Abstract
Acute kidney injury (AKI) can progress to chronic kidney disease (CKD) in the setting of maladaptive repair characterized by tubular atrophy, inflammation, and fibrosis. Programmed cell death is a key driver of proximal tubule (PT) loss, yet how immune infiltration promotes tubular injury and death remains incompletely understood. Using a mouse model of maladaptive repair, we integrated bulk and single-cell RNA sequencing with immunohistochemistry and protein analyses to define immune-epithelial interactions during AKI-to-CKD transition. Injured kidneys exhibited loss of healthy PTs, expansion of injured PT subsets, and late-stage T cell accumulation. Apoptotic and necroptotic signaling pathways were markedly upregulated, particularly in VCAM1+ PT cells. Cell-cell interaction analysis identified macrophage-derived Cxcl16 as the dominant chemokine mediating recruitment of Cxcr6+ T cells. Genetic deletion of Cxcr6 reduced renal T cell accumulation, cytotoxic effector expression, and activation of apoptotic (cleaved caspase-3, Bax) and necroptotic signaling (MLKL, phospho-MLKL) in PT cells. Accordingly, Cxcr6−/− mice displayed preserved PT differentiation, reduced fibrosis, and improved renal function. Together, these findings identify Cxcr6+ T cells as key mediators of immune-driven tubular cell death during maladaptive repair and suggest that targeting the CXCL16-CXCR6 axis may mitigate tubular injury and slow AKI-to-CKD progression.
Similar content being viewed by others
Introduction
Acute kidney injury (AKI) is a common clinical condition characterized by a rapid decline in renal function, often resulting from ischemic, toxic, or inflammatory insults. While the kidney is able to recover from mild AKI, severe or recurrent injury can lead to maladaptive repair, ultimately contributing to the development of chronic kidney disease (CKD) [1]. The transition from AKI to CKD is a complex process involving immune cell infiltration, persistent inflammation, tubular atrophy, and eventual tubulointerstitial fibrosis [2]. A key feature of maladaptive repair is the induction of programmed cell death, including apoptosis [3], necroptosis [4], ferroptosis [5], and pyroptosis [6] in proximal tubule (PT) cells, which exacerbates tissue loss and promotes CKD progression.
T cells have emerged as important mediators of kidney injury [7,8,9], yet the mechanisms governing their recruitment and cytotoxic activity in the injured kidney remain incompletely understood. Chemokines and their receptors are central regulators of immune cell trafficking, and CXCL16, which binds the receptor CXCR6, is expressed on T cells, including CD8+ T cells [10,11,12]. CXCR6+ T cells can be recruited to sites of inflammation, where they contribute to tissue damage via cytotoxic and pro-inflammatory pathways, potentially amplifying PT cell death [13].
In this study, we investigated the role of Cxcl16-Cxcr6 signaling in maladaptive kidney repair following AKI, focusing on its contribution to PT cell programmed death. Using single-cell RNA sequencing (scRNA-seq) and in vivo models of ischemia/reperfusion injury (IRI), we characterized interactions between CXCL16-expressing myeloid cells and CXCR6+ T cells, and evaluated how CXCR6 deficiency impacts T cell recruitment, tubular cell death, and renal functional recovery. Our findings provide novel mechanistic insights into how immune-mediated cell death drives AKI-to-CKD progression and highlight potential therapeutic targets to preserve renal function.
Methods
Animal surgery and experimental protocol
All animal protocols were approved by the Yale University Institutional Animal Care and Use Committee (IACUC; Protocol ID #: 20493). All methods were performed in accordance with the relevant guidelines and regulations.
C57BL/6J wild-type (WT) and Cxcr6-/- male mice (Jackson Laboratory, Strain #:005693) (age 9–10 weeks) were used in this work. All mice were maintained on a 12-h light and 12-h dark cycle with free access to standard rodent chow and water before and after surgery. Due to the substantial gender difference in susceptibility to IRI injury between male and female mice [14], male mice were exclusively used to reduce total numbers of mice required for statistical analysis. Before surgery, all mice were subjected to anesthesia by intraperitoneal injection with ketamine (100 mg/kg) and xylazine (10 mg/kg) as well as phosphate-buffered saline (1× PBS) and subcutaneous injection with Etiqa buprenorphine (3.25 mg/kg) to avoid dehydration and postoperative pain, respectively. To establish the unilateral IRI (U-IRI) model, mouse abdomen was opened, and warm renal ischemia was induced using a nontraumatic microaneurysm clip (FST Micro Clamps) on the left renal pedicle for 17 min on a 37 °C warming pad, leaving the right kidney intact. Blood was collected by retroorbital bleeding on day 1 after U-IRI for serum KIM-1 analysis. Mice were sacrificed 1 day (n = 5 mice/genotype) or 14 days (n = 8 mice/genotype) after U-IRI, and kidneys were harvested for RNA, protein, and histology analysis. Age-matched uninjured mice were sacrificed as baseline controls (n = 8 mice for each genotype). To determine the function of the injured kidney, the mouse dorsal flank was opened, and nephrectomy of the right kidney was performed on day 14 after U-IRI. Blood was collected on days 1, 3, 7, and 14 after contralateral nephrectomy for blood urea nitrogen (BUN) measurement using Stanbio™ BUN liquid reagent for diagnostic set (Thermo Fisher Scientific) and serum creatinine measurement using LC-MS/MS by Yale O’Brien Kidney Center.
ELISA of serum kidney injury molecule-1 (KIM-1) level
Mouse blood was collected at the indicated time points. Serum KIM-1 concentrations were measured using the mouse TIM-1/KIM-1/HAVCR Quantikine ELISA Kit (R&D Systems) according to the manufacturer’s instructions.
Histology
Kidneys were fixed in 10% formalin and embedded in paraffin processed for histology. To quantify tubular casts, deparaffinized kidney sections were stained with hematoxylin and eosin (H&E). For detection of collagen, deparaffinized kidney sections (5 µm) were rehydrated, stained with Picrosirius red in 1.3% picric acid for 1 h. Slides were then scanned at the Yale Pathology Tissue Services core facility and processed using ImageScope software. The percent area of cast or Picrosirius red staining was quantified using ImageJ (National Institutes of Health, NIH).
Immunohistochemistry (IHC) and Immunofluorescence (IF)
Formalin-fixed and paraffin-embedded kidney sections were deparaffinized, rehydrated, antigen retrieved, and stained with primary monoclonal antibodies against CD3ε and CD8α (#99940 and #98941, respectively; Cell Signaling Technology), SOX9 (#82630, Cell Signaling Technology), or biotinylated Lotus Tetragonolobus Lectin (LTL) (#B-1325-2, Vector Laboratories). After washing with TBST, the sections were incubated with biotinylated secondary antibody (Vector Laboratories) followed by VECTASTAIN Elite ABC system (Vector Laboratories). DAB (Vector Laboratories) and hematoxylin (Vector Laboratories) were used as the chromogen and the nuclear counterstain, respectively. IHC staining for TUNEL was conducted by Yale Pathology Tissue Services (YPTS). Whole-slide scanning was conducted by YPTS. LTL-positive area in the cortex was quantified by ImageJ. The number of CD3ε-, CD8α-, TUNEL-, or SOX9-positive cells was counted per high-power field (HPF) from six independent fields per kidney.
