Abstract
Brain disorders caused by large effect mutations in single genes often present unexplained large symptom diversity, even among carriers of the same mutation. Here we examined genetic interactions as a possible explanation for this diversity for SNAREopathies, a group of common neurodevelopmental disorders caused by de novo genetic variation in genes that together drive secretion of chemical signals in the brain. SNAREopathies are characterized by a striking phenotypic diversity, including different types/degrees or absence of seizures, developmental delay and intellectual disability. Here, we test the hypothesis that large phenotypic diversity is caused by non-linear genetic interactions between two or more functionally related genes by combining validated SNAREopathy mouse models and comparing phenotypic diversity between single and double mutants at the synaptic, network, system and behavioral level. Single Stxbp1 and Snap25 mutant animals showed EEG- and motor abnormalities, but no seizures, as reported before. In contrast, double mutants exhibited extreme diversity in seizure phenotypes. Some mice had lethal generalized seizures, frequent and complex epileptiform EEG activity and thalamic hyper-excitability as indicated by increased cFos staining, while other mice of the same genotype showed no detectable abnormalities, no increased cFos staining and a normal life span. The surviving double mutant mice showed phenotypes not more severe than single mutants at the synaptic, network, and behavioral level. Finally, we present a theoretical framework to quantitatively explain our findings and extrapolate the conclusions to symptoms diversity in human patients. Taken together, this study shows that haploinsufficiency at two interacting loci leads to extreme phenotypic diversity at the systems level. These findings provide a proof of concept for how modifying genes in the patient genome enhance phenotypic diversity.
Similar content being viewed by others
Introduction
Developmental and epileptic encephalopathies (DEEs) are a heterogeneous group of severe childhood disorders characterized by different types of seizures and epileptiform activities, intellectual disability (ID) and severe neurodevelopmental delays [1, 2]. DEEs are characterized by large genetic and phenotypic diversity, with hundreds of genes implicated as causal, and each gene associated with multiple phenotypes [3,4,5,6]. The genetic diversity implies the involvement of several cellular pathways in the etiology of DEEs. Sub-grouping genes to specific cellular pathways could be used as a strategy to resolve this heterogeneity and elucidate disease mechanisms more efficiently. This led to classification of channelopathies, synaptopathies, and recently also SNAREopathies [5, 7, 8]. However, striking phenotypic diversity remains within such DEE sub-groups, even among patients with mutations in the same gene, suggesting that additional explanations are required. Several studies have suggested influences of interacting factors such as modifying genes, epigenetic factors, environmental factors, and stochastic processes [9,10,11]. However, experimental tests of such suggestions in controlled laboratory conditions are scarce.
SNAREopathies are caused by mutations in genes encoding the neuronal SNARE (soluble N-ethylmaleimide sensitive factor attachment protein receptor)-complex and its interactors, which together form the essential machinery for the secretion of chemical signals in the brain. Most SNAREopathies are caused by de novo missense or loss-of-function variants in one of the SNARE genes and several molecular disease mechanisms have been suggested including haploinsufficiency, dominant-negative, gain-of-function and recessive mechanisms [8, 12,13,14,15,16,17,18]. Despite the fact that SNARE-genes work together in a single, integrated molecular machine, a striking phenotypic diversity among SNAREopathy patients is evident [8, 15, 17, 18]. However, the incidence of pathogenic mutations in individual SNARE genes in the population is low, with an estimated incidence of one in 30,000 individuals for the most reported SNAREopathy to date, STXBP1 related disorder [15, 17,18,19,20]. This severely limits the power to identify modifying genes and epigenetic or environmental factors in population studies.
In the present study we examined how genetic interactions influence phenotypic diversity in controlled laboratory conditions using validated mouse models for SNAREopathies. We selected two SNARE genes, Stxbp1 and Snap25, firmly associated with DEEs and based on many molecular studies, expected to genetically interact. We established a panel of four mouse models for single and combined haploinsufficiency on the same genetic background to minimize other sources of genetic variation and tested the phenotypic consequences of gene interactions on the cellular level by assessing synaptic transmission in cultured neurons, on the network level by assessing spontaneous synchronous network activity (SSA) in acute brain slices and excessive neuronal activation by c-Fos staining in brain sections, and on the system level, using ECoG-video monitoring and cognitive assessments. We found robust support for multiplicative genetic interactions at the highest level of organization, but not at the lower (cellular) levels. Generalized seizures, complex epileptiform activities and brain hyper-excitability were observed in double mutants, but never in single mutants. Based on these data we generated a theoretical framework and defined three modes of gene interactions that can be experimentally tested: common pathway, additive- and multiplicative- interaction, partially based on previous studies in invertebrates [21, 22].
Materials and methods
Study design
The study had multilevel design: in silico, in vitro, ex vivo and in vivo. The study followed the ARRIVE guidelines (https://arriveguidelines.org/arrive-guidelines). We generated, behaviorally and electrographically characterized Stxbp1 single, Snap25 single and double mutants and compared them to wildtype littermates (controls). We performed standardized battery of behavioral tests for assessment of different aspects of behavior. For behavioral analysis and ECoG recording, we used in total 79 mice. Experiments were performed in several batches and data were pooled. Ex vivo experiments were performed on the brain slices from mice. For Ca2+-imaging of the brain slices, 34 mice were used, and between 3–5 replicates per animal were analyzed. C-Fos immunohistochemistry was performed on brain slices from 21 mice. In vitro analysis of electrophysiological properties of neurons was performed in two batches of mice (n = 31 and 23 per batch). All experiments were performed by researcher unaware of the animal’s genotype.
Ethics approval
All methods were performed in accordance with the relevant guidelines and regulations. All animal experiments were approved by the local animal research committee and complied with the European Council Directive (86/609/EEC).
Subjects
Control (Stxbp1+/+Snap25+/+), Snap25 single mutant (Stxbp1+/+Snap25+/-), Stxbp1 single mutant (Stxbp1+/-Snap25+/+) and double mutants (Stxbp1+/-Snap25+/-) mice were generated by mating male congenic C57BL6/J Stxbp1+/- mice [13, 23] with female Snap25+/- mice also bred on C57BL6/J genetic background, backcrossed for 20 generations [24]. All animals were kept in standard husbandry conditions on a 12 h light-dark cycle with food and water available ad libitum. Animals aged around 8 weeks were separately housed on sawdust in the standard Makrolon type II cages. In total 200 mice were used. Additionally, previously published spontaneous behavior data from 14 control and 15 Snap25 single mutants were reanalyzed and represented here [25]. Those data are publicly available and can be found on the following address: http://syli.cz/urn:lsid:public.sylics.com:experiment:5B6A-9DE7-G724.
Behavioral phenotyping
Behavioral experiments were performed on male mice between 8 and 16 weeks of age. In total three batches of Stxbp1Snap25 mice were used: batch 1 and 2 each consisted of 6 controls and 6 double mutants; batch 3 consisted of 8 controls, 8 Stxbp1 single-, 8 Snap25 single- and 8 double- mutant mice. In all three batches, we assessed spontaneous behavior in an automated home-cage environment (PhenoTyper model 3000, Noldus Information Technology, Wageningen, The Netherlands). Spontaneous behavior was automatically monitored for two and a half days in the PhenoTyper. Spontaneous home-cage behavior is highly dimensional aspect of behavior leading to separation of 90 separate parameters divided into 6 different categories [26]: kinematics, activity, sheltering, habituation, dark-light index and light-dark phase transition. Kinematic parameters describe specific elements of animal’s behavior related to movement characteristics, such as short and long movement segments and short and long arrest segments. Activity bouts describe mouse behavior on sub-minute scale and cumulatively during the period of days of spontaneous behavior monitoring. Sheltering behavior describes the tendency of mice to sleep inside the shelter [26, 27]. Habituation ratio evaluates changes in activity over days by taking the ratio for respective parameters of dark phase 3 over dark phase 1. Dark-light indexes assess difference in the behavior during the dark and light phase of cycle, by taking the ratio for respective parameters of dark phase over the light phase. The effect of light/dark phase transition on spontaneous behavior was assessed by analysis of change in activity parameters during the periods surrounding phase transition. To increase the validity of the conclusions based on spontaneous behavior of double mutants and Stxbp1 single and Snap25 single mutants, the data from two previously published studies performed in the same laboratory on Stxbp1 single and Snap25 single mutants and their respective controls [13, 25] were pooled together with results from the current study and statistical differences were represented by different colors in Fig. 4A, and Supplementary Table 1. Additionally, animals from batches 1–3 were separately analyzed and their results were shown in Fig. 4B–F.
The observation of spontaneous behavior was followed by the assessment of discrimination- and reversal learning using the CognitionWall task in the same automated home-cage [28]. After testing in the PhenoTyper, animals acclimated to the new housing for one week before further testing.
The batch 1 and batch 2 consisted of 12 control- and 12 double mutant- mice. We used standard behavioral battery for assessment of vision, muscle strength, motor coordination, anxiety, learning and memory, as previously described [13]. All tests were performed during the light phase with the least stressful tests done at the beginning, and at least 1 day apart. Briefly, the vision was assessed using the vision test; muscle strength was assessed measuring grip strength; motor coordination and motor learning were assessed on the rotarod. We used three anxiety-related paradigms: elevated plus maze test, open field and dark-light box. Fear-conditioning experiment evaluated associative learning and memory in mice. With Barnes maze test we tested spatial learning, memory and reversal learning. Short-term (working) memory was measured using T-maze spontaneous alteration task. Detailed protocols for these behavioral tests can be found in [13].
The third batch of haploinsufficiency mutant mice was tested for spontaneous behavior in the Phenotyper followed by the fear conditioning and Barnes maze, since results from the first two tested batches of animals showed most phenotypical changes in these two paradigms. Detailed protocols for these behavioral tests can be found in [13].
