Abstract
Interbreeding between modern humans and archaic hominins, including Neanderthals and Denisovans, occurred as modern humans migrated outside of Africa. Here, we report on evidence of adaptive introgression from archaic hominins within genomic regions associated with circadian rhythm cycling, chronotype, and sleep using 76 worldwide modern human populations from the Human Genome Diversity Project and 1000 Genomes Project. We identified 265 independent segments suggestive of adaptive introgression, where 22 of these segments show evidence of positive selection. We tested for evidence of a latitudinal cline within 35 core haplotypes, finding no clear latitude gradient, and identified the likely archaic donor for each of these haplotypes. We found that several genes with evidence of adaptive introgression are associated with affective disorders, chronotype, and respiratory diseases. Lastly, many of the variants are eQTLs for several genes that are significantly enriched in immunity pathways.
Similar content being viewed by others
Introduction
A little over a decade ago, human evolutionary history was completely reshaped by the discovery that anatomically modern humans interbred with our archaic cousins, the Neanderthals1, and their enigmatic sister species, the Denisovans2. These were not isolated events, and evidence has been uncovered that there were several admixture events occurring sporadically over thousands of years and across diverse geographic areas3,4,5,6,7,8,9. Signatures of these events are left in our genome with estimates that every non-African alive today has on average just below 2% of their DNA shared with Neanderthals10. Denisovan signatures in modern humans are generally lower, on average below 1%11,12. However, some Oceanic populations have been noted to have nearly 5% of their DNA composed of Denisovan-introgressed regions2,8,13.
A number of these archaic introgressed regions are believed to be adaptive and have been brought to elevated frequencies in modern human populations. Several of the most notable examples are variants within the EPAS1 gene in Tibetan populations that confer adaptation to high altitude environments derived from Denisovans14, immunity and HLA-controlling regions likely giving rise to disease resistance to modern humans as they expanded into new territories after leaving Africa15,16,17,18, and the various skin, hair, and keratin linked regions introgressed from Neanderthals that have been highlighted in a number of studies4,19,20,21. Recently, the discovery of introgressed variants within genes involved in circadian rhythm and chronotype expression has become a new area of focus20,21,22,23,24. Single nucleotide polymorphisms (SNPs) identified to be associated with chronotype and several other sleep-associated traits such as daytime napping, narcolepsy, willingness to get up in the morning, and sleep duration have been suggested to be the product of admixture from Neanderthals20,23. In a meta-analysis using previously published genome-wide association study (GWAS) data identifying archaic introgressed loci, genetic variants associated with being a morning person were shown to be shared with the Altai Neanderthal21. Recent analysis using a combination of previously published archaically-introgressed variants found that modern humans and archaic hominins differed in their circadian rhythm genes, including alternative splicing events and regulatory divergence24.
The circadian rhythm is the cyclic oscillator of a 24-h period, which has remained relatively conserved across most of the animal kingdom25 and has been proposed to be a core controller of sleep and wake cycles26,27,28,29,30. To do this, the suprachiasmatic nucleus, located in the hypothalamus, uses external light stimuli to reset itself along a day-night cycle29. Output from the suprachiasmatic nucleus goes to the ventral subparaventricular zone and regulates this information into daily cycles of wakefulness and sleep25,29, that then falls across the natural 24-h circadian rhythm. In addition to controlling sleep and wake cycles, several review articles have highlighted the link between circadian rhythm and gastrointestinal processes31,32 and immune function33,34,35. The preference for how late someone stays awake is influenced by chronotype, that is, morning people who tend to go to bed and rise earlier, and those who show an evening preference and go to bed and rise later30,36. Three different GWAS datasets have independently identified four genes that support associations with chronotype: PER2, RGS16, FBXL13, and AK530. Additional studies increased the number of chronotype-linked variants which were associated with previously unidentified genes, such as PER1, CRY1, ARNTL37, and ARL14EP38.
For this study we used the gnomAD 1000 Genomes Project and Human Genome Diversity Project (1KGP + HGDP) phased call set39 to identify regions of archaic introgression in a worldwide population dataset (n = 76 populations). Specifically, we used SPrime7 to identify archaic segments in these populations with an interest in introgressed markers within genes showing circadian oscillations or variants previously associated with chronotype and sleep phenotype expression. We utilised four high coverage archaic genomes, including Denisova 32,40, the Altai Neanderthal3, the Vindija Neanderthal41, and the Chagyrskaya Neanderthal42 to accomplish this, and to also see whether different archaic populations may have been the donor population to core haplotypes found in our data.
Our main research goal for this study was to determine whether any variants that overlapped with genes that displayed a circadian cycling component, or variants that have been previously described to influence chronotype or sleep, show evidence of adaptive introgression in modern humans. Adaptive introgression is the increase in fitness of a population due to the introduction of novel loci after an admixture event. Our first hypothesis is that like prior studies, evidence of a latitude cline would be evident in genes or variants that are connected to circadian cycling or chronotype22,23,24,43,44. Archaic populations lived in various environments, some of which were higher latitude2,45,46,47. We also hypothesise that because these archaic populations were adapted to these environments that are typically characterised by strong seasonality and annual differences in photoperiod, some of the archaic variants influencing circadian rhythmicity could have increased in frequency in modern human populations due to natural selection as they also moved to high latitude environments. It is possible that higher latitude variants are likely associated with more versatile circadian cycling to combat the extreme seasonal changes in light availability. We further predict that these archaic groups will be associated with a higher prevalence of variants that enhance serotonin synthesis, such as within the DDC gene48, supporting adaptations to strong seasonal variations in light.
Results
Introgressed regions
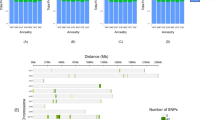
Our analysis was able to recover a large number of variants reported in prior studies, showing the effectiveness of our pipeline, as outlined in the Supplementary Material. In total, we recovered 64,834 putative archaic variants overlapping the gene coordinates listed in the Circadian Genome Database (CGDB)49 and 71 introgressed variants that were also found in the chronotype and sleep-associated trait data from the NHGRI-EBI GWAS Catalogue50. We identified 265 independent, non-overlapping segments in our 62 non-African populations (Table S1), where each segment overlaps at least one variant or window associated with circadian rhythm, chronotype, or a sleep phenotype as described by a GWAS study, or overlaps a gene coordinate listed in the CGDB, and the variant has an atypically high archaic allele frequency of ≥40% in at least one population within our analysis. Within the Americas, there are 127 segments that pass this criterion, 123 in East Asia, 44 in Europe, 12 in the Middle East, 88 in Oceania, and 16 in South Asia (Table S1). We were also able to recover from these segments previously documented introgression patterns, including both the Chagyrskaya and Vindija Neanderthals being more closely related to the introgressed Neanderthal DNA in modern humans41,42, and higher levels of Denisovan ancestry in Oceanic populations relative to other modern human groups2 (Fig. 1A–D). Additionally, we mapped the density of the SNPs from these 265 independent segments across the genome, which can be seen in Fig. 1E. Table S2 includes all variants with archaic allele frequencies ≥40%, highlighting a total of 1729 variants that are found within 303 genes and intergenic regions.
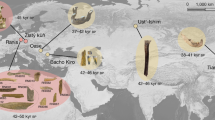
Putative archaic variants recovered in our pipeline overlapping the 265 independent segments provided in Table S1. Colours denote map_arch labelling51 and are representing the percentage of total recovered variants for the. A Altai Neanderthal, B Chagyrskaya Neanderthal, C Vindija Neanderthal, and D Denisovan. The population codes refer to: African (AFR), American (AMR), East Asian (EAS), European (EUR), Middle Eastern (EAS), Oceanic (OCE), and South Asian (SAS). The plots were generated using the ggplot2 package136 in R116. The density of SNPs for each of the 265 independent segments is shown in E, with low density plotted in green and high density in red. The plot was generated using rMVP137. The chromosome lengths are limited to the largest SNP position in our analysis found per chromosome.
Core haplotypes and evidence of positive selection
We further filtered Table S2 to search for core haplotypes, and the results of this can be seen in Table S3_A, comprising 35 regions of interest for further exploration. After running our tests of selection, 22 (~62%) had multiple tests showing evidence of selection, with the highest scoring region being the SUSD1 (sushi domain containing 1) core haplotype in the Melanesians with four tests highlighting selection (Table S3_A). Additionally, JAK1 (Janus kinase 1) in the Melanesians, DNAAF10 (dynein axonemal assembly factor 10) in the Chinese Dai in Xishuangbanna, China (CDX), RPSAP11-ENSG00000261572 (ribosomal protein SA pseudogene 11) in the Papuans, TSPAN11 (tetraspanin 11) in the Kinh in Ho Chi Minh City, Vietnam (KHV), and ENSG00000258312 in the Peruvian in Lima Peru (PEL) all had three tests showing potential evidence of selection (Table S3_A). There were 8 segments that had at least four tests suggesting positive selection within the general introgressed segment, but outside of the core haplotype (Table S3_A). We note, however, that a number of these introgressed segments are quite large (>1 megabase (Mb) in size) and the likely target area of selection falls outside of our identified core haplotypes. For this reason, we have opted to only discuss in detail results where the evidence of selection falls within our identified core haplotype regions. The results from our tests of positive selection can be seen in Tables S3_A-S3_I for our core haplotypes.