Cleaved caspase 3, phospho-MLKL, KIM-1, VCAM-1, LTL, CXCR6, and F4/80 were detected by IF using primary antibodies against, cleaved caspase 3 (clone: 5A1E, Cell Signaling Technology), MLKL (phospho S345) (#37333, Cell Signaling Technology), KIM-1 (#AF1817, Novus Biologicals), VCAM-1 (#32653, Cell Signaling Technology), Ki67 (#12202, Cell Signaling Technology), TUNEL-fluorescein (#11684795910, Roche), LTL-fluorescein (#L32480, Thermo Fisher Scientific), CXCR6 (#NLS1102, Novus Biologicals), and F4/80 (#MCA497, Bio-Rad), respectively. The sections were counterstained with DAPI (Sigma-Aldrich) and mounted with VECTASHIELD® HardSet™ Antifade Mounting Medium. The fluorescence images were obtained by confocal microscopy (Zeiss LSM 880) using tile-scanning. The percentage of KIM-1- and VCAM-1-positive area was quantified using ImageJ.
In vitro cell culture
Isolation and culture of bone marrow-derived macrophages (BMMs)
BMMs were isolated from Wild-type or myeloid differentiation factor 88/TIR-domain-containing adapter-inducing interferon-β (Myd88−/−;Trif−/−) mice using our recently reported protocol [15, 16]. Briefly, bone marrow cells were flushed from the femurs of 8–10-week-old mice, and red blood cells were lysed in RBC lysis buffer (Thomas Scientific). Remaining cells were incubated in RPMI 1640 medium (Life Technologies) supplemented with 10% FBS (Life Technologies), 10 μM macrophage-colony stimulating factor (M-CSF) (GenScript), 1% glutamine (Life Technologies), 1% MEM vitamin (Life Technologies), and 1% penicillin/streptomycin (Life Technologies) in regular tissue culture dishes. After 24 h, nonadherent cells (containing the majority of BMMs) were transferred to a new petri dish and incubated for an additional 7 days (with one medium change on day 3) to generate naïve BMMs.
Cultured BMMs were detached with 2 mM EDTA in PBS and plated in 6-well plates in the above mentioned complete media for 24 h. Wilt-type BMMs were treated with one of the following agent: 500 µM H2O2, 25 ng/mL GM-CSF (R&D Systems), 100 ng/mL interferon γ (R&D Systems), 20 ng/mL TNFα (R&D Systems), or 20 ng/mL IL-1β (R&D Systems) for 6 h. In a separated experiment, wild-type BMMs were treated with TNFα (20 ng/mL) or IL-1β (20 ng/mL) in the presence or absence of a selective IKKα and IKKβ inhibitor, ACHP (1 µM) (R&D Systems) for 6 h. Myd88−/−;Trif−/− BMMs were treated with TNFα (20 ng/mL) or IL-1β (20 ng/mL) for 6 h. After treatment, cell lysates were prepared with RLT buffer (Qiagen) supplemented with fresh β-mercaptoethanol for RNA extraction.
Culture of mouse proximal tubule cells (MPTs) and primary cultured renal cells (PCRCs)
Pathogen-free immortalized mouse proximal tubule (MPT) cells were plated in 6-well plates in Dulbecco’s modified Eagle medium (DMEM) (Gibco) supplemented with 10% FBS and antibiotic-antimycotic (Gibco) for 24 h. Cells were then treated for 6 h with one of the following: MPT cell debris, 500 µM H₂O₂, TNF-α (20 ng/mL), IFN-γ (100 ng/mL). Following treatment, cells were lysed in RLT buffer supplemented with freshly added β-mercaptoethanol for RNA extraction.
PCRCs were isolated from Wild-type or Myd88−/−;Trif−/− mice using our recently reported protocol [17]. Briefly, kidneys were harvested and minced into 2 mm3 cubes with a razor blade, and digested in Liberase (5 mL per kidney, 0.5 mg/mL; Roche), supplemented with DNase I (100 µg/mL, Roche) and MgCl2 (0.1%) in PBS for 30 min, with gentle pipetting every 10 min. The digested tissue was passed through a 70 µm cell strainer (Falcon) into a 50 mL conical tube, rinsed with 45 mL ice-cold PBS, and centrifuged at 1000 rpm for 5 min at 4 °C. The resulting pellet, containing single cells and tubular fragments, was resuspended with 3 mL red blood cell lysis buffer (Alfa Aesar) and incubated at room temperature for 3 min to remove residual erythrocytes. Cells were then cultured in complete medium consisting of DMEM supplemented with 10% FBS and Antibiotic-Antimycotic solution. After an initial medium change 48 h after seeding, cultures were maintained for an additional 3 days. PCRCs were subsequently trypsinized and replated in 6-well plates and allowed to adhere for 24 h.
In a separate experiment, MPT cells or PCRCs were treated with TNF-α (20 ng/mL) or IL-1β (20 ng/mL) in the presence or absence of the selective IKKα/IKKβ inhibitor ACHP (1 µM) for 6 h. Following treatment, cells were lysed in RLT buffer supplemented with freshly added β-mercaptoethanol for RNA extraction.
Quantitative PCR analysis
RNA from whole kidney or cell lysate was extracted with RNeasy Mini kit (Qiagen) and reverse transcribed using the iScript cDNA synthesis kit (Bio-Rad). Gene expression levels were determined by quantitative real-time PCR using the iCycler iQ (Bio-Rad) and normalized to Hprt1. Primer sequences are provided in Supplementary Table 1. Data were expressed using the comparative threshold cycle (ΔCT) method, and the relative mRNA levels were presented by 2−ΔCT or ΔΔCT.
Western blot analysis
Kidney lysates from wild-type and Cxcr6−/− mice were prepared using radioimmunoprecipitation assay (RIPA) lysis and extraction buffer (Thermo Fisher Scientific) supplemented with 1× protease and phosphatase inhibitor cocktail (Roche). Protein concentrations were determined using the Bio-Rad Protein Assay. Equal amounts of protein (50 μg per sample) were separated by SDS–PAGE on 10% or 12% polyacrylamide gels (Bio-Rad) and transferred to Immobilon PVDF membranes (Millipore). Membranes were blocked with 5% nonfat milk or 5% bovine serum albumin (BSA) in Tris-buffered saline with 0.1% Tween-20 (TBST) for 2 h at room temperature and incubated overnight at 4 °C with primary antibody against cleaved caspase-3 [clone 5A1E (#9664) or #9661, Cell Signaling Technology]. After TBST washes, membranes were incubated for 1 h at room temperature with horseradish peroxidase (HRP)-conjugated secondary antibody (Thermo Fisher Scientific). Protein bands were visualized using the ECL detection system (Thermo Fisher Scientific) and imaged with the LI-COR Odyssey FC Imager. Membranes were subsequently washed with TBS, stripped in Restore PLUS Western Blot Stripping Buffer (Thermo Fisher Scientific) for 15 min, rewashed with TBST, and re-blocked. The same membranes were sequentially re-probed with primary antibodies against Bax (clone D3R2M, Cell Signaling Technology), MLKL (clone E7V4W, Cell Signaling Technology), phospho-MLKL (Ser345, clone EPR9515(2), Abcam), RIP3 (clone D4G2A, Cell Signaling Technology), and HSP90 (clone C45G5, Cell Signaling Technology), following the same protocol. Band intensity was quantified using ImageJ.
