Abstract
As a crucial kinase in the host antiviral innate immune signaling pathway, TANK-binding kinase 1 (TBK1) is strictly regulated by various posttranslational modifications. Previous studies have demonstrated that the stability of TBK1 can be compromised through autophagy; however, the precise mechanisms involved in the regulation of TBK1 degradation remain unclear. In this study, we revealed that the E3 ubiquitin ligase Stub1 can inhibit the production of type I interferon (IFN-I) by targeting TBK1, thereby preventing host antiviral responses. Mechanistically, TBK1 is targeted for degradation by chaperone-mediated autophagy (CMA), which depends on its three typical motifs. This process relies on the interaction between TBK1 and Stub1. Simultaneously, Stub1 catalyzes the polyubiquitination of TBK1 at lysine 344 (K344), which is linked to K27. The ubiquitinated TBK1 is recognized by heat shock cognate protein 70 (HSC70/HspA8), resulting in autophagic degradation via CMA mediated by LAMP2A. Compared with wild-type mice, Stub1-deficient mice exhibit increased resistance to VSV and HSV-1 infection, accompanied by increased expression levels of type I IFN. Overall, our findings reveal a TBK1-Stub1 axis in the RIG-I and cGAS-STING pathways, highlighting the effects of CMA on host antiviral innate immune responses.
Similar content being viewed by others
Introduction
The innate immune response serves as the first line of defense against pathogenic microorganisms [1]. During a viral infection, host cell pattern recognition receptors (PRRs), including Toll-like receptors (TLRs) [2], RIG-I-like receptors (RLRs) [3, 4], NOD-like receptors (NLRs) [5, 6], C-type lectin receptors (CLRs) [7], cyclic GMP-AMP synthase and stimulator of interferon genes (cGAS-STING) pathways [8], and AIM2-like receptors (ALRs) [9], recognize various pathogen-associated molecular patterns (PAMPs), such as viral DNA, dsRNA, ssRNA, or surface glycoproteins, to initiate host antiviral responses. In the RIG-I-MAVS [10, 11] and cGAS-STING signaling pathways, a series of complex cascade reactions are activated by kinases, such as TBK1 and IKKƐ, leading to the production of IFN-I [12], which binds IFN receptors (IFNARs) to trigger the expression of IFN-stimulated genes (ISGs), thereby initiating host antiviral responses.
Previous studies have demonstrated that TBK1 plays a crucial role in IFN-I production. Upon viral infection, TBK1 is activated, leading to the phosphorylation and translocation of IFN regulatory factor 3 (IRF3), which in turn results in the transcriptional activation of IFNs [13]. TBK1 activity is tightly regulated by various posttranslational modifications, including phosphorylation [14], ubiquitination [15], acylation, dimerization [16], and acetylation [17]. For example, TBK1 can be degraded through various E3 ubiquitin ligase-mediated ubiquitin-proteasome systems involving NLRP4, DTX4 [18], TRIP [19], TRIM27 [20], USP38 [21], NLRP14 [22], and TRAF3IP3 [23], which impedes IFN-I production.
Autophagy is a highly conserved intracellular degradation process that aims to break down cytoplasmic components, organelles, and invading microorganisms [24, 25]. There are three forms of autophagy: macroautophagy [26], microautophagy [27], and chaperone-mediated autophagy (CMA) [28]. Accumulating evidence has shown that the autophagic substrate, also known as the “cargo”, can be translocated to the lysosome for degradation through one of these three forms of autophagy. TBK1 has also been reported to play a significant role in regulating autophagy. For example, TBK1 is required for the assembly of autophagic complexes, leading to the formation of the FIP200-ULK1-ATG13 complex [29]. In addition, active TBK1 enhances selective autophagy by phosphorylating cargo receptors such as OPTN, p62, NDP52, and TAX1BP1 [30]. Recently, numerous studies have shown that the production of IFN-I can be subtly regulated by the selective autophagic degradation of key PRRs, including aptamers, and transcription factors (such as RIG-I [31], cGAS [32], AIM2 [33], MAVS [34], STING [35], TRIF [36], and IRF3 [37]). Furthermore, the E3 ubiquitin ligase NEDD4 has been found to catalyze the polyubiquitination of TBK1. Polyubiquitinated TBK1 is recognized by the cargo receptor NDP52 and then degraded through selective autophagy [38].
As a crucial E3 ligase, Stub1 (STIP1 homology and U-box-containing protein 1) not only plays important roles in immune cell differentiation, maturation, and tumor immunity but also contributes to antiviral immune responses through the regulation of the degradation of key molecules in the RIG-I signaling pathway [39, 40]. Stub1 has two key functional regions: an N-terminal tetrapeptide repeat (TPR) domain and a highly conserved C-terminal U-box domain. Previous studies have shown that the TRR domain of Stub1 interacts with molecular chaperone proteins, such as heat-shock protein 70 (HSP70) and HSP90, whereas the U-box domain of Stub1 has E3 ligase activity and is involved in protein degradation via the ubiquitin-proteasome system (UPS) [41, 42]. However, whether Stub1 affects host antiviral immunity via chaperone-mediated autophagy remains unclear.
Here, we revealed a mechanism by which Stub1 negatively regulates the production of IFN and enhances the replication of vesicular stomatitis virus (VSV) and herpes simplex virus type 1 (HSV-1). Stub1 binds to TBK1 and catalyzes its K27 polyubiquitination through its E3 ligase activity. Polyubiquitinated TBK1 is recognized by HSC70 through the KFDKQ motif of TBK1, ultimately leading to chaperone-mediated autophagic degradation. Our findings reveal that Stub1 acts as a “molecular brake” to regulate the type I IFN response by targeting TBK1 through chaperone-mediated autophagy.
Results
Stub1 negatively regulates type I IFN signaling
To assess the effects of Stub1 on RIG-I- and cGAS-STING-mediated type I IFN signaling pathways, HeLa cells were transfected with the indicated luciferase reporters and then infected with the RNA virus vesicular stomatitis virus (VSV) or the DNA virus herpes simplex virus type I (HSV-1). Our results indicate that ectopic expression of Stub1 significantly inhibits VSV infection-mediated IFN-β production (Fig. 1A–C) as well as that of HSV-1 (Fig. 1E–G). In addition, Stub1 inhibited the activation of IFN-β induced by intracellular (IC) poly(I:C) as well as poly (dA: dT) treatment (Fig. S1A–D). Furthermore, we found that the overexpression of Stub1 inhibited the activities of promoters, including those of IFN-α, IFN-stimulated response element (ISRE), and the IFN-stimulated factor ISG54, induced by VSV and HSV-1 infection in a dose-dependent manner (Fig. S1E–J). Consistently, Stub1 inhibited the phosphorylation of IRF3 during VSV and HSV-1 infection (Fig. 1D, H).
HEK293T cells were transfected with an IFN-β luciferase reporter and a Renilla-TK reporter, together with increasing amounts (0, 100, 200, and 400 ng) of a plasmid expressing Myc-Stub1 for 12 h, and then the cells were infected with VSV (0.1 MOI) (A) or HSV-1 (10 MOI) (E) for another 12 h. Luciferase activities were then analyzed. HEK293T cells were transfected with increasing amounts (0, 100, 200, or 400 ng) of a plasmid expressing Myc-Stub1 for 12 h, after which the cells were infected with VSV (0.1 MOI) (B) or HSV-1 (10 MOI) (F) for another 12 h, and the mRNA levels of Ifnb1 were subsequently analyzed via qPCR. C, G ELISA of IFN-β in the supernatant shown in (B, F). HeLa cells were transfected with a plasmid expressing Myc-Stub1 for 12 h, after which the cells were left uninfected or infected with VSV (0.1 MOI) (D) or HSV-1 (10 MOI) (H). Then, IRF3, phosphorylated IRF3, the VSV protein, and GAPDH were analyzed by immunoblotting. The band intensity of the IRF3-p/IRF3 immunoblotting result was quantified via ImageJ. I HEK293T-stub1+/+ and HEK293T-stub1−/− cells were transfected with an IFN-β Luc reporter, a Renilla-TK reporter, and an empty vector or a plasmid expressing Flag-Stub1. The Luc activity of the IFN-β promoter-reporter was detected. J HEK293T-stub1+/+ and HEK293T-stub1−/− cells were transfected with an empty vector or a plasmid expressing Flag-Stub1. The mRNA levels of Ifnb1 in the cells were subsequently analyzed via qPCR. The results are presented relative to those of untreated cells transfected with an empty vector. K ELISA of IFN-β in the supernatant shown in (J). HEK293T-stub1+/+ and HEK293T-stub1−/− cells were infected with VSV (0.1 MOI) (L) or HSV-1 (10 MOI) (M) for 12 h and then subjected to immunoblot analysis with antibodies against IRF3, phosphorylated IRF3, VSV protein, HSV-1 protein, and Stub1. N An IFN sensitivity assay was performed to test the inhibitory effects of Stub1 on the replication of VSV-GFP (0.1 MOI) or HSV-1-GFP (10 MOI). VSV-GFP and HSV-1-GFP replication was analyzed via fluorescence microscopy, and quantitative analysis was performed by measuring the proportion of GFP-positive cells. O Luciferase activity of the ISG56-Luc reporter in HEK293T cells transfected with a Renilla-TK reporter together with different amounts (0, 100, 200, and 400 ng) of a plasmid expressing Myc-Stub1 and then stimulated with IFN-α (500 IU/mL) for 12 h. Ns not significant (P > 0.05), *0.01 < P < 0.05, **P < 0.01, and ***P < 0.001 (two-tailed Student’s t test). The data are representative of three independent experiments with three biological replicates (the mean ± standard deviation of triplicate assays or are representative of three independent experiments with similar results) (A–O).
To further validate these observations, a stub1 gene knockout (KO) HEK293T cell line (293T-stub1−/−) was generated via CRISPR-Cas9 to examine the function of Stub1 in vitro. We found that VSV or HSV-1 infection increased IFN-β production in Stub1-deficient cells (Fig. 1I–K). Moreover, the phosphorylation levels of IRF3 in HEK293T-stub1−/− cells were greater than those in HEK293T-stub1+/+ cells infected with VSV or HSV-1, although the total protein levels of IRF3 were not affected (Fig. 1L, M). To further demonstrate the role of Stub1 in type I IFN signaling, an IFN sensitivity assay was performed. As shown in Fig. 1N, Stub1 inhibited the replication of VSV-GFP and HSV-1-GFP by suppressing the production of IFN-I. To test whether Stub1 has a potential effect on the IFN signaling pathway, we assessed the effect of Stub1 on ISG56 promoter activity in IFN-treated cells. The results demonstrated that Stub1 did not affect IFN-induced ISG56 promoter activity (Fig. 1O).
