, and green and red bars are negative and positive control gRNAs (Non-Target and MFN2 gRNA). Grey bars indicate gRNA selections of significant interest. Markers indicate individual plate replicate scores. b Fluorescent 20x confocal images showing mitochondrial aggregation of a sampling of syngRNA transfected U2OS cells and controls. Most of the cells show altered mitochondrial morphology similar to MFN2 mutations. c Scatterplot and Pearson’s R correlation of replicates for each syngRNA used in these experiments. Mito Path Scorep of the replicates are on each axis, and MFN2 gRNA was left out to better visualize the other syngRNAs. Each dot is an average of 531 ± 232 cells per replicate set. d Heat map showing the validation gRNAs and their Mito Path ScoreR derived from 48 individual binary classification models (blue wild type-like and red MFN2 mutant-like, scaled to the min and max per model, range is ~0.2–0.8).
We next explored the biology of the two highest scoring genes, NDUFAF4 and TMEM11. Polyclonal antibody staining was performed to reveal the localization of the proteins in WT U2OS cells (Fig. S7a). We noticed that the NDUFAF4 was symmetrically perinuclear but the TMEM11 was more polarized and aggregated in WT cells. Next, syngRNAs were nucleofected into the U2OS parental cell line for polyclonal founder cell populations. The cutting analyses showed between 64 and 90% of the reads were out-of-frame as a result from indels of the gRNA (data not shown). When applying the same polyclonal antibodies, we noted a 44.7% reduction of NDUFAF4 and a 23.2% reduction of TMEM11, respectively, in populations with those gRNAs. Interestingly, CRISPR editing of NDUFAF4 reduced TMEM11’s protein expression, while in the TMEM11 knockout, there was no effect on NDUFAF4’s protein expression. MFN2 knockout cells had reductions in both NDUFAF4 and TMEM11 protein expression (Fig. S7b). The co-reduction in protein expression levels implies previously unknown interactions between these genes.
We used the polyclonal founder pools to create isogenic clones of the NDUFAF4 and TMEM11 KO U2OS cells. We investigated the extent of genomic DNA alteration for isogenic TMEM11- and NDUFAF4-edited lines (clone E9 and F1, respectively). Both isogenics resulted in >94% out-of-frame indels in the genes, with the remaining reads appearing as substitutions, likely due to sequencing error. Not a single intact read was detected in these isogenic samples (Fig. S8a). Secondly, gene expression levels from ∆∆Ct analysis of each gene’s mRNA versus GAPDH were examined. In line with their genotype, NDUFAF4 F1 isogenic cells showed a 99.9% reduction of mRNA compared with controls, while TMEM11 E9 had a 96% reduction. For comparison, polyclonal MFN2 cells had only 57.2% reduction (Fig. S8b).
We next took the two isogenic clones, NDUFAF4 F1, TMEM11 E9, and the parental WT-Control line into a 96-well plate-based functional assay. We added the mitochondrial superoxide detector, mtSOX Deep Red to our earlier staining panel (Hoechst, MitoTracker, and TMRM) to assess reactive oxygen species (ROS) levels from mitochondrial superoxide in NDUFAF4- and TMEM11-KO cells compared to WT-controls (Fig. S9). The NDUFAF4 mutant had reduced TMRM intensity compared with the control, whereas incredibly elevated Mitotracker intensity (p = 6.7 × 10-6). The TMEM11 mutant had modestly increased MitoTracker intensity when compared to the WT control (overall p = 1.9 × 10−5, Fig. S9). Both isogenic knockouts had enhanced mito superoxide when compared to the control (p ≤ 6.4 × 10−5, p = 6.7 × 10−6 Fig. S9). These experiments indicate NDUFAF4 and TMEM11 mutants have mitochondrial morphological and oxidative impacts in U2OS cells.
Among the genes investigated, TMEM11-edited cells had the most penetrant MFN2 mutant-like phenotype. Previously, TMEM11 siRNAs (small interfering RNAs) have been shown to reduce mitochondrial tubules, while in Drosophila melanogaster’s TMEM11 ortholog (Pmi) mutations had defects in mitochondrial fission, and its phenotype could not be reversed by decreases in OPA1 or MFN234,35. Although there are few links to mitochondrial related genes, TMEM11 has been linked to MTX2, a gene known to be involved in mitochondrial trafficking in neurons (Fig. S10)37. TMEM11 has also been identified as an outer mitochondrial membrane protein that complexes with BNIP3 and BNIP3L to aid in mitophagy and is required for maintenance of normal mitochondrial morphology38. This is in line with our finding that TMEM11 knockout results in a high Mito Path Score.
It is worth noting that three of the NADH ubiquinone oxidoreductase-related genes produced significant Mito Path ScoreP values. These genes are known to be interconnected, and their protein network includes TIMM8A and MRPS18C (Fig. S10). Variants in NDUFAF4/7 are known to cause infantile mitochondrial encephalopathy (MC1DN15) and Leigh Syndrome39,40,41, while NDUFS5 has been correlated with Left Ventricular Reverse Remodeling (LVRR) in patients with non-ischemic dilated cardiomyopathy (NIDCM)42. NDUFS5 has also been linked to AIFM1, another gene found in these analyses43. TIMM8A variants are associated with Mohr-Tranebjaerg syndrome (deafness-dystonia-optic neuronopathy syndrome), and the gene is part of the mitochondrial protein import system44,45. MRPS18C is part of the mitochondrial ribosomal small subunit and is the least studied of the genes for which knockdown led to disrupted mitochondrial morphology46.
We also examined whether overexpression of normal (not mutant) MFN2 could rescue the mutant phenotype caused by TMEM11 or MRPS18C disruption. In addition to regular U2OS WT cells (with endogenous MFN2), we also introduced the syngRNAs into U2OS cells that had been pre-selected for MFN2 overexpression. Neither TMEM11 nor MRPS18C was rescued by the overexpression of MFN2, consistent with previous findings47. Curiously, MFN2 overexpression somewhat exacerbated the phenotype of TMEM11-edited cells (Fig. S11).
Scanning mutagenesis of SLC25A19 and ATAD3A
In addition to gene knockdown, we evaluated the effect of variants in two genes which are relevant to mitochondria and known to cause human disease: the transmembrane protein SLC25A19 and the ATPase ATAD3A. We interrogated a “scanning” gRNA library comprised of 224 gRNAs targeting SLC25A19 (142 gRNAs) and ATAD3A (82 gRNAs)48. The program inDelphi49 was used to design gRNAs tiled across both genes (Fig. 3a) with predicted in-frame editing consequences, leading to non-synonymous mutations. Using Raft-Seq, we again undertook pooled image-based analyses of U2OS cells which contained the scanning mutagenesis library. In total, 4128 cells were captured and genotyped. The isogenic clone MFN2V459fs/WT was again used as the MFN2-mutant control, while the WT-like control consisted of library-containing cells with no Cas9 cutting (no Dox). The Mito Path ScoreR was used on the primary dataset, and the results revealed positions where gRNAs produced indels that impacted mitochondrial morphology (Fig. 3b). The gRNA cut sites were also plotted along the cDNA of the respective gene, such that the disrupted domains are highlighted (Fig. S12), but this revealed no clear areas of cutting enrichment or depletion, likely resulting from frameshift mutations at multiple sites.
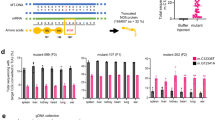
a Diagram of cDNA and domain architecture for two genes (ATAD3A and SLC25A19). gRNA cut sites are shown tiled across each gene from the scanning mutagenesis library. b Volcano plot results from the scanning mutagenesis library in these two genes. The x-axis shows the Mito Path ScoreR where higher values indicate mitochondrial phenotypes more like MFN2-mutant cells. −log p-value score determined by Mann-Whitney U test against Non-Target gRNAs. Dashed line indicates p-value threshold where FDR is 0.2. c Results from arrayed validation of the top eight gRNAs identified in the scanning mutagenesis. Genes selected for validation ranked by Mito Path ScoreP. Green denotes non-target, Blue anti-hits, and Red MFN2 gRNA. Overall Kruskal-Wallis p = 4.43 × 10−8, stars indicate significance as *=p ≤ 0.05 and **=p ≤ 0.01, and gray dots show individual replicate scores. d Pie charts showing deletion/insertion representation of the eight syngRNA bulk samples taken just before phenotyping. e Inner mitochondrial membrane-bound representations of SLC25A19 and ATAD3A proteins. For SLC25A19, the upper and lower dashed lines represent the inner mitochondrial membrane bilayer, with the intermembrane space above and matrix below. For ATAD3A, the intermembrane space is depicted above the dashed line. Key residues for both proteins are labeled.
