Abstract
Objective
This study aims to evaluate the diagnostic value of machine learning-based MRI imaging in differentiating benign and malignant prostate cancer and detecting clinically significant prostate cancer (csPCa, defined as Gleason score ≥7) using systematic review and meta-analysis methods.
Methods
Electronic databases (PubMed, Web of Science, Cochrane Library, and Embase) were systematically searched for predictive studies using machine learning-based MRI imaging for prostate cancer diagnosis. Sensitivity, specificity, and area under the curve (AUC) were used to assess the diagnostic accuracy of machine learning-based MRI imaging for both benign/malignant prostate cancer and csPCa.
Results
A total of 12 studies met the inclusion criteria, with 3474 patients included in the meta-analysis. Machine learning-based MRI imaging demonstrated good diagnostic value for both benign/malignant prostate cancer and csPCa. The pooled sensitivity and specificity for diagnosing benign/malignant prostate cancer were 0.92 (95% CI: 0.83–0.97) and 0.90 (95% CI: 0.68–0.97), respectively, with a combined AUC of 0.96 (95% CI: 0.94–0.98). For csPCa diagnosis, the pooled sensitivity and specificity were 0.83 (95% CI: 0.77–0.87) and 0.73 (95% CI: 0.65–0.81), respectively, with a combined AUC of 0.86 (95% CI: 0.83–0.89).
Conclusion
Machine learning-based MRI imaging shows good diagnostic accuracy for both benign/malignant prostate cancer and csPCa. Further in-depth studies are needed to validate these findings.
Similar content being viewed by others
Introduction
Prostate cancer has become the second most common malignancy and the fifth leading cause of cancer-related death among men globally, with nearly 1.4 million new cases and 375,000 deaths annually [1]. In Western countries, the incidence of prostate cancer is higher due to early screening using prostate-specific antigen (PSA), and prostate cancer ranks first among urological malignancies in China [2, 3].
Current methods for early screening and diagnosis of prostate cancer include PSA testing, digital rectal examination, transrectal ultrasound, magnetic resonance imaging (MRI), and biopsy. While PSA is a key biomarker, its limited sensitivity/specificity, diagnostic gray zone (4–10 ng/ml), and susceptibility to confounding factors challenge its reliability [4, 5]. Digital rectal examination relies heavily on subjective tactile interpretation, increasing missed diagnoses, whereas transrectal ultrasound exhibits lower sensitivity than MRI for early-stage detection. MRI, despite radiologists’ experience-dependent accuracy, has emerged as the preferred imaging modality due to its superior diagnostic efficacy for cancer detection and grading [6].
The integration of artificial intelligence (AI) in medicine has significantly advanced prostate cancer diagnostics by enabling rapid data synthesis and automated analysis of imaging features. In recent years, many studies have used artificial neural network-based image texture analysis techniques to extract image texture features from prostate lesions, revealing the heterogeneity of prostate cancer tissue through the distribution and changes of these features [7,8,9]. Several studies have shown that combining MRI with machine learning significantly improves the diagnostic performance for prostate cancer [10, 11]. For example, Kwon et al. [12] used a random forest algorithm to construct a three-class model for prostate cancer lesion identification based on T2-weighted imaging (T2WI), diffusion-weighted imaging (DWI), apparent diffusion coefficient (ADC), and dynamic contrast-enhanced (DCE) sequences from 204 patients with 330 suspicious lesions, achieving an area under the curve (AUC) of 0.82. Zhang et al. [13] integrated support vector machine (SVM) with KNN and Zernik feature selection methods to discriminate prostate cancer lesions, training their model with MR images from 271 lesions and achieving an accuracy rate of over 80%. Ellmann et al. [14] used the XGBoost algorithm to analyze 195 prostate cancer lesions from 124 patients, obtaining a high accuracy rate (AUC = 0.913). These studies suggest that machine learning-based MRI imaging has significant diagnostic value and clinical application potential for prostate cancer diagnosis.
However, there is currently a lack of systematic evaluations of the diagnostic value of machine learning-based MRI imaging for prostate cancer. Therefore, we conducted this systematic review and meta-analysis to include studies assessing the diagnostic accuracy of machine learning-based MRI in detecting prostate cancer and csPCa (defined as Gleason score ≥7 vs. Gleason score <7), aiming to provide evidence-based insights for clinical practice.