Video monitoring, simultaneous radiotelemetric video, ECoG recordings
Video monitoring of mice from batch 4, age: between 8 and 10 weeks old, have been performed in their home cages (2 control, 4 Snap25 single-, 6 Stxbp1 single-, and 6 double mutants). Mice were video monitored for at least 24 h. After video monitoring, seven mice from batch 4 and 13 mice from batches 1–3 were implanted with ECoG transmitters. In total: 4 control, 5 Stxbp1 single-, 4 Snap25 single-, and 7 double- mutants were implanted with ECoG transmitters (ETA-F10; specification: https://www.datasci.com/products/implantabletelemetry/mouse-(miniature)/eta-f10) as previously described in Kovačević et al. [13]. Briefly, mice were anesthetized with isoflurane (3% isoflurane/oxygene, flow 0.8 l/min) and immobilized in the stereotaxic instrument. After administration of lidocaine (2%, s.c.), a small incision was made on the skull allowing the recording electrode to be positioned above the motor cortex (2.2 mm anterior, 1 mm lateral) and ground electrode above the cerebellum (6 mm posterior, 1 mm lateral) using the stainless screws. The transmitter was placed in the abdominal subcutaneous pocket. The incision was closed with suture material. All animals received pre- and post-operative analgesic treatment with buprenorphine (0.05 mg/kg, s.c.). Animals were daily checked during recovery period of at least 7 days. During ECoG recordings (for at least 24 h), implanted animals were placed on the DSI receiver board (www.datasci.com) in front of an infrared camera allowing synchronous recording of behavior and ECoG signals.
Video and ECoG data were time matched at the beginning and at the end of recording. ECoG data were visually inspected and analyzed using the Event Classifier application within Neuroarchiver tool in LWDAQ software (Open Source Instruments, Inc.). The application classified the 1s-segments of EEG according to several metrics (asymmetry, intermittency, coherence, power, coastline and spikiness) enabling that similar patterns cluster together [29].Video analysis was performed independently of ECoG data analysis and later matched with ECoG data. Sleep states were identified from the video recordings by researcher. ECoG signal that corresponds to the sleep state were selected and divided in one-minute epochs. Epochs were visually checked and those that contained artifacts were excluded from analysis. Spectral power analysis during the sleep episodes was performed using open-source software package Chronux (http://chronux.org/) for signal processing of neural time-series data [30]. We used routine mtspectrumc which is applicable for analysis of continuous-valued data using the moving-window with 2 s time-width and 1 s step-size. Data were sampled at 1 kHz and high-pass filtered at 1 Hz and low-pass filtered at 200 Hz. Average spectral power was calculated as a mean value of relative power (expressed as the ratio of total power) for all 1-min episodes per animal. Power per frequency band was calculated as the average relative power within the frequency ranges: 1–4 Hz for delta band, 5–8 Hz for theta band, 9–12 Hz for alfa band. Spectrogram were made using our own written script in Matlab.
c-Fos staining
Mice, around 3 months old, were sacrificed either by cervical dislocation or overdose of avertin (2,2,2-tribromoethyl alcohol). Animals sacrificed by avertin were perfused with 4% paraformaldehyde (PFA) in 0.1 M phosphate buffered saline (PBS.) In the present study, 5 Snap25 single mutants, 10 double mutants and 6 controls were used for c-Fos analyses. In addition, we re-analyzed data from 5 Stxbp1 single mutants obtained in our laboratory and compared these to the controls from the same experiment. All brains were removed and post-fixed in 4% PFA in 0.1 M PBS. After overnight cryoprotection in 30% sucrose solution in 0.1 M PBS, brains were blocked in the coronal plane, frozen on dry ice and sectioned at 50 µm on a cryostat. To reveal c-Fos expression levels in the brain, free-floating sections were incubated in 0.3% H2O2 in 0.1 M PBS for 10 min. After three rinses in 0.1 M PBS, sections were incubated in 0.1 M PBS containing 5% normal goat serum, 0.25% TritonX-100 and a c-Fos antibody raised in rabbit (Santa Cruz, sc-52; 1:800/1:500) or c-Fos antibody raised in rat (SySy, 226017; 1:1000) and left for overnight incubation (up to 96 h) at 4 °C. Sections were washed with 1xPBS and incubated at RT for 1 h in biotinylated goat anti-rabbit (#65-6140, Invitrogen; 1:400) and anti-rat-Ab secondary antibodies (# 31830, ThermoFisher Scientific, 1:400). Sections were washed three times with 0.1 M PBS and incubated at room temperature (RT) for one and a half of hour in biotinylated goat anti-rabbit (#65-6140, Invitrogen; 1:400) or anti-rat-Ab secondary antibodies (# 31830, ThermoFisher Scientific, 1:400). After three rinses with 0.1 M PBS, the sections were incubated at RT for 1,5 h in avidin-biotin peroxidase complex (Vectastain ABC, Vector Laboratories; 1:800). To visualize the peroxidase labeling, sections were processed with a DAB/nickel substrate working solution (DAB Peroxidase Substrate, SK-4100; Vector Laboratories) for 7 min at RT. After rinsing with 0.1 M PBS, sections were mounted on gelatin-coated slides, dehydrated, and put on coverslips.
Sections were imaged using a Leica bright-field microscope at 5x and 10x magnification. Several brain regions were selected for analysis: prefrontal cortex (PFC), primary motor cortex and somatosensory cortex, hippocampus (CA1, CA3, and dentate gyrus (DG)) and thalamus (Thal). Each brain region of interest was identified using a standard mouse brain atlas (Paxinos and Franklin). c-Fos immunoreactive nuclei were counted using predefined threshold values in ImageJ software. Labelled cells were counted bilaterally, averaged and normalized to the size of area and expressed as a relative compared to appropriate control.
Calcium imaging in developing brain slices
Calcium imaging of spontaneous synchronous activity (SSA) in the developing PFC- brain slices was performed according to previously published protocol by Dawitz et al. and Pires et al.. Briefly this method contains several steps: slice preparation, dye-loading slices, imaging and analysis. In total for the calcium imaging experiment, 9 controls, 8 Stxbp1 single-, 9 Snap25 single- and 8 double- mutants were used. Animals (P14 ± 1) were decapitated and brains were rapidly removed and placed in cold oxygenated aCSF (artificial Cerebrospinal Fluid) [31, 32]. PFC- brain slices were cut (300 μm thick) using Microm HM 650 V and transferred into a slice holder containing oxygenated recovery-aCSF (rACSF). After 1 h of recovery period, slices were transferred to a staining-chamber filled with 1 ml experimental ACSF (eACSF) and heated to 34 °C. Slices were incubated in eACSF containing Fura2-AM dye (ThermoFisher scientific, F1201) between 20 and 45 min depending on the age of the animal. After incubation, slices were transferred to coated recording chambers and approximately 1 ml of eACSF was added. Recording chambers with slices were kept in a large, humidified interface chamber to recover for at least one hour before recording. Network imaging was performed on a two-photon laser-scanning microscope (Trimscope LaVision Biotec). Slices were heated to 37 °C and constantly perfused with oxygenated eACSF. Using a Hamamatsu C9100 EM-CCD camera as a detector, two time-lapse movies (4000 frames each) in PFC-ROIs were acquired with a sampling frequency of 7.65 Hz (450µmX525µm, binning 2×2). The first was to evaluate baseline network activity and the second the effect of incubation with Gabazine (10 μM, Hellobio, SR95531) for 10 min. For the detection of the somas, a z-stack ± 20μm around the central plane with step size of 1μm thickness was acquired after each recording.
To analyze calcium-imaging data, custom-built Matlab® (Mathworks) scripts were used (EvA methodology) [33]. Neurons were semi-automatically detected using the z-stacks of the network being imaged. Finally, the events in individual traces were then analyzed individually and on a network level and three different categories of neurons were derived: silent neurons (those neurons where no activity was detected), active neurons and synchronized neurons (active neurons whose activity is synchronized with other neurons in network) [33]. Several different parameters were extracted and statistically analyzed: percentage of active cells, frequency of active cells, percentage of synchronously active cells and the frequency of synchronously active cells.
Cell culture and electrophysiology
Hippocampal microisland cultures were prepared from controls, Stxbp1 single-, Snap25 single-and double mutants at embryonic day 18. Briefly, hippocampi were isolated, collected in ice-cold Hank’s balanced salt solution (HBSS; Sigma, H9394) supplemented with 10 mM HEPES and digested with 0.25% trypsin (Life Technologies, 15090-046) at 37 °C for 20 min. After trituration, cells were resuspended in Neurobasal medium supplemented with B-27, 1% HEPES, 0.25% GlutaMAX, and 0.1% Penicillin-Streptomycin (all Invitrogen) and plated in a 6-well plate at a density of 7000 cells/well on top of pre-grown mouse glia islands on 30-mm coverslips (first experiment, see below) or in a 12-well plate at a density of 2500 cells/well on top of pre-grown rat glia islands on 18 mm coverslips (second experiment) to achieve autaptic cultures.
Two independent electrophysiological experiments were performed. In the first experiment, electrophysiological measurements were performed in autaptic cultures grown for 10–14 days in vitro as previously described in Ruiter et al. [34]. The recordings were performed at room temperature (RT) using an EPC10 amplifier (HEKA) with the recording program Patchmaster v2x73.5 (HEKA). Traces were filtered with a 3 kHz Bessel low-pass filter and data were acquired at 20 kHz. The series resistance was compensated to 70%. Borosilicate glass pipettes (resistance between 3 to 5 MΩ) were filled with 136 mM KCl, 17.8 mM HEPES, 1 mM EGTA, 4 mM ATP-Na, 4.6 mM MgCl2, 15 mM Creatine Phosphate and 50 U/mL phosphocreatine kinase (pH 7.4). External solution contained the following (in mM): 10 HEPES, 10 glucose, 140 NaCl, 2.4 KCl, 2 MgCl2 and 2 CaCl2 (pH = 7.4, 300 mOsmol). Recordings were done in voltage clamp, with the holding potential kept at −70 mV. Evoked excitatory postsynaptic currents (EPSC) were induced by raising the holding voltage to 0 mV for 2 ms. The size of the readily releasable pool (RRP) was assessed by hypertonic sucrose application (500 mM, 3.5 s). Sucrose was dissolved into the external solution. Application to the cells was done using a custom-made barrel system, controlled by SF-77B perfusion fast step (Warner Instruments) via digital output switches from the EPC10.
