Abstract
BRAFV600-mutant melanoma relies on hyperactivation of the MAPK/ERK pathway for tumorigenesis, with BRAF/MEK inhibitors (BRAFi/MEKi) improving patient outcomes. However, therapeutic resistance frequently emerges, and male patients show poorer responses and outcomes, partially linked to androgen receptor (AR) overexpression. Here, we uncover a mechanistic link between AR signaling and autophagic resistance in BRAFV600-mutant melanoma. We show that BRAFi treatment upregulates AR expression, which induces cytoprotective autophagy through transcriptional activation of DRAM1, a key autophagy-related gene. Functional studies reveal that AR-driven autophagy confers resistance to BRAFi by enhancing cellular survival under therapeutic stress. Our findings establish AR-regulated autophagy as a critical resistance mechanism and provide preclinical evidence for combining AR-targeting PROTAC degrader ARV110 with autophagy inhibitors to overcome BRAFi resistance.
Similar content being viewed by others
Introduction
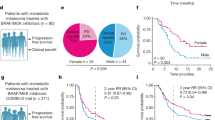
Melanoma is a cutaneous malignancy with somatic BRAFV600 mutations occurring in approximately 50% of cases. These mutations trigger constitutive activation of the MAPK/ERK signaling pathway, driving aggressive tumor development [1]. The advent of BRAF/MEK-targeted therapies (BRAFi/MEKi) has significantly improved clinical outcomes for patients. However, therapeutic resistance commonly emerges, leading to disease relapse within 7–24 months [2]. Notably, male patients exhibit notably higher resistance to BRAFi/MEKi and worse prognoses compared to female counterparts [3, 4]. Intriguingly, while melanoma is not traditionally classified as a sex hormone-responsive malignancy, it shows significant sex-based differences in its biology, prognosis and drug resistance [3, 5, 6]. A potential explanation for this gender-related discrepancy lies in divergent sex hormone signaling pathways between males and females [7]. Indeed, the androgen receptor (AR), which displays heterogeneous expression across melanoma tissues, has been linked to enhanced melanoma cell proliferation, metastatic capacity, and tumorigenesis [8, 9]. Clinical cohort analyses further reveal that significantly higher AR expression in male patients than in females, a finding associated with impaired response to BRAFi/MEKi targeted therapy [10]. Despite these observations, the underlying mechanisms through which AR drives resistance to BRAF/MEK-targeted agents remain largely uncharacterized.
Autophagy maintains cellular homeostasis by degrading and recycling cytotoxic protein aggregates and dysfunctional organelles via the lysosome, a conserved quality-control mechanism critical for stress adaptation [11]. Substantial studies have shown that autophagy contributes to the resistance against BRAFi/MEKi therapy in BRAF-mutant melanoma patients through several pathways, and was associated with reduced partial response rates and shorter progression-free survival [12, 13]. Notably, emerging evidence also highlights sex-dependent variations in autophagic activity within cancer [14, 15], with sex steroid hormones and their receptors mediating these disparities. For example, ERα can regulate the transcription of Bcl-2 and ULK1, two autophagy-related genes in breast cancer [16, 17]. Bioinformatics analyses have identified 12 autophagy genes transcriptionally controlled by ERα and 19 genes by ERβ [18], underscoring the pivotal roles of ERα/β in modulating autophagy initiation and tumor progression across malignancies like colon and lung cancer [19,20,21,22]. However, in striking contrast to estrogenic pathways in these tumors, other sex hormones, especially androgen-dependent autophagy, remain largely uninvestigated in melanoma.
In this study, we demonstrate how AR-regulated autophagy mechanistically contributes to BRAFi resistance in BRAFV600 mutant melanoma patients. We found that BRAF inhibitors upregulate AR expression in BRAFV600mut melanoma cells, a process that confers resistance to these inhibitors by inducing cytoprotective autophagy. Through functional screening, we identified DRAM1 as a key transcriptional target of AR mediating this autophagic response. These findings reveal the molecular basis of clinical resistance in BRAFV600 mutant melanoma and provide strong preclinical evidence for combining AR antagonists with autophagy inhibitors as a therapeutic strategy in this patient population.
Methods
Cell lines and cultures
Melanoma cell lines SH-4 (CRL-7724), SK-MEL-2 (HTB-68), A375(CRL-1619), WM-266-4(CRL-1676) and SK-MEL-28 (HTB-72) were purchased from ATCC. WM793 (WM793-01-0001) was sourced from Rockland Immunochemicals (Limerick, PA). A375, SH-4 were cultured in high-glucose DMEM (Gibco); SK-MEL-28 and SK-MEL-2 were maintained in EMEM (Gibco). WM793, WM1366 and IPC298 were cultured in RPMI-1640 (Gibco). All basal media were supplemented with 10% FBS (Gibco #10099-141) and 1% penicillin/streptomycin (Gibco, #15140122). All cell lines were authenticated by STR profiling and were routinely verified negative for Mycoplasma.
Cell manipulations and treatments
AR overexpression (Addgene # 213819) and TF-LC3 (EGFP-RFP-LC3, Addgene #22418) lentiviral vectors were packaged in HEK293T cells using third-generation plasmids (psPAX2/pMD2.G; Addgene #12260, #12259) via Lipofectamine 3000 (Invitrogen)-mediated transfection [23]. Melanoma cells were selected by hygromycin (300 μg/ml; Invitrogen) or puromycin (2 μg/ml; Invitrogen) for 6 days after 48 h post-transduction.
si-DRAM1: cells at 50% confluency in 6-well plates were transfected with siRNA using Lipofectamine RNAiMAX (Invitrogen, #13778150) in Opti-MEM (Gibco, #31985070). Media were replaced after 6 h, and knockdown efficiency was assessed 48 h post-transfection. The siRNA sequences for DRAM1 were: 5′- AUAAAUCCAAAAAUACCACUC-3′, and the siRNA sequences for the negative control were brought from Santa Cruz (sc-37007).
BRAF inhibitor (BRAFi)-resistant cell line (BR) establishment: Parental (P) cells were exposed to Dabrafenib (Selleck) in a stepwise dose-escalation protocol: initial concentration at 0.5 nM, doubled weekly for 4 weeks to reach 1 µM. Resistant (BR) cells were maintained in 1 µM Dabrafenib for ≥4 weeks prior to experiments.
