Abstract
Primary cilia function as critical sensory organelles that mediate multiple signaling pathways, including the Hedgehog (Hh) pathway, which is essential for organ patterning and morphogenesis. Disruptions in Hh signaling have been implicated in supernumerary tooth formation and molar fusion in mutant mice. Cilk1, a highly conserved serine/threonine-protein kinase localized within primary cilia, plays a critical role in ciliary transport. Loss of Cilk1 results in severe ciliopathy phenotypes, including polydactyly, edema, and cleft palate. However, the role of Cilk1 in tooth development remains unexplored. In this study, we investigated the role of Cilk1 in tooth development. Cilk1 was found to be expressed in both the epithelial and mesenchymal compartments of developing molars. Cilk1 deficiency resulted in altered ciliary dynamics, characterized by reduced frequency and increased length, accompanied by downregulation of Hh target genes, such as Ptch1 and Sostdc1, leading to the formation of diastemal supernumerary teeth. Furthermore, in Cilk1−/−;PCS1–MRCS1△/△ mice, which exhibit a compounded suppression of Hh signaling, we uncovered a novel phenomenon: diastemal supernumerary teeth can be larger than first molars. Based on these findings, we propose a progressive model linking Hh signaling levels to sequential changes in tooth patterning: initially inducing diastemal supernumerary teeth, then enlarging them, and ultimately leading to molar fusion. This study reveals a previously unrecognized role of Cilk1 in controlling tooth morphology via Hh signaling and highlights how Hh signaling levels shape tooth patterning in a gradient-dependent manner.
Similar content being viewed by others

Introduction
The primary cilium is a microtubule-based organelle that extends from the surface of most mammalian cells. They act as sensory structures that transduce extracellular signals into intracellular responses, influencing fundamental cellular functions during development and homeostasis.1,2 Primary cilia mediate multiple signaling pathways, including Hedgehog (Hh),3,4 Wnt,5 Fibroblast growth factor,6 Hippo,7 mTOR,8,9 platelet-derived growth factor,10 and Notch11 pathways. Of these, Hh signaling is most intimately linked to ciliary function and is essential for the patterning of various organs.3 Aberrant Hh signaling can lead to supernumerary teeth and molar fusion.12,13,14
Supernumerary teeth in mice are defined as additional teeth that develop beyond the typical complement of one incisor and three molars per quadrant. Based on their location and developmental origin, supernumerary teeth are simply categorized into three types: supernumerary incisors, supernumerary teeth in the toothless diastema region, and supernumerary molars located lingual or distal to the normal molars.15 During wild-type molar development, two transient tooth germs, referred to as the anterior diastemal rudiment (R1) and the posterior diastemal rudiment (R2), appear in the diastema region between the incisor and molars. R1 emerges around embryonic day 12.5 (E12.5) and regresses shortly thereafter, while R2 appears at E13.5 and typically regresses by E14.5 through incorporation into the developing first molar (M1), which initiates just distal to R2.14,16,17,18,19 This integration illustrates that M1 originates from the fusion of multiple tooth primordia, namely the R2 and M1. The second (M2) and third molars (M3) subsequently develop in a posterior sequence. However, in certain mutant mice, the R2 bud fails to regress and instead develops into a supernumerary tooth, which precedes the normal formation of M1 and M2. Thus, the diastemal supernumerary tooth corresponds to R2. Many mutant mouse models with impaired Hedgehog signaling have been reported to develop R2, including Gas1−/−, Shh⁺/⁻; Sostdc1⁺/⁻, Sostdc1−/−, and mice with deletions of Shh enhancers such as MRCS1△/△ and PCS1–MRCS1△/△ mice.12,14,20,21 PCS1–MRCS1△/△ mice, lacking PCS1 and MRCS1, two oral-specific cis-regulatory enhancer elements of Shh (Sonic Hedgehog), exhibit R2 in both the maxilla and mandible.12
Molar fusion in mice refers to a developmental anomaly in which two or more adjacent molar tooth germs, most commonly M1 and M2, fail to undergo proper segmentation and instead develop as a single fused molar. Other Hh signaling mutants, such as Krt14-Cre;Shhfl/−, Krt14-Cre;Smofl/fl, and Sostdc1−/− mice, also develop molar fusion.13,14,22,23,24,25,26
Given the role of primary cilia in Hh signaling, we investigated Cilk1 (Ciliogenesis-associated kinase 1) to understand its contribution to tooth patterning. Cilk1 is a key regulator of ciliary turnaround processes. It localizes at ciliary tips via interactions with the IFT-B complex.27 Cilk1 regulates the undocking of kinesin-II from intraflagellar transport (IFT) particles and coordinates the turnaround of the IFT machinery by phosphorylating kinesin-II subunits or other components of the IFT complex.28,29,30,31,32,33 This regulatory function is supported by the abnormal accumulation of IFT components at the ciliary tip in Cilk1-deficient cells.34 Mutations in Cilk1 result in elongated cilia and impair ciliary function, leading to defects in Hh signaling and are associated with ciliopathies, a group of syndromes that arise from impairments in sensory and signaling activities of the primary cilia and affect nearly all organs in the body.34,35,36 Cilk1 is critical for the proper retrograde trafficking and ciliary exit of Smoothened (Smo), a ciliary G protein–coupled receptor that mediates Hh signaling. In wild-type cells, stimulation with the Smoothened agonist (SAG) facilitates Smo entry into the primary cilium, resulting in downstream activation of the Hh pathway. By contrast, Cilk1-deficient cells exhibit excessive accumulation of Smo within the primary cilium even under unstimulated conditions, without corresponding activation of Hedgehog signaling.34 While loss-of-function mutations in primary cilia-related genes such as Ift88, Ofd1, and Kif3a cause severe developmental defects, including cleft palate and abnormal tooth patterning,37,38,39 the specific impact of Cilk1 on tooth development remains unclear.
In this study, we examined Cilk1 expression in developing tooth germs and analyzed tooth patterning in both conventional and conditional Cilk1-deficient mice. Our findings reveal that loss of Cilk1 results in R2 formation, altered ciliary morphology, and reduced Hh signaling activity. Furthermore, we generated Cilk1−/−; PCS1–MRCS1△/△ mice, which exhibited R2 enlargement and molar fusion, reinforcing the link between Hh signaling activity and tooth patterning.
Results
Cilk1 deletion leads to diastemal supernumerary tooth formation
To investigate the spatiotemporal expression of Cilk1 during tooth development, we performed RNA in situ hybridization. Cilk1 expression was detected in both the epithelium and mesenchyme of the mandibular M1 from E13.5 to E16.5 (Fig. 1a–d). To further confirm this pattern, we analyzed recently published single-cell RNA-sequencing (scRNA-seq) data,40,41 which demonstrated that Cilk1 was broadly expressed across both epithelial and mesenchymal compartments at E14.5 and E16.5 (Supplementary Fig. S1b, j). Immunohistochemistry staining also showed the presence of primary cilia in most cells within both dental epithelium and mesenchyme (Fig. 3e, g), confirming that Cilk1 is closely associated with primary cilia during tooth development, as in other organs.
Cilk1 expression in developing molars and developmental anomalies in Cilk1−/− mice. a–d Cilk1 expression (small brown dots) in frontal sections of the developing first molar (M1) of wild-type mice at E13, E14, E15.5, and E16.5. Cilk1 is weakly expressed in dental epithelium and dental mesenchyme. e–l Tooth germs dissected from maxilla (Mx) and mandible (Mn) of wild-type (WT) and Cilk1−/− mice at E14.5 and E16.5. Diastemal supernumerary tooth (R2) is observed in Cilk1−/− mice at E14.5 (f, h). M1 and second molar (M2) are observed in wild-type mice at E16.5 (i, k), while R2 and M1 are observed in Cilk1−/− mice (j, l). R2 is similar or smaller in size than M1 in both maxilla and mandible of Cilk1−/− mice at E16.5. m Wild-type mice show the fused palate at E18.5. n Cilk1−/− mice show cleft palate (asterisk) at E18.5 with two palatal shelves (PS). o–t Cilk1−/− embryo shows multiple lobules (black arrowheads) in the tongue at E12.5. Scale bars, a–d: 50 μm, e–l: 0.25 mm, m–t: 1 mm
In Cilk1−/− mice, R2 formation was observed at E14.5 and E16.5. By E14.5, both wild-type and Cilk1−/− mice had a single molar. However, the Cilk1−/− molar was significantly smaller (Fig. 1e–h). At E16.5, two molars were visible in wild-type mice, whereas Cilk1−/− mice exhibited an abnormally smaller mesial molar, indicative of R2 formation (Fig. 1i–l). This aberrant tooth patterning was observed in 100% of Cilk1−/− mice in both the maxilla and mandible. In frontal sections of wild-type mice, R2 regressed, and M1 consistently reached more advanced developmental stages than M2 at corresponding time points. In contrast, in Cilk1−/− mice, R2 survived and progressed earlier than M1 (Supplementary Fig. S2a–f”).
