Abstract
Type 2 diabetes is a global health burden driven by genetic and environmental factors. Continuous glucose monitoring (CGM) can effectively guide lifestyle interventions in non-diabetic. However, predefined CGM metrics fail to fully capture the dysglycemic information contained in the high-dimensional time-series CGM data. This study employed deep learning to learn dysglycemia features from CGM data associated with diabetes and derived a digital biomarker of dysglycemia, validated against traditional dysglycemic biomarkers and diabetes polygenic risk score (PRS). Output of the deep learning model, called the deep learning-score, was significantly associated with multiple existing dysglycemic biomarkers and PRS of diabetes (P = 0.007). Moreover, existing CGM metrics were not associated with prevalent diabetes after adjusting for the deep learning-score, while the deep learning-score remained significantly associated with prevalent diabetes (P < 0.001) in a regression analysis. This digital biomarker demonstrated potential for providing dynamic feedback on dysglycemia and improving long-term intervention adherence.
Similar content being viewed by others
Introduction
Type 2 diabetes (T2D) is a complex disease resulting from the combined effects of environmental and genetic factors1. It is estimated that by 2045, the number of patients with diabetes will reach 783 million, and T2D will account for approximately 90% of all diabetes patients2. T2D is associated with a range of microvascular and macrovascular complications, and it is one of the leading causes of death globally2,3. The Diabetes Prevention Program has demonstrated that adopting healthy lifestyle interventions can reduce the risk of T2D by 58% compared to the control groups4. Large-scale genome-wide association studies have identified hundreds of genetic risk loci for T2D, elucidating the significant contribution of genetic factors to T2D pathogenesis5,6. Early identification of high-risk populations based on T2D genetic risk, followed by lifestyle interventions, represents an effective preventive strategy7. However, lifestyle interventions often reported poor adherence that hindered the effectiveness of T2D prevention8,9.
Dynamic feedback is an effective behavior change technique that could improve adherence to a healthy lifestyle10,11. For instance, interventions using wearable devices can facilitate dynamic feedback on daily step counts, having shown significant improvements in physical activity12,13,14. The primary metabolic characteristic of T2D is dysglycemia refers to hypoglycemia, hyperglycemia, and abnormal glucose variability, stemming from β-cell dysfunction and insulin resistance15. Providing dynamic feedback on the degree of dysglycemia could enhance adherence to healthy lifestyle interventions. However, traditional methods for assessing dysglycemia, such as fingerstick blood tests and glycated hemoglobin (HbA1c) measurements, provide only a snapshot of glucose levels at a specific time due to invasiveness.
Minimally invasive continuous glucose monitoring (CGM) devices can continuously record blood glucose at intervals of 5–15 min over several days16. CGM is regarded as the roadmap for 21st-century diabetes therapy. With the growing availability and decreasing costs of CGM, applications of CGM in non-diabetic individuals are receiving significant attention17,18. CGM provides dynamic monitoring of multiple dysglycemic metrics, including hypoglycemia, hyperglycemia, and glucose variability. However, an excessive number of dysglycemia metrics may complicate clinical interpretation19. Moreover, reliance on any single CGM metric may fail to comprehensively reflect the overall degree of dysglycemia.
The multiplicity of CGM metrics stems from the fact that CGM data is high-dimensional time-series data. Each dimension—representing glucose values at different time points—contains partial information about dysglycemia. Various linear or nonlinear combinations of these dimensions give rise to numerous CGM metrics. Deep learning (DP), a class of algorithms capable of learning task-related features directly from high-dimensional data without predefined features20. DP have been effectively employed to derive digital biomarkers from types of high-dimensional medical data, including photoplethysmography and electrocardiograms21,22.
In this study, we employed DP to learn dysglycemia features from CGM data associated with diabetes and derived a digital biomarker of dysglycemia. We validated this digital biomarker across independent datasets by examining its correlations with established dysglycemic biomarkers, including conventional CGM metrics and HbA1c. Given that dysglycemia represents a core feature of elevated diabetes risk, we utilized the T2D polygenic risk score (PRS) as a genetic proxy for T2D susceptibility to validate that the deep learning-derived digital biomarker of dysglycemia reflects T2D risk.