Cell proliferation assays: cells (1 × 10³/well) were treated with Dabrafenib (0.1-1000 nM) for 120 h. Cell viability was evaluated using the ATP-dependent luminescence assay with CellTiter-Glo® 2.0 reagent (Promega), and measurements were performed on an EnVision microplate reader (PerkinElmer). Dose-response curves and IC50 values were generated via four-parameter logistic regression (GraphPad Prism v9.0).
Clonogenicity: cells (1000/well, 6-well plates) were treated with indicated compounds, fixed with methanol after 14 days, stained with 1% crystal violet (Sigma), and analyzed using ImageJ v1.53.
Western blot
Cancer cells or tumor samples were homogenized and lysed in RIPA buffer (Merk, #20-188) supplemented with protease inhibitors (Selleck) and PMSF. To separate the cytoplasmic and nuclear components, NE-PER™ Nuclear and Cytoplasmic Extraction Reagents (Invitrogen) were employed. After the extraction, the protein concentration was determined using the Bradford assay (Invitrogen). Subsequently, an equal amount of proteins (20 µg) was separated by 4–12% SDS-PAGE (Invitrogen) followed by immunoblotting analysis. Antibodies were used as follows: AR (#5153), β-actin (#4970), Lamin B (#17416), LC3 (#12741) and p62 (#23214) were brought from Cell Signaling Technology. DRAM1 (ab64739) was obtained from abcam.
RT-qPCR
Total RNA was isolated from cancer cells or tumor samples using TRIzol™ reagent (Invitrogen), treated with DNase and quantified by using a NanoDrop spectrophotometer. For the synthesis of first-strand cDNA, SuperScript™ IV reverse transcriptase (Invitrogen) was applied. The q-PCR reactions, with a total volume of 20 μl were carried out using PowerUp™ SYBR Green Master Mix (Applied Biosystems) on a QuantStudio™ 6 Flex system. Relative quantification was performed through the ΔΔCt method with gene expression normalized to GAPDH. Strict quality control measures were implemented throughout the entire procedure. Primers used in the study are provided as follows: AR F: -GACGACCAGATGGCTGTCATT-; R: -GACGACCAGATGGCTGTCATT-; DRAM1 F: -TCAAATATCACCATTGATTTCTGT-; R: -GCCACATACGGATGGTCATCTCTG-; GAPDH F: -GAACATCATCCCTGCCTCTACT-; R: -CCTGCTTCACCACCTTCTTG-;
Transmission electron microscopy (TEM)
TEM assay was performed as described previously [23]. Cellular specimens were immobilized in 2.5% glutaraldehyde/0.1 M sodium cacodylate, with secondary fixation using 1% OsO4/1.5% K4[Fe(CN)6] (EMS) for 3 h. Sequential ethanol dehydration (50-100%) preceded embedding in EMbed 812 resin (EMS) with thermal polymerization. Ultrathin sections (60 nm; Leica UC7) underwent dual-contrast staining: 2% uranyl acetate (5 min) followed by Reynolds’ lead citrate (1 min). Final imaging was acquired on a Philips CM-12 TEM (FEI) operating at 80 kV, strictly following standard EM processing protocols.
Evaluation of fluorescent TF-LC3 puncta
TF-LC3, TF-LC3/AR-OE stable cell lines or TF-LC3/AR-OE stable cell lines transfected with si-DRAM1 were used to measure the TF-LC3 puncta. Images acquisition by using a Zeiss LSM710 confocal microscope with environmental control (37 °C, 5% CO₂).
Dual luciferase reporter gene assay
The DRAM1 promoter region (−2000 bp) and its mutant (core AR-binding site disrupted) were cloned into pGL3-Basic (Promega). Melanoma cells (5 × 10⁵/well) in 6-well plates received co-transfection of pGL3-DRAM1 (WT or mutant) with pRL-TK normalization vector (Promega) via Lipo3000 (Invitrogen). At 48 h post-transfection, dual-luciferase activities were assessed using Promega’s Dual-Luciferase® system (Promega) on a GloMax® Navigator. Relative luciferase activity (RLA) was calculated as Firefly/Renilla signal ratio and compared between AR-modulated (overexpression or knockdown) and control cells to assess AR-dependent transcriptional regulation.
Chromatin immunoprecipitation (ChIP) assay
Melanoma cells underwent formaldehyde crosslinking (1%, Sigma #F8775; 10 min) followed by glycine quenching (125 mM). Nuclear lysis in ChIP buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 0.5% NP-40). Chromatin was fragmented to ~500 bp using a Covaris S220 sonicator (20% duty cycle, 200 cycles/burst, 15 cycles). Lysates were immunoprecipitated overnight at 4 °C with 5 μg anti-AR (CST, #5153) or IgG control (Proteintech, #30000-0-AP), followed by 3 h incubation with Protein G Magnetic Beads (CST, #9006). After crosslink reversal and DNA purification (Tiangen, #DP214), AR occupancy at the DRAM1 promoter was quantified via qPCR (SYBR Green) with primers targeting predicted response elements (ΔCt normalized to Input DNA).
Tissue microarrays (TMAs) and IHC staining
Human metastatic melanoma (pretreatment/posttreatment) tissue microarray (ZL-MEL963) was sourced from Shanghai Ouwei Biotech (Ouwei, Shanghai) with full patient consent. This study was approved by the Institutional Review Board at Huashan Hospital, Fudan University in Shanghai, China. The patient cohort included paired pre- and post-treatment patient samples from 30 patients with resectable stage III melanoma. All patients received neoadjuvant dabrafenib (150 mg BID) and trametinib (2 mg QD) for 8 weeks before surgical resection, followed by adjuvant therapy with the same agents for up to 12 months (Table 1).
The commercial tissue microarray provider supplies both frozen tissue samples and their corresponding FFPE sections for this patient cohort. For the extraction of RNA and protein (used in subsequent RT-qPCR and Western blot analyses of patient-derived samples), we exclusively used the frozen tissue samples, as they better preserve the integrity of nucleic acids and proteins compared to FFPE sections.