Additional craniofacial abnormalities, including cleft palate and lobulated tongue, were detected in Cilk1−/− mice (Fig. 1m–t), consistent with previous reports.42 These defects were observed in 100% of Cilk1−/− mice, reinforcing the role of Cilk1 in craniofacial and tooth development.
Due to perinatal lethality, the final tooth pattern was assessed at postnatal day 0 (PN 0) (Fig. 2a–d). At this stage, R2 was retained in both the maxilla and mandible of Cilk1−/− mice, as confirmed by micro-CT images. Sagittal and occlusal sections revealed that R2 was smaller than M1, indicating an altered developmental trajectory. In frontal sections of molars at E15.5 and PN 0, the dental epithelium of R2 in Cilk1−/− mice differed from M1 in wild-type mice (Fig. 3a–d, Supplementary Fig. S2g−j). Notably, a thickened dental lamina was observed in mandibular R2 (Supplementary Fig. S2j). Ameloblasts and odontoblasts were well observed in both Cilk1−/− and wild-type mice (Supplementary Fig. S2k, l).
Diastemal supernumerary teeth in Cilk1−/− mice. a–d Micro-computed tomography (micro-CT) sections, histologic images, and 3-dimensional reconstructed tooth germs at PN 0. First molar (M1) is larger than second molar (M2) in wild-type mice in both maxilla (a–a”’) and mandible (b–b”’). A smaller-sized diastemal supernumerary tooth (R2) is observed on the mesial side of M1 in Cilk1−/− mice in both maxilla (c–c”’) and mandible (d–d”’). Scale bars, a–d, a’–d’: 0.25 mm
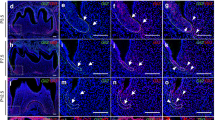
Alterations in primary cilia and gene expression in developing molars in Cilk1−/− mice. a–d Frontal histologic sections of first molar (M1) of wild-type (WT) and diastemal supernumerary tooth (R2) of Cilk1−/− mice at E15.5. The dental epithelium of Cilk1−/− mice is different in shape from that of wild-type mice. e–h Immunohistochemistry of Arl13b in tooth germs at the cap stage. Arl13b highlights the cilia (white arrows) in the dental epithelium (Epi) and mesenchyme (Mes) of wild-type and Cilk1−/− tooth germs at E15.5. DAPI was used for counterstaining. i, j Quantification of ciliary length and ciliated cell percentages in the dental epithelium and mesenchyme of wild-type M1 and Cilk1−/− R2. Primary cilia are significantly elongated in both the dental epithelium and mesenchyme of R2 (i). Percentage of ciliated cells in dental epithelium and mesenchyme is significantly decreased in R2 of Cilk1−/− mice (j). Scale bars: a–d: 500 μm, e–h: 20 μm. **P < 0.01
Cilk1 loss increases primary cilia length and reduces Hh signaling activity
The length of primary cilia and Hh signaling activity were significantly altered in the developing tooth germ of Cilk1−/− mice. To determine whether the observed changes in tooth pattern were associated with numerical and morphological alterations in primary cilia, we quantified cilia frequency and length in frontal sections of developing molars at E15.5.
Immunohistochemistry staining using Arl13b, a specific ciliary marker, confirmed the presence of primary cilia in the majority of cells within both dental epithelium and mesenchyme (Fig. 3g), consistent with previous findings.43 In Cilk1−/− mice, primary cilia were significantly elongated, whereas their overall frequency was markedly reduced in both epithelial and mesenchymal cells (Fig. 3i, j). These findings indicate that Cilk1 plays a critical role in regulating both ciliary length and frequency during tooth development.
Mesenchymal Cilk1 is essential for tooth patterning
To determine whether primary cilia in the epithelium or mesenchyme are critical for tooth patterning, we generated tissue-specific Cilk1−/− mice (Fig. 4).
Loss of Cilk1 in mesenchyme leads to changes in tooth pattern. a–j Sagittal sections of the maxilla (Mx) and mandible (Mn)of Krt14-Cre;Cilk1fl/fl and Wnt1-Cre;Cilk1fl/fl mice at PN 0. Krt14-Cre;Cilk1fl/fl shows a normal tooth pattern with first molar (M1) and second molar (M2). Wnt1-Cre;Cilk1fl/fl shows smaller diastemal supernumerary tooth (R2), M1 and M2. i, j R2 is clearly observed in Wnt1-Cre;Cilk1fl/fl mice by the immunofluorescent staining against Krt14. DAPI was used for counterstaining. At E16.5, X-gal–positive cells were absent in R26R control mice (k), present in the oral and dental epithelium of Krt14-Cre;R26R mice (l) and observed in the oral and dental mesenchyme of Wnt1-Cre;R26R mice (m) confirming tissue-specific Cre recombinase activity. Scale bars: a–h: 250 μm, i, j: 500 μm, k–m: 200 μm
First, Krt14-Cre;Cilk1fl/fl mice, which lack Cilk1 in the epithelium, exhibited no apparent phenotypic changes in either gross morphology or tooth patterning at PN 0 (Fig. 4a, c, e, g, i). In contrast, Wnt1-Cre;Cilk1fl/fl mice, in which Cilk1 is deleted in cranial neural crest-derived mesenchyme, developed R2 in both the maxilla and mandible (Fig. 4b, d, f, h, j). Interestingly, M2 development was observed in Wnt1-Cre;Cilk1fl/fl mice at PN 0; however, M3, which develops from PN 4 in wild-type mice, could not be assessed due to perinatal lethality. Immunostaining with anti-Krt14 antibody further confirmed the presence of R2 in Wnt1-Cre;Cilk1fl/fl mice (Fig. 4j).
The tissue-specific gene knockout efficiency of Krt14-Cre and Wnt1-Cre transgenic mice has been well characterized in previous studies. The Wnt1-Cre line has been shown to activate Cre recombinase in approximately 96% of neural crest cell lineages, including both premigratory and migratory populations.44 Similarly, the Krt14-Cre line has been reported to mediate gene deletion in nearly all keratinocytes of the skin and oral epithelium.45,46 To confirm tissue-specific Cre recombinase activity in this study, X-gal staining was performed at E16.5 in Krt14-Cre;R26R and Wnt1-Cre;R26R embryos (Fig. 4k–m). In Krt14-Cre;R26R mice, X-gal–positive cells were detected in the oral and dental epithelium, whereas in Wnt1-Cre;R26R mice, X-gal–positive cells were observed in the oral and dental mesenchyme. These patterns confirm epithelial- and mesenchymal-specific Cre activity in the respective transgenic mouse lines. PCR analysis was also performed using epithelial tissues isolated from the tongue and palate of Krt14-Cre;Cilk1fl/fl mice at PN 3, confirming that exon 6 of Cilk1 was efficiently and completely deleted in the oral epithelium by Krt14-Cre–mediated recombination (Supplementary Fig. S3a–c). Additionally, while Krt14-Cre;Cilk1fl/fl mice were viable beyond birth, Wnt1-Cre;Cilk1fl/fl mice exhibited cleft palate, lobulated tongue, and perinatal lethality (Fig. 6d). These findings strongly indicate that mesenchymal primary cilia play a critical role in tooth patterning.
Cilk1 −/− tooth germs form calcified teeth
The Cilk1−/− mice exhibit perinatal lethality, preventing the assessment of the final tooth pattern and dental hard tissue formation. To investigate the role of Cilk1 in the later stages of tooth development, mandibular molars at the cap stage were isolated from both wild-type and Cilk1−/− embryos at E14.5 and transplanted under the renal capsule of adult wild-type mice. After 4 weeks, fully calcified teeth developed from the transplanted tooth germs of both wild-type and Cilk1−/− mice (Fig. 5a–j).