Results
Dataset characteristics
Characteristics of the primary dataset and validation dataset are shown in Table 1. In both the primary and validation datasets, the age and HbA1c levels of patients with diabetes were significantly higher than those of individuals without diabetes.
DP model performance
We evaluated the performance of the DP model in terms of sensitivity, specificity, and AUROC. Considering that each participant contributed multiple days of data, the performance evaluation of the DP model was conducted on an “individual-level”, averaging the DP-scores for each participant. In the primary test dataset, the AUROC of the DP model to predict diabetes was 0.99 (95% CI: 0.97–1.00; Supplementary Table 1). In the validation dataset, the AUROC of the DP model was 0.89 (95% CI: 0.80–0.97; Supplementary Table 1). Calibration plots showed good agreement between observed and predicted probability of T2D, and the Brier score was 0.044 in the validation dataset (Supplementary Fig. 1). The accurate differentiation between diabetic and non-diabetic populations demonstrates that the DP model has successfully learned diabetes-related dysglycemic features from CGM data. The distribution of the DP score in the validation is shown in Supplementary Fig. 2. As expected, because most participants were non-diabetic, the DP score exhibited a left-skewed distribution with a right-tailed trend. These results indicate that the DP could reflect the population-level risk of diabetes.
Association of DP-score with HbA1c and CGM metrics
In the validation dataset, results revealed a significant association between the DP-score and HbA1c (β = 0.60, 95% CI: 0.50–0.70, P < 0.001). After adjusting for age, sex, and BMI, the DP-score was still associated with HbA1c (\(\beta\) = 0.48, 95% CI: 0.38–0.59, P < 0.001). We conducted an analysis to evaluate the association between the DP-score and glycemic measures, including mean glucose, glucose variability, and specifically the mean amplitude of glucose excursions (MAGE). DP-score was significantly associated with both MAGE (β = 0.91, 95% CI: 0.87–0.96, P < 0.001) and mean glucose (β = 0.61, 95% CI: 0.51–0.70, P < 0.001). Same results were observed in the primary test dataset (Supplementary table 2 and table 3). The association between the DP-score and conventional biomarkers of dysglycemia suggested that the DP-score captures information about dysglycemia in CGM data.
The AUROC (area under the receiver operating curve) and AUPRC (area under the precision-recall curve) of the DP-score and the commonly used CGM metrics were compared based on logistic regression in the validation dataset (Supplementary table 8). Results showed that both the AUROC and AUPRC of the DP-score were higher than those of CGM metrics, indicating that the deep learning approach was able to capture more information on glycemic dysregulation from high-dimensional CGM time-series data and thereby achieve more accurate identification of patients with diabetes. The classification threshold that maximized the sum of sensitivity and specificity was calculated. At this threshold, the sensitivity was 0.75 (95% CI: 0.48–0.93) and the specificity was 0.90 (95% CI: 0.85–0.93). The AUPRC of the DP-score is 0.56 (95% CI: 0.30–0.78).
Moreover, we analyzed the correlations of the DP-score with Time above Range (TAR), Time in Range (TIR), and Time below Range (TBR) in diabetic patients from both the test and validation sets. The results showed that the DP score was significantly correlated with TAR (β = 0.67, 95% CI: 0.50–0.83, P < 0.001) and TIR (β = -0.68, 95% CI: -0.84– -0.50, P < 0.001), but not with TBR. Considering the clinical implications of TIR, TAR, and TBR, our findings suggest that the DP score reflects glycemic control capacity and hyperglycemia risk in patients with diabetes, but does not capture hypoglycemia risk.
We compared the effect size of the DP-score and conventional CGM metrics for prevalent T2D in the validation dataset (Supplementary table 4 and table 5). The DP-score was significantly associated with prevalent T2D (Odds ratio (OR) of DP-score: 3.61, 95% CI: 2.36–5.91, P < 0.001). Even after adjusting for age, sex, and BMI, the association between the DP-score and prevalent T2D remained significant (OR of DP-score: 2.77, 95% CI: 1.71–4.77, P < 0.001). Moreover, even after adjusting for MAGE and mean glucose, the DP-score remained significantly associated with prevalent T2D (OR of DP-score: 3.46, 95% CI: 1.24–0.21, P = 0.018), whereas MAGE and mean glucose did not exhibit significance (P = 0.838, P = 0.467). These findings indicated that the DP-score provides additional information about diabetic dysglycemia beyond what is captured by traditional CGM metrics.