Bulk RNA sequencing (RNA-seq) data analysis of mouse IRI kidneys
Total RNA was extracted from control kidneys (n = 4) and injured kidneys (n = 5) collected on day 14 post-IRI using the RNeasy Mini kit (Qiagen) supplied with on-column RNase-free DNase I treatment (Qiagen). Library preparation was performed using Illumina’s standard PolyA enrichment protocol by the Yale Center for Genome Analysis. Sequencing was carried out on the NovaSeq platform with 100-bp paired-end reads at a depth of 50 million reads per sample. Raw reads were aligned to the mouse genome (mm10) with STAR [18], and gene expression levels were quantified with FeatureCounts [19]. Differential expression analysis between the different groups was conducted with DESeq2 [20]. Genes with an adjusted P < 0.05 were considered significantly differentially expressed. DEGs with a log2 fold change >0.25 were subjected to gene set enrichment analyses using ClusterProfiler and KEGG R packages [21].
Single cell-RNA sequencing (scRNA-seq) data analysis of mouse IRI kidneys
We re-analyzed our previously published scRNA-seq dataset (GSE197626) from the U-IRI mouse kidney model (on days 14 and 30) and control kidney using the Seurat v5.1.0 R package [22, 23]. Cells of poor quality were excluded based on the following criteria: <500 unique genes or <1000 counts (likely cell fragment), or >7500 unique genes (potentially cell duplet). Additionally, cells with mitochondrial gene content exceeding 50% and low-complexity cells, such as red blood cells with <0.8 log10 genes per UMI counts, were removed. Only genes expressed in at least five cells were retained for further analysis. Doublets were identified and excluded using the DoubletFinder R package [24]. Confounding factors, including mitochondrial genes, were removed before downstream clustering [25]. After applying quality control filters, a total of 30,323 cells remained, with a median of 5301 counts per cell and a sequencing depth of 23,438 genes across all cells. The dataset was normalized, scaled, and integrated using reciprocal principal component analysis (RPCA) on the 2000 most variable genes. The top 30 principal components were chosen for cell clustering and neighbors finding with k.param = 20. Dimensionality reduction was performed by retaining the top 30 principal components, and cell clustering was carried out with k.param = 20. Uniform manifold approximation and projection (UMAP) was used for two-dimensional visualization. Marker genes for each cluster were identified through differential expression analysis using the Wilcoxon rank-sum test, considering genes expressed in at least 25% of cells either within or outside the cluster. This analysis identified 30 distinct cell clusters based on kidney and immune cell lineage-specific marker expression. Cell interaction analysis was performed using CellChat v2 R package [26]. Finally, the relative expression levels of Ccl6, Ccr2, Cxcl16, and Cxcr6 were analyzed across all the cell clusters and time points. Gene set enrichment analysis across multiple PT clusters was analyzed using the singleseqgset R package.
Statistical analysis
All in vitro experiments were repeated at least three times. Data were expressed as means ± standard deviation (SD). Two-group comparison was performed by two-tailed unpaired Student’s t test. Multigroup comparison was performed by one-way analysis of variance (ANOVA) for group mean comparison followed by Tukey’s multiple comparison test for subgroup comparison. Correlation of gene expression was performed by Pearson correlation coefficient R with a two-tailed P value. All statistical analyses were performed using Prism 10 (GraphPad Software). A value of P < 0.05 was considered statistically significant.
Results
Upregulation of apoptosis and necroptosis during AKI-to-CKD transition
The transition from AKI to CKD is commonly associated with tubular cell death and progressive kidney atrophy [22]. To identify the predominant cell death mechanisms involved, we first performed bulk RNA-seq analysis of kidneys harvested 14 days after IRI. Multiple genes associated with cell death pathways were upregulated in IRI kidneys compared with controls (Fig. 1A). Pathway enrichment analysis revealed apoptosis and necroptosis as the top-enriched cell death pathways (Fig. 1B). To validate these findings, we assessed protein expression of key regulators of apoptosis and necroptosis at the whole-kidney level. Cleaved caspase-3 and Bax showed minimal or modest induction on day 1 but were markedly increased on day 14 post-IRI. In contrast, RIPK3, MLKL, and phosphorylated MLKL were strongly upregulated as early as day 1 and remained highly expressed on days 14 and 28 (Fig. 1C, D). These results suggest that necroptosis is engaged throughout initial injury and maladaptive repair, whereas apoptosis is selectively upregulated at the time of kidney atrophy during the AKI-to-CKD transition. IF co-staining with TUNEL reveals that the majority of TUNEL-positive tubular cells co-express either cleaved caspase-3 or phosphorylated MLKL, confirming the presence of both apoptotic and necroptotic cell death following U-IRI (Fig. 1E and Supplementary Fig. 2). To assess whether cell death pathways were specifically upregulated in PT cells, we re-analyzed our scRNA-seq dataset from mouse kidneys subjected to U-IRI at days 14 and 28 [22]. Consistent with previous reports, we observed a decline in healthy PT populations accompanied by an expansion of injured PT and T cell populations (Supplementary Fig. 3A, B). At the single-cell level, KEGG pathway enrichment analysis confirmed a significant upregulation of apoptosis in injured PT cells, along with activation of natural killer (NK) cell–mediated cytotoxicity, T cell receptor signaling, and cytokine-receptor interactions (Fig. 1F). These findings indicate that PT cells, particularly VCAM1+ subsets, are subject to both apoptosis and immune cell–mediated necroptosis during the AKI-to-CKD transition.
A, B Wild-type mice were subjected to unilateral ischemia/reperfusion injury (IRI), and the injured kidneys were collected on day 14 post-injury for bulk RNA-seq analysis. Age-matched healthy kidneys served as controls (CTRL). A Differentially expressed genes (DEGs) were identified using DEseq2 and visualized in a heatmap (n = 4 CTRL and n = 5 IRI kidneys). B KEGG pathway enrichment analysis of upregulated DEGs in IRI kidneys. Cell death pathways (apoptosis, necroptosis, ferroptosis, pyroptosis, and autophagy) are highlighted in the bubble plot. C, D Wild-type mice were subjected to unilateral IRI, and the injured kidneys were harvested on days 1, 14, and 28 post-injury. Age-matched healthy kidneys served as controls (CTRL). C Western blot analysis of cleaved caspase 3, Bax, phospho-MLKL, MLKL, RIP3, and HSP90 (loading control, re-probed after stripping) from whole kidney lysates (each lane represents an individual kidney). The full length uncropped original Western blots were provided in Supplementary Fig. 1. D Densitometric quantification of protein bands. Expression of cleaved caspase-3, Bax, phospho-MLKL, MLKL, and RIP3 was normalized to HSP90. n = 6 kidneys/group. One-way ANOVA: P = 0.0004 (cleaved caspase 3), P < 0.0001 (Bax), P = 0.0034 (phospho-MLKL), P = 0.0003 (MLKL), and P = 0.0138 (RIP3). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison as compared to CTRL. E Wild-type mice were subjected to U-IRI, and the injured kidneys were harvested on day 14 post-injury. Control (CTRL) kidneys were obtained from healthy uninjured mice. Kidney sections were immunofluorescence-stained for cleaved caspase 3 (upper panel, red), phosphorylated MLKL (pMLKL; lower panel, red), TUNEL (green), and DAPI (blue). Individual channels are shown in Supplemental Fig. 2. Original magnification, ×63. Scale bar: 50 μm. (F) Wild-type mice were subjected to unilateral (IRI), and the injured kidneys were collected on days 14 and 30 post-injury for single-cell RNA sequencing analysis as shown in Supplementary Fig. 2A and 1B. KEGG pathway enrichment analysis was performed on healthy and injured proximal tubule (PT) cells and visualized as a heatmap.