We generated a haplotype network, a plot of genetic relationships between populations and associated frequencies of specific haplotypes, and ancestral recombination graphs, coalescence-based genealogies for mutations and recombination events leading to the diversity seen in modern samples at specific loci, for SUSD1 and present this in Fig. 2. As expected, we can see a number of Oceanic populations clustering with the Neanderthal haplotypes in this region, complementing our identification of the core haplotype for this gene being found with the highest frequencies in Oceanic populations (Tables S2 and S3). Our results also support our contour plot51 (Fig. 2C), which visualises the degree of sharing of the core haplotype segment to each archaic population in our analysis. Here, all of the Neanderthal samples have a very high degree of sharing to the SUSD1 core haplotype region in the Melanesians, with match/mismatch ratios over 90%, further illustrating that our haplotype network is displaying accurate relationships. Lastly, our ancestral recombination graph supports our findings of the archaic origins of the SUSD1 core haplotype, where we have an archaic-like branching event signified by a derived mutation shared with archaic populations originating around 1,000,000 years ago followed by a long, non-recombining branch (Fig. 2B).
The A SUSD1 haplotype network was generated using PopArt v1.7126 using the median joining function for the top 50 haplotypes and their ties, the archaic haplotypes, and Oceanic haplotypes with frequencies ≥2. The number of mutations along each edge between nodes is shown in brackets while the archaic haplotypes are labelled with arrows. The networks demonstrate that the haplotypes within this region largely cluster by genetic and geographic similarity and relationships can be attributed to similar levels of archaic ancestry. B Ancestral recombination graph for chr9:114830551 (rs72762055) in the Oceanic populations (Melanesians and Papuans) relative to the Yoruba in Ibadan, Nigeria (YRI), which displays an archaic-like, derived branching event followed by rapid expansion through the Oceanic groups. The ancestral recombination graph was generated using Relate54. C Contour plot of the SPrime7 segment where the SUSD1 core haplotype was found is shown as a white crosshair. The heat map represents the number of genome-wide segments at each match/mismatch ratio for the high-coverage archaic samples in our analysis found in the Melanesians. The segment is clearly introgressed from Neanderthals with very low matching to the Denisova 3 sample, but all of the Neanderthals have a match/mismatch ratio over 0.9 showing no one sample is the putative donor. The plot was generated using the kde2d function from MASS package120 in R116 using modified scripts provided by Zhou & Browning51.
In total, we ran 10 tests of selection using a combination of haplotype-specific, per-site, and windowed analyses to best understand if any of our core haplotypes displayed evidence of selection. The most sensitive statistic to signal selection was extended haplotype homozygosity (EHH)52, showing that 28 archaic core haplotype main alleles displayed longer stretches of homozygosity relative to the non-archaic allele (Table S3_B; Fig. 3B). High FST values53 highlighted that 26 core haplotypes were extremely genetically differentiated relative to the mean FST across the genome. The FST values of the core haplotype main alleles averaged over 9.2 times higher than the genome-wide mean FST, and the mean core haplotype FST was nearly 3 times higher relative to the genome-wide mean (Table S3_C). Relate’s54 selection statistic illustrated that 6 core haplotype segments contained at least one variant that was at the top 1% of its population’s empirical distribution (Table S3_F). The number of segregating sites by length (nSL)55 normalised per-site values indicated only two core haplotype main alleles, one from SUSD1 and the other from ENSG00000289087, both in Oceanic populations, that were at the top 1% of their genome-wide distributions (Table S3_D), while RAiSD56 identified only one core haplotype segment, CEACAM1-LIPE-AS1 (CEA cell adhesion molecule 1; LIPE antisense RNA 1) in the Papuans, as having variants at the top 1% of their distribution (Table S3_E). The other three statistics we explored, saltiLASSI57, Tajima’s D58, and cross-population number of segregating sites by length (XP-nSL)59, along with the normalised windowed nSL analysis, did not show any evidence of selection in the core haplotypes (Table S3_A). To test our approach for accuracy, we applied our methodology to known instances of selection in the human genome and readily identified these segments in our protocol (Supplementary Material). Therefore, we suggest that the same signals can be freely applied to our core haplotypes and generalised introgressed segments, where multiple lines of evidence are clear indications of selection.
The A SUSD1 main core haplotype allele, chr9:114810102 (rs10981228, shown as a red diamond), 12 variants within the core haplotype (green dots), and several additional variants in the SPrime7 located archaic segment (blue dots) all showing evidence of selection using the normalised nSL statistic55, where each of these variants were at the top 1% of the genome-wide distribution (red dashed) within the Melanesians. While they are not the highest scoring variants along this locus, it is clear that the high number of variants surpassing the genome-wide threshold suggests that the archaic haplotype is likely being selected for in this population. The plot was generated using the locuszoomr package138 in R116. B EHH52 decay curve for the archaic allele (red) and non-archaic allele (blue) for a 300,000 bp window around rs10981228. The archaic, derived allele has a considerably larger homozygous region compared to the non-archaic allele, highlighting probable selection acting on the archaic allele. C Rapid increase in allele frequency suggesting selection is acting on rs10981228. Both EHH and allele frequency plots were generated using ggplot136 in R.
As described above, the core haplotype with the most tests indicating selection was in the SUSD1 segment found in the Melanesians. We show in Fig. 3A that 13 variants in this core haplotype are at the top 1% of the genome-wide normalised nSL55 distribution. When examining the normalised, windowed nSL results, the core haplotype region was within the top 5% of windows across the genome for the Melanesians (Table S3_D). In the Papuans, the general introgressed segment identified by SPrime7 that contained this core haplotype had several windows that scored at the top 1% of the distribution (Table S3_D), although the closest top scoring window was approximately 241,000 base pairs (bp) away from the start of our core haplotype region and overlaps a different gene (SHOC1; shortage in chiasmata 1), which was not part of our analysis. Additionally, we plot in Fig. 3B the EHH52 measured decay for the SUSD1 main archaic allele relative to the non-archaic allele, illustrating the exceptional long range haplotype homozygosity in this segment for the archaic allele. Lastly, in Fig. 3C, we show the rapid increase in allele frequency for chr9:114810102 (rs10981228), the main core haplotype archaic allele in the Melanesians. In short, the probable evidence for selection in this region is high.
No evidence of latitude cline within core haplotypes
Several prior studies have provided evidence that circadian rhythm or chronotype associated genes and variants will often exhibit a latitudinal cline. Specifically, phenotypes related to chronotype based on latitude have been identified in modern human populations43,44 and have also been described in regions introgressed from archaic populations20,22,24. The link between latitude and circadian rhythm has been established in plants, animals, and insects to varying degrees60,61,62. However, an article examining the clock gene PER2 (period circadian regulator 2) found no evidence of a latitude-based gradient in their analysis of modern humans across 5 continents63. We tested if any evidence of a latitudinal cline could be found in our general introgressed segments that contain a core haplotype (Table S3_A). We extracted the maximum archaic allele frequency from each population that intersected our coordinates of interest and evaluated the correlation of these frequencies with latitude. Our results support the idea that there is no clear latitude signature displayed (Fig. 4; Table S4; Figs. S1–S62), like with previous analysis of PER263. In general, the correlations between absolute latitude and maximum archaic allele frequency were low. Only 10 of these regions have significant p-values (p ≤ 0.05), and the relationship is not very strong with the highest Pearson’s correlation coefficient (r) being 0.4885 in TLR1 (toll like receptor 1) (Table S4). Most of the relationships are also counter to our hypothesis that higher latitude modern human populations would have higher frequencies of archaic alleles. Higher latitude groups, such as the British from England and Scotland (GBR) and Finnish in Finland (FIN) have lower allele frequencies than many populations from middle and low latitude regions. This suggests that other mechanisms rather than latitude alone are driving archaic allele frequencies.
Frequency distribution maps generated by extracting the maximum archaic allele frequency from each core haplotype region in each population in our analysis. Here we display a subset of these with interesting geographic patterns. Both A CEACAM1-LIPE-AS1 and B JAK1 high frequency alleles are nearly exclusively found in Oceanic populations. C AMIGO2 and D CCR9 exhibit higher frequencies within Asia and Oceania. E RN7SL423P-ENSG00000232337 shows a band across South Asia into Europe of high-frequency variants. Our analysis displays no clear latitudinal cline within our core haplotypes with evidence of positive selection. The plots were generated with the rnaturalearth139, sf140,141, and ggplot2136 packages in R116.
