Abstract
Renal cell carcinomas (RCCs) have multiple subtypes that are difficult to distinguish using imaging alone. This study characterises renal tumour microstructure using diffusion MRI (dMRI) and the Vascular, Extracellular and Restricted Diffusion for Cytometry in Tumours (VERDICT)-MRI framework. Patients were prospectively recruited from the RIM trial (ClinicalTrials.gov: NCT07173140, 20/11/2024). Fourteen patients with 17 renal tumours (including benign and various RCC subtypes) underwent dMRI using nine b-values (0–2500 s/mm²). A three-compartment VERDICT model was fitted with a self-supervised neural network. Compared to simpler dMRI models, VERDICT more accurately captured the diffusion data in tumour and healthy tissue. VERDICT revealed significant differences in intracellular volume fraction between cancerous and normal tissue, and in vascular volume fraction between vascular and non-vascular regions. A feature selection method identified a reduced 4 b-value protocol (b = [70, 150, 1000, 2000]), cutting scan time by over 30 min, enabling more efficient imaging in larger cohorts.
Similar content being viewed by others
Introduction
Renal cancer accounts for roughly 2% of all cancer diagnoses and deaths worldwide1. Over 90% of primary renal cancers are renal cell carcinoma (RCC), arising from the renal tubule epithelium2. The 2022 WHO classification subdivides RCC into 21 different histotypes of which the commonest are clear-cell RCC (ccRCC) (~70%), papillary RCC (pRCC) (~10%) and chromophobe (chRCC) (~5%)3. Benign tumours of the kidney can also occur, notably oncocytoma and angiomyolipoma4. The main diagnostic imaging for renal tumours is computed tomography (CT), which can diagnose RCC with a sensitivity of over 95% and a specificity of around 90%5. While different tumour subtypes exhibit typical imaging features on both CT and MRI, there is considerable overlap between certain types e.g., oncocytoma, clear cell and chRCC6. With the exception of the identification of “macroscopic fat” in lipid-rich angiomyolipomas, there are currently no imaging biomarkers with sufficient accuracy to differentiate benign and malignant tumours in clinical practice7. A historical reluctance to use biopsy due to the risks, sampling error and a potentially low negative predictive value have meant that rates of benign histology following nephrectomy remain as high as 31%8.
Therefore, new imaging biomarkers are required that are capable of accurately defining both lesion subtype and metastatic risk. MRI techniques offer valuable insights into tumour biology; particularly diffusion-weighted (DW) MRI, which can access restricted diffusion via the apparent diffusion coefficient (ADC). DW-MRI has been used to differentiate RCC subgroups9,10,11 and has proven useful in characterising indeterminate small renal lesions, whether inflammatory or malignant, as both may exhibit restricted diffusion12. However, whilst most clinical DW-MRI studies focus on the ADC, it conflates multiple physiological parameters, reducing its specificity for identifying disease mechanisms, with studies showing varying performance in RCC subtype characterisation13,14,15. The intravoxel incoherent motion (IVIM) model, which separates diffusion in tissue from pseudo-diffusion in blood vessels, has demonstrated enhanced performance by capturing tumour vascularity16,17,18, as flow compartments play a significant role in microstuctural imaging of the kidney19. While IVIM improves upon the uni-compartmental ADC, it still oversimplifies the complex underlying tissue microstructure.
The Vascular, Extracellular and Restricted Diffusion for Cytometry in Tumours (VERDICT)-MRI framework combines a specific diffusion acquisition protocol with a three-compartment biophysical model of the DW-MRI signal in tumour tissue20. Previously, VERDICT has shown effective tumour characterisation in colorectal21, prostate22,23 and brain tumours24. Specifically, in prostate cancer where the problem of lesion over-treatment is similar to that in renal tumours, VERDICT improves differentiation between benign and low-grade malignant lesions compared to the ADC25,26. In this work, we trial VERDICT-MRI for renal tumours for the first time, in a cohort of 14 patients with 17 renal masses. The patients participated in a specific DW acquisition to obtain rich datasets for modelling analysis, and a kidney-specific VERDICT model was identified and fitted to this data using a self-supervised deep learning method27. The acquisition of comprehensive datasets took 46 min; a clinically impractical acquisition time. Traditionally, optimised protocols are identified via the Fisher information, a computationally expensive and model-restricted approach28. We investigated recursive feature elimination methods that have been used previously to identify optimal VERDICT imaging protocols29,30, with a model-free approach adaptable to different tasks.