In the second experiment, electrophysiological measurements were performed in autaptic cultures grown for 13–16 days in vitro as explained in Meijer et al. [35]. Whole-cell voltage clamp recordings (Vm = −70 mV) were performed at RT using an Axopatch 200B amplifier, and Digidata 1440 A and Clampex 10 for signal acquisition (Molecular Devices). Action potentials were induced by stepping to + 30 mV for 1 ms. Data were acquired at 10 kHz with 2 kHz filtering with a low-pass Bessel filter. Series resistance was compensated by 70–80% (bandwidth 7.52 Hz). Borosilicate glass pipettes (resistance of 2–4.5 MΩ) were filled with (in mM): 136 KCl, 17.8 HEPES, 1 EGTA, 0.6 MgCl2x6H2O, 4 ATP-Mg, 0.3 GTP-Na, 12 phosphocreatine di-potassium-salt and 50 U/mL phosphocreatine kinase. External solution contained the following (in mM): 10 HEPES, 10 glucose, 140 NaCl, 2.4 KCl, 4 MgCl2, and 2 CaCl2 (pH = 7.30, 300 mOsmol). Traces were excluded when series resistance was higher than 15 MΩ or leak current was larger than 300pA. GABAergic traces were recognized by prolonged EPSC decays and excluded. Analysis was performed in MATLAB R2019a (Mathworks) using available scripts (viewEPSC, downloaded from Github user vhuson). The RRP was estimated by back extrapolation from the cumulative EPSC charge during a 40 Hz 100 action potential train as in Meijer et al..
Immunocytochemistry
Neurons were fixed on DIV10-DIV14. Coverslips were fixed at RT with 2% PFA diluted in culture medium (Neurobasal with 2% ml B-27, 1 M HEPES, 0.26% Glutamax, 14.3 mM β-mercaptoethanol, 20 U/ml penicillin, 20 μg/ml streptomycin) for 10 min followed by another fixation with 4% PFA diluted in PBS for 10 min. After washing with PBS, the neurons were permeabilized for 5 min at RT with a solution of 0.5% Triton X-100 diluted in PBS. The neurons were then blocked with a solution containing 4% goat serum and 0.1% Triton X-100 for 30 min at RT. The coverslips were then incubated for 2 h at RT with a blocking solution containing α-VGLUT antibody (1/1000; Guinea pig, Millipore AB5905) and α-MAP2 antibody (1/5000; chicken, Abcam Ab5392) or a solution containing α-VGAT antibody (1/500; Rabbit, Synaptic System 131002) and α-MAP2 antibody (1/200; chicken, Abcam Ab5392). Cells were washed with PBS, and then transferred for 1 h at RT to a blocking solution containing the secondary antibodies Alexa Fluor-568 α-Chicken (1/1000; goat, Invitrogen A11041) and Alexa fluor-647 α-Guinea pig (1/4000; goat, Invitrogen A21450) or a solution containing the secondary antibodies Alexa Fluor-568 α-Chicken (1/1000; goat, Invitrogen A11041) and Alexa fluor-488 α-Rabbit (1/5000; goat, Invitrogen A11008). Cells were washed with PBS before being mounted on glass slides with FluorSave (Calbiochem, 345789). Coverslips were examined on Leica SP8 inverted confocal microscope with 40x oil immersion Objective (NA = 1.4).
Survival rate assessment
The assessment of survival rate per genotype was performed in twelve E18-litters bred for electrophysiological experiments (caesarian section) and ten nests bred for behavioral experiments (natural birth). All pups from E18-nests were collected and genotyped before culturing and the results analyzed for the expected genotype ratio. In addition, genotyping results of ten nests bred for behavioral experiments were performed after weaning, at 3–4 weeks of age and the survival rate of animals was compared to those at E18. Following behavioral and/or video monitoring experiments, genotypes were conformed at the time of sacrifice.
Statistics
All statistical analyses were performed using IBM SPSS statistic 24 (IBM corporation, Armonk, NY, USA) and GraphPad Prism®, version 8.3.1. If the normality and homoscedasticity criteria were met, data were analyzed using parametric tests (t-test, ANOVA, repeated measure ANOVA). If the normality and homoscedasticity criteria were not met, nonparametric tests were performed (Mann Whitney U-test) or data were ln-transformed. There were no significant differences of behavioral measures between batches. Therefore, data from different batches were analyzed together. Outliers were removed from analysis using ROUT method, with Q set to 1%. Tukey’s posthoc tests were performed if ANOVA showed significant effect. The genotype effect in CognitionWall DL/RL task was assessed by performing log-rank test on two Kaplan Meier survival curves. An error probability lower than p < 0.05 was accepted as statistically significant through the study. For all given level of analysis of PhenoTyper spontaneous behavior data, statistical analysis was based on estimated false discovery rate (FDR), P-values were corrected by minimum positive FDR with a threshold set at 5%.
Electrophysiology data were analyzed using linear mixed-effects model (LMM) in R (version 4.4.2). Genotype was included as a fixed effect, while batch was modeled as a random effect to account for variability across experimental batches. For the merged dataset combining two laboratories, an additional random effect for lab ID was included to control for inter-laboratory variation.
All response variables were log-transformed in R prior to model fitting to better satisfy the assumption of normally distributed residuals. For datasets containing zeros, a constant of 0.01 nA (EPSC amplitude) or 0.01 Hz (for mEPSC frequency) was added before transformation. Models were fitted using the lmerTest package in R [36], which provides p-values for fixed effects using Satterthwaite’s degrees of freedom approximation.
The general model structure was as follows:
For lab-merged datasets, the model included an additional random effect:
The reference genotype group was the wildtype. Estimated changes in mutant groups were calculated by back-transforming the log-scale coefficients using the following formula:
where β is the estimated coefficient for the genotype effect on the log-transformed scale.
Model assumptions were assessed through visual inspection of Q-Q plots.
Results
Stxbp1/Snap25 double mutants show extremely diverse seizure phenotypes
Two presynaptic DEE genes, STXBP1 and SNAP25, were selected to test the consequences of gene-gene interactions, given their established functional relationship in SNARE-complex assembly. We generated three haploinsufficiency mouse models by heterozygous inactivation of 1) single Stxbp1 (‘Stxbp1 single mutants’), 2) Snap25 (‘Snap25 single mutants’) gene and 3) combined haploinsufficient inactivation of Stxbp1 and Snap25 (‘double mutants’) mice [23, 24]. These single mutants have been established to mimic the human situation (haploinsufficiency, construct validity) and have been characterized before, describing phenotypes in the same domains/directions as human symptoms (face validity), which can be reversed by medication prescribed to human patients [13]. Array-based genetic analysis of 11,000 SNP probes (miniMUGA [37], confirmed a homogeneous genomic background for all experimental groups (96.8% of SNPs consistent with C57BL/6J sub-strain, Supplementary Table 5 and Supplementary Fig. 1), with small contributions of 0.1% and 0.4% of flanking regions of 129 strain background around the deletion site, for the Stxbp1 and Snap25 locus, respectively, as reported before for Stxbp1 single mutants [13].
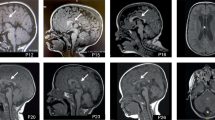
In E18 embryos, obtained by caesarian section for synapse physiology experiments (see below), all four genotypes were obtained, with a trend towards a reduced fraction for double mutants (18% vs expected Mendelian ratio of 25%, p = 0.093, χ2, labelled A in Fig. 1A). At the age of three/four weeks, when naturally born animals bred for behavioral experiments were weaned and genotyped, the distribution of genotypes significantly deviated from expected, with double mutants representing only 12% of animals, approximately half of the expected number (p = 0.041, χ2, Fig. 1A). Concomitantly, animal care takers reported lethal generalized seizures prior to and during weaning in these nests (Supplementary Video 1). Following the three/four weeks genotyping, sudden unexpected death of epilepsy (SUDEP) was detected in two additional double mutants older than 6 weeks. Together, this led to an even more significant deviation of the expected number of double mutants (10% instead of 25%, p = 0.031, labelled B in Fig. 1A). Further breeding of these double mutants was restricted by the local ethical committee regulations. Strikingly, surviving double mutants had no reported seizures, normal vision, normal muscle strength, normal motor coordination and ability to acquire motor skills, but 16% lower body weight compared to their controls (Supplementary Fig. 2 and Supplementary Table 1). These data suggest that at least half of double mutants died before the age of eight weeks due to severe seizures, while the surviving double mutants had no apparent phenotypes, except a lower body weight.
A Number of mice in litters with experimental time-line. The number of mice was calculated from 12 litters at E18 and 10 litters at age of three weeks (P3w). The number of mice per genotype in genotyped and weaned litters was counted again at the age of 6 weeks, after video monitoring. Exact number of mice per genotype in 12litters (E18) and 10 litters (P3w/P > 6w) is shown in pie-charts. † represents detected sudden unexpected death event in epilepsy (SUDEP). B Video monitoring of single-housed mice revealed different types of behavioral events: generalized seizure, clonic attack, jump, twitch, myoclonus and body extension. The aberrant behavioral events were analyzed in controls (n = 2), Stxbp1 single mutants (n = 5), Snap25 single mutants (n = 4) and double mutants (n = 6). C Average number of twitches and jumps detected in mice during 12 h of video monitoring. D Number of clonic attacks and generalized seizures detected in controls (n = 4), Stxbp1 single mutants (n = 4), Snap25 single mutants (n = 4) and double mutants (n = 7) during 12 h of video monitoring. E Representative ECoG trace of tonic-clonic generalized seizures detected in double mutant mouse with corresponding behavioral state pictogram. F Model of epileptic diversity represents the diversity of epileptic phenotype according to multiplicative model of interaction. We assume that most severe phenotype corresponds to lethal generalized seizures; moderate phenotype corresponds to clonic-tonic seizures and mild phenotype corresponds to low-incidence of SWDs. G 3D representation of different epileptiform events library. Event Classifier application in Neuroarchiver software distinguished three different types of epileptiform activities: SWDs, sharp spikes and spike-slow waves. H-J Number of different epileptiform events during 24 h in controls, Stxbp1 single-, Snap25 single- and double mutants represented as natural logarithm of (number of events +1). H Number of SWDs (I) Number of spike-slow waves and J Number of sharp spikes. K Representation of ECoG traces and corresponding time-frequency spectogram for control, Stxbp1 single-, Snap25 single- and double mutants. L Average relative power during the sleep episodes for all four genotypes. M Quantification of average relative power during the sleep in two frequency bands: delta band (1–4 Hz) and alfa band (9–12 Hz). Shift of relative power to higher frequency bands for all four genotypes is represented as the significant increase of power in alfa frequency band independent on the genotype. Panels represent mean value ± SEM.
