Abstract
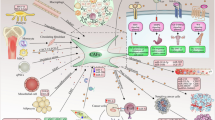
Adenoid cystic carcinoma (ACC) is a malignant epithelial neoplasm that arises in secretory glands and commonly metastasizes to the lungs, remaining the primary cause of patient mortality. Here, we observed that S100A9, a calcium-binding protein, was significantly upregulated in highly metastatic ACC cells and markedly enriched in their derived exosomes. Single-cell RNA sequencing was employed to comprehensively map the tumor microenvironment (TME), revealing robust intercellular communication between epithelial cells and cancer-associated fibroblasts (CAFs). Mechanistically, S100A9-enriched exosomes reprogrammed normal fibroblasts, resulting in the acquisition of an activated CAF phenotype characterized by the upregulation of FAP and α-SMA expression, and enrichment of IL-17, TNF-α, and NF-κB signaling pathways. Functionally, these exosome-educated CAFs promoted the epithelial–mesenchymal transition in ACC cells and facilitated lung metastasis in vivo via an IL-17-dependent signaling axis. Overall, these findings establish exosomal S100A9 as a crucial mediator of TME reprogramming, suggesting that targeting the S100A9–IL-17 axis may serve as a promising therapeutic strategy for disrupting ACC lung metastasis.

Similar content being viewed by others
Introduction
Adenoid cystic carcinoma (ACC) is a malignant epithelial neoplasm of glandular origin that predominantly occurs in the head and neck region. Salivary adenoid cystic carcinoma (SACC) represents the most common malignancy of the minor salivary glands and is among the most frequent cancers of the major salivary glands. Additionally, ACC involves the lacrimal and ceruminous glands, as well as other anatomical sites, including the nasal cavity, paranasal sinuses, trachea, and larynx [1, 2]. Despite the local control achieved through surgery and radiotherapy, distant metastasis, predominantly pulmonary metastasis, remains the primary cause of mortality and poor prognosis. Indeed, the 10-year survival rate for patients with distant metastases is approximately 29.0% [3]. Given the paucity of effective systemic therapies for refractory or metastatic ACC, largely due to an incomplete understanding of metastatic mechanisms [4], elucidating the biological drivers of ACC lung metastasis is imperative for developing novel therapeutic interventions.
Emerging evidence implicates the tumor microenvironment (TME) as a critical determinant of metastasis. Within the TME, cancer-associated fibroblasts (CAFs) constitute the dominant stromal component, contributing to tumorigenesis, metastasis, and therapy resistance by remodeling the extracellular matrix (ECM), promoting angiogenesis, secreting growth factors and cytokines, and suppressing anti-tumor immunity [5]. Although single-cell RNA sequencing (scRNA-seq) has begun to reveal the cellular heterogeneity of ACC [6], the precise intercellular crosstalk between ACC cells and stromal fibroblasts that promotes pulmonary metastasis remains poorly defined.
Intercellular communication within the TME is critically mediated by exosomes, which are small extracellular vesicles containing biomolecules such as proteins and nucleic acids [7]. Distinct from soluble factor-mediated signaling, exosomes facilitate an “exosomecrine” mechanism, allowing tumor cells to transfer bioactive cargo to both local stromal cells and distant organs, thereby protecting signaling molecules from degradation. While tumor-derived exosomes are known to modulate recipient cell phenotypes to establish a prometastatic environment [8, 9], the specific exosomal cargo that mediates the ACC–fibroblast interactions to promote lung metastasis remains largely unexplored.
S100A9, a proinflammatory calcium-binding protein, orchestrates premetastatic niche formation and therapeutic resistance in malignancies such as breast cancer and melanoma [10, 11]. Therapeutically, S100A9 inhibitors have been shown to attenuate fibrosis [12] and significantly prolong progression-free survival in patients with metastatic castration-resistant prostate cancer [13]. Furthermore, S100A9 can be packaged into exosomes to drive disease progression in hematological malignancies [14]. However, the contribution of exosomal S100A9 to ACC lung metastasis remains unclear. In this study, we integrated scRNA-seq analysis with in vitro and in vivo experiments to characterize the function of exosomal S100A9. We demonstrated that exosomal S100A9 reprogrammed fibroblasts to facilitate ACC lung metastasis, thereby identifying the S100A9–CAF axis as a potential therapeutic target.
Results
S100A9 is upregulated in ACC and associated with lung metastasis
To screen for genes involved in ACC lung metastasis, we performed mRNA microarray analysis to compare the gene expression profiles of SACC-83 and SACC-LM cell lines (Fig. 1A). To determine the prometastatic role of the exosomes, we isolated the corresponding exosomes (SACC-83-exo/SACC-LM-exo). Their characteristic cup-shaped morphology, size, and concentration were subsequently confirmed (Fig. S1A, B), and the expression of exosomal markers (HSP70, TSG101, CD63, and CD9) was validated by Western blotting (Fig. S1C). Proteomic profiling revealed 149 upregulated proteins and 398 downregulated proteins in SACC-LM-exo (fold change ≥2 or ≤0.5, p < 0.05) (Fig. 1B). Integration with the transcriptomic data revealed that S100A9 expression was consistently elevated in both tumor cells and exosomes, which was further validated by Western blotting (Fig. 1C). Gene Ontology (GO) analysis of SACC-83-exo and SACC-LM-exo revealed enrichment of vesicle-mediated transport, component biogenesis, and protein localization (Fig. 1D). Additionally, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed enrichment in cancer-related and ECM-receptor interaction pathways (Fig. 1E).
