Abstract
Cas13a-based diagnostic systems have been widely utilized for the detection of RNA targets. However, without preamplification such systems are difficult to realize ultrasensitive (RT-PCR level) single pot RNA detection. Here, we found that double strand RNA can effectively activate the trans-cleavage activity of Cas13a RNP, while the cleavage rates of dsRNA by activated Cas13a RNP are very low. In addition, specially designed RNA-Nanocircle has limited ability to activate Cas13a RNP, but this activation is restored once the circular structures are cleaved and become linear. Based on this original method to control trans-cleavage activity of Cas13a RNP, we developed a Cas13a autocatalytic biosensing system assisted by RNA-Nanocircles, which allows one target RNA to activate numerous Cas13a RNPs. With this approach we show ultrasensitive detection of 1aM of synthetic RNA targets without preamplification within 15 min. The clinical utility of this biosensor was validated by monitoring miRNA-21 levels in plasma samples from colorectal cancer patients. This innovative approach highlights the potential of Cas13a-based biosensors in precision oncology, offering a rapid, non-invasive, and ultrasensitive method for RNA biomarker detection in liquid biopsies.
Similar content being viewed by others
Introduction
Cas13a is a programmable RNA-guided ribonuclease1,2. When bound to a target RNA, Cas13a displays both sequence-specific RNA target cleavage (cis-cleavage) and collateral cleavage (trans-cleavage) characteristics1,2. The trans-cleavage is realised by the HEPN catalytic site revealed upon Cas13a activation which is capable of non-specific cleavage of single-strand RNA (ssRNA)2. This can be exploited to create sequence-specific RNA biosensors; however basic Cas13a biosensing systems which recognise the presence of RNA targets by cleaving ssRNA fluorescent quenched reporters show limited sensitivity in the pM range3,4. Thus, nucleic acid preamplification techniques have been required to realize ultrasensitive (aM level) detection of RNA targets in Cas13a systems, such as recombinase polymerase amplification (RPA) in the specific high-sensitivity enzymatic reporter unlocking (SHERLOCK) system3,5 and Loop-mediated Isothermal Amplification (LAMP) in the diagnostics with coronavirus enzymatic reporting (DISCoVER) system6. These nucleic acid amplification methods require specialised laboratories/equipment, and they also suffer from false positive readings7,8 and amplicon contamination9. The requirement for amplification currently hinders the creation of convenient, low-cost assays with the requisite specificity and sensitivity for nucleic acid analysis in clinical settings. Therefore, there is a demand for sequence-specific detection methods of RNA targets that do not rely on amplification.
Up until now, two primary types of approaches for detecting RNA targets without preamplification have been reported. One type of approach is to integrate diverse signal amplification techniques, such as polydisperse droplet digital assay10, mobile phone microscopy11, nanoenzyme12, or field-effect transistors13 with CRISPR/Cas systems. These methods either require specialised instrumentation or are difficult to realize a point-of-care setting. Another type of approach is to employ the Cas tandem system, where the first Cas protein performs target recognition, and a second Cas protein realises cascade signal amplification; examples include Cas13-Csm63,14, Cas13-Cas1415, and Cas13-Cas12 systems16,17. In comparison with a basic Cas13a system, where one target activates one Cas13a RNP, the Cas tandem systems are able to achieve aM sensitivity but only with 2-3 hour incubation time3,16. However, in point-of-care test (POCT) setting, the aM level sensitivity at less than 15–20 min incubation time is expected. Furthermore, tandem systems are not single pot reactions which critical for end-user friendly sensors. An autocatalytic reaction such as that reported in OUR PAPER18 is not only single-pot but is also expected to further increase biosensing performance, because one target will, in principle, be able to activate unlimited Cas RNPs18,19,20. To date, a Cas13 autocatalytic reaction system has not been reported.
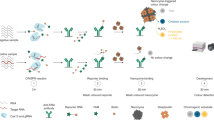
In this study, we investigated fundamental properties of Cas13a nuclease, including the ability of ssRNA, dsRNA, and circular RNA to activate the Cas13a RNP as well as the trans-cleavage activity of Cas13a on ssRNA, dsRNA, and circular RNA substrates. We found that dsRNA is able to activate a Cas13a RNP, while such activated Cas13a RNP was not able to cleave dsRNA. In addition, circular RNA could activate Cas13a only to a very limited degree, while after linearization the activation property has been recovered. Based on these findings, we developed an innovative type of autocatalytic biosensor (termed “autosensor”) based on Cas13a. Its key component is a molecular construct called “RNA-Nanocircle” which consists of a dsRNA region acting as a replicate activation trigger for Cas13a, and a short ssRNA linker which controls the activation property of dsRNA. The RNA-Nanocircle facilitates the autocatalysis reaction in the following way. In the presence of a target RNA molecule, one Cas13a RNP is activated, which continues to cleave the ssRNA region of the supplied RNA-Nanocircles to release the dsRNA triggers, which act as surrogates of the original detection targets. These triggers then activate new Cas13a RNPs. In this way, one target can theoretically activate an unlimited number of Cas RNPs in the reaction system. We show here that the optimized Cas13a autosensor is able to detect 1aM miRNA targets from clinical CRC samples within 15 min. Our autosensor provides an original platform for point-of-care testing of RNA targets without preamplification.