Cilk1-deficient tooth germs form fully calcified teeth. a, b Fully calcified teeth formed from mandibular E14.5 developing molars of wild-type (WT) and Cilk1−/− mice within the renal subcapsular layer over a 4-week period. c, d Micro-CT reconstructions showing distinct differences in tooth pattern between wild-type and Cilk1−/− mice. In wild-type samples, tooth size progressively decreases from left to right, whereas in Cilk1−/− calcified teeth, the central tooth is the largest. Wild-type samples show the first, second, and third molars (M1, M2, M3) from left to right, whereas Cilk1−/− samples show the diastemal supernumerary tooth (R2), M1, and M2. e, f Comparative images showing that root (asterisks) length appears shorter in Cilk1−/− calcified teeth than in wild-type teeth. g, h Micro-CT sections demonstrating well-formed enamel (white arrows), dentin (white arrowheads), and surrounding bone (yellow arrowheads) in both wild-type and Cilk1−/− teeth. i, j Histological analysis confirming the presence of enamel space (ES), dentin (D), dentinal tubules, predentin (PD), and odontoblasts (OB) in both genotypes. k–p Representative images of Krt14-Cre;Cilk1fl/fl mice at PN 8 and PN 4 weeks, showing organic enamel (black arrows) and secretory ameloblasts (black arrowheads) at PN 8 and fully mineralized enamel (yellow arrows) at PN 4 weeks, similar to Krt14-Cre;Cilk1fl/+ mice. q Crown volume measurements of calcified teeth indicate that the central tooth in Cilk1−/− teeth is the largest, supporting the identification of the left tooth as R2. r The relative crown volume of R2 to M1 (R2/M1 ratio) in Cilk1−/− teeth is significantly smaller than the M1 to M2 volume ratio (M1/M2) in wild-type teeth. s Root length is shorter in Cilk1−/− teeth compared to wild-type controls. Scale bars: a–h: 250 μm, i, j: 50 μm, k, l: 300 μm, m, n: 200 μm. **P < 0.01
First, the final tooth pattern differed between wild-type and Cilk1−/− calcified teeth. In wild-type samples, tooth size progressively decreased from left to right (Fig. 5a, c), whereas in Cilk1−/− calcified teeth, the central tooth was the largest (Fig. 5b, d). Crown volume measurements further confirmed that, in all Cilk1−/− samples, the left tooth was consistently smaller than the central tooth (Fig. 5q), supporting the identification of the left tooth as R2. Moreover, the relative crown volume of R2 to M1 (R2/M1 ratio) in Cilk1−/− teeth was significantly smaller than the M1 to M2 volume ratio (M1/M2) observed in wild-type samples (Fig. 5r). This Cilk1−/− tooth pattern is consistent with tooth patterns detected in Cilk1−/− embryos at E16.5 and PN 0 (Figs. 1j, l and 2c, d) and mirrors the tooth pattern with R2 observed in many mutant mice.12,47,48 These findings indicate that R2 in Cilk1−/− embryos was retained and successfully developed into calcified teeth.
Second, dental hard tissue formation was unaffected by Cilk1 deletion, though root length was shorter in Cilk1−/− teeth compared to wild-type (Fig. 5f, h, s). Micro-CT analyses revealed well-formed enamel, dentin, and surrounding bone in all teeth derived from both wild-type and Cilk1−/− tooth germs (Fig. 5g, h). Histological examination further confirmed the presence of dentinal tubules, pre-dentin, and odontoblasts in both wild-type and Cilk1−/− teeth (Fig. 5i, j). These results suggest that Cilk1 is not required for the formation of enamel and dentin, implying that ameloblasts and odontoblasts can differentiate and function in the absence of Cilk1. Consistent with these results, Krt14-Cre;Cilk1fl/fl mice exhibited ameloblasts, odontoblasts, enamel, and dentin at PN 8 and mineralized enamel and dentin by PN 4 weeks, mirroring the normal mineralization process observed in wild-type mice (Fig. 5k–p). These findings indicate that despite Cilk1 deletion in epithelial cells, the differentiation and functional maturation of ameloblasts proceed normally, highlighting the non-essential role of Cilk1 in enamel formation.
Tooth patterning is dependent on Hh pathway activity
Previous studies have reported that compound mutant mice, such as Shh+/−;MRCS1△/△, Shh+/−;MFCS1△/△, and Shh+/−;MFCS4△/△, generated by crossing Shh+/− mice with various Shh enhancer-deletion mouse lines, exhibit a more pronounced reduction in Hedgehog signaling activity.21,49 In this study, to further investigate the impact of Hh signaling activity on tooth patterning, PCS1–MRCS1△/△ mice were crossed with Cilk1−/− mice to generate Cilk1−/−;PCS1–MRCS1△/△ mice. In Cilk1−/− mice, Hh signaling was impaired in Shh-responsive cells, whereas PCS1–MRCS1△/△ mice exhibited reduced Shh expression within Shh-producing cells due to the deletion of oral-specific Shh enhancers.12 In Cilk1−/−;PCS1–MRCS1△/△ mice, Hh signaling is likely simultaneously compromised in both Shh-responsive and Shh-producing cells (Supplementary Fig. S5).
Similar to Cilk1−/− mice, Cilk1−/−;PCS1–MRCS1△/△ mice exhibited perinatal lethality. Additionally, cleft palate and lobulated tongue were observed in Cilk1−/−;PCS1–MRCS1△/△ mice, resembling the phenotypes seen in Wnt1-Cre;Cilk1fl/fl and Cilk1−/− mice (Fig. 6a–f).
Enlargement of R2 and molar fusion in compound mutant mice. a–f Frontal sections of the head at eye level in various genotypes at PN 0, showing cleft palate (asterisks) in Wnt1-Cre;Cilk1fl/fl, Cilk1–/– and Cilk1–/–;PCS1–MRCS1△/△ mice. g–r Micro-CT and histologic images in PCS1–MRCS1△/△ and Cilk1–/–;PCS1–MRCS1△/△ mice at PN 0. Diastemal supernumerary teeth (R2) are detected in both the maxilla (Mx) and mandible (Mn) of PCS1–MRCS1△/△. m–r Cilk1–/–;PCS1–MRCS1△/△ mice exhibits fused molar in maxilla and R2 and first molar (M1) in mandible
R2 was also observed in both the maxilla and mandible of Cilk1−/−;PCS1–MRCS1△/△ mice at PN 0 (Fig. 6g–r). Moreover, molar fusion was detected in the maxilla but not in the mandible of Cilk1−/−;PCS1–MRCS1△/△ mice (Fig. 6m–r).
To further investigate the molecular basis of these alterations, RNA-sequencing was performed on maxillary molars at the cap stage, which are crucial stages for tooth pattern formation. Differential gene expression analysis based on RNA-sequencing revealed 85 significantly downregulated genes in Cilk1−/− compared to wild type and 105 in Cilk1−/−;PCS1–MRCS1△/△ mice. Among these, 43 genes were commonly downregulated in both mutant mice (Fig. 7a, Supplementary Table S2). On the other hand, 41 genes were commonly upregulated in both mutant mice compared to wild-type mice (Fig. 7a, Supplementary Table S1). Functional enrichment analysis of the commonly downregulated genes, compared with wild type, identified significant enrichment of the “Hedgehog signaling pathway” in KEGG pathway analysis and “Hedgehog family protein binding” in Gene Ontology molecular functions (Fig. 7b). Ptch1, Gli1, Sostdc1, Hhip, and Foxf2, which are known targets of the Hedgehog (Hh) signaling pathway, were among the 43 commonly downregulated genes (Supplementary Fig. S4, Supplementary Table S2). Differential gene expression analysis at E14.5 revealed a progressive reduction in their expression levels from Cilk1−/− mice to Cilk1−/−;PCS1–MRCS1△/△ mice, compared to wild-type controls (Fig. 7c, Supplementary Table S2). This marked reduction in mRNA expression of key Hh target genes was further validated by qPCR (Fig. 7d). These results suggest that Cilk1 is essential for modulating Hh signaling activity during tooth development and that its absence leads to a downregulation of Hh pathway activity, contributing to the observed alterations in tooth patterning. Additionally, RNA-sequencing analysis also confirmed the deletion of Cilk1 exon 6 in Cilk1−/− mice and Cilk1−/−;PCS1–MRCS1△/△ mice (Supplementary Fig. S6).
