Abstract
The emergence of coinfection with Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia, and Trypanosoma evansi is an important problem that endangers human health, animal quality and public sanitary security. These pathogenic protozoans play important roles in establishment of similar clinical signs of diseases in humans, pigs, sheep, and rabbits, including fever, diarrhea, hepatitis, encephalitis, and reproductive disorders. Therefore, a rapid and specific diagnostic method to simultaneously detect these five pathogens is urgently required. Here, we developed a TaqMan-probe-based quantitative real-time polymerase chain reaction (qPCR) for the simultaneous detection of these five pathogens for the first time. Specific primers and probes were designed targeting the G3PDH gene of Toxoplasma gondii, the NC5 gene of Neospora caninum, the ADF gene of Eimeria stiedai, the GDH gene of Giardia lamblia, the COX1 gene of Trypanosoma evansi, and a TaqMan-probe-based pentaplex qPCR assay capable of simultaneously detecting these five pathogens was developed. The assay showed strong specificity, with no cross-reactivity detected against nucleic acids from other control pathogens. The assay demonstrated high sensitivity, with the lower limit of quantification (LLOQ) of 10 copies per reaction for the recombinant plasmid standards pTgG3PDH, pNcNC5, pEsADF, pGlGDH, pTeCOX1, and the limit of detection (LOD) as low as 1.1 copies. The standard curves exhibited excellent linearity (correlation coefficient values of 0.996, 0.996, 0.996, 0.992, and 0.996, respectively) and high amplification efficiencies (95.534%, 96.203%, 107.818%, 100.851%, and 104.487%, respectively). The assay also exhibited excellent repeatability and reproducibility, with inter- and intra-assay coefficient of variation (CV) ranging from 0.07% to 2.13%. The anti-interference test showed that high-concentration nucleic acids did not interfere with low-concentration nucleic acids, thus ensuring excellent detection results even in complex samples. Among 210 clinical samples, the assay detected Toxoplasma gondii in 2.38%, Neospora caninum in 2.38%, Eimeria stiedai in 10%, and Giardia lamblia in 36.19%. The coinfection rates were 8.1% for Eimeria stiedai & Giardia lamblia. This TaqMan-probe-based pentaplex qPCR assay offers a rapid, sensitive, and specific tool for the simultaneous detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia, and Trypanosoma. It is of great significance to safeguard human health.
Similar content being viewed by others
Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi are all pathogenic protozoans. These pathogenic protozoa can cause similar disease symptoms in humans, sheep, pigs, rabbits and other animals, such as fever, diarrhea, hepatitis, encephalitis and reproductive disorders. Toxoplasma gondii can cause disease in humans, with clinical outcomes ranging from asymptomatic presentations to ocular disease1,2, cerebral, neurological symptoms3, reproductive disturbances4, and congenital toxoplasmosis in the fetus, such as abortion, hydrocephalus, cerebral calcifications, chorioretinitis, lymphadenopathy and microcephaly5,6,7,8. It is important to note that toxoplasmosis can even cause fatal or high-risk severe diseases in individuals with compromised immune systems, such as AIDS patients9 and organ transplant recipients10,11,12,13,14. Neospora caninum infection is an important cause of abortion15,16. Infection with Eimeria stiedai can lead to hepatic coccidiosis17,18. Giardia lamblia is associated with gastroenteritis and malabsorption syndrome19,20. Trypanosoma evansi is a pathogenic protozoan found in the plasma and hematopoietic organs of animals21,22 and humans23, which is responsible for fever causing Surra in a variety of mammalian hosts over a wide geographical area24,25. Therefore, rapid and sensitive detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi is of great significance.
Currently, the diagnosis of parasitic diseases mainly relies on traditional microscopic examination methods. However, this method is cumbersome, time-consuming, labor-intensive, and often lacks sensitivity and specificity. This may result in cases with indistinct parasitic morphological characteristics being overlooked, which could potentially cause environmental pollution and endanger the safety of the inspectors26.
The detection of pathogen-specific antibody in immunological method is the main diagnostic tool27,28, but this method is complicated to operate or has problem in terms of specificity and sensitivity. In addition, for immunosuppressed individual, serological testing may result in false negative or false positive result due to individual immune status or reactivation of toxoplasmosis29. Serological assays often suffer from significant limitations such as cross-reactivity with closely related organisms and prolonged antibody persistence even after successful treatment, both of which can lead to false-positive results.
One common nucleic acid-based detection method is the polymerase chain reaction (PCR). Due to its high sensitivity and the availability of commercial kits, it has become the gold standard for pathogen monitoring. However, PCR-based methods are usually cumbersome and time-consuming because both the traditional agarose gel electrophoresis and the screening process of amplification products require a significant amount of time and effort. The sensitivity of conventional PCR detection methods is relatively low. Moreover, due to factors such as opening test tubes, conventional PCR methods are prone to contamination and are also susceptible to the influence of inhibitors that may exist in the samples30,31,32,33,34.
The loop-mediated isothermal amplification (LAMP) technique is simple and rapid to operate. This technique is carried out under constant temperature conditions. Minor contamination in the environment may lead to the occurrence of false positive results, thereby affecting the accuracy of the experiment. The detection of LAMP products relies on turbidity observation, but in some cases, its sensitivity and accuracy may not be high enough35.
TaqMan-probe-based quantitative real-time PCR (qPCR) is a promising tool for detecting and quantifying the target genomes of infectious agents in different samples (blood, tissue, body fluid, feces). Its principle is to achieve this by continuously measuring the cumulative fluorescence signal during the amplification reaction. Due to the advantages of simplicity in operation, rapidity and convenience, high sensitivity, good repeatability, and low contamination rate, it is widely used in various fields such as medical testing, drug efficacy assessment, gene expression research, transgenic research, gene detection, animal and plant detection, food detection, and pathogen detection. For example, TaqMan-probe-based single-plex qPCR assay for Toxoplasma gondii36, Neospora caninum37, Giardia lamblia38, Dictyocaulus filaria39, and Clostridium piliforme40 have been reported previously.
Multiplex qPCR involves the detection of multiple target genes within a single tube. To distinguish the fluorescence signals of these target genes, it is impossible to achieve this by using SYBR Green I dye to label the amplification products. Therefore, the probe method is commonly used for multiplex qPCR41. That is, for each target gene, specific probes are designed, and by using the fluorescent groups labeled on different probes and combining with the detection capabilities of different channels of the instrument, real-time quantitative detection of multiple targets can be achieved. Recently, a TaqMan-probe-based duplex qPCR assay has been reported for the detection of Toxoplasma gondii and Neospora caninum42.
The emergence of coinfection with Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi is a serious problem, which poses a threat to human health, animal quality and public health security. They are prone to mixed infections or secondary infections, making clinical differentiation extremely difficult. To address these challenges, we designed specific primers and probes targeting the G3PDH gene of Toxoplasma gondii, the NC5 gene of Neospora caninum, the ADF gene of Eimeria stiedai, the GDH gene of Giardia lamblia, and the COX1 gene of Trypanosoma evansi. We developed a TaqMan-probe-based pentaplex qPCR assay which can simultaneously detect Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. This provides an efficient and convenient tool for the prevention and control of parasitic diseases.