Despite a lack of a clear latitudinal pattern, we found evidence of geographic grouping of some core haplotypes. For instance, the CEACAM1-LIPE-AS1 and JAK1 frequency maps show a clear bias towards Oceania, with allele frequencies greater than 40%, while these segments are nearly, or entirely, absent in the rest of our populations (Fig. 4A, B). Similar patterning can be seen in the AMIGO2 (adhesion molecule with Ig like domain 2) frequency map, where elevated allele frequencies are mostly isolated to Asia and Oceania (Fig. 4C). We find high allele frequencies in Oceanic and South Asian populations in the CCR9 (C-C motif chemokine receptor 9) map (Fig. 4D), which also has some moderately elevated allele frequencies within Europe and several parts of the Americas. There is also a South Asia to Europe band of high-frequency alleles found in the RN7SL423P-ENSG00000232337 (RNA, 7SL, cytoplasmic 423, pseudogene) segment (Fig. 4E). The allele frequencies and latitude for each population for all core haplotype regions are given in Table S4.
Association of archaic variants with complex traits and diseases
We wanted to explore potential associations of the archaic variants within the segments reported in Table S2 to better understand the implications of our results. In total, we identified 714 putative archaic SNPs that were found at frequencies ≥40% in our results that also had genome-wide significant p-values in the IEU OpenGWAS project database64,65. These variants and associations are provided in Table S5. There are 75 SNPs with 138 associations at genome-wide significance within our core haplotypes (Table S5). Our results demonstrated a high level of pleiotropy, often with many of the same variants found to be associated with more than one of the 358 significant traits identified in our analyses. Some of these relationships are elaborated on in the Supplementary Material. Variant annotation for functional consequences using SNPnexus66,67 is provided in Table S6. One of our hypotheses was that archaic variants would be significantly associated with serotonin traits, as it has been found before to support adaptations to strong seasonal variations in light. However, our results were largely negative, and we found no significant trait associations with genes related to serotonin in our results, which we describe in detail in the Supplementary Material. Gene ontologies (GO)68,69 obtained from BioMart70 for genes related to serotonin are provided in Table S7.
SUSD1 has been previously linked with calcium ion and protein binding, along with being integral to membrane in our GO analysis68,69 (Table S7). Only one SUSD1 SNP was identified in SNPnexus66,67, chr9:114810102 (rs10981228), however, it was an intronic mutation that had no suspected impact on protein function (Table S6). This same mutation has been significantly negatively associated with haematocrit, haemoglobin concentration, and total testosterone71,72 in the OpenGWAS project database64,65 (Table S5). Similarly, JAK1 has no mutations that change any amino acids (Table S6), and it has been associated with protein binding (Table S7), but there are no genome-wide significant variants identified for this core haplotype (Table S5). DNAAF10 has 7 SNPs where the archaic allele is genome-wide significant for 12 individual traits including being negatively associated with various vein-related traits and systolic blood pressure while also being positively associated with height and waist-hip ratios73,74 (Table S5). None of these were amino acid changing (Table S6) nor was the gene associated with any GO terms (Table S7). TSPAN11 had 24 SNPs listed in SNPnexus and they did not change the amino acid (Table S6). Additionally, it was listed as being integral to membrane in our GO analysis (Table S7). ENSG00000258312 had no genome-wide significant SNPs (Table S5) or any listed GO terms (Table S7) but it had two variants listed in SNPnexus, however, they had no impact on the amino acid (Table S6). Lastly, RPSAP11-ENSG00000261572 had no genome-wide significant variants, notable SNPs, or GO terms in our analyses (Tables S5–S7).
Core haplotype alleles are genome-wide significant for chronotype and other sleep traits
Over twenty percent of our genome-wide significant variants are associated with chronotype (Table S5), spanning three separate chromosomes and 11 genes. ENSG00000286749, one of our core haplotypes, contains 101 variants that are significantly positively associated with being a morning person73, with 11 of these found directly within the core region. This finding confirms prior analysis which documented various archaically derived variants are associated with a morning preference in modern humans21,24. Conversely, the maximum archaic alleles for the LINC01107 (long intergenic non-protein coding RNA 1107) and LINC01937 (long intergenic non-protein coding RNA 1937) core haplotypes are significantly negatively associated with being a morning person73, confirming prior research finding an archaic haplotype also on chromosome 2 that was associated with being an evening person20. Some research has linked latitude and chronotype43,44. For example, in adolescents, preference for being a morning person occurred at higher latitudes within Europe74, while evidence for a higher likelihood of evening preference was found in both Turkish75 and Brazilian populations43. In summary, our results indicate that archaic alleles can have different directions of effect on chronotype, as reported in previous studies20,21,24, but no clear latitude-chronotype link is evident in our results. This mirrors our results above where no obvious signature of a latitude cline could be found in our core haplotypes.
Five variants in our results are negatively associated with sleep duration. All these variants are found overlapping directly or intergenic to the SGCZ gene (sarcoglycan zeta) on chromosome 8 (Table S5). Additionally, we have evidence of one variant within the SLC4A10 gene (solute carrier family 4 member 10) on chromosome 2 that is negatively associated with daytime napping76 (Table S5), indirectly supporting the results from SGCZ as the archaic alleles in both regions yield reduced levels of total sleep. Both findings are supported in prior research that found archaic loci associated with sleep duration and daytime napping behaviours20,23, however, our variants were not described in their datasets, so it is unclear if the direction of the effect is the same.
Archaic alleles and affective disorders
The connection between circadian rhythm and mental health disorders, such as bipolar disorder and schizophrenia, has been discussed before77,78. Previous research found enrichment for schizophrenia-associated loci in modern humans after the human-archaic split79, however, this association has been contested when two recent studies described they found no such connection21,23. Our results highlight 21 SNPs associated with schizophrenia, and one of our core haplotypes, ENSG00000286749, overlaps 11 variants that are negatively associated with schizophrenia80, suggesting the archaic allele may have a protective role against this trait (Table S5). An additional 8 variants within the genes ALPK3 (alpha kinase 3), SEC11A (SEC11 homolog A, signal peptidase complex subunit), and ZNF592 (zinc finger protein 592) show similar patterning80. Prior analysis also noted that archaic variants were protective against schizophrenia21. Further, a study analysing the genomes of patients with schizophrenia found that higher proportions of Neanderthal DNA was linked with lowered symptom severity81. Studies have illustrated that bipolar disorder and schizophrenia have a genetic correlation of approximately 0.6, which is likely due to sharing common risk alleles82. On chromosome 15, we have evidence of 6 variants that are protective against schizophrenia that are also negatively associated with bipolar disorder83. Taken together, these results show that archaic alleles in modern humans could be mutually protective against some linked affective disorder onsets.
Several core haplotypes are significantly associated with inflammatory and respiratory diseases
Many inflammatory and respiratory diseases have illustrated circadian cycling. For instance, asthma flare-ups are worse during nighttime suggesting that they follow circadian patterning84, and circadian rhythms have been shown to have an involvement with certain versions of dermatitis85. We identified 27 variants overlapping our core haplotypes spanning four different chromosomes that are significantly linked with inflammatory or respiratory illnesses in our dataset (Table S5). TLR1 has been described before as having haplotypes derived from archaic hominins17,86,87,88,89. The maximum archaic allele for our TLR1 core haplotype, rs5743592 (Table S3), in conjunction with 30 other variants in the same region, have been demonstrated to be protective against asthma, various allergy phenotypes, and eczema73,90,91, which contrasts with previous research describing that archaic haplotypes overlapping TLR1 may cause allergic phenotypes in modern humans17. Intriguingly, a study has highlighted the role neutrophils may have in mitigating airway inflammation in response to allergies and asthma by triggering cytokine production92. Potential support for this conclusion can also be seen in our results where the same variants associated with lowered incidence of asthma share a positive association with neutrophil concentrations93. Lastly, we also have evidence for one variant, rs11757612 on chromosome 6, an intergenic region between ENSG00000197251 and ENSG00000096433, where the archaic allele is protective against coeliac disease94 and chronic hepatitis B72. In summary, our results highlight the role archaic alleles may play in mitigating severe responses to asthma, allergic responses, and several other inflammatory diseases.
Introgressed variants and Type-2 diabetes risk
Our results highlighted over 50 genome-wide significant markers overlapping 9 genes on chromosomes 11 and 12 associated with Type-2 diabetes (Table S5). Disrupted circadian rhythms are believed to be at least partially responsible for diabetes onset. For instance, mouse models with lab-delayed circadian rhythms and removal of the pancreatic clock led to a diabetic phenotype95. Review articles have also provided evidence suggestive of a role for disturbed rhythms in the liver and pancreas ultimately manifesting in atypical glucose metabolisms96,97. In total, 27 variants from Biobank Japan72 and 21 variants from the FinnGen Biobank94 in our results show directions of effect indicative of the archaic alleles of these variants being protective against Type-2 diabetes. However, an earlier study demonstrated that a Neanderthal haplotype within the SLC16A11 gene (solute carrier family 16 member 11), which was not found to be one of our adaptively introgressed genes in our results, is associated with elevated Type-2 diabetes risk98, so it is possible that archaic alleles can impact risk in both directions.