Progressive Hedgehog signaling reduction in mutant mice. a Venn diagrams showing the number of differentially expressed genes in Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice compared to wild-type (WT) controls at E14.5. Among the significantly downregulated genes, 43 are commonly shared between the two mutant groups. A total of 41 genes are commonly upregulated in both Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice compared to WT. b Functional enrichment analysis of the 43 commonly downregulated genes showing the significant enrichment of the “Hedgehog signaling pathway” in KEGG and “Hedgehog family protein binding” in Gene Ontology (GO) molecular functions. c Heatmaps display the normalized count levels of genes in the Cilk1−/−, Cilk1−/−;PCS1–MRCS1△/△ and wild-type maxillary tooth germs from RNA-sequencing. Cilk1 and Hedgehog targets genes, such as Gli1, Ptch1, Hhip, Foxf2 and Sostdc1, are progressively downregulated from WT to Cilk1−/− and further in Cilk1−/−;PCS1–MRCS1△/△ mice. Fold changes are calculated relative to wild-type controls. d Quantitative PCR analysis showing reduced expression of Hedgehog pathway target genes in Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice compared to WT mice at E14.5. **P < 0.01, ***P < 0.001
To examine the expression patterns of genes involved in the Wnt-Shh-Sostdc1 negative feedback loop, which is known to regulate tooth patterning,13 we performed RNA in situ hybridization to detect expression changes in Lef1, Ptch1, and Sostdc1 in sagittal sections of Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice at E15.5 (Fig. 8a–l). Ptch1 and Sostdc1, key Hh pathway targets, were weakly expressed in the dental epithelium and completely absent in the dental mesenchyme of R2 in Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice, compared to wild-type mice (Fig. 8d–i). Conversely, Lef1 expression remained similar among wild-type, Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ tooth germs (Fig. 8j–l). Collectively, these findings suggest that the downregulation of Hh signaling activity is a key driver of R2 formation in Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice.
Altered spatial gene expression patterns in tooth germs of Cilk1 mutant mice. a–c The sagittal histologic sections of wild-type, Cilk1−/−, and Cilk1−/−;PCS1–MRCS1△/△ mice. R2 in Cilk1−/−, and Cilk1−/−;PCS1–MRCS1△/△ is smaller than first molar (M1) in wild-type mice at E15.5. d–l Ptch1 and Sostdc1 expression were observed in the dental epithelium and mesenchyme of the first molar in wild-type mice, but weakened and absent in the dental epithelium (white asterisks) and mesenchyme (black asterisks), respectively, of R2 in Cilk1−/− and Cilk1−/−;PCS1–MRCS1△/△ mice. Lef1 is expressed in all tooth germs and shows non-significant changes between wild-type, Cilk1−/−, and Cilk1−/−;PCS1–MRCS1△/△ mice. Black dotted lines indicate the boundary between epithelium and mesenchyme. Yellow dotted lines indicate the region between M1 and M2
Quantitative analysis of R2 formation revealed a 41.6% penetrance in the maxillary quadrants and 34.6% in the mandibular quadrants of PCS1–MRCS1△/△ mice. In contrast, Wnt1-Cre;Cilk1fl/fl mice exhibited an 83.3% penetrance in the maxillary quadrants and 50% in the mandibular quadrants, while Cilk1−/− mice consistently showed 100% penetrance in both the maxilla and mandibular quadrants. In Cilk1−/−;PCS1–MRCS1△/△ mice, R2 formation and molar fusion were observed in 54.5% and 45.5% of maxillary quadrants, respectively, while the mandibular phenotype exclusively consisted of R2 formation (100% penetrance) (Fig. 9a, b). These findings indicate that as Hh pathway activity decreases, the frequency of R2 formation and molar fusion increases.
Tooth pattern changes along a gradient of Hedgehog signaling activity. a, b Quantification of each phenotype penetrance in various genotypes. Cilk1–/– mice display 100% penetrance of R2 in both maxilla and mandible. In Cilk1–/–;PCS1–MRCS1△/△ mice, R2 formation and molar fusion are observed in 54.5% and 45.5% of maxilla, respectively, while the mandible exclusively exhibits R2 formation in all cases. While Krt14-Cre;Cilk1fl/fl exhibited a WT-like phenotype with no changes in tooth patterning, Wnt1-Cre;Cilk1fl/fl, PCS1–MRCS1△/△, Cilk1–/–, and Cilk1–/–;PCS1–MRCS1△/△ all displayed patterns of R2 formation. However, a molar fusion phenotype was observed only in the maxilla of Cilk1–/–;PCS1–MRCS1△/△. c, d Graph showing the relative size of M1 to second molar (M1/M2 ratio) in wild-type and Krt14-Cre;Cilk1fl/fl mice and the relative size of R2 to M1 (R2/M1 ratio) in Wnt1-Cre;Cilk1fl/fl, Cilk1–/–, and Cilk1–/–;PCS1–MRCS1△/△ mice at PN 0. The size ratio of R2 relative to M1 increases significantly and progressively from Wnt1-Cre;Cilk1fl/fl mice to Cilk1−/− mice, and further to Cilk1−/−;PCS1–MRCS1△/△ mice. The mandibular R2/M1 ratio in Cilk1–/–;PCS1–MRCS1△/△ mice ranges from 0.90 to 1.16, whereas the maxillary R2/M1 ratio exhibits broader variability, ranging from 0.90 to 2.61. e A hypothetical model for tooth pattern formation influenced by Hh signaling: exploring how R2 and M1 interact. **P < 0.01
Furthermore, as Hh pathway activity decreased, the relative size of R2 to M1 (R2/M1 ratio) progressively increased. The R2/M1 ratio significantly increased across Wnt1-Cre;Cilk1fl/fl mice, Cilk1−/− mice, and Cilk1−/−;PCS1–MRCS1△/△ mice. In the mandible of Cilk1−/−;PCS1–MRCS1△/△ mice, the R2/M1 ratio ranged from 0.90 to 1.16 (Fig. 9d), and the R2/M1 ratio in the maxilla of Cilk1−/−;PCS1–MRCS1△/△ mice varied widely between 0.90 and 2.61 (Fig. 9c), suggesting that as Hh signaling activity decreases, R2 becomes increasingly larger relative to M1. Notably, R2 exceeded the size of M1 at low levels of Hh activity, indicating a gradient-dependent relationship between Hh pathway suppression and tooth patterning abnormalities.
Discussion
Cilk1 and craniofacial development
The significance of primary cilia in craniofacial development is well established, as mutations in primary cilia-related genes in both humans and mice frequently result in cleft lip, cleft palate, oligodontia, or supernumerary teeth. In humans, mutations in EVC, EVC2, IFT88, KIF3A, OFD1, and IFT121 have been implicated in these craniofacial defects.50,51,52,53,54,55 Similarly, in mice, deletions of Evc, Evc2, Fuz, Ift88, or Kif3a frequently cause a combination of cleft palate, tongue agenesis, mandibular hypoplasia, and R2 formation.47,48,56,57,58,59,60 While cleft palate and lobulated tongue had previously been reported in Cilk1-deficient mice,42 the present study provides the first evidence of R2 formation in this mutant. Additionally, R2 was observed in Wnt1-Cre;Ift88fl/fl and Wnt1-Cre;Kif3afl/fl mice, whereas Krt5-Cre;Ift88fl/fl mice exhibited no changes in tooth patterning.48,61 These findings, together with our results of R2 formation in Wnt1-Cre;Cilk1fl/fl but not in Krt14-Cre;Cilk1fl/fl mice, further indicate that mesenchymal primary cilia play a critical role in tooth patterning.
Cilk1 and the role of primary cilia in tooth development
Molar tooth germ of Cilk1−/− mice, when transplanted into the subcapsular layer of the kidney, formed the calcified teeth and surrounding bone at 4 weeks after transplantation. These Cilk1-deficient teeth showed significantly shorter root length compared to wild-type teeth. This is consistent with previous results that Ift80, or Ift140-deficient mice, which also have defects in primary cilia, show significantly short roots,62,63 suggesting an important role of primary cilia in tooth root growth.
The importance of primary cilia in dental hard tissue formation has been previously reported. Tooth germs from Wnt1-Cre;Kif3afl/fl mice, when transplanted into the kidney capsule, demonstrated the formation of odontoblasts, but with significantly thinner dentin and no enamel,64 and Ift80, Ift88, or Ift140-deficient mice showed thin dentin or enamel formation.61,62,63 Our findings in the present study contrast with observations from previous studies. Enamel, dentinal tubules, pre-dentin, dentin, surrounding bone, and odontoblasts in dental pulp were observed in these Cilk1−/− teeth. Additionally, Krt14-Cre;Cilk1fl/fl mice exhibited calcified teeth without any phenotypic changes. These findings suggest that Cilk1 might not be essential for ameloblasts and odontoblasts differentiation and their hard tissue formation. However, analyzing the shape of the crown and root, as well as the volume of hard tissue, in teeth developed through culture in the kidney is challenging due to spatial constraints within the renal capsule. Therefore, the precise role of Cilk1 in hard tissue formation should be further investigated.