We found that our VERDICT model for renal tissue outperformed ADC and IVIM at describing the diffusion-weighted data and underlying microstructure. The VERDICT intracellular volume fraction was significantly higher in cancer than normal kidney tissue and revealed similar trends to those predicted by histology, whilst the vascular volume fraction was statistically higher in vascular tumours than in non-vascular. Finally, our dual-network feature selection method identified an economical acquisition protocol for clinical use, reducing the scan time by 32 min and maintaining diagnostic power comparable to the full protocol.
Results
Patient demographics
Fourteen patients with 17 tumours (15 of which had histological confirmation and were included in the analysis) underwent MRI examination. The patients had a mean age of 66 ± 11 years, and a 1:3.6 female:male ratio (Table 1).
Histological validation of the VERDICT model
ccRCC and pRCC demonstrated higher cell density in comparison to chRCC and oncocytoma (Fig. 1). Vascularity, measured by CD34 staining, grouped tumours into a vascular group of tumours (ccRCC, 12.6% positivity; oncocytoma, 10.4%; and chRCC, 7.1%) and non-vascular tumours (pRCC, 2.8%).
We observe that ccRCC and pRCC have a higher number of cells in the same area (180,000 mm2) of tumour in comparison to chRCC and oncocytoma, which agrees with the literature that they are more cellular tumour types.
VERDICT, IVIM and ADC model fitting of normal and cancerous tissue
The VERDICT, IVIM and ADC models were fitted to the data from ROIs in tumour and normal kidney in Fig. 2. In both low and high-grade renal tumours, the mean-squared error (MSE) of the VERDICT fit is lower than ADC and IVIM in both normal and cancerous tissue, suggesting that it describes the data most accurately.
Signal prediction vs DWI data for tumour and normal renal tissue via VERDICT vs. IVIM vs. ADC for a high-grade tumour and benign oncocytoma. The DW signal data are plotted as the points, the curve represents the model’s signal prediction and the MSE is calculated as the difference between the true and predicted value.
VERDICT and ADC parameter estimates
Parametric maps were generated for the VERDICT parameters (intracellular volume fraction; \({f}_{{IC}}\), extracellular-extravascular volume fraction; \({f}_{{EES}}\), vascular volume fraction; \({f}_{{VASC}}\), cell radius; \(R\)) and ADC, for each of the patient groupings (Fig. 3). \({f}_{{IC}}\) and ADC ranged between 0.06 and 0.21 (standard deviation (SD): 0.03) and 2.05 and 2.76 (SD: 0.2), respectively, for benign renal parenchyma across all patients. \({f}_{{IC}}\) discriminated between cancerous from normal tissue (0.32 vs. 0.12; p < 0.01), as did ADC (1.6 vs. 2.3; p < 0.05) (Fig. 5a, b). The \({f}_{{IC}}\) for oncocytomas (benign) was not significantly different to normal tissue (0.17 vs. 0.12; p = n.s.). There were trends towards higher \({f}_{{IC}}\) and \(R\) in the high-grade uRCC (\({f}_{{IC}}\) = 0.59) and pRCC (\({f}_{{IC}}\) = 0.55) compared to low grade tumours (\({f}_{{IC}}\) = 0.21 ± 0.07) (Figs. 3 and 4). A similar trend was seen with ADC.
We observe higher and for more cellular tumours, (uRCC, pRCC and ccRCC), and elevated in more vascular masses (ccRCC, oncocytoma). ADC is lower for uRCC, pRCC and chRCC, but not for the other groups.