Materials and methods
Ethics statement
Clinical trial number: not applicable. This study was conducted with the approval of the Animal Ethics Committee of National Institutes for Food and Drug Control [Approval ID: 2019 (A) 141].
Design of primers and probes and synthesis of recombinant plasmids
The sequences of the G3PDH gene from Toxoplasma gondii43 (GenBank: LN714499、AF179871), the NC5 gene from Neospora caninum43 (GenBank: LN714488、X84238), the ADF gene from Eimeria stiedai (GenBank: JF358017), the GDH gene from Giardia lamblia (GenBank: CP110919), and the COX1 gene from Trypanosoma evansi44 (GenBank: MW513373) were aligned with target sequences using the National Center for Biotechnology Information (NCBI) Basic Local Alignment Search Tool (BLAST) (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Sequences showing high similarity or consistency with the target genes were downloaded and analyzed for specific and conserved regions using DNASTAR software (7.1 version) and Primer Express Software Version 3.0 (Thermo Fisher Scientific, Waltham, MA, USA). The conserved regions were selected for the synthesis of plasmid standards: pTgG3PDH, pNcNC5, pEsADF, pGlGDH and pTeCOX1, and the copy number of them were calculated with the formula {plasmid copy number/µL = [6.02 × 1023 × plasmid concentration (ng/µL) × 10− 9]/[plasmid length × 660]}45. Specific primers and probes for the TaqMan-probe-based pentaplex qPCR assay were designed, with the 5′ end labeled with FAM, VIC, ABY, JUN, and CY5 fluorescent reporter dyes, and the 3′ end labeled with the corresponding QSY or MGB fluorescence quenching groups. Primers and probes demonstrating strong specificity and high sensitivity were identified through experimental screening (Table 1).
Parasites and clinical samples
This laboratory maintains Strongyloides stercoralis, Aspiculuris tetraptera, Syphacia obvelata, Syphacia muris, and Balantidium coli. Clinical samples collected in 2019–2023 were preserved at − 80℃ in our laboratory. The blood and tissue samples from sheep and rabbits were mainly collected in Jilin province and Beijing of China. DNA preserved at − 20℃ was used as PCR template. All positive samples were identified by conventional PCR in our laboratory and confirmed with DNA sequencing (Takara Biomedical Technology Co., Ltd., Beijing, China).
DNA extraction
According to the manufacturer′s instructions, total DNA was extracted from clinical samples using the DNeasy Blood & Tissue Kit (Qiagen, Germany). Parasite DNA was extracted using the QIAamp DNA Mini Kit (Qiagen, Germany). The purity, yield and extraction efficiency (EE) of DNA were monitored by using Xeno Internal Positive Control (IPC) DNA spiking.
Optimization of TaqMan-probe-based pentaplex qPCR
The primer and probe concentration, and annealing temperatures of TaqMan-probe-based pentaplex qPCR, were optimized to set up the optimal reaction system and conditions for simultaneous detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. In a brief, the TaqMan-probe-based pentaplex qPCR reactions were performed in 20 µL volumes, including 10 µL of TaqPath™ ProAmp™ Multiplex Master Mix (Thermo Fisher Scientific, Waltham, MA, USA), 3 µL of primers/probes mix (0.22 µM of forward primer GZQTgG3PDH-F, 0.22 µM of reverse primer GZQTgG3PDH-R, and 0.06 µM of TaqMan-probe GZQTgG3PDH-P for Toxoplasma gondii, 0.22 µM of forward primer GZQNcNC5-F, 0.22 µM of reverse primer GZQNcNC5-R, and 0.06 µM of TaqMan-probe GZQNcNC5-P for Neospora caninum, 0.22 µM of forward primer GZQEsADF-F, 0.22 µM of reverse primer GZQEsADF-R, and 0.06 µM of TaqMan-probe GZQEsADF-P for Eimeria stiedai, 0.22 µM of forward primer GZQGlGDH-F, 0.22 µM of reverse primer GZQGlGDH-R, and 0.06 µM of TaqMan-probe GZQGlGDH-P for Giardia lamblia, and 0.44 µM of forward primer GZQTeCOX1-F, 0.44 µM of reverse primer GZQTeCOX1-R, and 0.12 µM of TaqMan-probe GZQTeCOX1-P for Trypanosoma evansi in final concentration), 5 µL of template DNA, and 2 µL of nuclease-free water. Amplification reactions were carried out using the QuantStudio™ 6 Real-time PCR System. Amplification procedure was as follows: 50℃ for 2 min (enzyme activate); 95℃ for 10 min (initial denature); 40 cycles were performed at 95℃ for 15 s (denature) and 60℃ for 1 min (anneal). The FAM, VIC, ABY, JUN, and CY5 fluorescence signals intensity were recorded at the end of each cycle. Quantification cycle (Cq) values were generated with the QuantStudio Design & Analysis Software under automated threshold setting. Positive controls, negative control, and internal controls (IPC) were contained in each assay.
Specificity of the TaqMan-probe-based pentaplex qPCR assay
DNA was extracted from Strongyloides stercoralis, Aspiculuris tetraptera, Syphacia obvelata, Syphacia muris, and Balantidium coli, as non-target genes. The recombinant plasmid standards of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi served as positive controls while nuclease-free water as a negative control, and XenoInternal Positive Control – VIC Assay as an internal control (IPC). The samples were detected using the established TaqMan-probe-based pentaplex qPCR assay.
Efficiency (E), lower limit of quantification (LLOQ), and limit of detection (LOD) of the TaqMan-probe-based pentaplex qPCR assay
To evaluate the linearity, dynamic range, amplification efficiency (E), lower limit of quantification (LLOQ), and limit of detection (LOD) of the TaqMan-probe-based pentaplex qPCR assay46,47,48,49, serial dilutions of DNA standards of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi were tested as templates. A final standard curve was generated based on the Cq value and the logarithm of standard copy number. The slope of the linear regression of a plot of Cq (y-axis) vs. the log of the target concentration (x-axis) is determined by the amplification efficiency (E)50, which is calculated as %E = 100 × (− 1 + 10 (−1/slope)).
Repeatability and reproducibility of the TaqMan-probe-based pentaplex qPCR assay
The recombinant plasmid standards pTgG3PDH, pNcNC5, pEsADF, pGlGDH and pTeCOX1 were diluted from 5 × 107 copies/µL to 5 × 103 copies/µL using a gradient of TE buffer. These dilutions were then mixed in equal volumes to achieve final concentrations ranging from 1 × 107 copies/µL to 1 × 103 copies/µL using the optimized TaqMan-probe-based pentaplex qPCR. Each dilution of the DNAs carried out in triplicate in the same day for the intra-assay repeatability, and each dilution of the DNAs were tested with three replicates in three different days during a week for the inter-assay reproducibility. The standard deviation (SD) value and the coefficient of variation (CV) of the Cq values were calculated with the formula CV = (Standard Deviation/Mean) × 100%.
Anti-interference test of the TaqMan-probe-based pentaplex qPCR assay
Plasmid concentrations of 1 × 108 copies/µL and 1 × 102 copies/µL were selected, and standard plasmid concentrations of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi were randomly combined. Three parallel samples were prepared for each experiment to observe changes in Cq values at low concentrations, assessing whether high concentrations affected amplification at low concentrations.
Simulation of coinfection by combining different concentration of standard samples
DNA standards of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi at the different concentration were randomly chosen and mixed as templates and detected using the TaqMan-probe-based pentaplex qPCR assay.