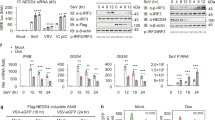
To study the effects of Stub1 on host antiviral responses, especially type I IFN production in vivo, Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice were generated and validated by sequencing and Western blot analysis (Fig. S2A–E). As demonstrated in Fig. 2A, B, primary bone marrow-derived macrophages from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice were treated with poly(I:C), poly (dA: dT), or infected with EMCV, VSV, PRV, or HSV-1, respectively. A comparison of primary bone marrow-derived macrophages (BMDMs) from Stub1fl/fl mice with those from Stub1fl/fl Lyz2-Cre mice revealed that the former presented reduced mRNA expression of Ifnb1 and Isg56. In accordance with these findings, the protein levels of IFN-β in the culture medium of primary bone marrow-derived macrophages from Stub1fl/fl Lyz2-Cre mice were also found to be considerably elevated.
A, B Primary bone marrow-derived macrophages isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice were transfected with poly(I:C) (1 μg/mL) or poly (dA: dT) (1 μg/mL) or infected with VSV (1 MOI), EMCV (1 MOI), HSV-1 (10 MOI), or PRV (1 MOI) for 12 h. The mRNA levels of Ifnb1 (A) and Isg56 (B) were analyzed via qPCR. C ELISA of IFN-β in the supernatant shown in (A, B). qPCR analysis of the mRNA levels of Ifnb1 (D), Isg56 (E), and Mx1 (F), the genomic copy numbers (G) and the viral titers measured by TCID50 assay of VSV (H) in peritoneal macrophages isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice infected with VSV (MOI of 1) for 0, 4, 8, or 12 h. qPCR analysis of the mRNA levels of Ifnb1 (I), Isg56 (J), and Mx1 (K) and the genomic copy numbers (L) and the viral titers measured by a TCID50 assay of HSV-1 (M) in peritoneal macrophages isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice infected with HSV-1 (MOI of 10) for 0, 4, 8, or 12 h. qPCR analysis of the mRNA levels of Isg56 (N) and Mx1 (O) in BMDMs isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice stimulated with IFN-α (500, 1000, or 2000 IU/mL) for 12 h. qPCR analysis of the mRNA levels of Isg56 (P) and Mx1 (Q) in peritoneal macrophages isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice stimulated with IFN-α (500, 1000, 2000 IU/mL) for 12 h. Data are representative of three independent experiments with three biological replicates (mean ± SD). NS not significant (P > 0.05), **P < 0.01, ***P < 0.001.
To determine whether the mRNA expression of Ifnb1 could be affected by Stub1, peritoneal macrophages isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice were infected with VSV or HSV-1 for 0, 4, 8, or 12 h. Compared with peritoneal macrophages derived from Stub1fl/fl mice, peritoneal macrophages from Stub1fl/fl Lyz2-Cre mice infected with the two viruses presented increased mRNA expression of Ifnb1, Isg56, and Mx1 at various time points (Fig. 2D–F, H–J). Furthermore, the viral genomic copy number and titer in peritoneal macrophages from Stub1fl/fl Lyz2-Cre mice were significantly lower than those in peritoneal macrophages from Stub1fl/fl mice (Fig. 2G, H, K–M). Consistent with the results shown in Fig. 1O, the results of the qPCR analysis revealed that Stub1 knockout did not affect the mRNA levels of IFN-induced Isg56 or Mx1 in either BMDMs or peritoneal macrophages isolated from Stub1fl/fl mice or Stub1fl/fl Lyz2-Cre mice after IFN-α treatment (Fig. 2N–Q). These collective results indicate that the knockout of Stub1 resulted in an increase in the production of type I IFNs and ISGs, thereby inhibiting viral replication.
To directly test whether the enhanced antiviral state resulting from Stub1 deficiency is mediated by IFN-I signaling, we isolated BMDMs from Stub1fl/fl mice or Stub1fl/fl Lyz2-Cre mice and performed IFNAR antibody blockade experiments. Treatment with anti-IFNAR antibodies abrogated the differences in IFN-β induction, ISG expression, and viral replication between Stub1-KO and wild-type macrophages (Fig. S3A–C), confirming the essential role of IFN-I signaling in this phenotype. Furthermore, we assessed the specificity of Stub1’s action. Stub1 deficiency enhanced Ifnb1 mRNA production induced by both the viral mimic poly(I:C) and the bacterial LPS, but did not affect the expression of the proinflammatory cytokine Tnf-α (Fig. S3D, E), indicating that Stub1 selectively inhibits the TRIF-TBK1-IRF3 axis without affecting the MyD88-NF-κB pathway, consistent with its TBK1-centric mechanism.
Stub1 suppresses type I IFN signaling by targeting TBK1
To clarify the underlying mechanisms by which Stub1 negatively regulates IFN-I production, we assessed the effect of Stub1 on IFN-β reporter activation induced by key molecules in the RIG-I and cGAS-STING signaling pathways in HEK293T cells. As shown in supplementary Fig. S3A–G, ectopically expressed Stub1 significantly inhibited the IFN-β reporter activation induced by RIG-I, MDA5, MAVS, TBK1, and cGAS+STING in a dose-dependent manner but not by IKKƐ or IRF3-5D (a constitutively active variant of IRF3) (Fig. S4A–G). In accordance with these results, qPCR analysis revealed that the ectopic expression of Stub1 inhibited the IFN-β reporter activation of Ifnβ1 induced by RIG-I, MDA5, MAVS, TBK1, and cGAS+STING but not by IKKε or IRF3-5D (Fig. 3A). Subsequently, HEK293T-stub1−/− and HEK293T-stub1+/+ cells were transfected with plasmids expressing RIG-I, MDA5, cGAS+STING, MAVS, TBK1, or IKKε and IRF3-5D. We found that IFN-β reporter activation was significantly increased by RIG-I, MDA5, MAVS, TBK1, and cGAS+STING in HEK293T-stub1−/− cells compared with that in HEK293T-stub1+/+ cells but not by IKKε or IRF3-5D (Fig. 3B). Taken together, these results indicate that Stub1 may negatively regulate IFN-I production by targeting TBK1.
A HEK293T cells were transfected with an IFN-β luciferase reporter and a Renilla-TK reporter and a plasmid expressing RIG-I, MDA-5, MAVS, TBK1, IKKε, IRF3-5D, or cGAS+STING, along with an empty vector or a plasmid expressing Flag-Stub1. The Luc activity of the IFN-β promoter-reporter was detected. B HEK293T-stub1+/+ and HEK293T-stub1−/− cells were transfected with an IFN-β luciferase reporter, a Renilla-TK reporter, and a plasmid expressing RIG-I, MDA-5, MAVS, TBK1, IKKε, IRF3-5D, or cGAS+STING. The luciferase activity of the IFN-β promoter reporter was detected. C HEK293T cells were transfected with plasmids expressing Myc-Stub1 alone or together with plasmids expressing Flag-RIG-I, MDA5, MAVS, TBK1, IKKε, IRF3, cGAS, or STING for 24 h. The cell lysates were harvested and used for Co-IP. Input and IP complexes were analyzed by Western blotting using antibodies against Flag and Myc. D HEK293T cells were transfected with plasmids expressing HA-Stub1 alone or together with a plasmid expressing Flag-TBK1. At 24 hpt, the cell lysates were harvested and used for reverse Co-IP. Input and IP complexes were analyzed by Western blotting using antibodies against Flag and HA. E His-TBK1 was incubated with GST or GST-Stub1 for 30 min, and the protein-protein interactions were then confirmed via a GST pull-down assay. F Bimolecular fluorescence complementation (BIFC) analysis of TBK1 and Stub1. HA-Stub1-VN, Stub1 fused with the N-terminus of Venus; Flag-TBK1-VC, TBK1 fused with the C-terminus of Venus. G, H Co-IP and immunoblot analysis of the interaction of endogenous Stub1 and TBK1 in THP-1 cells infected with VSV (0.1 MOI) (F) or HSV-1 (10 MOI) (G) for 12 h. Co-IP was subsequently performed with an anti-stub1 antibody. IgG was used as a negative control. I THP-1 cells were mock-infected or infected with VSV (0.1 MOI) or HSV-1 (10 MOI) for 12 h. The subcellular localization of endogenous Stub1 and TBK1 was analyzed via fluorescence microscopy. Ns not significant (P > 0.05), *0.01 < P < 0.05, **P < 0.01, ***P < 0.001 (two-tailed Student’s t test). The data represent three independent experiments with three biological replicates (means ± SDs) or three independent experiments with similar results (A–I).
To further identify the targets of Stub1, we tested the interactions between stub1 and RIG-I, MDA5, MAVS, TBK1, IKKε, IRF3, cGAS, and STING. Coimmunoprecipitation (co-IP) results revealed that Stub1 interacted with RIG-I, MAVS, and TBK1 but not with the other proteins (Fig. 3C). Because RIG-I and MAVS have been investigated by other researchers [39, 40], we focused on TBK1 in this study. As shown in Fig. 3D, Stub1 coimmunoprecipitated with TBK1 when the indicated proteins were co-expressed in HEK293T cells. To determine whether Stub1 physically interacted with TBK1 in vitro, we performed pull-down assays and found that His-TBK1 could be pulled down by GST-Stub1 but not by GST (Fig. 3E). We further demonstrated the interaction between Stub1 and TBK1 via bimolecular fluorescence complementation analysis (BIFA) (Fig. 3F). In addition, we also found that endogenous Stub1 interacted with endogenous TBK1 in HEK293T cells following both VSV infection and HSV-1 infection (Fig. 3G, H). Interestingly, when the lysosomal inhibitor CQ is added under conditions conducive to viral infection, the interaction between Stub1 and TBK1 is enhanced. These findings demonstrate that Stub1 plays a role in stabilizing TBK1 expression under such conditions (Fig. 3G, H). Immunofluorescence staining also revealed that Stub1 colocalized with TBK1 in the cytoplasm (Fig. 3I). To further substantiate that Stub1 limits IFN-I production primarily by targeting TBK1, we employed three distinct viral infection models alongside knockdown of key signaling molecules. Notably, while knockdown of RIG-I or MAVS did not abolish the inhibitory effect of Stub1 on IFN-β production induced by VSV, EMCV, or HSV-1, knockdown of TBK1 almost completely abolished this inhibition (Fig. S5A–G). These results reinforce the conclusion that Stub1 suppresses type I interferon production predominantly via TBK1.