Identification of T cell homing signals during AKI-to-CKD transition
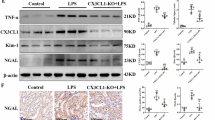
Previous studies have shown that maladaptive repair is characterized by late T cell infiltration which strongly correlates with kidney atrophy [22, 27]. To identify the signals mediating T cell recruitment, we performed cell-cell interaction analysis using the CellChat R package, focusing on chemokine-chemokine receptor interactions in injured kidneys. This analysis identified Ccl6-Ccr2 and Cxcl16-Cxcr6 as the top ligand-receptor pairs mediating interactions between T cells and other cell types (Fig. 2A). Specifically, Cxcl16-Cxcr6 was the predominant homing signal for CD8+ T cells (Fig. 2B), whereas Ccl6-Ccr2 primarily mediated recruitment of CD4+ and naïve T cells (Fig. 2C and Supplementary Fig. 3C). Further analysis revealed that F4/80+ macrophages were the main source of Cxcl16 driving CXCR6+ CD8+ T cell recruitment during the AKI-CKD transition (Fig. 2D); whereas myeloid cells broadly supported CD4+ T cell recruitment via Ccl6-Ccr2 signaling (Supplementary Fig. 3D). Expression analysis confirmed that Cxcl16 was predominantly expressed by Adgre1+ (F4/80+) macrophages and cDC2s, while Cxcr6 was broadly expressed in naïve T cells, CD4+ T cells, CD8+ T cells, and NKT cells (Fig. 2E). In addition, injured PT cells also upregulate Cxcl16, albeit at lower levels than macrophages and dendritic cells (Supplemental Fig. 3E), suggesting that tubular cells contribute as an auxiliary source of CXCL16, likely reinforcing T cell recruitment within the inflamed microenvironment. Consistent with these predictions, immunofluorescence staining showed that F4/80+ macrophages were located in close proximity to CXCR6+ cells 14 days after U-IRI (Fig. 2F). To further test the role of macrophages in Cxcl16 production, we analyzed day 14 U-IRI kidneys from Ccr2−/− mice, in which macrophage and dendritic cell recruitment is reduced by 30% and 50%, respectively [28]. Loss of Ccr2 resulted in a 33% reduction in Cxcl16 expression (Fig. 2G), supporting the hypothesis that recruited macrophages are a major source of Cxcl16.
Wild-type mice were subjected to unilateral ischemia/reperfusion injury (U-IRI), and the injured kidneys were collected on day 14 and 30 post-injury for single-cell RNA sequencing analysis as shown in Supplementary Figs. 2A and 1B. A–C Ligand-receptor interaction analysis using CellChat R package. The contribution of each ligand-receptor pair to the overall signaling pathway (CCL and CXCL) in overall T cells (A), CD8+ T cell (B), and CD4+ T cell (C) was computed and visualized in a bar graph, respectively. D The cell-cell communication mediated by Cxcl16-Cxcr6 pair was visualized in a chord hierarchy plot. E The distribution and relative expression of top pairs of ligand-receptor genes were visualized using a dot plot. PT proximal tubule, TAL thick ascending limb, DCT distal convoluted tubule, CNT connecting tubule, PC principal cell, CD-IC collecting duct-intercalated cell, EC-AEA endothelia cell-afferent/efferent arteriole, EC-PTC peritubular endothelia cell, MyoF myofibroblast, Infil. Neut infiltrating neutrophil, Inflam. Neut inflammed neutrophil, Degranul. Neut degranulated neutrophil, Infil. Mac infiltrating macrophage, Resid. Mac resident macrophage, pDC plasmacytoid dendritic cell, cDC conventional dendritic cell, T T cells, NK natural killer cells, B B cells, Prolif proliferating cells. F Kidney cryosections were immunofluorescence-stained with CXCR6 (red), F4/80 (identifies macrophages, green), and DAPI (identifies nuclei, blue) at day 14 after unilateral IRI. Original magnification, ×400. Scale bar: 20 μm. Asterisks (*) indicate representative macrophages adjacent to CXCR6⁺ cells (denoted by #). G Wild-type and Ccr2−/− mice were subjected to unilateral IRI. IRI and contralateral (CL) kidneys were harvested 30 days after IRI. Quantitative PCR for Cxcl16 was performed on whole-kidney mRNA. Two-way ANOVA: P < 0.0001 (injury and genotype factors) and P = 0.0007 (interaction). ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. Wild-type bone marrow-derived macrophages (BMMs, H), MPT cells (J), and primary cultured renal cells (PCRCs, K) were treated with PBS (control), interleukin (IL-1β, 10 ng/mL), or tumor necrosis factor (TNF-α) (20 ng/mL) with or without the NF-κB inhibitor ACHP (1 μM) for 6 h. Quantitative PCR for Cxcl16 was performed on BMM mRNA. n=3 independently treated experiments. One-way ANOVA: P < 0.0001. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. Myd88−/−;Trif−/− BMMs (I) and PCRCs (L) were treated with PBS (control), IL-1β (20 ng/mL), or TNF-α (20 ng/mL) for 6 h. Quantitative PCR for Cxcl16 was performed on BMM mRNA. n = 3 independently treated experiments. One-way ANOVA: P < 0.0001 (BMM) and P = 0.0011 (PCRC). *P < 0.05 and ****P < 0.0001 by Tukey’s multiple comparison. ns, not statistically significant.
We next sought to define the pathway regulating Cxcl16 expression in macrophages and injured PT cells. Since IL-1β, TNF-α, and IFN-γ have been reported as CXCL16 inducers [29, 30], we stimulated naïve BMMs and MPT cells with these cytokines. Both IL-1β and/or TNF-α markedly increased Cxcl16 expression (4–5.6 fold, respectively), whereas IFN-γ, H2O2, or GM-CSF had no or mild effect (Supplementary Fig. 4). Because IL-1β and TNF-α activate NF-κB, we tested the role of NF-κB signaling by treating BMMs, MPT cells, and primary cultured renal cells (PCRCs) with these cytokines in the presence of the IKKα/IKKβ inhibitor ACHP. IKK inhibition suppressed Cxcl16 induction by ~32% in BMMs (Fig. 2H), ~50% in MPT cells (Fig. 2J), and 32% in PCRCs (Fig. 2K), respectively. Moreover, BMMs from Myd88/Trif-deficient mice, which lack the MyD88 scaffold required for IL-1β/IL-1R1-mediated NF-κB activation [30], failed to upregulate Cxcl16 in response to IL-1β but responded normally to canonical TNF-α/TNFR1 activation (Fig. 2I); similar results were observed in PCRCs (Fig. 2L).