Validation of the TaqMan-probe-based pentaplex qPCR assay
Ten clinical samples of rabbits with diarrhea were simultaneously detected using both the developed TaqMan-probe-based pentaplex qPCR assay in this study and the conventional PCR. The conventional PCR primers were synthesized by Takara Biomedical Technology Co., Ltd., Beijing, China (Table 2). The total volume of the conventional PCR reaction was 50 µL, consisting of 5 µL of 10×Buffer (Mg2+ Plus), 4 µL of dNTP Mixture, 0.5 µL of forward primer (10 µM), 0.5 µL of reverse primer (10 µM), 0.25 µL of EX Taq (5 U/µL) (Takara Biomedical Technology Co., Ltd., Beijing, China), 2 µL of template DNA, 37.75 of nuclease-free water. Amplification was carried out on a Verity 96 Thermal Cycler Instrument (Applied Biosystems) using the following program: 94℃ for 5 min, 1 cycle; 95℃ for 30 s, 60℃ for 30 s, 72℃ for 1 min, 35 cycles; 72℃ for 10 min, 1 cycle. The PCR amplification products were detected using electrophoresis on 3% agarose gel and visualized under UV light after nucleic acid staining. Detection results from the developed TaqMan-probe-based pentaplex qPCR assay were compared with the conventional PCR.
Clinical performance of the TaqMan-probe-based pentaplex qPCR assay
The TaqMan-probe-based pentaplex qPCR assay was employed to investigate the detection rates of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi in 10 samples of rabbits with diarrhea, and 200 samples of sheep without any symptoms. Samples were collected in Jilin province and Beijing of China between June 2019 and September 2023. Nucleic acids were isolated with the DNA Extraction Kit. The constructed plasmid served as the positive control, while nuclease-free water as a negative control, and XenoInternal Positive Control – VIC Assay as an internal control (IPC). Detection rates were analyzed following the acquisition of assay results for all clinical samples.
Statistical analysis
Statistical analysis was conducted using GraphPad Prism 8 software (GraphPad Software, La Jolla, CA). Data are expressed as the mean ± standard deviation of three replicates. Correlation coefficient (R2) values and amplification efficiencies (E) of standard curves were calculated and visualization of TaqMan-probe-based pentaplex qPCR amplification curves was done by QuantStudio Design & Analysis Software (version: 1.4.1).
Results
Specificity of the TaqMan-probe-based pentaplex qPCR assay
As shown in Fig. 1, Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi produced typical amplification curves, while no amplification curves or Cq values were observed for Strongyloides stercoralis, Aspiculuris tetraptera, Syphacia obvelata, Syphacia muris, and Balantidium coli, demonstrating the high specificity of the method.
Specificity of the TaqMan-probe-based pentaplex qPCR assay for simultaneously detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. Amplification curves represent samples positive for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi detected using the TaqMan-probe-based pentaplex qPCR assay; negative samples include Aspiculuris tetraptera, Strongyloides stercoralis, Syphacia obvelata, Syphacia muris, and Balantidium coli, and nuclease-free water was used as negative control.
Efficiency (E) of the TaqMan-probe-based pentaplex qPCR assay
As shown in Fig. 2, fluorescence signals were detected in three parallel samples for each concentration gradient. Five standard curves were plotted, with the logarithm of the number of starting templates on the x-axis and the Cq values on the y-axis.
Establishment of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia, and Trypanosoma evansi standard curves with abscissa as logarithm of copy number and ordinate as Cq value, the correlation coefficient (R2) were 0.996, 0.996, 0.996, 0.992, and 0.996, the slopes of the equation were − 3.434, − 3.416, − 3.148, − 3.302, and − 3.219, the intercepts were 38.298, 40.457, 37.072, 38.099, and 39.264, and the amplification efficiencies (E) were 95.534%, 96.203%, 107.818%, 100.851%, and 104.487%, respectively.
The standard formulas for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia, and Trypanosoma evansi were y = − 3.434x + 38.298 (Fig. 2A, y-axis is Cq value and x-axis is the logarithm of concentration), y = − 3.416x + 40.457 (Fig. 2B), y = − 3.148x + 37.072 (Fig. 2C), y = − 3.302x + 38.099 (Fig. 2D), and y = − 3.219x + 39.264 (Fig. 2E), respectively.
The correlation coefficient of standard curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi were 0.996, 0.996, 0.996, 0.992, and 0.996, respectively, suggesting that the correlation between each quantified concentration group of standards is reliable.
The amplification efficiency (E) of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi were 95.534%, 96.203%, 107.818%, 100.851%, and 104.487%, respectively, regarded as satisfactory for the TaqMan-probe-based pentaplex qPCR assay.
With the correlation coefficient values close to 1 and amplification efficiencies within the optimal range from 90% to 110%, these results indicated a strong linear correlation between the template DNA concentrations and the corresponding Cq values (Fig. 2).
This demonstrates the reliability of the TaqMan-probe-based pentaplex qPCR developed in this study for quantitative analysis of nucleic acids in amplified samples.
Standard curves of the TaqMan-probe-based pentaplex qPCR assay for simultaneously detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. (A) Toxoplasma gondii; (B) Neospora caninum; (C) Eimeria stiedai; (D) Giardia lamblia; (E) Trypanosoma evansi; (F) All standard curves. Standard curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi for DNA standards at concentrations ranging from 1 × 108 to 1 × 103 copies per reaction.
- In Figure 2, please place Figure 2C and Figure 2D behind Figures 2A and 2B.- In Figure 2, please place Figure 2C and Figure 2D behind Figures 2A and 2B.Lower limit of quantification (LLOQ), and limit of detection (LOD) of the TaqMan-probe-based pentaplex qPCR assay
The 10-fold gradient dilution of the DNA standard were tested using the TaqMan-probe-based pentaplex qPCR assay (Fig. 3). The lower limit of quantification (LLOQ) of the method was 1 × 101 copies/µL for simultaneously detecting Toxoplasma gondii (Fig. 3A), Neospora caninum (Fig. 3B), Eimeria stiedai (Fig. 3C), Giardia lamblia (Fig. 3D), and Trypanosoma evansi (Fig. 3E), indicating that the TaqMan-probe-based pentaplex qPCR assay was highly sensitivity. The 2-fold gradient dilution of the DNA standard was tested by the TaqMan-probe-based pentaplex qPCR assay (Fig. 4; Table 3).