A Heatmap and hierarchical clustering of eleven selected mRNAs that were differentially expressed between SACC-LM cells and SACC-83 cells are shown. Red, high relative expression; green, low relative expression. B Heatmap and hierarchical clustering of ten selected proteins that were differentially expressed between SACC-83-exo and SACC-LM-exo. C Western blot analysis of the S100A9 expression level in SACC-83 cells, SACC-LM cells, and their corresponding exosomes, with β-actin and HSP70 serving as loading controls. Enrichment analysis of GO terms (D) and KEGG pathways (E) for differentially expressed proteins between SACC-83-exo and SACC-LM-exo based on the 4D mass spectrometry proteomics data. BP biological process, CC cellular component, MF molecular function. F Representative images of S100A9, FAP, and α-SMA expression and quantification in 56 human ACC tissue samples and 6 normal salivary gland tissue samples. Protein expression was quantified as an IHC score. The data points represent the individual patient samples scored. Scale bar = 50 μm. G Pearson correlation analysis of the FAP and α-SMA IHC scores with the S100A9 IHC score in all ACC tissues. H S100A9 expression in plasma exosomes derived from 9 salivary ACC patients with localized tumors and 5 with metastatic tumors. I Kaplan–Meier curves of overall survival (OS) and disease-free survival (DFS) in ACC patients after surgery stratified by S100A9 expression (cutoff: median). Data are presented as the mean ± SD. p values were calculated using Kruskal–Wallis test (F upper and middle), one-way ANOVA (F lower), Pearson correlation analysis (G), Mann–Whitney U test (H), and log-rank test (I). ns not significant; *p < 0.05, **p < 0.01, ***p < 0.001.
IHC analysis displayed elevated expression of S100A9, fibroblast activation protein (FAP), and α-smooth muscle actin (α-SMA) in primary ACC tumor tissues compared with normal salivary glands, with notably higher expression in patients with lung metastasis (Fig. 1F). Correlation analysis indicated that S100A9 expression was positively associated with FAP and α-SMA expression in 56 ACC samples (Fig. 1G), suggesting a potential role for S100A9 in tumor–CAF cross-talk. Clinically, high S100A9 expression correlated with advanced disease stage and lung metastasis (Table 1). To investigate the role of circulating S100A9 in plasma exosomes, we collected plasma samples from a cohort of 14 salivary ACC patients, comprising 9 with localized tumors and 5 with metastatic tumors. Plasma-derived exosomes were isolated and subsequently verified by TEM and Western blot analysis (Fig. S1D, E). We then measured S100A9 expression in these isolated exosomes and observed higher levels in patients with metastatic tumors than in those with localized disease (Fig. 1H). These findings suggest that circulating exosomal S100A9 may serve as a potential biomarker for predicting lung metastasis in ACC. We observed no significant difference in overall survival (OS), likely attributable to the indolent nature of ACC and the limited sample size. However, patients with high S100A9 expression exhibited significantly shorter disease-free survival (DFS) than those with low expression (Fig. 1I).
Single-cell analysis reveals CAF diversity and potential crosstalk with ACC cells in the tumor microenvironment
To comprehensively delineate the cellular landscape and investigate tumor–stroma interactions within the ACC tumor microenvironment, we performed scRNA-seq on five ACC tumor samples and two normal salivary gland samples (Fig. 2A). After stringent quality control, a total of 67,007 cells were retained and subjected to unsupervised clustering. This process resulted in the identification of 17 distinct clusters (Fig. S2A), which were subsequently annotated into 8 major cell types on the basis of the expression of canonical marker genes (Figs. 2B and S2B): epithelial cells, fibroblasts, smooth muscle cells, pericytes, NK/T cells, myeloid cells, endothelial cells, and B/plasma cells. The ACC tumors exhibited cellular heterogeneity, with a diverse cellular composition that varied between normal salivary glands and ACC tumors (Fig. 2B).
A UMAP visualization of 67,007 cells from 5 ACC tumors (ACC 1, ACC 3, ACC 6, ACC 7, and ACC 8) and 2 normal salivary glands (Norm 1 and Norm 2), colored by sample origin. B UMAP plot showing 8 major cell types annotated based on canonical marker genes (left). Stacked bar chart showing the normalized cell proportion of each cell type in tumor (n = 5) versus normal (n = 2) samples (right). C UMAP plot of 27,663 reclustered epithelial cells, colored by Seurat clusters (0–14). D InferCNV heatmap of epithelial cells, with normal epithelial cells as reference (top), revealing distinct CNA patterns (gains in red, losses in blue) in malignant cells. E UMAP plot of epithelial cells showing the calculated EMT score, with higher scores (red) indicating an EMT-like phenotype. F UMAP plot of epithelial cells annotated into functional subtypes based on canonical markers (Supplementary Table 1), CNA profiles (D), and EMT scores (E). G CellChat circle plots quantifying global intercellular communication. The left panel shows the total number of interactions, and the right panel shows the interaction strength between all major cell types. H UMAP plot of the reclustered fibroblast population, colored according to the Seurat cluster (0–8). I UMAP plot of the fibroblast population annotated into five functional subtypes (apCAFs, iCAFs, mCAFs, vCAFs, and nCAFs). J UMAP plot of the fibroblast population, highlighting the FAP⁺α-SMA⁺ CAF subpopulation (dark blue) against other CAFs (light blue). K Focused CellChat analysis showing the interaction number (left) and strength (right) between grouped epithelial subtypes (EMT-state vs. other) and fibroblast subtypes (FAP⁺α-SMA⁺ vs. other), highlighting a strong connection between “EMT-state epithelial cells” and “FAP+α-SMA+ CAFs”.
A total of 27,663 epithelial cells were isolated and subjected to reclustering analysis, resulting in 15 subclusters (Figs. 2C and S2C). To distinguish malignant cells from their normal counterparts, we inferred chromosomal copy number alterations. This analysis identified malignant cell clusters characterized by significant large-scale genomic gains and losses (Fig. 2D), allowing for the deconvolution of malignant and nonmalignant epithelial cells within the UMAP (Fig. S2D). Within this malignant population, we calculated enrichment scores for the hallmark epithelial–mesenchymal transition (EMT) pathway (Figs. 2E and S2E). Cells exhibiting high EMT scores were annotated as a distinct “EMT-state epithelial cell” functional subtype (Fig. 2F).