Association of DP-score with PRS of T2D
DP-score was significantly associated with PRS of T2D (\(\beta\) = 0.19, 95% CI: 0.07–0.30, P = 0.002), and the PRS explained 3.07% of variance of the DP-score (Supplementary table 6). After adjusting for age, sex, and BMI, the DP-score was associated with the PRS (\(\beta\) = 0.14, 95% CI: 0.04–0.25, P = 0.007), suggesting that the DP-score captures genetic risk of T2D.
Dynamic trajectory of DP-score
To characterize the dynamic nature of the DP-score, we analyzed non-diabetic individuals with consecutive 7-day CGM monitoring (223 participants, 1561 days) to visualize DP-score trajectories (Fig. 1). Among them, 212 participants (95.07%) exhibited at least one transition between DP-score quartiles during the observation period, with an average daily transition rate of 63.53% to different quartiles (Fig. 1). The trajectory patterns of DP-score revealed substantial day-to-day variability in dysglycemia among non-diabetic individuals, underscoring the critical need for dynamic monitoring to capture these fluctuations.
The dynamic trajectory of the DP-score was examined by dividing the selected population based on the 25, 50, and 75th percentiles of the DP score into four groups: Q1 (first quartile), Q2 (second quartile), Q3 (third quartile), and Q4 (fourth quartile). For each day, the number of individuals transitioning from one group to another was calculated, and the width of the lines in the visualization was proportional to the number of transitions.
However, 11 participants (4.93%) persistently remained in the highest DP-score quartile (Q4) throughout all 7 days. Notably, their polygenic risk scores (PRS) for T2D were significantly elevated compared to other participants (P = 0.007), suggesting this subgroup may represent a high-risk population for T2D (Fig. 2).
Subgroup A is the participants who remained in the highest quarter (Q4) of DP-score for all 7 days. Subgroup B is the participants who transitioned at least once during 7 days.
Visual explanation of DP model
To shed light on the mechanisms underlying how the DP-score captures dysglycemia, we utilized the Grad-CAM method to generate a coarse localization map highlighting important regions in the CGM data for the measurement of the DP-score. As depicted in Fig. 3, the important regions were predominantly distributed during specific time intervals, from 04:30 to 05:30, 06:30 to 08:00, and 12:00 to 14:00 (Fig. 3). These intervals correspond to fasting glucose and postprandial glucose levels, which are not only dysglycemic features but also key components of T2D diagnosis according to ADA Diagnosis Criteria23. These findings suggested that DP-score is highly interpretable for assessing the degree of dysglycemia.
The red line represents the average glucose of all participants in the validation dataset. Red shaded regions represent the standard deviation of the glucose level. Blue shaded regions represent the weight of relevant regions for DP-score.
Discussion
In this study, we have discovered a digital biomarker of dysglycemia from CGM signals using a deep learning model. Results demonstrated consistent correlations between this digital biomarker and conventional dysglycemic biomarkers across multiple independent datasets. Furthermore, our findings revealed associations of this biomarker with genetic risk of T2D, indicating its potential as a digital biomarker of dysglycemia associated with T2D risk.
Studies have identified multiple CGM metrics that correlate with diabetic dysglycemia. For example, postprandial hyperglycemia is one of the earliest abnormalities of glucose homeostasis associated with T2D24. Glucose variability was associated with insulin resistance and T2D risk25. However, a single CGM metric cannot fully capture the comprehensive dysglycemic information contained in high-dimensional CGM time-series data. This study utilized a deep learning model to extract dysglycemia features from CGM data and derived a digital biomarker. Visual explanations revealed that fasting glucose and postprandial glucose played pivotal roles in shaping the DP-score. The elevation of fasting and postprandial glucose levels is associated with insulin resistance and diminished insulin secretion, which are central to the pathogenesis of T2D. These findings indicate that the deep learning model effectively integrated multiple key features of diabetic dysglycemia present in CGM signals into the DP-score. Moreover, the DP-score provides additional information about diabetic dysglycemia beyond traditional CGM metrics. Adoption of this digital biomarker for assessing dysglycemia offers dual advantages: it not only reduces the complexity associated with interpreting multiple CGM metrics but also enables more precise evaluation of dysglycemia.