Cilk1, Hh signaling, and tooth patterning
Mutations in primary cilia-related genes often result in similar phenotypic alterations due to their central role in Hh signaling.3,4,60,64,65 Prior studies have established a strong correlation between Cilk1 and Hh signaling in various organs, including the central nervous system, smooth muscle patterning, and palatal fusion.28,66,67 The present study extends this correlation to tooth development by demonstrating a significant reduction in Hh signaling activity in Cilk1-deficient tooth germs.
Different tooth patterns arise depending on Hh signaling activity. For example, PCS1–MRCS1△/△ mice and MRCS1△/△;MFCS4△/△ mice, in which Shh enhancers were removed, develop R2.12,21 In contrast, Krt14-Cre;Shhfl/− and Krt14-Cre;Smofl/fl mice exhibit molar fusion.22,23 Furthermore, inhibition of Hh signaling through anti-Shh antibodies or Hh pathway inhibitors has been shown to induce supernumerary tooth formation and molar fusion.13,26
Phenotypic alterations in tooth pattern can emerge or become more pronounced in compound mutant mice carrying mutations in two different genetic backgrounds. For instance, Sostdc1+/−;Shh+/− mice, generated by crossing Sostdc1+/− and Shh+/− mice with normal tooth patterns, exhibited R2.14 Additionally, Wnt10a+/− and Wnt10b−/− mice exhibit a normal tooth pattern, and Wnt10a+/−;Wnt10b−/− mice develop R2.68 However, it has remained unclear whether molar fusion can be induced in compound mutant mice combining different mutational backgrounds, all of which have R2. In this study, we demonstrated that Cilk1−/−;PCS1–MRCS1△/△ mice developed fused molars in the maxilla. This synergistic effect is due to the simultaneous suppression of Hh signaling in both Shh-producing and Shh-responsive cells (Supplementary Fig. S6). The progressive suppression of Hh signaling from wild-type to Cilk1−/− mice and further to Cilk1−/−;PCS1–MRCS1△/△ mice was validated by differential gene expression analyses, including RNA-sequencing and quantitative PCR. These findings strongly suggest that R2 formation and molar fusion occur progressively as Hh signaling activity declines.
Our study revealed distinct differences in susceptibility to R2 formation and molar fusion between the maxilla and mandible. R2 formation was more prevalent in the maxilla than in the mandible of Wnt1-Cre;Cilk1fl/fl and PCS1–MRCS1△/△ mice. In contrast, a previous study reported a higher prevalence of R2 in the mandibular quadrants (92%) compared to the maxillary quadrants (58%) in MRCS1△/△;Shh+/− mice, which lack a distinct Shh enhancer.21 A similar discrepancy has been documented in Spry2 and Spry4 mutant mice, which are negative regulators of the FGF signaling pathway. Among 55 Spry2−/− mice analyzed, 97% exhibited R2 formation in the mandible, whereas fewer than 5% showed R2 in the maxilla. In contrast, Spry4−/− mice showed R2 formation in 17% of maxillary and only 3% of mandibular quadrants.69,70 These findings underscore the need for further systematic investigations into the frequency and distribution of R2 formation across jaw regions. Additionally, in Cilk1−/−;PCS1–MRCS1△/△ mice, molar fusion was exclusively observed in the maxilla, consistent with findings in Evc and Evc2-deficient mice. Evc-deficient mice exhibit molar fusion with a penetrance of 100% in the maxilla but only 17% in the mandible.58 Similarly, Evc2 deficiency results in molar fusion restricted to the maxilla, while only R2 formation develops in the mandible.60 Comparable jaw-specific differences have also been reported in Sostdc1−/− mice. Sostdc1 encodes a Wnt signaling inhibitor and is a downstream target of the Hedgehog pathway. In Sostdc1−/− mice, molar fusion occurs exclusively in the maxilla without accompanying R2, whereas in the mandible, R2 formation is significantly more frequent than molar fusion.71,72 The mechanisms underlying these regional differences are unknown but may be related to gene expression and signaling differences between the maxilla and mandible. One plausible explanation for the consistent maxilla-specific occurrence of molar fusion involves differential expression of Wnt inhibitors. Previous study has shown that the Wnt antagonists Dkk2 and Wif1 are expressed at significantly higher levels in the mandibular molar mesenchyme compared to the maxillary counterpart in wild-type embryos.73 It was proposed that these elevated levels may contribute to the more pronounced mandibular dental defects observed in Wnt1-Cre;Bmp4fl/fl mice. By extension, in Cilk1−/−;PCS1–MRCS1△/△ and Sostdc1−/− mice, the mandibular enrichment of alternative Wnt inhibitors such as Dkk2 and Wif1 may partially compensate for the loss of Sostdc1, thereby reducing the likelihood of molar fusion in the mandible.
Progressive enlargement of diastemal supernumerary tooth under Hh signaling suppression
Diverse tooth patterns have been reported in many mutant mice, but understanding the mechanism of tooth patterning with significantly reduced Hh or Wnt signaling activity remains challenging. For example, both Gas1+/– mice and ShhGFP/+ mice develop a normal tooth pattern, while approximately 80% of Gas1–/– mice exhibit R2. Surprisingly, the penetrance of R2 formation decreases to 38% in Gas1–/–;ShhGFP/+ mice.20 Similarly, both Wnt10a+/− and Wnt10b−/− mice have a normal tooth pattern, whereas 80% of Wnt10a+/−;Wnt10b−/− mice develop R2. Remarkably, once more, Wnt10a−/−;Wnt10b−/− mice, despite displaying significantly smaller molars, maxilla, mandible, and body length, exhibit a normal-like tooth pattern. Since Wnt10a−/−;Wnt10b−/− mice retain three molars per jaw quadrant, with the most mesial molar notably larger than the next molar, their tooth pattern was classified as lacking R2.68
Comparable patterns were observed in this study. Wnt1-Cre;Cilk1fl/fl, PCS1–MRCS1△/△, and Cilk1−/− mice exhibited R2, which were consistently smaller than M1 within the same quadrant. However, in Cilk1−/−;PCS1–MRCS1△/△ mice, the R2 was equal to or larger than M1. The size ratio of R2 relative to M1 increased significantly and progressively from Wnt1-Cre;Cilk1fl/fl mice to Cilk1−/− mice, and further to Cilk1−/−;PCS1–MRCS1△/△ mice.
These findings suggest that a reduction in Hh signaling activity promotes R2 formation and progressively increases the relative size of R2. Under conditions of markedly diminished Hh signaling, R2 may exceed the size of M1. Reanalyzing results from Gas1–/–;ShhGFP/+ mice and Wnt10a−/−;Wnt10b−/− mice, the most mesial molar in each quadrant is likely an R2 rather than M1. Moreover, the normal-like tooth pattern observed in these mice should not be interpreted as a rescued phenotype, but rather as an exacerbated phenotype.
A progressive model from diastemal supernumerary tooth to molar fusion
In this study, we identified a unique tooth pattern characterized by R2 enlargement. In some Cilk1−/−;PCS1–MRCS1△/△ mice, the R2 in the maxilla was approximately 2.5 times larger than M1 and comparable in size to the fused molar. These findings suggest that extreme enlargement of R2 may serve as a precursor to molar fusion. We propose a progressive model in which the R2 retains, enlarges, and subsequently fuses with M1 as Hh signaling activity declines.
This progressive model can be analyzed by the Wnt-Shh-Sostdc1 negative feedback loop, which is the previously suggested mechanism for tooth pattern formation.12,13,14 In wild-type mice, the R2 present at E13 degenerates due to inhibitors (Sostdc1) secreted by M1, resulting in the three-molar pattern. However, in mice with diminished Hh signaling, the R2 retains rather than regressing, as the reduced activity of mediators (Shh signaling) in M1 leads to lower inhibitor levels. As Shh signaling activity continues to decline, the mutual inhibitory effects between developing R2 and M1 weaken, allowing the R2 to expand. With further Hh pathway suppression, inhibitors are lost in all molars, leading to a blurred boundary between adjacent molars and eventual molar fusion (Fig. 9e). A broader spectrum of tooth pattern diversity in other signaling pathways, in addition to the Hh pathway, will provide further support for the progressive model.