Moreover, Co-IP revealed that Stub1 interacted with the ULD (ubiquitin-like domain) of TBK1 through its U-box domain (Fig. S6A–D). Collectively, these results suggest that Stub1 suppresses type I IFN signaling by targeting TBK1.
Stub1 promotes the autophagic degradation of TBK1
Stub1 can promote the degradation of RIG-I [15] and MAVS [40]. We first assessed the effects of Stub1 on the degradation of RIG-I, TBK1, and MAVS in HEK293T cells. We found that increasing amounts of Stub1 significantly decreased the protein levels of RIG-I and TBK1 but not those of MAVS (Fig. 4A). Furthermore, we did not observe significant Stub1-induced MAVS degradation in HEK293T and HeLa cell systems (Fig. S7A–C). This suggests that Stub1’s regulatory function may be highly context-dependent, potentially varying across cell types, species, or specific cellular states. However, the mRNA level of TBK1 remained unchanged (Fig. 4C), suggesting that Stub1 promotes TBK1 protein degradation (Fig. 4B). The CHX-chase assay results revealed that, compared with the control, Stub1 overexpression reduced the TBK1 half-life in HEK293T cells (Fig. 4D). Furthermore, the knockout of Stub1 resulted in TBK1 stability during VSV infection (Fig. 4E). The CHX-chase assay results revealed that the degradation rate of TBK1 in 293T-stub1−/− cells was lower than that in WT cells (Fig. 4F, G).
A HEK293T cells were transfected with increasing amounts (0, 0.5, 1, and 2 µg) of a plasmid expressing HA-Stub1, and the protein was harvested for immunoblot analysis. B, C HEK293T cells were transfected with Flag-TBK1, together with increasing amounts (wedge) of HA-Stub1 expression vector, and the protein was harvested for immunoblot analysis. Flag-TBK1 expression was normalized to that of GAPDH (below) (B). C Right, RT-PCR analysis of TBK1 mRNA; GAPDH mRNA served as a loading control. D Immunoblot analysis of protein extracts from HEK293T cells transfected with Flag-TBK1 or HA-Stub1 and treated with cycloheximide (CHX) (100 μg/mL) for the indicated durations (0, 4, 8, or 12 h). Flag-TBK1 expression was normalized to that of GAPDH (below). E Protein lysates of HEK293T-stub1+/+ and HEK293T-stub1−/− cells infected with VSV (MOI = 0.1) at the indicated time points were immunoblotted with the indicated antibodies. F, G Immunoblot analysis of protein extracts from HEK293T-stub1+/+ and HEK293T-stub1−/− cells treated with cycloheximide (CHX) (100 μg/mL) for the indicated durations. H Immunoblot analysis of protein lysates of HEK293T cells transfected with vectors expressing Myc-Stub1-WT, Myc-Stub-K31A, Myc-Stub-H261A, or the Myc-Stub1-DM mutant, together with plasmids encoding Flag-TBK1. I Analysis of the Luc activity of the IFN-β promoter reporter in HEK293T cells transfected with vectors expressing Myc-Stub1-WT, Myc-Stub-K31A, Myc-Stub-H261A, or the Myc-Stub1-DM mutant together with plasmids encoding Flag-TBK1. J HEK293T cells were transfected with a plasmid encoding Flag-TBK1 together with plasmids encoding EV or Myc-Stub1, followed by treatment with MG132 (10 μM), 3MA (10 mM), CQ (50 μM), or bafilomycin A1 (Baf A1) (0.2 μM) for 6 h. The cell lysates were then analyzed by immunoblotting. Flag-TBK1 expression was normalized to that of GAPDH (below). K Protein lysates of WT, BECLN1 KO, ATG5 KO, and ATG7 KO 293T cells transfected with plasmids encoding EV or HA-Stub1. The data represent three independent experiments with three biological replicates (means ± SDs) or three independent experiments with similar results (A–K). Statistical significance was determined by two-tailed Student’s t test or one-way ANOVA followed by the Bonferroni post hoc correction, as appropriate. *P < 0.05, **P < 0.01, ***P < 0.001.
To investigate whether the enzyme activity of Stub1 is required to promote TBK1 degradation, we generated four plasmids expressing catalytically inactive Stub1 mutants, namely, Stub1-wt, Stub1-K31A, Stub1-H261A, and Stub1-DM (double mutants K31A and H261A). Stub1-K31A has a point mutation in the TPR chaperone-binding domain, which prevents it from binding to chaperones; consequently, its ability to bind substrates is greatly reduced. Stub1-H261A has a point mutation in the U-box domain of Stub1 and, therefore, is deficient in E3 ubiquitin ligase activity; thus, its ability to ubiquitinate the substrate for degradation is diminished [43]. We observed that the TBK1 protein was no longer degraded when only TBK1 and Stub1-DM were co-expressed in the cells (Fig. 4H). These results suggest that Stub1 may inhibit IFN-I production through its enzymatic activity. The results of the dual-luciferase reporter gene assay demonstrated that Stub1 inhibits TBK1-mediated IFN-β promoter activity through its E3 ubiquitin ligase enzymatic activity and chaperone binding ability (Fig. 4I). Our research revealed that when Stub1 undergoes a single-site mutation, it cannot fully restore TBK1 expression. However, when the molecular chaperone-binding site of Stub1 is mutated, and the autophagy-related protein Beclin1 is simultaneously knocked out, TBK1 expression is fully restored (Fig. S7D). Therefore, we hypothesize that Stub1 may promote the ubiquitination of TBK1 through its E3 ubiquitin ligase activity, leading to its degradation via the autophagy pathway. Notably, when the E3 ubiquitin ligase activity site of Stub1 is mutated, and the molecular chaperone protein HSC70 is simultaneously knocked out, TBK1 expression is fully restored (Fig. S7E). Therefore, we hypothesize that other E3 ubiquitin ligases may promote TBK1 K27-linked ubiquitination, followed by interaction with HSC70 to achieve degradation. To establish a direct correlation between phenotype and the TBK1-CMA axis, BMDMs were isolated from Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice. The Stub1-WT gene and its double-site inactivating mutant Stub1-DM were reintroduced into the cells via lentivirus, and TBK1 expression was detected via Western blot analysis. The results demonstrated that TBK1 expression recovered to stable levels following Stub1 deletion. Reoverexpression of wild-type Stub1 led to a significant reduction in TBK1 expression, whereas overexpression of the Stub1-DM plasmid, which carries an enzyme active site mutation, resulted in the restoration of TBK1 expression to stable levels (Fig. S7F).
Protein degradation primarily occurs via two pathways: the proteasome-dependent pathway and the lysosome-dependent pathway [44]. To further investigate Stub1-mediated TBK1 degradation, Stub1 and TBK1 were co-expressed in HEK293T cells, which were then treated with a proteasome inhibitor (MG132), a lysosomal inhibitor (chloroquine (CQ)), a phosphatidylinositol 3-kinase inhibitor (3-methyladenine (3-MA)), or an autophagosome-lysosome fusion inhibitor, bafilomycin A1 (Baf-A1). As shown in Fig. 4J, only CQ treatment suppressed TBK1 degradation. Consistently, Stub1 was still able to trigger TBK1 degradation after Stub1 was overexpressed in ATG5-, ATG7-, and Beclin1-KO cells (Fig. 4K). We also incorporated inhibitors such as CQ and MG132 into time-dependent CHX-mediated degradation assays. The results revealed that TBK1 protein expression recovery was observed exclusively in chloroquine-treated cells (Fig. S7G). Taken together, these results suggest that Stub1 specifically degrades TBK1 via the lysosomal pathway during viral infection.
Stub1 promotes the autophagic degradation of TBK1 by enhancing K27 polyubiquitination at the K344 site of TBK1
The E3 ubiquitin ligase activity of Stub1 has been implicated in TBK1 degradation. To determine whether Stub1 directly ubiquitinates TBK1, we examined the effect of Stub1 overexpression on TBK1 polyubiquitination and observed a significant increase. Further analysis revealed that Stub1 specifically promotes K27-linked polyubiquitination of TBK1, with no apparent effect on other types of ubiquitin linkages (Fig. S8A). We then conducted in vitro ubiquitination experiments to elucidate the role of Stub1 in the ubiquitination process of TBK1. The smeared bands detected by ubiquitin antibodies following Western blotting indicated that polyubiquitin chains of different molecular weights had formed (Fig. S8B). In contrast, no smeared bands were detected in the reaction containing E1, E2, or His-TBK1 but lacking GST-Stub1 (Fig. S8B), indicating that Stub1 is a specific ubiquitin ligase for TBK1. Stub1 carrying mutations in the ubiquitin ligase active site, such as Stub1-H261R or Stub1-DM, lost ubiquitin ligase activity, as no diffuse bands of polyubiquitin were detected in these reactions (Fig. S8C), supporting the importance of Stub1 enzyme activity for TBK1 ubiquitination. In addition, we found that TBK1 underwent K27-linked polyubiquitination approximately 4 h after viral infection, which was markedly enhanced by the overexpression of Stub1 (Fig. 5A). We further explored the role of the enzymatic activity of Stub1 in promoting the K27-linked polyubiquitination of TBK1 and revealed that the overexpression of Stub1 and Stub1-K31A, but not Stub1-H261A, promoted the K27-linked polyubiquitination of TBK1 (Fig. 5B, C). Consistently, the depletion of Stub1 resulted in decreased K27-linked polyubiquitination of TBK1 during VSV infection (Fig. 5D). To further confirm that endogenous TBK1 undergoes covalent modification via K27-linked polyubiquitin chains, we immunoprecipitated endogenous TBK1 from Stub1fl/fl and Stub1fl/fl Lyz2-Cre BMDMs. Western blot analysis of immunoprecipitates with a K27-linkage-specific antibody revealed robust infection-induced K27-linked polyubiquitination of TBK1 in control cells, which was significantly attenuated in Stub1-deficient macrophages (Fig. 5E). In contrast, Stub1 deficiency had no significant effect on K48-linked ubiquitination. Together, these data provide direct endogenous evidence that Stub1 specifically mediates K27-linked polyubiquitination of TBK1.