Results
Double strand RNA capable of triggering activation of Cas13a RNP
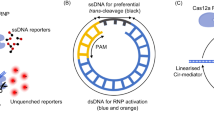
To date, Cas13a has been reported to be effectively activated by ssRNA (Fig. S1)1. To explore alternative triggers of Cas13a (Fig. 1A), we investigated the previously unexplored types of single-stranded nucleic acids targets such as single-stranded DNA (ssDNA) and phosphorothioate-modified single strand RNA (ps-ssRNA). As shown in Fig.1B & Fig. S2A, ssDNA was unable to activate Cas13a, while a slight signal increase was observed on ps-ssRNA (7.5%). A range of double-stranded nucleic acids were also investigated (Fig.1C & Fig. S2B), including double-stranded RNA (ssRNA-cRNA), RNA-DNA hybrid trigger (ssRNA-cDNA), double-stranded DNA (ssDNA-cDNA), and ps-ssRNA-cDNA hybrid trigger. In comparison with ssRNA trigger, the dsRNA target shows a significant level of Cas13a activation (53%), while the RNA-DNA hybrid target shows limited activation (14.3%), and no activation was observed for double-stranded DNA and ps-ssRNA-cDNA hybrid targets. Therefore, our explorations reveal that while ps-ssRNA and RNA-DNA hybrid target produce limited activation, double-stranded RNA (dsRNA) unexpectedly emerged as a highly effective activator of Cas13a.
A Schematic of a single strand or double strand trigger of Cas13a RNP; (B) Investigation of different types of single strand trigger, including single strand RNA (ssRNA), phosphorothioate-modified single strand RNA (ps-ssRNA) and single strand DNA (ssDNA) (n = 3); (C) Investigation of different types of double strand trigger, including double strand RNA (ssRNA-cRNA), RNA-DNA hybrid trigger (ssRNA-cDNA), double strand DNA (ssDNA-cDNA), and ps-ssRNA-cDNA hybrid trigger (ps-ssRNA-cDNA) (n = 3); (D) Schematic of the FRET approach for the investigation of the dsRNA Cas13a activation mechanism. t-RNA represents target RNA, c-RNA represents complementary RNA; (E) FRET measurements shed light on the dsRNA Cas13a activation mechanism confirming the binding of t-RNA on the gRNA of Cas13a (n = 3). (*P < 0.05, **P < 0.005, ***P < 0.001).
To investigate the Cas13a RNP activation mechanism by dsRNA, a Forster Resonance Energy Transfer (FRET) approach was applied21. To this aim, as shown in Fig. 1D, the two strands in dsRNA were labelled on one end with a fluorophore and a quencher (t-RNA with FAM, c-RNA with BHQ1); in such conditions limited background fluorescence signal was observed. The interaction of this dsRNA with Cas13a RNP is expected to involve binding of the t-RNA to gRNA of Cas13a RNP, and release of the cRNA, which should increase the FRET signal. This was verified in Fig. 1E, where a low fluorescence signal of dsRNA was observed, while the signal for ssRNA-FAM was significantly higher. The fluorescence signal of Cas13a RNP upon interaction with dsRNA was comparable to that of ssRNA-FAM, confirming the binding of t-RNA on the gRNA of Cas13a. This recognition event was further demonstrated using AlphaFold 322 (Fig. S2C). Upon the interaction between dsRNA target with Cas13a RNP, the target RNA was recognized by Cas13a RNP, leading to the release of cRNA.
Activated Cas13a RNP incapable of trans-cleaving dsRNA
Published reports indicate that Cas13a RNP is capable of trans-cleavage of ssRNA (Fig. S3)2. However, there has been relatively little investigation into the trans-cleavage activity of Cas13a on dsRNA substrates explored in this section (Fig. 2A). As shown in Fig. 2B, agarose gel electrophoresis shows that dsRNA is not cleaved by activated Cas13a. To further confirm this finding, a double-stranded fluorescent reporter was applied in a CRISPR/Cas13a biosensing system (Fig. 2C), and no significant fluorescence increase was seen, demonstrating that dsRNA cannot be cleaved by activated Cas13a.
A Schematic of the trans-cleavage activity of Cas13a RNP on dsRNA. B Investigation of the trans-cleavage activity of Cas13a on dsRNA and ssRNA substrates using gel electrophoresis assay. 1) 10 bp ladder; 2) dsRNA; 3) CRISPR/Cas13a treated dsRNA; 4) ssRNA; 5) CRISPR/Cas13a treated ssRNA. C Investigation of the trans-cleavage activity of Cas13a on ssRNA and dsRNA substrates using fluorescent reporters (n = 3). (*P < 0.05, **P < 0.005, ***P < 0.001).
