Abstract
Liver metastasis is the primary cause of mortality in colorectal cancer (CRC) patients. To decipher the underlying mechanisms, we performed single-cell RNA sequencing (scRNA-seq) on paired primary colorectal tumors, adjacent tissues and liver metastases from three CRC liver metastasis (CRLM) patients, alongside colorectal tumors and adjacent tissues from three non-metastatic CRC patients. Our analysis revealed a significant enrichment of Enolase 2-expressing (ENO2⁺) cancer cells in CRLM patients compared to their non-metastatic counterparts. Functional characterization, supported by bioinformatics and murine models, demonstrated that ENO2⁺ cancer cells exhibit enhanced epithelial-mesenchymal transition (EMT) and are critical drivers of CRLM. Mechanistically, the ENO2 protein directly binds to macrophage migration inhibitory factor (MIF) within cancer cells, stabilizing MIF by inhibiting its C-terminus of Hsc70-Interacting Protein (CHIP)-mediated ubiquitination and degradation. This ENO2-MIF interaction activates MIF signaling, fostering robust tumor cell-macrophage crosstalk that promotes M2 macrophage polarization, which is validated by spatial transcriptomics showing the colocalization of ENO2⁺ cancer cells and M2 macrophages. Crucially, both organoid and in vivo models confirmed that ENO2 in CRC cells is essential for inducing M2 macrophage polarization via the MIF pathway, thereby facilitating liver metastasis. Knockout of ENO2 significantly suppressed tumor growth and liver metastasis in mouse models. An inhibitor of the ENO2-MIF interaction, pyrithioxin, can effectively reduce the burden of liver metastasis in mice. Collectively, our findings identify ENO2 as a key driver of CRLM by stabilizing MIF to orchestrate M2 macrophage polarization, highlighting the ENO2-MIF axis as a promising therapeutic strategy for CRLM.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) ranks as the third most prevalent malignancy worldwide, with over 1.9 million new cases annually and mortality predominantly driven by distant metastasis.1 Among metastatic sites, the liver is the most common and lethal destination, accounting for >70% of CRC-related deaths.2 Current first-line systemic therapies, including anti-epidermal growth factor receptor and anti-vascular endothelial growth factor targeted agents, have yielded limited survival benefits in colorectal liver metastasis (CRLM) patients, especially for the vast majority with microsatellite stable (MSS) CRC, which exhibits primary resistance to single-agent immune checkpoint blockade.3 Despite multimodal advances, including surgical resection, systemic chemotherapy, and emerging immunotherapies, the 5-year survival for CRLM remains below 20%.4,5 This grim prognosis underscores an urgent need to decipher the molecular drivers of hepatic tropism and immune evasion, key biological features enabling metastatic outgrowth.
The liver metastatic niche represents a dynamically remodeled ecosystem where crosstalk between disseminated cancer cells and stromal components dictates metastatic efficiency.6,7 Tumor-associated macrophages (TAMs), particularly the immunosuppressive M2-polarized subset, are critical architects of this pro-metastatic milieu.8,9 They facilitate angiogenesis, extracellular matrix remodeling, and T-cell dysfunction through cytokine networks (e.g., interleukin-10, transforming growth factor-beta) and checkpoint ligand expression.10 Preclinical studies have shown that targeting TAM repolarization or depletion effectively inhibits CRLM, yet clinical translation of macrophage-targeted therapies (e.g., colony-stimulating factor 1 receptor inhibitors) has been hampered by modest single-agent efficacy and lack of predictive biomarkers.11 Recent single-cell RNA sequencing (scRNA-seq) landscapes of CRLM have corroborated the dominance of mannose receptor C type 1-positive (MRC1⁺) C-C motif chemokine ligand 18-positive (CCL18⁺) M2-like macrophages in metastatic lesions, revealing their spatial coordination with cancer-associated fibroblasts and metabolically rewired phenotypes.12,13 While these studies illuminate the immunosuppressive endpoint of CRLM progression, a fundamental question persists: What tumor-derived signals initiate and sustain this pathological reprogramming? Notably, the hepatic metastatic niche exhibits a profoundly more immunosuppressive landscape compared to primary CRC tumors, with markedly higher infiltration of M2-like TAMs and functionally exhausted T cells, which is widely considered the core driver of treatment failure in metastatic disease. Identifying the ‘instructive signals’ from metastatic cancer cells that hijack stromal cells remains a critical gap.
Intriguingly, cancer cells exhibit metabolic plasticity during metastasis, often coupling glycolytic reprogramming with non-metabolic ‘pleiotropic functions’.14 Enolase 2 (ENO2), a glycolytic enzyme catalyzing the conversion of 2-phosphoglycerate to phosphoenolpyruvate, exemplifies this duality. Beyond its canonical role in adenosine triphosphate (ATP) generation, ENO2 is dysregulated in neuroendocrine tumors, lung cancer, and CRC, where it correlates with chemoresistance.15,16 Mechanistically, ENO2 can regulate gene transcription via nuclear translocation and histone deacetylase (HDAC) inhibition.15 ENO2 also stabilizes oncogenic kinases through protein-protein interactions.17 However, its potential as a master regulator of immune-microenvironment crosstalk in CRLM is entirely unexplored. Whether ENO2 coordinates metastatic niche formation via direct interaction with immune modulators represents a compelling biological hypothesis.
To address this, we leveraged paired scRNA-seq of primary CRC tumors, adjacent normal tissues, and liver metastases from CRLM patients compared with non-metastatic patients. Combined with functional validation, we identified ENO2⁺ cancer cells as metastasis-specific drivers of epithelial-mesenchymal transition (EMT). Mechanistically, we discovered a novel ENO2-macrophage migration inhibitory factor (MIF) protein complex: ENO2 directly binds to MIF in CRC cells, stabilizes MIF by inhibiting CHIP-mediated ubiquitination and degradation, thus activating MIF signaling to promote tumor-macrophage crosstalk and induce M2 polarization. A novel ENO2-MIF protein complex activates tumor-macrophage interactions and induces M2 polarization, while the ENO2-MIF complex inhibitor pyrithioxin can effectively intervene in CRLM. Our work establishes ENO2 as a pleiotropic orchestrator of CRLM, bridging intracellular signaling with immune reprogramming through direct protein interaction, which unveiled a targetable vulnerability for metastatic CRC.