Archaic-derived eQTLs are significantly enriched in immune pathways
Next, we evaluated if our archaic variants are associated with gene expression levels in any tissues and obtained this list of eQTLs using the SNP2GENE function from FUMA GWAS99,100. Table S8 reports eQTL results for the variants listed in Table S5, where we identified 621 archaic alleles showing significant associations with gene expression in multiple tissues. Table S9 reports eQTL results (n = 59) for the variants overlapping core haplotypes listed in Table S3_A. This supports prior analysis suggesting that archaic variants associated with circadian rhythm expression have a higher likelihood of being eQTLs than expected relative to other introgressed variants24.
Links between circadian rhythm oscillations and immune function have been described previously33,34,35, and our analysis of the eQTLs reinforces these ties. We were interested in evaluating if genes with genome-wide significant variants (Table S5), core haplotype genes (Table S3_A), or genes regulated by archaic eQTLs (Tables S8 and S9) are enriched in specific pathways or GO terms68,69. A general overview of these results is explained in the Supplementary Material. We found one significant Reactome 2022 pathway101 in the results for our larger eQTL set, the chemokine receptors bind chemokines pathway (Table S12), which is associated with immunity. When analysing the eQTLs within just core haplotypes, we find four significant pathways within our ShinyGO102 results related to immune system function and response, specifically, significant hits regarding viral interactions and chemokine and cytokine pathways (Table S13). Additional support for this comes from our Reactome and KEGG pathway103 Enrichr104,105,106 results from just core haplotype eQTLs, showing significant pathways that include Toll-like receptor cascades, interleukin-10 signalling, and viral protein interaction with cytokine and cytokine receptor as some of our top results (Table S14). GO term analysis shows the two most significant enrichments for biological processes within the core haplotype eQTLs are inflammatory response and cellular response to bacterial lipopeptide and the GO molecular function results have significant enrichment in Toll-like receptor binding and cytokine receptor activity (Table S14). In summary, eQTLs that are the result of archaic introgression are significantly enriched within immune response pathways, biological processes, and molecular functions linked with immunity, particularly those found within our core haplotypes. Our results are in line with many previous studies suggesting that introgressed archaic loci are associated with immunity15,16,17,18,86,107,108,109.
Discussion
Our paper has documented high archaic contributions to over 300 genes and intergenic segments that fall within 265 independent windows within global non-African populations. We were able to expand on previous analyses by investigating the extent of adaptive introgression within genes displaying circadian rhythm cycling and variants associated with circadian rhythm, chronotype, or sleep phenotypes using 76 worldwide populations, where previous studies have focused mostly on Eurasian populations from the 1KGP. Many of our reported genes show well documented signatures of introgression from archaic samples into modern humans, including an abundance of immunity-associated loci, complex traits including schizophrenia and bipolar disorder, and sleep associated phenotypes. We utilised GWAS summary statistics to understand the effects our archaic alleles have on associated traits. Within our results, we identified 1729 variants that have allele frequencies of at least 40%, and are directly matched to an archaic allele, of which, 714 are genome-wide significant SNPs based on GWAS analysis. Additionally, we also note that within our genome-wide significant results, we were able to obtain 621 eQTLs expressed in a variety of tissues. We explored in detail 35 regions that we consider to be core haplotypes. From these, 22 core haplotypes displayed evidence of positive selection within modern human populations from at least two selection tests, providing leverage to the idea that these specific regions were direct products of adaptive introgression.
The adaptive introgression of variants within genes that display circadian rhythm cycling or are associated with circadian rhythm, chronotype, or sleep phenotypes likely contributed to both advantageous and complex effects in modern human populations. We explored several hypotheses within our paper, including whether adaptively introgressed segments with evidence of positive selection would display previously described signatures of a latitudinal cline. We did not find definitive evidence of this within our core haplotype regions, instead finding clearer signals that these regions cluster more closely based on geographic similarities and previously described archaic ancestry patterns. Additional hypotheses addressed the possibility of higher latitude variants being associated with more versatile circadian cycling to combat the extreme seasonal changes in light availability and that variants associated with serotonin synthesis would be apparent in our results. We found that archaic variants are associated with both a morning and an evening preference, contributing to research on the role of chronotype phenotypes in modern humans derived from archaic populations. However, more work needs to be done to understand the significance of these findings and how they relate to previous discussions on the role of chronotype and latitude. Additionally, our findings for evidence of adaptive introgression of serotonin-associated segments were largely inconclusive, and future research should explore this hypothesis in more detail. However, our findings suggest a complex and context-dependent relationship between archaic introgression, circadian regulation, and affective disorders, where some introgressed alleles may offer protection against conditions like schizophrenia and bipolar disorder. Further, the obvious ties to immune pathways, and some evidence of digestive traits such as coeliac disease in our results, showcase clear ties to the adaptive introgression of genes that show circadian cycling.
Our study has several limitations. First, is that SPrime’s accuracy drops when a population has less than 15 samples for analysis7. This is unfortunately the case for many of the populations within the HGDP sample set. A consequence of this is that some of our windows may represent false positives. Additionally, small sample sizes are associated with more uncertainty in allele frequency estimates, and this may result in high archaic allele frequencies in some of the HGDP samples, where other geographically similar populations with adequate sample sizes, such as in the 1KGP populations, do not show such high frequencies. While several introgressed variants show frequencies close to fixation, many of these variants are also seen in populations with sample sizes over 15 and at allele frequencies greater than the typical introgressed archaic background frequencies20,110, along with passing our authentication parameters, suggesting that the introgressed segment is correct. Overall, it is important to be cautious about the interpretation of the archaic allele frequencies of some of the HGDP samples due to their very small sample sizes.
A second limitation is SPrime’s masking of modern human segments found in an African reference panel7, which has been shown to limit the detection power of archaic sequences in populations outside of the reference111. Therefore, we may be removing variants that would have passed our filters due to being shared with our reference population. An extension of this is our filtering thresholds were quite stringent. Because we were looking for signatures of adaptive introgression, all the variants discussed here have ≥ 40% allele frequency, which means most archaic alleles will fail this filtration step. On one hand, we can clearly exhibit instances of adaptive introgression regarding circadian rhythm and chronotype-associated variants in modern humans due to admixture with archaic hominins, on the other hand, we also removed many other very interesting segments worth exploring that may have elevated allele frequencies relative to typical archaically-introgressed levels.
In our study, we explored the potential implication of the archaic variants by downloading information on the association of these variants with complex traits and diseases from the IEU OpenGWAS project64,65. Importantly, we retrieved information on the direction of effect of the archaic variants, which is relevant to understand their putative impact. However, it should be noted that the interpretation of the direction of effects for some traits is not straightforward without specific information about the definition of the phenotypes studied. It is also important to point out that although the archaic variants show genome-wide significant effects for a wide range of traits, this does not necessarily imply that they are the causal variants. In principle, these associations may be driven by other variants in linkage disequilibrium with the archaic variants. Identifying causal effects would require fine-mapping and functional validation for dozens of regions and traits, which is beyond the scope of this paper. Lastly, we did not attempt to understand why some of the core haplotypes described in our analysis may be under selection and leave this potential analysis for future work.
In conclusion, this study significantly advances our understanding of how archaic introgression has influenced modern human circadian rhythms and sleep patterns. By identifying a broad array of introgressed genes and intergenic segments linked to circadian functions across diverse global populations, we provide new insights into the evolutionary pressures that shaped these traits. The clear evidence of positive selection in several of these regions underscores their adaptive value. Yet, this work also leaves room to explore future hypothesis driven work to disentangle the role clinal adaptation has played in the evolution of circadian rhythms in the human lineage. This research paves the way for future studies to explore the intricate connections between circadian rhythms, mental health, and immune function, potentially leading to innovative approaches in chronomedicine and personalised healthcare.
Materials and methods
Modern human, Neanderthal, and Denisovan VCF files
Our modern human samples came from the previously published, phased gnomAD 1KGP + HGDP call set39. This unique dataset compiles the high-resolution data from the HGDP (n = 51 populations) and 1KGP (n = 25 populations) all mapped to GRCh38 (hg38) coordinates. Following the SPrime7 protocol outlined by Zhou and Browning51, we used the Yoruba in Ibadan, Nigeria (YRI) population from the 1KGP (n = 106) as the outgroup for our analyses and combined them with each target population and removed any non-biallelic SNPs and duplicated variants using BCFtools v1.13112. We updated all known variant IDs using the dbSNP database113 annotation files for matching abilities in downstream analyses. The archaic VCFs and their associated mask file locations are available in the data availability statement.