We next ran a small set of multi-feature analyses (Fig. S13) on the results from the variant screening and selected ten of the highest-ranked gRNAs that scored consistently across the models (Fig. S14). The highest scoring gRNAs were then taken into plate-based validation experiments (Table S6). Using individual syngRNAs, we attained an average of 92% cutting and evaluated the Mito Path ScoreP for the set of SLC25A19 and ATAD3A variants (Fig. 3c, overall Kruskal-Wallis amongst plate replicates p = 4.43 × 10−8). SLC25A19 syngRNAs, which insert indels near V139 and V162, as well as the 5’ UTR at positions 1465, 1239, 1469, and 1560, all had significantly perturbed mitochondrial phenotypes. Likewise, ATAD3A indels at S343 produced a strongly perturbed phenotype. These results were as expected since those gRNAs had the strongest phenotypes in the microraft-based experiments. Images of the mitochondrial morphology of these variants highlight their disruption (Fig. S15).
Next, we analyzed the genomic consequences using bulk samples taken from the same cell lines used for the phenotypic analysis. Generally, the gRNAs brought about in-frame indels in just under half of the cells (as purposely designed by inDelphi). However, the specific insertions and deletions exhibited wide variation (Fig. 3d). Additionally, we employed AlphaFold2 to generate the proteins’ 3D structures within the mitochondrial membrane, since SLC25A19 and ATAD3A both produce mitochondrial membrane proteins50. The high Mito Path ScoreP mutations tended to fall near the membrane leaflet interface for SLC25A19, and at a single loop just before the ATPase domain in ATAD3A (Fig. 3e). While these variants did not induce as pronounced a perturbed phenotype as in MFN2, they appeared to have a subtle, yet robust, effect on mitochondrial morphology, comparable to the NDU knockdowns in the mitochondrial related gRNA screen. (Fig. S16 for comparison).
Underlying details of the multi-feature mitochondrial pathogenicity phenotype
To characterize the mitochondrial associated phenotype that Mito Path ScoreR represents, we assessed the individual contribution of its constituent features for their individual discriminative power. This section therefore, evaluates the phenotype itself. Each 20x confocal image of a cell was segmented in InCarta on the Hoechst-stained nucleus, and then a 12 µm region around the nucleus was expanded to account for the rest of the cell body. Pixels were measured separately for MitoTracker and TMRM intensity, producing 66 numeric features. These features were standardized and ranked by the Kolmogorov-Smirnov (KS) statistic comparing the labeled controls, MFN2V459fs/WT versus the WT-like No Cas9 (No Dox) condition (Fig. 4a). The integrated intensity of the perinuclear mitochondria (termed Nuclei Total Intensity) emerged as the most discriminative single feature (Fig. 4b), while texture features, which measure the variation of pixel intensity within a region of the cell, also scored a high KS statistic (Fig. 4c). MitoTracker features made up the top 3, and TMRM (depolarization of the mitochondria) came in as the 4th and 5th highest. However, no single feature adequately separated the labeled controls. Simple linear combinations of pairs of features also did not improve the separation between controls (Fig. 4d), but a multi-feature machine learning model did (Figs. 4e, S3). Random Forest classifiers demonstrated greater discriminatory power overall compared to Logistic Regressions but also produced overly optimistic AUC scores when inferring on data seen during training. Logistic Regressions produced more conservative AUC scores under the same conditions.
a. Twenty of the most discriminative features between MFN2V459fs/WT positive controls and No Cas9 (No Dox) negative controls in the mitochondrial gene knockout screening data as determined by the Kolmogorov-Smirnov test statistic. Bars are color-coded according to the wavelength from which each feature was derived. Every bar shown had a KS p-value near 0. b–e Cumulative probability histograms from the full dataset with the red line showing MFN2V459fs/WT cells and the blue line showing No Cas9 cells. The stated AUCs are for the full dataset (training and test). b Cumulative probability histogram segmented according to the most discriminative feature: MitoTracker nuclei total intensity. c The second most discriminative feature: MitoTracker cell intensity spreading. d. The product of the top two features. e One of the 12-feature logistic regressions from the Mito Path ScoreR ensemble. f, g Scatter plots and the Pearson’s R correlation between the Mito Path ScoreR and each of the top three principal components for MitoTracker and TMRM. The top loadings for each principal component are included. The percentage listed after the PC1, 2, and 3 are the percent explained variance for that PC.
Next, we decomposed our highly dimensional dataset using principal component analysis (PCA) for the MitoTracker and TMRM features (Fig. 4f, g). Cell-based comparisons against the PCs are also included (Fig. S17). For MitoTracker, the first 3 PCs explained 69% of the total variance. MitoTracker PC1 happened to strongly correlate (Pearson r = 0.68, p = 2.8e-64) with the Mito Path ScoreR, indicating that the morphological disturbance caused by the MFN2 mutation represented a large fraction in the data overall (31% variance explained), but not all of it. The top 3 feature loadings into PC1 were all perinuclear mitochondrial features that were related to the ‘texture’ or variation among the MitoTracker pixel intensities. The TMRM PCs were also moderately correlated to the Mito Path ScoreR (Pearson R = 0.49, 0.35, and 0.32), indicating that the variance in TMRM was not as well linked to the MFN2 phenotypic signature—likely because TMRM indicates mitochondrial activity and not morphology. Two of the top three feature loadings for TMRM PC1 were “Max Intensity”. This is the brightness of the strongest mitochondrial pixel in the TMRM wavelength and could be interpreted as the activity of the cells’ most active mitochondria. Importantly, these results emphasize that our analyses do not depend on resolving the mitochondrial network but are sufficiently powered by coarse measurements (Fig. S3). All these analyses were performed on data spanning the entirety of the microraft-based experiments and encompassed a large set of batch variations, making them more generalizable than if taken from a single experiment.
After characterizing the Mito Path ScoreR and using it to identify which perturbations best resemble mitochondrial aggregation found in MFN2 mutations, we set out to address if there were other genetic knockouts of interest in our data set that had mitochondrial phenotypes unlike mutant MFN2. This approach required a new scoring system from above.
Unique mitochondrial morphologies caused by knockdown of mitochondrial genes
Thus far, we had only identified dysmorphic mitochondrial phenotypes that resembled MFN2-mutant pathology. We also sought to identify genes which altered mitochondrial morphology in ways that were distinct from that of MFN2-mutants. Unlike the previous method of scoring phenotypes using classifiers trained on WT and MFN2 mutant cells, here we trained classifiers solely on WT controls—as to perform anomaly detection. This approach enabled us to utilize our existing phenotypic data to identify potentially pathogenic variants whose phenotype is not characterized primarily by the aggregated mitochondria of the MFN2 mutant cells. The combined outputs of several anomaly detection models (Anomaly ScoreR) pinpointed mitochondrial-related genes that, when silenced, resulted in anomalous cell phenotypes compared to cells that were nucleofected with a Non-Target gRNA (Fig. 5a). Among the most anomalous of these gRNAs were PLGRKT, COX7B2, MRPL39, and AIFM2. Although these gRNAs did not lead to aggregated mitochondria as in MFN2 mutants, they significantly affected mitochondrial morphology (Fig. 5b, additional images in Fig. S18).
a Volcano plot listing the Anomaly ScoreR on the x-axis, and the -log p-Value on the y-axis (p-value derived by T-test, with cutoff based on FDR of 0.2). Red is MFN2, Blue represents Non-Targets, Green represents genes that were selected for validation in the primary screen for their MFN2-mutant phenotype, and Yellow represents genes previously selected for their benignity. b Fluorescent 20x confocal images of anomalous single cells on microrafts: Blue = Hoechst, Green = MitoTracker, Yellow=MitoTracker/TMRM colocalization. c Venn Diagram showing genes above the FDR threshold by the Anomaly ScoreR (blue), and the Mito Path ScoreR from Fig. 1 (orange). Genes listed in the center are significant for both metrics. Bolded genes had strong Mito Path ScoreP in the Fig. 2 Validation set. d Previously validated gRNAs ranked by their Anomaly ScoreP instead of their Mito Path Scorep. NDU genes listed as Brown, with NDUFS5 and FAM195A containing two different gRNAs. Asterisks indicate p ≤ 0.001 (p-values derived by Mann-Whitney U-test comparing against Non-Target). e UMAP displaying the position of the single most anomalous gRNA per gene. Non-target gRNAs surround the region indicated (black dots), and several genes at the extremes of each region are labeled. Different genes are colored based on hierarchical clustering.