Results
Single-cell atlas reveals enhanced malignancy in liver metastatic lesions
To delineate the cellular ecosystem of CRLM, we performed scRNA-seq on 15 tissue specimens obtained from 6 patients: 3 with non-metastatic CRC (CRC-nM) and three with CRLM. For CRLM patients, paired samples included primary colorectal tumors (CRC), adjacent intestinal tissues and liver metastases (LM). The clinicopathological characteristics of the patients are summarized in Fig. 1a.
Single-Cell Atlas Reveals Enhanced Malignancy in Liver Metastatic Lesions. a Study design: scRNA-seq of tissues from 3 non-metastatic CRC patients and 3 CRLM patients. Clinicopathological features are summarized. (Created with BioRender.com). b Composition of major cell types across liver metastasis (LM), adjacent tissues (CRC-adjacent), and primary CRC tissues in three CRLM patients. c UMAP visualization of 11 cell types annotated by marker genes (Supplementary Table 1). d Tissue origin distribution of annotated cells. e Subclustering of epithelial cells showing cancer cell enrichment in tumors. f Proportional abundance of epithelial subpopulations. g CNV scores in cancer cells from primary (CRC) versus metastatic (LM) sites. h Volcano plot displaying differentially expressed genes from scRNA sequencing data, comparing cells from liver metastases (LM) to those from primary colorectal cancers (CRC). i Cell proliferation signature in metastatic versus primary cancer cells. j Hypoxia‑related gene expression in liver metastasis. Box plot showing hypoxia signature scores in LM versus CRC cells
In the three CRLM patients, unsupervised clustering identified epithelial cells as the dominant compartment within tumor tissues at both primary and metastatic sites (Fig. 1b–d), and further subclustering of epithelial cells revealed cancer cell enrichment in tumors versus adjacent tissues (Fig. 1e, f). Critically, cancer cells from liver metastases exhibited multifaceted evidence of enhanced malignancy compared to their primary site counterparts. Elevated genomic instability was evidenced by significantly higher copy number variation (CNV) scores (Fig. 1g), accompanied by transcriptional upregulation of proliferation drivers and hypoxia-responsive genes (Fig. 1h). These molecular alterations converged into functionally amplified phenotypes, with metastatic cells demonstrating markedly increased proliferation capacity and hypoxia adaptation (Fig. 1i, j). Collectively, our single-cell atlas delineates the cellular ecosystem of CRLM and uncovers increased malignancy, marked by higher genomic instability, proliferative capacity, and hypoxic adaptation in liver metastatic lesions compared to primary tumors.
Module-based discovery of metastasis-primed cancer cell subpopulations
To uncover metastasis-primed cancer cell subpopulations, we performed a module-based analysis on single-cell RNA-seq data. Uniform Manifold Approximation and Projection (UMAP) dimensionality reduction of tumor cells revealed eight distinct mechanical subgroups in both metastatic and primary lesions (Fig. 2a, b, Supplementary Fig. 1a). These subgroups exhibited varying degrees of malignancy as assessed by CNV scoring (Fig. 2c, Supplementary Fig. 1b, c).
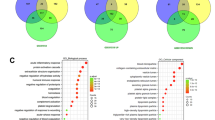
Module-Based Discovery of Metastasis-Primed Cancer Cell Subpopulations. a, b UMAP dimensionality reduction analysis was performed on tumor cells, and the distribution of eight mechanical subgroups in metastatic and primary lesions was displayed. c Evaluate the malignancy level of eight cancer cell subpopulations through CNV scoring, with macrophages (MPs) with less mutation load as negative control. d Cluster and reduce the dimensionality of cancer cells based on the module analysis model and compare them with the specific subtypes of metastatic lesions shown in (a, b); positive scoring cells under ‘Module 11’ are distributed in cancer cells 1/2/3, which better reflects the characteristics of metastatic cancer cells. e Analyzing the correlation between genes in the ‘Module 11’ gene set and the prognosis of colorectal cancer using a multifactorial Cox regression model in the TCGA-COAD dataset. f, g UMAP dimensionality reduction analysis was performed on tumor cells from 6 primary lesion tissues. The distribution of six subgroups in the primary lesions of metastatic patients and non-metastatic patients was displayed. h, i Analysis of the potential characteristics of six subpopulations of tumor cells using typical malignant features of tumor cells; the cancer cell 3 subgroup shows a clear tendency toward EMT
We next applied a module analysis model to cluster cancer cells and compared the results with the metastasis-associated subgroups identified above. Cells positively scoring for ‘Module 11’ were predominantly enriched in cancer cell subgroups 1, 2, and 3, which collectively exhibited characteristics of metastatic cancer cells (Fig. 2d, Supplementary Fig. 2). Importantly, several genes within ‘Module 11’ were significantly correlated with poor prognosis in CRC patients from the The Cancer Genome Atlas (TCGA) Colon Adenocarcinoma (COAD) dataset, as determined by a multifactorial Cox regression model (Fig. 2e).
We further analyzed the single-cell profiles of in situ lesions in metastatic and non-metastatic patients (Supplementary Fig. 3a–d). Focusing on primary lesions, UMAP dimensionality reduction identified six subgroups, whose distribution differed between metastatic and non-metastatic patients (Fig. 2f, g). Evaluation of classical malignant features across these subgroups revealed that cancer cell subgroup 3 displayed a pronounced epithelial-mesenchymal transition (EMT) signature, suggesting its potential role in initiating metastasis (Fig. 2h, i). Together, these findings suggest that the ‘Module 11’ gene set conforms to the expression characteristics of metastatic CRC cells and pinpoint a distinct cancer cell subpopulation (subgroup 3) with EMT features that may represent a metastasis-primed state in primary tumors.