Introgression identification and matching to archaic sequences
To identify variants that are likely due to admixture between archaic hominins and modern humans we used SPrime7 with all recommended settings according to the original paper. SPrime is an archaic-reference-free software that uses a scoring parameter to identify segments in modern humans considered to be inherited from archaic hominins. These segments are kept by the software if they are above the recommended scoring threshold of 150,000. Because the modern human genomes are mapped to the newer hg38 modern human genome coordinates and the Neanderthal and Denisovan genomes are mapped to the older GRCh37 (hg19) modern human genome coordinates, we lifted over the SPrime output files using the UCSC LiftOver Linux executable114 to hg19. To ensure that LiftOver had not mapped any variants incorrectly, we discarded any variants that had jumped chromosomes. We used a secondary software, map_arch51 to match our results to the archaic alleles. This software takes the SPrime output file, an archaic VCF, and the associated archaic mask file to create a new file showcasing whether the modern human variant identified by SPrime matches, mismatches, or is not comparable with the archaic genome of interest. Next, we used BCFtools112 to generate allele frequencies for each of the modern human populations and merged these with our output using the dplyr package115 in R v4.1.2116. We extracted the putatively introgressed segments identified in our analysis from each population, combined them together with populations from the same region according to the gnomAD sample meta table, and reduced them to the minimum number of non-overlapping segments using the GenomicRanges package117 in R.
Chronotype and circadian rhythm datasets
We were interested in identifying if genomic regions showing circadian rhythm oscillations or regions associated with sleep and chronotype traits in modern humans had any signatures of introgression from archaic hominins. Our analysis focused on compiling genes and variants linked with circadian rhythm or chronotype expression from previously reported datasets to test for these signatures. The CGDB contains over 70,000 genes that display circadian oscillating behaviours identified in eukaryotic organisms49. We downloaded the genes found on modern human autosomes (n = 1236) from the CGDB along with variants and introgressed segments published by Dannemann and Kelso20, McArthur et al.21, Dannemann et al.23, and Velazquez-Arcelay et al.24, all of which connected with circadian rhythms and chronotype expression because of archaic introgression. Additionally, we extracted all hits from the NHGRI-EBI GWAS Catalogue50 showing associations with chronotype, circadian rhythm, or sleep phenotypes at genome-wide significant p-values of p = 5 × 10−8 or less. Since some studies will report windows suggestive of introgressed haplotypes, and others focus on just reporting variants, we opted to normalise our analysis by extracting either previously reported variants or variants found within previously described windows. Because some studies used in our analyses use different genome builds (hg19 vs. hg38), we report all our results in hg19 coordinates to match that of the archaic samples. We extracted these variants from our results by matching the variant rsIDs using the dplyr package115 in R116.
Adaptive introgression and identifying core haplotypes
We were interested in identifying if any archaic variants present in the modern human genome were brought to elevated frequencies due to adaptive introgression, focusing on genes with a detectable circadian oscillation component or loci previously linked with chronotype, circadian rhythms, or sleep in GWAS studies. SPrime is sensitive enough to detect adaptive introgression in the human genome7. To test for this, we followed the recommended workflow of selecting identified archaic segments that have 30 or more markers in the identified segment and have a match/mismatch ratio (number of matches divided by the total number of matches and mismatches) of >50% for the Neanderthals and >40% for the Denisovan. After filtering out segments that failed this step, we used snpEff118 to annotate our VCFs with gene names that overlap our variants and the variant functional consequences, followed by merging this information with our population files using the rsIDs. Next, we used BEDTools v2.30.0119 to intersect our population files with every other population to identify regional signatures and repeated this for each population for each archaic.
Browning and colleagues7 identified two highly probable regions of adaptive introgression per population using SPrime, and we applied a similar methodology to identify possible targets of adaptive introgression in each modern human autosome that were related to circadian rhythm or chronotype. For each SPrime-identified segment that passed our thresholds, we located core haplotypes by finding variants that were the top archaic allele frequency within the segment with allele frequencies of at least 40%, the allele matched at least one archaic sample, and the variant intersected one of our genes of interest from the CGDB49, or matched a variant described previously as being significantly associated with circadian rhythm or chronotype20,21,23,24,50. Once we identified these variants, we then preferentially selected all variants directly adjacent to this core variant that had an allele frequency no more than 5% below the maximum archaic allele frequency variant. This allowed us to identify what we believe are core haplotypes introgressed from archaic populations related to circadian rhythm, chronotype, or sleep expression. Due to oftentimes small sample size and the effects of drift, we focused our analysis of the core haplotypes from the 1KGP populations only, along with the Papuan and Melanesian samples from the HGDP, to test for signatures of Denisovan ancestry.
Archaic donor populations
The ratio of the number of matches and mismatches can be compared to identify whether introgressed segments are from Neanderthals or Denisovans7,51. We tested for this by taking segments with 30 or more variants where segments that are believed to be of Neanderthal affinity will have match/mismatch ratios greater than 60% to a Neanderthal sample coinciding with a match/mismatch ratio below 40% for the Denisovan. Similarly, segments likely of Denisovan origin will have a match/mismatch ratio more than 40% to the Denisovan and below 30% with the Neanderthals. To identify which of these segments are clearly introgressed from populations similar to one archaic sample relative to the others we first excluded any segments that did not pass the adaptive introgression thresholds described above. When a remaining segment passed the donor thresholds and had a segment match/mismatch ratio more than 5% higher relative to the other three archaic samples, we inferred that the putative archaic donor is closest to that archaic population. If the match/mismatch ratio is within 5% relative to the other archaic samples, we consider that to be inconclusive and is of archaic affinity generally. We applied these calculations to our core haplotypes. To visualise the number of segments at specific match/mismatch ratios within a chromosome containing a core haplotype between the different Neanderthal samples and Denisovans, we generated contour plots based on the scripts provided by Zhou and Browning51 in R116 using the kde2d function from MASS120 for core haplotypes with evidence of positive selection within their core region.
Tests for selection
To give additional weight to our analysis and identify core haplotypes with evidence of adaptive introgression, we ran a variety of tests to detect selection across the genome. We first calculated FST using the Weir and Cockerham method53 (--weir-fst-pop) using VCFtools v0.1.16121 with 100,000 bp sliding windows (--fst-window-size 100000) with a 10,000 bp step (--fst-window-step 10000). Since FST can be sensitive to allele frequency122 and sample size123, the comparison population for each of our core haplotype populations was the YRI for the 1KGP populations, and the Yoruba from the HGDP for the Oceanic populations, which appropriately sample size matched the target and reference populations. Additionally, we calculated per site FST values with no additional parameters (--weir-fst-pop). Following this, we calculated Tajima’s D58 values using VCFtools in 100,000 bp non-overlapping windows (--TajimaD 100000).
Next, we applied a number of methods that are useful for detecting selection on haplotypes, which rely on identifying reductions of homozygosity due to distance from a specified region. We used selscan v2.0.3124 to examine population specific statistics that track this decay, implementing EHH52 (--ehh) and nSL55 (--nsl). The former has been shown to be an excellent first pass to determine selection even in comparison to more modern approaches125 and the latter is robust to many different demographic models and selection scenarios while still maintaining high accuracy. We calculated EHH for our maximum archaic allele frequency variants with a 300,000 bp window (--ehh-win 300000) directly adjacent to this variant to visualise haplotype decay patterns for both the archaic and non-archaic alleles at each locus. We inferred selection to have occurred when the archaic allele haplotype was larger relative to the non-archaic allele, measured by the drop of the EHH score below 0.25 in both directions from the main core allele. Per-site ancestral and derived allele nSL scores were obtained for each core haplotype in each target population. We also applied a method based on cross-population comparisons (XP-nSL)59, using the target population and the YRI population as reference (--xpnsl). Selscan was then used to normalise the data for both nSL and XP-nSL, first per-site, and second, in 100,000 bp windows and placing them into 10 quantile bins for all windows containing at least 10 variants per window (norm --nsl --bp-win --winsize 100000 --qbins 10 --min-snps 10; norm --xpnsl --bp-win --winsize 100000 --qbins 10 --min-snps 10) as described in a prior publication59.
Following this, we applied two modern composite likelihood methods, RAiSD v2.956 and saltiLASSI v1.2.157. RAiSD examines for evidence of selection by identifying regions that satisfy three criteria, namely, a localised reduction in polymorphic loci, elevated linkage disequilibrium around a specific mutation, and site frequency spectrum changes of derived variants, which are compiled into the μ statistic. In contrast, saltiLASSI looks for deviations from expected values regarding the haplotype frequency spectrum to infer selection. saltiLASSI outputs three statistics: m, reporting the number of high frequency haplotypes, where m = 1 signifies a hard sweep and m > 1 suggests a soft sweep, A, which measures the width of a sweep to gauge the strength and timing of a selection event, and Λ, which characterises the haplotype frequency spectrum relative to other haplotypes in the region. Lastly, we also included Relate’s (v1.2.1) tree-based approach, which measures selection via the speed at which a mutation expands throughout a population relative to other lineages within the same group54.