Video observations of mice in their home cage confirmed frequent generalized and clonic seizures in some double mutant mice, but not others (Fig. 1B). The incidence of seizures varied substantially among double mutants (compare the two double mutant individuals in Fig. 1B). Postmortem genetic analyses produced no evidence for unexpected de novo genetic deficits in the most severely affected animals (Supplemental Table 5). Stxbp1 single mutants and double mutants showed twitches (n = 20.2 ± 2.6 / 12 h) and jumps (n = 7.4 ± 1.3 / 12 h, Fig. 1C), as reported earlier for Stxbp1 single mutants [12, 13]. The incidence of these two types of behaviors was lower in double mutants (n = 4.5 ± 1.9 and n = 1.0 + 0.4 for twitches and jumps per 12 h, respectively, Fig.1C). Except for these abnormalities, overall development of single Stxbp1 and Snap25 mice was normal, as reported before [13, 25, 38, 39]. Taken together, the reduced incidence of twitches and jumps and the occurrence of clonic, lethal seizures in a subset of double mutant mice suggests large phenotypic diversity regarding behavioral manifestation of epilepsy in double mutant mice.
Next, we combined video with electrocorticography (ECoG) monitoring at six to twelve weeks of age using implanted electrodes in 19 mice from new litters. These recordings confirmed generalized seizures in a subset of double mutant mice, but not others (Fig. 1D, E). The generalized seizures began with clusters of spike-slow waves (interictal spikes) accompanied with clonic seizures; the seizure progressed to a full generalized seizure and stopped with the postictal suppression and behavioral immobility (Fig. 1E). Strikingly, clonic and generalized seizures were observed only in a subset of double mutant mice, while other double mutant mice showed no generalized seizures during 24 h of recording (Fig. 1D). Hence, double mutant mice show extreme phenotype diversity in behavioral and electrographic abnormalities related to seizures, as opposed to the smaller, distinct variation in the single mutants (Fig. 1F).
Analysis of ECoG recordings revealed a collection of diverse patterns of epileptiform activity in several experimental groups: slow-wave discharges (SWDs), sharp spikes and spike-slow waves (Fig. 1G–J). SWDs were observed before in Stxbp1 single mutants [13] and typically accompanied by behavioral twitches [13] (Fig. 1B). The number of SWDs was higher in Stxbp1 single mutants compared to control, Snap25 single mutants and double mutants (F(3,15) = 4.501, p = 0.019, post hoc: p = 0.003, p = 0.015 and p = 0.039, respectively, Fig. 1H). The lower number of SWDs in double mutant mice compared to single Stxbp1 mice confirms the previous conclusion that a higher phenotypic diversity exists for epilepsy-related phenotypes in double mutant mice compared to single mutant and control mice.
The number of spike slow waves was significantly higher in double mutant mice compared to control mice, Snap25 single and Stxbp1 single mutants (F(3,15) = 4.913, p = 0.014, post hoc: p = 0.004, p = 0.020 and p = 0.026, respectively, Fig. 1I). When the spike-slow waves occurred during the awake state, they were accompanied by clonic seizures in double mutant mice. Sharp spikes were observed mainly during sleep and the number of sharp spikes did not differ between genotypes (Fig. 1J). Hence, more severe electrographic seizures were observed in double mutant mice compared to Stxbp1 single- and Snap25 single- mutants. Taken together, the diversity in epileptic phenotype observed in double mutant mice is characterized by higher probability of low incidence SWDs but also with exacerbated seizure activity represented with increased lethality, generalized seizures and increased number of spike-slow wave epileptiform discharges in a subset of double mutants, while others showed mild/no phenotypes.
Finally, power spectral analysis during the sleep episodes (Fig. 1K) revealed a shift in relative power towards higher frequency bands for all three genotypes. A decrease in the relative power in the delta band (F(3,4) = 3.846, p = 0.113) and an increase in the relative power in the alfa band (F(3,4) = 26.72, p = 0.0042) were observed for all three mutant groups, independent on the genotype (Fig. 1K–M).
c-Fos expression is increased in the thalamus of surviving double mutants
To corroborate the observed seizure activity, c-Fos immunoreactivity was used as a marker for excessive neuronal activity in the brain [40] (Fig. 2). Increased c-Fos expression was observed in cortical brain regions of Stxbp1 single mutants as shown before [13], in the surviving double mutants (p = 0.026) and a strong trend was observed in Snap25 single mutants (p = 0.058) (Fig. 2A, B and E). Furthermore, only in double mutants, c-Fos expression was significantly increased in the thalamus (Fig. 2C and E; p = 0.026) and a strong trend was observed in hippocampal regions (Fig. 2D and E; p = 0.058). Thus, c-Fos expression data confirms genetic interaction between the two genotypes in thalamus and hippocampus of the double mutants, but not in cortical regions, possibly because double mutants with most excessive cortical excitability had already died (see above) prior to c-Fos expression analysis.
A - D Representatives of c-Fos expression in controls and double mutants in the prefrontal cortex (PFC), cortex, thalamus (Thal) and hippocampus. E Quantification of c-Fos expression in 5 brain regions expressed as relative number of c-Fos positive cells in Stxbp1+/- single BL6 mutants (n = 5), Snap25 single mutants (n = 6) and double mutants (n = 10) compared to appropriate control.
Surviving double mutants show impaired cognition, like Stxbp1 mice
Different aspects of learning and memory were assessed in surviving animals of the four experimental groups. To assess associative learning and memory, the fear conditioning test was performed (Fig. 3A). After one pairing session between shock and tone, contextual memory was assessed by placing animals in the training context during the next day (Fig. 3B). Snap25 single mutants showed a similar percentage of freezing as controls (p = 0.639). Stxbp1 single mutants showed a significantly lower percentage of freezing in the training context compared to controls (p = 0.001, Fig. 3C), as shown before13 and compared to Snap25 single mutants (p = 0.010). Double mutants showed similar effects as Stxbp1 single mutants (p < 0.001 and p = 0.007, respectively, Fig. 3B, C). Supplementary Table 1 lists all statistical tests. Exposure of mice to the new context resulted in ~10% of time freezing in control mice and Snap25 single mutants due to general fear (Fig. 3B). Stxbp1 single mutants and double mutants showed significantly lower percentage of freezing compared to controls (p = 0.014 and p = 0.020, respectively, Fig. 3C and Supplementary Table 1) and Snap25 single mutants (p = 0.048 and p = 0.060, respectively, Fig. 3C). Cued memory was assessed by tone exposure of animals in a new context Fig. 3B. Stxbp1 single mutants and double mutants showed a significantly lower percentage of freezing compared to controls (p = 0.009 and p = 0.013 Fig. 4C and Supplementary Table 1) and Snap25 single mutants (p = 0.044 and p = 0.104, respectively, Fig. 3C). Taken together, fear conditioning experiments suggest a strong and similar impairment in the contextual and cued fear memory in Stxbp1 single mutants and surviving double mutants. No evidence was observed for stronger phenotypes in surviving double mutant mice than for single mutants.
A Fear conditioning test protocol. B Percentage of freezing duration per trial for control, Stxbp1 single-, Snap25 single- and double mutants. C Quantification of percentage of freezing time during exposure to the training context during the second day and exposure to the new context and tone in new context during the third day. Stxbp1 single- and double- mutants showed decreased percentage of freezing during those three trials. D Barnes maze test protocol. E Latency to find a platform during the acquisition phase was similar for all tested groups. F Hole visits in target octant during the first probe trial was significantly increased for Stxbp1 single- and double- mutants. G Latency to find target hole during reversal phase was significantly longer for double mutants during reversal phase learning. Panels represent mean value ± SEM.
A Statistical analysis of 90 parameters of spontaneous behavior separated into 6 categories: kinematics, activity, sheltering, habituation, dark-light index and phase transition. Three annuli show statistical analysis for double mutants, Stxbp1 single- and Snap25 single- mutants compared to their control groups, respectively from periphery to the center. The full list of parameters and statistical analysis can be found in Supplemental Tables 2–4. B-C Kinematic parameters: short movement number during the light and long movement number during the light were significantly decreased in Stxbp1 single- and double- mutants compared to control group. D Activity parameter: On shelter zone duration during the dark was significantly decreased in Stxbp1 single- and double- mutants compared to control groups. E Sheltering parameter: Long shelter visit number during light was significantly decreased in double mutants compared to Snap25 single mutants and controls. F Habituation: On shelter zone habituation ratio was significantly decreased in Stxbp1 single- and double- mutants compared to control group. Panels B - F represent mean value ± SEM.
The Barnes maze test was used to assess spatial learning, memory and reversal learning (Fig. 3D). During the acquisition phase, mice were trained to locate the escape hole and the time needed to escape the aversive environment was assessed. No significant effects of genotypes on the escape latency were observed, although latencies tended to be higher for Stxbp1 single mutants and double mutants compared to controls (F(3,208) = 2.613, p = 0.061, Fig. 3E and Supplementary Table 1). During the probe trial (to assess spatial memory one day after the last acquisition training), Stxbp1 single- and double- mutants showed a higher probability of hole visits in the target octant compared to controls and Snap25 single mutants (p = 0.042 and p = 0.033, respectively, Fig. 3F and Supplementary Table 1). Impaired behavioral flexibility was already described in Stxbp1 single mutants [13] and a similar phenotype was observed in surviving double mutant mice.
During the reversal phase, the location of the escape hole was changed to the opposite side of the maze. Similar to the trend observed during the acquisition phase, double mutants needed more time to find the new escape hole compared to controls and Snap25 single mutants (p = 0.036 and 0.011, respectively Fig. 3G) and Stxbp1 single mutants showed a trend towards longer escape latencies compared to controls and Snap25 single mutants (p = 0.010 and 0.057, respectively Fig. 3G). These data show that single Stxbp1 and surviving double mutants preserved the learned response stronger than single Snap25 mutants and control mice, suggesting impaired behavioral flexibility and deficits in the reversal learning in single Stxbp1 and double mutants. Again, no evidence was observed for stronger phenotypes in surviving double mutant mice compared to single mutants.
Attention and working memory were assessed using the spontaneous alteration task in the T maze (Supplementary Fig. 3A). This test is based on the natural tendency of mice to visit the previously not visited arm [41]. Double mutants showed a similar percentage of alterations as their controls (t(22) = −0.963, p = 0.346, Supplementary Fig. 3B and Supplementary Table 1), suggesting normal attention and hippocampal-dependent short-term memory.