ENO2 drives metastatic progression and serves as a therapeutic target
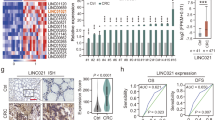
Building upon the identification of metastasis-associated signatures, we distilled 10 candidate genes from the ‘Module 11’ and cancer cell 3 profiles (Fig. 3a, Supplementary Fig. 4a, b). Among these, only ENO2 emerged as a significant prognostic determinant, with high expression correlating with markedly reduced survival in CRC patients (Fig. 3b). Clinical validation in the Nanjing and Guangzhou cohorts confirmed concurrent ENO2 upregulation in primary tumors and liver metastases compared to normal tissues (Fig. 3c, Supplementary Fig. 4c, d), establishing its negative correlation with patient survival (Fig. 3d). The correlation between ENO2 expression and CRLM was validated using the GSE41258 dataset and TNMplot online tool18 (Supplementary Fig. 5a–c). Functional interrogation in C57BL/6 J mouse models revealed profound metastasis-suppressing effects of ENO2 ablation. Subcutaneous xenograft assays demonstrated significantly attenuated tumor growth in the ENO2-knockout groups, as evidenced by reduced tumor volume and weight (Fig. 3e). Crucially, splenic injection models showed suppressed hepatic colonization in ENO2-deficient cells, with metastatic nodules reduced by >50% (Fig. 3f).
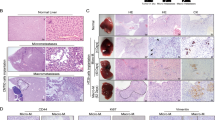
ENO2 Drives Metastatic Progression and Serves as a Therapeutic Target. a Ten candidate genes were screened from the ‘Module 11’ gene set and ‘Cancer cell 3’ characteristic genes. b Correlation analysis of 10 candidate genes and prognosis of colorectal cancer based on multivariate Cox regression in the TCGA-COAD database. c Detection of the correlation between ENO2 expression and colorectal cancer progression using immunohistochemistry in multiple independent cohorts. d Using three independent sample cohorts from Nanjing and Guangzhou, Kaplan-Meier analysis was conducted to investigate the association between ENO2 expression and the prognosis of colorectal cancer. e Observation of the tumorigenic ability of ENO2 knockout DLD-1 cells in a C57BL/6 J mouse subcutaneous tumor model and comparison of the weight and volume of the tumor tissues. f Using the intrasplenic injection model in C57BL/6 J mice, we observed the number of liver metastatic nodules formed by ENO2-KO DLD-1 cells compared with wild-type cells. g Using pharmacological inhibition of ENO2 (ENOblock) to validate the potential of targeting ENO2 to inhibit CRLM. h Multiplex immunofluorescence of EMT markers (E-cadherin, vimentin) in control and ENO2-modulated cancer cells
Therapeutic relevance was further underscored by pharmacological inhibition of ENO2 (ENOblock), which effectively blocked its pro-metastatic function (Fig. 3g). Multiplex immunofluorescence confirmed the mechanistic link between ENO2 and EMT induction, showing characteristic E-cadherin loss and vimentin gain in ENO2-expressing tumors (Fig. 3h). This suggests that targeting ENO2 has certain clinical application prospects.
ENO2 orchestrates M2 macrophage polarization through direct interaction with MIF
To investigate how cancer cells shape the tumor microenvironment (TME), we analyzed cell-cell communication using CellChat. The analysis revealed that ENO2+ cancer cells were major signaling hubs within the TME, with the MIF signaling pathway exhibiting particularly strong and specific outgoing/incoming signals (Fig. 4a, Supplementary Fig. 6a–d). Focusing on MIF-mediated interactions, we found that ENO2+ cancer cells showed significantly enhanced communication with macrophages compared to ENO2- cancer cells (Fig. 4b, c, Supplementary Fig. 6e, f). This computational prediction aligned with our observation that macrophages in metastatic lesions exhibited a higher M2 polarization tendency than those in primary or non-metastatic tumors (Fig. 4d).
ENO2 Orchestrates M2 Macrophage Polarization through Direct Interaction with MIF. a Analysis of the strength and potential ways in which various cell types participate in microenvironment signal communication based on CellChat; ENO2+ cancer cells exhibit strong microenvironment signals, while the MIF pathway shows significant differences in the transmission/reception of signals in ENO2+ cancer cells. b, c The interaction strength between ENO2-/+ cancer cells and different cell types was analyzed under the involvement of the MIF signaling pathway, with significant differences observed in the interaction with macrophages. d Compared to in situ or non-metastatic in situ lesions, macrophages in metastatic lesions and metastatic in situ lesions have a higher tendency toward M2 polarization. e Co-IP detection confirms the interaction between endogenous ENO2 and MIF. Immunoprecipitation was performed using anti-MIF or anti-ENO2 antibodies in HCT-116 (left) and DLD-1 (right) cell lysates, followed by immunoblotting. IgG served as a negative control. f Co‑IP analysis of endogenous MIF and ENO2 in DLD-1 cells, confirming their endogenous interaction in cancer cells. g GST pull‑down assay demonstrating a direct interaction between MIF and ENO2 in vitro. Recombinant GST‑ENO2 or GST alone was incubated with Myc‑tagged MIF. Proteins bound to glutathione beads were analyzed by immunoblotting with anti‑Myc and anti‑GST antibodies. h, i Molecular docking simulations predicted the key binding interface between ENO2 and MIF, and domain-specific mutants were used in mutual IP analysis to demonstrate complete elimination of binding after domain disruption
To elucidate the mechanistic basis, we identified direct physical binding between ENO2 and MIF through co-immunoprecipitation (Co-IP) coupled with mass spectrometry (Fig. 4e, f, Supplementary Fig. 7a). This interaction was direct, as demonstrated by an in vitro glutathione S-transferase (GST) pull-down assay using purified recombinant proteins (Fig. 4g). To define the molecular interface, we performed molecular docking simulations, which predicted a specific binding mode between ENO2 and MIF (Fig. 4h). Subsequent Co-IP experiments using domain-specific mutants of both proteins validated these predictions, showing that disrupting the predicted interface completely abrogated their binding (Fig. 4i). These results establish a direct physical interaction between ENO2 and MIF in cancer cells and suggest that this interaction underpins the enhanced MIF-mediated signaling from ENO2+ cancer cells to macrophages, promoting an M2-polarized immunosuppressive niche.
ENO2 stabilizes MIF by inhibiting ubiquitin-mediated degradation to activate pro-metastatic signaling
We next investigated the functional consequence of the ENO2-MIF interaction. A proximity ligation assay (PLA) confirmed the endogenous ENO2-MIF interaction in situ within cancer cells (Fig. 5a). Functionally, knockdown of ENO2 in DLD-1 cells led to a marked decrease in MIF protein levels, whereas its overexpression in HCT-116 cells increased MIF abundance, suggesting that ENO2 positively regulates MIF steady-state levels (Fig. 5b).