To infer evidence of regions putatively under selection we applied an outlier approach, where for each statistic we combined the data genome-wide for each population and identified values equal or greater than the top 1% of the empirical distribution. For two-sided tests we calculated the top and bottom 1% values from the genome-wide empirical distribution. Tajima’s D is a two-sided test where positive values may suggest increased heterozygosity or population expansion and negative values may suggest increased homozygosity or populations reducing in size58. Additionally, nSL55 and XP-nSL59 also are two-sided tests, where positive values indicate long range homozygosity in favour of the derived allele (nSL) or target population (XP-nSL), and negative values suggest long, homozygous haplotypes in favour of the ancestral allele (nSL) or reference population (XP-nSL), respectively. All variants and/or regions that matched or exceeded these 1% thresholds for both one- and two-sided tests we took as potentially being under evidence of selection. Lastly, normalising the selscan124 outputs we also looked for evidence of our general introgressed segment or core haplotypes containing windows at the top 5% and top 1% of the empirical distribution of windows genome-wide.
Haplotype networks
Haplotype networks are used to visualise genotype relationships within species to help answer questions revolving around biogeography, genealogical relationships, or population histories126 for the genetic region of interest. After identifying the core haplotypes with evidence of positive selection within their core region (Table S3_A) we generated a haplotype network for SUSD1, the highest scoring segment in our selection tests, to help illustrate the relationships between these sequences. To do this, we first removed all heterozygous sites from the archaic samples. These samples are not phased and removing heterozygous sites de facto phases the genome, because at homozygous sites the allele is the same for both parents. Further, archaic samples have demonstrated lengthy stretches of homozygosity41,42,127,128, and prior analyses have generated haplotype networks including archaic samples without phasing and relying on runs of homozygosity in the archaic samples to generate accurate relationships14.
To generate the SUSD1 haplotype network, we used BCFtools112 to remove 11,315 variants (0.000153% of the total variants) from the Altai Neanderthal, 15,089 variants (0.000204% of the total variants) from the Chagyrskaya Neanderthal, 15,618 variants (0.000211% of the total variants) from the Vindija Neanderthal, and 17,103 variants (0.000231% of the total variants) from the Denisovan, from chromosome 9, respectively, after filtering out heterozygous sites. This meant that our filtering led to the loss from the core haplotype of three variants (0.000153% of the total variants) from the Altai and Chagyrskaya Neanderthals, 10 variants (0.000511% of the total variants) from the Vindija Neanderthal, and no variants lost from the Denisovan. Following this, we merged the modern human and archaic SUSD1 core haplotype VCFs, removed any missing sites using BCFtools, and removed monomorphic sites using PLINK2129. This left us with 430 SNPs across a 27,265 bp region for our haplotype analysis.
We used PGDSpider v2.1.1.5130 to generate FASTA format files from the VCFs and then aligned the FASTA files using MUSCLE v3.8.31131. MEGA v11.0.13132 was used to convert the aligned FASTA files into NEXUS format133, where we imported these into DnaSP6 v6.12.03134 to assign population sequence sets and generate haplotype files. We generated our haplotype networks using PopART v1.7126 using the median-joining function after uploading our subset haplotype files and associated traits file (counts of each population found within each haplotype of interest). For simplicity, we grouped the populations based on their region/superpopulation (i.e., AFR/African, AMR/American) in our haplotype file, traits file, and on the haplotype network itself due to the large number of subpopulations we have in our joined dataset. In total, there were 668 haplotypes in the SUSD1 core region. To generate coherent networks, we selected the 50 most frequent haplotypes, and their ties, along with the haplotypes associated with the archaic samples, and further preferentially selected all Oceanic haplotypes with frequencies ≥2 that were not already included in the top 50 haplotypes to help us visualise possible relationships to the populations our core haplotype was identified in in more detail. This left us with 53 haplotypes to generate our network with.
Ancestral recombination graphs
Ancestral recombination graphs, which visualise coalescence relationship trees, showing both mutations and lineages broken up by recombination135, were generated using Relate54. We regenerated VCF files for the core haplotypes using all modern human samples in our dataset (n = 4091) using the phased 1KGP + HGDP callset39. We converted the VCFs to the proper format using Relate’s built in commands, including the RelateFileFormats --mode ConvertFromVcf and PrepareInputFiles.sh operations. The PrepareInputFiles script removes non-biallelic SNPs, determines the ancestral allele, and filters variants based on the genomic masks. After filtering, we were left with 404 SNPs for analysis within the core haplotype. We ran Relate in parallel (RelateParallel.sh) using the standard input parameters of 1.25e-8 for the mutation rate and an effective population size of 30,000. We selected trees of interest based on the criteria from the original Relate paper, where signatures of archaic introgression show as a long, non-recombining branch that begins before the split date of modern humans, Neanderthals and Denisovans, which subsequently has rapid proliferation through the modern human population of interest recently. We selected branches with origins ~1,000,000 years ago and the derived variant matches the archaic allele. We used Relate to extract subpopulation trees (RelateExtract --mode SubTreesForSubpopulation) and plotted mutational trees (TreeViewMutation.sh) using the built in Relate functions.
OpenGWAS trait associations
We evaluated significant associations of archaic variants with common traits and diseases. GWAS summary statistics including effect and non-effect alleles, beta values, standard errors, and p values for all genome-wide significant markers were curated from the IEU OpenGWAS project64,65 to determine trait directionality. For genome-wide significant variants where the archaic allele did not match the effect allele, we converted beta to the correct allele by flipping the sign and changing the non-effect allele to represent the proper allele.
Regulation of gene expression on archaic variants
We explored if the archaic variants showing significant associations to complex traits and diseases play a role in the regulation of gene expression. We extracted eQTL information for all the archaic variants with significant trait associations in our OpenGWAS results (n = 714) using FUMA GWAS’s SNP2GENE function99,100 and repeated this analysis again for just significant variants that were found inside of our core haplotypes (n = 75).
Variant annotations and gene ontology
Annotation for all variants for consequence information in our results (n = 1729) was done using SNPnexus v466,67. We were also interested to see if any genes were associated with specific ontologies and compiled GO Consortium data68,69 results for our genes with evidence of adaptive introgression (n = 303). We downloaded this information from BioMart using the GRCh37 Release 11270.
Enrichment analysis
To complement our ontology analysis, we wanted to determine if any of our genes were significantly enriched in any pathways. To do this, we input the set of genes showing genome-wide significant associations with traits or diseases in our OpenGWAS64,65 results (n = 113) through ShinyGO102 to examine for enrichment within the KEGG database103 and through Enrichr104,105,106 to explore enrichment in the following databases: Reactome 2022101, KEGG 2021 human, GO biological processes, and GO molecular functions68,69. We repeated this analysis for just the genes that had archaic variants with significant OpenGWAS results from the core haplotypes (n = 19). Finally, we also performed enrichment analysis focused on the set of genes regulated by the eQTLs identified by FUMA GWAS99,100, both for the complete set of archaic variants with significant OpenGWAS results (n = 334), and the subset of variants overlapping our core haplotypes (n = 53).
Data availability
Our main SPrime results files used to create the analyses are publicly available without restriction at https://zenodo.org/records/15271373. The SPrime protocol, and subsequent links to relevant software used in the protocol are available at https://doi.org/10.1016/j.xpro.2021.100550. The phased 1KGP + HGDP call set, and associated metadata tables were acquired through the relevant reference. The Altai Neanderthal, Vindija Neanderthal, and Denisovan mask and VCF files can be downloaded directly at http://cdna.eva.mpg.de/neandertal/Vindija/. The Chagyrskaya Neanderthal VCF file and mask files can be found at http://ftp.eva.mpg.de/neandertal/Chagyrskaya/. All selection software used in our analyses are available via download links with their respective reference.
References
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Vernot, B. & Akey, J. M. Resurrecting surviving Neandertal lineages from modern human genomes. Science 343, 1017–1021 (2014).
Vernot, B. & Akey, J. M. Complex history of admixture between modern humans and Neandertals. Am. J. Hum. Genet 96, 448–453 (2015).
Kuhlwilm, M. et al. Ancient gene flow from early modern humans into Eastern Neanderthals. Nature 530, 429–433 (2016).
Browning, S. R., Browning, B. L., Zhou, Y., Tucci, S. & Akey, J. M. Analysis of human sequence data reveals two pulses of archaic Denisovan admixture. Cell 173, 53–61 (2018).
Jacobs, G. S. et al. Multiple deeply divergent Denisovan ancestries in Papuans. Cell 177, 1010–1021 (2019).
Villanea, F. A. & Schraiber, J. G. Multiple episodes of interbreeding between Neanderthals and modern humans. Nat. Ecol. Evol 3, 39–44 (2019).
Li, L., Comi, T. J., Bierman, R. F. & Akey, J. M. Recurrent gene flow between Neanderthals and modern humans over the past 200,000 years. Science 385, eadi1768 (2024).
Qin, P. & Stoneking, M. Denisovan ancestry in East Eurasian and native American populations. Mol. Biol. Evol 32, 2665–2674 (2015).
Sankararaman, S., Mallick, S., Patterson, N. & Reich, D. The combined landscape of Denisovan and Neanderthal ancestry in present-day humans. Curr. Biol 26, 1241–1247 (2016).
Skov, L. et al. Detecting archaic introgression using an unadmixed outgroup. PLoS Genet 14, e1007641 (2018).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014).