A VERDICT fIC and B ADC in normal kidney tissue, benign oncocytomas and cancerous tumours. The fIC can discriminate cancer from normal tissue (p < 0.01), as can ADC with p < 0.05. C VERDICT fVASC and D IVIM fIVIM in vascular vs. non-vascular tumours – the fVASC can discriminate these (p < 0.05).
The tumours were grouped into vascular (ccRCC, oncocytoma and chromophobe tumours), and non-vascular (uRCC, pRCC) based on the unpaired histological analysis and degree of enhancement demonstrated on nephrographic-phase CT for each patient.
The \({f}_{{VASC}}\) was significantly higher in the vascular tumours than in non-vascular (mean ± SD: 0.29 ± 0.05 vs. 0.13 ± 0.02, p < 0.05; Fig. 5d), but there was no significance for \(f\)(0.33 ± 0.07 vs. 0.23 ± 0.03, p = n.s.; Fig. 5c).
Deep learning protocol optimisation
The deep learning feature selection method produced a protocol of four optimally-informative b-values: 70, 150, 1000, and 2000 s/mm2. In Fig. 5, we reproduce our results with this protocol, demonstrating the quality of the parameter maps is largely preserved, with some smoothing. We plot the differences between these maps and the original and find that they are generally small with more variation outside of the tumour. We also find the MSE between the signal predictions and DW data remains similar to Fig. 2 in a benign and cancerous tumour. In Supplementary Fig. 3, we recreate Fig. 4 with the optimised protocol, finding similar statistical significance to the full protocol. The optimised protocol has an acquisition time of ~14 min, a reduction of over 30 min from the full protocol.
In (A) we show the maps produced using the identified shorter protocol, and in (B) we plot the difference between these maps and the original. We observe that they maintain the map features with some smoothing, and more variation is seen outside the tumour region. In (C) we plot the signal prediction vs. DWI data for a high-grade tumour and benign oncocytoma, observing that the MSE between the data and predictions remains similar to that seen with the full protocol.
Discussion
In this work, we create a model for renal tumours within the VERDICT-MRI framework for the first time, in a cohort of 14 patients. We acquire rich datasets of nine b-values to develop and fit a three-compartment VERDICT model for the kidney using self-supervised deep learning. We show that VERDICT provides a better description of the data than IVIM and ADC and obtain statistically significant differences between tissue types in the intracellular and vascular volume fractions. Finally, we find that our results hold when using a significantly shortened protocol of four b-values, demonstrating that this technique can be translated to the clinic with ease.
In Fig. 2, we show that the predicted signal from VERDICT matches the DW data more closely in terms of MSE than IVIM or ADC in both tumour and benign tissue. This suggests VERDICT provides a better description of the underlying microstructure, agreeing with previous studies where VERDICT outperforms both models in the prostate20,23. We note the MSE is lower in tumour regions than benign; we hypothesise that this is because the model is developed to describe the DW signal in tumour tissue specifically, therefore describes those features more accurately.
Histologically, ccRCC and pRCC demonstrated higher cellularity than chRCC and benign oncocytoma (Fig. 1), while chRCC and oncocytoma appear similar, explaining the challenge of differentiating these tumours on imaging. This also agrees with our results in Supplementary Fig. 2, where we note that for \({f}_{{IC}}\), ADC and \(D\), our results overlap considerably for chRCC and oncocytoma. The higher \({f}_{{IC}}\) estimates in cancerous tumours, particularly the high-grade tumours, compared to the normal renal parenchyma agrees with prior knowledge that cancerous tumours are typically hypercellular31. Benign oncocytomas could not be discriminated from either normal tissue or cancer at a statistically significant level with either or ADC however, highlighting their confounding nature.
The unique microstructural information available via VERDICT compared to IVIM and ADC permitted comparison of tumour vascularity. Informed by our prior knowledge from CT imaging we grouped tumours into a vascular and non-vascular group based on their CT-enhancement characteristics and showed the \({f}_{{VASC}}\) to be significantly higher in the vascular tumours group. Inferring enhancement from DWI and VERDICT could potentially avoid contrast administration clinically, and further studies into diffusion time dependence could lead to the extraction of specific microstructural measurements32.