To map the intercellular communication networks, we performed CellChat analysis on the entire dataset. This revealed frequent and robust signaling interactions between the identified epithelial cells and fibroblasts (Fig. 2G). Given the pivotal role of fibroblasts in EMT-driven tumor metastasis, we performed a detailed subclustering of the fibroblast population. This analysis identified 9 distinct fibroblast subclusters (Figs. 2H and S2F), which were subsequently classified into five functional subtypes on the basis of their marker gene expression: antigen-presenting CAFs (apCAFs), matrix CAFs (mCAFs), inflammatory CAFs (iCAFs), vascular CAFs (vCAFs), and nonspecific CAFs (nCAFs) (Fig. 2I). Feature plot and violin plot analyses confirmed that while the key CAF activation markers FAP and α-SMA were expressed across several CAF subsets and that their distributions were not entirely consistent, their expression was predominantly concentrated and markedly higher within the mCAF subpopulation (Figs. 2J and S2G, H).
A subsequent focused CellChat analysis demonstrated that CAFs coexpressing FAP and α-SMA (FAP⁺α-SMA⁺ CAFs), which largely comprised the mCAF subpopulation, displayed the strongest and most frequent interactions with “EMT-state epithelial cells” (Fig. 2K). This key finding, highlighting specific crosstalk between activated, matrix-remodeling fibroblasts and malignant cells undergoing EMT, was further validated in an independent, external scRNA-seq dataset (GSE216852). The external dataset analysis replicated our findings, identifying analogous cell subtypes (Fig. S3A–I) and confirming both the mCAF expression of FAP/α-SMA (Fig. S3J) and the specific, strong interaction between mCAFs and EMT-state epithelial cells (Fig. S3K). To visually validate this inferred crosstalk within the native tissue architecture, we performed multiplex immunohistochemistry (mIHC). This analysis of ACC tissues confirmed the intimate spatial proximity between FAP+ stromal populations (CAFs) and Pan-CK+ malignant epithelial cells, providing a morphological basis for the direct interactions inferred by our scRNA-seq analysis (Fig. S4).
Collectively, these findings highlight a robust and specific interaction network between FAP⁺α-SMA⁺ CAFs and malignant tumor cells in an EMT state, underscoring the importance of identifying key regulatory factors mediating this tumor–fibroblast crosstalk in ACC.
S100A9 in exosomes mediates fibroblast activation
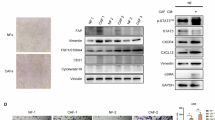
To assess the relationship between S100A9 and fibroblast activation, primary CAFs and matched NFs were isolated from ACC tumors and paired normal salivary glands (Fig. S5A). Compared with NFs, CAFs exhibited significantly higher expression of fibrosis markers (FAP and α-SMA), confirming their activated phenotype (Fig. 3A, C). Western blot analysis confirmed this difference in multiple samples (Fig. S5B). Exosome uptake was verified by PKH26 labeling and confocal microscopy (Fig. S5C). Consistent with the increased metastatic potential of the donor cells, compared with SACC-83-derived exosomes, SACC-LM-derived exosomes induced marked increases in FAP and α-SMA expression in NFs (Figs. 3A, C and S5B). To explore the underlying mechanism, we established S100A9 overexpression models in SACC-83 cells and NFs. The successful overexpression of S100A9 (in cell lysates, exosomes, and NFs) was confirmed by qPCR and Western blotting (Fig. S6A–D). Using these models, we found that compared with their respective controls, both S100A9-enriched exosomes (S100A9-exo) and the S100A9 overexpression plasmid significantly increased the expression of FAP and α-SMA in NFs (Figs. 3B, C and S5B). All three treatments (SACC-LM-exo, S100A9-exo, and S100A9 overexpression plasmids) significantly increased the contraction and migration abilities of NFs compared with those of the control groups (SACC-83-exo, Vector-exo, and control plasmid treatments, respectively) (Fig. 3D–G). Furthermore, S100A9-exo treatment increased the mRNA levels of fibrosis markers, proinflammatory cytokines (IL-6, IL-8, and IL-1β), and matrix metalloproteinases (MMP2 and MMP9) (Fig. 3H). These molecules are known to contribute to a tumor-promoting microenvironment.
A Representative IF images and quantification of the mean fluorescence intensity (MFI) of α-SMA and FAP in NFs, CAFs, and NFs treated with different types of exosomes (SACC-83-exo and SACC-LM-exo). Scale bar = 50 μm. B Representative IF images and MFI quantification of α-SMA and FAP in NFs treated with different types of exosomes (Vector-exo and S100A9-exo) or transfected with plasmids (NFVector and NFS100A9). Scale bar = 50 μm. C Western blot analysis of α-SMA and FAP expression in NFs treated as indicated in (A) and (B). D Representative images and quantification of collagen contraction in NFs, CAFs, and NFs treated with exosomes as in (A). Scale bar = 2 mm. E Migration of NFs, CAFs, and NFs treated with exosomes as in A was assessed by Transwell assay. Scale bar = 50 μm. F Representative images and quantification of collagen contraction in NFs treated with different exosomes and plasmids as in (B). Scale bar = 2 mm. G Migration of NFs treated with different exosomes and plasmids as in B was assessed by Transwell assay. Scale bar = 50 μm. H Relative mRNA levels of IL-6, IL-8, IL-1β, MMP2, and MMP9 in NFs treated with PBS (Control), Vector-exo, or S100A9-exo. I Representative IF images and MFI quantification of α-SMA and FAP in NFs incubated with exosomes from SACC-LM cells treated with siControl or siS100A9 (two independent siRNAs, −2 and −3). Scale bar = 50 μm. J Collagen contraction ability of NFs treated as in (I). Scale bar = 2 mm. K Representative IF images and MFI quantification of α-SMA and FAP in S100A9-exo-treated NFs with or without the inhibitor paquinimod. Scale bar = 50 μm. L Collagen contraction ability of NFs treated as in (K). Scale bar = 2 mm. Data are presented as the mean ± SD. p values were calculated using two-tailed Student’s t test for two-group comparisons and one-way ANOVA for multi-group comparisons. *p < 0.05, **p < 0.01, ***p < 0.001.