As expected, the Anomaly ScoreR revealed potentially pathogenic phenotypes that were not identified using the Mito Path ScoreR (Fig. 1). Figure 5c displays the name of the gene identified by either Anomaly ScoreR, Mito Path ScoreR, or both. The overlapping region includes the validated gene NDUFAF4 and eight others that show dysmorphic mitochondrial signatures by both anomaly detection and binary classification models (Fig. 5c). In addition, five genes were found to be significant only by the Anomaly ScoreR: APOOL, MCEE, NIT1, PHB, SLC16A7. Notably, APOOL has been shown to regulate mitochondrial cristae morphology51. While the link between some of these genes and mitochondrial dynamics has not yet been published, knocking them out in U2OS cells is sufficient to affect mitochondrial morphology.
Anomaly detection models were also trained on the arrayed data from Fig. 2, and their combined outputs (Anomaly ScoreP) were used to validate potentially pathogenic phenotypes without relying on the MFN2-mutant signature (Fig. 5d). According to Anomaly ScoreP, TIMM8A and MRPS18C emerged as the top mitochondrial-anomalous genes, and AIP and METTL17 became more significant than they were according to Mito Path ScoreP.
An alternative to our supervised approaches of binary classification and anomaly detection is the unsupervised algorithm Uniform Manifold Approximation and Projection (UMAP)52. We selected one gRNA for each gene based on its maximal difference from the Non-Target gRNA, and then UMAP embedded the gRNAs based on their mitochondrial features (Fig. 5e). Non-Target gRNAs were grouped together while other subclusters emerged. Examining the outliers within these clusters provides another avenue to identifying potentially interesting morphologies observed in our phenotypic data.
Discussion
We undertook a pooled image-based phenotype analysis of nearly 6000 gRNAs aimed at perturbing mitochondria. Using high-quality single-cell images, we assessed mitochondrial morphology (with a general mitochondrial dye) and polarization state (with TMRM). Our analysis was guided by a known genetic mutation in the gene MFN2, which served as a reference to define the abnormal mitochondrial signature, without biases to a particular pre-conceived phenotype like aggregation. The assay worked successfully, with a spiked-in control demonstrating high fidelity enrichment. Notably, Drp1 (DNM1L) was present in the gRNA library and is known to promote fission (in opposition to MFN2). The Mito Path ScoreR of two Drp1 gRNAs mirrored MFN2’s increase, but in the opposite direction relative to the non-targeting controls. We found several gRNAs that could significantly perturb mitochondrial morphology along the MFN2-mutant axis and undertook arrayed experiments to validate the effects of the gRNAs independent of the primary raft-based experiment. TMEM11 and TIMM8A were some of the strongest validated genes, as well as a suite of three NADH ubiquinone oxidoreductase-related genes, NDUFAF4, NDUFAF7, and NDUFS5. In addition to screening pools of mitochondrial gRNAs using Raft-Seq, we also focused on two specific genes-of-interest that are under-studied, yet important to mitochondrial function and dynamics—SLC25A19 and ATAD3A. We found several sites in those genes that, when mutated, also led to mitochondrial perturbation. Since no substitutions within this gene were produced by the library, there are no previously understood pathogenic variants nested in the screen itself. Details on the nature of the phenotype that identify all these genes involve specific mitochondrial patterns, which are examined in Fig. 4. Finally, we re-analyzed the cellular phenotypes to look for anomalous mitochondrial morphologies, extending beyond MFN2-mutant specific morphologies, and uncovered additional genes like APOOL, MCEE, NIT1, PHB, and SLC16A7. Several gene knockouts identified by their resemblance to MFN2-mutant morphology did not resurface as anomalous, likely because anomaly detection methods presuppose a multi-dimensional response space. This makes significance more difficult to achieve compared to MFN2-mutant versus wild-type classification, which collapses the response space to a single dimension. Overall, this manuscript offers causal links between gene knockouts and gene-specific variants into the assembly and maintenance of mitochondrial morphology.
As a pooled phenotyping platform, Raft-Seq has several strengths which allowed for this dataset and discoveries. Data is retrieved on a per-cell basis, rather than in bulk format, as is the case with enrichment or FACS-based screens23,26. It is most similar to In Situ Sequencing, that has been recently pioneered in a genome-wide phenotyping effort53. In that paper, every cell is genotyped after stripping the stains from imaging and doing several rounds of sequencing. In contrast, Raft-Seq only selects cells of interest after imaging, focusing the sequencing resources on a subset of cells. It also means that any type of sequencing can be performed, DNA seq, as is done in this paper to discover the gRNA, but also RNA seq in the style of FACS-Seq, on the captured wells. Raft-Seq also enables the selection of live single cells with various indel mutations from the primary phenotyping, and several isogenic clones are available for SLC25A19, ATAD3A, TMEM11, and NDUFAF4 upon request. Additionally, it is worth mentioning the advent of tools in the realm of machine learning-enabled variant prediction, including AlphaMissense and CPT54,55. While such tools excel in predicting substitution variant effects, they face limitations in their ability to predict the functional consequences of indel missense variants. The intricate and subtle nature of such variant effects also poses a challenge for conventional methods like survival or reporter assays, demanding an image-based approach.
Since we are interested in elucidating genes that may lead to mitochondrial-related neuropathy, we focused most of the manuscript on MFN2-mutant like phenotypes. Much work, including our own, has found that MFN2-induced changes in mitochondrial fusion lead to neurodegeneration13. For this reason, we used the MFN2-mutant cells to define the signature of the mitochondrial perturbation we were looking for, which is a combination of a fairly coarse aggregation phenotype and other more subtle features about mitochondrial morphology and TMRM polarization in each cell. But since the phenotype data is single images linked to a gRNA, many other approaches could be taken. In Fig. 5, we used anomaly detection to train classifiers on just the WT-like cells and scored how distant other gRNAs are from this “central” point. This could help define a mitochondrial ‘space’ that represents multiple dimensions but is still grounded in the WT-like state and supervised from there. Although we also showed an unsupervised UMAP projection of the dataset, this manifold is not aware that cells can be wild-type or perturbed, so it may miss the directions in mitochondrial ‘space’ that are of interest. The single-cell data is available for brand new interpretations of the phenotype and links with genotype (Table S3).
Another technical note is the use of two different scoring systems in the primary microraft-based experiments compared with the syngRNA plate-based experiments. We have both a Mito Path ScoreR (raft) and Mito Path ScoreP (plate), as well as a similar pair for anomaly detection. There are two reasons for this. First, the raft topology makes it such that the physical size of the cells is slightly different than in a plate. Also, the structure of the raft is slightly concave, so the cells do not sit as flat as they do on a plate. More importantly, all the gRNAs recovered from the microrafts were from cells treated with only Dox to induce Cas9 CRISPR cutting, whereas all the cells in the syngRNA plates were nucleofected with gRNA+Cas9 ribonuclear protein complexes. The nucleofection appeared to change the morphology and affected the controls and conditions alike but made cross-comparisons between raft and plate experiments challenging. For this reason, the two scoring systems were necessary.
These plate-based validation assays provided an opportunity to assess the accuracy of Raft-Seq’s variant calling under different FDRs and determine an optimal FDR for variant calling in future screens. Some of the thirty gRNAs selected from the primary screen fell below our initial FDR (Fig. 1d) and thus were not expected to validate. The proportion of the thirty selected gRNAs that were expected to validate to those that did validate in our plate-based assays was used to extrapolate a true FDR of 0.121 for the primary screen.
