Abstract
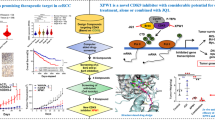
Programmed cell death protein 1 (PD-1) and its ligand programmed cell death ligand-1 (PD-L1) are key mediators of immune checkpoint blockade therapy in clear cell renal cell carcinoma (RCC). However, immune evasion and primary resistance often limit their efficacy, highlighting the need for improved strategies. Here, we identified dipeptidyl peptidase 9 (DPP9) as a critical regulator of PD-L1 expression in ccRCC. Pharmacological inhibition of DPP9 with 1G244 restores T cell cytotoxicity and enhances checkpoint blockade efficacy. Mechanistically, DPP9 disrupts the BRISC-SHMT2 complex, enhancing BRISC-mediated deubiquitination and stabilization of IFNAR1, which activates the JAK/STAT pathway and drives PD-L1 transcription. 1G244 reverses this process by reducing DPP9 interacting with SHMT2, promoting IFNAR1 ubiquitination and degradation, thereby reducing PD-L1 levels and restoring T cell-mediated cytotoxicity. Moreover, the combination of 1G244 and anti-CTLA-4 therapy further enhanced antitumor immunity, highlighting a potential synergistic therapeutic strategy. Collectively, our findings define a novel DPP9–BRISC–SHMT2 regulatory axis in PD-L1 transcriptional control and identify 1G244 as an alternative combinatorial strategy to enhance the efficacy of cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options








Similar content being viewed by others
Data availability
The original MS proteomics data, which have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository (www.ebi.ac.uk/pride) with the dataset [48] identifiers PXD059150 and PXD059151.
References
Xue G, Li X, Kalim M, Fang J, Jiang Z, Zheng N, et al. Clinical drug screening reveals clofazimine potentiates the efficacy while reducing the toxicity of anti-PD-1 and CTLA-4 immunotherapy. Cancer Cell. 2024;42:780–96.
Chuah S, Lee J, Song Y, Kim HD, Wasser M, Kaya NA, et al. Uncoupling immune trajectories of response and adverse events from anti-PD-1 immunotherapy in hepatocellular carcinoma. J Hepatol. 2022;77:683–94.
Riaz N, Havel JJ, Makarov V, Desrichard A, Urba WJ, Sims JS, et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell. 2017;171:934–49.
Kortlever RM, Sodir NM, Wilson CH, Burkhart DL, Pellegrinet L, Brown Swigart L, et al. Myc cooperates with Ras by programming inflammation and immune suppression. Cell. 2017;171:1301–15.
Kataoka K, Shiraishi Y, Takeda Y, Sakata S, Matsumoto M, Nagano S, et al. Aberrant PD-L1 expression through 3’-UTR disruption in multiple cancers. Nature. 2016;534:402–6.
Johnson DC, Taabazuing CY, Okondo MC, Chui AJ, Rao SD, Brown FC, et al. DPP8/DPP9 inhibitor-induced pyroptosis for treatment of acute myeloid leukemia. Nat Med. 2018;24:1151–6.
Hollingsworth LR, Sharif H, Griswold AR, Fontana P, Mintseris J, Dagbay KB, et al. DPP9 sequesters the C terminus of NLRP1 to repress inflammasome activation. Nature. 2021;592:778–83.
Podos SD, Ferguson EL. Morphogen gradients: new insights from DPP. Trends Genet. 1999;15:396–402.
Chang K, Chen Y, Zhang X, Zhang W, Xu N, Zeng B, et al. DPP9 stabilizes NRF2 to suppress ferroptosis and induce sorafenib resistance in clear cell renal cell carcinoma. Cancer Res. 2023;83:3940–55.
Karlow JA, Pehrsson EC, Xing X, Watson M, Devarakonda S, Govindan R, et al. Non-small cell lung cancer epigenomes exhibit altered DNA methylation in smokers and never-smokers. Genomics Proteomics Bioinformatics. 2023;21:991–1013.
Saso K, Miyoshi N, Fujino S, Sasaki M, Yasui M, Ohue M, et al. Dipeptidyl peptidase 9 increases chemoresistance and is an indicator of poor prognosis in colorectal cancer. Ann Surg Oncol. 2020;27:4337–47.
Zhou Y, Chen Y, Xuan C, Li X, Tan Y, Yang M, et al. DPP9 regulates NQO1 and ROS to promote resistance to chemotherapy in liver cancer cells. Redox Biol. 2024;75:103292.
Robinson KS, Toh GA, Rozario P, Chua R, Bauernfried S, Sun Z, et al. ZAKα-driven ribotoxic stress response activates the human NLRP1 inflammasome. Science. 2022;377:328–35.