Collectively, our findings highlight the critical role of Cilk1 in primary cilia function and the importance of Hh signaling during tooth development. We uncover a new insight into tooth patterning dynamics: R2 can be larger than M1. Furthermore, we propose a progressive model where R2 formation, R2 enlargement, and ultimately tooth fusion occur sequentially in response to the degree of Hh pathway suppression.
Materials and methods
Animals
All animal experiments were performed under approved protocols of Yonsei University Health System Institutional Animal Care and Use Committee (YUHS-IACUC). All mice used in this study were euthanized via CO2 exposure at selected embryonic and postnatal days. A mix of males and females was assigned without considering the sex, and at least one individual from each sex was included in each group after screening for the genotype with PCR. All mice were maintained in the C57BL/6 background. PCS1–MRCS1△/△ mice with a deletion of a 70 kb region from PCS1 to MRCS1 were previously described.12 Cilk1−/−, Cilk1fl/fl, Wnt1-Cre, and Krt14-Cre mice were described previously.22,28,42,66,74,75 In brief, Cilk1fl/fl mice were generated by crossing Cilk1tm1a(KOMP)Mbp mice with Rosa26-FLP1 mice, which express FLP1 recombinase under the control of Rosa26. To generate Cilk1−/− mice, Cilk1fl/fl mice were crossed with EIIa-Cre transgenic mice, enabling Cre-mediated deletion of Cilk1 exon 6 in germ cells. Cilk1–/– mice were subsequently crossed with PCS1–MRCS1△/△ mice to generate Cilk1–/–;PCS1–MRCS1△/△ mice. For tissue-specific deletion, Cilk1fl/fl mice were crossed with Krt14-Cre and Wnt1-Cre transgenic mice to obtain epithelial- and mesenchyme-specific Cilk1 knockout mice, respectively (Supplementary Fig. S8).
Section in situ hybridization
Heads were dissected from wild-type mouse embryos at E13, E14, E15.5, and E16.5 on cold DEPC-PBS and fixed with 4% PFA in DEPC-PBS overnight at 4 °C with rocking. After being rinsed in DEPC-PBS, the samples were decalcified for 3 h at 4 °C using 10% EDTA in DEPC-PBS. Decalcified samples were washed with DEPC-PBS, followed by a saline wash. Samples were dehydrated with a graded ethanol series. Embryo samples were embedded in paraffin and sectioned at 5 µm thickness. Advanced Cell Diagnostics (Newark, USA) designed and manufactured antisense Cilk1 RNA probes. The RNAscope® 2.5 High Definition (HD) assay-brown (Advanced Cell Diagnostics) was used for in situ hybridization in accordance with the manufacturer’s instructions for 322452 (FFPE sample preparation and pretreatment) and 322310 (RNAscope® 2.5 HD Detection Reagent-brown). Mm-Cilk1 (1177181, targeting NM_001163780.1, nucleotide 1358-2353), Mm-Ptch1 (402811, targeting NM_008957.2, nucleotide 2260-3220), Mm-Sostdc1 (313151, targeting NM_025312.3, nucleotide 22-1385), Mm-Lef1 (441861, targeting NM_175263.4, nucleotide 406-1623) RNA probes were used.
Histological and immunohistochemistry staining
Heads were fixed with 4% PFA at 4 °C for 24 h and decalcified in 10% ethylenediaminetetraacetic acid (EDTA) at 4 °C for 24 h. The tissues were then dehydrated through a graded ethanol series, embedded in paraffin, and sectioned at 5 or 7 µm thickness. For histological analysis, sections were stained with hematoxylin and eosin using standard protocols. For immunohistochemistry staining, sections underwent antigen retrieval by incubation in citrate buffer (pH 6.0) at 90 °C for 1 h, followed by blocking with 5% bovine serum albumin (BSA) in phosphate-buffered saline with Tween 20 (PBST) at room temperature for 40 min. Sections for primary cilia staining were incubated overnight at 4 °C with Arl13b antibody.76 Sections for epithelial staining were incubated overnight at 4 °C with a primary anti-cytokeratin 14 antibody (1:200, LS-B8270, LSBio), washed with PBST, and then incubated for 1 h at room temperature with an Alexa Fluor-conjugated secondary antibody (1:200, Invitrogen). Following antibody incubation, sections were washed and counterstained with DAPI (6-diamidino-2-phenylindole, Sigma) to visualize the nuclei. Images were captured using a fluorescence microscope (Leica Microsystem) or a Zeiss LSM 900 confocal microscope. The ciliary length (n = 24 per group) was measured using the Zeiss Zen 3.3 (blue edition) program in a single plane (7 µm thickness) of confocal microscopy Z-stack images. The percentage of ciliated cells was determined by counting DAPI-positive nuclei and Arl13b-positive primary cilia in sections (n = 4 per group).
Single-cell RNA-sequencing analysis
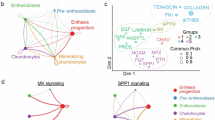
The scRNA-seq datasets for E14.5 and E16.5 mouse mandibular molars were processed, analyzed, and visualized using publicly available Gene Expression Omnibus data (GSE189381 and GSE162413).40,41 We imported the pre-processed count matrices into the Biomage (https://biomage.net/)-hosted instance of Trailmaker® (https://app.trailmaker.parsebiosciences.com/). Data handling and cell clustering were described previously.77 All cell clusters were manually annotated based on existing literatures.40,41 The differentially expressed genes in each cluster were listed in the cluster map and heatmap previously.77
Three-dimensional (3D) reconstruction using computed tomography
The dissected embryo heads were stained with phosphotungstic acid (P4006-10G, Sigma-Aldrich) and scanned by a micro-computed tomography (CT) system (Skyscan 1173; Bruker, Kontich, Belgium) using 90 kV, 88 μA, 360° rotation with a step of 0.3°, pixel size of 6 μm and an aluminum filter (1.0 mm of thickness) and reconstructed by the NRecon software (Bruker). The reconstructions were visualized and converted to 3D volumes using the software 3D Slicer.78 Crown volume (n = 5 per group) and root length (n = 5 per group) were measured directly from the 3D reconstructed images.
RNA-sequencing analysis
Maxillary molars were dissected from mouse embryos at E14.5 (n = 6 per biological replicate, and n = 4 for wild-type and Cilk1–/– group, and n = 3 for Cilk1–/–;PCS1–MRCS1△/△ group) in DEPC-PBS. The 0.5 mm zirconia beads were used to homogenize tooth germ samples in TRIZOL® (Invitrogen) using the Bullet Blender® homogenizer (Next Advance, USA). Chloroform was used to phase separate the total RNA, and isopropyl alcohol was used to precipitate it. After being cleaned with 75% ethanol, the RNA pellet was eluted using RNase-free water. NanoDrop (Thermo Fisher Scientific, USA) was used to measure the concentration and quality of RNA. After the library was built and sequenced on an Illumina platform, paired-end reads were produced, quality-controlled, and mapped to the reference genome (mm10). The number of reads mapped to each gene was quantified using FeatureCounts v1.5.0-p3, and FPKM values were calculated based on gene length and read counts. Differential expression analysis between the two conditions was performed using the edgeR package (3.22.5), with read counts adjusted by a scaling normalization factor. P values were adjusted using the Benjamini–Hochberg method, and a corrected Padj value of 0.05 and an absolute fold change of 1.5 were set as the threshold for significance. Gene Ontology (GO) enrichment analysis of differentially expressed genes was carried out using ShinyGo 0.77 (http://bioinformatics.sdstate.edu/go/). GO terms with corrected P values below 0.05 were considered significantly enriched.
Statistical analysis
The percentage of ciliated cells was analyzed using the Mann–Whitney U test, while the ciliary length and M1/M2 ratio were analyzed using Student’s t-tests (two-tailed). The R2/M1 ratio was assessed using one-way analysis of variance (ANOVA) with Tukey’s test for multiple comparisons. Every graph was visualized with Prism 8.0 (GraphPad, San Diego, CA, USA). All data are presented as mean ± SEM. P value less than 0.05 was considered a statistically significant difference. P value larger than 0.05 was considered a statistically non-significant difference.
Transplantation of tooth germs into the renal subcapsular layer
Developing molars (n = 5 per group) were isolated from mandibles of mouse embryos at E14.5, transplanted into the renal subcapsular layer of C57BL/6 adult male mice, and harvested after 4 weeks. All surgical procedures were performed under anesthesia administered intraperitoneally. No immunosuppressive medication was used.