Together, these results indicate that following kidney injury, recruited macrophages as the predominant source of Cxcl16, with injured PT cells contributing as an auxiliary source. Both cell types upregulate Cxcl16 in response to IL-1β and TNF-α via NF-κB-dependent signaling, facilitating CXCR6+ T cell recruitment to the interstitium and contributing to maladaptive kidney repair.
CXCR6 promotes T cell accumulation during tubule atrophy
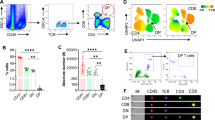
To evaluate the role of CXCL16-CXCR6 signaling in T cell recruitment during AKI-to-CKD transition, we performed U-IRI in wild-type and Cxcr6−/− mice. Serum KIM-1 levels, a marker of acute PT injury, were equally elevated in both groups 1 day post-U-IRI (Fig. 3A), indicating equivalent initial injury. Consistent with the scRNA-seq data, wild-type mice showed a significant increase in Cxcr6 mRNA 14 days after U-IRI, whereas Cxcr6 expression was undetectable at both time points in kidneys from Cxcr6−/− mice (Fig. 3B). IHC staining for CD3ε and CD8α demonstrated a decrease in total T cells and CD8α+ cytotoxic T cells in Cxcr6−/− kidneys on day 14 after U-IRI as compared to wild-type kidneys (Fig. 3C, D). Consistent with reduced T cell accumulation, whole-kidney mRNA expression of Cd3e, Cd4, and Cd8a was significantly decreased in Cxcr6−/− at this time point (Fig. 3E), suggesting that CXCR6 deficiency is associated with reduced recruitment of both CD8α+ and CD4+ T cell populations. In comparison, Cxcr6 expression was low to absent in Foxp3+ regulatory T cell (Treg) and macrophage clusters (Supplementary Fig. 5A). Consistent with this, whole-kidney mRNA expression of canonical Treg markers (Foxp3, Ctla4, and Il2ra) and macrophage marker Adgre1 (F4/80) did not differ significantly wild-type and Cxcr6−/− injured kidneys (Supplementary Fig. 5B and Fig. 3E), consistent with the finding that Cxcr6 is not expressed on macrophage, monocyte, DC, or Treg populations in the injured kidney (Fig. 2E) [28]. These results indicate that CXCL16-CXCR6 signaling specifically promotes T cell recruitment without affecting macrophage accumulation or Treg-mediated reparative response [31, 32].
Wild-type and Cxcr6−/− mice were subjected U-IRI. A Serum KIM-1 (sKIM-1) levels of mice were determined by ELISA in control and injured mice on day 1 after U-IRI. n = 11–16 mice for baseline/genotype and n = 23, 24 mice for IRI/genotype. Two-way ANOVA: P = 0.4519 (genotype), P < 0.0001 (injury factor), and P = 0.4475 (interaction). ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. B Healthy control (CTRL) or injured (IRI) kidneys were harvested 14 days after unilateral IRI. Quantitative PCR for Cxcr6 was performed on whole-kidney mRNA. Two-way ANOVA: P < 0.0001 (genotype), P = 0.0055 (injury factor), and P = 0.0055 (interaction). **P < 0.01 and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. C Kidney sections were immunostained with CD3ε (left panel) and CD8α (right panel) (representative images shown). Scale bars, 100 µm. D Quantitation of CD3ε- and CD8α-cell numbers per high power field (HPF), as in (C). n = 4 CTRL kidneys/genotype and n = 8 IRI kidneys/genotype. Two-way ANOVA (injury and genotype interaction): P = 0.0036 (CD3ε) and P = 0.0442 (CD8α). **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns, not statistically significant. E Quantitative PCR for Cd3e, Cd4, Cd8a, and Adgre1 was performed on whole-kidney mRNA. n = 8 mice/genotype. Two-way ANOVA (injury and genotype interaction): P < 0.0199 (Cd3e), P = 0.0255 (Cd4), P = 0.0305 (Cd8a), and P = 0.3698 (Adgre1). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. F, G Kidney sections were immunofluorescence-stained with KIM-1 (identifies injured PT, green), CD3ε (identifies T cell, red, F), CXCR6 (magenta, G), and DAPI (identifies nuclei, blue). Original magnification, ×40. Scale bar: 20 μm.
CXCR6 promotes PT cell death via apoptosis and necroptosis
In wild-type injured kidneys, CD3ε+ and CXCR6+ T cells were frequently localized around KIM-1+ PT, whereas markedly fewer CD3ε+ T cells were observed in Cxcr6−/− kidneys (Fig. 3E–G). To investigate the functional role(s) of Cxcr6-expressing T cells in promoting cell death pathway activation, we assessed markers of apoptosis and necroptosis. The increase in cleaved caspase-3 and Bax protein levels seen in the early phase of tubule atrophy in wild-type mice (day 14) was reduced by 23% and 41%, respectively, in Cxcr6−/− kidneys (Fig. 4A, B). Similarly, Ripk3, Trp53, and Mlkl (mRNA) as well as MLKL and phospho-MLKL (protein) were all reduced on day 14 in the absence of Cxcr6 as compared to wild-type injured kidneys (Fig. 4A–C). In contrast, one day after injury, T cell markers (Cd3e, Cd4, and Cd8a) were not upregulated in either genotype, and Cxcr6 deletion had no effect on their expression (Supplementary Fig. 7A). Similarly, phosphorylated MLKL protein levels and necroptosis-related or injury-related transcripts (Mlkl, Ripk3, and Havcr1) were not altered by Cxcr6 deletion (Supplementary Fig. 7B–D), consistent with unchanged serum KIM-1 levels (Fig. 3A). These data indicate that CXCR6 does not contribute to injury initiation, but rather promotes persistent immune-driven damage during the maladaptive repair phase.
Wild-type and Cxcr6−/− mice were subjected unilateral ischemia/reperfusion injury (IRI). Healthy control (CTRL) or injured (IRI) kidneys were harvested 14 days after IRI. A Western blot analysis of cleaved caspase 3, Bax, phospho-MLKL, MLKL, and HSP90 (loading control, re-probed after stripping) from whole kidney lysates (each lane represents an individual kidney). The full length uncropped original Western blots were provided in Supplementary Fig. 6. B Densitometric quantification of protein bands. Expression of cleaved caspase-3, Bax, phospho-MLKL, and MLKL was normalized to HSP90. n = 5 CTRL and n = 7 IRI kidneys/genotype. Two-way ANOVA (injury and genotype interaction): P = 0.1096 (cleaved caspase 3), P = 0.0027 (Bax), P = 0.0061 (phospho-MLKL), and P = 0.0278 (MLKL). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. C Quantitative PCR was performed on whole-kidney mRNA. n = 8 mice/genotype. Two-way ANOVA (injury and genotype interaction): P = 0.0030 (Mlkl), P < 0.0001 (Ripk3), P = 0.2288 (Trp53), P = 0.0943 (Fasl), P = 0.0187 (Gzmb), P < 0.0001 (Tnf), and P = 0.2642 (Il1b). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. D Kidney sections were immunofluorescence-stained as indicated (representative images shown): LTL (green), pMLKL (red), and DAPI (blue) (left panel); KIM-1 (green), pMLKL (red), and DAPI (blue) (middle panel); and KIM-1 (green), cleaved caspase 3 (red), and DAPI (blue) (right panel). Original magnification, ×40. Scale bar: 50 μm. E Kidney sections were immuno-stained with TUNEL (representative images shown). Scale bar: 50 μm. Arrows indicate TUNEL-positive tubular cells. F Quantitation of TUNEL-positive tubular cell per high power field (HPF), as in (E). n = 3 CTRL kidneys/genotype and n = 6 IRI kidneys/genotype. Two-way ANOVA (genotype and injury interaction): P = 0.0429. **P < 0.01 and ***P < 0.001 by Tukey’s multiple comparison. ns not statistically significant.