We next examined the impact of S100A9 loss. After knocking down S100A9 in SACC-LM cells using siRNA (Fig. S6E, F), we observed that exosomes derived from these cells had a significantly reduced capacity to activate fibroblasts, as evidenced by a reduction in FAP and α-SMA expression, and a decrease in the capacity for collagen contraction (Fig. 3I, J). Similarly, the S100A9 inhibitor paquinimod suppressed fibroblast activation induced by S100A9-exo (Fig. 3K, L). Collectively, these findings establish a potential role for S100A9-enriched exosomes in promoting fibroblast activation and stromal remodeling.
Exosomal S100A9 potentiates lung metastasis and activates fibroblasts via the IL-17–TNF–NF-κB axis
To evaluate the contribution of S100A9-exo to organotropic metastasis, a lung premetastatic niche model was established in vivo (Fig. 4A). Tail vein injection of S100A9-exo into NOD/SCID mice resulted in elevated expression of fibroblast activation markers (FAP and α-SMA) in lung tissue (Fig. 4B). Elevated levels of fibronectin, MMP9, and VEGFR1 were observed in lung tissue from S100A9-exo-injected mice, suggesting the establishment of a premetastatic microenvironment (Fig. 4C). Crucially, tail vein injection of S100A9-exo significantly increased ACC lung metastasis in immunodeficient mice (Fig. 4D, E). NFs were treated with Vector-exo or S100A9-exo, harvested, and subjected to high-throughput RNA sequencing to investigate the downstream molecular events. The results demonstrated significant activation of the IL-17, TNF, and NF-κB signaling pathways, along with enrichment in chemokine receptor binding (Fig. 4F, G). Previous studies have consistently revealed that NF-κB activation is a crucial regulatory node in the IL-17 signaling cascade and acts synergistically with TNF-α [15, 16]. Treatment with S100A9-exo significantly upregulated the mRNA expression levels of IL-17 and TNF-α in these educated NFs (Fig. 4H). Furthermore, the IKK inhibitor BAY-117082 effectively blocked NF-κB activation (Fig. 4I) and consequently abolished the effect of S100A9-exo on fibroblast activation and inflammatory cytokine expression (Fig. 4J). Metabolomic analysis further highlighted metabolic reprogramming in the S100A9-exo group, characterized by enhanced nucleotide metabolism (purine and pyrimidine) and lipid metabolism (sphingolipid and glycerophospholipid) (Fig. 4K). Collectively, these results establish that S100A9-exo stimulates the production of IL-17 and TNF-α, thereby inducing NF-κB signaling and promoting fibroblast activation, which may facilitate the formation of a prometastatic microenvironment.
A Schematic of the two independent in vivo experimental models. Top (premetastatic niche model): NOD/SCID mice (n = 8 per group) were intravenously injected with exosomes daily for 7 days. Lung tissues were collected on day 7 for analysis. Bottom (metastasis model): NOD/SCID mice (n = 7 per group) were pretreated with daily exosome injections for 7 days, followed by intravenous injection of SACC-83 cells on day 8. The lungs were collected for analysis 8 weeks later. B IHC analysis of FAP and α-SMA in lung tissues from NOD/SCID mice following 7 days of exosome stimulation (n = 8 per group). Scale bar = 200 μm. C IF staining of fibronectin, MMP9, and VEGFR1 in serial sections of the same cohort (n = 8 per group). Scale bar = 200 μm. D Formation of lung metastases in mice after stimulation with Vector-exo or S100A9-exo, as represented by bioluminescence signals (left) and normalized photon flux (right) (n = 7 per group). E Representative images (left) and quantification of metastatic tumor nodules (right) in lung sections from NOD/SCID mice. Scale bar = 100 μm. (n = 7 per group). F GO annotation of differentially expressed mRNAs with the top 20 enrichment factors. G KEGG pathway enrichment analysis of the differentially expressed mRNAs by RNA sequencing. H Relative IL-17 and TNF-α mRNA levels in fibroblasts after incubation with corresponding exosomes. I Western blot analysis of levels of cytoplasmic p65 and p-p65 and nuclear p-p65 in NFs following treatment with Vector-exo or S100A9-exo in the presence or absence of BAY-117082. β-actin and histone H3 were used as loading controls for cytoplasmic and nuclear fractions, respectively. J Quantification of the mRNA expression of FAP, α-SMA, inflammatory markers, and cytokines in NFs treated with Vector-exo or S100A9-exo, with or without BAY-117082. K Metabolomic analysis showing altered pathways in S100A9-exo-treated NFs. Data are presented as the mean ± SD for (H, J) and as the mean ± SEM for (B right, C right, D right). p values were calculated using two-tailed Student’s t test for (B right, C right, D right, H, J), and Mann–Whitney U test for (E right). **p < 0.01, ***p < 0.001.
Activated fibroblasts promote ACC cell epithelial–mesenchymal transition and lung metastasis through the secretion of IL-17
To assess the impact of S100A9-exo-activated fibroblasts on SACC-83 cell malignancy, we established an indirect co-culture system. Following 48 h of exosome treatment, activated fibroblasts were seeded into the upper chamber of the Transwell insert (0.4 μm), while SACC-83 cells were simultaneously seeded into the lower chamber (Fig. 5A). Following co-culture, the “educated” SACC-83 cells were harvested and subjected to functional assays. These cells exhibited increased migration, invasion, viability, proliferation, and colony formation abilities (Fig. 5B–E). Moreover, co-culture with S100A9-exo-pretreated fibroblasts induced an EMT phenotype in SACC-83 cells, characterized by the downregulation of E-cadherin and upregulation of N-cadherin (Fig. 5F). In vivo, injection of these co-cultured SACC-83 cells significantly enhanced the metastatic potential of SACC-83 cells in a mouse model (Fig. 5G, H). Collectively, results from both the direct S100A9-exo treatment (Fig. 4D, E) and the fibroblast co-culture model (Fig. 5G, H) consistently demonstrate that exosomal S100A9 promotes ACC lung metastasis, underscoring its pivotal role in metastatic progression.