A healthy lifestyle, including regular physical activity and a balanced diet, is the first-line approach to preventing T2D26. Prior studies have shown that providing dynamic feedback can be an effective behavior change technique, enhancing adherence to healthy lifestyle practices10,11. The DP-score could serve as a tool to alert individuals about degree of dysglycemia, prompting them to adopt healthy lifestyle interventions. Its dynamic nature allows for continuous reflection of dysglycemia, motivating individuals to adhere healthy lifestyle.
As the earliest measurable risk factor for T2D, PRS enables early identification of high-risk populations, allowing offset this risk by maintaining personalized lifestyle interventions. For example, a study observed that a sharp reduction of 23% in T2D risk was associated with an increase in diet quality among participants in the highest PRS group27. Significant correlation between DP-score and T2D PRS indicated the DP-score’s potential for early identification of high-risk populations. Specifically, the dynamic trajectory of DP-score in non-diabetic individuals revealed that those maintaining DP-score in the highest quarter exhibited greater genetic risk than those of other participants, suggesting this subgroup needs to adopt lifestyle interventions.
Although CGM was originally developed for diabetes management, its application in non-diabetic populations has garnered increasing attention in recent years. Research in this field primarily focuses on utilizing CGM as a health education tool for non-diabetic individuals to provide lifestyle guidance regarding diet and physical activity28. For instance, one study employed artificial intelligence to develop personalized postprandial glucose response prediction models based on CGM data, enabling personalized dietary recommendations29. A series of clinical studies have demonstrated that CGM technology can improve glycemic control in non-diabetic populations through diet intervention facilitation30,31. The dysglycemia of digital biomarker identified in our study may serve as an evaluative measure for glycemic control in non-diabetic individuals, providing dynamic feedback on healthy lifestyle intervention efficacy and thereby further enhancing adherence. Emerging non-invasive CGM technologies will significantly improve the cost of CGM and promote the application of this CGM-derived digital biomarker for T2D prevention32.
This study has several limitations. While we validated the deep learning model across two CGM datasets from distinct populations and observed a high degree of consistency in the results, the validation dataset contained only a limited number of T2D patients, which may constrain statistical power and limit generalizability. Additional validation using larger and more diverse datasets is therefore warranted. The dynamic trajectory analysis of the DP score provided a descriptive depiction of the progressive patterns of glycemic dysregulation in non-diabetic individuals; future work should incorporate more rigorous quantitative approaches to better characterize these complex dynamic processes.
In conclusion, this study employed deep learning to derive a digital biomarker of dysglycemia from CGM data, which was associated with the genetic predisposition to T2D. This digital biomarker demonstrates potential for providing dynamic feedback on glycemic control efficacy following lifestyle interventions in non-diabetic populations, and improving long-term intervention adherence through quantitative monitoring.
Methods
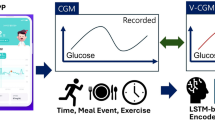
We collected two independent CGM datasets (primary dataset and validation dataset) to train and validate a DP model. The primary dataset, sourced from the A1C-DERIVED AVERAGE GLUCOSE study, comprised data used for constructing the DP model33. We recruited 289 participants and collected their CGM data as the validation dataset to validate the generalizability of the DP model. Given that a circadian cycle spans 24 h, daily CGM data served as input for the DP model, yielding a DP-score ranging from 0 to 1. A higher DP-score indicates both a greater probability of diabetes and more severe dysglycemia.
As a digital biomarker for dysglycemia, the “DP-score” was validated in 3 steps1: by validating DP model performance for the identification of prevalent T2D2, by revealing associations between the DP-score and dysglycemic biomarkers, and3 by testing associations between the DP-score and PRS of T2D. Figure 4 shows a summary of the study design.
Summary of methods.