The limit of detection (LOD) for this experiment was defined as the lowest concentration of the target analyte, which can be detected with a 95% detection rate. Follow-up experiments indicated that the detection rate of samples at 2 copies/µL for Toxoplasma gondii, 1.5 copies/µL for Neospora caninum, 1.5 copies/µL for Eimeria stiedai, 1.8 copies/µL for Giardia lamblia, and 1.1 copies/µL for Trypanosoma evansi, was 100%, 100%, 100%, 100%, 100% of 20 replicates, respectively. Thus, the reliable LODs of TaqMan-probe-based pentaplex qPCR are 2, 1.5. 1.5. 1.8, and 1.1 copies per reaction for detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. We set the cutoff line for positivity at 38, meaning that the samples with a Cq value less than or equal to 35 (≤ 35) are regarded as positive, those with Cq value greater than 35 but less than or equal to 38 (35 < and ≤ 38) were considered invalid, and those with Cq value greater than 38 (> 38) were considered negative. The criteria were established based on two considerations. First, the LOD of the method was 2 copies/µL for Toxoplasma gondii, 1.5 copies/µL for Neospora caninum, 1.5 copies/µL for Eimeria stiedai, 1.8 copies/µL for Giardia lamblia, and 1.1 copies/µL for Trypanosoma evansi, with a corresponding Cq value of ཞ 35. Second, although some samples at 1.1 copies/µL were detectable, their Cq values were ཞ 35.29. All subsequent experiments were conducted in accordance with these criteria. Briefly, if there was a specific S-shaped curve and the Cq value was ≤ 35, the result was determined to be positive for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi nucleic acid. If there was no Cq value and no specific fluorescence amplification curve, the result was judged as negative for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi nucleic acid. If the Cq value was between 35 and 38 (35 < Cq value < 38) and there was a specific fluorescence amplification curve, the result was interpreted as suspicious for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi nucleic acid, and the suspicious samples were resampled for DNA extraction and retesting. If the Cq value was < 38, the result was interpreted positive; otherwise, the result was considered negative.
Lower limit of quantification (LLOQ) of the TaqMan-probe-based pentaplex qPCR assay for simultaneously detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. (A) Toxoplasma gondii; (B) Neospora caninum; (C) Eimeria stiedai; (D) Giardia lamblia; (E) Trypanosoma evansi. Amplification curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi for DNA standards at concentrations ranging from 1 × 108 to 1 × 101 copies per reaction.
Limit of detection (LOD) of the TaqMan-probe-based pentaplex qPCR assay for simultaneously detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. (A) Amplification curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi using serial dilutions of DNA standards 14.5, 15, 15, 18, and 11 copies/µL. (B) Amplification curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi using serial dilutions of DNA standards 7.25, 7.5, 7.5, 9, and 5.5 copies/µL. (C) Amplification curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi using serial dilutions of DNA standards 3.6, 3.75, 3.75, 4.5, and 2.75 copies/µL. (D) Amplification curves of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi using serial dilutions of DNA standards 2, 1.5, 1.5, 1.8, and 1.1 copies/µL.
Repeatability and reproducibility of the TaqMan-probe-based pentaplex qPCR assay
As shown in Table 4, the TaqMan-probe-based pentaplex qPCR assay had excellent intra-assay repeatability with the CV of the intra-assay from 0.07% to 0.59% for Toxoplasma gondii, from 0.07% to 0.75% for Neospora caninum, from 0.19% to 2.10% for Eimeria stiedai, from 0.28% to 1.43% to Giardia lamblia, and from 0.33% to 0.95% for Trypanosoma evansi. The TaqMan-probe-based pentaplex qPCR assay had the coefficient of variation of the inter-assay from 0.40% to 0.75% for Toxoplasma gondii, from 0.22% to 0.80% for Neospora caninum, from 0.46% to 1.49% for Eimeria stiedai, from 0.50% to 2.13% for Giardia lamblia, and from 0.26% to 0.89% for Trypanosoma evansi. The results showed that both intra- and inter-assay coefficient of variation were less than 3%, indicating good repeatability (intra-assay) and reproducibility (inter-assay) of TaqMan-probe-based pentaplex qPCR.
Anti-interference test of the TaqMan-probe-based pentaplex qPCR assay
The recombinant plasmids of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi were selected for TaqMan-probe-based pentaplex qPCR amplification at high concentrations of 1 × 108 copies/µL and low concentrations of 1 × 102 copies/µL. Changes in the thresholds of the amplification curves were observed to assess any potential interference. As shown in Table 5, no interference was detected in the amplification of any one high-concentration pathogen against the other four low-concentration pathogens, nor among any four high concentration pathogens against the low-concentration pathogens.
Assessment of coinfection in the TaqMan-probe-based pentaplex qPCR assay
In coinfection analysis, standard plasmids were used to simulate mixed infections commonly observed in clinical samples (Fig. 5). The results demonstrated that the TaqMan-probe-based pentaplex qPCR assay was capable of detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi simultaneously.
Coinfection simulation experiments with Toxoplasma gondii (Tg), Neospora caninum (Nc), Eimeria stiedai (Es), Giardia lamblia (Gl) and Trypanosoma evansi (Te). (A) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 108, 1 × 107, 1 × 106, 1 × 105, and 1 × 104 copies/µL; (B) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 108, 1 × 106, 1 × 107, 1 × 104, and 1 × 105 copies/µL; (C) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 105, 1 × 108, 1 × 107, 1 × 104, and 1 × 106 copies/µL; (D) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 106, 1 × 108, 1 × 104, 1 × 105, and 1 × 107 copies/µL; (E) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 104, 1 × 105, 1 × 108, 1 × 106, and 1 × 107 copies/µL; (F) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 104, 1 × 106, 1 × 108, 1 × 105, and 1 × 107 copies/µL; (G) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 107, 1 × 104, 1 × 105, 1 × 106, and 1 × 108 copies/µL; (H) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 105, 1 × 107, 1 × 106, 1 × 104, and 1 × 108 copies/µL; (I) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 106, 1 × 105, 1 × 103, 1 × 108, and 1 × 104 copies/µL; (J) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 105, 1 × 103, 1 × 104, 1 × 108, and 1 × 106 copies/µL; (K) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 104, 1 × 105, 1 × 102, 1 × 103, and 1 × 101 copies/µL; (L) Coinfection amplification curves of Tg, Nc, Es, Gl, and Te using serial dilutions of DNA standards 1 × 102, 1 × 104, 1 × 101, 1 × 105, and 1 × 103 copies/µL.
Validation of the TaqMan-probe-based pentaplex qPCR assay
A comparison was conducted between the TaqMan-probe-based pentaplex qPCR and the conventional PCR using ten diarrhea samples. Table 6 demonstrated that the developed TaqMan-probe-based pentaplex qPCR results align with those of the conventional PCR, indicating its efficacy in concurrently detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi.
Clinical performance of the TaqMan-probe-based pentaplex qPCR assay
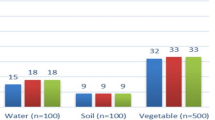
The TaqMan-probe-based pentaplex qPCR developed in this study was used to analyze 210 clinical samples. Figure 6 illustrated that the individual infection rates for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, and Giardia lamblia, as 2.38% (5 out of 210), 2.38% (5 out of 210), 10% (21 out of 210), and 36.19% (76 out of 210), respectively. The coinfection rates were 8.1% (17 out of 210) for Eimeria stiedai & Giardia lamblia. The above results demonstrated that the developed TaqMan-probe-based pentaplex qPCR assay had an excellent ability to detect coinfections.
Discussion
In this study, we developed a novel TaqMan-probe-based pentaplex qPCR assay for simultaneously detecting Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. As far as we know, this is the first report on the TaqMan-probe-based pentaplex qPCR assay for the simultaneous detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi.