ENO2 Stabilizes MIF by Inhibiting Ubiquitin-Mediated Degradation. a Proximity ligation assay (PLA) demonstrating the interaction between endogenous ENO2 and MIF in cells. Nuclei were stained with DAPI. b Western blot analysis of MIF protein levels upon knockdown of ENO2 (left, DLD-1 cells) or overexpression of ENO2 (right, HCT-116 cells) compared to non-targeting controls. c Western blot analysis of MIF protein levels in HCT-116 cells overexpressing ENO2 or DLD-1 cells with ENO2 knockdown; both were treated with the proteasome inhibitor MG132. d Western blot analysis of MIF protein levels under various conditions: knockdown of ENO2 (left, DLD-1 cells), knockdown of CHIP, or overexpression of ENO2 (right, HCT-116 cells) and CHIP as indicated. The corresponding expression levels of ENO2, CHIP, and MIF are shown. e Western blot analysis assessing the polyubiquitination level of wild-type or a putative mutant Flag-tagged ENO2 protein in cells treated with MG132. f MIF protein levels in cells expressing wild-type or mutant Flag-tagged ENO2. g Western blot analysis of MIF downstream signaling pathway proteins in ENO2-modulated cancer cells. The phosphorylation levels of STAT3 (p-STAT3) and P65 (p-P65), along with their total protein levels, were examined. β-Actin served as a loading control
Given that MIF is known to be regulated by ubiquitin-mediated proteasomal degradation, we explored whether ENO2 stabilizes MIF by interfering with this process. Treatment with the proteasome inhibitor MG132 restored MIF levels in ENO2-knockdown cells and further elevated them in ENO2-overexpressing cells (Fig. 5c, Supplementary Fig. 8). In MG132-treated cells, ENO2 knockdown significantly enhanced MIF polyubiquitination, while ENO2 overexpression exerted the opposite effect (Fig. 5d).
Domain-mapping experiments confirmed that this stabilization requires intact binding interfaces, as ENO2 catalytic domain mutants failed to protect MIF from ubiquitination (Fig. 5e, f, Supplementary Fig. 7b, c). Finally, we examined the impact on downstream oncogenic signaling. Modulation of ENO2 expression correspondingly altered the phosphorylation levels of STAT3 and NF-κB subunit P65 (Fig. 5g), two key pathways activated by MIF. These data demonstrate that ENO2 binds to and stabilizes the MIF protein by antagonizing CHIP-mediated polyubiquitination and subsequent proteasomal degradation, thereby potentiating MIF-dependent activation of STAT3 and NF-κB signaling in cancer cells.
ENO2 induces M2 macrophage polarization to drive liver metastasis
To investigate whether the ENO2-MIF axis functionally reshapes the tumor immune microenvironment, we first analyzed their spatial relationship. Spatial transcriptomic analysis revealed a significant co-localization between ENO2+ cancer cells and M2-like macrophages in metastatic lesions (Fig. 6a, Supplementary Fig. 9a). Quantitative correlation analysis across multiple ST samples confirmed a consistent and significant positive correlation (all P < 0.001) between these two populations (Supplementary Fig. 9b). This spatial association prompted us to test whether ENO2-expressing cancer cells could directly induce M2 polarization.
ENO2 Induces M2 Macrophage Polarization to Drive Liver Metastasis. a Spatial feature plot showing the deconvoluted proportions of ENO2+ cancer cells and M2 macrophages in ST_liver4. The color scale indicates the predicted cell type weights per spot. Flow cytometry analysis of the macrophage markers CD86 and CD206 in co-culture systems (b) and xenograft tumors (c). d Representative flow cytometry plots showing the expression of CD206 and CD86 on macrophages co-cultured with patient-derived organoid (PDO) models. PDOs were transduced with ENO2 knockout sgRNA (Sg-ENO2 PDO) or ENO2 overexpression vector (ENO2 PDO) compared with their controls (Sg-NC PDO and Vector PDO). e, f Investigation of the effect of ENO2 expression on the formation of liver metastases in a splenic implantation model using C57BL/6 J mice. M2 polarization inhibitor (LPS), MIF inhibitor (ISO-1), M2 polarization agonist (IL-4), and MIF overexpression were used in the rescue experimental system
Using in vitro co-culture systems, we found that macrophages co-cultured with ENO2-knockdown cancer cells displayed increased M1 marker (CD86) and decreased M2 markers (CD206), whereas those co-cultured with ENO2-overexpressing cells showed the opposite trend (Fig. 6b, Supplementary Fig. 10a, b). This ENO2-dependent M2 polarization was recapitulated in vivo, as evidenced by a higher proportion of CD206+ macrophages in xenograft tumors derived from ENO2-overexpressing cells compared to controls (Fig. 6c, Supplementary Fig. 10c). The critical role of cancer cell-intrinsic ENO2 was further confirmed using patient-derived organoids (PDOs); macrophages co-cultured with ENO2-knockout PDOs had a reduced M2 phenotype, while those co-cultured with ENO2-overexpressing PDOs exhibited enhanced M2 polarization (Fig. 6d, Supplementary Fig. 10d, e).
We then assessed the functional impact of this axis on metastasis in vivo. In a splenic implantation-liver metastasis model using C57BL/6 J mice, ENO2 overexpression in cancer cells significantly promoted liver metastasis formation (Fig. 6e, f). Importantly, this pro-metastatic effect was abolished by treatment with the MIF inhibitor ISO-1 or the M2 polarization inhibitor LPS. Conversely, the attenuated metastasis resulting from ENO2 knockdown was rescued by either administering the M2 agonist IL-4 or enforcing MIF overexpression in cancer cells (Fig. 6e, f), and immunohistochemistry confirmed the abundance of M2 polarization between groups (Supplementary Fig. 10f, g). These results demonstrate that ENO2+ cancer cells, through secreting MIF, drive M2 macrophage polarization in the tumor microenvironment, which in turn facilitates the formation of liver metastases.