A Schematic diagram of exosome-modulated fibroblasts regulating tumor cells and generating a feedback loop for adjacent tumor cells. The fibroblasts were pretreated with Vector-exo or S100A9-exo for 48 h; these activated fibroblasts were subsequently co-cultured with SACC-83 cells for follow-up experiments. B CCK-8 assay of SACC-83 cells co-cultured with pretreated fibroblasts was performed every 24 h. C Migration and invasion of these co-cultured samples were assessed by the Transwell assay. Scale bar = 200 μm. D Representative images and quantification of colony number in colony formation assays after 10 days of co-culture. E Representative fluorescence images and quantification displaying EdU incorporation (green) in the co-cultured samples. Nuclei were counterstained with DAPI (blue). Scale bar = 100 µm. F Western blot analysis of E-cadherin and N-cadherin expression in co-cultured SACC-83 cells, with GAPDH used as a loading control. G Formation of lung metastases in mice after injection of SACC-83 cells harvested from co-culture with exosomes/fibroblasts, as represented by bioluminescence signals (left) and normalized photon flux (right) (n = 7 per group). H Representative images (left) and quantification of metastatic tumor nodules (right) in H&E-stained lung sections from NOD/SCID mice (n = 7 per group). Scale bar = 100 μm. GO (I) and KEGG (J) enrichment analyses of differentially expressed genes in SACC-83 cells co-cultured with the indicated fibroblasts, determined by RNA sequencing. K ELISA quantification of IL-17 protein levels in conditioned medium (CM) from NFs treated with Vector-exo (Vector-exo-NF-CM) or S100A9-exo (S100A9-exo-NF-CM). Control IgG or an anti-IL-17 neutralizing antibody (IL-17 Ab) was added to S100A9-exo-NF-CM as indicated. L Western blot analysis of N-cadherin, E-cadherin, MMP9, and MMP2 expression in SACC-83 cells. The cells were co-cultured with S100A9-exo-activated fibroblasts, with either control IgG or an IL-17 neutralizing antibody added to the co-culture medium. M Western blot analysis of EMT markers in SACC-83 cells treated with increasing concentrations of rhIL-17A. Data are presented as the mean ± SD for (B, C right, D, E right, K) and as the mean ± SEM for (G right). p values were calculated using two-way ANOVA for (B), two-tailed Student’s t test for (C–E, G right, K), and Mann–Whitney U test for (H right). *p < 0.05, **p < 0.01, ***p < 0.001.
To identify the molecular mediators driving tumor–fibroblast interactions, we performed RNA sequencing on SACC-83 cells co-cultured with Vector-exo- or S100A9-exo-treated fibroblasts. GO and KEGG pathway enrichment analyses revealed that the IL-17 signaling pathway was the most significantly enriched pathway in the S100A9-exo group (Fig. 5I, J). ELISA analysis confirmed elevated IL-17 levels in the conditioned medium of S100A9-exo-treated NFs, which were effectively depleted by an anti-IL-17 neutralizing antibody (IL-17 Ab) (Fig. 5K). Crucially, Western blot analysis revealed that blocking IL-17 with the neutralizing antibody reversed the EMT phenotype induced by the activated fibroblasts, as evidenced by the restoration of E-cadherin expression and the suppression of N-cadherin, MMP2, and MMP9 (Fig. 5L). Furthermore, direct treatment with recombinant human IL-17A (rhIL-17A) significantly induced EMT in SACC-83 cells in a dose-dependent manner (Fig. 5M). These results indicate that S100A9-exo-activated fibroblasts regulate SACC-83 cell epithelial–mesenchymal transition and lung metastasis through the secretion of IL-17.
Discussion
Despite significant therapeutic advances in ACC, the long-term prognosis for patients remains unsatisfactory because of an insufficient understanding of the underlying mechanisms of lung metastasis [17]. The interplay between malignant tumor cells and their TME plays a critical role in promoting tumor progression [18] and contributing to drug resistance [19]. This crosstalk is driven by cell-to-cell interactions, soluble factors (cytokines and chemokines), or exosomes [20]. In ACC, exosomes have been previously shown to promote premetastatic niche formation by differentially regulating pulmonary microvascular endothelial cell functions [8, 9, 21].
Our proteomic analysis revealed that exosomes derived from different ACC cell lines display distinct proteomic profiles, which aligns with increasing evidence that exosomal properties are dynamic and reflect a tumor’s aggressive phenotype [22]. Proteins that were differentially expressed in SACC-LM-exo were significantly associated with “pathways in cancer” and “extracellular matrix organization”. We focused on proteins consistently overexpressed in both the aggressive cell line and its exosomes, suggesting that these proteins exert effects both intracellularly and extracellularly via exosomal secretion. Through this integrated analysis, we identified an upregulation of S100A9. Although S100A9 typically functions as a heterodimer with S100A8 (calprotectin), it is also known to promote tumor development through independent mechanisms [23]. As S100A8 levels remained unchanged, we prioritized S100A9 for further investigation, particularly given its well-established role in chronic inflammation and tumor promotion [24]. Clinically, we confirmed its relevance, observing that elevated S100A9 levels correlated with frequent lung metastasis and advanced clinical stages.