Compared with the prior technology51,52,53,54,55, this study has the following characteristics:
-
ⅰ)
According to this study, the G3PDH gene, NC5 gene, ADF gene, GDH gene and COX1 gene are selected as target detection genes for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi, and after multiple gene sequence alignment, the highly conserved regions of G3PDH, NC5, ADF, GDH and COX1 genes were selected to design the primers and probes after eliminating the regions with very low homology and many mutation sites. However, because the purpose of the study is to simultaneously detect the abovementioned five different pathogen targets in the same detection system (that is, five fluorescence channels including FAM, VIC, ABY, JUN, and CY5 are used), if the detection system for each target is not suitable, the Cq value of the detection is too different, and/or there is interference between each primer and probe, the fluorescence inhibition will affect detection rate in the detection process. In view of this, through creative efforts, we further designed and determined the primer and probe sets with similar amplification efficiency for the detection of the nucleic acid of the five pathogens in the short and highly conserved sequences of the five targets. At the same time, the ratio of primers to fluorescent probes in the detection system was adjusted during qPCR amplification. Furthermore, the amplification efficiency of different pathogen nucleic acids was consistent. TaqMan-MGB probe is a unique probe firstly reported in 200056. In this study, we developed a TaqMan-MGB probe to differentiate Trypanosoma evansi from other parasites. The primers and probes have been optimized, with high sensitivity. A positive control is provided to facilitate the differentiation of false-negative samples. The specificity is high. The primers are designed based on the highly conserved region of Trypanosoma evansi and will not have cross-reactions with the DNA of other parasites.
-
ⅱ)
When coinfections of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi occurs, it is time-consuming, laborious and costly to determine the presence of coinfections of multiple pathogens by single qPCR alone. This study provides a method for simultaneous detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi in the same detection system. By targeting the G3PDH gene of Toxoplasma gondii, the NC5 gene of Neospora caninum, the ADF gene of Eimeria stiedai, the GDH gene of Giardia lamblia and the COX1 gene of Trypanosoma evansi, the specific probes labeled with fluorescent emission groups can be used to determine the coinfection of multiple pathogens at one time, which is time-saving and labor-saving. And it can significantly reduce the detection cost. This method provided by our study has high specificity, and there is no nonspecific amplification curve for any other non-target species, which can ensure the accuracy of detection.
-
ⅲ)
This study provides a method for rapid detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. The assay was qualitatively, quantitatively and highly sensitive (the lower limits of detection for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi was 2, 1.5, 1.5, 1.8 and 1.1 copies per reaction, respectively). It has the characteristics of high specificity, excellent repeatability and reproducibility (the coefficient of variation value of intra-assay and inter-assay is less than 3%). Furthermore, this assay has shown eight logs of dynamic range with five different pathogens simultaneously, and has extremely high sensitivity. It is superior to other reports57,58,59.
-
ⅳ)
The clinical performance of the developed TaqMan-probe-based pentaplex qPCR was evaluated in two hundred and ten. clinical samples to verify its practicality and usefulness. The results demonstrate that five, five, twenty-one and seventy-six samples were positive for Toxoplasma gondii, Neospora caninum, Eimeria stiedai, and Giardia lamblia, respectively. Furthermore, coinfections with two kinds of pathogens are also found, which may elevate immunosuppression and inflammation, and thus increase the possibility of secondary infection by other pathogens and further exacerbate these diseases60,61,62,63,64. In this study, among the clinical samples, only seventeen double positive samples of Eimeria stiedai/Giardia lamblia were detected using the TaqMan-probe-based pentaplex qPCR assay and confirmed using the microscopic examination, TaqMan-probe-based single-plex qPCR, conventional PCR, and sequencing methods. This suggests that Eimeria stiedai/Giardia lamblia were still prevalent in China, which were consistent with the results of other reports65.
However, the experiments have some limitations. First, although the primers and probes in this study were designed for highly conserved regions of G3PDH of Toxoplasma gondii, NC5 of Neospora caninum, ADF of Eimeria stiedai, GDH of Giardia lamblia, and COX1 of Trypanosoma evansi, the potential for gene mutations, deletions, and genetic recombination over time leads to off-target primers and probes, which further results to false negative results. Second, the specimen size was relatively small. Validation of the TaqMan-probe-based pentaplex qPCR method in more samples is needed.
In summary, we developed a novel TaqMan-probe-based pentaplex qPCR assay for the simultaneous detection of Toxoplasma gondii, Neospora caninum, Eimeria stiedai, Giardia lamblia and Trypanosoma evansi. The developed TaqMan-probe-based pentaplex qPCR assay has the advantages of high sensitivity and strong specificity, high throughput, convenient, fast, and cost-effectiveness. It is of great significance to safeguard human health, animal quality and public sanitary security. The TaqMan-probe-based pentaplex qPCR assay has great practical significance in controlling the quality of animal derived biological products for human use and in the field of import and export animal inspection and quarantine, and deserves a board application.
Data availability
Study data are available from the corresponding author upon request.
References
Miokovic, A. P., Ratkovic, M. & Gadzo, A. P. Toxoplasmosis in the outer retina. Rom J Ophthalmol. 68 (2),198–201. https://doi.org/10.22336/rjo.2024.37 (2024).
Bourdin, C., Busse, A., Kouamou, E., Touafek, F., Bodaghi, B., Le Hoang, P., Mazier, D., Paris, L., Fekkar, A.. PCR-based detection of Toxoplasma gondii DNA in blood and ocular samples for diagnosis of ocular toxoplasmosis. J. Clin. Microbiol. 52 (11), 3987–91. https://doi.org/10.1128/JCM.01793-14 (2014).
Desmettre, T. Toxoplasmosis and behavioural changes. J. Fr. Ophtalmol. 43 (3), e89–e93. https://doi.org/10.1016/j.jfo.2020.01.001 (2020).
Aral Akarsu, G., Elhan, H. A. & Akarsu, C. Retrospective evaluation of Toxoplasma gondii seropositivity in fertile and infertile women. Mikrobiyol Bul. 45 (1), 174–180 (2011).
Dao, V. T., Anagnostou, A., Schlösser, R., Rochwalsky, U., Groß, U., Hoehl, S., Kempf, V. A. J., Besier, S. First description of congenital toxoplasmosis after maternal coinfection with Toxoplasma gondii and severe acute respiratory syndrome coronavirus 2: A case report. J Med Case Rep. 17 (1), 121. https://doi.org/10.1186/s13256-023-03855-8 (2023).
Wahdini, S., Sari, I. P., Oswari, H. & Kurniawan, A. Unspecific congenital toxoplasmosis in a two-month-old baby. Acta Biomed. 94 (S1), e2023144. https://doi.org/10.23750/abm.v94iS1.14308 (2023).
Modrzejewska, M., Patalan, J., Kulik, U. & Czeszyńska, M. B. Ocular manifestation of congenital toxoplasmosis, clinical implication - case report. Ginekol Pol. 87(3), 226–30. https://doi.org/10.17772/gp/61990 (2016).
Londoño-Martinez, J. C., Velasco-Velasquez, S., Cordero-Lopez, S., Osorio, M. F., Celis-Giraldo, D., Thibodeau, J., Baird, I., McLeod, R., Gomez-Marin, J. Evaluation of the acceptability of point of care diagnostic test for prenatal toxoplasmosis (translational research phase III). J. Infect Public Health 16 (1), 15–24. https://doi.org/10.1016/j.jiph.2022.11.023 (2023).