We next examined cytotoxic effector molecules expressed by CD8+ T cells. Fasl, Gmzb, and Tnf were all significantly increased in wild-type injured kidneys at day 14 post-U-IRI, but their expression levels were reduced by 48%, 69%, and 48%, respectively in the absence of Cxcr6 (Fig. 4C). In contrast, the increase in Il1b, which is primarily expressed by myeloid cells, was equivalent in wild-type and Cxcr6−/− mice on day 14 after U-IRI, consistent with unchanged Adgre1 expression (Fig. 4C).
Morphologically, cleaved caspase-3 was primarily localized to a subset of KIM-1-positive PT cells in wild-type injured kidneys (Fig. 4D, right panel), while phospho-MLKL was detected at the membranes of KIM-1-positive PT cells and other distal tubular cells (Fig. 4D, middle panel), but was largely absent from LTL-positive “healthy” PT cells in wild-type injured kidneys (Fig. 4D, left panel). In Cxcr6−/− injured kidneys, KIM-1+, cleaved caspase-3+, and phospho-MLKL+ cells appeared less frequently (Fig. 4D). Lastly, tubular TUNEL positivity was significantly reduced in Cxcr6−/− kidneys compared with wild-type kidneys after IRI (Fig. 4E, F), providing quantitative support for reduced cell death in the absence of Cxcr6. Together, these findings suggest that CXCR6+ T cells promote both apoptotic and necroptotic PT cell injury during AKI-to-CKD transition.
Recruitment of CXCR6+ cells impairs functional renal recovery
Compared to wild-type kidneys, the expression levels of KIM-1 (Havcr1) and VCAM-1 (Vcam1) were both significantly lower in Cxcr6−/− injured kidneys (Fig. 5A–C). In addition, Cxcr6−/− kidneys exhibited preserved expression of the PT differentiation marker LTL (Fig. 5D, E), decreased cast accumulation (Fig. 5F, G), diminished expression of the dedifferentiation marker SOX9 (Fig. 5H–J), and attenuated interstitial fibrosis on day 14 post-U-IRI (Supplementary Fig. 8). However, the Cxcr6 null kidneys exhibited no difference in tubular proliferation or kidney weight on day 14 after U-IRI as compared to wild-type kidneys (Supplementary Figs. 9 and 10). To determine if the preservation of PT differentiation seen in Cxcr6−/− kidneys resulted in improved kidney function, we performed contralateral nephrectomy on day 14 post-U-IRI and measured BUN and serum creatinine afterwards (Fig. 5K). One day after nephrectomy (day 15), Cxcr6−/− mice exhibited significantly less rise in both BUN and serum creatinine demonstrating that the function of the injured kidney was better preserved in these mice (Fig. 5L, M). While GFR slowly improved in both groups over the next 2 weeks, the Cxcr6−/− mice consistently exhibited lower BUN and creatinine values. These findings indicate that CXCR6+ T cells exacerbate PT injury and reduce the degree of recoverable renal function during the AKI-to-CKD transition.
A–D Wild-type and Cxcr6-/- mice were subjected unilateral ischemia/reperfusion injury (IRI). The injured (IRI) kidneys were harvested 14 days after U-IRI. A Kidney sections were immunofluorescence-stained with KIM-1 (green), VCAM-1 (red), and DAPI (blue). Original magnification, ×40. Scale bar: 200 μm. B Fluorescence positivity was quantified using ImageJ. n = 4 CTRL kidneys/genotype and n = 8 IRI kidneys/genotype. Two-way ANOVA (injury and genotype interaction): P = 0.0623 (KIM-1) and P = 0.0060 (VCAM-1). *P < 0.05, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. C Quantitative PCR for Havcr1 and Vcam1 was performed on whole-kidney mRNA. n = 8 mice/genotype. Two-way ANOVA (injury and genotype interaction): P < 0.0001 (Havcr1) and P = 0.0006 (Vcam1). ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. D Kidney sections were immune-stained with LTL (dark gray) (representative images shown), and LTL positive area was quantified in (E). F Kidney sections were stained for H&E (representative images shown), and cast area was quantified in (G). Green arrows in D indicate LTL-positive tubule. Red arrows in D and black arrows in F indicate tubular cast. Scale bar: 200 μm. n = 8 IRI kidneys/genotype. *P < 0.05 and **P < 0.01 by unpaired t test. H Kidney sections were immunohistochemistry-stained for SOX9 (representative images shown). Scale bar: 50 μm. Arrows indicate SOX9-positive tubular cells. I Quantitation of SOX9-positive tubular cell per high power field (HPF), as in (H). n = 4 CTRL kidneys/genotype and n = 8 IRI kidneys/genotype. Two-way ANOVA (genotype and injury interaction): P = 0.0032. *P < 0.05 and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. J Quantitative PCR for Sox9 was performed on whole-kidney mRNA. n = 8 mice/genotype. Two-way ANOVA (injury and genotype interaction): P = 0.0151. **P < 0.01 and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant. K Scheme of experimental design. Wild-type and Cxcr6−/− mice were subjected U-IRI, followed by contralateral nephrectomy (CL-NX) on day 14 after U-IRI. The blood was withdrawn for blood urea nitrogen (BUN in L) and serum creatinine (SCr in M) at the indicated time points. n = 14, 15 mice/genotype. Two-way ANOVA (injury and genotype interaction): P < 0.0001 (BUN and SCr) by two-way ANOVA. *P < 0.05, ***P < 0.001, and ****P < 0.0001 by Tukey’s multiple comparison. ns not statistically significant.
Discussion
Our findings highlight a central role for CXCL16-CXCR6 signaling in mediating T cell recruitment during AKI-to-CKD transition. The accumulation of CXCR6+ T cells in the injured kidney suggests that persistent immune activation contributes to prolonged tubular damage, interstitial fibrosis, and impaired renal function. Mechanistically, macrophage-derived CXCL16 serves as a dominant chemotactic signal for CXCR6-expressing T cells, which in turn promote PT injury through cytotoxic mechanisms involving both apoptosis and necroptosis.