The genes found to strongly perturb mitochondrial morphology are fairly unstudied, such as TMEM11, NDUFAF4, TIMM8A, and MRPS18C. Some of them are known to cause disease in humans, and the data presented here could indicate that they might disrupt axons and cause degeneration, as is known to happen with MFN2-neuropathy13. Interestingly, the genes SLC25A51, FAM195A, AIP, and METTL17 were seen to be strong candidates from microraft-based screening experiments. However, in the validation (plate-based) experiments, their phenotype was weaker (Fig. 2b). Surprisingly, when their data was re-examined as part of the anomaly detection scoring in Fig. 5d, they were among the strongest results, with AIP specifically being almost as ‘anomalous’ as MFN2-mutants and both FAM195A gRNAs showing up strongly as well. SLC25A51 is a mammalian mitochondrial NAD+ transporter, so it may also relate to the NDU findings56. METTL17 and MRPS18C are both involved in mitochondrial ribosomes and in yeast mitochondrial stress response57. AIP is involved in mitochondrial protein import, as is TIMM8A58. Further studies were also conducted on isogenic mutants of TMEM11 and NDUFAF4, which showed extensive changes in MitoTracker-labeled mitochondria, as well as increased mitochondrial superoxide. While the imaging helped identify mitochondrial morphological differences, how these changes map onto underlying metabolic alterations require further investigation.
It is also worth mentioning that several gRNAs showed strong effects that were not discussed above. These include APOPT1, FASTKD5, STARD3, OGDHL, that were pulled from the original screening set but didn’t show as strong signals compared to MFN2-mutant. Like SLC25A51, they did show significant differences from the Non-Target gRNAs, and so are another validated set of genes that perturb mitochondrial morphology. Finally, since most gRNAs did validate in at least one of the two methods for testing them (either in a way similar to MFN2 or in a dimension away from Non-Target), it is worth listing that the majority of the following genes are also likely to perturb mitochondrial morphology but have not been independently tested with syngRNAs. They include ALDH9A1, ASAH2, C6ORF203, CASP8, COX7A1, COX7B2, CPT1B, CRAT, CRY1, DNAJC28, GSR, HCCS, HIGD2A, ISOC2, MRPL30, MRPL39, MRPL55, MSRB3, NAGS, PLGRKT, SLC25A44, and TOMM40L.
Methods
Culture and transfection
Human osteosarcoma U2OS cell line (U2OS, ATCC HTB-96) with doxycycline-inducible Cas913 was cultured in McCoy’s 5 A Modified Medium (16600082, Gibco, Gaithersburg, MD, USA) supplemented with 10% tetracycline-free fetal bovine serum (FBS) (PS-FB3, Peak Bio, Pleasanton, CA, USA). Human embryonic kidney (HEK) 293 T cells (CRL-11268, ATCC) line was cultured in Dulbecco’s Modified Eagle’s Medium (11965-092, Gibco) supplemented with 10% FBS (26140079, Gibco), 1% Penicillin-Streptomycin (15140122, Gibco), and 1% non-essential amino acids (11140050, Gibco). Both lines were maintained at 37 °C, 5% CO2, and were monitored daily for overall cell health and confluency. Cells were grown in either T75 or T150 tissue culture flasks and were passaged at 90% confluency. Trypsinization utilized 0.25% Trypsin-EDTA 1x (25200056, Gibco) on the cells for 5 min in the 37 °C incubator, and the cell suspensions were collected and centrifuged for 3 min at 1200 RPM. Cells were counted with either a BioRad TC20 automated cell counter or hemacytometer. Cells were passaged at a 1:10 ratio and were always maintained at a cellular representation of (# gRNAs * 1000 * rF), where rF was the representation factor between 2 and 5 for different sub-pools of the library. Passage number was recorded at each passage, and cells were not allowed to exceed 10 passage events before freshly thawed cells were introduced. Testing for mycoplasma was performed bi-annually. Experiments in this paper utilized 100x100 µm single and quad reservoir CellRaft Cytosort raft plates (CS-100S and CS-100Q, Cell Microsystems Inc., NC, USA). Prior to plating, raft plates were rinsed 3 times with 1xPBS and allowed to incubate at 37 °C, 5% CO2 for 3 minutes between each rinse. Plating densities for each were as follows; Quad 48,000 cells total (12,000/quadrant), Single 32,000 cells total. These wells were then brought up in 300–500 µl of their respective medias. Post-plating, raft plates were allowed to incubate for 12–14 h prior to staining/imaging. Isogenic clone MFN2V459fs/WT was spiked into the screen at a frequency of 0.2%.
Lentivirus production and titering
Lentiviruses carrying the gRNA libraries were packaged into 8 × 106 HEK 293 T cells/well in a 6-well plate (TPP 92006, MIDSCI, Fenton, MO, USA), using the TransIT Lentivirus transfection reagent (MIR 6600, Mirus Bio, Madison, WI, USA). Our mitochondrial-related gRNA libraries were designed from Brunello gRNA in seven sub-pools with 4-gRNAs per gene and 5% Non-Target (Table S1)59. These gRNAs were assembled into plasmids by Genome Engineering and Stem Cell Center (GESC@MGI) at Washington University in St. Louis, and their representations were confirmed. The gRNA plasmid libraries were combined with VSVg (Plasmid #8454, Addgene, Watertown, MA, USA) and psPAX2 (Plasmid #12260, Addgene) plasmids in a mass ratio of 0.5/0.5/1.0 for a total of 2 µg, before being combined with Opti-MEM media (31985070, Gibco) and TransIT reagent according to the manufacturer’s instructions for each well of a 6-well plate. Roughly 200 µl of packaging solution was added to each well of a 6-well plate and incubated at 37 °C and 5% CO2 for 48–60 h. Media was collected, centrifuged to remove cell debris, and passed through a 0.22 µm PES sterile filter (SLGP033RS, EMD Millipore, St. Louis, MO, USA) attached to a 5 mL BD Luer-Lok™ Syringe sterile, single use (309646, BD Biosciences, Franklin Lake, NJ, USA). One-third of the total media volume of Lenti-X Concentrator (631232, Takara Bio, Shiga, Japan) was added to the viral media and refrigerated for 30 min at 4 °C before being centrifuged for 45 min at 1500 × g and 4 °C. Supernatant was removed, and the concentrated lentivirus was resuspended in 1/10th initial media volume with 1xPBS (10010023, Gibco). Lentivirus was aliquoted and stored at −80 °C in 50 µl aliquots.
Lentivirus was titered by serially infecting U2OS wild type cells with 1/5, 1/25, and 1/125 dilutions with 1 µg/µl Polybrene infection/transfection reagent (TR-1003-G, Sigma) for 24 h. Cells were trypsinized with 0.25% Trypsin (25200056, Gibco) for five minutes in the 37 °C incubator, and DNA was extracted with in-house extraction buffer to run qPCR (quantitative polymerase chain reaction) for titer determination. IDT PrimeTime Gene Expression Master Mix (1055770, IDT, Coralville, IA, USA) was used for qPCR, along with primers and probes to detect Albumin and Psi, to determine titer against a lentivirus control standard. An ABI 3100 Cycler was used to run qPCR and determine titer.
CRISPR/Cas9 gRNA library infection and induction
U2OS gRNA library cell lines were created by lentiviral infection at MOI = 0.1, which was performed in T75 flasks. Lentivirus was added to U2OS iCas9 cells along with 10 µg/ml Polybrene (TR-1003-G, Sigma, St. Louis, MO, USA), and allowed to infect cells in a 37 °C and 5% CO2 incubator for twenty-four hours. Lentivirus media was removed after 24 h, and the cells were allowed to rest for another twenty-four hours before being selected for seven days with Puromycin at 8 µg/ml (73342, STEMCELL Technologies, Cambridge, MA, USA). Surviving cells were expanded, frozen down, and submitted for bulk NGS (Next Generation Sequencing) to ensure representation of specific guides. STR profiling, to confirm cell type, was performed using NGS-based analysis by the GESC@MGI. Representation of the gRNAs in the cell lines was performed by bulk PCR and NGS analysis to confirm presence of all gRNAs and their rF (representation factor).