ENO2 recruits HSP90 to antagonize CHIP-mediated ubiquitination and degradation of MIF
To further elucidate how ENO2 stabilizes MIF, we sought to identify its molecular interactors. Mass spectrometry analysis of ENO2 complexes identified the chaperone heat shock protein 90 (HSP90) as a binding partner (Fig. 7a), which was confirmed by reciprocal Co-IP of endogenous proteins (Fig. 7b, c). Intriguingly, Co-IP experiments revealed that ENO2 enhanced the physical association between HSP90 and MIF, suggesting that ENO2 acts as a scaffold to recruit HSP90 to MIF (Fig. 7d).
ENO2 recruits HSP90 to antagonize CHIP-mediated ubiquitination and degradation of MIF. a Representative liquid chromatography-tandem mass spectrometry (LC-MS/MS) spectrum identifying HSP90 as an ENO2-interacting protein, with matched peptide fragments highlighted. Reciprocal Co-IP assays confirming the interaction between endogenous ENO2 and HSP90. HCT-116 (b) and DLD-1 (c) cell lysates were immunoprecipitated with anti-HSP90 or anti-ENO2 antibodies and immunoblotted with the indicated antibodies. IgG served as a negative control. d Co-IP analysis showing that ENO2 enhances the association between HSP90 and MIF. DLD-1 cells were transfected with the indicated plasmids or shRNAs and treated with MG132 prior to immunoprecipitation using an anti-Flag antibody. Immunoblot analysis of MIF protein levels following knockdown (DLD-1, e) or overexpression (HCT-116, f) of ENO2 and/or CHIP, indicating that ENO2 stabilizes MIF in a CHIP-dependent manner. g Ubiquitination assay demonstrating that ENO2 depletion promotes CHIP-mediated ubiquitination of MIF. DLD-1 cells were treated with MG132, followed by immunoprecipitation of MIF and immunoblotting for ubiquitin. h Expression validation of wild-type and lysine-mutant MIF constructs (K32R, K66R, K77R) co-transfected with Flag-CHIP in DLD-1 cells. i Ubiquitination assays identifying lysine residues critical for CHIP-mediated ubiquitination of MIF. DLD-1 cells were co-transfected with HA-tagged ubiquitin, Flag-CHIP, and Myc-tagged wild-type or mutant MIF constructs, treated with MG132, immunoprecipitated with anti-Myc antibody, and immunoblotted with anti-HA antibody
Since the E3 ligase CHIP is known to target HSP90-client proteins for degradation,19 we hypothesized that ENO2 might protect MIF by antagonizing CHIP. Consistent with this, the reduction in MIF levels upon ENO2 knockdown was rescued by concurrent CHIP depletion (Fig. 7e), while ENO2 overexpression-induced MIF accumulation was suppressed by co-expressing CHIP (Fig. 7f). Mechanistically, ENO2 knockdown markedly enhanced CHIP-mediated polyubiquitination of MIF (Fig. 7g).
To define the ubiquitination site, we generated MIF mutants in which candidate lysines (K32, K66, K77) were substituted. Ubiquitination assays revealed that mutation at K66, but not K32 or K77, substantially abolished CHIP-mediated polyubiquitination of MIF (Fig. 7h, i), identifying K66 as the primary site. Taken together, these results demonstrate that ENO2 recruits HSP90 to MIF, forming a tripartite complex that competitively inhibits the E3 ligase CHIP, thereby suppressing CHIP-mediated ubiquitination of MIF at K66 and its subsequent proteasomal degradation.
Identification and validation of pyrithioxin as an ENO2-MIF interaction inhibitor and its anti-metastatic effect in vivo
To translate our mechanistic findings into a potential therapeutic strategy, we sought to identify a small-molecule inhibitor targeting the ENO2-MIF interaction through an in silico screening approach. A library of 6,723 compounds was sequentially filtered through absorption, distribution, metabolism, excretion, and toxicity (ADME/T) evaluation, multi-level molecular docking, and molecular mechanics with generalized Born and surface area solvation (MM-GBSA) binding free energy rescoring (Fig. 8a). Among the top candidates, pyrithioxin dihydrochloride exhibited one of the most favorable predicted binding affinities (Fig. 8b). In validation experiments, the selected compounds, including pyrithioxin, effectively disrupted the interaction between ENO2 and MIF in Co-IP assays and increased MIF ubiquitination (Fig. 8c). Molecular docking suggested that pyrithioxin binds stably at the ENO2-MIF interface (Fig. 8d), a prediction further supported by stable root-mean-square deviation (RMSD) and favorable residue-specific root-mean-square fluctuation (RMSF) profiles in molecular dynamics simulations (Fig. 8e).
Identification and validation of pyrithioxin as an ENO2-MIF interaction inhibitor and its anti-metastatic effect in vivo. a Workflow of the in silico screening strategy. A total of 6723 compounds were subjected to ADME/T evaluation, multilevel docking, MM-GBSA rescoring, and binding assays to identify candidate compounds with high affinity. b MM-GBSA binding free energy (ΔG_bind) values of the top-ranked compounds. Pyrithioxin dihydrochloride showed one of the most favorable binding energies. c Co-IP assay showing that compounds (#1, #2, and #3) disrupt the interaction between ENO2 and MIF. Flag-ENO2 was immunoprecipitated, and ubiquitination of MIF was detected in the presence of MG132 in DLD-1 cells. d Molecular docking model illustrating the binding mode of pyrithioxin within the ENO2–MIF interface, highlighting key interacting residues. e Molecular dynamics simulations of the ENO2–pyrithioxin complex showing RMSD changes over time and residuewise RMSF analysis, indicating stable binding. f Schematic of the in vivo experimental design. MC38 cells were injected on day 0, followed by oral gavage of PBS or pyrithioxin three times per week until sacrifice on day 21. g Representative images of liver metastases, H&E staining (2× and 10×), in vivo bioluminescence imaging, and quantitative analysis of total flux and number of liver metastatic nodules in mice treated with 0, 10, or 20 mg/kg pyrithioxin. h Immunoblot analysis of MIF, p-STAT3/STAT3, and p-P65/P65 levels in tumor tissues from DMSO- or pyrithioxin-treated groups, with β-actin as a loading control
We next evaluated the anti-metastatic efficacy of pyrithioxin in vivo using a liver metastasis model (Fig. 8f). Oral administration of pyrithioxin (10 or 20 mg/kg) significantly inhibited the formation of liver metastases compared to the vehicle control, as evidenced by a reduced bioluminescent signal and fewer metastatic nodules on the liver surface (Fig. 8g). Histological analysis confirmed a marked decrease in metastatic burden within the liver parenchyma. Consistent with the role of the ENO2-MIF axis in activating STAT3 and NF-κB signaling, tumor tissues from pyrithioxin-treated mice showed reduced levels of MIF and decreased phosphorylation of STAT3 and P65 (Fig. 8h). These results identify pyrithioxin as a novel inhibitor of the ENO2-MIF interaction and demonstrate its potent anti-metastatic activity in vivo, highlighting the therapeutic potential of targeting this protein-protein interface.