Lastly, in Fig. 5 we recreate our results with the optimised protocol identified via deep-learning-based feature selection and show consistency with those achieved with the full protocol. Along with Supplementary Fig. 3, these results demonstrate that similar diagnostic performance can be achieved whilst reducing the acquisition time by over 30 min, enabling the clinical translation of this work. This result also motivates the development of VERDICT models for other organs, as we demonstrate how the use of a few comprehensive datasets for model development can be used to identify clinical acquisitions.
This work is limited primarily by the small patient cohort, resulting in few patients per tumour subtype, which limits the statistical significance of these results. This is in part due to the long acquisition protocol; with the identified economical protocol, we hope to soon be able to trial VERDICT on a much larger number of kidney patients with a variety of subtypes. Additionally, whilst some histological analysis has been conducted, these are not paired samples with the imaging data, meaning that the imaging biomarkers cannot be fully validated histologically. Future studies will include paired histology and DWI for more in-depth analysis, such as quantitative correlations between fIC and cellularity and fVASC and CD34-based vascularity. We also note some field inhomogeneities in the images due to the EPI sequence used; future work will investigate distortion correction methods33. Finally, further information about vascularity could be extracted via the separation of vascularity into two distinct contributors – perfusion and tubular flow – or incorporation of spatial profiling of the renal parenchyma34,35,36,37. Such studies propose significant extensions to our current model, therefore we leave this avenue for future work.
In conclusion, this work develops a VERDICT-MRI model for renal tumours, and analyses results in a cohort of 14 patients. We show that VERDICT provides better data description and enhanced microstructural information over ADC, such as differences between patient groups in intracellular and vascular volume fractions. We exploit novel deep learning advances for both the model fitting approach to ensure robustness, and to identify a significantly shorter acquisition protocol. We find that we can extract equivalent diagnostic performance with our abbreviated protocol as with the original, allowing for smooth clinical translation of the technique.
Methods
Patient cohort
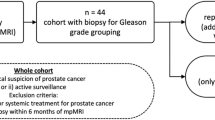
The RIM study (ClinicalTrials.gov: NCT07173140, 20/11/2024) was approved by the UCL Research Ethics Committee (07/Q0502/15) and all patients provided informed written consent. Patients were identified at the Royal Free Hospital Renal Cancer Specialist Multi-Disciplinary Team Meeting (SMDT). Patients were recruited prospectively and were eligible to participate if they had a diagnosis of a T1 solid renal tumour made on standard of care imaging, were able to have an MRI and had not had a recent biopsy within the last three months. Histology from biopsy or surgical histology was available for all patients, performed as standard-of-care.
VERDICT-MRI acquisition
VERDICT-MRI was performed on a 3T MRI system (Ingenia; Philips, Best, Netherlands), using a 16-channel body coil (SENSE XL Torso, Philips, Best, Netherlands), using a pulsed gradient spin-echo (PGSE) sequence with echo-planar imaging (EPI) readout in the coronal imaging plane. The acquisition was informed by the optimised protocol for prostate38, with additional measurements to obtain rich datasets for analysis and emphasis placed on lower b-values to reflect the vascularity of renal microstructure. The imaging parameters were as follows: repetition time (TR), 2000–3349 ms; field of view, 220 × 200 mm; voxel size, 1.25 × 1.25 × 5 mm; no interslice gap; acquisition matrix, 176 ×176. The VERDICT acquisition protocol for kidney is: b = [70, 90, 150, 500, 1000, 1500, 2000, 2200, 2500] s/mm2; δ = [4.8, 4.8, 4.8, 12.0, 12.0, 26.3, 16.8, 16.8, 21.4] ms; Δ = [27.0, 27.0, 27.0, 34.0, 34.0, 47.0, 37.5, 37.5, 43.5] ms. For each of the nine combinations of b/δ/Δ, we used the minimum possible echo time (TE), giving TEs of 54–87 ms. For each b-value, images were acquired in three orthogonal directions which were then spherically-averaged, and we acquired a separate image for each TE, resulting in 18 image volumes each with 14 slices. The acquisition time was approximately 46 min.