Data collection and preprocessing
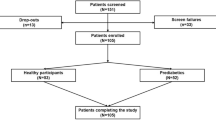
The primary dataset consisted of 507 participants whose age ranged from 18 to 70 years. CGM (Medtronic Minimed, Northridge, CA) was used to continuously monitor blood glucose, which measures glucose levels every 5 min over 3 days. After excluding days with missing data from the primary dataset, 502 participants and 3113 days remained. Subsequently, individuals in the primary dataset were randomly divided into a training dataset (n = 351, 70%) and a test dataset (n = 151, 30%). The training dataset was utilized for training DP model, whereas the test dataset was employed to evaluate the performance of the DP model. 80% of the training dataset was used to train DP model, and 20% of the training dataset was used to tune the hyperparameters of DP. Performance of DP model was evaluated in the test dataset.
In order to validate generalizability of DP model, we recruited 289 participants (validation dataset) in the physical examination center at Chongqing Fifth People’s Hospital between 1 April 2020 and 31 October 2020. The participants began wearing the CGM device following their physical examination. The diabetes status of the recruited participants was obtained from the electronic health records system using the ICD-10 code (E11–E14). The CGM device used was the Abbott FreeStyle Libre, which monitored blood glucose levels every 15 min over a period of 14 days. To address concerns regarding the accuracy of glucose level detection, the CGM data from the first and last three days were excluded, as studies have indicated reduced accuracy during these periods34. Four participants were excluded due to sensor failure, resulting in a total of 264 participants and 1759 days of CGM data remaining. To standardize the number of blood glucose values per day in the primary dataset with that of the validation dataset, the time intervals of blood glucose values in the primary dataset were adjusted to 15 min using a method that involved filtering out two out of every three blood glucose values.
The study was observational in design. Participants were masked to the CGM data. The study protocol was approved by the ethics committees of Chongqing Fifth People’s Hospital in accordance with the principles of the Declaration of Helsinki. Written informed consent was obtained from each participant before recruitment.
DP model development
A residual convolutional deep neural network DP model was constructed to distinguish participants with T2D from those without T2D. The DP model consisted of 31 convolutional layers and accepted raw CGM data in one day (96 glucose values per day) as input. Each residual connection contained two convolutional layers, and a total of 15 residual connections were utilized. The filter length of convolutional layers was 11, with the number of filters starting at 32 and doubling every four shortcut connections. After each convolutional layer, batch normalization, rectified linear activation, and dropout with a probability of 0.2 were applied. Finally, the fully connected layer used SoftMax to output probabilities. The weights of the DP model were initialized as described by He et al.35. Adam optimizer with the default parameters36, and a mini batch size of 128 was employed. The learning rate began at 0.001 and was multiplied by a factor of 0.1 when the loss of the training dataset ceased to improve for two consecutive epochs.
We used gradient-weighted class activation mapping (Grad-CAM) to identify the important CGM regions for the decision of DP model37. Grad-CAM utilizes the gradients of the output probability with respect to the final convolutional layer to generate a weighted localization map, which highlights the significant regions in the CGM data contributing to the output probability of the DP model. This weighted localization map was generated using the validation dataset.
Evaluation of DP model performance
The predictive performance of the DP model was evaluated using the area under the receiver operating curve (AUROC), sensitivity, and specificity. The DeLong method was used to calculate the confidence interval of AUROC38. Additionally, the 95% confidence intervals of sensitivity and specificity were estimated through bootstrapping with 1000 iterations.
Calculation of PRS
PRS aggregates genetic variants contributing to T2D risk into a single quantitative measure39. All blood samples were genotyped with Illumina Infinium Global Screening Array. After quality control procedures, 468,322 variants and 289 participants remained. Subsequently, imputation resulted in a total of 17,154,889 variants. C + T method was used to calculate PRS of T2D. The single nucleotide polymorphisms (SNPs) and their respective effect sizes were derived from a large-scale genome-wide association study conducted in a Japanese population5. The SNPs with linkage disequilibrium > 0.2 and P value > 5e-8 were filtered out, a total of 339 SNPs remained. Linkage disequilibrium was estimated in a whole genome sequencing data from 2638 Chinese population. PRS was calculated by PRSice-2 (version 2.3.3)40.