To assess discrimination- and reversal learning, we performed a 4-day automated home-cage task, the CognitionWall test [28] Supplementary Fig. 3C and Supplementary Table 1. During the discrimination-learning phase (DL), animals should learn to earn food rewards by passing through the correct hole of the three holed CognitionWall placed inside the PhenoTyper. During the reversal-learning (RL) task animals should suppress previously learned response and learn that passing through the other hole is rewarded. During the discrimination and reversal learning tasks, all mice showed similar distribution of entries made to reach the criterion of 80% correct entries (p = 0.264 and p = 0.766, Supplementary Fig. 3D–G and Supplementary Table 1). Thus, the results from the Cognition Wall test suggest a normal discrimination and reversal learning in all four experimental groups.
Surviving double mutants show anxiety-related behaviors like Stxbp1 mice
The anxiety-related phenotypes in double mutant mice were tested using a classical anxiety-related paradigm, the elevated plus maze test (EPM). In this test, double mutants showed anxiety-related behaviors represented by significantly less time spent on the open arms (t(22) = 2.561, p = 0.018) and a lower percentage of visits to the open arms (t(22) = 3.216, p = 0.004) accompanied with a mild, but significant increase of total distance moved compared to their controls (t(22) = −2.813, p = 0.040), (Supplementary Fig. 4A–C and Supplementary Table 1). Double mutants did not show increased anxiety in the open field test and in the dark-light box test (Supplementary Fig. 4D–I and Supplementary Table 1). Taken together, these data show mild anxiety-related behavior accompanied with hyper-activity detected in the elevated plus maze test, comparable to Stxbp1 single mutant mice.
Surviving double mutants show spontaneous behaviors like Stxbp1 mice
Analysis of spontaneous behavior was performed with surviving animals in all experimental groups in the automated home-cage environment (PhenoTyper) enriched with a shelter [26]. Double mutants showed several behaviors that were altered to a similar extent as the single Stxbp1 mutants, especially in Kinematics (parameters 1–26), but also a few abnormalities in other spontaneous behaviors (Fig. 4A–F, Supplementary Table 2–4). Snap25 single mutants showed very few significantly altered (kinematic) behaviors (Fig. 4A). Taken together, spontaneous behavior analysis indicates that surviving double mutants showed no evidence for super-additivity, but only phenotypes like the strongest single mutant (Stxbp1).
GABA inhibition in developing cortex is abnormal in double mutants, like single Stxbp1 mice
Spontaneous Synchronous Activity (SSA) is essential for the correct development of neural circuits [42]. SSA has been recently characterized in the mPFC and it has been shown that GABA blockade at the end of the second postnatal week can partially restore the SSA, which is present at earlier times [32]. We measured SSA in the developing prefrontal cortex of all experimental groups at 2 weeks of age, by monitoring Ca2+-transients using two-photon calcium imaging in acute brain slices. The role of GABA in SSA was assessed by adding gabazine at concentrations that block both the phasic and tonic activity of GABAA receptors (Fig. 5A, B). On average, approximately 40% of all neurons had spontaneous Ca2+-transients with an overall frequency of 0.0063 Hz in all groups, independent of the genotype (percentage of active cells: F(3,90) = 1.630, p = 0.188 and frequency of active cells: F(3,90) = 1.577, p = 0.200, Supplementary Fig. 5A, B). The majority of active cells was synchronously active; the percentage of synchronously active cells and the frequency of their activity did not differ between genotypes during baseline recording (F(3,90) = 0.600, p = 0.617 and F(3,90) = 2.058, p = 0.111, respectively, Supplementary Fig. 5A, B). Blockade of GABAA receptors by gabazine did not significantly affect the percentage of active cells in any of the experimental groups (F(1,90) = 1.614, p = 0.207, Supplementary Fig. 5A) but did show an overall trend towards increased frequency of active cells (F(1,90) = 3.463, p = 0.067, Supplementary Fig. 5B and Supplementary Table 1). Gabazine affected the percentage of SSA-participating cells and the frequency of SSA: a significant increase of the percentage of SSA was observed for control and Snap25 single slices (p = 0.045 and p = 0.014, respectively Fig. 5C), but not for Stxbp1 single and double mutant slices (p = 0.145 and p = 0.827 Fig. 6D). The frequency of SSA in control slices was significantly increased after application of gabazine (p = 0.022, Fig. 5D). On the other hand, the application of gabazine did not affect the frequency of SSA in Stxbp1 single and double mutant slices. Thus, the frequency of SSA in brain slice from Stxbp1 single mutants was significantly lower than in brain slices from control and Snap25 single mutants after application of gabazine (p = 0.016 and p = 0.018 Fig. 5D). These data show that GABA inhibits SSA in brain slices from control and Snap25 single mutants but that this inhibitory effect of GABA was absent in brain slices from Stxbp1 single- and double- mutants.
A Pictorgram representation of experimental design. PFC-brain slices of mice, age postnatal day 14 (P14) were loaded with Fura-2 AM and imaged under two-photon microscope for 8 mins to assess baseline network activity. The effect of gabazine was assessed during the following 8 min of recording. B Contour map of Fura2-AM ester bulk-loaded cells and representative traces of cells in the PFC brain slices. Active neurons are shown in blue, silent neurons are shown in black and synchronized neurons are shown in red. Scale bar is 50 μm. C - D The percentage C and frequency D of SSA cells at the baseline and after application of gabazine in the PFC- brain slices from control, Stxbp1 single-, Snap25 single- and double- mutants (n = 24, n = 27, n = 25, n = 17, respectively). Panels C and D represent mean value ± SEM.
A Typical evoked responses (EPSC) in control-, Stxbp1 single-, Snap25- single and double- mutant glutamatergic neurons in 2 mM / 2 mM Ca2+/Mg2+ concentrations. B Normalized EPSC per week. C Representative traces of 500 mM sucrose responses. D Normalized readily releasable pool (RRP) of glutamatergic neurons calculated from hypertonic sucrose responses. E Typical inhibitory evoked response (IPSC) in control, Stxbp1 single-, Snap25 single- and double- mutants GABA-ergic neurons in 2 mM / 2 mM Ca2+/Mg2+ concentrations. F Normalized IPSC per week. G Representative traces of 500 mM sucrose responses in GABA-ergic neurons. H Normalized RRP in GABA-ergic neurons. I Normalized EPSC per week in control-, Stxbp1 single -, Snap25 single- and double- mutant glutamatergic neurons in 2 mM / 4 mM Ca2+/Mg2+ concentrations. J Normalized paired pulse ratio (PPR) per week per group for paired-pulse intervals of 20 ms, 50 ms and 100 ms. K Cumulative charge during train-stimulation (40 Hz, 100 pulses) with back-extrapolation line of last 20 pulses used for estimation of RRP. L Normalized synaptic rundown of EPSC during first ten pulses of 40 Hz train-stimulation.
Double mutants show reduced synaptic transmission like single Stxbp1 neurons
To assess effects of Stxbp1 and Snap25 gene interaction on basic synaptic function, we performed two independent electrophysiological experiments in single (autaptic) hippocampal neurons in culture. Experiments were performed over 5–9 independent experimental weeks. One set of experiments was carried out in the same laboratory using the same mouse lines used for the system- and behavioral analyses (Figs. 1–5), (Fig. 6I–L) in Amsterdam, and one set of experiments were carried out in a different laboratory, in Copenhagen (Fig. 6A–H). Both laboratories have maintained independent strains of Stxbp1 and Snap25 mice for >15 years. MiniMUGA analysis revealed the presence of 129-strain SNPs in all samples flanking the deletion sites for the two genes (Supplementary Table 5 and Supplementary Fig. 1). The dominant genetic background of mice used in Copenhagen was the C57BL/6JBomTac sub-strain (90% consistent SNPs, Supplementary Table 5 and Supplementary Fig. 2), while mice used in Amsterdam were original C57BL/6J sub-strain (96.8% consistent SNPs, Supplementary Table 5 and Supplementary Fig. 1). This slight difference in genomic background, in addition to environmental factors and subtle differences in experimental procedure between laboratories, may contribute to variation in electrophysiological phenotypes between the labs. Such differences – as well as random differences between preparations within the same laboratory – result in a complex data structure, where data are nested within the preparation and recording laboratory, which can lead to inflation of significance (Type I error) if not treated properly [43]. To account for the clustered/nested data structure, we analyzed data using a linear mixed-effects model (LMM), where the preparation and the laboratory were modeled by random factors. All data were log-transformed, to better satisfy the assumption of normally distributed residuals.
Both labs measured the amplitude of evoked EPSCs in glutamatergic neurons upon stimulation with a single action potential. When analyzed separately, the Copenhagen dataset showed that both Stxbp1 single mutants and Snap25/Stxbp1 double mutants significantly reduced EPSC amplitude (p = 0.0043 and p = 0.030, respectively, Supplementary Table 6). The Amsterdam dataset was somewhat smaller but also showed a trend toward reduced EPSC amplitude in Snap25/Stxbp1 double mutants (Fig. 6I). When combining the two datasets using a LMM, double heterozygotes and Stxbp1 single mutants exhibited a significant reduction (p = 0.008, and p = 0.020, respectively) in EPSC amplitude. The estimated reduction was 16% in Snap25 single mutants (non-significant), 30% in Stxbp1 mutants and 34% in Snap25/Stxbp1 double mutants (Supplementary Table 6).
Spontaneous (miniature) release events were analyzed in the Copenhagen laboratory and showed broadly changes that correlated with EPSC (Supplementary Figure 6A–C), with mEPSC frequencies significantly reduced in double mutant (−52%, p = 0.0054) and Stxbp1 mutants (−53%, p = 0.0027, Supplementary Table 6). mEPSC amplitudes were slightly reduced as well (5–10%, Supplementary Figure 6)
The decreased glutamatergic transmission can be caused by fewer glutamatergic synapses, decreased synaptic efficacy (release probability) and/or by decreased size of the readily releasable pool (RRP). To discriminate between these possibilities, we tested in vitro synapse formation. Morphological analysis revealed no consistent differences in the dendritic length, synaptic size and synaptic density between single Stxbp1 and double mutant VGLUT(+) neurons compared to control neurons (Supplementary Fig. 7 and Supplementary Table 1). The size of RRP was assessed by application of hypertonic sucrose (Fig. 6C, D) in the Copenhagen lab and was significantly decreased in single Stxbp1, Snap25 and double mutant neurons (Stxbp1:−44%, p < 0.001; Snap25: −32%, p = 0.031; double mutants: −35%, p = 0.0069, respectively, Fig. 6D). Taken together, these data suggest that the decreased glutamate release is mainly due to a decreased size of the RRP. We also note that there is no additive (or multiplicative) effect when combining the Snap25 and Stxbp1 mutants.