For IHC staining, FFPE sections (4 μm) were deparaffinized and pressurized antigen retrieval (citrate buffer, pH 6.0, 125 °C/3 min), and peroxidase blockade (3% H₂O₂). After incubation with anti-DRAM1 antibody (PA5-20336, Invitrogen) or IgG control overnight. HRP-conjugated secondary antibodies were incubated for another 1 h. Chromogenic development employed DAB substrate with hematoxylin counterstaining, followed by acid alcohol differentiation and neutral gum mounting. DRAM1 expression quantification utilized histoscore algorithms integrating staining intensity and positive cell percentage, following CAP-accredited pathology protocols.
Synergistic combination studies
Synergistic drug combinations were identified through a high-throughput matrix screening approach. Cells were seeded in 384-well plates, followed by compound addition in a matrixed format with serial dilutions as indicated. After 5 days, cell viability was assessed using CellTiter-Glo assays. Dose-response curves were fitted using a 4-parameter logistic model, and synergistic interactions were analyzed with the Combenefit™ software based on the Loewe additivity model. Statistical significance was determined by comparing heterologous combination synergy scores against their respective self-crosses using a two-sample t test, a score above 10 is considered significant (p < 0.05).
Animal xenograft evaluation
All animal procedures were conducted in accordance with the protocols approved by the Institutional Animal Care and Use Committee at Fudan University. A375R xenograft models were generated using established protocols [23] with modifications: Male BALB/c nude mice (6-week-old, Shanghai Laboratory Animal Center) received subcutaneous flank inoculation of 5 × 10⁶ A375R cells in Matrigel/RPMI 1640 (1:1 v/v). When tumors reached a size of ~80 mm3, mice were randomly assigned into four groups (n = 4/group): Oral vehicle (PBS), ARV110 (10 mg/kg, oral gavage), 3-MA (25 mg/kg, i.p.) and combination therapy once daily. Terminal endpoints at day 21 post-implantation involved CO₂ euthanasia followed by tumor harvest for weighing and immunoblotting.
Statistical analyses
All the experiments were performed at least three times independently. Student’s t test was used to determine the statistical difference between the two groups. For comparisons among more than two groups, a one-way analysis of variance (ANOVA) followed by Tukey’s post-hoc test was performed unless otherwise mentioned. Detailed information on the statistical methods applied for each experiment is in the corresponding figure legends. Statistical analysis was performed using Graphpad Prism software. p < 0.05 was considered significant.
No statistical methods were employed to predetermine sample size, and no animals or samples were excluded from the study. The sample size was determined based on established precedents in prior literature to ensure sufficient statistical power. Randomization was performed to allocate all samples/cells/mice into the control and treatment groups for every experiment. The experimental design did not incorporate blinding during data collection and analysis.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations, including the Declaration of Helsinki for human research and the ARRIVE guidelines for animal experimentation. Commercial human melanoma tissue microarrays (ZL-MEL963) were obtained from Shanghai Ouwei Biotech. The company confirmed that all tissue specimens were collected in accordance with the highest ethical standards and with full informed consent from donors. We adhere to standard clinical care protocols and enforce strict donor privacy protection. All human tissue collection was conducted under HIPAA-approved procedures. All animal experimental procedures were approved by the Institutional Animal Care and Use Committee at Fudan University, Shanghai, China (Approval No.: 2025-HSYY-134). Informed consent for the collection, analysis and research use of tissue samples and associated clinical data was obtained from all human tissue donors, as verified by the commercial tissue microarray provider.
Results
AR upregulation promotes BRAFi resistance in melanoma
Adaptive resistance to BRAF inhibitors (BRAFi) can arise either as an acute response or as a consequence of chronic BRAFi exposure. To explore the role of AR in BRAFi resistance, we first analyzed the acute response by treating six patient-derived melanoma cell lines with dabrafenib (DAB) for 48 h. Although only three cell lines (A375, WM793 and SK-MEL-28) exhibited baseline AR protein expression, DAB treatment induced AR upregulation across all cell lines, independent of their mutational background or sex of origin (Fig. 1A). Based on the AR expression, we selected A375 (female-derived) and WM793 (male-derived) for subsequent studies. We treated A375 and WM793 with multiple BRAF and MEK inhibitors (BRAFi/MEKi) and found the significant upregulation of AR mRNA expression (Fig. 1B). To confirm AR expression following chronic BRAF inhibitor (BRAFi) treatment, we established BRAFi-resistant A375 (A375R) and WM793 (WM793R) melanoma cell lines. We found that AR expression was upregulated in these resistant cells (Fig. 1C and supplementary Fig. 1A, B). To further evaluate the clinical relevance of AR expression and BRAFi resistance, we compared the AR expression levels in matched lesions excised from the same patient before (B) and after (A) BRAFi/MEKi therapy. Consistently, AR expression was markedly upregulated after BRAFi/MEKi therapy in both male and female patients, indicating its potential role as a compensatory resistance mechanism independent of sex (Fig. 1D). In addition, activated AR (nuclear/cytoplasmic AR) were increased upon 48 h, 72 h and 7 days DAB treatment in A375 and WM793 melanoma cell lines (Fig. 1E, F and supplementary Fig. 1C, D). To systematically interrogate the functional networks underlying these findings, we generate A375 and WM793 melanoma cell lines with stably overexpressed AR (AR-OE) and treated with DAB for 120 h. We found that cell viability was drastically improved in A375 AR-OE and WM793 AR-OE cells (Fig. 1G, H). Consistent with these findings, clonogenic assays demonstrated that AR overexpression increases melanoma cell colony formation, and DAB treatment partially reverses this effect (Fig. 1I, J). Collectively, these results indicate that the development of BRAF inhibitor (BRAFi) resistance in melanoma cell lines is closely linked to elevated AR expression levels.