Materials and methods
Search strategy
Following the PRISMA 2020 statement [15], we systematically searched four electronic databases (PubMed, Web of Science, Cochrane Library, and Embase) from inception to February 15, 2025. The search strategy in English databases was as follows: “Magnetic Resonance Imaging” OR “MRI” AND “Prostate” OR “Prostate Cancer” AND “Machine Learning” OR “Radiomics.” Keywords were connected using “OR” or “AND.” Additionally, target literature was obtained by reviewing the references of included studies.
Eligibility
Inclusion Criteria: (1) Populations (P): participants with prostate cancer; (2) Interventions (I): intervention involving MRI examination of the prostate in all participants; (3) Controls (C): healthy individuals or those without prostate cancer or other malignancies; (4) Outcomes (O): diagnostic results (benign/malignant and csPCa) presented in a 2 × 2 test performance table, with csPCa defined as Gleason score ≥7 [16]; (5) Study design (S): machine learning studies evaluating the diagnostic value of MRI for primary prostate cancer, published in peer-reviewed journals in Chinese or English.
Exclusion Criteria: (1) Non-human studies; (2) Conference articles, case reports, systematic reviews, etc.; (3) Insufficient outcome information for data analysis; (4) Duplicate reports; (5) Studies without full-text availability.
Literature screening and data extraction
Two researchers independently screened the literature according to the inclusion and exclusion criteria. Initial screening was performed by reading titles and abstracts, followed by full-text review for potentially eligible studies. Disagreements were resolved by consulting a third researcher. After literature screening, data extraction was performed independently by two researchers using a standardized data extraction form. Extracted information included study characteristics, demographic features of participants, true positives (TP), false positives (FP), true negatives (TN), false negatives (FN), and study duration.
Quality assessment
The quality of diagnostic accuracy studies was assessed using the QUADAS-2 tool developed by Whiting et al. at the University of York [17]. This tool evaluates studies in four domains: patient selection, index test, reference standard, and flow and timing. Risk of bias was categorized as “high risk,” “low risk,” or “unclear risk.”
Statistical analysis
Stata 16.0 software was used for statistical analysis. Pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), and their 95% confidence intervals (CIs) were calculated. Summary receiver operating characteristic (SROC) curves were plotted, and the AUC was calculated. Heterogeneity was assessed using Q-tests or I² statistics; I² < 50% or P > 0.1 indicated homogeneity, while I² > 50% or P ≤ 0.1 indicated significant heterogeneity. Publication bias was assessed using funnel plots, with P < 0.05 indicating statistical significance.
Results
Basic characteristics of included studies
A total of 1392 studies were identified through systematic database searches (Fig. 1). After excluding 928 duplicates and 247 irrelevant studies, 217 studies were reviewed in full. Ultimately, 12 studies were included in the meta-analysis [18,19,20,21,22,23,24,25,26,27,28,29].
Literature selection flowchart.
Table 1 presents the basic characteristics of the included studies. The 12 studies were published between 2019 and 2023. Five studies were from China, two from the United States, and the remaining studies were from the United Kingdom (n = 1), Canada (n = 1), Italy (n = 1), Germany (n = 1), and New Zealand (n = 1). Of the 12 studies, 11 were retrospective and one was prospective in design; 11 were single-center studies and one was a multicenter study. The data collection period ranged from 2008 to 2019. A total of 3474 patients were included in the meta-analysis, with ages ranging from 30 to 90 years. All diagnoses of prostate cancer were confirmed by histopathological examination. Among the 12 studies, five assessed the diagnostic value of machine learning-based MRI imaging for differentiating benign and malignant prostate cancer, while the remaining seven evaluated its diagnostic value for csPCa. Eleven studies used T2WI sequences for feature extraction and lesion segmentation, and seven used DWI sequences for feature extraction. Notably, all studies utilized at least two MRI sequences, highlighting the advantage of biparametric and multiparametric approaches in feature extraction and lesion identification for prostate cancer MRI imaging.
Quality assessment of included studies
The quality of the included studies was assessed using the QUADAS-2 tool. The results showed that the included studies had high quality with low potential bias (Figs. 2, 3).
Risk of bias and applicability concerns graph.
Risk of bias and applicability concerns summary.