Exosomal S100A9 is involved in tumor progression, such as by orchestrating NF-κB pathway activation in chronic lymphocytic leukemia [14] and enhancing the stemness of colorectal cancer (CRC) cells [25]. In addition, S100A9 has shown potential as a biomarker for tumor diagnosis and prognosis in acute myeloid leukemia and CRC [26]. Our in vivo results align with these findings, showing that S100A9-overexpressing cell-derived exosomes significantly increased ACC lung metastasis in an “exosomecrine” manner. Recent single-cell analyses have suggested that characterizing the ACC stromal microenvironment is key to understanding its biology and that the most prevalent cell population in the ACC stroma is fibroblasts [27, 28]. Our scRNA-seq analysis revealed strong interactions between EMT-state epithelial cells and CAFs. CAFs are known to remodel the ECM, induce metabolic reprogramming, and promote invasion and metastasis [29]. For example, in pancreatic ductal adenocarcinoma (PDAC), CAFs that have undergone fibroblast-to-myofibroblast transition can form metastatic clusters and “hijack” cancer cells to distant organs [30]. Given that tumor cells can reprogram CAFs and that S100A9 is known to modulate myeloid cells [31], we hypothesized that exosomal S100A9 cargo might be a key factor in fibroblast reprogramming.
Validating this hypothesis, our investigation revealed that both SACC-LM-exo and S100A9-exo are capable of activating NFs toward a CAF-like phenotype, characterized by high expression of fibrogenic genes (FAP and α-SMA) and enhanced contractility. To confirm that S100A9 itself was the critical mediator of this reprogramming, we utilized genetic and pharmacological approaches. Corroborating our genetic knockdown findings, inhibition of S100A9 using paquinimod significantly diminished this S100A9-exo-induced CAF activation profile. These findings confirm that exosomal S100A9 is indeed the key functional cargo driving fibroblast activation.
To explore the mechanism by which S100A9-enriched exosomes activate fibroblasts, transcriptomic profiling of S100A9-exo-treated NFs revealed significant enrichment of the IL-17, TNF-α, and NF-κB signaling pathways. While NF-κB-driven CAF activation is an established mechanism [32], current observations suggest that it may be downstream of cytokine signaling. IL-17 is a key inflammatory mediator [33] and is known to facilitate fibroblast differentiation [34], with NF-κB serving as a critical player in its signaling cascade [35]. Furthermore, TNF-α and IL-17 are known to cooperate [16, 36], with TNF-α inducing transcription and IL-17 stabilizing mRNAs [37, 38]. These findings align with the literature, supporting a model in which S100A9-enriched exosomes first act as an upstream trigger to stimulate IL-17 and TNF-α expression. These two cytokines, in turn, cooperatively converge to promote enhanced NF-κB activity, leading to the subsequent activation of matrix CAFs in ACC.
We subsequently observed that co-culturing SACC-83 cells with S100A9-exo/fibroblast enhanced the EMT phenotype and invasive capabilities of the SACC-83 cells. Notably, KEGG pathway analysis of these tumor cells also revealed the involvement of the IL-17 signaling pathway, implicating that IL-17 is a key factor in this feedback loop. These findings align with those of studies showing that IL-17 secreted from TME can facilitate tumor progression [39, 40]. Validation experiments confirmed this role, as recombinant IL-17A alone promoted EMT in ACC tumor cells, whereas an anti-IL-17 neutralizing antibody reversed the CAF-induced effects. Thus, disrupting this vicious cycle involving S100A9–CAF–IL-17 may help reshape the TME and suppress tumor progression.
While our study highlights the significance of exosomal S100A9 in ACC progression, further validation in larger, multicenter clinical cohorts is warranted to reinforce the statistical robustness of our clinical and scRNA-seq findings. Additionally, although our tail vein injection models effectively recapitulate the colonization phase of metastasis, the development of stable spontaneous metastasis models remains an essential future goal to fully capture the entire metastatic cascade. Nevertheless, our data provide solid evidence identifying exosomal S100A9 as a central mediator of ACC tumor microenvironment remodeling.
In summary, our results suggest a positive feedback loop in the ACC TME (Graphical Abstract). Tumor-derived exosomal S100A9 induces the transformation of NFs to activated CAFs, which reshape the TME. In turn, these activated CAFs promote the EMT and metastasis of ACC cells through the secretion of IL-17. This vicious cycle within the TME ultimately perpetuates the progression of ACC. Our findings indicate that targeting the S100A9–IL-17 axis may offer a promising therapeutic strategy for ACC lung metastasis.
Materials and methods
Tissue samples and cell lines
ACC specimens (n = 56) and normal salivary gland (NG) tissues (n = 6) were obtained from patients undergoing radical resection at the Department of Oral and Maxillofacial Surgery, Hospital of Stomatology, Sun Yat-sen University (2015–2024). No patients received preoperative chemotherapy or radiotherapy, and all diagnoses were histopathologically confirmed.
The SACC-83 cell line was established from sublingual glandular ACC, while the highly metastatic SACC-LM cell line was isolated in vivo following tail vein injection of SACC-83 cells into immunodeficient mice [41]. Both cell lines were authenticated by STR profiling, confirmed to be mycoplasma-free by PCR, and used within 15 passages post-thaw. ACC cells were cultured in RPMI-1640 medium with 10% fetal bovine serum (FBS) (Gibco, Grand Island, NY, USA).
Primary normal fibroblasts (NFs) and cancer-associated fibroblasts (CAFs) were isolated from paired normal gland and ACC tumor tissues, respectively, following established protocols [42]. Fibroblasts were maintained in DMEM/F12 supplemented with 10% FBS, 100 U/mL penicillin, and 100 µg/mL streptomycin (Gibco) at 37 °C in 5% CO₂. Conditioned media from exosome-treated NFs are designated S100A9-exo-NF-CM or Vector-exo-NF-CM.
Microarray expression profiling and mRNA high-throughput sequencing
Gene expression profiling of SACC-83 and SACC-LM cells was performed using the SBC Human 4 × 180 K ceRNA microarray (Shanghai Biotechnology Corporation, Shanghai, China). For high-throughput sequencing, total RNA was isolated from Vector-exo-NF, S100A9-exo-NF, and SACC-83 cells co-cultured with exosome-treated fibroblasts. Sequencing was conducted on the Illumina platform by Shanghai Biotechnology Corporation.