Ayisi, N. K., Wiredu, E. K., Sata, T., Nyadedzor, C., Tsiagbe, V. K., Newman, M., Cofie, C. N., Taneguchi, K. T-lymphocytopaenia, opportunistic infections and pathological findings in Ghanaian AIDS patients and their sexual partners. East. Afr Med J.74 (12), 784–791 (1997).
Butani, L. & Tancredi, D. Outcomes of kidney transplants from toxoplasma-positive donors: An organ procurement and transplant network database analysis. Transpl Int. 37, 13203. https://doi.org/10.3389/ti.2024.13203 (2024).
Wesołowski, R., Pawłowska, M. & Mila-Kierzenkowska, C. The medical relevance of toxoplasma infections in terms of the safety of blood recipients under immunosuppression-a meta-analysis. Microorganisms. 11(8), 1980. https://doi.org/10.3390/microorganisms11081980 (2023).
Garg, A., McKay, B. R., Francisconi, C. L. M. & Muni, R. H. Toxoplasmosis chorioretinitis mimicking cytomegalovirus retinitis in an immunocompromised pediatric patient following bone marrow transplantation. GMS Ophthalmol Cases. 12, Doc14. https://doi.org/10.3205/oc000201 (2022).
Lindell, R. B., Wolf, M. S., Alcamo, A. M., Silverman, M. A., Dulek, D. E., Otto, W. R., Olson, T. S., Kitko, C. L., Paueksakon, P., Chiotos, K. Case report: Immune dysregulation due to Toxoplasma gondii reactivation after allogeneic hematopoietic cell transplant. Front Pediatr. 9, 719679. https://doi.org/10.3389/fped.2021.719679 (2021).
Adekunle, R. O., Sherman, A., Spicer, J. O., Messina, J. A., Steinbrink, J. M., Sexton, M. E., Lyon, G. M., Mehta, A. K., Phadke, V. K., Woodworth, M. H. Clinical characteristics and outcomes of toxoplasmosis among transplant recipients at two US academic medical centers. Transpl Infect Dis. 23 (4), e13636. https://doi.org/10.1111/tid.13636 (2021).
Quinn, H. E., Ellis, J. T. & Smith, N. C. Neospora caninum: a cause of immune-mediated failure of pregnancy? Trends Parasitol. 18 (9), 391–4. https://doi.org/10.1016/s1471-4922(02)02324-3 (2002).
Sánchez-Sánchez, R., Vázquez-Calvo, Á., Fernández-Escobar, M., Regidor-Cerrillo, J., Benavides, J., Gutiérrez, J., Gutiérrez-Expósito, D., Crespo-Ramos, F. J., Ortega-Mora, L. M., Álvarez-García, G. Dynamics of Neospora caninum-associated abortions in a dairy sheep flock and results of a Test-and-Cull Control Programme. Pathogens. 10 (11), 1518. https://doi.org/10.3390/pathogens10111518 (2021).
Omata, Y., Sueda, M., Koyama, T., Tanabe, S., Uzuka, Y., Sarashina, T., Makino, S., Maeda, R., Saito, A., Mikami, T. Identification and the role of soluble antigens detected in bile from Eimeria stiedai-infected rabbits. J. Parasitol. 87 (2), 287–91. https://doi.org/10.1645/0022-3395 (2001).
Chen, X. D., Xie, J., Wei, Y., Yu, J. F., Cao, Y., Xiao, L., Wu, X. J., Mao, C. J., Kang, R. M., Ye, Y. G. Immune modulation of Th1/Th2/Treg/Th17/Th9/Th21 cells in rabbits infected with Eimeria stiedai. Front Cell Infect Microbiol. 13, 1230689. https://doi.org/10.3389/fcimb.2023.1230689 (2023).
Bashar, S., Das, A., Erdem, S., Hafeez, W. & Ismail, R. Severe gastroenteritis from Giardia lamblia and Salmonella saintpaul co-infection causing acute renal failure. Cureus. 14 (5), e25288. https://doi.org/10.7759/cureus.25288 (2022).
Ogundare, O. T., Olalemi, A. & Triumphant, E. O. Microbial health risks of Cryptosporidium parvum and Giardia lamblia in tropical coastal water in Araromi, Nigeria. Turkiye Parazitol Derg. 48 (2), 82–8. https://doi.org/10.4274/tpd.galenos.2024.69733 (2024).
Rjeibi, M. R., Ben Hamida, T., Dalgatova, Z., Mahjoub, T., Rejeb, A., Dridi, W., Gharbi, M. First report of surra (Trypanosoma evansi infection) in a Tunisian dog. Parasite. 22, 3. https://doi.org/10.1051/parasite/2015004 (2015).
Ereqat, S., Nasereddin, A., Al-Jawabreh, A., Al-Jawabreh, H., Al-Laham, N., Abdeen, Z. Prevalence of Trypanosoma evansi in livestock in Palestine. Parasit Vectors. 13 (1), 21. https://doi.org/10.1186/s13071-020-3894-9 (2020).
Van Vinh Chau, N., Buu Chau, L., Desquesnes, M., Herder, S., Phu Huong Lan, N., Campbell, J. I., Van Cuong, N., Yimming, B., Chalermwong, P., Jittapalapong, S., Ramon, Franco, J., Tri Tue, N., Rabaa,, M. A., Carrique-Mas J., Pham Thi Thanh, T., Tran Vu Thieu, N., Berto, A., Thi Hoa, N., Van Minh Hoang, N., Canh Tu, N., Khac Chuyen, N., Wills, B., Tinh Hien, T., Thwaites, G. E., Yacoub, S., Baker, S. A clinical and epidemiological investigation of the first reported human infection with the zoonotic parasite Trypanosoma evansi in Southeast Asia. Clin. Infect. Dis. 62 (8), 1002–8. https://doi.org/10.1093/cid/ciw052 (2016).
Desquesnes, M., Dargantes, A., Lai, D. H., Lun, Z. R., Holzmuller, P., Jittapalapong S. Trypanosoma evansi and surra: a review and perspectives on transmission, epidemiology and control, impact, and zoonotic aspects. Biomed Res Int. 2013, 321237. https://doi.org/10.1155/2013/321237 (2013).
Kim, J., Álvarez-Rodríguez, A., Li, Z., Radwanska, M. & Magez, S. Recent progress in the detection of surra, a neglected disease caused by Trypanosoma evansi with a One Health impact in large parts of the tropic and sub-tropic world. Microorganisms. 12(1), 44. https://doi.org/10.3390/microorganisms12010044 (2013).
Ndao, M. Diagnosis of parasitic diseases: Old and new approaches. Interdiscip Perspect Infect Dis. 2009, 278246. https://doi.org/10.1155/2009/278246 (2009).
Veletzky, L., Eberhardt, K. A., Hergeth, J., Stelzl, D. R., Zoleko Manego, R., Kreuzmair, R., Burger, G., Mischlinger, J., McCall, M. B. B., Mombo-Ngoma, G., Adegnika, A. A., Agnandji, S. T., Matsiegui, P. B., Lell, B., Kremsner, P., Mordmüller, B., Tappe, D., Ramharter, M.. Analysis of diagnostic test outcomes in a large loiasis cohort from an endemic region: Serological tests are often false negative in hyper-microfilaremic infections. PLoS Negl Trop Dis. 18(3), e0012054. https://doi.org/10.1371/journal.pntd.0012054 (2024).