A Protein lysates of HEK293T cells transfected with plasmids expressing HA-K27-Ub together with an EV or Myc-Stub1 expression vector, followed by VSV (MOI = 0.1) infection for various durations, were subjected to immunoprecipitation with an anti-TBK1 antibody and immunoblot analysis with the indicated antibodies. B Co-IP and immunoblot analysis of HEK293T cells transfected with Flag-TBK1 and HA-K27-Ub, together with vectors for WT Myc-Stub1 or its mutants. Protein lysates were harvested after CQ (50 μM) treatment (6 h) for immunoblotting with the indicated antibodies. C HEK293T cells were transfected with vectors for WT Myc-Stub1 or its mutants, followed by SeV (MOI = 0.1) infection for 12 h. Protein lysates were immunoprecipitated with an anti-TBK1 antibody and immunoblotted with the indicated antibodies. D Co-IP and immunoblot analysis of HEK293T-stub1+/+ and HEK293T-stub1−/− cells transfected with Flag-TBK1 and HA-K27-Ub, together with vectors for WT Myc-Stub1 or its mutants. Protein lysates were harvested after CQ (50 μM) treatment (6 h) for immunoblotting with the indicated antibodies. E Co-IP analysis of the ubiquitination of endogenous TBK1 in BMDMs from Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice that were mock infected (mock) or infected with VSV for 12 h. F Analysis of the ubiquitination sites of the ULC domain of TBK1 (above). Co-IP and immunoblot analysis of protein extracts of HEK293T cells transfected with vectors expressing WT Flag-TBK1 or its mutants (K323R, K341R, K344R, K372R), HA-K27-Ub and EV or vector for Myc-Stub1, in the presence of CQ (50 μM) for 6 h (below). G HEK293T cells were transfected with plasmids expressing WT Flag-TBK1 or its indicated mutants, together with plasmids for HA-Stub1, in the presence of CQ (50 μM) for 6 h, followed by immunoprecipitation with anti-Flag beads and immunoblotting with an anti-HA antibody. H Immunoblot analysis of protein lysates of HEK293T cells transfected with HA-Stub1 together with plasmids expressing WT Flag-TBK1 or the indicated mutants. The data are representative of three independent experiments with three biological replicates (means ± SDs) or represent three independent experiments with similar results (A–H).
We overexpressed Stub1, TBK1, and a series of ubiquitin (Ub) mutants in HEK293T cells: wild-type ubiquitin (WT-Ub), K27 point mutation ubiquitin (K27R-Ub), ubiquitin lacking lysine (K0), K27-only ubiquitin (K27-only Ub), and tandem ubiquitin (Ub2, Ub4)—the latter specifically designed to mimic short K27-linked chains in HEK293T cells. The stability of the TBK1 protein was assessed in total cell lysates via Western blot analysis. TBK1 was enriched via immunoprecipitation and analyzed for K27-linked ubiquitination by immunoblotting with a K27-linked-specific antibody. The results revealed that K27-only Ub (but not the ubiquitin-deficient mutants K27R or K0) significantly enhanced Stub1-mediated TBK1 degradation and induced pronounced K27-linked polyubiquitination. Preformed K27-linked dimeric ubiquitin (Ub2) was sufficient to drive degradation, indicating that short K27-specific conjugates constitute the minimal degradation signal (Fig. S8D). The temporal relationship between TBK1 activation and ubiquitination remains to be elucidated. Bone marrow-derived macrophages were isolated from Stub1fl/fl mice and subsequently infected with VSV at multiple time points. The present study detected TBK1 phosphorylation and ubiquitination and revealed that significant phosphorylation occurred 4 h post-VSV infection. Concurrently, TBK1 K27 ubiquitination also began to increase (Fig. S8E). On the basis of these findings, we propose that TBK1 phosphorylation may alter its conformation, thereby promoting K27 ubiquitination and subsequent degradation, which could modulate TBK1-mediated signaling pathways.
Notably, Stub1 interacts with the ULD domain of TBK1. To gain insight into the mechanism by which Stub1 regulates TBK1 ubiquitination, we analyzed and identified four lysine residues (K323, K341, K344, and K372) within the TBK1 ULD domain and generated mutants ranging from lysine (K) to arginine (R) (Fig. 5F). Our results revealed that Stub1 was unable to facilitate K27-linked polyubiquitination in the TBK1 K344R mutant (Fig. 5F). Furthermore, we discovered that the TBK1 K344R mutant did not interact with Stub1 (Fig. 5G) and that Stub1 was unable to promote the degradation of the TBK1 K344R mutant (Fig. 5H). To address the functional importance of ubiquitination at the K344 site of TBK1 in type I IFN signaling, we overexpressed Stub1 with a TBK1 or TBK1 K344R mutant in TBK1 KO cells and found that Stub1 failed to inhibit TBK1-mediated IFN production after VSV or HSV-1 infection when it was restored with the TBK1 K344R mutant in TBK1 KO cells (Fig. S8F, G). Collectively, these findings suggest that Stub1 specifically promotes the K27-linked polyubiquitination of TBK1 at K344.
Stub1 promotes TBK1 degradation through chaperone-mediated autophagy
To investigate the underlying mechanisms of the regulation of TBK1 degradation by Stub1, cells co-expressing Stub1 and TBK1 were treated with different indicated inhibitors. As shown in Fig. 4J, only the lysosomal inhibitor CQ, but not the autophagosome formation inhibitor 3-MA or the autophagosome-lysosome fusion inhibitor Baf-A1, suppressed TBK1 degradation, indicating that Stub1 promotes TBK1 degradation independent of macroautophagy. In addition, analysis of the human TBK1 amino acid sequence revealed the presence of three canonical putative KFERQ-like motifs (Fig. S9A, B). To confirm which CMA motif is key to Stub1-mediated TBK1 degradation, we constructed three CMA motif mutant plasmids of TBK1 (M1-VAALA, M2-QAAKA, and M3-KAAKA) and experimentally demonstrated that the KFDKQ motif regulates Stub1-mediated TBK1 degradation by CMA (Fig. S9C).
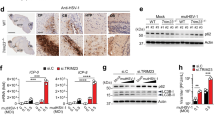
We inferred that Stub1 may promote TBK1 degradation in a CMA-dependent manner. Lysosome-associated membrane protein 2A (LAMP2A) and HSC70 are central proteins involved in CMA. Previous data have shown that HSC70 recognizes and interacts with the CMA motif of target proteins, whereas LAMP2A, located on the lysosome membrane, facilitates the transport of target proteins into lysosomes for degradation [45]. After the overexpression of TBK1, HSC70, and Stub1 in cells, the coimmunoprecipitation results revealed that TBK1 interacted with HSC70 and that Stub1 increased their interaction (Fig. 6A). Consistently, we also noted that endogenous HSC70 interacted with TBK1 during VSV and HSV-1 infection (Fig. 6B, C). BIFC results also confirmed the interaction between HSC70 and TBK1 (Fig. 6D). To determine whether the K27 chain of TBK1 is essential for HSC70 binding, we examined the interaction between TBK1-WT and TBK1-K344R with HSC70. We found that after the K27 ubiquitin-binding site of TBK1 was mutated, TBK1 no longer interacted with HSC70 (Fig. 6E). To further demonstrate that HSC70 specifically recognizes K27-linked polyubiquitination of TBK1, we performed in vitro binding assays. The results showed that HSC70 only bound to K27-ubiquitinated TBK1, but not to K48-ubiquitinated TBK1 (Fig. S10A). Subsequently, to investigate whether other heat shock proteins are involved, we also conducted in vitro binding experiments. The results indicated that only HSC70—and not HSP70 or HSP90—specifically bound to K27-ubiquitinated TBK1 (Fig. S10B), underscoring the specificity of HSC70 in recognizing TBK1 during chaperone-mediated autophagy (CMA). We noted that the overexpression of HSC70 increased Stub1-mediated TBK1 degradation. Additionally, VER-155008, an HSC70 inhibitor that inhibits CMA [46], was used to treat cells. We observed that Stub1-regulated TBK1 degradation was greatly reduced in VER-155008-treated cells (Fig. 6F). Notably, the overexpression of HA-HSC70 and Stub1 significantly improved the degradation of TBK1 in HEK293T-hsc70−/− cells (Fig. 6G), although the knockout of HSC70 did not affect the interaction between TBK1 and Stub1 (Fig. 6H). However, Stub1 knockout significantly attenuated the interaction between TBK1 and HSC70 (Fig. 6I). In addition, compared with that in HEK293T-hsc70+/+ cells, the expression level of TBK1 in the lysosomes of HEK293T-hsc70−/− cells was significantly lower after VSV infection (Fig. 6J). Furthermore, the confocal microscopy results also revealed that TBK1 does not localize to lysosomes following VSV infection in HEK293T-hsc70−/− cells (Fig. 6K). These results suggest that Stub1 may promote TBK1 degradation through CMA.
A HEK293T cells were transfected with Flag-TBK1, Myc-Stub1, and HA-HSC70 plasmids for 24 h, and then the lysates were immunoprecipitated with an anti-Flag antibody and analyzed by immunoblotting. B, C THP-1 cells were infected with VSV or HSV-1 for the indicated times and then subjected to immunoprecipitation with a TBK1 antibody before immunoblot analysis of endogenous TBK1 and HSC70 protein levels. D BIFC analysis of TBK1 and HSC70. HA-HSC70-VN, HSC70 fused with the N-terminus of Venus; Flag-TBK1-VC, TBK1 fused with the C-terminus of Venus. E Co-IP and immunoblot analysis of HEK293T cells transfected with Flag-TBK1 or Flag-TBK1-K344R and HA-HSC70. F HEK293T cells were transfected with a Flag-TBK1 plasmid with or without HA-HSC70 plasmids and then treated with VER-155,008 (5 μM, 10 μM, or 25 μM). TBK1 protein levels were analyzed by immunoblotting. G HEK293T-hsc70−/− cells were transfected with Flag-TBK1, Myc-Stub1, and HA-HSC70 plasmids for 24 h, after which the lysates were analyzed by immunoblotting. H HEK293T-hsc70+/+ and HEK293T-hsc70−/− cells were transfected with Flag-TBK1 and HA-Stub1 plasmids for 24 h, after which the lysates were immunoprecipitated with an anti-Flag antibody and analyzed by immunoblotting in the presence of CQ (50 μM) for 6 h. I HEK293T-stub1+/+ and HEK293T-stub1−/− cells were transfected with Flag-TBK1 and HA-HSC70 plasmids for 24 h, and then the lysates were immunoprecipitated with an anti-Flag antibody and analyzed by immunoblotting in the presence of CQ (50 μM) for 6 h. J HEK293T-hsc70+/+ and HEK293T-hsc70−/− cells were infected with VSV for 0, 4, 8, or 12 h in the presence of CQ (50 μM). Lysosomes were subsequently isolated and enriched before the proteins were analyzed by immunoblotting. K Fluorescence images showing the colocalization of TBK1 (green) with HSC70 (cyan) on lysosomes (red) in hsc70+/+ cells, whereas TBK1 (green) does not colocalize with lysosomes (red) in hsc70−/− cells; original magnification ×600. The data are representative of three independent experiments with three biological replicates (means ± SDs) or represent three independent experiments with similar results (A–K).