Staining and microscopy
Cells were stained with the following vital dye suite: Hoechst 33342 (4.05 µM, Hoechst, Thermo Fisher H3570), MitoTracker Deep Red (MitoTracker, 0.5 µM, Thermo Fisher M2246), and Tetramethylrhodamine methyl ester (TMRM, 0.1 µM, Thermo Fisher I34361) for 30 min at 37 °C and 5% CO2. A master batch of stain was made on the day of screening and kept away from light to avoid batch effects. Post-staining, media was removed, and raft plates were rinsed once with media (1 mL of media per quadrant or 5 mL of media for Single raft plates). Final media volume of raft plates prior to scanning was 2 mL (Single) or 500 µl/quadrant (Quad). The plates were placed in a Cell Microsystems plate adapter and imaged on Molecular Devices INCell 6500HS Confocal microscope at 20x 0.45 NA utilizing live cell chamber capabilities (5% CO2, 37 °C temperature). A custom protocol was developed to maximize fields and field of view overlap (12%). Exposure times for Hoechst (405 nm) and TMRM (561 nm) averaged 0.15 s while MitoTracker Deep Red (642 nm) averaged 0.05 s. Confocality was used in the 405 and 642 nm wavelengths, and plates were scanned in two halves to expedite image tracing, classification, and picking turnaround time.
For 40x imaging, cells were plated as described in the validation section on 96 well plates (89606, ibidi USA, Fitchberg, WI, USA). After plating the cells for imaging, the 96-well plate was incubated at 37 °C and 5% CO2 overnight. The plate was stained with 4.05 µM Hoechst 33342, 0.83 µg/ µL MitoTracker Green, 0.1 µM TMRM, and 10 µmol/L mtSOX Deep Red (MT16, Dojindo Laboratories, Rockville, MD) for 30 min at 37 °C and 5% CO2 as before. A master batch of stain was made on the day of imaging and kept away from light to avoid batch effects across plates. The 96-well plate was imaged on an ImageXpress Confocal HT.ai High-Content Imaging System (Molecular Devices, San Jose, CA, USA) microscope with a 40x water APO Lambda objective while utilizing live cell chamber capabilities.
Image QC and segmentation
The in-house custom software “FIVTools” was used to perform quality control and calibrations (https://gitlab.com/buchserlab/fivetools). Semi-manual focus checks ensured that out-of-focus fields were excluded from the analysis. Molecular Devices software, InCarta, was used to generate segmentation masks, delineating regions of interest corresponding to nuclei, cell body, and mitochondria. Tracing was performed for the nuclei according to Hoechst, and mitochondria were traced in both MitoTracker and TMRM-based fluorescent images to account for mitochondrial morphology and membrane potential, respectively. For the cell body mask, the nuclei mask was expanded by 12 µm using collar. A network segmentation algorithm was used to find mitochondrial puncta, likewise within the 12 µm radius of the cell. Once subcellular masks are generated, InCarta can compute several pixel-based measurements on a single-cell basis.
Quantitative image analysis
A graphical illustration of the process is also provided in Fig. S19. A full list of these features is available in Table S2. Each feature was identified by the subcellular mask, the measurement computed, and the fluorescent wavelength. These cell-based features were joined with image quality annotations and raft position mapping in FIVTools. The resulting data is hierarchically organized. Quantitative image analysis, downstream machine learning, and statistics are similar for both raft-based and plate-based experiments. However, single cells and their genotype can only be identified within the context of the nested experimental that differs between raft-based and plate-based experiments (important for data aggregation schemes that influence data normalization). In both raft-based and plate-based experiments, single cells are data-table rows. In the former case, the hierarchy proceeds top-down from experimental ID, plate, then raft. In the latter case, the hierarchy proceeds from experimental ID to plate, well, field of view (FOV), and finally object ID. The resulting data was used to analyze single cells for post-tracing quality control using Tibco’s Spotfire Analyst software. The inclusion criteria for cells taken to downstream analyses were based on nuclear area, nuclear form factor, and distance of the cell from the edge of the raft. Furthermore, cellular intensity, nuclear intensity, and nuclear area were used to exclude any rafts that had non-cellular debris or dead nuclei (Fig. S1). Quality checked cells were used to identify or validate genetic variants of interest based on machine learning models trained on the mask-defined, pixel-based subcellular features.
Machine learning
The filtered cell data table was imported into a Jupyter notebook running Python (version 3.8.13) within Microsoft’s Visual Studio Code application. In a single raft-seq experiment (mid screen), the analysis served to identify cells of interest to be captured for genotyping. In compiled experiments (raft and plate-based), the prediction of Mito Path Score was equivalent to calling variants as likely pathogenic or benign based on their phenotype. In all cases, the task was a binary classification such that the features computed by the InCarta segmentation constituted the predictor variables, and the genotype of each cell was the target variable. The resulting prediction scores of an ensembled machine learning model established the basis for identifying potentially pathogenic phenotypes in single-cells, and subsequent statistics determined the gRNA as causative or not.
Batch effects mitigation
Due to spatial and temporal differences between experiments and conditions, small, artifactual variations (batch effects) between replicates can emerge at multiple scales. Several computational methods exist for correcting batch effects in single-cell RNA-seq data. However, many of these rely on initial dimensionality reduction techniques that would obscure the richness of the data and make biological explanations of image-based phenotypes difficult. Other methods, such as Empirical Bayes Batch Correction or Surrogate Variable Analysis correct batch effects directly on cell features but make potentially inappropriate assumptions due to the complexity of Raft-Seq’s multi-levelled batch effects. Instead, batch effects were managed explicitly in the experimental design. Consistent settings and protocols were maintained across screening experiments, making inter-plate variation—likely due to temporal factors—the primary source of batch effects. By treating each plate as a discrete unit of variation and applying a simple standard scaler (plate-wise mean and standard deviation), we normalize cell-level data in a way that preserves biological signal while minimizing technical noise.
Feature selection
Features that were traced in Hoechst or brightfield, global and background features, and most mitochondrial mask features were excluded from the training data (Hoechst and Brightfield do not include useful information, background and global features are near proxies for the position of a cell on the plate which ‘leaks’ target variable information, and mitochondrial mask features consist of a small number of pixels and are noisy). Features were further selected to minimize multicollinearity by thresholding for inter-feature correlations and maximize information content by thresholding for intra-feature variance across cells. Features were also selected for their importance weights according to an L1-regularized linear support vector machine. To prevent overfitting, models were constrained to a set number of features per fit, with 12 features providing a balance between over- and under-fitting.
Ensemble generation
To generate ensemble models, we randomly sampled the (above-filtered) feature space, fit a model, and then iterated until the filtered feature space was sampled. Typically, each model utilized the same hyperparameters, selected from some initial tuning. Once all models had been generated, each was deployed on the entire dataset to infer pathogenicity or anomaly prediction scores for each cell. These prediction scores provided a final opportunity to assess model performance by plotting a cumulative probability histogram of the models’ predictions on different admixtures of labeled cell populations that were withheld from training data. A reliable model would not only rank mutant cells highly and WT cells lowly but would also rank mixed populations according to the expected proportion of WT and mutant cells. We then took a soft-voting approach to ensemble models by taking an unweighted average of the prediction scores. This mean score served as the metric to call variants, the Mito Path Score.
Cell capture and DNA extraction
Raft capture is detailed previously13. Briefly, semi skirted 96-well PCR plates (1402-9200, USA Scientific, Ocala, FL, USA) were prepped by dispensing 5 µl of extraction buffer (molecular grade water with 10 mM Tris-HCl (pH 8.0), 2 mM EDTA, 200 µg/mL Proteinase K, and 0.2% TritonX-100) into each well using an Integra 12 channel dispenser and the plates are sealed prior to isolations/picking. Once a 96-well PCR plate is isolated, it is placed in a BioRad T100 thermocycler (65 °C for 15 min, 95 °C for 5 min) to lyse the cell and extract DNA. PCR plates are spun down at 2000 RPMs for 1 min and stored at 4 °C prior to downstream NGS applications. Each 96-well plate is manually checked under a dissection microscope to verify successful isolations in each well. Subsequent isolations require disinfection of the PCR wand and loading additional custom picklists with the rafts of interest.