Isolation of oral epithelium from tongues and palates in Krt14-Cre;Cilk1 fl/fl mice
Mucosa from the tongue and palate of Krt14-Cre;Cilk1fl/fl mice at PN 3 were dissected and incubated in Dispase II (1.2 U/mL in PBS) for 30 min. After incubation, the tissues were washed in DMEM supplemented with 10% FBS, and the oral epithelium was carefully separated from the underlying mesenchyme. gDNA was extracted from these epithelia, and genotyping PCR was carried out across three individuals.
RNA extraction and quantitative RT–PCR
Total RNA was extracted from E14.5 embryonic tooth germs (n = 6 per biological replicate and triplicate per group) using TRIzol reagent (Qiagen). One microgram of RNA was converted to complementary DNA (cDNA) using the First-Strand cDNA Synthesis Kit (Takara) according to the manufacturer’s instructions. Quantitative reverse transcription PCR was conducted on a real-time PCR system using SYBR Premix Ex Taq™ (Takara). Gene expression was analyzed using the following primer pairs: Gli1 (forward: GAGGTTGGGATGAAGAAGCA, reverse: GTGCCAATC-CGGTGGAGTCAGACCC), Ptch1 (forward: GTGGACGAGTGAGTCGAGAAT, reverse: GGAGTGCTGAGTCCAGGTGT), Sostdc1 (forward: AGGCAGGCATTTCAGTAGCA, reverse: CTGGCCGTCCGAAATGTA), and Hhip (forward: GGCTCTGTCGAAACGGCTAC, reverse: GGCACTTGTTCGGTCTGACA). Gapdh was used as the internal control, with primer sequences as follows: forward: AACAGCAACTCCCACTCTTC, reverse: CCTGTTGCTGTAGCCGTATT. Relative gene expression was determined using the 2−ΔΔCt method with QuantStudio Design and Analysis software.
Data availability
The raw and processed bulk RNA-Seq data associated with this study have been deposited in the NCBI GEO database under the accession number GSE290878.
References
Christensen, S. T., Pedersen, L. B., Schneider, L. & Satir, P. Sensory cilia and integration of signal transduction in human health and disease. Traffic 8, 97–109 (2007).
Hampl, M. et al. Role of primary cilia in odontogenesis. J. Dent. Res. 96, 965–974 (2017).
Huangfu, D. et al. Hedgehog signalling in the mouse requires intraflagellar transport proteins. Nature 426, 83–87 (2003).
Goetz, S. C. & Anderson, K. V. The primary cilium: a signalling centre during vertebrate development. Nat. Rev. Genet. 11, 331–344 (2010).
Simons, M. et al. Inversin, the gene product mutated in nephronophthisis type II, functions as a molecular switch between Wnt signaling pathways. Nat. Genet. 37, 537–543 (2005).
Neugebauer, J. M., Amack, J. D., Peterson, A. G., Bisgrove, B. W. & Yost, H. J. FGF signalling during embryo development regulates cilia length in diverse epithelia. Nature 458, 651–654 (2009).
Kim, M., Kim, M., Lee, M. S., Kim, C. H. & Lim, D. S. The MST1/2-SAV1 complex of the Hippo pathway promotes ciliogenesis. Nat. Commun. 5, 5370 (2014).
Boehlke, C. et al. Primary cilia regulate mTORC1 activity and cell size through Lkb1. Nat. Cell Biol. 12, 1115–1122 (2010).
Wu, D. et al. Intestinal cell kinase (ICK) promotes activation of mTOR complex 1 (mTORC1) through phosphorylation of Raptor Thr-908. J. Biol. Chem. 287, 12510–12519 (2012).
Schneider, L. et al. PDGFRalphaalpha signaling is regulated through the primary cilium in fibroblasts. Curr. Biol. 15, 1861–1866 (2005).
Ezratty, E. J. et al. A role for the primary cilium in Notch signaling and epidermal differentiation during skin development. Cell 145, 1129–1141 (2011).
Seo, H. et al. Upstream enhancer elements of Shh regulate oral and dental patterning. J. Dent. Res. 97, 1055–1063 (2018).
Cho, S. W. et al. Interactions between Shh, Sostdc1 and Wnt signaling and a new feedback loop for spatial patterning of the teeth. Development 138, 1807–1816 (2011).
Ahn, Y., Sanderson, B. W., Klein, O. D. & Krumlauf, R. Inhibition of Wnt signaling by Wise (Sostdc1) and negative feedback from Shh controls tooth number and patterning. Development 137, 3221–3231 (2010).
Zhang, H., Gong, X., Xu, X., Wang, X. & Sun, Y. Tooth number abnormality: from bench to bedside. Int. J. Oral. Sci. 15, 5 (2023).
Viriot, L. et al. The presence of rudimentary odontogenic structures in the mouse embryonic mandible requires reinterpretation of developmental control of first lower molar histomorphogenesis. Int. J. Dev. Biol. 44, 233–240 (2000).
Prochazka, J. et al. Migration of founder epithelial cells drives proper molar tooth positioning and morphogenesis. Dev. Cell 35, 713–724 (2015).
Peterkova, R., Lesot, H. & Peterka, M. Phylogenetic memory of developing mammalian dentition. J. Exp. Zool. B Mol. Dev. Evol. 306, 234–250 (2006).
Prochazka, J. et al. Patterning by heritage in mouse molar row development. Proc. Natl. Acad. Sci. USA 107, 15497–15502 (2010).
Seppala, M. et al. Gas1 regulates patterning of the murine and human dentitions through sonic Hedgehog. J. Dent. Res. 101, 473–482 (2022).
Sagai, T. et al. SHH signaling directed by two oral epithelium-specific enhancers controls tooth and oral development. Sci. Rep. 7, 13004 (2017).
Dassule, H. R., Lewis, P., Bei, M., Maas, R. & McMahon, A. P. Sonic hedgehog regulates growth and morphogenesis of the tooth. Development 127, 4775–4785 (2000).
Gritli-Linde, A. et al. Shh signaling within the dental epithelium is necessary for cell proliferation, growth and polarization. Development 129, 5323–5337 (2002).
Kassai, Y. et al. Regulation of mammalian tooth cusp patterning by ectodin. Science 309, 2067–2070 (2005).
Ohazama, A. et al. Lrp4 modulates extracellular integration of cell signaling pathways in development. PLoS ONE 3, e4092 (2008).
Mao, C. et al. Revitalizing mouse diphyodontic dentition formation by inhibiting the sonic hedgehog signaling pathway. Dev. Dyn. 251, 759–776 (2022).
Satoda, Y. et al. BROMI/TBC1D32 together with CCRK/CDK20 and FAM149B1/JBTS36 contributes to intraflagellar transport turnaround involving ICK/CILK1. Mol. Biol. Cell 33, ar79 (2022).
Chaya, T., Omori, Y., Kuwahara, R. & Furukawa, T. ICK is essential for cell type-specific ciliogenesis and the regulation of ciliary transport. EMBO J. 33, 1227–1242 (2014).
Burghoorn, J. et al. Mutation of the MAP kinase DYF-5 affects docking and undocking of kinesin-2 motors and reduces their speed in the cilia of Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 104, 7157–7162 (2007).
Oh, Y. S. et al. Ciliopathy-associated protein kinase ICK requires its non-catalytic carboxyl-terminal domain for regulation of ciliogenesis. Cells 8, 677 (2019).
Wang, E. J., Gailey, C. D., Brautigan, D. L. & Fu, Z. Functional alterations in ciliogenesis-associated kinase 1 (CILK1) that result from mutations linked to juvenile myoclonic epilepsy. Cells 9, 694 (2020).
Liang, Y. et al. FLA8/KIF3B phosphorylation regulates kinesin-II interaction with IFT-B to control IFT entry and turnaround. Dev. Cell 30, 585–597 (2014).
Broekhuis, J. R., Verhey, K. J. & Jansen, G. Regulation of cilium length and intraflagellar transport by the RCK-kinases ICK and MOK in renal epithelial cells. PLoS ONE 9, e108470 (2014).
Nakamura, K. et al. Anterograde trafficking of ciliary MAP kinase-like ICK/CILK1 by the intraflagellar transport machinery is required for intraciliary retrograde protein trafficking. J. Biol. Chem. 295, 13363–13376 (2020).
Reiter, J. F. & Leroux, M. R. Genes and molecular pathways underpinning ciliopathies. Nat. Rev. Mol. Cell Biol. 18, 533–547 (2017).
Yang, Y., Roine, N. & Makela, T. P. CCRK depletion inhibits glioblastoma cell proliferation in a cilium-dependent manner. EMBO Rep. 14, 741–747 (2013).