Bauernfried S, Scherr MJ, Pichlmair A, Duderstadt KE, Hornung V Human NLRP1 is a sensor for double-stranded RNA. Science 2021;371:eabd0811.
Robinson KS, Teo DET, Tan KS, Toh GA, Ong HH, Lim CK, et al. Enteroviral 3C protease activates the human NLRP1 inflammasome in airway epithelia. Science 2020;370:eaay2002.
Harapas CR, Robinson KS, Lay K, Wong J, Moreno Traspas R, Nabavizadeh N, et al. DPP9 deficiency: an inflammasomopathy that can be rescued by lowering NLRP1/IL-1 signaling. Sci Immunol. 2022;7:eabi4611.
Huang M, Zhang X, Toh GA, Gong Q, Wang J, Han Z, et al. Structural and biochemical mechanisms of NLRP1 inhibition by DPP9. Nature. 2021;592:773–7.
Sharif H, Hollingsworth LR, Griswold AR, Hsiao JC, Wang Q, Bachovchin DA, et al. Dipeptidyl peptidase 9 sets a threshold for CARD8 inflammasome formation by sequestering its active C-terminal fragment. Immunity. 2021;54:1392–1404.
Clark KM, Kim JG, Wang Q, Gao H, Presti RM, Shan L. Chemical inhibition of DPP9 sensitizes the CARD8 inflammasome in HIV-1-infected cells. Nat Chem Biol. 2023;19:431–9.
Wolf C, Fischer H, Kühl JS, Koss S, Jamra RA, Starke S, et al. Hemophagocytic lymphohistiocytosis-like hyperinflammation due to a de novo mutation in DPP9. J Allergy Clin Immunol. 2023;152:1336–44.
Matheeussen V, Waumans Y, Martinet W, Van Goethem S, Van der Veken P, Scharpé S, et al. Dipeptidyl peptidases in atherosclerosis: expression and role in macrophage differentiation, activation and apoptosis. Basic Res Cardiol. 2013;108:350.
Wu JJ, Tang HK, Yeh TK, Chen CM, Shy HS, Chu YR, et al. Biochemistry, pharmacokinetics, and toxicology of a potent and selective DPP8/9 inhibitor. Biochem Pharmacol. 2009;78:203–10.
Kikuchi S, Wada A, Kamihara Y, Okazaki K, Jawaid P, Rehman MU, et al. DPP8 selective inhibitor tominostat as a novel and broad-spectrum anticancer agent against hematological malignancies. Cells 2023;12:1100.
Bettecken A, Heß L, Hölzen L, Reinheckel T Dipeptidyl-aminopeptidases 8 and 9 regulate autophagy and tamoxifen response in breast cancer cells. Cells 2023;12:2031.
Rabl J, Bunker RD, Schenk AD, Cavadini S, Gill ME, Abdulrahman W, et al. Structural basis of BRCC36 function in DNA repair and immune regulation. Mol Cell. 2019;75:483–97.
Walden, Tian M, Ross L, Sykora RL, Byrne UM, Hesketh EL DP, et al. Metabolic control of BRISC-SHMT2 assembly regulates immune signalling. Nature. 2019;570:194–9.
Yan K, Li L, Wang X, Hong R, Zhang Y, Yang H, et al. The deubiquitinating enzyme complex BRISC is required for proper mitotic spindle assembly in mammalian cells. J Cell Biol. 2015;210:209–24.
Zheng H, Gupta V, Patterson-Fortin J, Bhattacharya S, Katlinski K, Wu J, et al. A BRISC-SHMT complex deubiquitinates IFNAR1 and regulates interferon responses. Cell Rep. 2013;5:180–93.
Donaghy R, Han X, Rozenova K, Lv K, Jiang Q, Doepner M, et al. The BRISC deubiquitinating enzyme complex limits hematopoietic stem cell expansion by regulating JAK2 K63-ubiquitination. Blood. 2019;133:1560–71.
Huttlin EL, Bruckner RJ, Navarrete-Perea J, Cannon JR, Baltier K, Gebreab F, et al. Dual proteome-scale networks reveal cell-specific remodeling of the human interactome. Cell. 2021;184:3022–40.e3028.
Garcia-Diaz A, Shin DS, Moreno BH, Saco J, Escuin-Ordinas H, Rodriguez GA, et al. Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell Rep. 2017;19:1189–1201.
Curran MA, Montalvo W, Yagita H, Allison JP. PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc Natl Acad Sci USA. 2010;107:4275–80.
Fan X. Recent highlights of cancer immunotherapy. Holistic Integrative Oncology. 2023;2:37.