DNA amplification and sequencing
Extracted DNA was amplified using KOD Hot Start Polymerase (71086-4, Millipore Sigma, St. Louis, MO, USA) following manufacturer’s instructions with primers designed in the U6 region of the gRNA CCIV plasmid. Amplification using KOD was optimized for 30 cycles. A nested PCR approach was used where a larger section of the U6 region was amplified and then performed PCR1 with DeepSeq primers (primers with a complimentary Illumina sequence for i7 and i5 sequencing primer attachment) nested inside that larger amplicon60. This allows the samples to bypass a costly cleanup step.
KOD Primers:
F | GACGGTACAGGCCAGACAAT |
R | ACTTTCCCAGTTTACCCCGC |
PCR1 DeepSeq Primers:
F | CACTCTTTCCCTACACGACGCTCTTCCGATCTTCTTGTGGAAAGGACGAAACAC |
R | GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTTTGTGGATGAATACTGCCATTTG |
These methods were expanded from a previous paper13. After amplification of the single cell gRNA amplicon, a second amplification was performed to attach a specific 6-base pair (bp) barcoded forward Illumina index primer to the existing forward DeepSeq tags for four 96-well plates with picked single U2OS cells. These primers were used in combination with 4x 96-well plates of reverse Illumina Index primers from the GESC@MGI to give a unique combination of forward and reverse primers to each well. This allowed the amplified products to be pooled and demultiplexed after next-generation sequencing. A mastermix containing the forward NGS primers, Bioline MyTaq HS Red Mix 2x (C755G97, Merdian Life Sciences, Memphis, TN, USA), and Nuclease Free Water (AM9937, Thermo Scientific, Waltham, MA, USA) was added to 384-well plates (1438-4700, USA Scientific) using a Multidrop Combi Reagent Dispenser (Model 836, Thermo Scientific) and then pooled with 2 µL of PCR1 product and reverse indexes using the Biomek I5 liquid handling system (Beckman Coulter, Indianapolis, IN, USA). After pooling of the indexed amplicons on the Biomek, 100 µL of pooled sample was cleaned up with 60 µl of AMPure XP beads (A63882, Beckman Coulter) according to the manufacturer’s protocol and normalized to the correct concentration using the NanoDrop One Spectrophotometer (Thermo Scientific, ND-ONE-W). Samples were then submitted to the Center for Genome Sciences and Systems Biology (Washington University in St. Louis) for either a 2x250 bp run or a 2x150 bp run, depending on amplicon size, performed using the Illumina MiSeq Platform. FastQ files were obtained from demultiplexed Illumina paired reads, then joined, trimmed, and aligned with the various pooled mito gRNA libraries. The resulting dataset showed the number of reads containing each of the 20-mer gRNAs for each well. From the primary screen, we preselected 10,000 cells. We do lose a percentage of the preselected 10,000 cells as they progress through various QC stages in the pipeline resulting in the final list of gRNAs with significant allele frequencies >80%. Approximately 10–15% of cells are lost from preselection to actual isolation on the Cell Microsystems instrument. Approximately 20-25% of cells are removed during QC/preliminary genotyping. Lastly, 25–30% of cells are omitted from the final list during the final genotyping QC leading to a single gRNA with greater than 80% single allele frequency per well. Pipeline optimization is in the works focusing on final genotyping. Typically, a majority of the screened cells were captured after adjusting for isolation and sequencing errors (these filters were applied in Tibco Spotfire), and a flat file as exported containing each picked raft and its assigned genotype.
Selecting gRNAs from primary analysis
Genes of interest were identified by generating p-values for each gRNA in the variant library using a one-sample t-test that compared the Mito Path ScoreR of each gRNA to the median score of all Non-Target gRNAs. We determined a threshold for significance by assuming a FDR of 0.2 and calculated Benjamini-Hochberg critical values for each gRNA. The largest p-value that was less than its corresponding critical value was used as the significance threshold. Genes considered for validation also had a minimum of 2 experimental replicates and 2 distinct gRNAs targeting the same gene. Thus, any gene targeted by a gRNA that had a significant p-value and a Mito Path ScoreR greater than the Non-Target comparator was considered a gene of interest. Some genes were selected for validation despite their lack of significance, enabling the post hoc assessment of Raft-Seq’s efficacy in identifying variants that successfully validated at different FDR.
Validation with synthetic gRNAs
Synthetic gRNAs (syngRNAs) (Table S4, S6) from genes of interest were ordered from Integrated DNA Technologies (IDT, Coralville, IA, USA). Primers complementary to 150 bp in either direction of the gRNA were designed and ordered through IDT. U2OS cells were grown in either T75 or T150 tissue culture flasks and were passaged at 90% confluency. RNP complexes consisting of 1 µl of syngRNA at 100 µM complexed with 10 µg of Cas9 protein and stored at −20 °C the day prior to nucleofection. Nucleofections were performed using 4D-Nucleofector Core and X unit (AAF-1003B, AAF-1003X, Lonza, Morristown, NJ, USA). In total, approximately 75,000 U2OS cells were nucleofected into small cuvettes (V4SP-3096, Lonza) with 20 µl of P3 Primary Cell line solution (V4SP-3096, Lonza) and RNP using the cell type program DS-150. Post-nucleofection, contents from the nucleofection were transferred to 12-well plates (TPP 92012, MIDSCI, Fenton, MO, USA) with 2 mL of McCoy’s media and were allowed to proliferate for 7 days, with media changes on the second and fifth days. The syngRNA cell conditions were randomly arrayed (to reduce batch variation by averaging out potential plate-related effects) across two 96-well black clear bottom tissue culture treated plates (6005182, Perkin Elmer, Shelton, CT, USA) using custom software and plated using the Biomek i5 liquid handling robot. After plating, bulk samples were taken from remaining cell suspensions, and genomic DNA extractions were performed for downstream NGS/cutting analysis. For the indel analysis, next generation sequencing reads for each bulk sample were aligned against gene specific reference sequences to catalogue causative deletions/insertions for each gRNA.
After plating the cells for imaging, the 96-well plates were incubated at 37°C and 5% CO2 overnight. The 96-well plates were stained with 4.05 µM Hoechst 33342, 0.5 µM MitoTracker Deep Red, and 0.1 µM TMRM for 30 min at 37 °C and 5% CO2 as before. A master batch of stain was made on the day of screening and kept away from light to avoid batch effects. The 96-well plates were imaged on Molecular Devices INCell 6500HS Confocal microscope at 20x 0.45 NA utilizing live cell chamber. Images were analyzed with InCarta as before, filtered, and compiled. Then, in a Jupyter notebook, multi-feature models were again trained, but using small aNNs (artificial neural networks with 2-3 dense layers) to distinguish between the Non-Target gRNA control and the MFN2 gRNA control (Fig. 2d, Table S2).
Isogenic cell line production
Conditioned McCoy’s media, recovered one week after cell culturing to help retain growth factors, was sterile-filtered (SCGP00525, Millipore Sigma, St. Louis, MO, USA), and 200 µl were dispensed across each 96-well round bottom tissue culture plate (TPP 92096) using an Integra 12-channel pipette. Next, 96-well tissue culture plates were allowed to warm prior to isolations. Custom picklists targeted the perturbed cells of interest and were loaded into array records in preparation for isolating/picking. CellRaft AIR System was used to isolate individual rafts into a 96-well flat bottom tissue culture plate. Post-isolation, lids were immediately placed onto the tissue culture plates and were promptly transferred to an incubator at 37 °C, 5% CO2. Media was added to tissue culture plates weekly (100 µl) to account for evaporation. Single cells on rafts were allowed to expand in culture for 2 weeks before they were imaged on Cytiva INCell 6500HS Confocal microscope at 4x in brightfield to evaluate growth. Once wells reached about 40% confluency, they were transferred and progressed through to larger vessels. Ten isogenic U2OS clones are available from SLC25A19 and ATAD3A, as well as TMEM11 and NDUFAF4.
