Abstract
This cross-sectional study explores the impact of FTO gene single nucleotide polymorphisms (SNPs) rs9939609 and rs1421085 on dietary habits contributing to obesity risk in Thai adults. The study enrolled 384 participants from Bangkok, categorized as non-obese (BMI < 25 kg/m2) or obese (BMI ≥ 25 kg/m2) based on WHO Asia Pacific Guidelines. Genotyping for FTO variants was performed using DNA from blood samples. While both SNPs adhered to Hardy–Weinberg equilibrium, the association between risk alleles and anthropometric measurements was not statistically significant. However, risk allele carriers showed significantly higher intakes of sugar and saturated fat compared to homozygous dominant individuals. In the obese group, the odds ratio for high-sugar intake was 2.22 (95% CI 1.13–4.37, p = 0.021) for rs9939609 risk allele carriers. For high-saturated fat intake, the odds ratio was 1.86 (95% CI 1.02–3.40, p = 0.041). Similar associations were observed for rs1421085. Risk allele carriers also exhibited significantly higher leptin levels (p < 0.043) and a positive correlation with myeloperoxidase levels (p < 0.038). These findings highlight the complex relationship between FTO risk alleles, increased consumption of sugar and saturated fat, and obesity-related parameters. The insights emphasize the importance of considering both genetic and dietary factors in obesity prevention strategies.
Similar content being viewed by others
Introduction
Obesity is a prominent global health issue characterized by the accumulation of excess body fat. This condition significantly increases the likelihood of developing metabolic disorders such as type 2 diabetes mellitus, hypertension, and cardiovascular diseases1. By 2030, over 1 billion individuals worldwide will be affected by obesity and its associated health complications. The prevalence of obesity in Thailand has been increasing over the past few decades. In the 2021 survey, approximately 42% of the adult population was classified as overweight or obese (BMI ≥ 25 kg/m2), with around 9% being obese (BMI ≥ 30 kg/m2)2. The increase in obesity prevalence over the past four decades has been attributed to changes in food habits and a more obesogenic environment resulting from industrialization and globalization3. Important causes of obesity include lifestyle, physical activity, and dietary consumption. Overconsumption of high-energy foods such as those containing high amounts of fat and sugar, combined with low physical activity, leads to an energy imbalance that causes obesity4 Although poor lifestyle has been identified as the primary cause of obesity, genetic factors also play a crucial role in its development5 Among the genes implicated in obesity, the fat mass and obesity-associated (FTO) gene stands out as one of the most important. FTO, located on chromosome 16, encodes an enzyme involved in adipocyte thermogenesis, energy homeostasis, and metabolic rate regulation, contributing significantly to body fat accumulation6. FTO genetic variants, including rs9939609 and rs1421085, have been correlated with high body mass index (BMI), increased adiposity, and a predisposition to obesity across diverse populations7,8 Moreover, FTO polymorphisms have been implicated in the onset of metabolic syndrome and type 2 diabetes mellitus, underscoring their involvement in obesity-related metabolic disorders9 In addition, a study examining the FTO gene in 12 families (comprising 83 individuals of Thai descent) revealed that associations with three single nucleotide polymorphisms (SNPs) situated in the first intron of FTO (rs1421085, rs17817449, and rs8043757) escalated the obesity risk by 2.82 times for rs142108510.
The effect of FTO variants on obesity risk extends beyond genetic predisposition, encompassing dietary behavior and nutrient intake. Carriers of the A allele of the rs9939609 polymorphism tend to prefer energy-dense foods, particularly those high in fat, resulting in increased body weight caused by altered FTO expression in the hypothalamus11.
In addition, associations among FTO variants, obesity, and dietary intake have been observed across diverse ethnic groups, suggesting an interplay between genetic predisposition and dietary factors in obesity development. Moreover, a study described the longitudinal effects of FTO on personality, brain function, and dietary habits among older individuals12 The FTO rs1421085 variant was associated with decreased brain function in the medial prefrontal cortex, potentially influencing a preference for dietary fat over time. Furthermore, studies in various populations, including Iranian adults who were overweight, revealed that rs9939609 AA carriers exhibited higher carbohydrate, calorie, and fat intakes than TT carriers, whereas Emirati participants with the same genotype showed higher carbohydrate but lower fat intake13 A recent study of older individuals also indicated an inverse relationship between serum leptin levels and the number of FTO C risk alleles, depending on distinct rs17817449 genotypes14 These findings suggest that FTO SNPs may influence weight gain by altering endocrine balance and are closely associated with food intake patterns.
The expression of FTO influences multiple brain regions involved in regulating energy balance and appetite. The FTO mutation rs9939609 is associated with not only obesity but also macronutrient intake, including carbohydrates, proteins, polyunsaturated fatty acids (PUFA), and saturated fatty acids (SAT fat)15,16 (A similar influence was expected to exist for FTO rs1421085, which exhibits high linkage disequilibrium with FTO rs9939609. Several studies have shown associations between FTO rs1421085 and both obesity-related traits and dietary macronutrient intake. In vivo and in vitro model studies have indicated that FTO rs1421085 can modify the binding of transcriptional repressors in nearby regions, thereby affecting the expression of genes related to adipocyte thermogenesis and food intake17,18.
Studies conducted in Western countries, such as the European nations, have consistently shown strong associations between FTO gene polymorphisms and obesity risk, particularly when coupled with high-calorie diets rich in sugar and fat 19,20. These studies provide a foundational understanding of the genetic and environmental contributors to obesity but may not fully reflect the dietary patterns and genetic diversity of other regions. In regions such as Asia, studies have demonstrated that while the prevalence of the FTO risk allele is lower compared to Western populations, the association between FTO polymorphisms and obesity remains significant21. However, the interaction between FTO variants and dietary factors, such as sugar and fat intake, has been less extensively explored in these populations. Moreover, exploring the association between diet and FTO variants in Indonesia could provide valuable insights into the management of obesity, given the prevalence of fatty Indonesian cuisine caused by the abundant use of coconut milk and palm oil. In Indonesia, dietary fat and SAT fat intakes rank among the highest globally, emphasizing the importance of understanding how FTO variants interact with dietary factors16 FTO rs9939609 has been linked to increased energy intake, which is likely due to its significant expression in the hypothalamus, where it is believed to influence appetite regulation mechanisms and potentially contribute to obesity development. Although many studies have examined the effect of genetic variants and hormonal dysregulation on appetite in individuals with obesity, fewer studies have examined their associations among those with normal weight. Within our country, limited research has focused specifically on the interaction between FTO polymorphisms and diets. In addition, a study examining the FTO gene in 12 families revealed that associations with three single nucleotide polymorphisms (SNPs) situated in the first intron of FTO (rs1421085, rs17817449, and rs8043757) escalated the obesity risk by 2.82 times for rs142108510.