In summary, we revealed a new mechanism of CRLM: ENO2 binds to MIF, inhibiting its CHIP-mediated ubiquitination degradation, thereby activating the MIF pathway, inducing M2 macrophage polarization, and driving immune suppression and metastatic proliferation. This mechanism of liver metastasis depends on the joint participation of ENO2+ cancer cells and M2 macrophages, and a novel inhibitor of the ENO2-MIF interaction, pyrithioxin, can effectively intervene in the process of liver metastasis. This study identifies ENO2 as a key target for blocking CRLM.
Discussion
CRLM remains a lethal clinical challenge, with its poor prognosis underscoring the urgent need to decipher the molecular drivers of metastatic progression.20 Leveraging single-cell RNA sequencing of anatomically paired primary and metastatic lesions compared with non-metastatic tissues, we identified ENO2 as a central orchestrator of CRLM. Considering that we did not observe a direct effect of ENO2 on promoting CRLM in nude mice, we are more inclined to believe that ENO2 promotes metastasis through intrinsic tumor reprogramming and extrinsic immune regulation. While ENO2 is canonically recognized as a glycolytic enzyme,21 our study reveals its non-metabolic ‘moonlighting’ function in CRLM pathogenesis. Beyond catalyzing phosphoenolpyruvate production,15 ENO2 physically binds and stabilizes MIF within cancer cells—activating p65 and STAT3 phosphorylation to amplify MIF signaling. This mechanism redefines ENO2 as a signaling scaffold that bridges tumor cell plasticity with immune evasion. Crucially, ENO2-driven MIF stabilization activates tumor-macrophage crosstalk, inducing immunosuppressive M2 polarization—a feature recently correlated with chemotherapy resistance in scRNA-seq studies.22 This suggests that ENO2 inhibition may also prevent local recurrence by attenuating EMT and immune suppression.
Our findings contextualize the immunosuppressive landscape of CRLM. Although prior work established the dominance of M2-like macrophages in metastatic niches,23,24 the tumor-derived signals instructing their polarization remain elusive. We demonstrate that ENO2⁺ cancer cells initiate this reprogramming through MIF, a pleiotropic cytokine known to trigger M2 differentiation via CD74/pERK pathways.25 Mechanistically, we revealed that ENO2 directly binds to MIF, and further recruits HSP90 to form a tripartite complex, which competitively inhibits E3 ligase CHIP-mediated K66-site polyubiquitination and proteasomal degradation of MIF. This explains the strong interaction between ENO2⁺ tumor clusters and M2 macrophages while accounting for broader immunosuppressive features such as NK cell suppression.26 Importantly, our paired-site scRNA-seq design revealed metastasis-specific ENO2 enrichment—a pattern undetectable in bulk analyses, resolving the paradox of minimal transcriptomic differences between primary and metastatic cancer cells.27
Therapeutic implications emerge from the druggable nature of the ENO2-MIF interface. Disrupting this axis could simultaneously target EMT and M2 polarization, offering advantages over single-pathway inhibitors. Clinically, ENO2⁺/MIF⁺ tumors may identify patients for tailored therapy, particularly those with chemoresistant CRLM.28 Existing MIF inhibitors show preclinical efficacy in other cancers29; our data justify their prioritization for CRLM. Contrary to assumptions of microenvironmental uniformity, our scRNA-seq atlas confirms dynamic TME remodeling during hepatic colonization.30 This heterogeneity necessitates precision strategies targeting metastasis-adaptive mechanisms such as the ENO2-MIF axis. Meanwhile, several limitations should be noted. First, our discovery scRNA-seq cohort included only 6 patients, and the prognostic and predictive value of ENO2 needs to be validated in large-scale, multi-center clinical cohorts. Second, all enrolled patients were treatment-naive without neoadjuvant chemoradiotherapy, so the impact of systemic treatment on the ENO2-MIF axis remains unclear, which may affect its clinical application in pre-treated patients.31 Third, our study focused exclusively on liver metastasis, and the role of the ENO2-MIF axis in other distant metastatic sites of CRC (e.g., lung, peritoneum) has not been explored. Finally, the specificity and clinical safety of the ENO2-MIF interaction inhibitor pyrithioxin need further optimization and validation in preclinical studies.
While our study focuses on a non-canonical, protein-stabilizing function of ENO2, its role as a glycolytic enzyme necessitates consideration of the broader metabolic and systemic context of CRLM patients.32 Tumor glycolytic flux, driven in part by enzymes such as ENO2, is influenced by and contributes to the systemic metabolic state, including potential cachexia and altered nutrient availability in advanced cancer patients. This metabolic reprogramming within the tumor may intersect with the mechanism described here. Furthermore, patient immune status, often modulated by systemic metabolism and nutrition, forms a critical backdrop for macrophage polarization. The ENO2-MIF axis we identified may therefore serve as a node connecting tumor-intrinsic glycolytic activity with extrinsic immune modulation within the metastatic niche. Also, we identified and validated pyrithioxin as a novel small-molecule inhibitor of the ENO2-MIF interaction, which effectively reduced liver metastasis burden in mice, providing a preclinical proof-of-concept for targeting this axis in CRLM, especially for MSS CRC patients who are insensitive to single-agent immune checkpoint blockade. For future research directions, high-priority areas include: large multi-center cohort validation of the clinical value of ENO2; optimization of the specificity and pharmacokinetic properties of pyrithioxin for systematic clinical trials; exploration of the crosstalk between ENO2’s canonical glycolytic enzyme activity and its non-canonical immune regulatory function; evaluation of the impact of neoadjuvant therapy on the ENO2-MIF axis; and investigation of the role of this axis in other CRC metastatic sites.