The cumulative charge released after 100 pulses at 40 Hz (measured in the Amsterdam laboratory) showed non-significant trend toward increased value in single Snap25 neurons compared to control neurons (74% increase, p = 0.072), while it did not differ in single Stxbp1 neurons and double mutant neurons compared to control neurons (Fig. 6K, Supplementary Table 6).
Synaptic transmission was also assessed in Stxbp1 single, Snap25 single and double mutant GABA-ergic neurons in the Copenhagen laboratory (Fig. 6E). No significant effect was observed for evoked IPSC, RRP size and mIPSC frequency and amplitude (Fig. 6F–H, Supplementary Fig. 6D–F and Supplementary Table 6), indicating impaired excitatory synaptic transmission, without affecting inhibitory synaptic function. These findings are consistent with data obtained from slice recordings in the somatosensory cortex of the same mouse line in Copenhagen [44]. However note that because fewer measurements were carried out in GABAergic neurons, the 95% Confidence Limits are quite wide (for the double mutant, IPSC-amplitudes were changed by −5% on average; 95% C.L: −42% to +55%; Supplementary Table 6); a moderate change could therefore be missed on statistical grounds.
Non-linear gene-gene interactions explain observed phenotypic diversity
To quantitatively explain the phenotypic variation in double mutants observed in this study and extrapolate these findings to the human situation, we defined three modes of gene-gene interaction to describe how phenotypic differences observed among individuals with a pathogenic variant in one gene depends on their genotypes at other loci. First, we assume that if two genes encode proteins that operate in the same pathway, a variant/mutant has a unique phenotypic spectrum characterized by a mean effect and a variance determined by environmental factors as well as background genetic diversity (Fig. 7A). If both variants affect the same pathway and are expressed in the same individual this would result in a phenotypic spectrum of the strongest single case, but not more severe or diverse than that (lack of additivity, Fig. 7, model B.1). In contrast, if two genetic variants from independent pathways lead to common disorders, the combination of variants results in summation of the phenotypic spectra (additive model, Fig. 7, model B.2.1 [45]). Finally, in cases where genes interact genetically or physically, genetic variation in two such genes may give rise to a multiplicative effect on the phenotypic spectra (‘epistasis’ or ‘super-additivity’ Fig. 7, model B.2.2, [46]). The last two scenarios can be distinguished by the different distributions and variances of their effects (phenotypes): if the effects of single gene variants follow a normal distribution (Fig. 7A), the additive model predicts that the effect of the combination of variants also follows a normal distribution, with a mean being the sum of the means of single variants and the variance being the sum of the variances of single variants. However, according to the multiplicative model, the distribution of the effect of the combination of two normally distributed variants is right-skewed, it approaches a lognormal distribution and has markedly larger variance compared to the additive model (Fig. 7). As a result of the increased variance, multiplicative interaction results both in more individuals with severe phenotypes, but also more individuals with mild phenotypes, compared to additive interactions (Fig. 7). Notably, this conclusion follows from simply combining the background variances (Fig. 7A) in a multiplicative model, without introducing new sources of variance. Hence, these quantitative models explain that the large phenotypic diversity in DEE-patients can be explained by multiplicative interactions between genes.
A Effects of pathogenic variation in two hypothetical genes (Α and Β) are represented as normally distributed severity of phenotypes that are shifted to the right relative to the normal (healthy) individuals, indicating pathogenicity. B Graphs from B.1 to B2.2. represent the predicted effects of the combination of variants in the two genes in one individual. B.1 Common pathway model: combinations of two variants in these genes are predicted to lead to phenotypes similar to the strongest single variant (αA or αB). The resulting distribution is similar, but not identical, to the distribution of the more severe of the two variants (αB); the distribution was identified by simulating pairs of normally distributed variables according to αA or αB, and selecting the maximal number. B.2.1 Additive interaction model: phenotypic severity in individuals with variants in two genes is represented as the summation of effects of single gene variants (αA + αB). If the effects of single gene variants follow a normal distribution (as represented in panel A), the effect of a combination of variants in two or more genes follows a normal distribution, according to an additive model of interaction. The vertical dashed line indicates 20% of most severe cases in an additive model with two hypothetical gene variants (αA and αB), called ‘moderate’ phenotype. The percentage of population with moderate phenotype according to each model is indicated. B.2.2 Multiplicative interaction model: phenotypic severity in individuals with variants in two genes is represented as the multiplication of the effects of single gene variants (αA * αB). If the effects of single gene variants follow the normal distribution, according to multiplicative model of interaction the effect of combination of the variants in more genes would be right-skewed, approaching lognormal distribution, with larger variance compared to additive model. The blank (non-filled) curves represent the effects of single gene variants, re-plotted from panel A. Vertical black dash-dash line is drawn to represent 20% of most severe cases of multiplicative model, called ‘severe’ phenotype. Vertical grey dash-dot-dash line is drawn to represent 10% of the least severe cases of multiplicative model, called ‘mild’ phenotype. The percentage of population with mild and severe phenotypes according to other models are represented within the respective graphs. Following parameters were used to generate a model: normal (mean = 0, s.d. = 0.1), variant in gene 1 (mean = 2, s.d. = 0.6), variant in gene 2 (mean = 4, s.d. = 1.4) on 1*106 random repetitions. Distribution fit was performed in MATLAB.
Discussion
In this study, we investigated how gene interactions influence phenotypic diversity and found strong evidence for multiplicative (epistatic) interaction between two SNAREopathy loci at the systems level: while single mutants showed mild phenotypes that were similar within each single mutant group, double mutants showed extreme diversity in phenotypes, ranging from lethal clonic and generalized seizures to mild phenotypes similar to the single mutants in the surviving double mutants. C-Fos staining showed a concomitant large variation in double mutants. However, at the synapse and network level we found no evidence for such interactions. At the behavioral/cognitive level, we could only study surviving animals (approximately half the population, the least severely affected half). These remaining double mutant animals showed phenotypes similar to the strongest of the two single mutants (Stxbp1 single mutants).
Epistatic interactions explain diversity in seizures and EEG-abnormalities
The incidence of (mild) seizure-like events among individual mice showed a normal distribution in Stxbp1 and Snap25 single mutants with more severe effects in Stxbp1 single mutants. This suggests that epileptic events in these mice are largely mediated by one factor (inactivation of one Stxbp1 or Snap25 allele, Fig. 7 model A), with Stxbp1 haploinsufficiency having a larger impact than Snap25 haploinsufficiency. Stxbp1 and Snap25 are known to work together to regulate neurotransmitter release and synaptic transmission [23, 24]. The extreme phenotypic diversity observed in double mutants is not consistent with the predicted phenotypic effects of variation in two genes acting in the same pathway (common pathway model, Fig. 7B1) and suggests multiplicative (epistatic) gene-gene interactions (Fig. 7, model B2.2.), and the involvement of distinct, interacting deficiencies caused by Stxbp1 and Snap25 haploinsufficiency.
Such distinct, interacting deficiencies may be explained in different, not mutually exclusive ways. First, although the two genes work together in neurotransmitter release, and also in the secretion of neuropeptides and neuromodulators [47, 48], haploinsufficiency may affect different aspects of these processes, in opposite direction or under different circumstances for the two genes. The current study did not reveal deficits in synaptic transmission for Snap25 haploinsufficiency in single neurons (Fig. 6), but previous studies have reported impairments in different aspects of synaptic transmission in Snap25 haploinsufficiency models which are distinct from the deficits for Stxbp1 haploinsufficiency detected in the present and the previous studies, (Fig. 6 and [39, 49,50,51]).
Second, Stxbp1 and Snap25 haploinsufficiency may have distinct, interacting effects on different populations of neurons and/or brain networks. Stxbp1 haploinsufficiency was shown to have different effects on hippocampal GABAergic and glutamatergic neurons (Fig. 6, [50]) or deficits specifically in GABAergic interneurons in the cortex [12], deficits in recruiting these interneurons [44] and glutamatergic inputs in the striatum [52]. On the other hand, the sporadic epileptiform events in single Snap25 mice were ascribed to increased calcium responsiveness of thalamic neurons and hyper-excitability of thalamo-cortical circuits [53, 54]. In the present study we found a trend towards an increase in c-Fos expression in the cortical regions of Snap25 single mutants and increased c-Fos expression in double mutants, confirming the cortical hyperexcitability. Interestingly, c-Fos expression was increased in the thalamus of double mutants, but not in the thalamus of both single mutants, suggesting that the thalamus is a critical brain region for genetic interaction. The idea that Stxbp1 and Snap25 haploinsufficiency may have distinct, interacting effects on different populations of neurons and/or brain networks is in line with the observation that multiplicative (epistatic) effects were not observed in individual neurons (Fig. 6) or networks (Fig. 5), but were pronounced at the system level (generalized seizures, only in the double mutants).
Third, in addition to their best characterized cellular functions in the regulated secretion of neurotransmitters and neuromodulators, Stxbp1 and Snap25 haploinsufficiency may affect other functions which may contribute to non-linear genetic interaction, especially during earlier developmental phases. For instance, the data from early postnatal network activity suggests deficits in the GABA shift in Stxbp1 mutants, but not Snap25 mutants (see also below). Furthermore, Snap25, but not Stxbp1 haploinsufficiency was reported to produce negative modulation of voltage-gated calcium channels [51, 55, 56] and both genes have crucial, not fully overlapping roles in neuronal viability [57].
Finally, epigenetic variation or stochastic processes during development may also contribute to interacting deficiencies detected on the system level. Taken together, we conclude that non-linear (multiplicative) genetic interaction of distinct aspects of regulated secretion, distinct neuronal populations/networks in the brain, with other cellular functions and/or epigenetic/stochastic effects, together explain the broad diversity in phenotypic manifestations in mutant mice.
No evidence for epistatic interactions in surviving double mutants
Across many behavioral domains, the behavioral phenotypes of single Stxbp1 mutants and surviving double mutants were similar: learning and memory, behavioral flexibility, anxiety and spontaneous behavior, while Snap25 single mutants behaved like control mice in all these domains. Single Stxbp1 mice and double mutant mice showed pronounced impairment in associative learning and memory and behavioral flexibility, lower body weight and normal motor coordination and muscle strength in line with previous studies in Stxbp1 haploinsufficient mice [12, 13]. Interestingly, subtle increases in activity in a habituated environment during the dark phase in single Stxbp1 mice was alleviated in double mutants. On the other hand, normal cognition, spontaneous behavior, motor coordination, muscle strength and anxiety found in single Snap25 mice are in line with previously reported lack of significant behavioral and cognitive impairments in Snap25+/- mice [25, 38]. Behavior was assessed in the surviving mice older than eight-weeks, after approximately 50% of double mutants have died (Fig. 1A). It is plausible that double mutants with most severe behavioral phenotypes were lost due to early lethality. However, the relationship between early lethality and later performance in behavioral tests of the surviving animals is unknown, precluding strong conclusions on which type of genetic interaction (Fig. 7) is most consistent with the behavioral data.