RT-qPCR to validate gene KOs
In addition to taking a sample for NGS cutting efficiency analysis and cell plating, a small subset of isogenic cells was also taken for RNA extraction and RT-qPCR analysis. RNA was extracted from samples using a Nucleospin RNA Extraction Kit (74095550, Machery-Nagel, Duren, Germany) according to protocol. Samples were prepped for RT-qPCR according to protocol using a qScript 1-Step SYBR Green qRT-PCR Kit (95087-200, Quantabio, Beverly, MA, USA), and primers from IDT were designed to span an exon-exon junction in each test gene to prevent DNA amplification. RT-qPCR was run on bulk transfected, bulk untransfected, and isogenic clones of U2OS RNA samples along with primers from the housekeeping gene GAPDH to normalize gene expression. Samples were run on an Applied Biosystems QuantStudio 3 Real-Time PCR system (Thermo Fisher, Waltham, MA, USA), and a ∆∆Ct analysis was implemented to determine gene expression fold change.
Antibody staining
Primary antibodies for validated gRNAs of TMEM11 (PA566537, Thermo Fisher) and NDUFAF4 (PA5-89771, Thermo Fisher) were added to synthetic gRNA transfected U2OS cells for 1 h at room temperature according to manufacturer’s instructions after fixation with 4% Paraformaldehyde (15714-S, Electron Microscopy Sciences, Hatfield, PA, USA) for 15 min, permeabilization with 0.5% Triton-X 100 (T8787-50ML, Sigma)) for 15 min, and blocking with 3% Bovine Serum Albumin (A4612, Sigma) for 1 h at room temperature. All samples were rinsed 3x with 1xPBS (J61196.AP, Thermo Fisher) after each step. Donkey anti-Rabbit igG (H + L) Secondary Antibody (A32790, Thermo Fisher) was applied for one hour at room temp to each primary stained group of cells and combined with Hoechst before being imaged.
Protein structure
Three-dimensional structures for ATAD3A and SLC25A19 proteins were estimated with the monomer model of AlphaFold2 using the full DBS sequence database50. The highest-ranked model for each protein was selected for transmembrane orientation prediction using the PPM 3.0 web server61. ATAD3A and SLC25A19 both associated with the inner mitochondrial membrane, and the N-termini of both proteins were constrained to reside within the transmembrane space18,62. Membrane-bound proteins and specific residues with significant Mito Path ScoreR were visualized in the PyMOL Molecular Graphics System v2.5.5 (Schrödinger, LLC).
Data availability
Data that support the findings of this study have been deposited in Figshare at the following links: Figshare private link - https://figshare.com/s/dff650e9f62f587621d2. Figshare DOI - https://doi.org/10.6084/m9.figshare.25534324.
Code availability
Abbreviations
- aNN:
-
Artificial Neural Network
- AUC:
-
The area under the ROC curve of sensitivity/specificity
- Dox:
-
Doxycycline used to induce Cas9 cutting
- gRNA:
-
guide RNA for CRISPR Cas9
- MFN2 :
-
Mitofusin2, a gene that regulates mitochondrial fusion
- MFN2-mutant like:
-
Cells with morphological features similar to MFN2-disrupted cells, specifically MFN2V459fs/WT
- Non-Target gRNA:
-
a scrambled sequence known to not target any human gene
- MitoTracker:
-
Vital stain that marks all mitochondria
- TMRM:
-
Tetramethylrhodamine methyl ester, a red-orange vital stain that is readily sequestered by active (depolarized) mitochondria
- syngRNAs:
-
synthetic gRNA used with Cas9 protein as an RNP
- U2OS:
-
Osteosarcoma sarcoma cell line, usually with a dox-inducible Cas9
- WT-like:
-
Cells with morphological features similar to wild-type cells. Usually No Cas9 (no doxycycline induction)
- CMT2A:
-
Charcot-Marie-Tooth Disease Type 2 A
- FDR:
-
False discovery rate
- CNS:
-
Central nervous system
- FACS:
-
Fluorescence-Activated Cell Sorting
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- siRNA:
-
small interfering RNA
- MOI:
-
multiplicity of infection
- STR:
-
short tandem repeat
References
Ma, K. et al. Mitophagy, mitochondrial homeostasis, and cell fate. Front. Cell Dev. Biol. 8, 531786 (2020).
Seo, B. J., Yoon, S. H. & Do, J. T. Mitochondrial dynamics in stem cells and differentiation. Int. J. Mol. Sci. 19, 3893 (2018).
Tait, S. W. G. & Green, D. R. Mitochondrial regulation of cell death. Cold Spring Harb. Perspect Biol. 5, a008706 (2013).
Pekkurnaz, G. & Wang, X. Mitochondrial heterogeneity and homeostasis through the lens of a neuron. Nat. Metab. 4, 802–812 (2022).
Suárez-Rivero, J. M. et al. Mitochondrial dynamics in mitochondrial diseases. Diseases 5, 1 (2016).
Ferree, A. & Shirihai, O. Mitochondrial dynamics: the intersection of form and function. Adv. Exp. Med Biol. 748, 13–40 (2012).
Tilokani, L., Nagashima, S., Paupe, V. & Prudent, J. Mitochondrial dynamics: overview of molecular mechanisms. Essays Biochem. 62, 341 (2018).
Chen, W., Zhao, H. & Li, Y. Mitochondrial dynamics in health and disease: mechanisms and potential targets. Sig Transduct Target Ther. 8, 333 (2023).
Pipis, M. et al. Natural history of Charcot-Marie-Tooth disease type 2A: a large international multicentre study. Brain 143, 3589 (2020).
Huang, P., Yu, T. & Yoon, Y. Mitochondrial clustering induced by overexpression of the mitochondrial fusion protein Mfn2 causes mitochondrial dysfunction and cell death. Eur. J. Cell Biol. 86, 289–302 (2007).
Zaman, M. & Shutt, T. E. The role of impaired mitochondrial dynamics in MFN2-mediated pathology. Front. Cell Dev. Biol. 10, 858286 (2022).
Franco, A., Walton, C. E. & Dang, X. Mitochondria clumping vs. mitochondria fusion in CMT2A diseases. Life 12, 2110 (2022).
Yenkin, A. L. et al. Pooled image-base screening of mitochondria with microraft isolation distinguishes pathogenic mitofusin 2 mutations. Commun. Biol. 5, 1–14 (2022).
Ghiselli, F. & Milani, L. Linking the mitochondrial genotype to phenotype: a complex endeavour. Philos. Trans. R. Soc. B 375, 20190169 (2020).
Yonova-Doing, E. et al. An atlas of mitochondrial DNA genotype–phenotype associations in the UK Biobank. Nat. Genet. 53, 982–993 (2021).
Dolce, V., Fiermonte, G., Runswick, M. J., Palmieri, F. & Walker, J. E. The human mitochondrial deoxynucleotide carrier and its role in the toxicity of nucleoside antivirals. Proc. Natl. Acad. Sci. USA 98, 2284–2288 (2001).
Palmieri, F., Scarcia, P. & Monn‚, M. Diseases caused by mutations in mitochondrial carrier genes SLC25: a review. Biomolecules 10, 655 (2020).
Bottega, R. et al. Functional analysis of the third identified SLC25A19 mutation causative for the thiamine metabolism dysfunction syndrome 4. J. Hum. Genet. 64, 1075–1081 (2019).
Desai, R. et al. ATAD3 gene cluster deletions cause cerebellar dysfunction associated with altered mitochondrial DNA and cholesterol metabolism. Brain 140, 1595–1610 (2017).
Zhao, Y. et al. ATAD3A oligomerization causes neurodegeneration by coupling mitochondrial fragmentation and bioenergetics defects. Nat. Commun. 10, 1–20 (2019).
Zhao, Y. et al. ATAD3A oligomerization promotes neuropathology and cognitive deficits in Alzheimer’s disease models. Nat. Commun. 13, 1–20 (2022).
Chen, L., Li, Y., Zambidis, A. & Papadopoulos, V. ATAD3A: A key regulator of mitochondria-associated diseases. Int. J. Mol. Sci. 24 https://doi.org/10.3390/ijms241512511 (2023).
Kishi, J. Y. et al. Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing. Nat. Methods 19, 1393–1402 (2022).
Funk, L. et al. The phenotypic landscape of essential human genes. Cell 185, 4634–4653.e22 (2022).
Feldman, D. et al. Pooled genetic perturbation screens with image-based phenotypes. Nat. Protoc. 17, 476–512 (2022).
Schraivogel, D. et al. High-speed fluorescence image-enabled cell sorting. Science 375, 315–320 (2022).