Progressive epithelial cell loss is a defining feature of AKI-to-CKD transition and CKD progression, and regulated forms of PT cell death, including necroptosis, ferroptosis, pyroptosis, and apoptosis, have emerged as central drivers of maladaptive repair. Inhibition of necroptosis improves renal outcomes in subtotal nephrectomy and unilateral ureter obstruction (UUO) models [33, 34]. Inhibition of ferroptosis using liproxstatin and pyroptosis using VX-765 reduces the inflammatory response and interstitial fibrosis and ameliorates renal function 14 days after bilateral IRI [5]. On the other hand, apoptosis can also promote loss of renal epithelial cells during chronic tubular atrophy, which can be activated by FasL/Fas/Bax-mediated mitochondrial injury, ER stress, aminoglycosides, and cisplatin [35]. Single-cell regulatory network inference and clustering (SCENIC) analysis of IRI kidneys revealed increased regulon activity of Myc in the Vcam1+/Ccl2+ “late injured” PT cluster, a factor known to be involved in tubular cell apoptosis [36]. We now show that injured PTs exhibit elevated apoptotic and necroptotic signaling, evidenced by increased cleaved caspase-3, Bax, MLKL, and phospho-MLKL expression. Although MLKL phosphorylation is not sufficient to definitively establish execution of necroptosis, its presence supports activation of necroptosis-associated signaling in these cells [37], suggesting that necroptotic cell death may occur in a subset of phosphorylated MLKL-positive tubules.
Necroptosis-driven necroinflammation has emerged as a key mechanism linking unresolved AKI to chronic inflammation and fibrosis. Unlike apoptosis, necroptosis culminates in MLKL-mediated plasma membrane disruption, resulting in the release of damage-associated molecular patterns (DAMPs) that amplify innate immune signaling [38,39,40]. In mouse models of IRI, RIPK3-MLKL-dependent necroptosis promotes inflammasome activation, macrophage recruitment, and long-term interstitial fibrosis [4]. In our U-IRI model, MLKL and phospho-MLKL induction, predominantly within injured PT cells, was markedly attenuated in Cxcr6−/− kidneys, accompanied by reduced VCAM-1 expression and improved structural and functional outcomes. These findings suggest that CXCR6-dependent immune cell recruitment sustains a necroinflammatory microenvironment that perpetuates tubular injury and fibrotic remodeling. Persistent necroptotic signaling within injured tubules may reinforce immune cell retention and promote fibroblast or myofibroblast activation through paracrine inflammatory cues, forming a feed-forward loop of maladaptive repair. Disruption of this loop in Cxcr6-deficient mice appears to mitigate progression from AKI to CKD. Although the specific pathways through which CXCR6+ T cells promote necroptosis remain to be fully defined, our data suggest that this effect is likely mediated indirectly via cytokine- and death receptor-dependent signaling, rather than through direct enzymatic activity.
Upstream of this process, NF-κB-dependent activation of macrophages plays a pivotal role in shaping the inflammatory milieu of the injured kidney [41]. We demonstrate that IL-1β and TNF-α induce CXCL16 expression predominantly in macrophages, with injured PT cells also contributing via NF-κB-dependent signaling, thereby linking innate immune activation within both immune and parenchymal compartments to adaptive immune recruitment. Consistent with this model, Cxcr6 deletion reduced expression of cytotoxic effector molecules, including Fasl, Gzmb, and Tnf, implicating CXCR6+ T cells as amplifiers of tubular cytotoxicity. Prior studies have shown that CXCR6+ T cells migrate to injured kidneys and promote fibrosis through production of IL-17A and PD-1 signaling in models of bilateral IRI, UUO, and DOCA/salt hypertension [42, 43]. Although direct pharmacologic targeting of CXCR6 remains challenging, our data highlight NF-κB-CXCL16 signaling in macrophages and injured tubular cells as a tractable upstream axis to limit pathogenic T cell recruitment and immune-mediated tubular injury.
Beyond CD8α+ T cells, CXCL16-CXCR6 signaling also contributes to the recruitment of CD4+ T cell populations. While CD8α T cells are classically associated with direct cytotoxicity toward injured tubular cells, CD4 T cells may contribute indirectly by sustaining inflammatory signaling, enhancing antigen presentation, and promoting profibrotic immune responses. T helper 17 (Th17) cells, in particular, exacerbate renal inflammation and injury [7], whereas Tregs facilitate injury resolution and tissue repair [31, 32]. Notably, Cxcr6 deletion reduced recruitment of conventional CD4+ T cells without affecting Treg abundance, potentially shifting the immune balance toward a reparative phenotype. This selective modulation of pathogenic, but not protective, T cell subsets may contribute to the improved structural and functional recovery observed in Cxcr6-deficient kidneys.
In conclusion, our study establishes macrophage-derived CXCL16 as the predominant chemokine driving recruitment of CXCR6+ T cells during the AKI-to-CKD transition, with injured PT cells providing an additional local source of CXCL16 within the inflamed microenvironment. Accumulation of CXCR6+ T cells promotes sustained apoptotic and necroptotic signaling in injured tubules, exacerbating fibrosis and limiting renal recovery. Targeting the CXCL16-CXCR6 axis, or its upstream inflammatory regulators, represents a promising strategy to disrupt maladaptive immune-epithelial crosstalk and preserve kidney function, warranting further investigation in human CKD.
Data availability
The scRNA-seq dataset analyzed in this study is available in the Gene Expression Omnibus (GEO) under accession number GSE197626. The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Coca SG, Singanamala S, Parikh CR. Chronic kidney disease after acute kidney injury: a systematic review and meta-analysis. Kidney Int. 2012;81:442–8.
Fu Y, Xiang Y, Wei Q, Ilatovskaya D, Dong Z. Rodent models of AKI and AKI-CKD transition: an update in 2024. Am J Physiol Renal Physiol. 2024;326:F563–83.
Yang X, Guan Y, Bayliss G, Zhao TC, Zhuang S. SET8 inhibition preserves PTEN to attenuate kidney cell apoptosis in cisplatin nephrotoxicity. Cell Death Dis. 2025;16:226.
Chen H, Fang Y, Wu J, Chen H, Zou Z, Zhang X, et al. RIPK3-MLKL-mediated necroinflammation contributes to AKI progression to CKD. Cell Death Dis. 2018;9:878.
Balzer MS, Doke T, Yang YW, Aldridge DL, Hu H, Mai H, et al. Single-cell analysis highlights differences in druggable pathways underlying adaptive or fibrotic kidney regeneration. Nat Commun. 2022;13:4018.
Cao Y, Chen X, Zhu Z, Luo Z, Hao Y, Yang X, et al. STING contributes to lipopolysaccharide-induced tubular cell inflammation and pyroptosis by activating endoplasmic reticulum stress in acute kidney injury. Cell Death Dis. 2024;15:217.
Lee K, Jang HR, Rabb H. Lymphocytes and innate immune cells in acute kidney injury and repair. Nat Rev Nephrol. 2024;20:789–805.
Dellepiane S, Leventhal JS, Cravedi P. T cells and acute kidney injury: a two-way relationship. Front Immunol. 2020;11:1546.
Sharma R, Kinsey GR. Regulatory T cells in acute and chronic kidney diseases. Am J Physiol Renal Physiol. 2018;314:F679–f698.
Di Pilato M, Kfuri-Rubens R, Pruessmann JN, Ozga AJ, Messemaker M, Cadilha BL, et al. CXCR6 positions cytotoxic T cells to receive critical survival signals in the tumor microenvironment. Cell. 2021;184:4512–4530.e4522.
Chia TY, Billingham LK, Boland L, Katz JL, Arrieta VA, Shireman J, et al. The CXCL16-CXCR6 axis in glioblastoma modulates T-cell activity in a spatiotemporal context. Front Immunol. 2023;14:1331287.