Stub1 promotes TBK1 localization in lysosomes
The substrate-HSC70 complex has been shown to interact with LAMP2A, which mediates the translocation of the substrate into the lysosome for degradation [47]. To assess the relationships among Stub1, HSC70, and LAMP2A, these three proteins were coexpressed in HEK293T cells. We found that Stub1 and HSC70 can synergistically promote TBK1 degradation (Fig. 7A). Co-IP results indicated that Stub1 interacted with TBK1, LAMP2A, and HSC70 (Fig. 7B). To further determine whether the interaction between Lamp2a and TBK1 occurs in the cytoplasm, lysosomes, or other vesicular compartments, we isolated lysosomes and performed co-IP using lysosomal extracts to detect the interaction between TBK1 and Lamp2a. The results indicate that TBK1 and Lamp2a interact within lysosomes and that Stub1 enhances this interaction (Fig. 7C). To definitively demonstrate the specific degradation of TBK1 by CMA, we established a LAMP2A knockout cell line via CRISPR-Cas9 technology. Cotransfection analysis of TBK1 and Stub1 in LAMP2A-deficient cells revealed that Stub1-mediated degradation of TBK1 was completely inhibited (Fig. 7D). This finding further corroborates the indispensable role of CMA in this specific process. Subsequent retransfection of LAMP2A restored TBK1 degradation, thereby reactivating Stub1’s ability to promote TBK1 protein degradation. This finding suggests that the lysosomal degradation specificity of TBK1 is achieved through the CMA pathway rather than through alternative mechanisms (Fig. 7D). Confocal microscopy results further revealed that TBK1 and LAMP2A exhibited distinct colocalization within lysosomes. In the absence of LAMP2A, there was a significant reduction in TBK1 colocalization with lysosomes (Fig. 7E). To investigate whether Stub1 directly promotes TBK1 degradation via the lysosomal pathway, we performed lysosomal isolation. Under basal conditions with Stub1 overexpression, we observed only a faint TBK1 signal during lysosomal isolation, which was consistent with rapid degradation upon entry. However, when cells were treated with chloroquine (CQ, 50–100 μM) to inhibit lysosomal hydrolases, a pronounced accumulation of TBK1 within lysosomes was evident. This CQ-induced accumulation was specific to the lysosomal pathway, as it was not recapitulated by the proteasome inhibitor MG-132. Furthermore, the lysosomal accumulation of TBK1, a well-established inducer of CMA, was markedly increased upon serum starvation. Critically, this accumulation was abrogated in LAMP2A knockout cells, in which TBK1 levels in lysosomes remained low even with CQ treatment. These data collectively demonstrate that TBK1 is targeted to the lysosome for degradation in a CMA-dependent manner (Fig. 7F). To further validate this finding, we performed a LAMP2A antibody blocking assay after lysosomal isolation. The results revealed that following LAMP2A antibody-mediated blockade of lysosomes, TBK1 failed to enter the lysosomal compartment (Fig. 7G). Interestingly, we also found that TBK1 can be transported to lysosomes only when Stub1 promotes the polyubiquitination of TBK1 (Fig. 7G). Overall, these findings prove that Stub1 facilitates the entry of TBK1 for degradation in lysosomes through CMA.
A HEK293T cells were transfected with Flag-TBK1, Myc-Stub1, GFP-LAMP2A, and HA-HSC70 plasmids before analysis by immunoblotting with the indicated antibodies. B HEK293T cells were transfected with Flag-TBK1, Myc-Stub1, GFP-LAMP2A, and HA-HSC70 plasmids for 24 h, and then the lysates were immunoprecipitated with an anti-Flag antibody and analyzed by immunoblotting in the presence of CQ (50 μM) for 6 h. C Flag-vector and Flag-stub1 were overexpressed in HEK293T cells. The cells were treated with CQ for 6 h prior to total protein extraction. Lysosomes were subsequently isolated for co-IP experiments to detect the interaction between TBK1 and Lamp2a. D Immunoblot analysis of TBK1 protein levels in Lamp2a+/+ and Lamp2a−/− cells cotransfected with plasmids encoding TBK1 and Stub1. Lamp2a−/− cells were subsequently reconstituted via transfection with a LAMP2A expression plasmid. E Fluorescence images showing the colocalization of TBK1 (green) with LAMP2A (cyan) on lysosomes (red) in Lamp2a+/+ cells, whereas TBK1 (green) does not colocalize with lysosomes (red) in Lamp2a−/− cells; original magnification ×600. F Stub1 was overexpressed in Lamp2a+/+ and Lamp2a−/− cells, the cells were treated with CQ or MG132 and starved, and then, the lysosomes were isolated and enriched before the proteins were analyzed by immunoblotting. G Isolated lysosomes were preincubated with a function-blocking anti-LAMP-2A antibody or control IgG. After being washed, they were incubated with purified TBK1 protein, Stub1 protein, and recombinant ubiquitin/E1/E2 proteins in CMA reaction buffer containing an ATP-regenerating system at 37 °C. Lysosomal protease inhibitors such as leupeptin were added. Lysosomes were reisolated via centrifugation, and TBK1 uptake was analyzed by immunoblotting. The data are representative of three independent experiments with three biological replicates (means ± SDs) or represent three independent experiments with similar results (A–G).
Stub1 deficiency augments host antiviral responses
Stub1fl/fl mice and Stub1fl/fl Lyz2-Cre mice were challenged with HSV-1 and VSV via intraperitoneal injection to further define the ability of Stub1 to inhibit type І IFN production and host antiviral responses in vivo. As shown in Fig. 8A, H, Stub1fl/fl Lyz2-Cre mice presented a high level of resistance to HSV-1 infection (Fig. 8A) in terms of overall survival compared with Stub1fl/fl mice and a significantly lower mortality rate caused by VSV infection in these mice (Fig. 8H). The ELISA results indicated that IFN-β and IL-6 production in the sera of Stub1fl/fl Lyz2-Cre mice was significantly greater than that in the sera of Stub1fl/fl mice (Fig. 8B, C, I, J). Consistently, we found that the mRNA levels of Ifnβ1 in the brains of Stub1fl/fl Lyz2-Cre mice were significantly greater than those in the brains of Stub1fl/fl mice after infection with HSV-1 for 48 or 72 h (Fig. 8D). Similarly, the mRNA levels of Ifnβ1 in the lungs of Stub1fl/fl Lyz2-Cre mice were significantly greater than those in the lungs of Stub fl/fl mice after infection with VSV for 48 and 72 h (Fig. 8K).
A Survival of Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice treated with HSV-1 (2 × 108 PFU/g) via intraperitoneal injection (n = 10 per group). ELISA analysis of IFN-β (B) and IL-6 (C) production in sera from Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice (n = 3 per group). D qPCR analysis of Ifnb1 mRNA expression in the brains of Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice after VSV administration (2 × 108 PFU/g) via intraperitoneal injection (n = 3 per group) for 48 and 72 h. E qPCR analysis of HSV-1 genomic DNA expression in the brains of Stub1 fl/fl and Stub1fl/fl Lyz2-Cre mice after HSV-1 administration (2 × 108 PFU/g) via intraperitoneal injection (n = 3 per group) for 48 and 72 h. F Determination of HSV-1 loads in the brain of Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice via the TCID50 assay. G Hematoxylin and eosin-stained images of brain sections from Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice infected with HSV-1 for 72 h. H Survival of Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice administered VSV (1 × 108 PFU/g) via intraperitoneal injection (n = 10 per group). ELISA analysis of IFN-β (I) and IL-6 (J) production in sera from Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice (n = 3 per group). K qPCR analysis of Ifnb1 mRNA expression in the brains of Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice after VSV administration (1 × 108 PFU/g) via intraperitoneal injection (n = 3 per group) for 48 and 72 h. L qPCR analysis of VSV genomic mRNA expression in lungs from Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice after VSV administration (1 × 108 PFU/g) via intraperitoneal injection (n = 3 per group) for 48 and 72 h. M Determination of VSV loads in the lungs of Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice via the TCID50 assay. N Hematoxylin and eosin-stained images of lung sections from Stub1 fl/fl and Stub1 fl/fl Lyz2-Cre mice infected with VSV for 72 h. O Scoring is based on the proportion of neutrophils among all cells: 0 points: Neutrophils absent; 1 point: Neutrophil count <5%; 2 points: Neutrophil count 5–50%; 3 points: Neutrophil count >50%. The data are presented as the means ± SDs of three independent experiments. Statistical significance was determined by two-tailed Student’s t test or one-way ANOVA followed by the Bonferroni post hoc correction, as appropriate. *P < 0.05, **P < 0.01, ***P < 0.001.
The HSV-1 DNA copy numbers and titers in the brains of Stub1fl/fl Lyz2-Cre mice were significantly lower than those in Stub1fl/fl mice (Fig. 8E, F), and the VSV copy numbers and titers in the lungs of Stub1fl/fl Lyz2-Cre mice were significantly lower than those in Stub1fl/fl mice after infection with VSV for 48 h and 72 h (Fig. 7L, M). We also observed less inflammatory cell infiltration and slight tissue injury in the brains of Stub1fl/fl Lyz2-Cre HSV-1 mice than in those of Stub1fl/fl mice after infection with HSV-1 (Fig. 8G). Fewer signs of severe inflammation and less pathological damage were observed in the lungs of Stub1fl/fl Lyz2-Cre mice than in those of Stub1fl/fl mice (Fig. 8N). Immune infiltration was scored on the basis of the proportion of neutrophils among total cells: 0 points: absence of neutrophils; 1 point: neutrophil count <5%; 2 points: neutrophil count 5%-50%; 3 points: neutrophil count >50%. The results are shown in Fig. 8O. Together, these results suggest that Stub1 suppresses IFN-I production and inhibits the antiviral innate response in the context of VSV and HSV-1 infections in vivo.