Abi-Rached, L. et al. The shaping of modern human immune systems by multiregional admixture with archaic humans. Science 334, 89–94 (2011).
Racimo, F., Sankararaman, S., Nielsen, R. & Huerta-Sánchez, E. Evidence for archaic adaptive introgression in humans. Nat. Rev. Genet. 16, 359–371 (2015).
Dannemann, M., Andrés, A. M. & Kelso, J. Introgression of Neandertal- and Denisovan-like haplotypes contributes to adaptive variation in human toll-like receptors. Am. J. Hum. Genet. 98, 22–33 (2016).
Vespasiani, D. M. et al. Denisovan introgression has shaped the immune system of present-day Papuans. PLoS Genet. 18, e1010470 (2022).
Sankararaman, S. et al. The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507, 354–357 (2014).
Dannemann, M. & Kelso, J. The contribution of Neanderthals to phenotypic variation in modern humans. Am. J. Hum. Genet. 101, 578–589 (2017).
McArthur, E., Rinker, D. C. & Capra, J. A. Quantifying the contribution of Neanderthal introgression to the heritability of complex traits. Nat. Commun 12, 4481 (2021).
Putilov, A. A., Dorokhov, V. B., Puchkova, A. N., Arsenyev, G. N. & Sveshnikov, D. S. Genetic-based signatures of the latitudinal differences in chronotype. Biol. Rhythm. Res. 50, 255–271 (2019).
Dannemann, M. et al. Neanderthal introgression partitions the genetic landscape of neuropsychiatric disorders and associated behavioral phenotypes. Transl. Psychiatry 12, 433 (2022).
Velazquez-Arcelay, K. et al. Archaic introgression shaped human circadian traits. Genome Biol. Evol. 15, evad203 (2023).
Archer, S. N. & Oster, H. How sleep and wakefulness influence circadian rhythmicity: effects of insufficient and mistimed sleep on the animal and human transcriptome. J. Sleep. Res. 24, 476–493 (2015).
Moore, R. Y. & Eichler, V. B. Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in the rat. Brain Res. 42, 201–206 (1972).
Dijk, D. J. & Czeisler, C. A. Contribution of the circadian pacemaker and the sleep homeostasis to sleep propensity, sleep structure, electroencephalographic slow waves, and sleep spindle activity in humans. J. Neurosci. 15, 3526–2538 (1995).
Achermann, P. & Borbély, A.A. Mathematical models of sleep regulation. Front. Biosci. 8, 683–693 (2003).
Saper, C. B., Scammell, T. E. & Lu, J. Hypothalamic regulation of sleep and circadian rhythms. Nature 437, 1257–1263 (2005).
Kalmbach, D. A. et al. Genetic basis of chronotype in humans: insights from three landmark GWAS. Sleep 40, 1–10 (2017).
Voigt, R. M., Forsyth, C. B. & Keshavarzian, A. Circadian rhythms: a regulator of gastrointestinal health and dysfunction. Expert Rev. Gastroenterol. Hepatol 13, 411–424 (2019).
Segers, A. & Depoortere, I. Circadian clocks in the digestive system. Nat. Rev. Gastroenterol. Hepatol. 18, 239–251 (2021).
Scheiermann, C., Kunisaki, Y. & Frenette, P. S. Circadian control of the immune system. Nat. Rev. Immunol. 13, 190–198 (2013).
Haspel, J. A. et al. Perfect timing: circadian rhythms, sleep, and immunity – an NIH workshop summary. JCI Insight 5, e131487 (2020).
Zeng, Y., Guo, Z., Wu, M., Chen, F. & Chen, L. Circadian rhythm regulates the function of immune cells and participates in the development of tumours. Cell Death Discov. 10, 199 (2024).
Roenneberg, T. et al. Epidemiology of the human circadian clock. Sleep Med. Rev. 11, 429–438 (2007).
Jones, S. E. et al. Genome-wide association analyses of chronotype in 697,828 individuals provides insights into circadian rhythms. Nat. Commun. 10, 343 (2019).
Burns, A. C. et al. Genome-wide gene by environment study of time spent in daylight and chronotype identifies emerging genetic architecture underlying light sensitivity. Sleep 46, zsac287 (2023).
Koenig, Z. et al. A harmonized public resource of deeply sequenced diverse human genomes. Genome Res. 34, 796–809 (2024).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Prüfer, K. et al. A high-coverage Neandertal genome from Vindija Cave in Croatia. Science 358, 655–658 (2017).
Mafessoni, F. et al. A high-coverage Neandertal genome from Chagyrskaya Cave. Proc. Natl. Acad. Sci. USA 117, 15132–15136 (2020).
Leocadio-Miguel, M. A. et al. Latitudinal cline of chronotype. Sci. Rep. 7, 5437 (2017).
Randler, C. & Rahafar, A. Latitude effects morningness-eveningness: evidence for the environment hypothesis based on a systematic review. Sci. Rep. 7, 39976 (2017).
Chen, F. et al. A late Middle Pleistocene Denisovan mandible from the Tibetan Plateau. Nature 569, 409–412 (2019).
Demeter, F. et al. A Middle Pleistocene Denisovan molar from the Annamite Chain of northern Laos. Nat. Commun. 13, 2557 (2022).
Yaworksy, P. M., Nielsen, E. S. & Nielsen, T. K. The Neanderthal niche space of Western Eurasia 145 ka to 30 ka ago. Sci. Rep. 14, 7788 (2024).
Bertoldi, M. Mammalian Dopa decarboxylase: structure, catalytic activity and inhibition. Arch. Biochem. Biophys. 546, 1–7 (2014).
Li, S. et al. CGDB: a database of circadian genes in eukaryotes. Nucleic Acids Res. 45, D397–D403 (2017).
Sollis, E. et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. 51, D977–D985 (2023).
Zhou, Y. & Browning, S. R. Protocol for detecting introgressed archaic variants with SPrime. STAR Protoc 2, 100550 (2021).
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Speidel, L., Forest, M., Shi, S. & Myers, S. R. A method for genome-wide genealogy estimation for thousands of samples. Nat. Genet. 51, 1321–1329 (2019).
Ferrer-Admetlla, A., Liang, M., Korneliussen, T. & Nielsen, R. On detecting incomplete soft or hard selective sweeps using haplotype structure. Mol. Biol. Evol. 31, 1275–1291 (2014).
Alachiotis, N. & Pavlidis, P. RAiSD detects positive selection based on multiple signatures of a selective sweep and SNP vectors. Commun. Biol. 1, 79 (2018).
DeGiorgio, M. & Szpiech, Z. A. A spatially aware likelihood test to detect sweeps from haplotype distributions. PLoS Genet. 18, e1010134 (2022).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Szpiech, Z. A., Novak, T. E., Bailey, N. P. & Stevison, L. S. Application of a novel haplotype-based scan for local adaptation to study high-altitude adaptation in rhesus macaques. Evol. Lett. 5, 408–421 (2021).
Hut, R. A., Paolucci, S., Dor, R., Kyriacou, C. P. & Daan, S. Latitudinal clines an evolutionary view on biological rhythms. Proc. Biol. Sci. 280, 20130433 (2013).
Bertolini, E. et al. Life at high latitudes does not require circadian behavioural rhythmicity under constant darkness. Curr. Biol. 29, 3928–3926.e3 (2019).
Muranaka, T., Ito, S., Kudoh, H. & Oyama, T. Circadian-period variation underlies the local adaptation of photoperiodism in the short-day plant Lemna aequinoctialis. iScience 25, 104634 (2022).
Cruciani, F. et al. Genetic diversity patterns at the human clock gene period 2 are suggestive of population-specific positive selection. Eur. J. Hum. Genet. 16, 1526–1534 (2008).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. elife 7, e34408 (2018).
Elsworth, B. et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv; https://doi.org/10.1101/2020.08.10.244293 (2020).
Chelala, C., Khan, A. & Lemoine, N. R. SNPnexus: a web database for functional annotation of newly discovered and public domain single nucleotide polymorphisms. Bioinformatics 25, 655–661 (2009).
Oscanoa, J. et al. SNPnexus: a web server for functional annotation of human genome sequence variation (2020 update). Nucleic Acids Res. 48, W185–W192 (2020).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 25, 25–29 (2000).
Gene Ontology Consortium et al. The Gene Ontology knowledgebase in 2023. Genetics 224, iyad031 (2023).
Harrison, P. W. et al. Ensembl 2024. Nucleic Acids Res. 52, D891–D899 (2024).
Ruth, K. S. et al. Using human genetics to understand the disease impacts of testosterone in men and women. Nat. Med. 26, 252–258 (2020).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
Randler, C. Morningness-eveningness comparison in adolescents from different countries around the world. Chronobiol. Int. 25, 1017–1028 (2008).
Mascal, E. et al. Effects of longitude, latitude and social factors on chronotype in Turkish students. Pers. Individ. Differ. 86, 73–81 (2015).