Sato T, Thorlacius H, Johnston B, Staton TL, Xiang W, Littman DR, et al. Role for CXCR6 in recruitment of activated CD8+ lymphocytes to inflamed liver. J Immunol. 2005;174:277–83.
Mabrouk N, Tran T, Sam I, Pourmir I, Gruel N, Granier C, et al. CXCR6 expressing T cells: functions and role in the control of tumors. Front Immunol. 2022;13:1022136.
Wei Q, Dong Z. Mouse model of ischemic acute kidney injury: technical notes and tricks. Am J Physiol Renal Physiol. 2012;303:F1487–1494.
Martino MM, Maruyama K, Kuhn GA, Satoh T, Takeuchi O, Müller R, et al. Inhibition of IL-1R1/MyD88 signalling promotes mesenchymal stem cell-driven tissue regeneration. Nat Commun. 2016;7:11051.
Shin NS, Marlier A, Xu L, Doilicho N, Linberg D, Guo J, et al. Arginase-1 is required for macrophage-mediated renal tubule regeneration. J Am Soc Nephrol. 2022;33:1077–86.
Melchinger I, Guo K, Li X, Guo J, Cantley LG, Xu L. VCAM-1 mediates proximal tubule-immune cell cross talk in failed tubule recovery during AKI-to-CKD transition. Am J Physiol Renal Physiol. 2024;327:F610–f622.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Xu L, Guo J, Moledina DG, Cantley LG. Immune-mediated tubule atrophy promotes acute kidney injury to chronic kidney disease transition. Nat Commun. 2022;13:4892.
Hao Y, Stuart T, Kowalski MH, Choudhary S, Hoffman P, Hartman A, et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. 2024;42:293–304.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–337.e324.
Isola JVV, Ocañas SR, Hubbart CR, Ko S, Mondal SA, Hense JD, et al. A single-cell atlas of the aging mouse ovary. Nat Aging. 2024;4:145–62.
Jin S, Plikus MV, Nie Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat Protoc. 2025;20:180–219.
Lee K, Gharaie S, Kurzhagen JT, Newman-Rivera AM, Arend LJ, Noel S, et al. Double-negative T cells have a reparative role after experimental severe ischemic acute kidney injury. Am J Physiol Renal Physiol. 2024;326:F942–f956.
Xu L, Sharkey D, Cantley LG. Tubular GM-CSF promotes late MCP-1/CCR2-mediated fibrosis and inflammation after ischemia/reperfusion injury. J Am Soc Nephrol. 2019;30:1825–40.
Darash-Yahana M, Gillespie JW, Hewitt SM, Chen YY, Maeda S, Stein I, et al. The chemokine CXCL16 and its receptor, CXCR6, as markers and promoters of inflammation-associated cancers. PLoS ONE. 2009;4:e6695.
Medzhitov R, Preston-Hurlburt P, Kopp E, Stadlen A, Chen C, Ghosh S, et al. MyD88 is an adaptor protein in the hToll/IL-1 receptor family signaling pathways. Mol Cell. 1998;2:253–8.
Kinsey GR, Sharma R, Huang L, Li L, Vergis AL, Ye H, et al. Regulatory T cells suppress innate immunity in kidney ischemia-reperfusion injury. J Am Soc Nephrol. 2009;20:1744–53.
Gandolfo MT, Jang HR, Bagnasco SM, Ko GJ, Agreda P, Satpute SR, et al. Foxp3+ regulatory T cells participate in repair of ischemic acute kidney injury. Kidney Int. 2009;76:717–29.
Zhu Y, Cui H, Xia Y, Gan H. RIPK3-mediated necroptosis and apoptosis contributes to renal tubular cell progressive loss and chronic kidney disease progression in rats. PLoS ONE. 2016;11:e0156729.
Imamura M, Moon JS, Chung KP, Nakahira K, Muthukumar T, Shingarev R, et al. RIPK3 promotes kidney fibrosis via AKT-dependent ATP citrate lyase. JCI Insight. 2018;3:e94979.
Sanz AB, Santamaria B, Ruiz-Ortega M, Egido J, Ortiz A. Mechanisms of renal apoptosis in health and disease. J Am Soc Nephrol. 2008;19:1634–42.
Gerhardt LMS, Liu J, Koppitch K, Cippà PE, McMahon AP. Single-nuclear transcriptomics reveals diversity of proximal tubule cell states in a dynamic response to acute kidney injury. Proc Natl Acad Sci USA. 2021;118:e2026684118.
M. Bader S, Cooney JP, Bhandari R, Mackiewicz L, Dayton M, Sheerin D, et al. Necroptosis does not drive disease pathogenesis in a mouse infective model of SARS-CoV-2 in vivo. Cell Death Dis. 2024;15:100.
Belavgeni A, Meyer C, Stumpf J, Hugo C, Linkermann A. Ferroptosis and necroptosis in the kidney. Cell Chem Biol. 2020;27:448–62.
Guerrero-Mauvecin J, Villar-Gómez N, Rayego-Mateos S, Ramos AM, Ruiz-Ortega M, Ortiz A, et al. Regulated necrosis role in inflammation and repair in acute kidney injury. Front Immunol. 2023;14:1324996.
Choi ME, Price DR, Ryter SW, Choi AMK. Necroptosis: a crucial pathogenic mediator of human disease. JCI Insight. 2019;4:e128834.
Ricardo SD, van Goor H, Eddy AA. Macrophage diversity in renal injury and repair. J Clin Investig. 2008;118:3522–30.
Liang Z, Tang Z, Zhu C, Li F, Chen S, Han X, et al. Intestinal CXCR6(+) ILC3s migrate to the kidney and exacerbate renal fibrosis via IL-23 receptor signaling enhanced by PD-1 expression. Immunity. 2024;57:1306–1323.e1308.
Wu Y, An C, Jin X, Hu Z, Wang Y. Disruption of CXCR6 ameliorates kidney inflammation and fibrosis in deoxycorticosterone acetate/salt hypertension. Sci Rep. 2020;10:133.
Acknowledgements
We are grateful to Dr. Ruslan Medzhitov from the Department of Immunobiology at the Yale School of Medicine for the generous gift of Myd88−/−;Trif−/− mice. We thank technician Hongmei Shi (Department of Internal Medicine/Section of Nephrology, Yale School of Medicine) for performing mouse surgery and preparing kidney frozen sections.
Funding
This work was supported by National Institutes of Health Grant R01 DK135689 (to LX) and S10 OD023598 (to the Yale Center for Advanced Light Microscopy Facility). LGC was supported by R01 DK093771. IM was supported by a National Institutes of Health Grant R25 DK121566 (to Dr. Shuta Ishibe from the Department of Internal Medicine/Section of Nephrology at the Yale School of Medicine).
Author information
Authors and Affiliations
Contributions
LX conceptualized and designed the studies. XL, IM, YC, JG, and LX performed experiments. XL, IM, YC, and LX analyzed the data. LX performed in silico RNA-seq analysis. LX supervised the overall project and wrote the original manuscript draft. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Dr Andre Samson
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X., Melchinger, I., Chen, Y. et al. CXCR6+ T cells promote apoptosis and necroptosis in proximal tubules during AKI-to-CKD transition. Cell Death Dis 17, 359 (2026). https://doi.org/10.1038/s41419-026-08644-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-026-08644-x