To the best of our knowledge, this is the first prospective study in Thais people that examines the interaction between FTO gene polymorphisms and elevated sugar and fat consumption in relation to obesity risk. Although similar studies have been conducted at a national level or in other regions, no previous research has focused on the unique dietary and genetic characteristics of the population in this particular region. Unlike earlier studies, which primarily relied on cross-sectional designs. For the first time, this study investigated the effect of FTO SNPs on food preferences in adult Thai people. It revealed intriguing interactions between the two SNPs that are well-known for being connected to obesity. This gap is noteworthy because genetic variants and hormonal imbalances related to appetite control may enhance the reward response to foods, particularly those rich in simple sugars and SAT fats22 (Our study explores FTO the role of polymorphisms in obesity, focusing on their impact on energy balance, dietary habits, and related metabolic disorders. We hypothesize that FTO risk allele carriers exhibit significant responses to dietary intake, motivating our investigation into their correlation with dietary consumption, anthropometry, and other obesity risk factors in both non-obese and obese groups.
Materials and methods
Study design and populations
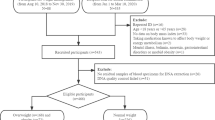
In the cross-sectional study design employed convenience sampling and was conducted between July 2023 and October 2023. Participants were recruited from the Laksi District, Bangkok, Thailand. Individuals were invited to participate voluntarily after receiving a thorough description of the study objectives. Participants were then categorized into non-obese and obese groups based on their BMI, following the World Health Organization Asia Pacific Guidelines for Asians.
Study procedure
The study recruited 384 adults aged 18–59 years following a set of inclusion and exclusion criteria. The inclusion criteria were as follows: individuals with a BMI ≥ 18.5 kg/m2 who were able to read and write independently, willing to participate, and signed a consent form. The exclusion criteria were as follows: pregnant or lactating individuals; those with severe chronic diseases or complications, metabolic conditions altering nutrition requirements, eating behavioral disorders, significant recent weight loss or participation in weight loss programs, and incomplete data, or those in whom blood samples could not be drawn.
Data collection spanned 14 weeks, including 1–2 weeks for public relations, 3–6 weeks for participant screening and collection, and 7–14 weeks for the experiment and data analysis. To clarify project details and be informed of the fasting period, participants were contacted before the study. Communication with participants included informing them of the study details and potential side effects, and interested volunteers registered via Google Forms.
Ethical considerations
The project was approved on June 23, 2023, by the Ethics Committee of the Faculty of Tropical Medicine, Mahidol University of Thailand with the ethical approval no. MUTM 2023-042-01. The study was conducted following the Declaration of Helsinki. Every participant provided informed consent.
Genetic analysis
The SNPs selection criteria included: (a) Minor Allele Frequency (MAF) greater than 5% in the reference population (e.g., 1000 Genomes Project, relevant Asian or Thai-specific databases) to ensure sufficient genetic variability. (b) Hardy–Weinberg Equilibrium (HWE): SNPs that did not deviate significantly from HWE (p > 0.05) in the control population were included to ensure data quality and consistency. (c) Functional relevance: SNPs were chosen based on prior evidence linking them to obesity and metabolic disorders, particularly in relation to the FTO gene. We prioritized SNPs previously reported in genome-wide association studies (GWAS) or literature focused on obesity risk. We used sequencing method to validate the HRM method. The results of showed sensitivity 100%, specificity 100% to Genotyping and Quality Control. Moreover, duplicate and blank samples were used as internal controls to confirm the consistency and reliability of the genotyping process.
DNA extraction
DNA was extracted from blood samples using the QIAamp DNA Blood Mini Kit (250) (QIAGEN, Hilden, Germany) following the manufacturer’s instructions. DNA concentration was measured using a Nano-Drop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
Primer design for high-resolution melting (HRM) assay
Primers were designed to detect FTO rs9939609 (T > A) and rs1421085 (T > C) variants using primer software NCBI Primer-BLAST. Each primer pair was designed to generate polymerase chain reaction (PCR) products with distinctive melting temperatures (Tm). Two primers sets were used: one allele-specific primers for the mutant and one for the wild-type template.
The primers of each FTO rs9939609 were as follows:
-
T F: 5′ GCGACTGCTGTGAATTTT 3′,
-
A F: 5′ GCGACTGCTGTGAATTTA 3′
-
R: 5′ TTTGCTTTTATGCTCTCCCA 3′
The primers of each FTO rs1421085 were as follows:
-
T F: 5′ CAGGTCCTAAGGCATGAT 3′
-
C F: 5′ CAGGTCCTAAGGCATGAC 3′
-
R: 5′ TGGCCCAGTGGGGAGAT 3′
PCR amplification and melting curve analysis
The assay conditions, primer concentrations, protocol, and detection parameters were optimized to enhance the sensitivity and specificity of the assay23 The multiplexed HRM assay was conducted in a total volume of 12.5 µL, comprising 6.25 µL of 2 × HRM Type-It mix (QIAGEN), varying concentrations of each primer, molecular-grade water, and 2.5 µL of the gDNA template (3–10 ng/µL). PCR amplification and melting curve analysis were performed using the Rotor-Gene Q (QIAGEN), with specific conditions tailored to the primer design. HRM analysis involved melting from 70 to 90 ℃, with readings taken at every 0.1 ℃ interval and a 2-s stabilization period. Each run included positive controls (gDNA with known mutations, confirmed by DNA sequencing) and negative controls (wild-type gDNA for each gene, confirmed by DNA sequencing). Data analysis was performed using the Rotor-Gene Q software. Experiments were conducted in duplicate.