Image analysis
The preprocessing pipeline included denoising using Marchenko–Pastur Principal Component Analysis (MP-PCA)39 (MrTrix3 ‘dwidenoise’40), and correction for Gibbs ringing41. We applied mutual-information rigid and affine registration42 and divided DW-MRI volumes by their matched b = 0 for normalisation. Regions of interest (ROIs) for tumour, simple cysts and normal renal parenchyma were drawn manually in ITKSnap v4.0 using co-registered T2-weighted images by a radiologist in-training with seven years’ experience in abdominal imaging43. Histological subtype and tumour WHO/ISUP (World Health Organisation/International Society of Urological Pathology) grade was determined from either lesion biopsy or surgery and compared to imaging tumour analysis. 10 unpaired surgical histological samples were of the main tumour types (ccRCC, pRCC, chRCC and oncocytoma) were stained with haematoxylin-and-eosin (H&E) and CD34 (a vascular endothelial marker) immunohistochemistry. Slides were scanned and the digital images analysed using the software QuPath44. Cell density and vascularity were used as histological validation of the VERDICT model.
VERDICT model
The VERDICT model for kidney has three compartments that characterise the diffusion of water molecules in the intracellular (IC), vascular (VASC) and extracellular-extravascular space (EES) in tumours, with no exchange between the components20,21.
The total normalised signal is:
where \({f}_{i}\) is the volume fraction and \({S}_{i}\) is the normalised signal from water molecules in population \(i\), where \(i=\)IC, VASC or EES. The vascular signal fraction, \({f}_{{VASC}}\), is computed as \(1-{f}_{{IC}}-{f}_{{EES}}\), since \(0\le {f}_{i}\le 1\) and \({\sum }_{i=0}^{1}{f}_{i}=1\), and \({S}_{0}\) is the signal with no diffusion weighting. Here \(b\) is the b-value, \(\Delta\) is the gradient pulse separation and \(\delta\) is the gradient pulse duration.
To represent renal tissue, the IC compartment was modelled as an impermeable sphere of radius \(R\) with IC diffusivity fixed at \({d}_{{IC}}=2\)µm2/ms and the EES compartment as Gaussian free diffusion with diffusivity \({d}_{{EES}}=2\)µm2/ms. The vascular compartment was randomly-oriented sticks with pseudo-diffusivity \({d}_{{VASC}}=50\)µm2/ms to reflect the high vascularity of kidney tissue (see Supplementary Fig. 1). Three free parameters were estimated: \({f}_{{IC}}\), \({f}_{{EES}}\) and \(R\) within biophysically-realistic parameter ranges: [0, 1], [0, 1], [0, 15 µm].
Model fitting
The renal VERDICT model is fit to the data using self-supervised deep learning, previously used in ref. 27 with the prostate model. These techniques remove the need for training data, instead extracting labels from the input data itself. They work to estimate the model parameters and reconstruct the signal, then calculate the training loss as the MSE between the predicted signal, S^, and the input signal S45 to find the optimal estimates. A fully connected neural network with three hidden layers, each with 18 neurons (i.e. the number of image volumes, a spherically-averaged b-value image with a corresponding b0), was implemented using PyTorch 1.12.1. The output layer is fed into the VERDICT model equation to generate the predicted signal S^. We used the ADAM optimiser, early stopping to mitigate overfitting, and a learning rate of 0.0001. We used dropout of p = 0.5 and constrained the parameter values to the ranges above using the PyTorch clamp function. Training and inference are performed simultaneously and took about 50 s for each masked dMRI data set.
Protocol optimisation
The protocol mentioned above provides rich DW datasets useful for model development and analysis, however the acquisition time of 46 min is not clinically feasible. We develop a dual-network deep learning approach based on similar work in literature29,30,46 to select the most informative subset from the full protocol of nine b-values.