Synthesis and characterization of RNA-Nanocircle
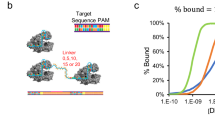
Since dsRNA is an effective activation trigger of Cas13a RNP, but not a trans-cleavage substrate of activated Cas13a RNP, we constructed a RNA-Nanocircle structure comprising a double-stranded RNA section and a short fragment of ssRNA. The RNA-Nanocircle was constructed by a circular ssRNA with a slightly shorter cRNA. The circular ssRNA (Cir-ssRNA) was synthesized by using a click chemistry method based on our previous work20 (Fig. S4), and it was characterised by agarose gel electrophoresis (Fig. 3A). As shown in Fig. 3A, Cir-ssRNA moves more slowly than linear ssRNA, confirming a difference in molecular conformation. In addition, reduced activation of Cas13a by Cir-ssRNA compared with linear ssRNA was observed (Fig. 3B), consistent with the circular conformation of Cir-ssRNA. After mixing Cir-ssRNA with its cRNA, Cir-dsRNA was formed. Lower fluorescence signals for Cir-dsRNA triggered Cas13a system were observed in Fig. 3C compared to the corresponding linearised structures-triggered Cas13a system, indicating that circular targets were locked by the linker with limited ability to trigger activation of Cas13a, and shorter linker length led to lower background signals.
A Evidence of circular single-stranded RNA formation through gel electrophoresis assay. 1). 10 bp ladder; 2). Linear ssRNA; 3) Circular ssRNA. B Exploring the ability of circular ssRNA (Cir-ssRNA) to trigger activation of Cas13a RNP (n = 3), L-x represents the ssRNA linker length. C Exploring the ability of circular dsRNA (Cir-dsRNA) to trigger activation of Cas13a RNP (n = 3), L-x represents the ssRNA linker length. D Schematic of fluorescent RNA-Nanocircle. E Background signal of fluorescent RNA-Nanocircle (n = 3), L-x represents the ssRNA linker length. F Investigation of the linker length of fluorescent RNA-Nanocircle in a CRISPR/Cas13a biosensing system (n = 3). (*P < 0.05, **P < 0.005, ***P < 0.001).
To open the locked circular targets, a fluorescent RNA-Nanocircle was established (Fig. 3D), where a fluorescent reporter (FAM) and a quencher (BHQ1) were placed in close proximity on both sides of the cRNA on the RNA-Nanocircle. Following the cleavage of the ssRNA region in the RNA-Nanocircle, a linearised dsRNA is produced, which unquenches the fluorophore, leading to restoration of fluorescence signals. The background fluorescence signal of different fluorescent RNA-Nanocircles with varying linker length was tested (Fig. 3E). We found that longer ssRNA linker length leads to higher fluorescence background due to increased fluorophore-quencher distance. Subsequently, the fluorescent RNA-Nanocircle was applied as a standard reporter in a CRISPR/Cas13a biosensing system (Fig. 3F). We found that increasing the linker length resulted in stronger signals, with L-5 identified as the optimal length (Fig. 3F). Afterwards, the stability of the fluorescent RNA-Nanocircle was also investigated (Fig. S5), and we found that the RNA-Nanocircle shows very high stability in the rCutSmart buffer and acceptable stability in human plasma within 15 min.
Assessing the fundamental characteristics of Cas13a-based autocatalytic biosensor
The operation of a RNA-Nanocircle-based Cas13a autocatalytic biosensor is illustrated in Fig. 4A. After introducing a target RNA, Cas13a RNP is activated to trans-cleave the ssRNA region of the RNA-Nanocircles. The linearized RNA-Nanocircles become surrogate targets able to activate new Cas13a RNP and sustain an autocatalytic loop20. Subsequently, we evaluated the biosensing performance of this Cas13a-autosensor. In comparison with a basic standard Cas13a biosensor, which shows a linear signal increase with increasing time, the Cas13a-autosensor shows an exponential signal increase (Fig. 4B), and it achieves the limit of detection of 1aM (Fig. 4C). To verify the versatility of Cas13a-autosensor, except LwaCas13a, LbuCas13a was selected, and all the key findings were demonstrated (Fig. S6).
A Mechanism of a Cas13a-autosensor. B Verification of autocatalysis in the Cas13a-autosensor (n = 3). C Investigation of the sensitivity of Cas13a-autosensor using short synthetic RNA targets (n = 3). (*P < 0.05, **P < 0.005, ***P < 0.001).
Application of Cas13a-based autocatalytic biosensor to CRC liquid biospy samples
Although single Cas RNP based Cas13a-autosensor (Cas13a-autosensor-1) realized the limit of detection of 1aM, its versatility was restricted to one specific RNA target matching the RNA-Nanocircle sequence. To improve the versatility, we devised a two Cas RNPs-based autocatalytic system (Cas13a-autosensor-2, Fig. 5A). In this system, first the target RNA activates Cas13a RNP-1 to trans-cleave RNA-Nanocircles matching gRNA in RNP-2 present in its proximity. This then produces linearized RNA-Nanocircles able to trigger activation of Cas13a RNP-2, which continue to trans-cleave the RNA-Nanocircles and drive the autocatalytic reaction system. In contrast to a single Cas13a-autosensor, the two Cas13a RNPs-based autosensor is able to detect different targets by only changing the gRNA of Cas13a RNP-1, but using the same RNA-Nanocircles matching gRNA in Cas13a RNP-2.