Discussion
TBK1 is an essential kinase in the host antiviral innate immune signaling pathway [48, 49], whose activity and stability are finely regulated by various posttranslational modifications, including ubiquitination [17, 29, 50, 51]. Ubiquitination can either activate TBK1 (e.g., K63‑linked chains) [52] or target it for degradation (e.g., K48‑linked chains) [15, 18, 23, 53]. In recent years, autophagy, particularly selective autophagy, has been shown to degrade multiple innate immune signaling molecules to maintain immune homeostasis [25, 32, 53, 54]. In this study, we revealed a novel negative regulatory mechanism for TBK1: the E3 ubiquitin ligase Stub1 catalyzes the K27‑linked polyubiquitination of TBK1, which subsequently directs TBK1 for degradation via chaperone‑mediated autophagy, thereby suppressing type I interferon production and promoting viral replication.
Although previous studies have shown that Stub1 inhibits antiviral immunity by promoting K48-linked ubiquitination and proteasomal degradation of RIG-I and MAVS [39, 40], its role in chaperone-mediated autophagy in relation to innate immunity remains poorly understood. Notably, while these works demonstrated Stub1-mediated MAVS degradation in BHK cells, we did not observe significant degradation of MAVS upon Stub1 overexpression in the human cell lines used in our study, such as HEK293T and HeLa cells. This observation suggests that the regulatory function of Stub1 may be highly context dependent, potentially varying with cell type, species, or specific cellular state. Our findings do not contradict these earlier reports but rather highlight the complexity of Stub1’s role in immune regulation. Importantly, in our experimental systems, we identified a distinct and potent mechanism by which Stub1 negatively regulates antiviral signaling by targeting TBK1, not MAVS, for chaperone-mediated autophagic degradation via K27-linked ubiquitination. We confirmed that the ectopic expression of Stub1 inhibited the activation of type I IFN signaling and enhanced viral infection, whereas the knockout of Stub1 expression in mice increased their resistance to VSV and HSV-1 infection. Therefore, our findings reveal a Stub1-TBK1 axis in the RLR and cGAS-STING pathways, highlighting the effects of CMA on host antiviral innate immune responses.
Autophagy has also been reported to play important roles in regulating type I IFN production and the innate immune response [54]. In the present study, Stub1 promoted TBK1 degradation. The E3 ubiquitin ligase Stub1 is a cytoplasmic protein with a highly conserved amino acid sequence across species. Previous studies have shown that Stub1 can degrade misfolded or chronologically expressed proteins via the ubiquitin-proteasome and autophagy-lysosome pathways [55]. Here, we found that only the lysosomal inhibitor CQ was able to repress Stub1-mediated TBK1 degradation, which is dependent on its E3 ligase activity. By comparing the amino acid sequences of those CMA motifs, we speculated that TBK1 may have three conserved CMA motifs (KFDKQ). These findings led us to hypothesize that Stub1 promotes TBK1 degradation via CMA. We subsequently confirmed that KFDKQ is the key motif for Stub1-mediated TBK1 degradation.
Critically, Stub1‑mediated TBK1 degradation is independent of canonical macroautophagy or the proteasome but occurs through CMA. A conserved KFERQ‑like motif (KFDKQ) within the TBK1 protein sequence is essential for its recognition by CMA. Mechanistically, Stub1‑catalyzed K27 ubiquitin chains enhance the recognition and binding of TBK1 by the chaperone HSC70, which in turn promotes the interaction of TBK1 with the lysosomal membrane protein LAMP2A, ultimately leading to its translocation into lysosomes for degradation. Our experiments confirmed that the knockout of HSC70 or LAMP2A or treatment with an HSC70 inhibitor effectively blocks Stub1‑mediated TBK1 degradation. This CMA‑dependent degradation pathway is activated during the later stages of viral infection, potentially serving as a feedback mechanism to prevent tissue damage caused by excessive immune responses.
We further validated the physiological relevance of the Stub1‑TBK1 axis in animal models. Mice with myeloid‑specific Stub1 knockout presented enhanced type I interferon responses, lower viral loads, milder histopathological damage, and higher survival rates upon VSV and HSV‑1 infection. These results clearly demonstrate that Stub1 acts as a “molecular brake” in antiviral immunity by targeting TBK1 for degradation in vivo.
In summary, our study elucidates for the first time the complete molecular mechanism by which Stub1 directs TBK1 to CMA for degradation via K27‑linked polyubiquitination (Fig. 9). This finding not only expands our understanding of the posttranslational regulatory network of TBK1 and reveals a new role for CMA in innate immune signal transduction but also provides a fresh perspective on the balance between viral infection and host immune homeostasis. Future investigations into the role of Stub1 in autoimmune diseases or chronic viral infections, as well as the therapeutic potential of targeting the Stub1-TBK1 interaction, represent promising directions for further research.
Stub1 binds to TBK1 and catalyzes its K27 polyubiquitination through its E3 ligase activity. Polyubiquitinated TBK1 is recognized by HSC70 through the KFDKQ motif of TBK1, ultimately leading to chaperone-mediated autophagic degradation.
Materials and methods
Cell lines
HeLa and HEK293T cells were purchased from the American Type Culture Collection (Manassas, VA) and cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS, 100 U/mL penicillin, and 100 mg/mL streptomycin at 37 °C with 5% CO2. The cell lines were routinely tested for mycoplasma contamination within the past year. Peritoneal macrophages and primary bone marrow-derived macrophages were isolated from mice 3 days after injection of thioglycolate (MERCK) and cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS), 100 U/mL penicillin, and 100 mg/mL streptomycin at 37 °C with 5% CO2.
Viruses
The VSV and VSV-GFP were kindly provided by Prof. Zhigao Bu (HVRI, China). HSV-1 was kindly provided by Prof. Hongbin Shu (Wuhan University, China), and GFP-HSV-1 was kindly provided by Prof. Diuqiu Liu (HVRI, China).
Mice
Stub1fl/+ mice were generated by Suzhou Cyagen Biosciences via CRISPR-Cas9-mediated gene editing. In summary, guide RNAs (gRNAs, 5′- CAAGCCCAGAGCTTGTAGAG-GGG-3′ and 5′- AAGGACAACAACCGGTAC AT-GGG-3′) were synthesized via in vitro transcription and subsequently purified. The gRNAs and Cas9 protein were mixed and injected into one-cell-stage fertilized eggs together with a targeting vector containing two LoxP sites flanking exons 2 and 3 of the Stub1 gene. The fertilized eggs that were injected were subsequently transplanted into pseudo-pregnant mice. The targeted genomes of the F0 mice were amplified via PCR and subsequently sequenced. The chimeras were crossed with wild-type C57BL/6 mice to obtain F1 Stub1fl/+ mice. Stub1fl/fl mice were crossed with Lyz2-Cre mice to generate Stub1fl/fl Lyz2-Cre mice. Age- and sex-matched littermates (8 weeks) were used throughout the study. The sex of the animals used is indicated in the respective figure legends. Murine genotypes were determined via PCR analysis of tail DNA via the following primers to amplify the wild-type (WT) allele (204 bp) and the floxed allele (259 bp): Stub1 forward: 5′- AGTGAGCTGCATTCATATCTCACC-3′, Stub1 reverse: 5′- ACCAGAAGC TAGTTGGTGATGG-3′. The mice were maintained and bred under specific pathogen-free (SPF) conditions. All the mice were generated and housed in specific pathogen-free (SPF) barrier facilities at the Harbin Veterinary Research Institute (HVRI) of the Chinese Academy of Agricultural Sciences (CAAS) (Harbin, China). All animal experiments were performed according to animal protocols approved by the Subcommittee on Research Animal Care at the HVRI, and all the mice used were less than 6 months old.
Generation of HEK293T-Stub1/HSC70/LAMP2A knockout cell lines
As previously described, CRISPR/Cas9 genomic editing for gene deletion was used [56]. HEK293T-stub1−/− cell lines were constructed via the CRISPR/Cas9 method. To create mammalian stub1−/− cells, one CRISPR guide RNA (sgRNA) sequence targeting the Stub1 locus in the genome was chosen on the basis of specificity scores (http://crispr.mit.edu/). The sgRNA sequence used was as follows: stub1−/− sgRNA, 5’- GGAGATGGAGAGCTATGATG -3’. The CRISPR/Cas9 system was subsequently introduced into HEK293T cells via the transfection reagent Lipofectamine 2000. Monoclonal cells were isolated via flow cytometry and validated through sequencing and Western blot analysis. Correctly knocked-out cells were amplified to establish stable knockout cell lines for subsequent experiments. The same method was used to construct the other two knockout cell lines.
Plasmids
HA-tagged full-length and deletion mutants of Stub1, HA-tagged TBK1 (HA-TBK1), deletion mutants of TBK1-encoding plasmids, GFP-tagged LAMP2A (pEGFP-N1-LAMP2A), and HA-tagged HSC70 (HA-HSC70) plasmids were constructed via standard molecular biology techniques. Plasmids expressing Flag-tagged RIG-I, MDA5, MAVS, TBK1, IKKε, and IRF3 have been previously described [51]. The IFN-β reporter and TK-Renilla reporter were obtained from Prof. Hong Tang. The primers used in this study are listed in Table 1.
Reagents and antibodies
Rabbit anti-Flag (F7425-2MG), mouse anti-Flag (F1804-1MG), rabbit anti-HA (SAB4300603), and mouse anti-HA (HS658-2ML) antibodies were purchased from Sigma-Aldrich (St. Louis, MO, USA). Poly (dA: dT) (P0883-10UN) and anti-Flag (M2) beads (M8823) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Mouse anti-GAPDH (60004-1-Ig), rabbit anti-TBK1 (28397-1-AP), recombinant human ubiquitin protein (Ag0260), recombinant human UBE1 protein (Ag8920), and recombinant human UBE2N protein (Ag0292) were purchased from Proteintech (Wuhan, China). Rabbit anti-phospho-TBK1 (5483), rabbit anti-IRF3 (11904), and rabbit anti-phospho-IRF3 (29047) antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA). Anti-LAMP2A antibody (AB125068), anti-LAMP1 antibody (ab208943), and anti-ubiquitin (linkage-specific K27) antibody (ab181537) were purchased from Abcam (Shanghai, China). IRDye 800CW goat anti-rabbit IgG (H + L) (925-32211) and IRDye 800CW goat anti-mouse IgG (H + L) (925-32210) were purchased from LI-COR (Lincoln, NE, USA). Alexa Fluor 488-conjugated goat anti-rabbit IgG (H + L) (A11008), Alexa Fluor 594-conjugated goat anti-mouse IgG (H + L) (A11032), Alexa Fluor 633-conjugated goat anti-rabbit IgG (H + L) (A21070), and a lysosome enrichment kit (89839) were purchased from Thermo Fisher Scientific (Waltham, MA, USA). Lyso-Tracker Red (C1046) was purchased from Beyotime.