Features were selected from 27 measurements from the DW images acquired along the three orthogonal directions per b-value. The non-DW measurements were excluded to prevent them from being chosen, as they would be included in the protocol with their matched DW. Each DW dataset was normalised and fed into the first network for feature selection. Each dataset was normalised and input into a Feature Scoring Network – a fully connected model with 27 input nodes, one hidden layer with 64 units (using ReLU activation and batch normalisation), and a 27-node output layer with a sigmoid activation. This network produced importance scores in the range [0,1] for each measurement of a dataset. The 12 highest-scoring features were selected for each dataset and passed into a Prediction Network with 12 input nodes, a hidden layer of 64 units (ReLU activation), and an output layer of 27 nodes, which aimed to reconstruct the full measurement set from the chosen subset of 12 measurements.
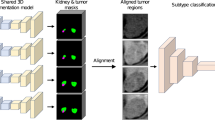
The two networks were trained in tandem for 100 epochs and the loss was computed as the MSE between the reconstructed full measurement and ground truth target values (the input 27 measurements). A learning rate of 1e−5 was used, with the ADAM optimiser and dropout to prevent overfitting. Upon training completion, we extracted the final optimal feature set by computing the average importance scores across all subjects and extracting a final protocol of 4 b-values (12 measurements in 3 directions). The method is outlined in Fig. 6.
We first remove the non-DW images from the image volumes, resulting in 27 measurements per patient. Our feature scoring network takes in the 27 measurements (27 input nodes), has hidden layers with 64 nodes and outputs 27 scores for the measurements. From these, the 12 measurements with the highest scores are selected. These are then used to reconstruct the target initial 27 measurements in the predictor network, and the loss is computed as the MSE between the output and the target. We evaluate on our full patient dataset, choosing b = 70, 150, 1000, 2000 as our final protocol. This gives a reduction in acquisition time of 32 min.
Statistical analysis
Data were analysed using Python 3.11 (SciPy 1.15.047) and statistical tests performed were Wilcoxon’s signed-rank tests with errors reported as standard deviation, unless stated otherwise. p-values are summarised in figures as: <0.0001, ****; 0.0001–0.001, ***; 0.001–0.01, **; 0.01–0.05, *.
Data availability
The datasets generated and/or analysed during the current study are not publicly available as participants in the pilot study consented to data use for research purposes, but not to public online sharing. Data are available from the corresponding author on reasonable request, in line with ethical guidelines.
Code availability
The underlying code for this study [and training/validation datasets] is not publicly available but may be made available to qualified researchers on reasonable request from the corresponding author.
References
Padala, S. A. et al. Epidemiology of renal cell carcinoma. World J. Oncol. 11, 79–87 (2020).
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Prim. 3, 17009 (2017).
Lobo, J. et al. WHO 2022 landscape of papillary and chromophobe renal cell carcinoma. Histopathology 81, 426–438 (2022).
Tamboli, P., Ro, J. Y., Amin, M. B., Ligato, S. & Ayala, A. G. Benign tumors and tumor-like lesions of the adult kidney. Part II: Benign mesenchymal and mixed neoplasms, and tumor-like lesions. Adv. Anat. Pathol. 7, 47–66 (2000).
Morshid, A., Duran, E. S., Choi, W. J. & Duran, C. A concise review of the multimodality imaging features of renal cell carcinoma. Cureus 13, e13231 (2021).
Metin, M., Aydın, H. & Karaoğlanoğlu, M. Renal cell carcinoma or oncocytoma? The contribution of diffusion-weighted magnetic resonance imaging to the differential diagnosis of renal masses. Medicine 58, 221 (2022).
Bellin, M. F. et al. Update on renal cell carcinoma diagnosis with novel imaging approaches. Cancers 16, 1926 (2024).
Kim, J. H. et al. Association of prevalence of benign pathologic findings after partial nephrectomy with preoperative imaging patterns in the United States from 2007 to 2014. JAMA Surg. 154, 225–231 (2019).