Anthropometric and body composition measurements
Anthropometric factors such as height, weight, hips, and waist were measured. BMI was calculated using weight in kilograms divided by height in meters squared (kg/m2). Using a plastic, non-stretchable tailor’s tape, the hip circumference (HC) and waist circumference (WC) were measured in cm. The HC was measured at the broadest circumference across the buttocks, whereas the WC was measured while the patient was standing, breathing properly, and wearing light clothing at a level halfway between the lower rib border and the iliac crest. Then, the waist–hip ratio (W/H) was determined by dividing the WC by the HC. Anthropometric measurements were made in accordance with the International Biological Program’s guidelines using standardized equipment. The percentage of muscle mass, fat mass, muscle mass, and metabolic components of body fat body, composition parameters including muscle mass percentage, fat mass, muscle mass, metabolic age, and basal metabolic age were measured using a body impedance analyzer model TANITA-SC330.
Physical activity assessment
The metabolic equivalent of the task (MET) value was used to calculate the amount of physical activity. Additionally, the activity status was assessed using the Global Physical Activity Questionnaire (GPAQ)24. The level of physical inactivity (PA) was classified into three groups: (a) Low PA: Less than 300 MET-minutes per week or inactive. (b) Moderate PA: Between 300 and 600 MET-minutes per week. (c) High PA: At least 600 MET-minutes per week of moderate-intensity activities (for at least 5 days a week), or 20 min of vigorous-intensity exercise per day (for at least 3 days in a typical week), or activities totaling 1500 MET-minutes per week25.
Laboratory analysis
To assess lipid profiles, fasting plasma glucose levels, and inflammation markers, blood samples were collected after a 12-h fasting period. Serum separation was performed after refrigerating blood samples at approximately 5 ℃, followed by storing the serum at − 80 ℃ for further analysis.
Lipid profile assessment
Blood samples were taken for serum biochemical tests, including measurements of low-density lipoprotein cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C), triglycerides (TGs), and total cholesterol (TC). Using the Stanbio Cholesterol, TG LiquiColor, USA, standard enzymatic methods were used to measure the plasma concentrations of TC and TG. After mixing 1000 µL of the reagent with 10 µL of the sample or standard, the mixture was incubated for 5 min at 37 ℃. The spectrophotometer was then used to measure the absorbance of the samples and the standard at 500 nm. For both TC and TG, the findings were computed by comparing them to the standard and applying the formula: (Absorbance sample/Absorbance standard) * 200. For HDL-C measurement, the CS-400 Auto-Chemistry Analyzer (Istanbul, Turkey) was used. The Friedewald formula was utilized to compute the LDL-C: LDL-C (mg/dL) = TC (mg/dL) − HDL-C (mg/dL) − TG (mg/dL)/5 (Chen Yet al. 2010).
Blood glucose assessment
Fasting blood glucose was measured using the standard glucose oxidase assay method of the Glucose LiquiColor, USA protocol. Participants fasted overnight (8–12 h), and while sitting, their blood was drawn the early morning of the following day (7–8 a.m.). The 10 µL of the blood samples was mixed with 1000 µL of the reagent and incubated for 5 min at 37 ℃. Spectrophotometry was performed at an absorbance of 500 nm. Moreover, the results were calculated by comparing them with the standard and using the following formula: (Absorbance sample/Abs standard) *100. HOMA-IR was calculated from fasting glucose (mg/dL) and insulin (mUI/L) as [fasting glucose (mg/dL) × fasting serum insulin (mU/L)/405)26,27.
Blood inflammation markers
Tumor necrosis factor-α, interleukin-6, and myeloperoxidase (MPO) were evaluated using the enzyme-linked immunosorbent assay (ELISA) method. The ELISA kit (MyBioSource, USA) was used to detect inflammation by this protocol. To summarize the protocol, 100 µL of the standard or samples were added to each well and then incubated for 90 min at 37 ℃. Subsequently, the liquid was removed, and 100 µL of biotinylated detection was added, and incubated for another hour at 37 ℃, followed by aspiration and washing three times. Then, 100 µL of horseradish peroxidase (HRP) was added and incubated for 30 min at 37 ℃, followed by another round of aspiration and washing, performed five times. Then, 90 µL of the substrate reagent was added and incubated for 15 min at 37 ℃. Finally, 50 µL of the stop solution was added, and the absorbance was immediately read at 450 nm. The results were calculated from the standard curve and expressed as pg/mL.
Blood analysis for hormone-related obesity
Leptin and insulin were detected using the ELISA test kit (Sigma Aldrich, USA). Briefly, 100 µL of the samples or standard was added into the well, and the mixture was incubated overnight (4 ℃) or 2.5 h at room temperature. Then, discarding and washing were done four times with buffer. Moreover, 100 µL of HRP was added, and the mixture was incubated for 45 min, followed by washing four times. In addition, 100 µL of the 3,3′,5,5′-Tetramethylbenzidine substrate was added to each well, which was then incubated for 30 min in the dark. Thereafter, 50 µL of the stop solution was added, and absorbance was read immediately at 450 nm.
Dietary intake analyses
Food consumption was estimated using the semiquantitative food frequency questionnaire (FFQ). Seventy-five meals and drinks that are often consumed in each of the participating districts in Bangkok, Thailand, were included in the semiquantitative FFQ. A previous study28 developed the semiquantitative FFQ. Based on nutrient component, main macronutrient, main food group, culinary food group, use, and obesity risk characteristics such as high fat, high carbohydrate, high-sugar, etc., the food group items in the semiquantitative FFQ were grouped into 12 important food groups. A checklist of food and beverage items, including portion sizes and frequency of consumption, was used in the semiquantitative FFQ survey. The checklist was divided into seven categories: never, seldom, 1–2 times per week, 3–4 times per week, 5–6 times per week, 1 time per day, and 2–3 times per day. INMUCAL-Nutrients V.4.0 was used to assess the amount of energy and nutrients in food. To help with the response, an in-person survey was performed using frequency cards and food model items. A high-sugar content was defined as > 77 g from the average sugar consumption in a previous study29. Moreover, high-sat fat content was defined as > 20 g (> 10% of 2000 kcal/day) from WHO recommendation30.
Statistical analysis
The power of the study was calculated to ensure that the sample size was sufficient to detect meaningful associations between genetic variance and obesity. Assuming a significance level (α) of 0.05 and a moderate effect size, with 412 participants, the study has a power of 80% (1 − β = 0.80). This means the study has an 80% probability of detecting an effect if one exists. The sample size was determined using the formula for estimating population proportion in cross-sectional studies. We used a 95% confidence interval (Z = 1.96), a 42.4% prevalence of obesity in the Thai population (2021 data, p = 0.424), and a 5% margin of error (d). These parameters yielded a required sample size of 374 participants. To account for potential data loss, we added 10%, resulting in a final sample size of 412 participants, ensuring adequate power and precision.