Dashti, H. S. et al. Genetic determinants of daytime napping and effects on cardiometabolic health. Nat. Commun. 12, 900 (2021).
Forni, D. et al. Genetic adaptation of the human circadian clock to day-length latitudinal variations and relevance for affective disorders. Genome Biol. 15, 499 (2014).
Walker II, W. H., Walton, J. C., DeVries, A. C. & Nelson, R. J. Circadian rhythm disruption and mental health. Transl. Psychiatry 10, 28 (2020).
Srinivasan, S. et al. Genetic markers of human evolution are enriched in schizophrenia. Biol. Psychiatry 80, 284–292 (2016).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Gregory, M. D. et al. Neanderthal-derived genetic variation in living humans relates to schizophrenia diagnosis, to psychotic symptom severity, and to dopamine synthesis. Am. J. Med. Genet. Part B: Neuropsychiatr. Genet. 186, 329–338 (2021).
Cardno, A. G. & Owne, M.J. Genetic relationships between schizophrenia, bipolar disorder, and schizoaffective disorder. Schizophr. Bull. 40, 504–515 (2014).
Mullins, N. et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet. 53, 817–829 (2021).
Scheer, F. A. J. L. et al. The endogenous circadian system worsens asthma at night independent of sleep and other daily behavioral or environmental cycles. Proc. Natl. Acad. Sci. USA 118, e2018486118 (2021).
Sun, N., Dai, D., Deng, S., Cai, X. & Song, P. Bioinformatics integrative analysis of circadian rhythms effects on atopic dermatitis and dendritic cells. Clin. Cosmet. Investig. Dermatol 16, 2919–2930 (2023).
Jagoda, E. et al. Detection of Neanderthal adaptively introgressed genetic variants that modulate reporter gene expression in human immune cells. Mol. Biol. Evol. 39, msab304 (2022).
Koller, D. et al. Denisovan and Neanderthal archaic introgression differentially impacted the genetics of complex traits in modern populations. BMC Biol. 20, 249 (2022).
Gao, Y. et al. A pangenome reference of 36 Chinese populations. Nature 619, 112–121 (2023).
Rong, S. et al. Large-scale functional screen identifies genetic variants with splicing effects in modern and archaic humans. Proc. Natl. Acad. Sci. USA 120, e2218308120 (2023).
Ferreira, M. A. et al. Shared genetic origin of asthma, hay fever, and eczema elucidates allergic disease biology. Nat. Genet 49, 1752–1757 (2017).
Mitchell, R. et al. MRC IEU UK Biobank GWAS pipeline version 2. [dataset]. https://doi.org/10.5523/bris.pnoat8cxo0u52p6ynfaekeigi (2019).
Patel, D. F. et al. Neutrophils restrain allergic airway inflammation by limiting ILC2 function and monocyte-dendritic cell antigen presentation. Sci. Immunol. 4, eaax7006 (2019).
Vuckovic, D. et al. The polygenic and monogenic basis of blood traits and diseases. Cell 182, 1214–1231.e11 (2020).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Marcheva, B. et al. Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 466, 627–631 (2010).
Kalsbeek, A., la Fleur, S. & Fliers, E. Circadian control of glucose metabolism. Mol. Metab. 3, 372–383 (2014).
Ferrell, J. M. & Chiang, J. Y. L. Circadian rhythms in liver metabolism and disease. Acta. Pharm. Sin. B 5, 113–122 (2015).
The SIGMA Type 2 Diabetes Consortium, et al. Sequence variants in SLC16A11 are a common risk factor for type 2 diabetes in Mexico. Nature 506, 97–101; (2014).
Watanabe, K., Taskese, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun 8, 1826 (2017).
Watanabe, K., Mirkov, M. U., de Leeuw, C. A., van den Heuvel, M. & Posthuma, D. Genetic mapping of cell type specificity for complex traits. Nat. Commun. 10, 3222 (2019).
Milacic, M. et al. The Reactome pathway knowledgebase 2024. Nucleic Acids Res. 52, D672–D678 (2024).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC. Bioinform. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr. A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc 1, e90 (2021).
Dannemann, M., Prüfer, K. & Kelso, J. Functional implications of Neanderthal introgression in modern humans. Genome Biol. 18, 61 (2017).
Enard, D. & Petrov, D. A. Evidence that RNA viruses drove adaptive introgression between Neanderthals and modern humans. Cell 175, 360–371 (2018).
Silvert, M., Quintana-Murci, L. & Rotival, M. Impact and evolutionary determinants of Neanderthal introgression on transcriptional and post-transcriptional regulation. Am. J. Hum. Genet 104, 1241–1250 (2019).
McCoy, R., Wakefield, J. & Akey, J. M. Impacts of Neanderthal-introgressed sequences on the landscape of human gene expression. Cell 168, 916–927.e12 (2017).
Chen, L., Wolf, A. B., Fu, W., Li, L. & Akey, J. M. Identifying and interpreting apparent Neanderthal ancestry in African individuals. Cell 180, 677–687 (2020).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, 1–4 (2021).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2006).
Hinrichs, A. S. et al. The UCSC Genome Browser database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: A grammar of data manipulation. Version 1.1.4. [software. https://cran.r-project.org/web/packages/dplyr/index.html (2023).
R Core Team. R: A language and environment for statistical computing. Version 4.1.2 [software]. https://www.R-project.org (2023).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Venables, W.N. & Ripley, B.D. Modern applied statistics with S (Springer, 2002).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Jakobsson, M., Edge, M. D. & Rosenberg, N. A. The relationship between FST and the frequency of the most frequent allele. Genetics 193, 515–528 (2013).
Willing, E.-M., Dreyer, C. & van Oosterhout, C. Estimates of genetic differentiation measured by FST do not necessarily require large sample sizes when using many SNP markers. PLoS ONE 7, e42649 (2012).
Szpiech, Z. A. selscan 2.0: scanning for sweeps in unphased data. Bioinformatics 40, btae006 (2024).
Klassmann, A. & Gautier, M. Detecting selection using extended haplotype homozygosity (EHH)-based statistics in unphased or unpolarized data. PLoS ONE 17, e0262024 (2022).
Leigh, J. W. & Bryant, D. PopART: full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
Sánchez-Quinto, F. & Lalueza-Fox, C. Almost 20 years of Neanderthal palaeogenetics: adaptation, admixture, diversity, demography and extinction. Philos. Trans. R. Soc. Lond. B. Biol. Sci 370, 20130374 (2015).
Villanea, F. A., Huerta-Sánchez, E. & Fox, K. ABO genetic variation in Neanderthals and Denisovans. Mol. Biol. Evol. 38, 3373–3382 (2021).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 1–16 (2015).
Lischer, H. E. L. & Excoffier, L. PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28, 298–299 (2012).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Maddison, D. R., Swofford, D. L. & Maddison, W. P. NEXUS: an extensible file format for systematic information. Syst. Biol. 46, 590–621 (1997).
Rozas, J. et al. DnaSP 6: DNA Sequence Polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Nielsen, R., Vaughn, A. H. & Deng, Y. Inference and applications of ancestral recombination graphs. Nat. Rev. Genet. 26, 47–58 (2025).
Wickham, H. ggplot2: Elegant graphics for data analysis. (Springer-Verlag, 2016).
Yin, L. et al. rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genom. Proteom. Bioinform. 19, 619–629 (2021).
Lewis, M. J. & Wang, S. locuszoomr: an R package for visualizing publication-ready regional gene locus plots. Bioinform. Adv 5, vbaf006 (2025).
Massicotte, P. & South, A. rnaturalearth. Version 1.0.1.9000 [software]. https://docs.ropensci.org/rnaturalearth/ (2024).
Pebesma, E. Simple features for R: standardized support for spatial vector data. R J. 10, 439–446 (2018).
Pebesma, R. & Bivand, R. Spatial data science: With applications in R. 1st ed. (Chapman and Hall/CRC, 2023).
Acknowledgements
C.K. discloses support for the research provided in this work by NSERC [grant number CGSD2-535025-2019]. D.S. and E.P. are supported by NSERC [grant numbers DGECR-2020-00176 and RGPIN-2024-04911, respectively]. M.M. is funded by the CGEn HostSeq/CIHR fellowship [CGE 185054] and the SickKids Restracomp Fellowship. This research was enabled in part by support provided by Compute Ontario (https://www.computeontario.ca/) and the Digital Research Alliance of Canada (https://alliancecan.ca/en/).
Author information
Authors and Affiliations
Contributions
C.K., D.S., E.P., B.V., and M.A.S. designed the study, ran data analysis, and wrote the manuscript. C.K. and G.D. collected the data. C.K., V.C.F.R., M.M., E.P., B.V., and M.A.S. devised the methodology. C.K. generated all figures and supplemental material. A.N. generated code for the analysis. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kendall, C., Nooranikhojasteh, A., Debortoli, G. et al. Adaptive introgression in modern human circadian rhythm genes. npj Biol Timing Sleep 2, 41 (2025). https://doi.org/10.1038/s44323-025-00060-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44323-025-00060-2