Mardani-Kataki, M., Beiromvand, M., Teimoori, A., Amari, A. & Tavalla, M. Is immuno-PCR better than ELISA test for detection of Toxoplasma gondii IgG antibody? Acta Parasitol. 67(2), 904–11. https://doi.org/10.1007/s11686-022-00537-1 (2022).
Mardani-Kataki, M., Beiromvand, M., Teimoori, A., Amari, A., Tavalla, M. Risk of reactivated toxoplasmosis in haematopoietic stem cell transplant recipients: a prospective cohort study in a setting withholding prophylaxis. Clin Microbiol Infect. 28(5), 733.e1–733.e5. (2022). https://doi.org/10.1016/j.cmi.2021.09.012 (2022).
Štajner, T., Vujić, D., Srbljanović, J., Bauman, N., Zečević, Ž., Simić, M., Djurković-Djaković, O. Detection of ocular Toxoplasma gondii infection in chronic irregular recurrent uveitis by PCR. Korean J Parasitol. 50(3), 229–31. https://doi.org/10.3347/kjp.2012.50.3.229 (2022).
Jia, L. J., Zhang, S. F., Liu, M. M., Qian, N. C. & Guo, H. P. Isolation, identification, and pathogenicity of Neospora caninum China Yanbian strain. Iran J Parasitol. 9(3), 394–401. https://doi.org/10.3347/kjp.2012.50.3.229 (2014).
Lee, S. E., Hong, S. H., Lee, S. H., Jeong, Y. I., Lim, S. J., Kwon, O. W., Kim, S. H., You, Y. S., Cho, S. H., Lee, W. J. Molecular analysis of single oocyst of Eimeria by whole genome amplification (WGA) based nested PCR. Exp Parasitol. 144, 96–9. https://doi.org/10.1016/j.exppara.2014.06.019 (2014).
Jia, L. J., Zhang, S. F., Liu, M. M., Qian, N. C., Guo, H. PCR detection of Giardia lamblia in stool: targeting intergenic spacer region of multicopy rRNA gene. Mol Cell Probes.14(3), 181–9. https://doi.org/10.1016/j.exppara.2014.06.019 (2014).
Njiru, Z. K., Gitonga, P. K. & Ndungu, K. The typing of Trypanosoma evansi isolates using mobile genetic element (MGE) PCR. Parasitol Res. 108 (6), 1583–7. https://doi.org/10.1007/s00436-010-2246-7 (2011).
Avendaño, C. & Patarroyo, M. A. Loop-mediated isothermal amplification as point-of-care diagnosis for neglected parasitic infections. Int. J. Mol. Sci. 21 (21), 7981. https://doi.org/10.3390/ijms21217981 (2020).
Mousavi, P., Mirhendi, H., Mohebali, M., Shojaee, S., Keshavarz Valian, H., Fallahi, S., Mamishi, S. Detection of Toxoplasma gondii in acute and chronic phases of infection in immunocompromised patients and pregnant women with real-time PCR assay using TaqMan fluorescent probe. Iran J Parasitol. 13 (3), 373–381 (2018).
Bandelj, P., Kušar, D., Šimenc, L., Jamnikar-Ciglenečki, U., Vengušt, G., Vengušt, D. Ž. First molecular detection of Neospora caninum in feces of Grey Wolf (Canis lupus) and Golden Jackal (Canis aureus) populations in Slovenia. Animals (Basel). 13 (19), 3089. https://doi.org/10.3390/ani13193089 (2023).
Marcial-Quino, J., Gómez-Manzo, S., Fierro, F., Vanoye-Carlo, A., Rufino-González, Y., Sierra-Palacios, E., Castillo-Villanueva, A., Castillo-Rodríguez, R. A., Rodríguez-Bustamante, E., Arreguin-Espinosa, R., Reyes-Vivas, H. Stem-loop RT-qPCR as an efficient tool for the detection and quantification of small RNAs in Giardia lamblia. Genes (Basel). 7 (12),131. https://doi.org/10.3390/genes7120131 (2016).
Gao, Z. Q. & Xing, J. Development and application of a TaqMan-MGB probe-based quantitative real-time polymerase chain reaction assay for the rapid detection of Dictyocaulus filaria. Front Vet Sci. 12, 1559088. https://doi.org/10.3389/fvets.2025.1559088 (2025).
Gao, Z. Q., Yue, B. F. & He, Z. M. Development and application of TaqMan MGB probe fluorescence quantitative PCR method for rapid detection of Clostridium piliforme. Zhonghua liu xing bing xue za zhi. 33(2), 226–8. https://doi.org/10.3760/cma.j.issn.0254-6450.2012.02.022 (2012).
Hodzic, E., Glavinic, A. & Wademan, C. A novel approach for simultaneous detection of the most common food-borne pathogens by multiplex qPCR. Biomol Biomed. 23 (4), 640–8. https://doi.org/10.17305/bb.2022.8693 (2023).
Truong, M. & Šlapeta, J. Analytical sensitivity of a multiplex quantitative PCR for Toxoplasma gondii and Neospora caninum. Parasitol Res. 122 (4), 1043–7. https://doi.org/10.1007/s00436-023-07796-5 (2023).
Ramaprasad, A., Mourier, T., Naeem, R., Malas, T. B., Moussa, E., Panigrahi, A., Vermont, S. J., Otto, T. D., Wastling, J., Pain, A. Comprehensive evaluation of Toxoplasma gondii VEG and Neospora caninum LIV genomes with tachyzoite stage transcriptome and proteome defines novel transcript features. PLoS One. 10(4), e0124473. https://doi.org/10.1371/journal.pone.0124473 (2015).
Kangethe, R. T., Winger, E. M., Settypalli, T. B., Datta, S., Wijewardana, V., Lamien, C.. Unger, H., Coetzer, T. H., Cattoli, G., Diallo, A. Low dose gamma irradiation of Trypanosoma evansi parasites identifies molecular changes that occur to repair radiation damage and gene transcripts that may be involved in establishing disease in mice post-irradiation. Front Immunol. 13, 852091. https://doi.org/10.3389/fimmu.2022.852091 (2022).
Cohen, R. N., van der Aa, M. A., Macaraeg, N., Lee, A. P., Szoka, F. C. Jr. Quantification of plasmid DNA copies in the nucleus after lipoplex and polyplex transfection. J. Control Release. 135(2), 166–74. https://doi.org/10.1016/j.jconrel.2008.12.016 (2009).
Bustin, S. A., Benes, V., Garson, J. A., Hellemans, J., Huggett, J., Kubista, M., Mueller, R., Nolan, T., Pfaffl, M. W., Shipley, G. L., Vandesompele, J., Wittwer, C. T. The MIQE guidelines: minimuminformation for publication of quantitative real-time PCR experiments. Clin Chem. 55(4), 611–22. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Forootan, A., Sjöback, R., Björkman, J., Sjögreen, B., Linz, L., Kubista, M. Methods to determine limit of detection and limit of quantification in quantitative real-time PCR (qPCR). Biomol Detect Quantif. 12, 1–6. https://doi.org/10.1016/j.bdq.2017.04.001 (2017).
Ruijter, J. M., Barnewall, R. J., Marsh, I. B., Szentirmay, A. N., Quinn, J. C., van Houdt, R., Gunst, Q. D., van den Hoff, M. J. B. Efficiency correction is required for accurate quantitative PCR analysis and reporting. Clin Chem. 67 (6), 829–42 https://doi.org/10.1093/clinchem/hvab052 (2021).