Diagnostic value for benign/malignant prostate cancer
A total of five studies reported the diagnostic value of machine learning-based MRI imaging for benign/malignant prostate cancer, involving 1398 participants. The meta-analysis showed that the pooled sensitivity and specificity for diagnosing benign/malignant prostate cancer were 0.92 (95% CI: 0.83–0.97; Fig. 4) and 0.90 (95% CI: 0.68–0.97; Fig. 4), respectively, with a combined AUC of 0.96 (95% CI: 0.94–0.98; Fig. 5). The heterogeneity was 95.68% and 97.52%, respectively. Additionally, the pooled PLR and NLR were 9.24 (95% CI: 2.36–36.12; Supplementary Fig. 1) and 0.08 (95% CI: 0.03-0.21; Supplementary Fig. 1), respectively.
Summary of sensitivity and specificity of machine learning-based MRI in PCa diagnosis.
Summary of AUC of machine learning-based MRI in PCa diagnosis.
Diagnostic value for csPCa
A total of seven studies reported the diagnostic value of machine learning-based MRI imaging for csPCa, involving 2076 participants. The meta-analysis showed that the pooled sensitivity and specificity for diagnosing csPCa were 0.83 (95% CI: 0.77–0.87; Fig. 6) and 0.73 (95% CI: 0.65–0.81; Fig. 6), respectively, with a combined AUC of 0.86 (95% CI: 0.83–0.89; Fig. 7). The heterogeneity was 73.96% and 87.02%, respectively. Additionally, the pooled PLR and NLR were 3.12 (95% CI: 2.34–4.15; Supplementary Fig. 2) and 0.23 (95% CI: 0.18-0.30; Supplementary Fig. 2), respectively.
Summary of sensitivity and specificity of machine learning-based MRI in csPCa diagnosis.
Summary of AUC of machine learning-based MRI in csPCa diagnosis.
Publication bias
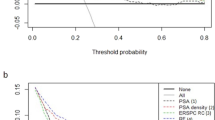
Publication bias was assessed for studies on the diagnosis of both benign/malignant prostate cancer and csPCa using machine learning-based MRI imaging. The results showed no significant publication bias among the included studies, with all P values greater than 0.05 (Supplementary Figs. 3, 4).
Discussion
In recent years, machine learning-based medical imaging analysis has shown great potential in prostate cancer diagnosis, but its clinical application still requires higher levels of evidence for validation. This study, through a systematic review and meta-analysis, comprehensively evaluated the diagnostic performance of machine learning models based on multi-modal MRI for differentiating benign and malignant prostate cancer and detecting csPCa for the first time. A total of 12 studies were included, involving 3474 patients, with five studies focusing on tumor benign/malignant diagnosis and seven studies on csPCa diagnostic accuracy. The results showed that machine learning models performed exceptionally well in prostate cancer benign/malignant diagnosis (combined AUC = 0.96), while the diagnostic efficacy for csPCa was relatively lower (combined AUC = 0.86). These findings provide important evidence for the application of machine learning in prostate cancer diagnosis and also reveal the limitations of current studies and future directions for improvement.
Our study found that machine learning-based MRI models demonstrated extremely high diagnostic performance in differentiating benign and malignant prostate cancer (sensitivity: 0.92, specificity: 0.90, AUC = 0.96), significantly outperforming the traditional PI-RADS scoring system, which has an AUC of approximately 0.80 [30]. This improvement may be attributed to the deep integration of multi-sequence MRI features (such as T2WI and DWI) by machine learning. For example, T2WI can clearly display the anatomical structure of the prostate, while DWI quantifies tissue cell density through the ADC. The combination of these two sequences can more comprehensively capture tumor heterogeneity [31, 32]. Despite the clinically meaningful AUC for csPCa diagnosis, the relatively low specificity may reflect several issues. First, tumors with Gleason scores ≥7 exhibit diverse MRI appearances, especially intermediate-grade cancers (Gleason 3 + 4), which overlap with benign hyperplasia or inflammation on imaging [33]. Moreover, only 11 studies utilized T2WI, and 7 studies employed DWI. The application of dynamic contrast-enhanced MRI (DCE) or emerging technologies, such as PSMA-PET fusion, was limited. This scarcity may restrict the model’s ability to identify microinvasive lesions. Additionally, nine studies relied on manual segmentation, which may introduce interobserver variability. Automated segmentation, used in only three studies, requires further validation. These factors could also be potential sources of the heterogeneity identified in this study.