Exosome proteomic mass spectrometry
Exosome proteomic sequencing services were provided by Wayen Biotechnologies (Shanghai, China). Peptides were separated using an EASY-nLC 1200 system (Thermo Scientific, Waltham, MA, USA) equipped with a C18 analytical column (1.9 µm, 75 µm × 20 cm) at a flow rate of 200 nL/min. Mass spectrometry analysis was performed on an Orbitrap Exploris 480 mass spectrometer coupled with a FAIMS Pro interface (Thermo Scientific) operating in Data Dependent Acquisition (DDA) mode. The full scan resolution was set to 60,000 (FWHM) with a scan range of m/z 350–1600. Precursor ions were fragmented via Higher-energy Collisional Dissociation (HCD) with a normalized collision energy of 30%. Raw data were processed using Proteome Discoverer 2.4 software (Sequest HT, Thermo Scientific). Downstream functional annotation and pathway enrichment analyses were conducted using Metascape (http://metascape.org) and the clusterProfiler R package. Terms with a p value < 0.05, minimum count of 3, and enrichment factor >1.5 were identified and grouped into clusters on the basis of their membership similarity.
Single-cell RNA sequencing
Sample collection
Tumor specimens were obtained from five patients with a preoperative diagnosis of ACC, which was subsequently confirmed by postoperative histopathological examination. Normal control salivary gland tissues comprising one parotid gland and one submandibular gland were collected from two individuals without ACC. Immediately following surgical resection, fresh tissue samples were preserved in tissue storage solution (Miltenyi Biotec, Bergisch Gladbach, Germany) on ice and transferred to the laboratory.
Sample preparation
Single-cell suspensions were prepared immediately from fresh tissues. Briefly, tissues were washed by DPBS (calcium- and magnesium-free PBS) and minced into small pieces. Enzymatic digestion was performed using collagenase I (2 mg/mL), collagenase II (2 mg/mL), collagenase IV (2 mg/mL), and dispase (0.2 mg/mL) at 37 °C for 15–25 min. The cell suspension was filtered through a 40-μm nylon cell strainer into a centrifuge tube. The filtrate was centrifuged at 500 × g for 5 min, and the supernatant was carefully discarded. The cell pellet was resuspended in RBC lysis buffer and incubated on ice for 5 min to remove red blood cells. The cells were then washed twice with DPBS by centrifugation at 400 × g for 5 min. The cell concentration and viability were assessed using a Cellometer Auto 2000 (Nexcelom Bioscience, Lawrence, MA, USA) via AO/PI staining.
Library construction and sequencing
Single-cell suspensions were loaded onto a 10x Genomics Chromium Controller to generate gel-beads-in-emulsion (GEMs). Libraries were constructed using the Chromium Next GEM Single Cell 3’ Reagent Kit v3.1. Following cell lysis and reverse transcription within GEMs, barcoded full-length cDNA was synthesized from polyadenylated mRNA. Post-GEM-RT cleanup was performed using Dynabeads MyOne Silane to remove biochemical reagents and primers. The cDNA was then amplified, enzymatically fragmented, end-repaired, and A-tailed. Subsequently, adapters were ligated, and sample index PCR was performed following size selection. The final libraries were subsequently purified using SPRIselect beads and sequenced on an Illumina NovaSeq X Plus platform.
Data preprocessing and quality control
Raw BCL files were converted to FASTQ format, aligned, and quantified using the Cell Ranger pipeline (version 3.1.0, 10x Genomics). Downstream analysis was performed using the Seurat R package (v5.3.0) [43]. Low-quality cells were filtered on the basis of the following criteria: (1) >10% mitochondrial RNA content; (2) <300 detected genes; or (3) > 20,000 unique molecular identifiers (UMI) or >7000 detected genes. Potential doublets were identified and removed using DoubletFinder (version 2.0.4) [44]. Following quality control, 67,007 high-quality cells were retained and merged into a single Seurat object. Data were normalized and scaled, and highly variable features were identified using the FindVariableFeatures function. Principal component analysis (PCA) was performed on the scaled data (RunPCA). Batch effects were corrected using the Harmony algorithm (version 1.2.3) [45]. Nonlinear dimensional reduction was performed using UMAP (RunUMAP). Cell clusters were identified via shared nearest neighbor (SNN)-based clustering (FindNeighbors and FindClusters, resolution = 0.2) using the top 15 principal components (PCs), as determined by an elbow plot. Visualization was optimized using UMAP and the SCP R package (version 0.5.6).
Differentially expressed genes analysis
Differentially expressed genes (DEGs) in each cell cluster were identified using the FindAllMarkers function in Seurat via the Wilcoxon Rank Sum test. The significance criteria were set at an adjusted p value < 0.05 and a |log2 fold change (FC)| >0.25.
Annotation of cell clusters and sub-clusters
The 17 identified clusters were annotated based on DEGs and canonical cell type markers. Epithelial cells and CAFs were subsetted and reclustered for fine-resolution analysis. Subtypes were annotated on the basis of their unique DEGs and the expression of well-established canonical markers (Supplementary Table 1). Epithelial cells were classified into EMT-state, acinar, myoepithelial, nonspecific, fibroblast-like, proliferating, mucous acinar, and nonmalignant epithelial cells. CAFs were categorized into antigen-presenting CAFs (apCAFs), matrix CAFs (mCAFs), inflammatory CAFs (iCAFs), nonspecific CAFs (nCAFs), and vascular CAFs (vCAFs).
Copy number alterations (CNA) analysis
To distinguish malignant cells from nonmalignant cells, copy number alterations (CNAs) in epithelial cells were estimated using the infercnv R package (version 1.20.0). Endothelial cells served as the normal reference. Cells were classified on the basis of their inferred malignancy scores.