The results were reported as means ± SDs for continuous variables and as percentages for categorical variables. Obesity was defined according to the World Health Organization Asia Pacific Guideline for Asians, categorizing individuals as non-obese (BMI < 25 kg/m2) or obese (BMI ≥ 25 kg/m2)31. The normality of data distribution was assessed using the Kolmogorov–Smirnov test. The distribution of genotypes was evaluated for Hardy–Weinberg equilibrium using the chi-square test. General data were compared using the chi-square test for categorical variables. Differences in body composition, blood biochemistry, and nutritional composition data between groups were analyzed using Student’s t-test. Pearson correlation coefficients were used to assess the correlation between blood biochemistry or nutritional composition and FTO variants. The association between sugar content and SAT fat content and FTO variant was examined by calculating the odds ratio with a 95% confidence interval, and analyses of association were conducted using the chi-square test and logistic regressions. All statistical analyses were performed using IBM SPSS Statistics version 23 (IBM Corp., Armonk, NY, USA), with a p < 0.05 considered statistically significant. STROBE cross sectional reporting guidelines was used.
Results
General characteristics according to the FTO genotype
The distribution of FTO rs9939609 and rs1421085 and the general characteristics of the study population are presented in Table 1. The minor allele frequencies (MAFs) were approximately 0.245 (rs9939609) and 0.248 (rs1421085) in the Thai population (data not shown). Because of the low MAF, all analyses were performed by applying the dominant model (risk allele carriers vs. homozygous dominant). The study group consisted of 384 adults, including 185 people who were non-obese and 199 who were obese. Among all participants, the number of female participants was higher than that of their male counterparts in both FTO SNPs. Moreover, the results showed that the homozygous dominant (TT) participants in both groups had a higher prevalence than risk allele carriers (TA + AA or TC + CC), but not significantly. Regarding age, the proportion of those aged 46–59 years among all homozygous dominant (TT) participants in the non-obese group was 26.49%, followed by those aged 18–35 years with 22.16%. Moreover, in the non-obese and obese groups, the number of homozygous dominant (TT) participants who finished a bachelor’s degree (22.16%) was higher than those who completed high school (13.51%). In addition, both FTO SNPs in the non-obese and obese groups showed higher non-smoking rates in the homozygous dominant (TT) group than in the risk allele group (TA + AA or TC + CC) and the same trend with exercise; the rate of non-exercising was higher in the homozygous dominant (TT) group (36.22%, 34.17%, 36.22%, and 31.16%). However, the homozygous dominant (TT) group in both the non-obese and obese groups had higher physical inactivity levels than risk allele group (TA + AA or TC + CC).
FTO frequencies, Hardy–Weinberg equilibrium, and influence of FTO variants on obesity risk
The frequencies of both FTO SNPs are presented in Table 2. The distributions of genotypes and alleles for rs9939609 and rs1421085 polymorphisms were compared between the non-obese and obese groups (Table 2). Genotypes of rs9939609 and rs1421085 polymorphisms were found to be in the Hardy–Weinberg equilibrium (p > 0.05) for both non-obese and obese groups. Our findings indicated that the genotypic and allelic frequencies of both polymorphisms did not differ significantly between the groups. However, the frequency of the rs9939609 (AA) genotype was higher in the obese group than in the non-obese group. Allelic analyses revealed that the prevalence of the rs9939609 (A) allele was higher in the obese group (0.43 vs. 0.39), with rs9939609 (A) allele carriers having a 1.18 times higher risk of developing this phenotype. Similarly, the frequency of the rs1421085 (CC) genotype was higher in the obese group than in the non-obese group. Furthermore, CC genotype carriers had a 1.17-fold increased risk of obesity. In addition, carriers of at least one rs1421085 (C) allele were 1.28 times more likely to develop this obesity phenotype.
Association of FTO variants with anthropometric parameters
The anthropometric and blood pressure values of the non-obese and obese groups according to the dominant FTO genotypes and both SNPs are presented in Table 3. The variables in homozygous dominant (TT) group in rs9939609 were higher in the obese group than in the non-obese group. Accordingly, the parameters in the non-homozygous dominant (TA + AA) group were higher in the obese group than in the non-obese group. In both the non-obese and obese groups, all parameters in the homozygous dominant (TA + AA) group of rs9939609 were higher than those in the homozygous dominant (TT) group; however, the difference was not significant. As the same results were observed in rs1421085, both the non-homozygous dominant (TC + CC) and homozygous dominant group (TT) groups showed significantly higher parameters in the obese group than in the non-obese group. However, if the genotypes were compared between the obese and non-obese groups, other parameters were higher in non-homozygous dominant (TC + CC) participants than in homozygous dominant (TT) participants, and the results were not significantly different.
Association of FTO variants with biochemical-, inflammation-, and hormone-related obesity
Data on the blood biochemistry, blood inflammation, and hormonal levels of non-obese and obese groups according to the dominant genotypes of FTO rs9939609 and rs1421085 are presented in Table 4. Regarding the homozygous dominant (TT) genotype in rs9939609, most variables, except TC and LDL-C levels, in the obese group were significantly higher than those in the non-obese group. In addition, the same trend in the non-homozygous dominant (TA + AA) group indicated that the parameters, except for TC and LDL-C, were significantly higher parameters in the obese group than in the non-obese group. However, for rs9939609 in both the non-obese and obese groups, all parameters were higher in the homozygous dominant (TA + AA) group than in the homozygous dominant (TT) group, and the results were not significantly different. As the same results were observed for rs1421085, both the non-homozygous dominant (TC + CC) and homozygous dominant (TT) groups showed that parameters were significantly higher in the obesity group than in the non-obese group, except for TC and LDL-C levels. Moreover, in the obese group, leptin levels were significantly higher in non-homozygous dominant (TC + CC) than in the homozygous dominant (TT) groups (p < 0.043). However, other parameters were higher in non-homozygous dominant (TC + CC) participants than in the homozygous dominant (TT) participants in the non-obese and obese groups, and the results were not significant.