A Schematics of RNA-Nanocircle assisted autocatalytic biosensing system using two Cas13a RNPs (Cas13-autosensor-2). B Investigation of the specificity of Cas13a-autosensor-2 for different interference sequences (n = 3), S represents specificity sequence. C The calibration curve of Cas13a-autosensor-2 for the detection of miRNA-21 in human plasma. D Testing the concentration of miRNA-21 in different tumor stages (Neg: 0; T1: 1.2fM; T2: 10.2pM; T3: 13.3pM; T4: 11.4pM). E Testing the concentration of miRNA-21 before and post treatment (T3 post treatment: 10.2 fM; T4 post treatment: 17.1 fM), Neg represents healthy human plasma. (*P < 0.05, **P < 0.005, ***P < 0.001).
To investigate whether any interfering sequences are able to activate the Cas13a-autosensor-2, a range of different sequences were added into Cas13a-autosensor-2 solution, and significant differences were observed between matched and mismatched targets, demonstrating exceptional specificity of Cas13a-autosensor-2 (Fig. 5B). Subsequently, the optimized Cas13a-autosensor-2 was applied to detect miRNA-21 in human colorectal cancer (CRC) plasma samples. Earlier reports indicate that miRNA-21 in plasma represents a potential clinical CRC biomarker, which has 90% specificity and sensitivity23,24, and its physiological values in healthy and CRC patients range from fM to nM25. We established the calibration curve by spiking different concentrations of miRNA-21 in healthy human plasma (Fig. 5C). The Cas13a-autosensor-2 was then applied to test the miRNA-21 concentrations at different tumor stages (Fig. 5D and Table S1), and significantly higher concentration of miRNA-21 was observed in all the CRC plasma samples (T1 to T4) in comparison with healthy human plasma (Neg). In addition, we observed that the concentration of miRNA-21 rises from femtomolar (fM) to picomolar (pM) levels as the tumour stage progresses from T1 to T2. Subsequently, it stabilizes within the picomolar range from T2 to T4. These miRNA-21 concentrations were also tested using RT-PCR, and similar trend was observed (Fig. S7). Post chemotherapy treatment, the concentration of miRNA-21 was found to be reduced from pM range to fM range (Fig. 5E). Taken together, these data confirm that Cas13a-autosensor is an effective approach for the monitoring of CRC progression and treatment. Table 1.
Discussion
In this study, we discovered original fundamental properties of Cas13a that dsRNA effectively triggers activation of Cas13a RNP, but activated Cas13a RNP is unable to trans-cleave dsRNA targets. Circular RNA, which has a dsRNA and ssRNA section (RNA-Nanocircle) with optimised lengths has limited ability to activate Cas13a, but activation is restored when circular RNA is linearised by trans-cleavage. Based on this finding, we established a RNA-Nanocircle-assisted Cas13a autocatalytic biosensing system. The original design changes the original activation pattern from one-to-one correspondence between RNA targets and activated Cas RNPs, to a single target activating an avalanche of activated Cas RNPs. This elegant design of a Cas13a-autosensor has been able to detect 1aM RNA targets within 15 min demonstrating its capability to realize RT-PCR level sensitivity in a POCT setting. It provides a platform for ultrasensitive and rapid detection of RNA targets without preamplification.
To further improve the versatility of the Cas13a-autosensor, a two-Cas13a RNPs-based autocatalytic sensing system was established, in which Cas13a RNP-1 recognizes the target RNA, and Cas13a RNP-2 and matching RNA-Nanocircles are responsible for the autocatalysis reaction. In contrast to a single Cas13a-based autosensor, which is only applicable to one specific target since the dsRNA trigger in the RNA-Nanocircle is identical to the genuine target RNA, the two Cas13a RNP-based autosensor is able to detect diverse targets by a simple replacement of gRNA of the Cas13a RNP-1. This two Cas13a RNP system transforms the standard Cas13a reaction system into an autocatalytic system with the addition of a universal Cas13a autosensor solution (Cas13a RNP2 and matching RNA-Nanocircle). The combination of RNA-Nanocircles and Cas13a RNP-2 mixtures can serve as an autonomous signal amplifier for all existing Cas13-based biosensing systems26,27. The optimized Cas13a-autosensor was successfully applied to monitor the miRNA-21 concentration from clinical CRC plasmas, demonstrating clinical applicability of this autosensor. The implementation of such assays in clinical practice would give the practitioners and patients much increased certainty that clinical decisions are optimally made and based on the state-of-the-art clinical evidence.