Willaredt, M. A. et al. A crucial role for primary cilia in cortical morphogenesis. J. Neurosci. 28, 12887–12900 (2008).
Garcia-Garcia, M. J. et al. Analysis of mouse embryonic patterning and morphogenesis by forward genetics. Proc. Natl. Acad. Sci. USA 102, 5913–5919 (2005).
Ferrante, M. I. et al. Oral-facial-digital type I protein is required for primary cilia formation and left-right axis specification. Nat. Genet. 38, 112–117 (2006).
Jing, J. et al. Spatiotemporal single-cell regulatory atlas reveals neural crest lineage diversification and cellular function during tooth morphogenesis. Nat. Commun. 13, 4803 (2022).
Hu, H. et al. Dental niche cells directly contribute to tooth reconstitution and morphogenesis. Cell Rep. 41, 111737 (2022).
Moon, H. et al. Intestinal cell kinase, a protein associated with endocrine-cerebro-osteodysplasia syndrome, is a key regulator of cilia length and Hedgehog signaling. Proc. Natl. Acad. Sci. USA 111, 8541–8546 (2014).
Hisamoto, M. et al. Developmental changes in primary cilia in the mouse tooth germ and oral cavity. Biomed. Res. 37, 207–214 (2016).
Hari, L. et al. Temporal control of neural crest lineage generation by Wnt/beta-catenin signaling. Development 139, 2107–2117 (2012).
Vasioukhin, V., Degenstein, L., Wise, B. & Fuchs, E. The magical touch: genome targeting in epidermal stem cells induced by tamoxifen application to mouse skin. Proc. Natl. Acad. Sci. USA 96, 8551–8556 (1999).
Hafner, M. et al. Keratin 14 Cre transgenic mice authenticate keratin 14 as an oocyte-expressed protein. Genesis 38, 176–181 (2004).
Tian, H. et al. Intraflagellar transport 88 (IFT88) is crucial for craniofacial development in mice and is a candidate gene for human cleft lip and palate. Hum. Mol. Genet. 26, 860–872 (2017).
Ohazama, A. et al. Primary cilia regulate Shh activity in the control of molar tooth number. Development 136, 897–903 (2009).
Sagai, T. et al. A cluster of three long-range enhancers directs regional expression in the epithelial linings. Development 136, 1665–1674 (2009).
Dodge, J. A. & Kernohan, D. C. Oral-facial-digital syndrome. Arch. Dis. Child 42, 214–219 (1967).
Curry, C. J. & Hall, B. D. Polydactyly, conical teeth, nail dysplasia, and short limbs: a new autosomal dominant malformation syndrome. Birth Defects Orig. Artic. Ser. 15, 253–263 (1979).
Levin, L. S. et al. A heritable syndrome of craniosynostosis, short thin hair, dental abnormalities, and short limbs: cranioectodermal dysplasia. J. Pediatr. 90, 55–61 (1977).
Susami, T., Kuroda, T., Yoshimasu, H. & Suzuki, R. Ellis-van Creveld syndrome: craniofacial morphology and multidisciplinary treatment. Cleft Palate Craniofacial J. 36, 345–352 (1999).
Baujat, G. & Le Merrer, M. Ellis-van Creveld syndrome. Orphanet J. Rare Dis. 2, 27 (2007).
Bureau, A. et al. Whole exome sequencing of distant relatives in multiplex families implicates rare variants in candidate genes for oral clefts. Genetics 197, 1039–1044 (2014).
Brugmann, S. A. et al. A primary cilia-dependent etiology for midline facial disorders. Hum. Mol. Genet. 19, 1577–1592 (2010).
Yuan, G. et al. Cleft palate and aglossia result from perturbations in Wnt and Hedgehog signaling. Cleft Palate Craniofacial J. 54, 269–280 (2017).
Nakatomi, M. et al. Evc regulates a symmetrical response to Shh signaling in molar development. J. Dent. Res. 92, 222–228 (2013).
Zhang, Z. et al. Fuz regulates craniofacial development through tissue specific responses to signaling factors. PLoS ONE 6, e24608 (2011).
Zhang, H. et al. Elevated WNT signaling and compromised Hedgehog signaling due to Evc2 loss of function contribute to the abnormal molar patterning. Front. Dent. Med. 3, https://doi.org/10.3389/fdmed.2022.876015 (2022).
Kudo, T. et al. Ift88 regulates enamel formation via involving Shh signaling. Oral. Dis. 29, 1622–1631 (2023).
Yuan, X., Cao, X. & Yang, S. IFT80 is required for stem cell proliferation, differentiation, and odontoblast polarization during tooth development. Cell Death Dis. 10, 63 (2019).
Li, G. et al. Essential role of IFT140 in promoting dentinogenesis. J. Dent. Res. 97, 423–431 (2018).
Liu, B., Chen, S., Cheng, D., Jing, W. & Helms, J. A. Primary cilia integrate hedgehog and Wnt signaling during tooth development. J. Dent. Res. 93, 475–482 (2014).
Dorn, K. V., Hughes, C. E. & Rohatgi, R. A Smoothened-Evc2 complex transduces the Hedgehog signal at primary cilia. Dev. Cell 23, 823–835 (2012).
Yang, Y. et al. Ciliary Hedgehog signaling patterns the digestive system to generate mechanical forces driving elongation. Nat. Commun. 12, 7186 (2021).
Shin, J. O. et al. Activation of sonic hedgehog signaling by a Smoothened agonist restores congenital defects in mouse models of endocrine-cerebro-osteodysplasia syndrome. EBioMedicine 49, 305–317 (2019).
Yoshinaga, K. et al. Effects of Wnt10a and Wnt10b double mutations on tooth development. Genes 14, 340 (2023).
Klein, O. D. et al. Sprouty genes control diastema tooth development via bidirectional antagonism of epithelial-mesenchymal FGF signaling. Dev. Cell 11, 181–190 (2006).
Marangoni, P. et al. Phenotypic and evolutionary implications of modulating the ERK-MAPK cascade using the dentition as a model. Sci. Rep. 5, 11658 (2015).
Murashima-Suginami, A. et al. Rudiment incisors survive and erupt as supernumerary teeth as a result of USAG-1 abrogation. Biochem. Biophys. Res. Commun. 359, 549–555 (2007).
Kim, J. et al. Shh plays an inhibitory role in Cusp patterning by regulation of Sostdc1. J. Dent. Res. 98, 98–106 (2019).
Jia, S. et al. Roles of Bmp4 during tooth morphogenesis and sequential tooth formation. Development 140, 423–432 (2013).
Jiang, X., Rowitch, D. H., Soriano, P., McMahon, A. P. & Sucov, H. M. Fate of the mammalian cardiac neural crest. Development 127, 1607–1616 (2000).
Paige Taylor, S. et al. An inactivating mutation in intestinal cell kinase, ICK, impairs hedgehog signalling and causes short rib-polydactyly syndrome. Hum. Mol. Genet. 25, 3998–4011 (2016).
Kim, Y. K., Kim, J. H., Yu, Y. S., Ko, H. W. & Kim, J. H. Localization of primary cilia in mouse retina. Acta Histochem. 115, 789–794 (2013).
Adasooriya, D. et al. Notum regulates the cusp and root patterns in mouse molar. Sci. Rep. 14, 13633 (2024).
Fedorov, A. et al. 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn. Reson. Imaging 30, 1323–1341 (2012).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIT) (NRF-2020R1A2C2005790, NRF-2023R1A2C1007510, RS-2023-00269830, RS-2024-00438542). The authors thank Jangki Jo and Chae Eun Lee (Yonsei University College of Dentistry) and Bioinformatics Collaboration Unit (BiCU) in the Yonsei Biomedical Research Institute (Yonsei University College of Medicine) for their technical assistance in this study.
Author information
Authors and Affiliations
Contributions
M.K., J.K.J., H.W.K., E.S.C. and S.W.C. contributed to the conception, design, data acquisition, and interpretation, and drafted the manuscript. D.A., S.K., J.K., J.S., S.P., S.J., S.J.M. and Y.B.P. contributed to data acquisition and interpretation. H.W.K., E.S.C. and S.W.C. supervised the study. All authors critically revised the manuscript and provided final approval of the version to be published.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kyeong, M., Jeong, JK., Adasooriya, D. et al. Progressive tooth pattern changes in Cilk1-deficient mice depending on Hedgehog signaling. Int J Oral Sci 17, 71 (2025). https://doi.org/10.1038/s41368-025-00405-4
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41368-025-00405-4