AR expression in (A) 6 melanoma cell lines treated with/without DAB (0.5 μM) for 48 h, C BRAFi resistance (R) or parental (P) A375 or WM793 cells, D matched lesions before (B) and after A BRAFi/MEKi therapy, patients are randomly selected from a total of 30. B AR mRNA expression in A375 and WM793 cells treated with various BRAF and MEK inhibitors (Vemurafenib: 0.5 μM, DAB: 0.5 μM, Encorafenib: 5 nM, Trametinib: 5 nM and Cobimetinib 5 nM) for 48 h. E AR subcellular distribution in A375 treated with DAB (0.5 μM) for indicated time points. LaminB1 and GAPDH are nuclear and cytoplasmic markers, respectively. F quantification of activated AR (nuclear/cytoplasmic, N/C AR) in (E), at each time point (48, 72 h, and 7 d), the N/C ratio of DMSO was normalized to 1 and data for DAB-treated cells are shown as fold-change relative to their respective DMSO control. The “DMSO Control” in column combines DMSO from all time points (set to 1). G, H Cell proliferation of A375 and WM793 cells treated with DAB at the indicated concentrations for 120 h. I, J Clonogenicity assays of cells from (G, H) treated with 0.5 μM DAB or DMSO. Mean ± SEM, n = 3, two-tailed t-test. *p < 0.05, **p < 0.01. For (G, H), EV vs AR-OE, **p = 0.0013 (A375), **p = 0.0027 (WM793) respectively, (EV Empty vector).
AR induces autophagy-associated BRAFi resistance in melanoma
To thoroughly explore the underlying mechanism of AR in BRAFi-resistance, we utilized A375 AR-OE and WM793 AR-OE melanoma cell lines. We observed that overexpression of AR led to a marked increase in the accumulation of LC3II and a degradation of SQSTM1/p62 (Fig. 2A). We further confirmed the regulatory function of AR in autophagy by transmission electron microscopy (TEM). The results showed that the AR overexpression leads to a substantial elevation of double membrane or single membrane structures (Fig. 2B). The observed elevation in LC3II levels might result from either enhanced autophagic flux or impaired degradation of LC3II. To distinguish between these possibilities, we utilized the TF-LC3 reporter system, which enables discrimination between autophagosomes (yellow puncta, exhibiting both EGFP and RFP signals) and autolysosomes (red puncta, retaining RFP fluorescence due to EGFP quenching in acidic lysosomal environments). Our analysis revealed a marked increase in both red (autolysosomes) and yellow (autophagosomes) puncta in AR-OE A375 cells (Fig. 2C). Moreover, treatment with the lysosome inhibitor chloroquine (CQ) revealed that AR overexpression enhanced CQ-induced LC3II accumulation. Conversely, it reduced the accumulation of p62, indicating that AR activation drives complete autophagic flux in melanoma cells (Supplementary Fig. 2A). To investigate the potential role of AR-mediated autophagy in BRAFi resistance, we treated AR-OE and AR-EV cells with DAB and/or the autophagy inhibitor 3-methyladenine (3-MA). 3-MA treatment substantially attenuated LC3II accumulation without altering AR expression levels (Fig. 2D). AR-OE cells exhibited a significantly higher IC50 to DAB, however, this resistance was dramatically reversed by 3-MA co-treatment (Fig. 2E). Clonogenic assays further confirmed these findings, showing significant reduction in colony formation when combining 3-MA with DAB compared to monotherapy (Fig. 2F). Moreover, in the AR-silenced WM379R cells, treatment with the autophagy agonist rapamycin (Rapa) significantly enhanced LC3II levels without affecting AR expression. In addition, Rapa effectively reversed the suppression of cellular proliferation and colony-forming capacity induced by AR-knockdown (Fig. 2G–I and supplementary Fig. 2B–G). Altogether, these results indicate that autophagy activation was involved in AR-induced BRAFi resistance.
A AR, LC3II and p62 protein level in A375 and WM793 cells with or without AR overexpression. B TEM analysis of autophagosomes and autolysosomes in A375 EV and A375 AR-OE cells (scale bars: 0.5 μm). A autophagosomes, AL autolysosomes, M: mitochondria and N nucleus. Manually counted. Mean ± SEM, n = 30, two-tailed Student’s t test, **p < 0.01. C A375 EV and A375 AR-OE cells stably expressing TF-LC3 were imaged using fluorescence microscopy (Scale bars: 10 μm). White boxed regions in the panels are enlarged. Representative images of autolysosomes (red dots) and autophagosomes (yellow dots) are shown. Manually counted. n = 30, two tailed Student’s t test, **p < 0.01, ***p < 0.001. Following treatment with DAB (0.5 μM) and/or 3-MA (2.5 mM), A375 EV and A375 AR OE cells were analyzed for D protein expression, E cell proliferation and F colony formation assays as indicated. WM793R cells transfected with si-AR or si-scramble were treated with DAB (0.5 μM) and/or Rapa (5 nM), G protein expression, H cell proliferation and I colony formation assays were measured as indicated. Mean ± SEM, n = 3, one-way ANOVA and Tukey’s post-hoc test. **p < 0.01. For E, H EV(DAB) vs AR OE(DAB) **p = 0.0018, AR OE(DAB) vs AR OE(DAB + 3MA) **p = 0.0052, si-scramble (DAB) vs si-AR(DAB) **p = 0.0037, si-AR(DAB) vs si-AR(DAB+Rapa) **p = 0.0014.
AR upregulates DRAM1 expression in melanoma
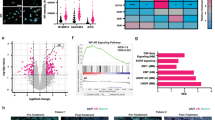
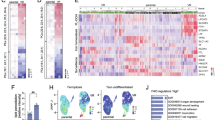
To systematically investigate how AR regulates autophagy in BRAFV600-mutant melanoma, we analyzed differentially expressed genes (DEGs) by using RNA-seq data from the Gene Expression Omnibus (GEO) dataset GSE232697. This dataset provides global transcriptomic profiles of three paired BRAFV600-mutant melanoma cell lines comparing AR-overexpressing (AR OE) and LacZ-expressing controls (CTRL), enabling identification of potential AR target genes. Consistently, genes associated with EGFR and TGF-β signaling pathways were significantly induced in AR-overexpressing cells as compared with controls. Among autophagy-related genes, DRAM1 (DNA damage-regulated autophagy modulator 1) emerged as one of the most prominently induced targets in this context (Fig. 3A). Next, we performed Venn diagram analysis using two additional GEO datasets, GSE255541 and GSE232693, to identify DEGs associated with adaptive resistance to BRAF inhibitors (BRAFi). These datasets comprise BRAFV600-mutant melanoma cell lines subjected to short-term dabrafenib treatment (acute BRAFi exposure) and BRAFi-resistant cell lines established through chronic BRAFi adaptation, with each group compared against their respective parental cell line controls. This cross-comparison revealed 111 overlapping DEGs shared between acutely treated and chronically adapted BRAFi-resistant cells. Strikingly, multiple autophagy-related genes were enriched in this overlapped gene set, with DRAM1 ranking among the top five most highly upregulated genes (Fig. 3B). Previous studies have shown that DRAM1 participates in autophagic regulation across multiple cancer types [24, 25]. Integrating the GEO data with evidence suggesting that AR and DRAM1 both contribute to autophagy promotion, we postulated that AR mediates DRAM1-dependent signaling to facilitate autophagy in melanoma. To corroborate this finding, matched tissue lesions excised from the same patients before and after BRAFi/MEKi therapy were prospectively collected. Quantitative analysis demonstrated significant upregulation of DRAM1 mRNA and protein expression following BRAFi treatment (Fig. 3C–E). In addition, DRAM1 protein and mRNA levels were upregulated in A375 AR-OE and WM793 AR-OE cells, while downregulated in AR-knockdown A375R and WM793R cells (Fig. 3F, G). Collectively, these results provide support for the hypothesis that AR drives DRAM1 expression in BRAF V600-mutated melanoma cells.