Rensvold, J. W. et al. Defining mitochondrial protein functions through deep multiomic profiling. Nature 606, 382–388 (2022).
DiPeso, L., Pendyala, S., Huang, H.Z., Fowler, D.M. & Hatch, E.M. Image-based identification and isolation of micronucleated cells to dissect cellular consequences. eLife 13, RP101579 (2025).
Schäfer, J. A., Bozkurt, S., Michaelis, J. B., Klann, K. & Münch, C. Global mitochondrial protein import proteomics reveal distinct regulation by translation and translocation machinery. Mol. Cell 82, 435–446.e7 (2022).
Cho, N. H. et al. OpenCell: endogenous tagging for the cartography of human cellular organization. Science 375, eabi6983 (2022).
Laine, R. F., Arganda-Carreras, I., Henriques, R. & Jacquemet, G. Avoiding a replication crisis in deep-learning-based bioimage analysis. Nat. Methods 18, 1136–1144 (2021).
Heckenbach, I. et al. Nuclear morphology is a deep learning biomarker of cellular senescence. Nat. Aging 2, 742–755 (2022).
Haghighi, M., Caicedo, J. C., Cimini, B. A., Carpenter, A. E. & Singh, S. High-dimensional gene expression and morphology profiles of cells across 28,000 genetic and chemical perturbations. Nat. Methods 19, 1550–1557 (2022).
Voeltz, G. K., Sawyer, E. M., Hajnó, G. & Prinz, W. A. Making the connection: How membrane contact sites have changed our view of organelle biology. Cell 187, 257–270 (2024).
Thoudam, T. et al. Noncanonical PDK4 action alters mitochondrial dynamics to affect the cellular respiratory status. Proc. Natl. Acad. Sci. USA 119, e2120157119 (2022).
Zwerger, M., Ho, C. Y. & Lammerding, J. Nuclear mechanics in disease. Annu. Rev. Biomed. Eng. 13, 397 (2011).
Zhao, Y. et al. Metaxins are core components of mitochondrial transport adaptor complexes. Nat. Commun. 12, 1–17 (2021).
Gok, M. O. et al. The outer mitochondrial membrane protein TMEM11 demarcates spatially restricted BNIP3/BNIP3L-mediated mitophagy. J. Cell Biol. 222, e202204021 (2023).
Saada, A. et al. C6ORF66 is an assembly factor of mitochondrial complex I. Am. J. Hum. Genet. 82, 32–38 (2008).
Baertling, F. et al. NDUFAF4 variants are associated with Leigh syndrome and cause a specific mitochondrial complex I assembly defect. Eur. J. Hum. Genet. 25, 1273–1277 (2017).
Sonsalla, G. et al. Direct neuronal reprogramming of NDUFS4 patient cells identifies the unfolded protein response as a novel general reprogramming hurdle. Neuron 12, 117–1132 (2024).
Iwahana, T. et al. Novel myocardial markers GADD45G and NDUFS5 identified by RNA-sequencing predicts left ventricular reverse remodeling in advanced non-ischemic heart failure: A retrospective cohort study. BMC Cardiovasc. Disord. 20, 1–12 (2020).
Salscheider, S. L. et al. AIFM1 is a component of the mitochondrial disulfide relay that drives complex I assembly through efficient import of NDUFS5. EMBO J. 41, e110784 (2022).
Tranebjærg, L. et al. Neuronal cell death in the visual cortex is a prominent feature of the X-linked recessive mitochondrial deafness-dystonia syndrome caused by mutations in the TIMM8a gene. Ophthalmic Genet. 22, 207–223 (2001).
Roesch, K., Curran, S. P., Tranebjaerg, L. & Koehler, C. M. Human deafness dystonia syndrome is caused by a defect in assembly of the DDP1/TIMM8a–TIMM13 complex. Hum. Mol. Genet. 11, 477–486 (2002).
Koc, E. C., Burkhart, W., Blackburn, K., Moseley, A. & Spremulli, L. L. The small subunit of the mammalian mitochondrial ribosome: identification of the full complement of ribosomal proteins present. J. Biol. Chem. 276, 19363–19374 (2001).
Rival, T. et al. Inner-membrane proteins PMI/TMEM11 regulate mitochondrial morphogenesis independently of the DRP1/MFN fission/fusion pathways. EMBO Rep. 12, 223–230 (2011).
Heydenreich, F. M., Miljuš, T., Milić, D. & Veprintsev, D. B. High-throughput site-directed scanning mutagenesis using a two-fragment PCR approach. Bio. Protoc. 10, 6787 (2020).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Weber, T. A. et al. APOOL is a cardiolipin-binding constituent of the mitofilin/MINOS protein complex determining cristae morphology in mammalian mitochondria. PLoS One 8, e63683 (2013).
McInnes et al. UMAP: Uniform Manifold Approximation and Projection. Journal of Open Source Software 3, 861 (2018).
Ramezani, M. et al. A genome-wide atlas of human cell morphology. Nat. Methods 22, 621–633 (2025).
Cheng, J. et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 2023, eadg7492 (2023).
Jagota, M. et al. Cross-protein transfer learning substantially improves disease variant prediction. Genome Biol. 24, 182 (2023).
Luongo, T. S. et al. SLC25A51 is a mammalian mitochondrial NAD+ transporter. Nature 588, 174–179 (2020).
Mota, M. N., Martins, L. C. & Sá-Correia, I. The identification of genetic determinants of methanol tolerance in yeast suggests differences in methanol and ethanol toxicity mechanisms and candidates for improved methanol tolerance engineering. J. Fungi 7, 90 (2021).
Yano, M., Terada, K. & Mori, M. AIP is a mitochondrial import mediator that binds to both import receptor Tom20 and preproteins. J. Cell Biol. 163, 45–56 (2003).
Doench, J. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Green, M. R. & Sambrook, J. Nested polymerase chain reaction (PCR). Cold Spring Harb. Protoc. 2019, pdb.prot095182 (2019).
Lomize, A. L., Todd, S. C. & Pogozheva, I. D. Spatial arrangement of proteins in planar and curved membranes by PPM 3.0. Protein Sci. 31, 209–220 (2022).
Gilquin, B. et al. The AAA+ ATPase ATAD3A controls mitochondrial dynamics at the interface of the inner and outer membranes. Mol. Cell Biol. 30, 1984–1996 (2010).
Acknowledgements
We would like to thank Caleb Valentine, Mariel Liebeskind, Alex Yenkin, Eileen Xu, Samah Nour, Kylan Kelley, Mallory Wright, Serena Elia, Diana Grigore, Josh Milbrandt, and Josh Langmade for their help with aspects of the project. Jared Zane and Uma Kaushik helped design some of the primers. We would also like to thank the Milbrandt, DiAntonio, and Mitra lab for their continued support. We would like to thank the McDonnell Genome Institute, specifically GESC@MGI helped with CRISPR advice and libraries, GTAC@MGI helped with sequencing, as well as WashU’s CGS. Cell Microsystems was very supportive and quick to lend a hand as needed. Finally, we would like to thank the Genetics and MGI administration and our maintenance and cleaning staff.
Author information
Authors and Affiliations
Contributions
C.L.K. and J.E.W., as first authors, contributed equally to this work. C.L.K., J.E.W., L.K.MA, G.W.B., and W.J.B. wrote the manuscript. W.J.B., J.C.B., J.E.W., and C.L.K. designed the experiments. Initial planning and ideas brought by W.J.B., J.D.M., R.D.M., J.E.W., and C.L.K. C.L.K., J.E.W., G.W.B., L.K.MA, J.C.B., M.A.V., P.P. performed the experiments. P.M.H. and D.P.M. performed protein structure analysis. J.E.W., C.L.K., G.W.B., L.K.MA, P.P., D.P.M. and W.J.B. analyzed the results. W.J.B., V.D.C., G.B. wrote some of the key software. All reviewed the paper and gave suggestions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kremitzki, C., Waligorski, J., Bachman, G. et al. Pathogenic morphological signatures of perturbations in mitochondrial-related genes revealed by pooled imaging assay. npj Imaging 3, 35 (2025). https://doi.org/10.1038/s44303-025-00097-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44303-025-00097-9