Lei, Y. et al. Diagnostic significance of diffusion-weighted MRI in renal cancer. Biomed. Res. Int. 172165 (2015).
Cova, M. et al. Diffusion-weighted MRI in the evaluation of renal lesions: preliminary results. Br. J. Radiol. 77, 51–857 (2004).
Sandrasegaran, K. et al. Usefulness of diffusion-weighted imaging in the evaluation of renal masses. AJR Am. J. Roentgenol. 194, 438–445 (2010).
Doğanay, S. et al. Ability and utility of diffusion-weighted MRI with different b values in the evaluation of benign and malignant renal lesions. Clin. Radiol. 66, 420–425 (2011).
de Silva, S. et al. The diagnostic utility of diffusion weighted MRI imaging and ADC ratio to distinguish benign from malignant renal masses: sorting the kittens from the tigers. BMC Urol. 21, 67 (2021).
Paudyal, B. et al. The role of the ADC value in the characterisation of renal carcinoma by diffusion-weighted MRI. Br. J. Radiol. 83, 336–343 (2010).
Hötker, A. M. et al. Use of DWI in the differentiation of renal cortical tumors. AJR Am. J. Roentgenol. 206, 100–105 (2016).
Mahmoud, A. R. et al. Role of intravoxel incoherent motion diffusion-weighted MRI in differentiation of renal cell carcinoma subtypes. Egypt J. Radiol. Nucl. Med. 55, 184 (2024).
Ding, Y. et al. Differentiating between malignant and benign renal tumors: do IVIM and diffusion kurtosis imaging perform better than DWI?. Eur. Radiol. 29, 6930–6939 (2019).
Ding, Y. et al. Comparison of biexponential and monoexponential model of diffusion-weighted imaging for distinguishing between common renal cell carcinoma and fat poor angiomyolipoma. Korean J. Radiol. 17, 853–863 (2016).
Liu, A. L. et al. REnal Flow and Microstructure AnisotroPy (REFMAP) MRI in normal and peritumoral renal tissue. J. Magn. Reson. Imaging 48, 188–197 (2018).
Panagiotaki, E. et al. Microstructural characterization of normal and malignant human prostate tissue with vascular, extracellular, and restricted diffusion for cytometry in tumours magnetic resonance imaging. Invest. Radiol. 50, 218–227 (2015).
Panagiotaki, E. et al. Noninvasive quantification of solid tumor microstructure using VERDICT MRI. Cancer Res. 74, 1902–1912 (2014).
Palombo, M. et al. Joint estimation of relaxation and diffusion tissue parameters for prostate cancer with relaxation-VERDICT MRI. Sci. Rep. 13, 1 (2023).
Sen, S. et al. Differentiating false positive lesions from clinically significant cancer and normal prostate tissue using VERDICT MRI and other diffusion models. Diagnostics 12, 1631 (2022).
Figini, M. et al. Comprehensive brain tumour characterisation with VERDICT-MRI: evaluation of cellular and vascular measures validated by histology. Cancers 15, 2490 (2023).
Singh, S. et al. Avoiding unnecessary biopsy after multiparametric prostate MRI with VERDICT analysis: the INNOVATE study. Radiology 305, 623–630 (2022).
Johnston, E. W. et al. VERDICT MRI for prostate cancer: intracellular volume fraction versus apparent diffusion coefficient. Radiology 291, 391–397 (2019).
Sen, S. et al. ssVERDICT: self-supervised VERDICT-MRI for enhanced prostate tumor characterization. Magn. Reson. Med. 92, 2181–2192 (2024).
Alexander, D. C. A general framework for experiment design in diffusion mri and its application in measuring direct tissue-microstructure features. Magn. Reson. Med. 60, 439–448 (2008).
Grussu, F. et al. Feasibility of data-driven, model-free quantitative MRI protocol design: application to brain and prostate diffusion-relaxation imaging. Front. Phys. 9, 752208 (2021).