Carlino MS, Larkin J, Long GV. Immune checkpoint inhibitors in melanoma. Lancet. 2021;398:1002–14.
Postow MA, Callahan MK, Wolchok JD. Immune checkpoint blockade in cancer therapy. J Clin Oncol. 2015;33:1974–82.
Curnock AP, Bossi G, Kumaran J, Bawden LJ, Figueiredo R, Tawar R, et al. Cell-targeted PD-1 agonists that mimic PD-L1 are potent T cell inhibitors. JCI insight 2021;6:e152468.
Yamaguchi H, Hsu JM, Yang WH, Hung MC. Mechanisms regulating PD-L1 expression in cancers and associated opportunities for novel small-molecule therapeutics. Nat Rev Clin Oncol. 2022;19:287–305.
Islam MK, Stanslas J. Peptide-based and small molecule PD-1 and PD-L1 pharmacological modulators in the treatment of cancer. Pharmacol Ther. 2021;227:107870.
Zhu Y, Chen M, Xu D, Li TE, Zhang Z, Li JH, et al. The combination of PD-1 blockade with interferon-α has a synergistic effect on hepatocellular carcinoma. Cell Mol Immunol. 2022;19:726–37.
Liang Y, Tang H, Guo J, Qiu X, Yang Z, Ren Z, et al. Targeting IFNα to tumor by anti-PD-L1 creates feedforward antitumor responses to overcome checkpoint blockade resistance. Nat Commun. 2018;9:4586.
Hiam-Galvez KJ, Allen BM, Spitzer MH. Systemic immunity in cancer. Nat Rev Cancer. 2021;21:345–59.
Yi M, Niu M, Xu L, Luo S, Wu K. Regulation of PD-L1 expression in the tumor microenvironment. J Hematol Oncol. 2021;14:10.
Wu M, Huang Q, Xie Y, Wu X, Ma H, Zhang Y, et al. Improvement of the anticancer efficacy of PD-1/PD-L1 blockade via combination therapy and PD-L1 regulation. J Hematol Oncol. 2022;15:24.
Griswold AR, Ball DP, Bhattacharjee A, Chui AJ, Rao SD, Taabazuing CY, et al. DPP9’s enzymatic activity and not its binding to CARD8 inhibits inflammasome activation. ACS Chem Biol. 2019;14:2424–9.
Zolg S, Donzelli L, Geiss-Friedlander R. N-terminal processing by dipeptidyl peptidase 9: cut and go! Biochimie. 2024;226:180–92.
Nguyen JMH, Zolg S, Geiss-Friedlander R, Gorrell MD. The multifunctional regulatory post-proline protease dipeptidyl peptidase 9 and its inhibitors: new opportunities for therapeutics. Cell Mol Life Sci. 2025;82:187.
Shimshon A, Dahan K, Israel-Gueta M, Olmayev-Yaakobov D, Timms RT, Bekturova A, et al. Dipeptidyl peptidases and E3 ligases of N-degron pathways cooperate to regulate protein stability. J Cell Biol 2024;223:e202311035.
Deutsch EW, Bandeira N, Perez-Riverol Y, Sharma V, Carver JJ, Mendoza L, et al. The ProteomeXchange consortium at 10 years: 2023 update. Nucleic Acids Res. 2023;51:D1539–48.
Funding
This work was supported in part by the National Natural Science Foundation of China (No. 81902614 to K.C.; No. 92357301, 32370726, and 81972396 to C.W.; No. 82473192, 82172741 to D.Y.; No. 32500513 to Y.C), the State Key Development Programs of China (No. 2022YFA1104200 to C.W.), the Natural Science Foundation of Shanghai (22ZR1406600 to C.W) and Science and Technology Research Program of Shanghai (No. 9DZ2282100 to C.W).
Author information
Authors and Affiliations
Contributions
W.Z., Y.W., and K.C. performed most of the experiments and contributed to writing and revising the manuscript. C.W., K.C., and D.Y. conceived the study, participated in its design and coordination and helped draft the manuscript. T.C., H.Z., Y.C., and S.Z. collected clinical samples. T.F. and W.W. analyzed part of the data. All the authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics
All animal experiments were approved by the Ethics Review Committee for Animal Experimentation of Fudan University Shanghai Cancer Center (No: IACUC-S2024-0273). Human tissue samples were obtained with written informed consent, and the study was approved by the Ethics Review Committee of Fudan University Shanghai Cancer Center (No: 050432-4-2108*).
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, W., Wang, Y., Feng, T. et al. DPP9 inhibition boosts antitumor immunity by disrupting BRISC-mediated PD-L1 expression in clear cell renal cell carcinoma. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01704-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01704-x