Association of FTO variants with nutrition consumption
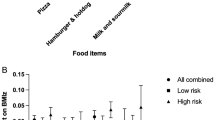
The nutritional composition of the non-obese and obese groups according to the dominant genotypes of FTO rs9939609 and rs1421085 is presented in Supplementary Table 1. Regarding the homozygous dominant (TT) genotype in rs9939609, all nutrition variables were higher in the obese group than in the non-obese group, and the results were not significant. However, the non-homozygous dominant (TA + AA) genotype indicated that the obese group had significantly higher energy, carbohydrate, sugar, protein, total fat, and SAT fat levels than the non-obese group. As the same results were obtained in rs1421085, the non-homozygous dominant (TC + CC) genotype showed significantly higher energy, carbohydrate, sugar, protein, total fat, and SAT fat levels in the obese group than in the non-obese group. However, in the homozygous dominant (TT) genotype, the carbohydrate content in the obese group was significantly higher than that in the non-obese group. Moreover, if compared with the homozygous dominant (TT) and non-homozygous dominant (TA + AA) genotypes of rs9939609 or TC + CC of rs1421085 in the non-obese and obese groups (Fig. 1A–D), in the obese group of both SNPs, the homozygous dominant (TT) group had significantly higher sugar levels than the non-homozygous dominant (TA + AA) or TC + CC (p < 0.010 and p < 0.011), respectively. In addition, Fig. 1E–H shows that in the obese group of rs9939609, the homozygous dominant (TT) group had a significantly higher sat fat content than the non-homozygous dominant (TA + AA) group (p < 0.044).
Comparison of sugar content between FTO rs1421085 (A and B) and rs993609 (C and D) and Sat fat content between FTO rs1421085 (E and F) and rs993609 (G and H) genotype in non-obese group and Obese group. The comparisons were realized by student t-test. A p < 0.05 was considered statistically significant.
Correlation of nutrition consumption with the FTO genotype and biochemical analysis with the FTO genotype
The correlation between nutrition composition and FTO rs9939609 and rs1421085 is shown in Supplementary Table 2. The positive correlation of sugar content, total fat content, and sat fat content with the FTO genotype was significant. Moreover, rs9939609 and rs1421085 were positively correlated with sugar content (r = 0.154, p = 0.003; r = 0.148, p = 0.005, respectively) and total fat content with FTO rs9939609 (r = 0.105, p = 0.045). Moreover, the sat fat content positively correlated with rs9939609 and rs1421085 (r = 0.124, p = 0.018; r = 0.113, p = 0.030, respectively). Supplementary Table 3 shows a significant positive correlation of MPO with rs1421085 and rs9939609 (r = 0.107, p = 0.038; r = 0.109, p = 0.034, respectively). Moreover, leptin showed a significant positive correlation with rs1421085 and rs9939609 (r = 0.111, p = 0.031; r = 0.113, p = 0.028, respectively).
Association of low sugar content and high-sugar content and low sat fat content and high-sat fat content with the FTO genotype
The association of the FTO genotype with low- and high-sugar content is shown in Supplementary Table 4. Significant differences were found in the obese groups of rs9939609 and rs1421085. In the obese group of rs9939609, the consumption of sugar increased 2.22 times (1.13–4.37) in the non-homozygous dominant (TA + AA) group compared with the homozygous dominant (TT) group. Moreover, in the obese group, risk allele carriers of rs1421085 (TC + CC) had sugar consumption increased 2.05 times (1.04–4.03) compared with the homozygous dominant (TT) group. Moreover, the association of the FTO genotype with low- and high-sat fat intake is shown in Supplementary Table 5. The results showed significant differences in the obese groups of rs9939609 and rs1421085. In the obese group, the sat fat consumption of the non-homozygous dominant (TA + AA) group increased 1.86 times (1.02–3.40) in rs9939609 and 1.94 times (1.06–3.55) in rs1421085 compared with those of the homozygous dominant (TT) group.
Discussion
Obesity is caused by a complex interplay between genetic makeup and environmental factors, such as low physical activity and unhealthy diets. Genetic effects on obesity depend on gene function. Genome-wide association studies have identified genetic variants linked to obesity, with the FTO being one of the most significant. FTO expression in the hypothalamus is crucial for regulating energy balance, adipogenesis, energy intake, and appetite control32. Moreover, DNA methylation changes are associated with various diseases, including obesity33.
This study involved a sample of adult populations to explore the effect of FTO variants on obesity-related risk factors, such as food consumption. FTO risk alleles (A allele of rs9939609 and C allele of rs1421085) demonstrated distinct effects on the intake of high-energy foods such as high-sugar and high-fat foods when compared between the non-obese and obese groups. Variants of both rs9939609 and rs1421085 showed a highly significant association with high BMI in the study of Mongolians34. In the present study, the homozygous variants of rs9939609 and rs1421085 were detected in 13.4% and 14.0% of the study populations, respectively. This value is close to the frequency of those genotypes in the Mongolian population34. The frequencies of FTO rs9939609 AA and rs1421085 CC genotypes were high in the obese group compared with those in the non-obese group, but not significantly. This may be due to the lower proportion of non-obese participants, as in the previous study35 (Sierra-Ruelas et al. 2022).
From the results of anthropometric analysis according to the homozygous wild type or comparison between the non-risk allele carriers in the obese and obese groups, all anthropometric parameters were higher in the obese group than in the non-obese group. However, when comparing genotypes, the results were not different, and the trend was higher in the risk allele group than in the dominant variance group. As the results were similar to previous findings36 a number of FTO SNPs were linked to HC, BMI, and total body weight. Interestingly, Zhang et al.37 found that certain FTO SNPs, such as the rs9939609 polymorphism, can affect body fat, fat-free mass, and muscle mass. Moreover, Rauhio et al.)38 analyzed premenopausal women and found that rs9939609 polymorphism was associated with body weight but not body composition or fat distribution. The AA group was heavier by approximately3.6 kg than the AT and TT groups. In addition, the BMI and FM of A-allele carriers were greater than those of non-carriers. In a Danish cohort, Andreasen et al.39 discovered that homozygous A-allele carriers weighed more than non-carriers, as seen by their BMI of 1.1 kg/m2 and 2.3 cm of WC.