Blumberg, S. B., Slator, P. J. & Alexander, D. C. Experimental design for multi-channel imaging via task-driven feature selection. In The Twelfth International Conference on Learning Representations (ICLR, 2024).
Wang, X. et al. Pathologic analysis of non-neoplastic parenchyma in renal cell carcinoma: a comprehensive observation in radical nephrectomy specimens. BMC Cancer 17, (2017).
Sigmund, E. E. et al. Cardiac phase and flow compensation effects on REnal Flow and Microstructure AnisotroPy MRI in healthy human kidney. J. Magn. Reson. Imaging 58, 210–220 (2023).
Gilani, N. ima et al. Characterization of motion dependent magnetic field inhomogeneity for DWI in the kidneys. Magn. Reson. Imaging 100, 93–101 (2023).
Stabinska, J. et al. Investigation of diffusion time dependence of apparent diffusion coefficient and intravoxel incoherent motion parameters in the human kidney. Magn. Reson. Med. 93, 2020–2028 (2025).
Stabinska, J. et al. Spectral diffusion analysis of kidney intravoxel incoherent motion MRI in healthy volunteers and patients with renal pathologies. Magn. Reson Med 85, 3085–3095 (2021).
Liu, M. M. et al. Estimation of multicomponent flow in the kidney with multi-b-value spectral diffusion. Magn. Reson. Med. 94(6), 2550–2566 (2025).
Kazerouni, A. S. et al. Quantifying tumor heterogeneity via MRI habitats to characterize microenvironmental alterations in HER2+ breast cancer. Cancers 14, 1837 (2022).
Panagiotaki, E. et al. Optimised verdict mri protocol for prostate cancer characterisation. In Proceedings of the International Society for Magnetic Resonance in Medicine (ISMRM), 2872 (ISMRM, 2015).
Veraart, J., Fieremans, E. & Novikov, D. S. Diffusion MRI noise mapping using random matrix theory. Magn. Reson. Med. 76, 1582–1593 (2016).
Tournier, J. D. et al. MRtrix3: a fast, flexible and open software framework for medical image processing and visualisation. Neuroimage 202, 116137 (2019).
Kellner, E., Dhital, B., Kiselev, V. G. & Reisert, M. Gibbs-ringing artifact removal based on local subvoxel-shifts. Magn. Reson. Med. 76, 1574–1581 (2015).
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23, S208–S219 (2004).
Yushkevich, P. A. et al. User-guided 3d active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. Neuroimage 31, 1116–1128 (2006).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Barbieri, S., Gurney-Champion, O. J., Klaassen, R. & Thoeny, H. C. Deep learning how to fit an intravoxel incoherent motion model to diffusion-weighted MRI. Magn. Reson. Med. 83, 312–321 (2020).
Blumberg, S. B. et al. Progressive subsampling for oversampled data - application to quantitative MRI. In MICCAI 421–431 (Springer Nature, 2022).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Acknowledgements
S.S. was supported by the EPSRC-funded UCL Center for Doctoral Training in Intelligent, Integrated Imaging in Healthcare (i4health) (EP/S021930/1) and the Department of Health's NIHR-funded Biomedical Research Centre at University College London Hospitals. R.L.H. was funded by NIHR UCLH BRC grants BRC1125/HEI/RH/110410 and BRC928/CN/RH/101330. E.P. was supported by EPSRC grants EP/N021967/1, EP/R006032/1.
Author information
Authors and Affiliations
Contributions
Conceptualisation was carried out by R.L.H. and E.P. Method development was done by S.S. and E.P., and data analysis was carried out by S.S. Data collection and curation was done by L.S., L.C., J.C., M.T., S.P. and R.L.H. Writing of the manuscript was led by S.S., with review/editing by D.A., R.L.H. and E.P. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sen, S., Smith, L., Caselton, L. et al. Dual deep learning approach for non-invasive renal tumour subtyping with VERDICT-MRI. npj Imaging 4, 2 (2026). https://doi.org/10.1038/s44303-025-00135-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44303-025-00135-6