While the Cas13a-autosensor demonstrates high sensitivity and clinical applicability, two technical limitations should be noted. First, the synthesis of fluorophore- and quencher-labelled RNA-Nanocircles involves multiple chemical and enzymatic steps, including click chemistry and exonuclease treatment. Although these procedures can be reliably performed in standard molecular biology laboratories, further optimization is required to simplify the workflow and reduce manufacturing costs for large-scale applications. Second, the RNA-Nanocircle exhibits excellent stability in buffer but only moderate stability in unprocessed human plasma, maintaining integrity for approximately 15 min. The incorporation of simple pretreatment steps, such as heat inactivation or Proteinase K digestion, or the introduction of chemical modifications to improve resistance against nucleases, could substantially enhance its stability in biological samples. Addressing these limitations will facilitate the practical translation of this autocatalytic CRISPR platform into deployable diagnostic assays.
Methods
Materials
LwCas13a (Magigen), Cas13a reaction buffer (Magigen), rCutSmart Buffer (NEB), NEBuffer 2.1 (NEB), agarose (ThermoFisher), SYBR Green RNA dye (ThermalFisher), 10 bp DNA ladder (ThermoFisher), 6X RNA loading dye (ThermoFisher), exonuclease T (NEB), copper sulfate (CuSO4) (Sigma, 209198), Tris(2-carboxyethyl) phosphine (TCEP) (Sigma, C4706), tris(benzyltriazolylmethyl) amine (TBTA) (ChemSupply, T2993), streptavidin coated magnetic particles (Spherotech, SVM-08-10), DNase/RNase free water (ThermoFisher), and phosphate buffered saline (PBS) (Sigma, 10 mM, pH = 7.4).
All DNA and RNA oligos are synthesized and modified by Gencefe Biotech Co. Ltd (Table 1).
Investigation of the basic properties of CRISP/Cas13a biosensing system
Standard CRISPR/Cas13a reaction mixture consisted of 40 nM of Cas13a protein, 20 nM of gRNA, 120 nM of 5U-reporters, and 1 mL of rCutSmart buffer.
Optimization of trigger ssRNA concentration: A variety of trigger ssRNA (0, 2.5, 5, 10, and 20 nM) were added into 100 μL of a standard reaction mixture and incubated at room temperature for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Optimization of gRNA to Cas13a ratio: A variety of gRNA-based reaction mixture was prepared (0, 5, 10, 20, 40, and 80 nM of gRNA), 40 nM of Cas13a protein, 120 nM of reporters, and 1X of rCutSmart buffer. Afterwards, 40 nM of trigger ssRNA was added into 100 μL of a standard reaction mixture and incubated at room temperature for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Optimization of reporter concentration: A variety of 5U-reporter solutions was prepared (0, 20, 40, 60, 120, 180 nM of reporter), 40 nM of Cas13a protein, 20 nM of gRNA, and 1X of rCutSmart buffer. Afterwards, 20 nM of trigger ssRNA was added into 100 μL of the standard reaction mixture and incubated at room temperature for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Optimization of buffers: Three different types of buffers were used as the reaction buffer, including Reaction buffer, NEB2.1, and rCutSmart buffer. After preparation of the standard reaction mixture, 20 nM of trigger ssRNA was added into 100 μL of the standard reaction mixture and incubated at room temperature for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Optimization of temperature: After preparation of the standard reaction mixture, 20 nM of trigger ssRNA was added into 100 μL of the standard reaction mixture and incubated at room temperature or 37 °C for two hours. The fluorescence signal was tested using anID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of the limit of detection of standard CRISPR/Cas13a biosensing system: After preparation of the standard reaction mixture, a variety of trigger ssRNA solutions (0, 1 pM, 10 pM, 100 pM, 1 nM, 10 nM, and 100 nM) were added into 100 μL of the standard reaction mixture and incubated at room temperature for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of the basic properties of single strand trigger for Cas13a RNP
Standard CRISPR/Cas13a reaction mixture was prepared by combining 40 nM of Cas13a protein, 20 nM of gRNA, 120 nM of 5U-reporter, and 1 mL of rCutSmart buffer.
Investigation of different single-strand trigger types was carried out as follows. 2 μL of 1 μM of trigger (ssRNA, ssDNA, and ps-ssRNA) was added into 100 μL standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of the basic properties of double strand trigger for Cas13a RNP
Standard CRISPR/Cas13a reaction mixture was prepared by combining 40 nM of Cas13a protein, 20 nM of gRNA, 120 nM of 5U-reporter, and 1 mL of rCutSmart buffer.