In blood biochemistry, inflammation, and hormonal analyses related to obesity, the results were higher in the risk allele group than in the dominant group; however, the results were not significant. However, when the parameters (lipid profiles, blood glucose, inflammation markers, and hormones) were compared between non-obese and obese groups according to the genotype, the results were significantly different. Numerous studies have shown an association between lipid profiles and the FTO genotype; however, no correlation was observed between FTO polymorphisms and TGs, TC, LDL-C, or BMI, which is consistent with the results of the present study40 Furthermore, Khella et al. showed that the anthropometric or biochemical parameters of individuals with different FTO rs9939609 genotypes were not significantly different in all genetic models (dominant, recessive, and additive), except for HDL-C. HDL-C levels were significantly lower in AA carriers41 However, higher MPO and leptin levels showed a positive correlation with both FTO SNPs. Obesity development is linked to the activation of MPO, a heme protein that is mostly produced in neutrophil granules. Moreover, leptin levels were significantly higher in the risk allele carriers of FTO rs1421085 than in the wild-type group. Similar to a previous study42, FTO risk allele carriers have higher body fat and BMI, which in turn raises serum leptin levels. FTO rs421085 polymorphism was found to be related to resting energy expenditure (REE). In addition, the control of REE was significantly influenced by leptin. We postulated that leptin may act as a mediator between FTO and REE. As previously mentioned, FTO and leptin may be related, and FTO polymorphism was found to be linked to the expression of the leptin gene43. In this study indicating the association of FTO variants with food intake, hunger and satiety regulation was hypothesized to be related to the postprandial plasma concentrations of leptin hormones44. White adipocytes synthesize and secrete leptin in the bloodstream. Leptin regulates food intake and energy expenditure through a range of central and peripheral activities, among other significant functions. A possible explanation for the high association observed between serum leptin concentrations and body fat mass in obesity could be the existence of an endogenous leptin resistance mechanism that limits its regulatory action. A study showed that the loss of function of FTO can induce leptin resistance45 Similarly, a previous study indicated that the risk allele of FTO variants was significantly associated with higher serum leptin concentrations independently of potential confounders including adiposity and effect to energy balance46.
Moreover, this study showed that the sugar, fat, and sat fat contents are associated with FTO variance. Moreover, high-sugar and sat fat intakes were significantly higher in the A risk allele of FTO rs9939609 and C risk allele of rs1421085 in the obese group than in wild-type genotypes. Similarly, Young et al.20 demonstrated a strong positive correlation between BMI and diet scores in foods rich in proteins, food weight, and sat fat. They discovered that in individuals with higher diet scores, FTO had an improved effect on BMI20. A study showed that genes predisposed to obesity may interact with SAT fats to promote weight gain, but not with monosaturated fats or PUFAs47. Consequently, high-fat diets may be primarily responsible for the obesity pandemic because of the increased palatability and high-energy content. Furthermore, over the past few decades, a rise in the consumption of refined carbohydrates and beverages sweetened with sugar has contributed to an increase in the prevalence of obesity48. Similarly, Daya et al. conducted a case–control study of obese and non-obese participants in Jakarta to evaluate the association between the FTO rs9939609 polymorphism and obesity risk and liking for fatty foods, and their results contradict our findings. They found that those with AT/AA genotypes consumed 5.98 times more dietary fat than people with TT genotypes22. In another study on Indonesian female teens, Susmiati et al.49 found that those with AT/AA genotypes ate more fried foods and ate fewer fruits than those with TT genotypes49. The FTO rs9939609 polymorphism is essential in appetite regulation, as confirmed by Cecil et al.50 The possible mechanism of this polymorphism is caused by a hyperphagic phenotype and a preference for high-energy foods. FTO is abundantly expressed in the hypothalamus, and the FTO A allele increases the risk of increasing food intake by decreasing the central processing of satiety51. Numerous studies have demonstrated that nutrition has a major effect on obesity development52. Recent studies have looked into the connection between gene variants and specific dietary nutrient intake in relation to obesity. Increased consumption of protein, carbohydrates, fats, and added sugars has been linked to excessive energy intake53. The FTO protein has 505 amino acids and dioxygenase that is dependent on alpha-ketoglutarate. Through oxidative demethylation, alkylated DNA and RNA are restored. It particularly demethylates N (6)-methyl adenosine (RNA, the most common internal messenger RNA (mRNA) modification in higher eukaryotes54. In blood and fibroblasts, the FTO transcripts with the A (risk) allele of rs9939609 were more prevalent than in those with the T allele55. Notably, individuals homozygous for the FTO rs9939609 AA allele have dysregulated orexigenic hormone acyl-ghrelin in appetite-regulating brain regions, which modulates the brain’s homeostatic and reward regions’ neural responses to food images56. In addition to the central effect, FTO variations may affect cellular metabolism. Because rs9939609 and rs1421085 (T > C) are in linkage disequilibrium, obesity may result from a disruption of AR1D5B-mediated regulation of Irx3 and Irx557. Consequently, the mitochondria’s whitening and browning programs change, and their mitochondrial thermogenesis decreases56 (Then, the possible mechanism discovered in this study showed the interaction of gene-diet and leptin hormone found in a previous study. Studies on obesity caused by a high-fat diet (HFD) have repeatedly linked it to FTO. The regulation of energy balance is mostly dependent on the hypothalamus, and obesity is a result of hypothalamic leptin resistance generated by HFD. FTO, an N6-methyladenosine (m6A) RNA methylation regulator, is a possible player in the development of leptin resistance45.
This study investigated the effect of FTO SNPs on food preferences in adult Thai people, revealing interactions between two SNPs well-known for their connection to obesity. While previous studies have examined genetic factors related to obesity in Thai populations, our research expands on these findings by exploring the interaction between FTO gene variants and dietary habits in a larger sample. The strength of this study lies in its examination of how genetic variance and dietary interaction may influence obesity development in Thai people, showing a positive correlation between high-energy food intake and genotype. However, some limitations should be considered when interpreting our results. The sample size, particularly in subgroup analyses, was relatively small, which may have contributed to the absence of significant differences in some analyses. Our study was conducted in Bangkok, which may not fully represent the genetic and dietary diversity of the entire Thai population. We emphasize that our findings should be interpreted cautiously when considering broader generalization. The cross-sectional nature of our study limits our ability to determine causal relationships between FTO gene polymorphisms and obesity-related outcomes. While we can identify associations, this design does not allow us to infer direct causality. Despite efforts to control for confounding variables, there may be other unmeasured factors influencing the observed associations. Our focus on specific FTO polymorphisms (rs9939609 and rs1421085) and their dietary interactions, while important, does not represent the full complexity of genetic influences on obesity. Future studies should consider a wider range of genetic variants and their potential interactions, using larger and more diverse samples across different regions of Thailand.