A Volcano plot depicting DEGs from the GSE232697 dataset, highlighting genes with significant changes in expression. B Venn diagram of the target genes from GSE255541 and GSE232693, and highlighting differentially expressed autophagy-related genes. C–E melanoma lesions from matched lesions before (B) and after (A) BRAFi/MEKi therapy C IHC staining of DRAM1 in three representative melanoma lesions. Scale bar, 50 μm. D, E DRAM1 protein and mRNA expression, Mean ± SEM, n = 30, two-tailed Student’s t test, **p < 0.0001. F DRAM1 expression in melanoma cells with AR overexpression. G DRAM1 expression in DAB-resistant melanoma cells with AR knockdown. Mean ± SEM, n = 3, two-tailed Student’s t test, **p < 0.01, ***p < 0.001.
AR regulates DRAM1 to promote both autophagy and BRAFi-resistance in melanoma
We transfected A375 and WM793 melanoma cells with Flag-tagged DRAM1, and found LC3 upregulation and p62 downregulation in DRAM1 overexpressed cells (Fig. 4A). In contrast, knockdown of DRAM1 in A375R and WM793R cell resulted in significant attenuation of LC3II accumulation and p62 degradation (Fig. 4B). Notably, DRAM1 knockdown markedly suppressed AR-induced LC3II upregulation and p62 downregulation (Fig. 4C). In addition, TF-LC3 fluorescence showed that increased red and yellow puncta in A375 AR OE cells were significantly down-regulated after DRAM1 knockdown (Fig. 4D). We further investigated the role of AR-DRAM1 axis in promoting BRAFi resistance in melanoma. We found the AR-OE induced drug resistance was remarkably reversed upon DRAM1 knockdown (Fig. 4E). In addition, clonogenic assays showed that DAB treatment in DRAM1 knockdown group led to significantly decrease in colony formation ability as compared to control group (Fig. 4F). Notably, DRAM1 knockdown in WM379R melanoma cells substantially attenuated DAB resistance, evidenced by a dramatic reduction in cell viability following DAB exposure (Fig. 4G). DRAM1 knockdown also profoundly impaired clonogenic survival, as demonstrated by significantly reduced colony-forming capacity in DRAM1-deficient WM379R cells compared to controls (Fig. 4H).
DRAM1, LC3 and p62 protein expression were measured in A A375 and WM793 cells transfected with Flag-DRAM1, B A375R and WM793R cells transfected with si-DRAM1 or si-scramble C A375 and WM793 AR-OE cells transfected with si-DRAM1. D A375 cells expressing TF-LC3, AR-OE-TF-LC3 and AR-OE-TF-LC3 cells transfected with si-DRAM1 were imaged by fluorescence microscopy (Scale bars: 10 μm). White boxed regions in the panels are enlarged. Representative images of autolysosomes (red dots) and autophagosomes (yellow dots) are shown. Manually counted. Mean ± SEM, n = 30, two tailed Student’s t test, **p < 0.01, ***p < 0.001. E, F cell proliferation and colony formation assays were measured in cells from (D) treated with DAB as indicated. Mean ± SEM, n = 3, one-way ANOVA and Tukey’s post-hoc test. **p < 0.01. G, H WM793R cells transfected with/without si-DRAM1 were incubated with DAB (0.5 μM), and cell proliferation and colony formation assays were measured. Mean ± SEM, n = 3, two-tailed Student’s t test, **p < 0.01. E EV vs AR-OE **p = 0.0017, AR-OE + si-scramble vs AR-OE+si-DRAM1 **p = 0.0032, G DAB + si-scramble vs DAB + si-DRAM1 **p = 0.0047.
AR induces DRAM1 transcription via the promoter androgen response element (ARE) binding
To analyze the AR-mediated direct transcriptional regulation of DRAM1, we first identified potential androgen response elements (AREs) within the DRAM1 5’-promoter region using the JASPAR database [26]. Computational analysis revealed two high-confidence AREs with robust predicted binding affinities (Fig. 5A). To validate functional AR binding, dual luciferase reporter assays were performed, demonstrating that AR directly interacts with the DRAM1 promoter and significantly enhances its transcriptional activation (Fig. 5B–E). Chromatin immunoprecipitation (ChIP) assays further corroborated these findings, showing pronounced enrichment of chromatin fragments harboring the identified AREs via anti-AR antibody pulldown compared to normal IgG controls. Notably, the enrichment positively correlated with AR overexpression and diminished upon AR knockdown (Fig. 5F–I). To establish the functional necessity of these AREs, we introduced point mutations (C/G→T and A/T→C substitutions) into the two core binding sites to disrupt AR-DNA interactions (Fig. 5J). Luciferase assays revealed that ARE mutations markedly attenuated AR-driven transcriptional activation (Fig. 5K, L). Consistent with this, AR knockdown substantially reduced promoter activity, while ARE mutations abolished this regulatory effect (Fig. 5M, N). Together, these data strongly suggest that AR upregulates DRAM1 transcription through direct binding to specific AREs in its 5’-promoter regions.