In conclusion, we unveil a pathogenic paradigm in which ENO2⁺ cancer cells stabilize MIF to induce M2 macrophage polarization, fueling a self-reinforcing cycle of immune suppression and metastatic outgrowth. This work shifts the focus from stromal cell reactiveness to tumor-instructive signaling in the metastatic niche, and identifies ENO2 as a dual functional regulator bridging tumor cell-intrinsic metabolic reprogramming and extrinsic immune niche remodeling. The ENO2-MIF axis we characterized represents a promising prognostic biomarker and actionable therapeutic target for intercepting CRLM progression and improving the dismal clinical outcomes of metastatic CRC patients.
Materials and methods
Sample collection
This study enrolled three non-metastatic CRC patients and three CRLM patients. All patients underwent preoperative imaging (CT/MRI) and colonoscopy, with postoperative pathology confirming primary colorectal adenocarcinoma (hepatocellular carcinoma excluded in CRLM cases). None of the patients received neoadjuvant radiotherapy or chemotherapy. During the 2–3 weeks before hospitalization, patients were guided in preoperative preparation, including standardized diet, exercise, etc. This was intended not only to prevent postoperative complications and enhance postoperative recovery but also to objectively strengthen the balance and comparability between different groups. From non-metastatic patients, matched tumor and adjacent tissues (n = 6 samples) were collected; from CRLM patients, primary tumor, adjacent intestinal tissue, and metastatic liver tissues (n = 9 samples) were obtained, totaling 15 samples for scRNA-seq. During postoperative follow-up, patient 2 experienced a recurrence at the 18th month. Written informed consent was acquired from all participants, and the study protocol was approved by the Ethics Committee of the First Affiliated Hospital of Nanjing Medical University (2022-SRFA-142).
Domain mapping experiment
The full-length ENO2 gene and its different domain truncations were amplified by PCR. The PCR products were digested with restriction enzymes (EcoRI and BamHI) and inserted into the pCMV-HA vector to construct HA-ENO2 (full-length), HA-ENO2-N, HA-ENO2-Cat, and HA-ENO2-C recombinant vectors. All recombinant vectors were verified by DNA sequencing. 293 T cells were seeded into 6-well plates at a density of 2 × 10⁵ cells/well and cultured overnight. The recombinant vectors (HA-ENO2 full-length or truncations) and pcDNA3.1-MIF vector were co-transfected into 293 T cells using Lipofectamine 3000 transfection reagent. The cells were cultured for 48 h. The cells were collected and lysed, and a Co-IP assay was performed using HA Tag antibody (same as Section 3.2). Western blotting was used to detect the binding of ENO2 truncations to MIF (MIF antibody was used for detection). The domain of ENO2 that binds to MIF was identified according to the presence or absence of MIF bands in the eluate.
In vitro co-culture assay
THP-1 cells were cultured in Roswell Park Memorial Institute 1640 medium (RPMI 1640) containing 10% fetal bovine serum (FBS). The cells were seeded into 6-well plates at a density of 1 × 10⁶ cells/well, and 100 ng/mL phorbol 12-myristate 13-acetate (PMA) was added. The cells were cultured in a 37 °C, 5% CO₂ incubator for 24 h to induce differentiation into macrophages. After induction, the medium was discarded, and the macrophages were washed twice with PBS. Transwell chambers (pore size 0.4 μm) were placed in 6-well plates. ENO2-overexpressing or -knockout HCT-116 cells (1 × 10⁵ cells/well) were seeded into the upper chamber (in Dulbecco’s Modified Eagle Medium (DMEM) containing 10% FBS). The induced THP-1 macrophages (5 × 10⁵ cells/well) were seeded into the lower chamber (in RPMI 1640 medium containing 10% FBS). The cells were co-cultured at 37 °C in a 5% CO₂ incubator for 48 h. A single culture group (only HCT-116 cells or only macrophages) was set as the control group. The macrophages in the lower chamber were collected, stained with CD68 and CD206 antibodies, and detected by flow cytometry to calculate the M2 macrophage positive rate.
Statistical analysis
Continuous variables were compared using Student’s t-test (normal distribution) or the Mann-Whitney U test (non-normal distribution). Categorical/ordinal data were analyzed by the chi-square test or Fisher’s exact test. Survival analysis employed the log-rank test with Kaplan-Meier curves. Multiple comparisons were adjusted using the Benjamini-Hochberg false discovery rate (FDR) method; FDR-adjusted p < 0.05 was considered statistically significant. Analyses were performed in R v4.1.2.
The remaining materials and methods are described in the Supplementary Materials.
Data availability
The expression matrix data were uploaded to the Gene Expression Omnibus (GEO) database under accession number GSE315534. All other data supporting the findings of this study are included in the published article and its supplementary information files.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J. Clin. 74, 229–263 (2024).
Dai, S. et al. miR-122/NEGR1 axis contributes colorectal cancer liver metastasis by PI3K/AKT pathway and macrophage modulation. J. Transl. Med. 22, 1060 (2024).
Bond, M. J. G. et al. First-line systemic treatment strategies in patients with initially unresectable colorectal cancer liver metastases (CAIRO5): an open-label, multicentre, randomised, controlled, phase 3 study from the Dutch Colorectal Cancer Group. Lancet Oncol. 24, 757–771 (2023).
Lebeck Lee, C. M. et al. A contemporary systematic review on liver transplantation for unresectable liver metastases of colorectal cancer. Cancer 128, 2243–2257 (2022).
Su, Y.-M. et al. Five-year survival post hepatectomy for colorectal liver metastases in a real-world Chinese cohort: Recurrence patterns and prediction for potential cure. Cancer Med. 12, 9559–9569 (2023).
Zhao, S. et al. Tumor-derived exosomal miR-934 induces macrophage M2 polarization to promote liver metastasis of colorectal cancer. J. Hematol. Oncol. 13, 156 (2020).
Zhao, S. et al. Highly-metastatic colorectal cancer cell released miR-181a-5p-rich extracellular vesicles promote liver metastasis by activating hepatic stellate cells and remodelling the tumour microenvironment. J. Extracell. Vesicles 11, e12186 (2022).