Despite these limitations, our study provides valuable insights into the relationship between FTO polymorphisms, dietary habits, and obesity-related parameters in a Thai population sample. These findings can serve as a foundation for future, more comprehensive studies in this area.
Conclusions
Our study provides important insights into the relationship between FTO polymorphisms, high-energy food intake, and obesity-related parameters in the Thai population. These findings serve as a foundation for developing targeted, personalized strategies that consider both genetic predisposition and dietary habits in addressing obesity. Future research should focus on establishing causal relationships, investigating underlying mechanisms, and expanding to larger, more diverse populations to further our understanding of gene-diet interactions in obesity risk.
Data availability
The datasets employed and/or examined in this study are accessible on the request from the corresponding author
References
Lin, X. & Li, H. Obesity: Epidemiology, pathophysiology, and therapeutics. Front Endocrinol. (Lausanne) 12, 706978 (2021).
Sakboonyarat, B. et al. Rising trends in obesity prevalence among Royal Thai Army personnel from 2017 to 2021. Sci. Rep. 12, 7726 (2022).
Hruby, A. & Hu, F. B. The epidemiology of obesity: A big picture. Pharmacoeconomics 33, 673–689 (2015).
Hill, J. O. et al. Energy balance and obesity. Circulation 126, 126–132 (2012).
Albuquerque, D. et al. The contribution of genetics and environment to obesity. Br. Med. Bull. 123, 159–173 (2017).
Popović, A. M. et al. FTO gene polymorphisms at the crossroads of metabolic pathways of obesity and epigenetic influences. Food Technol. Biotechnol. 61, 14–26 (2023).
Inandiklioğlu, N. & Yaşar, A. Association between rs1421085 and rs9939609 polymorphisms of fat mass and obesity-associated gene with high-density lipoprotein cholesterol and triglyceride in obese Turkish children and adolescents. J. Pediatr. Genet. 10, 9–15 (2021).
Ağagündüz, D. & Gezmen-Karadağ, M. Association of FTO common variant (rs9939609) with body fat in Turkish individuals. Lipids Health Dis. 18, 212 (2019).
Song, Y. et al. Polymorphisms of fat mass and obesity-associated gene in the pathogenesis of child and adolescent metabolic syndrome. Nutrients 15, 2643 (2023).
Chuenta, W. et al. Common variations in the FTO gene and obesity in Thais: A family-based study. Gene 558, 75–81 (2015).
Rahimlou, M. et al. Fat mass and obesity-associated gene (FTO) rs9939609 (A/T) polymorphism and food preference in obese people with low-calorie intake and non-obese individuals with high-calorie intake. BMC Nutr. 9, 143 (2023).
Chuang, Y. F. et al. FTO genotype and aging: Pleiotropic longitudinal effects on adiposity, brain function, impulsivity and diet. Mol. Psychiatry 20, 133–139 (2015).
Saber-Ayad, M. et al. The FTO genetic variants are associated with dietary intake and body mass index amongst Emirati population. PLoS One 14, e0223808 (2019).
Harbron, J. et al. Fat mass and obesity-associated (FTO) gene polymorphisms are associated with physical activity, food intake, eating behaviors, psychological health, and modeled change in body mass index in overweight/obese Caucasian adults. Nutrients 6, 3130–3152 (2014).
Melhorn, S. et al. FTO genotype impacts food intake and corticolimbic activation. Am. J. Clin. Nutr. 107, 145–154 (2018).
Al-Jawadi, A. A. et al. Association of FTO rs1421085 single nucleotide polymorphism with fat and fatty acid intake in Indonesian adults. BMC Res. Notes 14, 411 (2021).
Rana, S. & Bhatti, A. A. Association and interaction of the FTO rs1421085 with overweight/obesity in a sample of Pakistani individuals. Eat. Weight Disord. 25, 1321–1332 (2020).
Laber, S. et al. Linking the FTO obesity rs1421085 variant circuitry to cellular, metabolic, and organismal phenotypes in vivo. Sci Adv 7, eabg0108 (2021).
Qi, Q. et al. FTO genetic variants, dietary intake and body mass index: Insights from 177,330 individuals. Hum. Mol. Genet. 23, 6961–6972 (2014).
Young, A. I. et al. Multiple novel gene-by-environment interactions modify the effect of FTO variants on body mass index. Nat. Commun. 7, 12724 (2016).
Chiang, K.-M. et al. Genome-wide association study of morbid obesity in Han Chinese. BMC Genet. 20, 97 (2019).
Daya, M. et al. Obesity risk and preference for high dietary fat intake are determined by FTO rs9939609 gene polymorphism in selected Indonesian adults. Asia Pac. J. Clin. Nutr. 28, 183–191 (2019).
Boonyuen, U. et al. Glucose-6-phosphate dehydrogenase mutations in malaria endemic area of Thailand by multiplexed high-resolution melting curve analysis. Malar. J. 20, 194 (2021).
Katayutanon, T. et al. Factors relating to physical activities of the elderly in a Village, Bangpla Subdistrict, Bangplee District, Samutprakarn Province. HCU J. 23, 1–14 (2019).
Adıgüzel, İ et al. Adaptation of the global physical activity questionnaire (GPAQ) into Turkish: A validation and reliability study. Turk. J. Phys. Med. Rehabil. 67, 175–186 (2021).
Sama, S. et al. Quantifying the homeostatic model assessment of insulin resistance to predict mortality in multi-organ dysfunction syndrome. Indian J. Crit. Care Med. 25, 1364–1369 (2021).
Ghosh, S. & Collier, A. Diagnosis, classification, epidemiology and biochemistry. In Churchill’s pocketbook of diabetes (eds Ghosh, S. & Collier, A.) 1–49 (Churchill Livingstone, 2012).
Nirdnoy, N. et al. Validation of a Thai semiquantitative food frequency questionnaire (semi-FFQ) for people at risk of metabolic syndrome. J. Health Popul. Nutr. 42, 13 (2023).