A similar conclusion can be reached for sleep phenotypes, with the only variation that phenotypes of the single mutants were similar. A shift of the spectral power to higher frequencies during sleep was found in Stxbp1 single, Snap25 single and double mutants. Hence, while for behavior, the phenotypic spectrum of Stxbp1 mutants is the strongest among the two single mutants, for sleep this spectrum is approximately the same for both single mutants and the double mutants behave similar to both single mutants.
A failing GABA-shift as a key determinant in Stxbp1 haploinsufficiency mice
The assessment of the spontaneous synchronous activity (SSA) in two-weeks-old mutants indicated that the normal shift from depolarizing to hyperpolarizing GABA action, observed in control mice and Snap25 single mutants, was not observed in Stxbp1 single- and double-mutants. SSA is important for the establishment and maturation of functional neural circuits [42, 58, 59] and the emergence of inhibitory GABA action is crucial for the termination of SSA [32]. GABA-shift abnormalities have been observed before in several other neurodevelopmental disorders, such as Fragile-X syndrome and Rett syndrome [60, 61]. The GABA-shift is mediated by several (external and internal) factors, including network activity and neuropeptide release [62]. Both processes depend on SNARE-dependent vesicle fusion. Therefore, it is plausible that initial dysregulation of network activity and/or neuropeptide release is a key determinant in Stxbp1 related disorders and potentially other SNAREopathies. Interestingly, while both Snap25 and Munc18/Stxbp1 are essential for neuropeptide release from dense core vesicles (DCV) in mature neurons [47, 48], immature Snap25 null mutant neurons showed substantial remaining DCV exocytosis, similar to immature wild type neurons, which was attributed to redundancy with other Qb/c SNAREs during early development [47]. This finding provides a plausible explanation for the fact that Snap25 single mutants showed a normal GABA-shift and multiplicative (epistatic) effects were not observed for spontaneous synchronous activity in slices. Taken together, our findings support the hypothesis that neurodevelopmental disorders including Stxbp1 related disorders are initiated at early developmental stages and involve abnormalities in GABA-shift and delayed cortical network development.
Genetic interaction effects may extrapolate to the human population
This study provides experimental evidence to conclude that non-linear genetic interactions can help explain symptom diversity (Fig. 7). Symptom diversity is a major, unexplained issue for SNAREopathies and other DEE and a major complication for the assessment of therapy success and the evaluation of new candidate therapies, e.g. in clinical trials. To establish proof of concept, the current study took a reductionistic approach, with a single ‘modifying gene’ of large effect size (heterozygous inactivation) interacting with a primary genetic deficit under conditions where other genomic variation and environmental factors are radically minimalized (inbred mouse lines and highly standardized laboratory conditions, respectively). In the human population, variation in these factors add complexity and the accumulation of two genetic variants of large effect size in single individuals is very rare. However, the clear evidence for genetic interaction in the current study can be extrapolated to the human population (Fig. 8). Interactions between a primary genetic deficit (pathogenic variant) and multiple (common) genetic variation is expected to produce similar effects on phenotypic diversity and may explain the high diversity observed among patients with mutations in the genes that work together. This idea is consistent with the concept of ‘genetic buffering’ [63]. According to the multiplicative model (Fig. 7B2.2), a fraction of individuals with pathogenic variants is only mildly affected, or even unaffected, due to the ‘buffering’ effect of certain other genetic variants in the genome. Furthermore, the conclusion that epilepsy and cognitive impairments are mediated by different cellular mechanisms explains the limited efficacy of antiepileptic drugs that act on one specific cellular mechanism and their inability to ameliorate developmental aspects of DEE. Finally, this study associates atypical GABA cortical network development with neurodevelopmental delay in SNAREopathy models, suggesting a novel predictive biomarker for future research, diagnostics and treatment design.
The model explains phenotypic diversity in the human population with one pathogenic variant and three common variants that interact genetically, physically with pathogenic variant. A Effects of three common gene variants (cv1, cv2 and cv3, green, blue and purple lines, respectively) and one pathogenic variant (pv, yellow curve) on the phenotypic severity are represented as normally distributed severity of phenotypes that are shifted to the right compared to the normal (healthy) individual (grey curve). Green, blue and purple curves represent the predicted effects of the multiplicative (pv*cv1, blue curve) and the additive (pv + cv2 and pv + cv3, green and purple curves, respectively) gene interactions between one common variant with pathogenic variant. B Distribution of the phenotypic severity in the human population is represented with pink curve. The pink curve shows the distribution of the phenotypic severity in population containing equal proportion of individuals with cv1, cv2 and cv3 interacting with pathogenic variant.
Data availability
Data are available: https://doi.org/10.34894/VOVDZQ (SciStore server @VU University).
References
Hamdan FF, Myers CT, Cossette P, Lemay P, Spiegelman D, Laporte AD, et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am J Hum Genet. 2017;101:664–85.
Scheffer IE, Berkovic S, Capovilla G, Connolly MB, French J, Guilhoto L, et al. ILAE classification of the epilepsies: Position paper of the ILAE Commission for Classification and Terminology. Epilepsia. 2017;58:512–21.
Carvill GL, Heavin SB, Yendle SC, McMahon JM, O’Roak BJ, Cook J, et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet. 2013;45:825–30.
Euro E-RESC, Epilepsy Phenome/Genome P, Epi KC. De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am J Hum Genet. 2014;95:360–70.
Heyne HO, Singh T, Stamberger H, Abou Jamra R, Caglayan H, Craiu D, et al. De novo variants in neurodevelopmental disorders with epilepsy. Nat Genet. 2018;50:1048–53.
McTague A, Howell KB, Cross JH, Kurian MA, Scheffer IE. The genetic landscape of the epileptic encephalopathies of infancy and childhood. Lancet Neurol. 2016;15:304–16.
Kullmann DM. Neurological channelopathies. Annu Rev Neurosci. 2010;33:151–72.
Verhage M, Sorensen JB. SNAREopathies: Diversity in mechanisms and symptoms. Neuron. 2020;107:22–37.
Vissers LE, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet. 2016;17:9–18.
McClellan J, King MC. Genetic heterogeneity in human disease. Cell. 2010;141:210–7.
Genç Ö, An J-Y, Fetter RD, Kulik Y, Zunino G, Sanders SJ, et al. Homeostatic plasticity fails at the intersection of autism-gene mutations and a novel class of common genetic modifiers. Elife. 2020;9:e55775.
Chen W, Cai ZL, Chao ES, Chen H, Longley CM, Hao S, et al. Stxbp1/Munc18-1 haploinsufficiency impairs inhibition and mediates key neurological features of STXBP1 encephalopathy. Elife. 2020;9:e48705.
Kovacevic J, Maroteaux G, Schut D, Loos M, Dubey M, Pitsch J, et al. Protein instability, haploinsufficiency, and cortical hyper-excitability underlie STXBP1 encephalopathy. Brain. 2018;141:1350–74.
Lammertse HCA, van Berkel AA, Iacomino M, Toonen RF, Striano P, Gambardella A, et al. Homozygous STXBP1 variant causes encephalopathy and gain-of-function in synaptic transmission. Brain. 2020;143:441–51.
Xian J, Parthasarathy S, McKeown S, Balagura G, Fitch E, Helbig K, et al. Assessing the landscape of STXBP1-related disorders in 534 individuals. Brain. 2021.
Kadkova A, Murach J, Ostergaard M, Malsam A, Malsam J, Lolicato F, et al. SNAP25 disease mutations change the energy landscape for synaptic exocytosis due to aberrant SNARE interactions. Elife. 2024;12:RP88619.
Stamberger H, Nikanorova M, Willemsen MH, Accorsi P, Angriman M, Baier H, et al. STXBP1 encephalopathy: A neurodevelopmental disorder including epilepsy. Neurology. 2016;86:954–62.
Balagura G, Xian J, Riva A, Marchese F, Ben Zeev B, Rios L, et al. Epilepsy course and developmental trajectories in STXBP1-DEE. Neurol Genet. 2022;8:e676.
López-Rivera JA, Pérez-Palma E, Symonds J, Lindy AS, McKnight DA, Leu C, et al. A catalogue of new incidence estimates of monogenic neurodevelopmental disorders caused by de novo variants. Brain. 2020;143:1099–105.
Abramov D, Guiberson NGL, Burré J. STXBP1 encephalopathies: Clinical spectrum, disease mechanisms, and therapeutic strategies. J Neurochem. 2021;157:165–78.
Phillips PC. Epistasis–the essential role of gene interactions in the structure and evolution of genetic systems. Nat Rev Genet. 2008;9:855–67.
Chou HH, Chiu HC, Delaney NF, Segre D, Marx CJ. Diminishing returns epistasis among beneficial mutations decelerates adaptation. Science. 2011;332:1190–2.
Verhage M, Maia AS, Plomp JJ, Brussaard AB, Heeroma JH, Vermeer H, et al. Synaptic assembly of the brain in the absence of neurotransmitter secretion. Science. 2000;287:864–9.
Washbourne P, Thompson PM, Carta M, Costa ET, Mathews JR, Lopez-Benditó G, et al. Genetic ablation of the t-SNARE SNAP-25 distinguishes mechanisms of neuroexocytosis. Nat Neurosci. 2002;5:19–26.
Corradini I, Donzelli A, Antonucci F, Welzl H, Loos M, Martucci R, et al. Epileptiform activity and cognitive deficits in SNAP-25(+/-) mice are normalized by antiepileptic drugs. Cereb Cortex. 2014;24:364–76.
Loos M, Koopmans B, Aarts E, Maroteaux G, van der Sluis S, Verhage M, et al. Sheltering behavior and locomotor activity in 11 genetically diverse common inbred mouse strains using home-cage monitoring. PLoS One. 2014;9:e108563.