A Bioinformatics prediction of putative AR binding sites (androgen response elements, AREs) in the 5’-promoter region of DRAM1 using the JASPAR database. Dual luciferase assays assessing DRAM1 promoter activity in (B, C) A375R and WM793R cells transfected with si-AR or D, E A375 and WM793 cells transfected with AR overexpression. ChIP assays using anti-AR antibody in (F, G) AR OE A375 or WM793 cells or H, I AR knockdown A375R and WM793R cells. J Schematic of mutated ARE sequences in the DRAM1 5’-promoter region. Dual luciferase assays in (K, L) AR OE A375 and WM793 cells or M, N AR knockdown A375R and WM793R cells transfected with wildtype (WT) or mutant (MT) DRAM1 promoter constructs. Mean ± SEM, n = 3, two-tailed Student’s t test. *p < 0.05; **p < 0.01; ***p < 0.001; ns no significance.
Targeting the AR and autophagy overcomes BRAFi resistance
Our findings establish the AR-DRAM1 axis as a pivotal driver of BRAF inhibitor (BRAFi) resistance in melanoma. To functionally characterize this pathway, BRAFi-resistant melanoma cell lines (A375R and WM793R) were treated with the AR-targeting PROTAC degrader ARV110. ARV110 treatment caused marked depletion of AR protein, accompanied by a great reduction in DRAM1 expression, attenuation of LC3II accumulation and p62 degradation (Fig. 6A, B). ARV110 monotherapy significantly suppressed the proliferation of resistant cells, with synergistic growth inhibition observed upon combination with autophagy inhibitor 3-MA (Fig. 6C, D).
A375R and WM793R cells were treated with/without ARV110 (10 nM) and/or 3-MA (2.5 mM), A mRNA, B protein expression and C cell proliferation were measured as indicated. n = 3, two-tailed student’s t-test, ARV110 vs ARV110 + 3-MA in A375R **p = 0.0028, in WM793R **p = 0.0019. D Loewe additivity excess heatmap for A375R cells treated with the combination of AV110 and 3-MA. Darker coloration signifies enhanced cell killing and stronger synergistic interaction. A score >10 is considered significant. E Tumor-bearing nude mice were treated with PBS, ARV110 (10 mg/kg, qd), 3-MA (25 mg/kg, qd) or ARV110 (10 mg/kg) + 3-MA (25 mg/kg) as indicated, and representative images of dissected tumors at the end of the treatment period. F–H Quantitative analysis of Tumor weight, tumor volume and body weight in mice. I H&E staining of major organs from all groups of mice. Scale bar, 1 mm. J Protein expression in xenograft tumors as indicated. For F, G, Mean ± SEM; n = 4 mice. one-way ANOVA and Tukey’s post-hoc test, ns no significance, *p < 0.05, **p < 0.01, ***p < 0.001.
In an orthotopic A375R melanoma model, immunodeficient mice received 21-day combination therapy with oral ARV110 (10 mg/kg, qd) and/or intraperitoneal injections of the autophagy inhibitor 3-MA (25 mg/kg, qd). Combination therapy exhibited substantially greater suppression of tumor growth when compared with monotherapies (Fig. 6D–F). Moreover, no significant body weight fluctuations or target organ toxicity (via H&E staining) were observed in mice receiving combination therapy versus controls, indicating favorable tolerability (Fig. 6G, H). WB analysis revealed that ARV110 potently downregulated AR levels, which in turn suppressed DRAM1 expression, dampened AR-dependent autophagic activity and cellular proliferation. The p62 degradation and LC3-II accumulation were further decreased in combination-treated tumors, consistent with dual AR/autophagy pathway blockade (Fig. 6I). These data highlight co-targeting AR and autophagy as a tolerable resistance-overcoming strategy in BRAFi-resistant melanoma.
Discussion
Therapeutic resistance persists as a major obstacle in oncology, even with advanced targeted therapies. Therapeutic resistance stems from two distinct mechanisms: (i) pre-existing intrinsic adaptations via expansion of resistant clones, and (ii) acquired resistance through therapy-induced genetic/epigenetic rewiring. Adaptive mechanisms mediated by signaling networks, which function as rapid-response pathways to maintain cellular homeostasis, represent promising targets for adjuvant therapeutic strategies. Our study in BRAF-mutant melanoma patients demonstrates that BRAFi activates AR, which in turn upregulates DRAM1 transcription. Activation of the AR-DRAM1 axis induces cytoprotective autophagy, thereby contributing to the adaptive resistance against BRAFi through this stress-responsive circuitry.
Sex disparities in cancer epidemiology persist across multiple non-reproductive malignancies (e.g., bladder, colorectal cancers, and glioblastoma), with males exhibiting higher incidence and mortality than females. In melanoma, clinical evidence also identifies sex as an independent prognostic factor, though its mechanistic basis remains poorly characterized [27]. These disparities arise from the interplay of three sex-linked biological factors: divergent immune surveillance dynamics, sex hormone signaling pathways (particularly estrogen-androgen axis variations), and X/Y chromosome-mediated transcriptional regulation [28, 29]. Recent studies have identified AR-mediated pathways as critical drivers of melanoma progression and therapeutic resistance. For instance, in male patients, AR signaling promotes metastasis through FUT4-dependent L1CAM fucosylation and the miRNA-539-3p/USP13/MITF-AXL regulatory axis [8, 30]. Our study reveals that BRAF inhibitors induce AR upregulation in both sexes of BRAFV600-mutant melanoma patients, driving resistance to BRAF/MEK inhibition. Importantly, acquired resistance to BRAF/MEKi is driven by therapy-induced AR upregulation independent of tumor-intrinsic baseline AR levels. Therefore, future studies should delineate functional crosstalk between intrinsic AR signaling and therapy-triggered AR dynamics in resistance mechanisms.
Sex differences in autophagy regulation also contribute to disease disparities in cancer, cardiovascular disorders, and neurodegeneration [14]. While estrogen receptors (ERα/β) are well-characterized autophagy modulators in malignancies, AR-mediated autophagy remains poorly explored beyond prostate cancer. In prostate cancer, AR exerts context-dependent effects on autophagy: it can suppress autophagy through the induction of Grp78/BiP [31, 32], or alternatively promote autophagy by upregulating TFEB [33]. Our research is the first to demonstrate that AR is upregulated and drives autophagy activation in BRAF inhibitor-resistant melanoma, thereby defining a novel mechanism of sex-dependent therapeutic resistance.