Investigation of different types of double strand trigger was carried out as follows. 2 μL of 1 μM of trigger (ssRNA, cRNA, dsRNA, cDNA, ssRNA/cDNA, dsDNA, ps-ssRNA/cDNA) was added into 100 μL of standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of trigger mechanism of dsRNA trigger for Cas13a RNP
To investigate the trigger activation mechanism of Cas13a, the FRET method was utilized. In brief, a reaction mixture was prepared by combining 40 nM of Cas13a protein, and 20 nM of gRNA in 1 mL of rCutSmart buffer. Afterwards, 2 μL of 1 μM of fluorescent dsRNA was added into 100 μL of the prepared reaction mixture. Subsequently, the fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM). For comparison, 2 μL of 1 μM of dsRNA or 2 μL of 1 μM of ssRNA-FAM was added into 100 μL of 1X rCutSmart buffer, and the fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of the basic trans-cleavage properties of Cas13a RNP
To investigate the trans-cleavage properties of Cas13a RNP on a single strand nucleic acid, a CRISPR/Cas13a reaction mixture was first prepared comprising 40 nM of Cas13a protein, 20 nM of gRNA, and 120 nM of reporter (RNA, DNA, RNA-DNA, 5 A, 5U, 5 C, and 5 G) in 1 mL of rCutSmart buffer. Subsequently, 2 μL of 1 μM of ssRNA trigger was added into 100 μL standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
To investigate the trans-cleavage properties of Cas13a RNP on dsRNA and ssRNA, a CRISPR/Cas13a reaction mixture was first prepared comprising 320 nM of Cas13a protein, 160 nM of gRNA, and 1 μM of dsRNA or ssRNA substrates in 1 mL rCutSmart buffer. Subsequently, 160 nM of ssRNA trigger was added into the standard reaction mixture and incubated at 37 °C for two hours. Afterwards, agarose gel electrophoresis was applied to evaluate the conformation of dsRNA or ssRNA. In brief, 5% agarose gel in 1×TAE buffer was prepared with SYBR Green DNA dye. 5 μL of dsRNA or ssRNA was premixed with 5 μL of 2X RNA gel loading dye and then loaded into gel for electrophoresis, which was carried out for 40 min at a constant voltage of 100 V. 4 μL of 10 bp DNA ladder was used for molecular weight reference. Gel images were visualized by using a Gel Doc + XR image system (Bio-Rad Laboratories Inc., USA) (Fig. S8).
To further investigate the trans-cleavage properties of Cas13a RNP on a fluorescent double strand nucleic acid, a CRISPR/Cas13a reaction mixture was first prepared comprising 40 nM of Cas13a protein, 20 nM of gRNA, and 120 nM of fluorescent dsRNA reporters in 1 mL of rCutSmart buffer. Subsequently, 2 μL of 1 μM of ssRNA trigger was added into 100 μL standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Synthesis and characterization of circular RNA
The synthesis approach of circular RNA was based on our previous approach (click chemistry)20. In brief, 0.4 mL of 0.5% w/v of streptavidin modified magnetic beads (0.74 μm) were first blocked with 1% BSA solution for 1 h to eliminate non-specific binding. Afterwards, 1 mL of 0.5 μM of biotinylated linear-ssRNA was incubated with the beads for 1 h following a PBS wash to remove the residual free linear-ssRNA. Subsequently, 1 mL of the click chemistry reaction solution (1.0 mM of CuSO4, 2.0 mM of TCEP, and 100 μM of TBTA) was added and incubated with the beads for 12 h at room temperature. After synthesis, the magnetic beads were collected and washed with PBS buffer to remove excess chemicals. Subsequently, 100 μL of 100 units/mL of Exonuclease T solution was added and incubated at 37 °C for 30 min to remove linear ssDNA. After washing with PBS buffer, the synthesized Cir-ssRNA was released from the streptavidin-modified magnetic beads by heat treatment at 95 °C for 30 min, and the supernatant was collected for further use. All the Cir-ssRNA used in this research are synthesized based on this approach. A Nanodrop system was utilized to test the concentration of synthesized Cir-ssDNA.
The formation of Cir-ssRNA was verified by using agarose gel electrophoresis. In brief, 5% agarose gel in 1×TBE buffer was prepared with SYBR Green DNA dye. 10 μL of Cir-ssRNA was premixed with 2 μL of 6X DNA gel loading dye and then loaded into gel for electrophoresis, which was carried out for 40 min at a constant voltage of 100 V. 5 μL of 10 bp DNA ladder was used for molecular weight reference. Gel images were visualized by using Gel Doc + XR image system (Bio-Rad Laboratories Inc., USA).
Investigation of the basic trigger ability of circular RNA to Cas13a RNP
To investigate the trigger ability of Cir-ssRNA and Cir-dsRNA, standard CRISPR/Cas13a reaction solution was first prepared (40 nM of Cas13a protein, 20 nM of gRNA, 120 nM of reporter, and 1 mL of rCutSmart buffer). Subsequently, 2 μL of 1 μM of Cir-ssRNA or Cir-dsRNA trigger was added into 100 μL of the standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Investigation of the trans-cleavage properties of Cas13a RNP on RNA-Nanocircle
The formation of fluorescent RNA-Nanocircle was conducted by mixing synthesized Cir-ssRNA with its corresponding fluorescent cRNA at the molar ratio of 1:1. Afterwards, the background of fluorescent RNA-Nanocircle (120 nM) in rCurSmart buffer was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
To investigate the trans-cleavage properties of Cas13a RNP on RNA-Nanocircle, a CRISPR/Cas13a reaction mixture was first prepared comprising 40 nM of Cas13a protein, 20 nM of gRNA, and 120 nM of fluorescent RNA-Nanocircle in 1 mL of rCutSmart buffer. Subsequently, 2 μL of 1 μM of ssRNA trigger was added into 100 μL standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
To investigate the stability of RNA-Nanocircle, 200 nM of fluorescent RNA-Nanocircle (L-5) was incubated in rCurSmart buffer, 10% human serum solution, or 10% saliva solution for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Evaluating the basic properties of Cas13a based autocatalytic biosensor
Cas13a-autosensor-1 standard reaction mixture was first prepared: 40 nM of Cas13a protein, 20 nM of gRNA, and 120 nM of fluorescent RNA-Nanocircle in 1 mL rCutSmart buffer. Subsequently, 2 μL of 1 μM of ssRNA trigger was added into 100 μL standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using ID5 plate reader (Ex 480 nM, and Em 520 nM).