The widespread application of artificial intelligence in clinical practice still faces the core challenge of insufficient generalizability, primarily due to the heterogeneous nature of medical data [34]. Variations in acquisition parameters, diverse imaging equipment, and the absence of standardized protocols for cross-institutional medical imaging data often lead to a decline in diagnostic performance when artificial intelligence models are validated across centers. This data shift, triggered by non-pathological features, has a particularly significant impact on radiomics-based machine learning models. A study by Bleker et al. [34] found that a radiomics-based biparametric MRI artificial intelligence model trained on single-center imaging data showed a substantial performance drop when validated on multicenter data (−28.1%, with AUC values of 0.594 vs. 0.816 for single-center validation, p = 0.002). In contrast, models trained on multicenter data did not experience significant performance degradation between multicenter and single-center validation (−12%, AUC: 0.750 vs. 0.660, p = 0.114) and even outperformed single-center models on multicenter data (AUC: 0.750 vs. 0.594, p = 0.012). Notably, several studies using both internal and external datasets for validation have demonstrated that models built on deep learning frameworks exhibit stronger data adaptability. These models maintain relatively stable diagnostic performance in both internal validation and external testing, suggesting that deep neural networks have unique advantages in autonomous feature extraction and adaptation to data distributions [35,36,37]. To optimize the generalizability of models in multicenter studies, the academic community has proposed using batch effect correction techniques to standardize cross-center data. This approach can effectively eliminate systematic biases caused by differences in scanning protocols and non-uniform feature standardization processes, thereby enhancing model robustness [38].
This study is the first to systematically analyze the diagnostic value of machine learning-based MRI for differentiating benign and malignant prostate cancer and detecting clinically significant prostate cancer (csPCa). However, several limitations exist. First, despite our systematic search strategy to identify eligible studies, the number of studies meeting the inclusion criteria was relatively low, likely due to the limited number of current studies using radiomics and machine learning for prostate cancer diagnosis. Additionally, most included studies were single-center and retrospective in design, which may introduce recall and confounding biases. This study’s findings should be interpreted with caution due to the inherent heterogeneity in machine learning methodologies across the included studies. Variations in algorithm architectures, preprocessing pipelines, and validation protocols may limit the generalizability of pooled results. Future work should prioritize harmonized reporting standards to mitigate algorithmic diversity and improve clinical translation. Moreover, due to the limited number of included studies, this meta-analysis was unable to explore subgroup analyses based on different variables.
Conclusion
In summary, this study demonstrates that artificial intelligence models built on MRI imaging data hold great potential and value for diagnosing prostate cancer. Although the included studies employed diverse methodologies, extracted MRI data from different centers, and utilized various algorithms to construct artificial intelligence models, our meta-analysis indicates that artificial models based on MRI imaging significantly outperform radiologists using the PI-RADS scoring criteria in diagnosing prostate cancer. However, the evidence presented by these studies should be interpreted with caution, as there is still a lack of multicenter evidence for artificial intelligence algorithms.
Data availability
All data generated or analysed during this study are included in this article. Further enquiries can be directed to the corresponding author.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Cao W, Chen HD, Yu YW, Li N, Chen WQ. Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020. Chin Med J (Engl). 2021;134:783–91.
Qi D, Wu C, Liu F, Gu K, Shi Z, Lin X, et al. Trends of prostate cancer incidence and mortality in Shanghai, China from 1973 to 2009. Prostate. 2015;75:1662–8.
Jamaspishvili T, Kral M, Khomeriki I, Student V, Kolar Z, Bouchal J. Urine markers in monitoring for prostate cancer. Prostate Cancer Prostatic Dis. 2010;13:12–9.
Rehman I, Evans CA, Glen A, Cross SS, Eaton CL, Down J, et al. iTRAQ identification of candidate serum biomarkers associated with metastatic progression of human prostate cancer. PLoS One. 2012;7:e30885.
Stabile A, Giganti F, Rosenkrantz AB, Taneja SS, Villeirs G, Gill IS, et al. Multiparametric MRI for prostate cancer diagnosis: current status and future directions. Nat Rev Urol. 2020;17:41–61.
Summers RM. Texture analysis in radiology: does the emperor have no clothes?. Abdom Radio (NY). 2017;42:342–5.
Avanzo M, Stancanello J, El Naqa I. Beyond imaging: the promise of radiomics. Phys Med. 2017;38:122–39.
Algohary A, Viswanath S, Shiradkar R, Ghose S, Pahwa S, Moses D, et al. Radiomic features on MRI enable risk categorization of prostate cancer patients on active surveillance: Preliminary findings. J Magn Reson Imaging. 2018: https://doi.org/10.1002/jmri.25983.