Maroteaux G, Loos M, van der Sluis S, Koopmans B, Aarts E, van Gassen K, et al. High-throughput phenotyping of avoidance learning in mice discriminates different genotypes and identifies a novel gene. Genes Brain Behav. 2012;11:772–84.
Remmelink E, Smit AB, Verhage M, Loos M. Measuring discrimination- and reversal learning in mouse models within 4 days and without prior food deprivation. Learn Mem. 2016;23:660–7.
Wright S, Hashemi K, Stasiak L, Bartram J, Lang B, Vincent A, et al. Epileptogenic effects of NMDAR antibodies in a passive transfer mouse model. Brain. 2015;138:3159–67.
Bokil H, Andrews P, Kulkarni JE, Mehta S, Mitra PP. Chronux: a platform for analyzing neural signals. J Neurosci Methods. 2010;192:146–51.
Dawitz J, Kroon T, Hjorth JJ, Meredith RM Functional calcium imaging in developing cortical networks. J Vis Exp. 2011.
Pires J, Nelissen R, Mansvelder HD, Meredith RM. Spontaneous synchronous network activity in the neonatal development of mPFC in mice. Dev Neurobiol. 2021;81:207–25.
Hjorth JJ, Dawitz J, Kroon T, Pires J, Dassen VJ, Berkhout JA, et al. Detection of silent cells, synchronization and modulatory activity in developing cellular networks. Dev Neurobiol. 2016;76:357–74.
Ruiter M, Kadkova A, Scheutzow A, Malsam J, Sollner TH, Sorensen JB. An electrostatic energy barrier for SNARE-Dependent spontaneous and evoked synaptic transmission. Cell Rep. 2019;26:2340–52 e5.
Meijer M, Dorr B, Lammertse HC, Blithikioti C, van Weering JR, Toonen RF, et al. Tyrosine phosphorylation of Munc18-1 inhibits synaptic transmission by preventing SNARE assembly. EMBO J. 2018;37:300–20.
Kuznetsova A, Brockhoff PB, Christensen RHB. lmerTest package: Tests in linear mixed effects models. J Stat Softw. 2017;82:1–26.
Sigmon JS, Blanchard MW, Baric RS, Bell TA, Brennan J, Brockmann GA, et al. Content and performance of the MiniMUGA genotyping array: A new tool to improve rigor and reproducibility in mouse research. Genetics. 2020;216:905–30.
Monteggia LM, Lin PY, Adachi M, Kavalali ET. Behavioral analysis of SNAP-25 and Synaptobrevin-2 Haploinsufficiency in mice. Neuroscience. 2019;420:129–35.
Orock A, Logan S, Deak F. Munc18-1 haploinsufficiency impairs learning and memory by reduced synaptic vesicular release in a model of Ohtahara syndrome. Mol Cell Neurosci. 2018;88:33–42.
Herrera DG, Robertson HA. Activation of c-fos in the brain. Prog Neurobiol. 1996;50:83–107.
Deacon RM, Rawlins JN. T-maze alternation in the rodent. Nat Protoc. 2006;1:7–12.
Blankenship AG, Feller MB. Mechanisms underlying spontaneous patterned activity in developing neural circuits. Nat Rev Neurosci. 2010;11:18–29.
Aarts E, Verhage M, Veenvliet JV, Dolan CV, van der Sluis S. A solution to dependency: using multilevel analysis to accommodate nested data. Nat Neurosci. 2014;17:491–6.
Dos Santos AB, Larsen SD, Guo L, Barbagallo P, Montalant A, Verhage M, et al. Microcircuit failure in STXBP1 encephalopathy leads to hyperexcitability. Cell Rep Med. 2023;4:101308.
Fisher RA. The correlations between relatives on the supposition of Mendelian inheritance. Trans R Soc Edinb. 1918;52:399–433.
Mani R, St Onge RP, Hartman JLT, Giaever G, Roth FP. Defining genetic interaction. Proc Natl Acad Sci USA. 2008;105:3461–6.
Arora S, Saarloos I, Kooistra R, van de Bospoort R, Verhage M, Toonen RF. SNAP-25 gene family members differentially support secretory vesicle fusion. J Cell Sci. 2017;130:1877–89.
Puntman DC, Arora S, Farina M, Toonen RF, Verhage M. Munc18-1 is essential for neuropeptide secretion in neurons. J Neurosci. 2021;41:5980–93.
Patzke C, Han Y, Covy J, Yi F, Maxeiner S, Wernig M, et al. Analysis of conditional heterozygous STXBP1 mutations in human neurons. J Clin Invest. 2015;125:3560–71.
Toonen RF, Wierda K, Sons MS, de Wit H, Cornelisse LN, Brussaard A, et al. Munc18-1 expression levels control synapse recovery by regulating readily releasable pool size. Proc Natl Acad Sci USA. 2006;103:18332–7.
Antonucci F, Corradini I, Morini R, Fossati G, Menna E, Pozzi D, et al. Reduced SNAP-25 alters short-term plasticity at developing glutamatergic synapses. EMBO Rep. 2013;14:645–51.
Miyamoto H, Tatsukawa T, Shimohata A, Yamagata T, Suzuki T, Amano K, et al. Impaired cortico-striatal excitatory transmission triggers epilepsy. Nat Commun. 2019;10:1917.
Watanabe S, Yamamori S, Otsuka S, Saito M, Suzuki E, Kataoka M, et al. Epileptogenesis and epileptic maturation in phosphorylation site-specific SNAP-25 mutant mice. Epilepsy Res. 2015;115:30–44.
Zhang Y, Vilaythong AP, Yoshor D, Noebels JL. Elevated thalamic low-voltage-activated currents precede the onset of absence epilepsy in the SNAP25-deficient mouse mutant coloboma. J Neurosci. 2004;24:5239–48.
Condliffe SB, Corradini I, Pozzi D, Verderio C, Matteoli M. Endogenous SNAP-25 regulates native voltage-gated calcium channels in glutamatergic neurons. J Biol Chem. 2010;285:24968–76.
Toft-Bertelsen TL, Ziomkiewicz I, Houy S, Pinheiro PS, Sørensen JB. Regulation of Ca2+ channels by SNAP-25 via recruitment of syntaxin-1 from plasma membrane clusters. Mol Biol Cell. 2016;27:3329–41.
Santos TC, Wierda K, Broeke JH, Toonen RF, Verhage M. Early Golgi abnormalities and neurodegeneration upon loss of presynaptic proteins Munc18-1, Syntaxin-1, or SNAP-25. J Neurosci. 2017;37:4525–39.
Kirkby LA, Sack GS, Firl A, Feller MB. A role for correlated spontaneous activity in the assembly of neural circuits. Neuron. 2013;80:1129–44.
Ben-Ari Y. Excitatory actions of gaba during development: the nature of the nurture. Nat Rev Neurosci. 2002;3:728–39.
He Q, Nomura T, Xu J, Contractor A. The developmental switch in GABA polarity is delayed in fragile X mice. J Neurosci. 2014;34:446–50.
Tang X, Jaenisch R, Sur M. The role of GABAergic signalling in neurodevelopmental disorders. Nat Rev Neurosci. 2021;22:290–307.
Peerboom C, Wierenga CJ. The postnatal GABA shift: A developmental perspective. Neurosci Biobehav Rev. 2021;124:179–92.
Hartman JLT, Garvik B, Hartwell L. Principles for the buffering of genetic variation. Science. 2001;291:1001–4.
Acknowledgements
We thank Sita van der Wal, Natasja Bos and Christian van der Meer for breeding mutant mice. We thank Joost Hoetjes and Dorte Lauritsen for genotyping mutant mice. We thank Rolinka van der Loo for help with organizing perfusions and surgeries.
Funding
The work was supported by the Independent Research Fund Denmark (8020-00228A to JBS), the Lundbeck Foundation (R277-2018-802 to JBS and MV, and R389-2021-1596 to JBS), the Novo Nordisk Foundation (NNF10OC0058298 to JBS) and BRAINSCAPES (NWO 024.004.012 to MV).
Author information
Authors and Affiliations
Contributions
J.K. performed all behavioral experiments (Figs. 1, 3, 4), EEG recordings (Fig. 1), most of the cFOS stainings (Fig. 2), the initial modelling (Figs. 7, 8), analyzed the data and wrote the manuscript together with M.V. and J.B.S. and input from all authors; S.H. performed the 1st batch of synapse physiology and staining experiments in Copenhagen (Fig. 6, Fig. S6, Fig. S7) and analyzed the data; J.P. performed all calcium imaging experiments (Fig. 5) and analyzed the data; A. K. performed the 1st batch of synapse physiology and staining experiments in Copenhagen (Fig. 6, Fig. S6, Fig. S7) and analyzed the data; Z.L. performed the Linear Mixed-Effects model on synapse physiology data (Supplementary Table 6); H.C.A.L. performed the 2nd batch of synapse physiology experiments in Amsterdam (Fig. 6) and analyzed the data; J.S.M performed the 1st batch of synapse physiology and staining experiments in Copenhagen (Fig. 6, Fig. S6, Fig. S7) and analyzed the data; M.Ö. performed the 2nd batch of synapse physiology experiments in Amsterdam (Fig. 6) and analyzed the data; K.W. performed the 1st batch of synapse physiology experiments in Copenhagen (Fig. 6) and analyzed the data; J.W. supervised breeding, analyzed breeding results and performed some of the of the cFOS stainings; J.B.S designed the project, obtained the funding, participated in the final modelling (Figs. 7, 8), supervised S.H., A.K., Z.L.,J.S.M. and K.W. and wrote the manuscript together with J.K. and M.V.; M.V. designed the project, obtained the funding, supervised J.K., J.P., H.C.A.L.,M.Ö. and J.W. and wrote the manuscript together with J.K. and J.B.S. and input from all authors. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors report no competing interests. At the start of the experiments described in this manuscript, J.K. was a full-time employee of Sylics (Synaptologics BV), a private, VU University spin-off company that offers mouse phenotyping services, also on the mice described in this manuscript; during later phases, J.K. was employed by VU University. Sylics had no influence on the scientific decisions made by her and others involved in her work. During the time of this study, M.V. participated in a holding that owned Sylics shares and has received consulting fees from Sylics. InnoSer NV acquired Sylics in 2023. M.V. continued to receive consulting fees from InnoSer.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kovačević, J., Houy, S., Pires, J. et al. Phenotypic diversity is caused by non-linear genetic interactions between two SNAREopathy genes. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03509-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03509-3