Statistical analysis
Logistic regression was utilized to examine the association between DP-score and prevalent T2D. Linear regression was applied to evaluate the correlation between DP-score and HbA1c, PRS of T2D and CGM metrics. To facilitate comparison between coefficients in both linear regression and logistic regression, all continuous variables were standardized using Z scores. A P-value of < 0.05 (2-tailed) was considered statistically significant. All analyses were done using Python (version 3.7) with the following libraries: keras (version 2.2.4); tensorflow (version 2.4.0); pandas (version 0.24.2); seaborn (version 0.9.0); sklearn (version 0.21.2), and R (version 3.6.2) with the following libraries: pROC (version 1.16.2); glmnet (version 4.1).
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The code that supports the findings of this study is available from the corresponding author upon reasonable request.
References
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nat., Nat. Publ. Group 536, 41–47 (2016).
Sun, H. et al. IDF Diabetes Atlas: global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diab. Res. Clin. Pract. 183, 109119 (2022).
Mulnier, H. E. et al. Mortality in people with Type 2 diabetes in the UK. Diabet. Med. 23, 516–521 (2006).
Reduction in the Incidence of Type 2 Diabetes with Lifestyle Intervention or Metformin. New England Journal of Medicine, Massachusetts Medical Society, 2002, 346: 393–403.
Ishigaki, K. et al. Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nature Genetics. Nat. Publ. Group 52, 669–679 (2020).
Vujkovic, M. et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nature Genetics. Nat. Publ. Group 52, 680–691 (2020).
Liu, J. et al. Polygenic Risk Score, Lifestyles, and Type 2 diabetes risk: a prospective chinese cohort study. Nutrients, Multidiscip. Digital Publ. Inst. 15, 2144 (2023).
Burgess, E., Hassmén, P. & Pumpa, K. L. Determinants of adherence to lifestyle intervention in adults with obesity: a systematic review. Clin. Obes. 7, 123–135 (2017).
Brickwood, K.-J. et al. Consumer-based wearable activity trackers increase physical activity participation: systematic review and meta-analysis. JMIR mHealth uHealth 7, e11819 (2019).
Michie, S. et al. The behavior change technique taxonomy (v1) of 93 hierarchically clustered techniques: building an international consensus for the reporting of behavior change interventions. Ann. Behav. Med. 46, 81–95 (2013).
Michie, S. et al. Effective techniques in healthy eating and physical activity interventions: A meta-regression[J]. Health Psychology, US: American Psychological Association, 2009, 28: 690–701.
Laranjo, L. et al. Do smartphone applications and activity trackers increase physical activity in adults? Systematic review, meta-analysis and metaregression. Br. J. Sports Med., BMJ Publ. Group Ltd Br. Assoc. Sport Exerc. Med. 55, 422–432 (2021).
Bravata, D. M. et al. Using pedometers to increase physical activity and improve health: a systematic review. JAMA 298, 2296 (2007).
Hodkinson, A. et al. Interventions using wearable physical activity trackers among adults with cardiometabolic conditions: a systematic review and meta-analysis. JAMA Netw. Open 4, e2116382 (2021).
Lima, J. E. B. F., Moreira, N. C. S. & Sakamoto-Hojo, E. T. Mechanisms underlying the pathophysiology of type 2 diabetes: from risk factors to oxidative stress, metabolic dysfunction, and hyperglycemia. Mutat. Res./Genet. Toxicol. Environ. Mutagenesis 874–875, 503437 (2022).
Klonoff D. C. Continuous Glucose Monitoring. DIABETES CARE, 28 (2005).
Keshet, A. et al. CGMap: characterizing continuous glucose monitor data in thousands of non-diabetic individuals. Cell Metab. 35, 758–769 (2023).
Hall, H. et al. Glucotypes reveal new patterns of glucose dysregulation. PLOS Biol., Public Libr. Sci. 16, e2005143 (2018).
Nguyen M., et al. A review of continuous glucose monitoring-based composite metrics for glycemic control. diabetes technology & therapeutics, Mary Ann Liebert, Inc., publishers, 2020, 22: 613–622.
Sze, V. et al. Efficient processing of deep neural networks: a tutorial and survey. Proc. IEEE 105, 2295–2329 (2017).