DRAM1, a p53 target gene, is an evolutionarily conserved lysosomal transmembrane protein found in lysosomes, peroxisomes, autophagosomes, and the endoplasmic reticulum [34, 35], with established roles in autophagy and apoptosis. Mechanistically, it promotes autophagic flux by facilitating autophagosome biogenesis and lysosome fusion [25], while regulating apoptosis through BAX stabilization-inhibiting its degradation and recruiting it to lysosomes to activate caspase-9/-3 signaling [24]. Intriguingly, its role in cancer is starkly context-dependent: acting as a tumor-suppressor via epigenetic silencing in squamous cell carcinomas and non-small cell lung cancer, whereas exerting oncogenic functions to drive initiation, progression, and metastatic dissemination in glioblastoma, hepatoblastoma, and lung adenocarcinoma [36,37,38]. Our study demonstrates that AR-mediated transcriptional activation of DRAM1 drives autophagic flux, providing a mechanistic basis for BRAFi resistance in melanoma. Through functional validation, we identified two specific AREs within the DRAM1 promoter as critical AR-binding sites, with mutagenesis of these motifs abolishing AR-dependent transcriptional activation. This AR/DRAM1 axis establishes a previously unrecognized autophagy-dependent resistance pathway, expanding the functional repertoire of AR in melanoma progression beyond canonical hormone-driven signaling and underscoring its therapeutic relevance in overcoming BRAFi resistance.
We recognized several critical gaps requiring further investigation. First, the mechanistic interplay between AR signaling and autophagic flux remains incompletely characterized, which could inform the development of precision therapeutic strategies to selectively modulate these pathways. Such advances may mitigate the systemic adverse effects frequently associated with conventional androgen-deprivation therapies and broad-spectrum autophagy inhibitors. Second, there remains an urgent need to elucidate how sex-specific biological variables-including hormonal axes, immune dimorphism, and sex chromosome biology-influence therapeutic outcomes, as these factors may critically shape treatment responses.
In summary, our findings demonstrate that BRAF/MEK-targeted therapy activates the AR-DRAM1 axis to drive therapeutic resistance in melanoma. Mechanistically, BRAFV600E inhibition induces AR overexpression and autophagic flux, making tumors more selectively sensitive to combined AR/autophagy blockade compared to normal tissues, a strategy that enhances BRAFi/MEKi efficacy with manageable toxicity profiles [30, 39,40,41]. This mechanistic insight supports ongoing clinical trials investigating AR inhibition alongside chemotherapy, targeted therapy, or immunotherapy in patients with advanced cancers (NCT04926181, NCT01974765, NCT03207529, NCT02684227, NCT02312557), including a phase I/II trial (NCT02257424) testing BRAFi/MEKi plus autophagy inhibitor HCQ (autophagy inhibitor) in BRAF-mutant melanoma. Our preclinical data suggested the clinical rationale for simultaneously targeting AR and autophagy pathways as a promising strategy to overcome BRAF inhibitor resistance in melanoma therapy.
Data availability
All data presented in this study were available from the corresponding author upon reasonable request.
References
Ahmed B, Qadir MI, Ghafoor S. Malignant melanoma: skin cancer-diagnosis, prevention, and treatment. Crit Rev Eukaryot Gene Expr. 2020;30:291–7.
Kakadia S, Yarlagadda N, Awad R, Kundranda M, Niu J, Naraev B, et al. Mechanisms of resistance to BRAF and MEK inhibitors and clinical update of US Food and Drug Administration-approved targeted therapy in advanced melanoma. Onco Targets Ther. 2018;11:7095–107.
Nosrati A, Wei ML. Sex disparities in melanoma outcomes: the role of biology. Arch Biochem Biophys. 2014;563:42–50.
Chhabra Y, Fane ME, Pramod S, Hüser L, Zabransky DJ, Wang V, et al. Sex-dependent effects in the aged melanoma tumor microenvironment influence invasion and resistance to targeted therapy. Cell. 2024;187:6016–34.e25.
Gamba CS, Clarke CA, Keegan TH, Tao L, Swetter SM. Melanoma survival disadvantage in young, non-Hispanic white males compared with females. JAMA Dermatol. 2013;149:912–20.
Clocchiatti A, Cora E, Zhang Y, Dotto GP. Sexual dimorphism in cancer. Nat Rev Cancer. 2016;16:330–9.
Rampen FH, Mulder JH. Malignant melanoma: an androgen-dependent tumour? Lancet. 1980;1:562–4.
Wang Y, Ou Z, Sun Y, Yeh S, Wang X, Long J, et al. Androgen receptor promotes melanoma metastasis via altering the miRNA-539-3p/USP13/MITF/AXL signals. Oncogene. 2017;36:1644–54.
Ma M, Ghosh S, Tavernari D, Katarkar A, Clocchiatti A, Mazzeo L. et al. Sustained androgen receptor signaling is a determinant of melanoma cell growth potential and tumorigenesis. J Exp Med. 2021;18:e20201137
Vellano CP, White MG, Andrews MC, Chelvanambi M, Witt RG, Daniele JR, et al. Androgen receptor blockade promotes response to BRAF/MEK-targeted therapy. Nature. 2022;606:797–803.
Miller DR, Thorburn A. Autophagy and organelle homeostasis in cancer. Dev Cell. 2021;56:906–18.
Fratta E, Giurato G, Guerrieri R, Colizzi F, Dal Col J, Weisz A, et al. Autophagy in BRAF-mutant cutaneous melanoma: recent advances and therapeutic perspective. Cell Death Discov. 2023;9:202.
Ma XH, Piao SF, Dey S, McAfee Q, Karakousis G, Villanueva J, et al. Targeting ER stress-induced autophagy overcomes BRAF inhibitor resistance in melanoma. J Clin Investig. 2014;124:1406–17.
Lista P, Straface E, Brunelleschi S, Franconi F, Malorni W. On the role of autophagy in human diseases: a gender perspective. J Cell Mol Med. 2011;15:1443–57.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402:672–6.
Dong L, Wang W, Wang F, Stoner M, Reed JC, Harigai M, et al. Mechanisms of transcriptional activation of bcl-2 gene expression by 17beta-estradiol in breast cancer cells. J Biol Chem. 1999;274:32099–107.