Liu L, Tian Z, Zhang Z, Fei B. Computer-aided detection of prostate cancer with MRI: technology and applications. Acad Radio. 2016;23:1024–46.
Wang S, Burtt K, Turkbey B, Choyke P, Summers RM. Computer aided-diagnosis of prostate cancer on multiparametric MRI: a technical review of current research. Biomed Res Int. 2014;2014:789561.
Kwon D, Reis IM, Breto AL, Tschudi Y, Gautney N, Zavala-Romero O, et al. Classification of suspicious lesions on prostate multiparametric MRI using machine learning. J Med Imaging (Bellingham). 2018;5:034502.
Zhang L, Li LY, Tang M, Huan Y, Zhang XL, Zhe X. A new approach to diagnosing prostate cancer through magnetic resonance imaging. Alex Eng J. 2021;60:897–904.
Ellmann S, Schlicht M, Dietzel M, Janka R, Hammon M, Saake M, et al. Computer-aided diagnosis in multiparametric MRI of the prostate: an open-access online tool for lesion classification with high accuracy. Cancers (Basel). 2020;12:2366.
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71.
Mehralivand S, Shih JH, Rais-Bahrami S, Oto A, Bednarova S, Nix JW, et al. A magnetic resonance imaging-based prediction model for prostate biopsy risk stratification. JAMA Oncol. 2018;4:678–85.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155:529–36.
Chen T, Li M, Gu Y, Zhang Y, Yang S, Wei C, et al. Prostate cancer differentiation and aggressiveness: assessment with a radiomic-based model vs. PI-RADS v2. J Magn Reson Imaging. 2019;49:875–84.
Wu M, Krishna S, Thornhill RE, Flood TA, McInnes MDF, Schieda N. Transition zone prostate cancer: Logistic regression and machine-learning models of quantitative ADC, shape and texture features are highly accurate for diagnosis. J Magn Reson Imaging. 2019;50:940–50.
Varghese B, Chen F, Hwang D, Palmer SL, De Castro Abreu AL, Ukimura O, et al. Objective risk stratification of prostate cancer using machine learning and radiomics applied to multiparametric magnetic resonance images. Sci Rep. 2019;9:1570.
Xu M, Fang M, Zou J, Yang S, Yu D, Zhong L, et al. Using biparametric MRI radiomics signature to differentiate between benign and malignant prostate lesions. Eur J Radio. 2019;114:38–44.
Min X, Li M, Dong D, Feng Z, Zhang P, Ke Z, et al. Multi-parametric MRI-based radiomics signature for discriminating between clinically significant and insignificant prostate cancer: Cross-validation of a machine learning method. Eur J Radio. 2019;115:16–21.
Antonelli M, Johnston EW, Dikaios N, Cheung KK, Sidhu HS, Appayya MB, et al. Machine learning classifiers can predict Gleason pattern 4 prostate cancer with greater accuracy than experienced radiologists. Eur Radio. 2019;29:4754–64.
Bleker J, Kwee TC, Dierckx RAJO, de Jong IJ, Huisman H, Yakar D. Multiparametric MRI and auto-fixed volume of interest-based radiomics signature for clinically significant peripheral zone prostate cancer. Eur Radio. 2020;30:1313–24.
Gong L, Xu M, Fang M, Zou J, Yang S, Yu X, et al. Noninvasive prediction of high-grade prostate cancer via biparametric MRI radiomics. J Magn Reson Imaging. 2020;52:1102–9.
Mehralivand S, Harmon SA, Shih JH, Smith CP, Lay N, Argun B, et al. Multicenter multireader evaluation of an artificial intelligence-based attention mapping system for the detection of prostate cancer with multiparametric MRI. AJR Am J Roentgenol. 2020;215:903–12.
He D, Wang X, Fu C, Wei X, Bao J, Ji X, et al. MRI-based radiomics models to assess prostate cancer, extracapsular extension and positive surgical margins. Cancer Imaging. 2021;21:46.
Bertelli E, Mercatelli L, Marzi C, Pachetti E, Baccini M, Barucci A, et al. Machine and deep learning prediction of prostate cancer aggressiveness using multiparametric MRI. Front Oncol. 2022;11:802964.
Hamm CA, Baumgärtner GL, Biessmann F, Beetz NL, Hartenstein A, Savic LJ, et al. Interactive explainable deep learning model informs prostate cancer diagnosis at MRI. Radiology. 2023;307:e222276.