Avram, R. et al. A digital biomarker of diabetes from smartphone-based vascular signals. Nature Medicine. Nat. Publ. Group 26, 1576–1582 (2020).
Al-Alusi, M. A. et al. A deep learning digital biomarker to detect hypertension and stratify cardiovascular risk from the electrocardiogram. npj Digital Medicine. Nat. Publ. Group 8, 1–9 (2025).
American Diabetes Association Professional Practice Committee. 2. Classification and diagnosis of diabetes: standards of medical care in diabetes—2022. Diab. Care 45, S17–S38 (2021).
Holst, J. J. et al. Loss of incretin effect is a specific, important, and early characteristic of Type 2. Diab J. Diab Care 34, S251–S257 (2011).
Kim, J. A. et al. Impact of Visit-to-Visit Fasting Plasma Glucose Variability on the Development of Type 2 Diabetes: a nationwide Population-Based Cohort Study[J]. Diabetes Care, 2610–2616 (American Diabetes Association, 2018).
Tuomilehto, J. et al. Prevention of Type 2 diabetes mellitus by changes in lifestyle among subjects with impaired glucose tolerance. N. Engl. J. Med., Mass. Med. Soc. 344, 1343–1350 (2001).
Zhuang P., et al. Effect of Diet Quality and Genetic Predisposition on Hemoglobin A1c and Type 2 Diabetes risk: gene-diet interaction analysis of 357,419 Individuals[J]. Diabetes Care, 2470–2479 (American Diabetes Association, 2021).
Holzer, R., Bloch, W. & Brinkmann, C. Continuous glucose monitoring in healthy adults—possible applications in health care, wellness, and sports. Sens., Multidiscip. Digital Publ. Inst. 22, 2030 (2022).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Ben-Yacov, O. et al. Personalized postprandial glucose response–targeting diet versus mediterranean diet for glycemic control in prediabetes. Diab. Care 44, 1980–1991 (2021).
Zahedani, A. D. et al. Digital health application integrating wearable data and behavioral patterns improves metabolic health. npj Digital Medicine. Nat. Publ. Group 6, 1–15 (2023).
Bent, B. et al. Engineering digital biomarkers of interstitial glucose from noninvasive smartwatches[J]. npj Digital Medicine. Nat. Publ. Group 4, 1–11 (2021).
Nathan, D. M. et al. Translating the A1C assay into estimated average glucose values. Diab. Care 31, 1473–1478 (2008).
Hoss, U. & Budiman, E. S. Factory-calibrated continuous glucose sensors: the science behind the technology. Diab. Technol. Therapeutics, Mary Ann. Liebert, Inc., publishers 19, S–44 (2017).
He K., et al. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification// 1026–1034 (2015).
Liu, L. et al. On the variance of the adaptive learning rate and beyond. 2019.
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization// 2017: 618–626.
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988).
Torkamani, A., Wineinger, N. E. & Topol, E. J. The personal and clinical utility of polygenic risk scores. Nat. Rev. Genet., Nat. Publ. Group 19, 581–590 (2018).
Choi, S. W., O’Reilly, P. F. PRSice-2: Polygenic Risk Score software for biobank-scale data[J]. GigaScience, Oxford Academic, 2019, 8(7).
Acknowledgements
This study was funded by the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2024ZD0531900 & 2024ZD0531903), the National Key R&D Program of China (2022YFB3203704), Suzhou Science and Technology Project (SKY2022025), and Kunshan Social Development Science and Technology Project (KS2214).
Author information
Authors and Affiliations
Contributions
All authors read and approved the final manuscript. J.S., Y.P., S.Y.L., and K.X.Z. conceptualized and designed the study. J.S., Y.F.L., Y.Z., and H.Y.F. collected data. J.S., Z.D.N., and H.Y.G. developed the algorithm, and J.W., J.N.X, Z.J.T., and H.N.P analyzed the data. All the authors drafted the manuscript. J.S. and K.X.Z. revised the manuscript critically.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shao, J., pan, Y., Xue, J. et al. A deep learning-derived digital biomarker of dysglycemia and its association with genetic risk of type 2 diabetes. npj Metab Health Dis 3, 46 (2025). https://doi.org/10.1038/s44324-025-00089-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44324-025-00089-8