To investigate the autocatalysis activity, 1 pM ssRNA trigger was added into 100 μL standard reaction mixture and incubated at 37 °C for one hour. The fluorescence signal was tested using ID5 plate reader (Ex 480 nM, and Em 520 nM).
To investigate the sensitivity, different concentrations of ssRNA (0–10 nM) was added into 100 μL of the standard reaction mixture and incubated at 37 °C for two hours. The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
Application of Cas13a-based autocatalytic biosensor for the detection of miRNA-21 in healthy human plasma and clinical human plasma samples
The Cas13a-autosensor-2 reaction mixture for miRNA-21 detection was prepared as follows: 40 nM of Cas13a protein was mixed with 20 nM of gRNA (miRNA-21) to form the Cas13a RNP1. In the meantime, 40 nM of Cas13a protein was mixed with 20 nM of gRNA (RNA-Nanocircle) to form the Cas13a RNP2. Afterwards, the prepared Cas13a RNP1 and Cas13a RNP2 were mixed with 120 nM of fluorescent RNA-Nanocircle in 1 mL of rCutSmart buffer to form the reaction mixture. The reaction mixture was stored at 4 °C before use.
To investigate the specificity, 2 μL of 1 μM of interference ssRNA was added into 100 μL reaction mixture for activating trans-cleavage of Cas13a and enabling the CRISPR/Cas biosensing reaction (37 °C). The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
To establish the calibration curve, 10 μL of spiked in sample was added into 90 μL standard reaction mixture for activating trans-cleavage of Cas13a and enabling the CRISPR/Cas biosensing reaction (37° C). The fluorescence signal was tested using an ID5 plate reader (Ex 480 nM, and Em 520 nM).
To evaluate the clinical application performance of Cas13a-autosensor, colorectal cancer (CRC) clinical samples were collected and provided by Health Precincts Biobank of UNSW. All patients provided written informed consent for participation in this study. These plasma samples were collected from patients with different tumor stages from T1 to T4. In addition, post chemotherapy (post sx) samples from T2 to T4 were also collected to evaluate the effectiveness of treatment. For the testing, 10 μL of CRC plasma samples was added into 90 μL standard reaction mixture for activating trans-cleavage of Cas13a and enabling the CRISPR/Cas biosensing reaction (37 °C). The fluorescence signal was tested using ID5 plate reader (Ex 480 nM, and Em 520 nM).
All human plasma experiments were approved by the UNSW Ethics Committee (UNSW HC210160). The Health Precincts Biobank samples were received from operates under ethics approval from the South Eastern Sydney Local Health District – Northern Sector HREC, reference number 2019/ETH12304.
Validation of miRNA-21 expression in human plasma using RT-qPCR
RNA was extracted from 200 µL of human plasma using the Invitrogen PureLink RNA Mini Kit (Cat. No. 12183018 A) following the manufacturer’s protocol. Briefly, plasma samples were lysed with Lysis Buffer treated with Proteinase K. The lysate was then passed through a Spin column, washed with Wash Buffer I and Wash Buffer II, and RNA was eluted with RNase-free water. RNA yield and quality was determined using Nanodrop.
Reverse transcription was performed using the miRCURY LNA miRNA PCR Starter Kit (Qiagen, Cat. No. 339320) according to the manufacturer’s instructions. The extracted RNA was normalized across various samples, and 2 ng/µL per sample was reverse transcribed to cDNA using the miRCURY LNA RT Kit.
Sybr based real-time PCR was conducted with the miRCURY LNA miRNA PCR Assay for miRNA-21-5p (YP00204230) as the target gene and miRNA-103a (YP00205727) as the reference gene.
qPCR reactions were set up in a total volume of 10 µL in triplicates, and amplification was performed using the following cycling conditions: initial denaturation at 95 °C for 2 min, followed by 40 cycles of 95 °C for 10 s and 60°C for 1 min, followed by a melt curve from 60 °C to 95 °C.
The relative expression levels of miRNA-21-5p were normalized to miRNA-103a using the ΔCt method.
Data availability
All data are available in the main text or the supplementary materials.
References
Liu, L. et al. The molecular architecture for RNA-guided RNA cleavage by Cas13a. Cell 170, 714–726. e10 (2017).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Gootenberg, J. S. et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360, 439–444 (2018).