Wang Z, Lu P, Liu S, Fu C, Ye Y, Yu C, et al. Comparison of the impact of rectal susceptibility artifacts in prostate magnetic resonance imaging on subjective evaluation and deep learning: a two-center retrospective study. BMC Med Imaging. 2025;25:61.
Turkbey B, Rosenkrantz AB, Haider MA, Padhani AR, Villeirs G, Macura KJ, et al. Prostate Imaging Reporting and Data System Version 2.1: 2019 Update of Prostate Imaging Reporting and Data System Version 2. Eur Urol. 2019;76:340–51.
Palumbo P, Manetta R, Izzo A, Bruno F, Arrigoni F, De Filippo M, et al. Biparametric (bp) and multiparametric (mp) magnetic resonance imaging (MRI) approach to prostate cancer disease: a narrative review of current debate on dynamic contrast enhancement. Gland Surg. 2020;9:2235–47.
Schoots IG, Haider MA, Punwani S, Padhani AR. MRI in prostate cancer screening: a review and recommendations, from the AJR special series on screening. AJR Am J Roentgenol. 2025;https://doi.org/10.2214/AJR.24.32588.
Bleker J, Yakar D, van Noort B, Rouw D, de Jong IJ, Dierckx RAJO, et al. Single-center versus multi-center biparametric MRI radiomics approach for clinically significant peripheral zone prostate cancer. Insights Imaging. 2021;12:150.
Hiremath A, Shiradkar R, Fu P, Mahran A, Rastinehad AR, Tewari A, et al. An integrated nomogram combining deep learning, prostate imaging-reporting and data system (PI-RADS) scoring, and clinical variables for identification of clinically significant prostate cancer on biparametric MRI: a retrospective multicentre study. Lancet Digit Health. 2021;3:e445–e454.
Netzer N, Weißer C, Schelb P, Wang X, Qin X, Görtz M, et al. Fully automatic deep learning in bi-institutional prostate magnetic resonance imaging: effects of cohort size and heterogeneity. Invest Radio. 2021;56:799–808.
Saha A, Hosseinzadeh M, Huisman H. End-to-end prostate cancer detection in bpMRI via 3D CNNs: effects of attention mechanisms, clinical priori and decoupled false positive reduction. Med Image Anal. 2021;73:102155.
Dereziński P, Klupczynska A, Sawicki W, Pałka JA, Kokot ZJ. Amino acid profiles of serum and urine in search for prostate cancer biomarkers: a pilot study. Int J Med Sci. 2017;14:1–12.
Funding
The project was supported by the Health Science and Technology Project of the Lianyungang Health Commission (202325); the Scientific Research Project of the Lianyungang Anti-Cancer Association (MS202312); Science and Technology Plan Project of Lianyungang (No. SF2311); Aging Health Research Project of Lianyungang (No. L202318); Health Science and Technology Project of Lianyungang (No. 202219); Medical Education Collaborative Innovation Fund of Jiangsu University (No. JDYY2023087). Funding agencies did not play a role in study design, data collection, analysis and interpretation, and manuscript writing.
Author information
Authors and Affiliations
Contributions
JXW and YS conceived of the study, and SBZ, JJL, KMS, DY, QZL, TZ, LX, and LG participated in its design and data analysis and statistics and YSZ and LZ helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
An ethics statement is not applicable because this study is based exclusively on published literature.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, Y., Zhang, L., Zhang, S. et al. Machine learning-based MRI imaging for prostate cancer diagnosis: systematic review and meta-analysis. Prostate Cancer Prostatic Dis 29, 159–166 (2026). https://doi.org/10.1038/s41391-025-00997-2
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41391-025-00997-2
This article is cited by
-
UroFusion-X: a unified multimodal deep learning framework for robust diagnosis, subtyping, and prognosis of urological cancers
npj Digital Medicine (2026)
-
Best of 2025 in prostate cancer and prostatic diseases
Prostate Cancer and Prostatic Diseases (2026)
-
Inter- and intra-rater variability of MRI-based lesion size measurements in active surveillance for prostate cancer: a multicentre study
European Radiology (2026)
-
A modified deep learning approach for seminal vesicle region localization in prostate MRI
Scientific Reports (2025)
-
AI-aided diagnostic performance for prostate MRI: systematic review and meta-analysis
Prostate Cancer and Prostatic Diseases (2025)