Dai, S. et al. miR-455/GREM1 axis promotes colorectal cancer progression and liver me tastasis by affecting PI3K/AKT pathway and inducing M2 macrophage pola rization. Cancer Cell Int. 24, 235 (2024).
Zheng, Z. et al. KAJF alleviated colorectal cancer liver metastasis by reshaping the gut microbiota and inhibiting M2-like macrophage polarization. Phytomedicine 142, 156766 (2025).
Mantovani, A., Marchesi, F., Malesci, A., Laghi, L. & Allavena, P. Tumour-associated macrophages as treatment targets in oncology. Nat. Rev. Clin. Oncol. 14, 399–416 (2017).
Voissière, A. et al. The CSF-1R inhibitor pexidartinib affects FLT3-dependent DC differentiation and may antagonize durvalumab effect in patients with advanced cancers. Sci. Transl. Med. 16, eadd1834 (2024).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459.e429 (2020).
Guo, D., Meng, Y., Zhao, G., Wu, Q. & Lu, Z. Moonlighting functions of glucose metabolic enzymes and metabolites in cancer. Nat. Rev. Cancer 25, 426–446 (2025).
Wang, C. et al. ENO2-derived phosphoenolpyruvate functions as an endogenous inhibitor of HDAC1 and confers resistance to antiangiogenic therapy. Nat. Metab. 5, 1765–1786 (2023).
Lv, C. et al. ENO2 promotes colorectal cancer metastasis by interacting with the LncRNA CYTOR and activating YAP1-induced EMT. Cells 11, 2363 (2022).
Shao, R. et al. Tumor-derived Exosomal ENO2 modulates polarization of tumor-associated macrophages through reprogramming glycolysis to promote progression of diffuse large B-cell lymphoma. Int. J. Biol. Sci. 20, 848–863 (2024).
Bartha, Á & Győrffy, B. TNMplot.com: A Web Tool for the Comparison of Gene Expression in Normal, Tumor and Metastatic Tissues. Int. J. Mol. Sci. 22, 2622 (2021).
Schulz, R. et al. Inhibiting the HSP90 chaperone destabilizes macrophage migration inhibitory factor and thereby inhibits breast tumor progression. J. Exp. Med. 209, 275–289 (2012).
Li, C. et al. Integrated omics of metastatic colorectal cancer. Cancer Cell 38, 734–747.e739 (2020).
Zheng, Y. et al. Insulin-like growth factor 1-induced enolase 2 deacetylation by HDAC3 promotes metastasis of pancreatic cancer. Signal Transduct. Target. Ther. 5, 53 (2020).
Wen, J. et al. Clusterin-mediated polarization of M2 macrophages: a mechanism of temozolomide resistance in glioblastoma stem cells. Stem Cell Res. Ther. 16, 146 (2025).
Han, S. et al. d-lactate modulates M2 tumor-associated macrophages and remodels immunosuppressive tumor microenvironment for hepatocellular carcinoma. Sci. Adv. 9, eadg2697 (2023).
Han, D. et al. The USP1-WDR48 deubiquitinase complex functions as a molecular switch regulating tumor-associated macrophage activation and anti-tumor response. Cell Death Differ. 33, 92–110 (2025).
Xie, S. et al. Elevated MIF identified by multiple cytokine analyses facilitates macrophage M2 polarization contributing to postoperative recurrence in chronic rhinosinusitis with nasal polyps. Rhinology 62, 432–445 (2024).
Tang, L. et al. Extracellular vesicles-derived hybrid nanoplatforms for amplified CD47 blockade-based cancer immunotherapy. Adv. Mater. 35, e2303835 (2023).
Li, R. et al. Single-cell transcriptomic analysis deciphers heterogenous cancer stem-like cells in colorectal cancer and their organ-specific metastasis. Gut 73, 470–484 (2024).
Zheng, Y. et al. Matrix stiffness triggers lipid metabolic cross-talk between tumor and stromal cells to mediate bevacizumab resistance in colorectal cancer liver metastases. Cancer Res. 83, 3577–3592 (2023).
Cohen Shvefel, S. et al. Temporal genomic analysis of homogeneous tumor models reveals key regulators of immune evasion in melanoma. Cancer Discov. 15, 553–577 (2025).
Deng, Y. et al. Comprehensive single-cell atlas of colorectal neuroendocrine tumors with liver metastases: unraveling tumor microenvironment heterogeneity between primary lesions and metastases. Mol. Cancer 24, 28 (2025).
Shi, M. et al. Genetic and microenvironmental evolution of colorectal liver metastases under chemotherapy. Cell Rep. Med. 5, 101838 (2024).
Majem, B. et al. Abstract PR008: Cross-talk between metastatic cells and host systems: neutrophil metabolic adaptation, immune profiling, and systemic metabolic shifts in tumor progression and cachexia. Cancer Res. 84, PR008–PR008 (2024).
Acknowledgements
This work was supported by the National Natural Science Foundation (Grant Numbers 82304221, 82273406, and 82473049); the China Postdoctoral Science Foundation (2022M721679); the Basic Research Program of Jiangsu Province (BK20230730); the Jiangsu Key Medical Discipline (General Surgery; Grant No. ZDXK202222); and the Jiangsu Provincial Association of Science and Technology, Youth Science and Technology Talent Recruitment Project (JSTJ-2023-051). The Core Facility of Jiangsu Provincial People’s Hospital provided assistance in detecting experimental samples. We would like to thank Researcher Bo Li (Immunology Frontier Research Center, Osaka University) for his assistance in the analysis based on the TCGA and GEO databases.
Author information
Authors and Affiliations
Contributions
YM S conceived the project and supervised all experiments. J.T., Z.C., D.Z., H.X., and C.T. performed the experiments. X.W. analyzed the data. Q.S., S.Y., Z.C., and Y.F. were responsible for clinical sample collection. K.J. and J.D. provided support with analysis techniques. J.T., X.W., and D.X. constructed the manuscript. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, J., Chen, Z., Zhang, D. et al. ENO2 drives tumor cell-induced M2 macrophage polarization to promote colorectal cancer liver metastasis. Sig Transduct Target Ther 11, 166 (2026). https://doi.org/10.1038/s41392-026-02732-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02732-2