Hua S, Kallen CB, Dhar R, Baquero MT, Mason CE, Russell BA, et al. Genomic analysis of estrogen cascade reveals histone variant H2A.Z associated with breast cancer progression. Mol Syst Biol. 2008;4:188.
Türei D, Földvári-Nagy L, Fazekas D, Módos D, Kubisch J, Kadlecsik T, et al. Autophagy regulatory network—a systems-level bioinformatics resource for studying the mechanism and regulation of autophagy. Autophagy. 2015;11:155–65.
Wei Y, Huang C, Wu H, Huang J. Estrogen receptor beta (ERβ) mediated-cyclinD1 degradation via autophagy plays an anti-proliferation role in colon cells. Int J Biol Sci. 2019;15:942–52.
Marzagalli M, Casati L, Moretti RM, Montagnani Marelli M, Limonta P. Estrogen receptor β agonists differentially affect the growth of human melanoma cell lines. PLoS ONE. 2015;10:e0134396.
Chatterjee SJ, Pandey S. Chemo-resistant melanoma sensitized by tamoxifen to low dose curcumin treatment through induction of apoptosis and autophagy. Cancer Biol Ther. 2011;11:216–28.
Yakimchuk K, Jondal M, Okret S. Estrogen receptor α and β in the normal immune system and in lymphoid malignancies. Mol Cell Endocrinol. 2013;375:121–9.
Bao Y, Ding Z, Zhao P, Li J, Chen P, Zheng J, et al. Autophagy inhibition potentiates the anti-EMT effects of alteronol through TGF-β/Smad3 signaling in melanoma cells. Cell Death Dis. 2020;11:223.
Guan JJ, Zhang XD, Sun W, Qi L, Wu JC, Qin ZH. DRAM1 regulates apoptosis through increasing protein levels and lysosomal localization of BAX. Cell Death Dis. 2015;6:e1624.
Geng J, Zhang R, Yuan X, Xu H, Zhu Z, Wang X, et al. DRAM1 plays a tumor suppressor role in NSCLC cells by promoting lysosomal degradation of EGFR. Cell Death Dis. 2020;11:768.
Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2024;52:D174–d82.
Joosse A, Collette S, Suciu S, Nijsten T, Patel PM, Keilholz U, et al. Sex is an independent prognostic indicator for survival and relapse/progression-free survival in metastasized stage III to IV melanoma: a pooled analysis of five European Organisation for Research and Treatment of Cancer randomized controlled trials. J Clin Oncol. 2013;31:2337–46.
Dong M, Cioffi G, Wang J, Waite KA, Ostrom QT, Kruchko C, et al. Sex differences in cancer incidence and survival: a pan-cancer analysis. Cancer Epidemiol Biomark Prev. 2020;29:1389–97.
Haupt S, Caramia F, Klein SL, Rubin JB, Haupt Y. Sex disparities matter in cancer development and therapy. Nat Rev Cancer. 2021;21:393–407.
Liu Q, Adhikari E, Lester DK, Fang B, Johnson JO, Tian Y, et al. Androgen drives melanoma invasiveness and metastatic spread by inducing tumorigenic fucosylation. Nat Commun. 2024;15:1148.
Jiang Q, Yeh S, Wang X, Xu D, Zhang Q, Wen X, et al. Targeting the androgen receptor leads to suppression of prostate cancer via induction of autophagy. J Urol. 2012;188:1361–8.
Bennett HL, Fleming JT, O’Prey J, Ryan KM, Leung HY. Androgens modulate autophagy and cell death via regulation of the endoplasmic reticulum chaperone glucose-regulated protein 78/BiP in prostate cancer cells. Cell Death Dis. 2010;1:e72.
Blessing AM, Rajapakshe K, Reddy Bollu L, Shi Y, White MA, Pham AH, et al. Transcriptional regulation of core autophagy and lysosomal genes by the androgen receptor promotes prostate cancer progression. Autophagy. 2017;13:506–21.
Crighton D, Wilkinson S, O’Prey J, Syed N, Smith P, Harrison PR, et al. DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell. 2006;126:121–34.
Mah LY, O’Prey J, Baudot AD, Hoekstra A, Ryan KM. DRAM-1 encodes multiple isoforms that regulate autophagy. Autophagy. 2012;8:18–28.
Zhang Y, Yao E, Liu Y, Zhang Y, Ding M, Liu J. et al. FUT2 facilitates autophagy and suppresses apoptosis via p53 and JNK signaling in lung adenocarcinoma cells. Cells. 2022;11:4031
Chen C, Liang QY, Chen HK, Wu PF, Feng ZY, Ma XM, et al. DRAM1 regulates the migration and invasion of hepatoblastoma cells via the autophagy-EMT pathway. Oncol Lett. 2018;16:2427–33.
Galavotti S, Bartesaghi S, Faccenda D, Shaked-Rabi M, Sanzone S, McEvoy A, et al. The autophagy-associated factors DRAM1 and p62 regulate cell migration and invasion in glioblastoma stem cells. Oncogene. 2013;32:699–712.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A, et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Investig. 2003;112:1809–20.
Poillet-Perez L, White E. Role of tumor and host autophagy in cancer metabolism. Genes Dev. 2019;33:610–9.
Samarkina A, Youssef MK, Ostano P, Ghosh S, Ma M, Tassone B, et al. Androgen receptor is a determinant of melanoma-targeted drug resistance. Nat Commun. 2023;14:6498.
Author information
Authors and Affiliations
Contributions
Study conception and design, ZD, BW, and JY; Data acquisition: ZD, BW, DW, JQ, and FG; Data analysis and interpretation, ZD and FG; Data collection, ZD, DW, and JQ; Paper preparation: ZD and BW; Paper review, ZD and FG. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
Written informed consent for the publication of identifiable human tissue images, including immunohistochemical staining images of melanoma tissue samples, was obtained from all corresponding human tissue donors in accordance with the ethical standards of the tissue microarray provider.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Dr Alberto Gandarillas
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhi, D., Wu, B., Yang, J. et al. Androgen receptor-dependent DRAM1 activation drives autophagic resistance to BRAF inhibitors in BRAFV600-mutant melanoma. Cell Death Dis 17, 265 (2026). https://doi.org/10.1038/s41419-026-08547-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-026-08547-x