Bustin, S. A., Ruijter, J. M., van den Hoff, M. J. B., Kubista, M., Pfaffl, M. W., Shipley, G. L., Tran, N., Rödiger, S., Untergasser, A., Mueller, R., Nolan, T., Milavec, M., Burns, M. J., Huggett, J. F., Vandesompele, J., Wittwer, C. T. MIQE 2.0: Revision of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments Guidelines. Clin Chem. MIQE 2.0: Revision of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments Guidelines. Clin Chem. 71, 634–51 https://doi.org/10.1093/clinchem/hvaf043 (2025).
Rutledge, R. G. & Cote, C. Mathematics of quantitative kinetic PCR and the application of standard curves. Nucleic Acids Res. 31 (16), e93. https://doi.org/10.1093/nar/gng093 (2003).
Li, Y., Zhu, Z. & Wang, J. Toxoplasmic encephalitis of basal ganglia with tumor-like features proven by pathogen-specific polymerase chain reaction and direct DNA sequencing. Neuropathology. 39 (5), 398–403. https://doi.org/10.1111/neup.12588 (2019).
Yi, X. L., Yang, W. H., Zheng, H. L., Cao, M. L., Xiong, J., Chen, W. C., Zhou, Y. J., Li F., Zhu, X. Q., Liu, G. H. Seroprevalence and molecular detection of Toxoplasma gondii and Neospora caninum in beef cattle and goats in Hunan province, China. Parasit Vectors. 17(1), 195. https://doi.org/10.1186/s13071-024-06283-9 (2024).
Mishra, V., Mitra, P., Barbuddhe, S., Thorat, Y., Chavan, K., Shinde, S., Chaudhari, S., Khan, W., Deshmukh, A. S.Serological and molecular detection of Toxoplasma gondii and Neospora caninum in free-ranging rats from Nagpur, India. Parasitol Res. 123(1), 63. https://doi.org/10.1007/s00436-023-08095-9 (2023).
Rayani, M., Hatam, G., Unyah, N. Z., Ashrafmansori, A., Abdullah, W. O., Hamat, R. A. Phylogenetic analysis of Giardia lamblia human genotypes in fars province, Southern Iran. Iran. J. Parasitol. 12 (4), 522–533 (2017).
Sánchez, E., Perrone, T., Recchimuzzi, G., Cardozo, I., Biteau, N., Aso, P. M., Mijares, A., Baltz T., Berthier,, D., Balzano-Nogueira L., Gonzatti, M. I. Molecular characterization and classification of Trypanosoma spp. Venezuelan isolates based on microsatellite markers and kinetoplast maxicircle genes. Parasit Vectors. 8, 536. https://doi.org/10.1186/s13071-015-1129-2 (2015).
Kutyavin, I., Afonina, I. A., Mills, A. & Hedgpeth, J. 3’-minor groove binder-DNA probes increase sequence specificity at PCR extension temperatures. Nucleic Acids Research. 28 (2), 655–61. https://doi.org/10.1093/nar/28.2.655 (2000).
Cao, Z., Zhang, K., Yin, D., Zhang, Q., Yu, Y., Wen, J., Ni, H. Clinical validation of visual LAMP and qLAMP assays for the rapid detection of Toxoplasma gondii. Front Cell Infect Microbiol. 12, 1024690. https://doi.org/10.3389/fcimb.2022.1024690 (2022).
Nabet, C., Brossas, J. Y., Poignon, C., Bouzidi, A., Paris, L., Touafek, F., Varlet-Marie, E., Sterkers, Y., Passebosc-Faure, K., Dardé, M. L., Piarroux, R., Denis, J. A. Assessment of droplet digital PCR for the detection and absolute quantification of Toxoplasma gondii: a comparative retrospective study. J. Mol. Diagn. 25 (7), 467–76. https://doi.org/10.1016/j.jmoldx.2023.03.006 (2023).
Shin, J. H., Lee, S. E., Kim, T. S., Ma, D. W., Chai, J. Y., Shin, E. H. Multiplex-touchdown PCR to simultaneously detect Cryptosporidium parvum, Giardia lamblia, and Cyclospora cayetanensis, the major causes of traveler’s diarrhea. Korean J. Parasitol. 54 (5), 631–6. https://doi.org/10.3347/kjp.2016.54.5.631 (2016).
Basso, W., Holenweger, F., Schares, G., Müller, N., Campero, L. M., Ardüser, F., Moore-Jones, G., Frey, C. F., Zanolari, P. Toxoplasma gondii and Neospora caninum infections in sheep and goats in Switzerland: seroprevalence and occurrence in aborted foetuses. Food Waterborne Parasitol. 28, e00176. https://doi.org/10.1016/j.fawpar.2022.e00176 (2022).
Nazari, N. et al. Serological survey of Neospora Caninum and Toxoplasma gondii co-infection in rodents in Northwestern Iran. Iran J Parasitol. 15 (2), 253–258 (2020).
Sun, L. X., Liang, Q. L., Nie, L. B., Hu, X. H., Li, Z., Yang, J. F., Zou, F. C., Zhu, X. Q. Serological evidence of Toxoplasma gondii and Neospora caninum infection in black-boned sheep and goats in southwest China. Parasitol Int. 75, 102041. https://doi.org/10.1016/j.parint.2019.102041 (2020).
Innes, E. A., Lundén, A., Esteban, I., Marks, J., Maley, S., Wright, S., Rae, A., Harkins, D., Vermeulen, A., McKendrick, I. J., Buxton, D. A previous infection with Toxoplasma gondii does not protect against a challenge with Neospora Caninum in pregnant sheep. Parasite Immunol. 23 (3), 121–132 (2001).
Hughes, J. M., Williams, R. H., Morley, E. K., Cook, D. A., Terry, R. S., Murphy, R. G., Smith, J. E., Hide, G. The prevalence of Neospora caninum and co-infection with Toxoplasma gondii by PCR analysis in naturally occurring mammal populations. Parasitology. 132 (Pt 1), 29–36. https://doi.org/10.1017/S0031182005008784 (2006).
Lee, M. R., Shin, H. E., Chung, B. S., Lee, S. E., Ju, J. W., Xu, L., Nan, C. L., Park, M. Y., Cho, S. H. Intestinal parasite infections among inhabitants in Yanbian Prefecture, Jilin Province, China. Korean J Parasitol. 55 (5), 579–82. https://doi.org/10.3347/kjp.2017.55.5.579 (2017).
Acknowledgements
Thank editor and reviewers for their helpful comments on this paper.
Funding
The author(s) declare that financial support was received for the research and/or publication of this article. This work was supported by the National Key Research and Development Program of China (Project No. 2022YFF0711003).
Author information
Authors and Affiliations
Contributions
Z-qG: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing – original draft, Writing – review & editing. RF: Funding acquisition, Investigation, Project administration, Resources, Supervision, Validation, Writing – review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gao, Zq., Fu, R. A novel TaqMan probe-based pentaplex qPCR assay for the simultaneous detection of five pathogenic protozoans. Sci Rep 15, 45702 (2025). https://doi.org/10.1038/s41598-025-28046-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28046-x