Witek, K. et al. A high-sugar diet consumption, metabolism and health impacts with a focus on the development of substance use disorder: A narrative review. Nutrients 14, 2940 (2022).
Liu, A. G. et al. A healthy approach to dietary fats: Understanding the science and taking action to reduce consumer confusion. Nutr. J. 16, 53 (2017).
Lim, J. U. et al. Comparison of world health organization and asia-pacific body mass index classifications in COPD patients. Int. J. Chron. Obstruct. Pulmon. Dis. 12, 2465–2475 (2017).
Huang, C. et al. Studies on the fat mass and obesity-associated (FTO) gene and its impact on obesity-associated diseases. Genes Dis. 10, 2351–2365 (2023).
Lan, N. et al. FTO—a common genetic basis for obesity and cancer. Front. Genet. 11, 559138 (2020).
Zhang, Q. et al. Relationship between fat mass and obesity-associated (FTO) gene polymorphisms with obesity and metabolic syndrome in ethnic Mongolians. Med. Sci. Monit. 24, 8232–8238 (2018).
Sierra-Ruelas, E. et al. The rs9939609 variant in FTO increases the risk of hypercholesterolemia in metabolically healthy subjects with excess weight. Lifestyle Genom. 15, 131–138 (2022).
Scuteri, A. et al. Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet. 3, e115 (2007).
Chen, Y. et al. A modified formula for calculating low-density lipoprotein cholesterol values. Lipids Health Dis. 9, 52 (2010).
Rauhio, A. et al. Association of the FTO and ADRB2 genes with body composition and fat distribution in obese women. Maturitas 76, 165–171 (2013).
Andreasen, C. H. et al. Low physical activity accentuates the effect of the FTO rs9939609 polymorphism on body fat accumulation. Diabetes 57, 95–101 (2008).
Franczak, A. et al. Association between FTO gene polymorphisms and HDL cholesterol concentration may cause higher risk of cardiovascular disease in patients with acromegaly. Pituitary 21, 10–15 (2018).
Khella, M. S. et al. The (FTO) gene polymorphism is associated with metabolic syndrome risk in Egyptian females: A case-control study. BMC Med. Genet. 18, 101 (2017).
Hebbar, P. et al. FTO variant rs1421085 associates with increased body weight, soft lean mass, and total body water through interaction with ghrelin and apolipoproteins in Arab population. Front. Genet. 10, 1411 (2019).
Najd Hassan Bonab, L. et al. The role of FTO variant rs1421085 in the relationship with obesity: A systematic review and meta-analysis. Eat. Weight Disord. Stud. Anorexia Bulimia Obes. 27, 3053–3062 (2022).
Speakman, J. R. The “fat mass and obesity related” (FTO) gene: Mechanisms of impact on obesity and energy balance. Curr. Obes. Rep. 4, 73–91 (2015).
Liu, S. et al. Hypothalamic FTO promotes high-fat diet-induced leptin resistance in mice through increasing CX3CL1 expression. J. Nutr. Biochem. 123, 109512 (2024).
Labayen, I. et al. Association between the FTO rs9939609 polymorphism and leptin in European adolescents: A possible link with energy balance control. The HELENA study. Int. J. Obes. (Lond) 35, 66–71 (2011).
Liu, J. et al. Genetic and environmental influences on nutrient intake. Genes Nutr. 8, 241–252 (2013).
Malik, V. S. et al. Sugar-sweetened beverages, obesity, type 2 diabetes mellitus, and cardiovascular disease risk. Circulation 121, 1356–1364 (2010).
Susmiati, S. et al. Association of fat mass and obesity-associated rs9939609 polymorphisms and eating behaviour and food preferences in adolescent Minangkabau girls. Pak. J. Nutr. 17, 471–479 (2018).
Cecil, J. E. et al. An obesity-associated FTO gene variant and increased energy intake in children. N. Engl. J. Med. 359, 2558–2566 (2008).
Melhorn, S. et al. FTO genotype impacts food intake and corticolimbic activation. Am J Clin Nutr 107, 145–154 (2018).
Jiao, J. The role of nutrition in obesity. Nutrients 15, 2556 (2023).
Doo, M. & Kim, Y. Obesity: Interactions of genome and nutrients intake. Prev. Nutr. Food Sci. 20, 1–7 (2015).
Gulati, P. & Yeo, G. S. H. The biology of FTO: From nucleic acid demethylase to amino acid sensor. Diabetologia 56, 2113–2121 (2013).
Lurie, G. et al. The obesity-associated polymorphisms FTO rs9939609 and MC4R rs17782313 and endometrial cancer risk in non-Hispanic white women. PLoS One 6, e16756 (2011).
Karra, E. et al. A link between FTO, ghrelin, and impaired brain food-cue responsivity. J. Clin. Invest. 123, 3539–3551 (2013).
Schneeberger, M. Irx3, a new leader on obesity genetics. EBiomedicine 39, 19–20 (2019).
Acknowledgements
We extend our sincere gratitude to the participants for their contribution, as well as to the staff of Laksi District, Bangkok, for their invaluable assistance in facilitating the study.
Funding
This research paper is supported by Specific League Funds from Mahidol University awarded to PP.
Author information
Authors and Affiliations
Contributions
PP designed experiments. SP & PP participated in all experiments. PP, UB, CC, NS, KK assisted in advice the experiment. UB assist for HRM assay. SP & PP draft manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Poosri, S., Boonyuen, U., Chupeerach, C. et al. Association of FTO variants rs9939609 and rs1421085 with elevated sugar and fat consumption in adult obesity. Sci Rep 14, 25618 (2024). https://doi.org/10.1038/s41598-024-77004-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-77004-6
Keywords
This article is cited by
-
Multiomics: the intersection of personalized nutrition in cardiometabolic diseases
Journal of Translational Medicine (2026)
-
The diet–gene interaction: FTO rs9939609 and its role in obesity and metabolic syndrome among Asian populations
Egyptian Journal of Medical Human Genetics (2026)
-
Nutrigenomics meets multi-omics: integrating genetic, metabolic, and microbiome data for personalized nutrition strategies
Genes & Nutrition (2025)
-
Lactiplantibacillus plantarum from Thai fermented pork with inulin ameliorates metabolic disturbances through proteomic mechanisms
Scientific Reports (2025)
-
FTO rs9939609 polymorphism is associated with dietary quality in Korean females
European Journal of Nutrition (2025)