Stimulants and virus infection
Stimulants were used at the following concentrations: poly (dA: dT), 10 μg/mL; poly(I:C), 10 μg/mL; or poly(I:C), 1 μg/mL or poly (dA: dT), 1 μg/mL. The cells were infected with VSV (multiplicity of infection [MOI] = 1), GFP-labeled VSV (MOI = 1), or HSV (MOI = 5) for the indicated times. The cell lysates were analyzed via qPCR or immunoblotting; the supernatants were analyzed via ELISA or the TCID50 assay. For in vivo studies, age- and sex-matched littermates were intraperitoneally injected with VSV [1 × 108 plaque-forming units per gram (PFU/g)]. The sera from the mice were collected for ELISA detection 12 h after VSV infection. The mice were euthanized 12 h after viral infection to obtain liver, lung, and spleen tissues for qPCR analysis. Lungs from control or virus-infected mice were dissected, fixed, and stained with hematoxylin-eosin via standard procedures and examined via light microscopy for histological changes. For survival experiments, Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice were intraperitoneally injected with VSV and then monitored for survival after viral infection.
Luciferase reporter assay
The luciferase activities were measured with a Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s instructions. The data were normalized for transfection efficiency by dividing firefly luciferase activity by Renilla luciferase activity.
RNA extraction and qPCR
Total RNA was extracted via TRIzol reagent (Invitrogen, California, America), and reverse transcription was accomplished via the PrimeScript™ RT Reagent Kit (Takara, Tokyo, Japan). Real-time PCR was conducted via TB Green™ Premix Ex Taq™ II (TaKaRa, Tokyo, Japan) in a typical 20 μL PCR mixture that included 10 μL of TB Green™ Premix Ex Taq™ II, 1–5 μL of template cDNA, and 0.4 mM of each PCR primer. The cycling conditions were 95 °C for 2 min, followed by 40 cycles at 95 °C for 5 s and 60 °C for 30 s, and the samples were run on a Stratagene Mx3000P Real-Time PCR System (Stratagene, America). The data were normalized according to the level of β-actin expression in each sample. All experiments were performed at least in triplicate. The qPCR primers used are listed in Table 2.
ELISA
The concentrations of IFN-β (PBL Interferon Source) in the cell culture supernatants and sera were measured via ELISA kits according to the manufacturer’s instructions.
Lysosome isolation experiment
Lysosomes were isolated via the Lysosome Enrichment Kit for Tissues and Cultured Cells (Cat. No. 89839) (Thermo Scientific) according to the manufacturer’s instructions.
Bimolecular fluorescence complementation assays
The C-terminus or N-terminus of yellow fluorescent protein (VENUS) was inserted into the eukaryotic expression plasmids of Stub1, TBK1, and HSC70. The relevant constructs were transfected into HEK293T cells, which were subsequently incubated for approximately 24–48 h. The fluorescence signals from the HEK293T cells were imaged via a Zeiss LSM-880 laser scanning fluorescence microscope (Carl Zeiss AG, Oberkochen, Germany) with a 63× oil objective.
IFN sensitivity test
HEK293T cells were transfected with increasing amounts of a plasmid expressing Stub1 and then infected with SeV for 12 h. The cell supernatants were collected and inactivated with ultraviolet (UV) light. MDBK cells were incubated with UV-inactivated cell supernatants for 24 h and then infected with VSV-GFP (MOI = 0.1) or HSV-1-GFP (MOI = 1) for 12 h. VSV-GFP and HSV-1-GFP replication was analyzed under a fluorescence microscope.
Coimmunoprecipitation and Western blot analysis
Coimmunoprecipitation and Western blot analysis were performed as previously described [57]. In brief, HEK293T cells transfected with the indicated plasmids for 24 h were lysed in lysis buffer (50 mM Tris-HCl, pH 7.4; 150 mM NaCl; 5 mM MgCl2; 1 mM EDTA; 1% Triton X-100; and 10% glycerol) containing 1 mM PMSF and 1 × protease inhibitor cocktail (Roche). Then, the cell lysates were incubated with anti-Flag (M2) beads at 4 °C overnight on a roller. The precipitated beads were washed five times with cell lysis buffer. For Western blot analysis, equal amounts of cell lysates and immunoprecipitates were resolved via 10–12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to a polyvinylidene difluoride membrane (Millipore, Stafford, VA, USA). After incubation with primary and secondary antibodies, the membranes were visualized with an Odyssey two-color infrared fluorescence imaging system (LI-COR, USA).
GST pull-down
For GST pull-down, Stub1 was cloned and inserted into pGEX-6p-1 with a C-terminal GST tag, whereas TBK1 was cloned and inserted into pET28a with a C-terminal His-tag. Both Stub1 and TBK1 were expressed in E. coli BL21, which was induced with 0.2 mM isopropyl-β-D-thiogalactopyranoside (IPTG). The bacterial lysate containing GST-Stub1 was incubated with 10 μL of glutathione Sepharose 4B beads (Cat. No. 45-000-139; GE Healthcare) for 3 h and purified by washing with 1 × PBS five times. Then, the beads were incubated with bacterial lysate containing His-TBK1 for another 3 h and purified by washing with PBS five times. The proteins eluted from the beads were determined by Western blotting with anti-GST and anti-His antibodies.
In vitro ubiquitination assay
Recombinant ubiquitin protein (Ag0260) and recombinant E1 (Ag8920) and E2 (Ag0292) proteins were purchased from Wuhan Proteintech Group. GST-Stub1 and His-TBK1 were expressed in E. coli BL21. Each reaction with a 30 µL final volume contained 50 ng of E1 protein, 50 ng of E2 protein, and 5 μg of ubiquitin together with reaction buffer (50 mM Tris-HCl (pH 7.4), 10 mM MgCl2, 5 mM dithiothreitol, 5 mM ATP, and 10% glycerol) in tubes containing beads that bind with His-TBK1 and GST-Stub1 proteins. The reactions were incubated in a thermomixer for 3 h at 30 °C with shaking, stopped by the addition of 1× SDS-PAGE loading buffer, and incubated for another 5 min at 100 °C. Five microliters of each reaction mixture were analyzed via electrophoresis on 12% SDS-PAGE gels. The incubated mixtures were detected by Western blotting using anti-ubiquitin (Abcam), anti-GST (Abmart), and anti-Flag (Abmart) antibodies.
Confocal microscopy and colocalization analysis
The cells were transfected with the indicated plasmids and then fixed for 10 min in 4% paraformaldehyde in 1 × phosphate-buffered saline (PBS) pH 7.4. The fixed cells were permeabilized for 15 min with 0.3% Triton X-100 in 1 × PBS and then blocked in 1 × PBS with 10% bovine serum albumin for 30 min. The cells were incubated with the appropriate primary antibodies and then stained with Alexa Fluor 594-labeled goat anti-rabbit immunoglobulin G and Alexa Fluor 488-labeled goat anti-mouse IgG. The subcellular colocalization was visualized via a Zeiss LSM-880 laser scanning fluorescence microscope (Carl Zeiss AG, Oberkochen, Germany) with a 63× oil objective. Zeiss processing system software was used to determine the degree of colocalization. Ch3-T1 denotes the 633 nm channel, Ch2 GaAsP-T2 denotes the 488 nm channel, and Ch1-T3 denotes the 405 nm channel (DAPI).
Histopathology analysis
To assess histological changes in the brain and lungs, Stub1fl/fl and Stub1fl/fl Lyz2-Cre mice were infected with VSV or HSV-1, and the brain and lungs were fixed in 10% formalin neutral buffer solution overnight. The tissues were embedded in paraffin blocks and then sectioned at a thickness of 4 μM for staining with hematoxylin and eosin following standard procedures. The results were analyzed via light microscopy.
Quantification and statistical analysis
Sample sizes were chosen on the basis of standard methods and previous experience in the laboratory to ensure adequate power to detect biologically relevant effects. The data are presented as the means ± standard deviations (SDs). The normality of the data distribution was assumed on the basis of the nature of the assays and prior experience with similar datasets. Variance similarity between compared groups was assessed visually from the data distribution plots. Statistical tests were chosen accordingly. Statistical analysis was conducted via unpaired Student’s t test for comparisons between two groups or one-way or two-way analysis of variance (ANOVA), followed by the Bonferroni post hoc correction for multiple comparisons. P values less than 0.05 were considered statistically significant. No formal randomization or blinding was applied to animal group allocation or outcome assessment, as all comparisons were made between genetically defined littermate groups under controlled experimental conditions.
Compliance with ethics requirements
All the animals were acclimated under standard laboratory conditions (ventilated room, 25 ± 1 °C, 60 ± 5% humidity, 12 h light/dark cycle) and had free access to standard water and food (SYXK-2013-100). All procedures were conducted in accordance with the “Guiding Principles in the Care and Use of Animals” (China). All animal experiments have been approved by the Science and Technology Ethics Committee of Harbin Veterinary Research Institute (IAVUV review approval number: 240813-02-GR). Qualification Certificate Number for Animal Experimenters: 20210880.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Aderem A, Ulevitch RJ. Toll-like receptors in the induction of the innate immune response. Nature. 2000;406:782–7.
Kawai T, Akira S. TLR signaling. Semin Immunol. 2007;19:24–32.
Loo YM, Gale M Jr. Immune signaling by RIG-I-like receptors. Immunity. 2011;34:680–92.
Su C, Tang YD, Zheng C. DExD/H-box helicases: multifunctional regulators in antiviral innate immunity. Cell Mol Life Sci. 2021;79:2.
Carneiro LA, Magalhaes JG, Tattoli I, Philpott DJ, Travassos LH. Nod-like proteins in inflammation and disease. J Pathol. 2008;214:136–48.
Zheng C. The emerging roles of NOD-like receptors in antiviral innate immune signaling pathways. Int J Biol Macromol. 2021;169:407–13.
Reis ESC, Yamasaki S, Brown GD. Myeloid C-type lectin receptors in innate immune recognition. Immunity. 2024;57:700–17.