Li, Y. et al. CRISPR/Cas systems towards next-generation biosensing. Trends Biotechnol. 37, 730–743 (2019).
Kellner, M. J. et al. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat. Protoc. 14, 2986–3012 (2019).
Chandrasekaran, S. S. et al. Rapid detection of SARS-CoV-2 RNA in saliva via Cas13. Nat. Biomed. Eng. 6, 944–956 (2022).
Healy, B. et al. The impact of false positive COVID-19 results in an area of low prevalence. Clin. Med. 21, e54 (2021).
Huang, X. et al. Developing RT-LAMP assays for rapid diagnosis of SARS-CoV-2 in saliva. EBioMedicine 75, 103736 (2022).
Stinson, L. F., Keelan, J. A. & Payne, M. S. Identification and removal of contaminating microbial DNA from PCR reagents: impact on low-biomass microbiome analyses. Lett. Appl. Microbiol. 68, 2–8 (2019).
Xue, Y. et al. PddCas: A polydisperse droplet digital CRISPR/Cas-based assay for the rapid and ultrasensitive amplification-free detection of viral DNA/RNA. Anal. Chem. 95, 966–975 (2022).
Fozouni, P. et al. Amplification-free detection of SARS-CoV-2 with CRISPR-Cas13a and mobile phone microscopy. Cell 184, 323–333. e9 (2021).
Broto, M. et al. Nanozyme-catalysed CRISPR assay for preamplification-free detection of non-coding RNAs. Nat. Nanotechnol. 17, 1120–1126 (2022).
Li, H. et al. Amplification-Free Detection of SARS-CoV-2 and Respiratory Syncytial Virus Using CRISPR Cas13a and Graphene Field-Effect Transistors. Angew. Chem. 134, e202203826 (2022).
Liu, T. Y. et al. Accelerated RNA detection using tandem CRISPR nucleases. Nat. Chem. Biol. 17, 982–988 (2021).
Sha, Y. et al. Cascade CRISPR/cas enables amplification-free microRNA sensing with fM-sensitivity and single-base-specificity. Chem. Commun. 57, 247–250 (2021).
Zhao, D. et al. CRISPR/Cas13a-triggered Cas12a biosensing method for ultrasensitive and specific miRNA detection. Talanta 260, 124582 (2023).
Zhang, Y. et al. An aM-level sensitive cascade CRISPR-Dx system (ASCas) for rapid detection of RNA without pre-amplification. Biosens. Bioelectron. 230, 115248 (2023).
Deng, F. et al. Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing. Nat. Commun. 15, 1818 (2024).
Shi, K. et al. A CRISPR-Cas autocatalysis-driven feedback amplification network for supersensitive DNA diagnostics. Sci. Adv. 7, eabc7802 (2021).
Deng, F. et al. Bifunctional circular DNA amplifier transforms a classic CRISPR/Cas sensor into an ultrasensitive autocatalytic sensor. (2023).
Deng, F. et al. Increasing trans-cleavage catalytic efficiency of Cas12a and Cas13a with chemical enhancers: Application to amplified nucleic acid detection. Sens. Actuators B: Chem. 373, 132767 (2022).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 1–3 (2024).
Kanaan, Z. et al. Plasma miR-21: a potential diagnostic marker of colorectal cancer. Ann. Surg. 256, 544–551 (2012).
Toiyama, Y. et al. Serum miR-21 as a diagnostic and prognostic biomarker in colorectal cancer. J. Natl. Cancer Inst. 105, 849–859 (2013).
Calvo-Lozano, O. et al. One-step and real-time detection of microRNA-21 in human samples for lung cancer biosensing diagnosis. Anal. Chem. 94, 14659–14665 (2022).
Shan, Y. et al. High-fidelity and rapid quantification of miRNA combining crRNA programmability and CRISPR/Cas13a trans-cleavage activity. Anal. Chem. 91, 5278–5285 (2019).
Zhou, T. et al. CRISPR/Cas13a powered portable electrochemiluminescence chip for ultrasensitive and specific MiRNA detection. Adv. Sci. 7, 1903661 (2020).
Acknowledgements
The authors acknowledge the support of the UNSW SHARP program of E.M. G and Early Career Fellowship of Cancer Institute NSW (2024/ECF1573) of F.D. The authors acknowledge the support of Prof Diangeng Li from Beijing Ditan Hospital, Capital Medical University.
Author information
Authors and Affiliations
Contributions
Conceptualization: F.D & E.G. Methodology: F.D., S.G., & R.S. Investigation: F.D., S.G., & R.S. Supervision: E.G. Writing-original draft: F.D., Writing-review & editing: F.D, S.G., R.S., Y.L. & E.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, F., Gulati, S., Sang, R. et al. Autocatalytic Cas13a biosensor enabled by RNA-nanocircles for ultrasensitive RNA detection. npj Biosensing 3, 5 (2026). https://doi.org/10.1038/s44328-025-00067-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44328-025-00067-6