Chen Q, Sun L, Chen ZJ. Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat Immunol. 2016;17:1142–9.
Gray EE, Winship D, Snyder JM, Child SJ, Geballe AP, Stetson DB. The AIM2-like receptors are dispensable for the interferon response to intracellular DNA. Immunity. 2016;45:255–66.
Yi L, Song J, Zhang Z, Li L, Wu Y, Xue M, et al. Palmitoyl-transferase 3 promotes mitochondrial antiviral signaling protein degradation by modulating its ubiquitination. Int J Biol Macromol. 2025;310:143609.
Zheng C. The emerging roles of the MARCH ligases in antiviral innate immunity. Int J Biol Macromol. 2021;171:423–7.
Takeuchi O, Akira S. Pattern recognition receptors and inflammation. Cell. 2010;140:805–20.
Tanaka Y, Chen ZJ. STING specifies IRF3 phosphorylation by TBK1 in the cytosolic DNA signaling pathway. Sci Signal. 2012;5:ra20.
Zhang C, Shang G, Gui X, Zhang X, Bai XC, Chen ZJ. Structural basis of STING binding with and phosphorylation by TBK1. Nature. 2019;567:394–8.
Huang J, Chen Z, Ye Y, Shao Y, Zhu P, Li X, et al. DTX3L enhances type I interferon antiviral response by promoting the ubiquitination and phosphorylation of TBK1. J Virol. 2023;97:e0068723.
You H, Zheng S, Huang Z, Lin Y, Shen Q, Zheng C. Herpes simplex virus 1 tegument protein UL46 inhibits TANK-binding kinase 1-mediated signaling. mBio. 2019;10:e00919-19.
Li X, Zhang Q, Ding Y, Liu Y, Zhao D, Zhao K, et al. Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat Immunol. 2016;17:806–15.
Cui J, Li Y, Zhu L, Liu D, Songyang Z, Wang HY, et al. NLRP4 negatively regulates type I interferon signaling by targeting the kinase TBK1 for degradation via the ubiquitin ligase DTX4. Nat Immunol. 2012;13:387–95.
Zhang M, Wang L, Zhao X, Zhao K, Meng H, Zhao W, et al. TRAF-interacting protein (TRIP) negatively regulates IFN-beta production and antiviral response by promoting proteasomal degradation of TANK-binding kinase 1. J Exp Med. 2012;209:1703–11.
Zheng Q, Hou J, Zhou Y, Yang Y, Xie B, Cao X. Siglec1 suppresses antiviral innate immune response by inducing TBK1 degradation via the ubiquitin ligase TRIM27. Cell Res. 2015;25:1121–36.
Lin M, Zhao Z, Yang Z, Meng Q, Tan P, Xie W, et al. USP38 inhibits type I interferon signaling by editing TBK1 ubiquitination through NLRP4 signalosome. Mol Cell. 2016;64:267–81.
Abe T, Lee A, Sitharam R, Kesner J, Rabadan R, Shapira SD. Germ-cell-specific inflammasome component NLRP14 negatively regulates cytosolic nucleic acid sensing to promote fertilization. Immunity. 2017;46:621–34.
Deng M, Tam JW, Wang L, Liang K, Li S, Zhang L, et al. TRAF3IP3 negatively regulates cytosolic RNA induced anti-viral signaling by promoting TBK1 K48 ubiquitination. Nat Commun. 2020;11:2193.
Ma Y, Galluzzi L, Zitvogel L, Kroemer G. Autophagy and cellular immune responses. Immunity. 2013;39:211–27.
Clarke AJ, Simon AK. Autophagy in the renewal, differentiation and homeostasis of immune cells. Nat Rev Immunol. 2019;19:170–83.
Zhang H, Baehrecke EH. Eaten alive: novel insights into autophagy from multicellular model systems. Trends Cell Biol. 2015;25:376–87.
Li WW, Li J, Bao JK. Microautophagy: lesser-known self-eating. Cell Mol Life Sci. 2012;69:1125–36.
Cuervo AM, Wong E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 2014;24:92–104.
Kumar S, Gu Y, Abudu YP, Bruun JA, Jain A, Farzam F, et al. Phosphorylation of syntaxin 17 by TBK1 controls autophagy initiation. Dev Cell. 2019;49:130–44.e6.
Richter B, Sliter DA, Herhaus L, Stolz A, Wang C, Beli P, et al. Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria. Proc Natl Acad Sci USA. 2016;113:4039–44.
Du Y, Duan T, Feng Y, Liu Q, Lin M, Cui J, et al. LRRC25 inhibits type I IFN signaling by targeting ISG15-associated RIG-I for autophagic degradation. EMBO J. 2018;37:351–66.
Chen M, Meng Q, Qin Y, Liang P, Tan P, He L, et al. TRIM14 inhibits cGAS degradation mediated by selective autophagy receptor p62 to promote innate immune responses. Mol Cell. 2016;64:105–19.
Yu T, Yang X, Fu Q, Liang J, Wu X, Sheng J, et al. TRIM11 attenuates Treg cell differentiation by p62-selective autophagic degradation of AIM2. Cell Rep. 2023;42:113231.
Jin S, Tian S, Luo M, Xie W, Liu T, Duan T, et al. Tetherin suppresses type I interferon signaling by targeting MAVS for NDP52-mediated selective autophagic degradation in human cells. Mol Cell. 2017;68:308–22.e4.
Pan M, Yin Y, Hu T, Wang X, Jia T, Sun J, et al. UXT attenuates the CGAS-STING1 signaling by targeting STING1 for autophagic degradation. Autophagy. 2023;19:440–56.
Yang Q, Liu TT, Lin H, Zhang M, Wei J, Luo WW, et al. TRIM32-TAX1BP1-dependent selective autophagic degradation of TRIF negatively regulates TLR3/4-mediated innate immune responses. PLoS Pathog. 2017;13:e1006600.
Xie W, Tian S, Yang J, Cai S, Jin S, Zhou T, et al. OTUD7B deubiquitinates SQSTM1/p62 and promotes IRF3 degradation to regulate antiviral immunity. Autophagy. 2022;18:2288–302.
Xie W, Jin S, Zhang C, Yang S, Wu Y, Zhao Y, et al. Selective autophagy controls the stability of TBK1 via NEDD4 to balance host defense. Cell Death Differ. 2022;29:40–53.
Zhou P, Ding X, Wan X, Liu L, Yuan X, Zhang W, et al. MLL5 suppresses antiviral innate immune response by facilitating STUB1-mediated RIG-I degradation. Nat Commun. 2018;9:1243.
Zhang Y, Hou P, He DC, Wang H, He H. RACK1 degrades MAVS to promote bovine ephemeral fever virus replication via upregulating E3 ubiquitin ligase STUB1. Vet Microbiol. 2021;257:109096.
Zhang M, Windheim M, Roe SM, Peggie M, Cohen P, Prodromou C, et al. Chaperoned ubiquitylation-crystal structures of the CHIP U box E3 ubiquitin ligase and a CHIP-Ubc13-Uev1a complex. Mol Cell. 2005;20:525–38.
Xu Z, Kohli E, Devlin KI, Bold M, Nix JC, Misra S. Interactions between the quality control ubiquitin ligase CHIP and ubiquitin conjugating enzymes. BMC Struct Biol. 2008;8:26.
Xu W, Marcu M, Yuan X, Mimnaugh E, Patterson C, Neckers L. Chaperone-dependent E3 ubiquitin ligase CHIP mediates a degradative pathway for c-ErbB2/Neu. Proc Natl Acad Sci USA. 2002;99:12847–52.
Rusilowicz-Jones EV, Urbe S, Clague MJ. Protein degradation on the global scale. Mol Cell. 2022;82:1414–23.
Yang Q, Wang R, Zhu L. Chaperone-mediated autophagy. Adv Exp Med Biol. 2019;1206:435–52.
Moradi-Marjaneh R, Paseban M, Moradi Marjaneh M. Hsp70 inhibitors: Implications for the treatment of colorectal cancer. IUBMB Life. 2019;71:1834–45.
Kaushik S, Cuervo AM. Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol. 2012;22:407–17.
Fitzgerald KA, McWhirter SM, Faia KL, Rowe DC, Latz E, Golenbock DT, et al. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat Immunol. 2003;4:491–6.
Liu S, Cai X, Wu J, Cong Q, Chen X, Li T, et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science. 2015;347:aaa2630.
Perry AK, Chow EK, Goodnough JB, Yeh WC, Cheng G. Differential requirement for TANK-binding kinase-1 in type I interferon responses to toll-like receptor activation and viral infection. J Exp Med. 2004;199:1651–8.
Liu H, Ye G, Liu X, Xue M, Zhou Q, Zhang L, et al. Vimentin inhibits type I interferon production by disrupting the TBK1-IKKepsilon-IRF3 axis. Cell Rep. 2022;41:111469.
Zhang R, Yang W, Zhu H, Zhai J, Xue M, Zheng C. NLRC4 promotes the cGAS-STING signaling pathway by facilitating CBL-mediated K63-linked polyubiquitination of TBK1. J Med Virol. 2023;95:e29013.
Zhu H, Zhang R, Yi L, Tang YD, Zheng C. UNC93B1 attenuates the cGAS-STING signaling pathway by targeting STING for autophagy-lysosome degradation. J Med Virol. 2022;94:4490–501.
Choi Y, Bowman JW, Jung JU. Autophagy during viral infection—a double-edged sword. Nat Rev Microbiol. 2018;16:341–54.
Chakraborty A, Edkins AL. CHIP: a co-chaperone for degradation by the proteasome and lysosome. Subcell Biochem. 2023;101:351–87.
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 2013;154:1380–9.
Li J, Hu L, Liu Y, Huang L, Mu Y, Cai X, et al. DDX19A senses viral RNA and mediates NLRP3-dependent inflammasome activation. J Immunol. 2015;195:5732–49.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grant no. 32400721).
Author information
Authors and Affiliations
Contributions
Changjiang Weng and Hongyang Liu conceived and coordinated the study; Hongyang Liu and Changjiang Weng wrote the paper; Hongyang Liu, Jimin Yu, Guangqiang Ye, Jiaxiu Gao, Li Huang, and Zhaoxia Zhang performed and analyzed the experiments. Changjiang Weng and Hongyang Liu designed, modified, and corrected the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, H., Yu, J., Ye, G. et al. E3 ubiquitin ligase Stub1 enhances viral replication by promoting TBK1 degradation through molecular chaperone-mediated autophagy. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01716-7
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01716-7