Computation of EMT score
EMT scores for epithelial clusters were calculated using the AddModuleScore function in Seurat. The gene signature was derived from the “HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION” gene set in the MSigDB database. Malignant epithelial cells exhibiting high EMT scores were defined as “EMT-state epithelial cells”.
Inference of cell–cell communication
To investigate the degree of crosstalk between epithelial cells and CAFs, the subpopulations were defined as follows: epithelial cells were categorized into “EMT-state” and “other” groups on the basis of EMT scores, and CAFs were stratified into FAP+/α-SMA+ (double-positive) and “other” CAFs on the basis of marker expression. Intercellular communication networks were then inferred using the CellChat R package (version 2.2.0) [46].
Overexpression and siRNAs
Lentiviral constructs, including HBLV-h-S100A9-3xflag-ZsGreen-PURO (for S100A9 overexpression), HBLV-ZsGreen-PURO (empty vector control), and HBLV-LUC-PURO, were obtained from Hanbio (Shanghai, China). Stable SACC-83 cells overexpressing S100A9 (S100A9 cells) and control cells (Vector cells) were established following antibiotic selection.
For knockdown experiments, SACC-LM cells were transiently transfected with small interfering RNAs (siRNAs) targeting S100A9 (siS100A9) or a scrambled negative control (siControl). All siRNAs were designed and synthesized by RiboBio (Guangzhou, China). Cells were seeded in six-well plates and transfected at 50%–60% confluence using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. Cells were harvested 48 h post-transfection. The sequences of all siRNAs used are listed in Supplementary Table 2.
Exosome isolation, labeling, and uptake
Exosomes were isolated from cell culture supernatants via ultracentrifugation combined with a sucrose cushion method [21]. The supernatants were clarified by centrifugation, concentrated using a filter (100 kDa, Millipore, Burlington, MA, USA), and ultracentrifuged at 100,000 × g for 70 min (Optima L-90K, Beckman Coulter, Brea, CA, USA). Exosomes were purified using a 30% sucrose/deuterium oxide (D2O) cushion. The exosome-enriched fraction was subsequently resuspended in PBS and stored at –80 °C. Exosomal protein concentrations were quantified using a Bradford protein assay kit (Beyotime, Shanghai, China).
For uptake assays, exosomes were labeled with PKH26 (Sigma-Aldrich, St. Louis, MO, USA). Fibroblasts were stained with Actin-Tracker Green (phalloidin-FITC, Beyotime), and nuclei were counterstained with DAPI (ZSGB-Bio, Beijing, China). Fibroblasts were incubated with labeled exosomes for 24 h prior to imaging using a confocal microscope (FV3000, Olympus, Tokyo, Japan).
Immunofluorescence
Fibroblasts seeded on glass coverslips were treated with exosomes for 48–72 h, fixed with 4% paraformaldehyde (PFA), and permeabilized with 0.1% Triton X-100. Formalin-fixed, paraffin-embedded (FFPE) sample sections were deparaffinized, rehydrated, and subjected to heat-induced antigen retrieval using citrate buffer. Following blocking with 5% bovine serum albumin (BSA) containing mouse IgG blocking reagent for 1 h at room temperature, the samples were incubated with primary antibodies (Supplementary Table 3) overnight at 4 °C. After being washed, the samples were incubated with fluorophore-conjugated secondary antibodies for 1 h in the dark. Nuclei were counterstained with DAPI (ZLI-9556, ZSGB-Bio). Images were acquired using a confocal microscope (FV3000, Olympus) or an inverted fluorescence microscope (IX83, Olympus).
RNA preparation and qPCR
Total RNA was extracted using TRIzol reagent (Invitrogen) following the manufacturer’s protocol. Subsequently, cDNA was synthesized using a cDNA Reverse Transcription Kit (Takara, Kusatsu, Japan). RT–qPCR was performed using SYBR Green assays on a LightCycler 96 system (Roche, Basel, Switzerland). Relative gene expression levels were calculated using the 2−ΔΔCt method and normalized to those of the endogenous control. Specific primer sequences are listed in Supplementary Table 4.
Colony formation, cell viability, and proliferation assays
For colony formation assays, cells were seeded into 6-well plates (500 cells/well) and cultured for 10 days. Colonies were fixed with 4% paraformaldehyde for 30 min and stained with 1% crystal violet (RBG1019, Roles-Bio, Guangzhou, China). Colony counts were quantified using ImageJ software.
Cell viability and proliferation were evaluated using the Cell Counting Kit-8 (CCK-8) (Dojindo Laboratories, Kumamoto, Japan) and 5-ethynyl-2’-deoxyuridine (EdU) incorporation assays (KGA9602-100, KeyGEN BioTECH, Nanjing, China), respectively, following the manufacturer’s instructions.
Western blot analysis
Total protein was extracted from cultured cells with RIPA lysis buffer (Beyotime). Protein concentrations were quantified using a BCA protein assay kit (Thermo Scientific). Proteins were separated by SDS–PAGE and transferred onto PVDF membranes. Immunoblotting was performed using specific antibodies listed in Supplementary Table 3. Protein bands were visualized using an enhanced chemiluminescence (ECL) kit (CWBio, Beijing, China).
Fibroblasts treated with the IKK inhibitor BAY-117082 (Sigma-Aldrich) were processed using Nuclear and Cytoplasmic Extraction Reagents (P0028, Beyotime) to isolate nuclear and cytoplasmic fractions. SACC-83 cells stimulated with recombinant human IL-17A (200-17, PeproTech, Cranbury, NJ, USA) were harvested for total protein extraction. Full and uncropped Western blots can be found in the Supplementary Information.
Transwell assays
Cell migration and invasion were assessed using Transwell inserts with 8-μm pores (Costar, Corning, NY, USA). For invasion assays, the upper surface of the membrane was precoated with Matrigel (BD Biosciences, San Jose, CA, USA). Following 48 h of treatment, cells were harvested and seeded at a density of 7 × 104 cells in 200 μL of serum-free RPMI-1640 medium into the upper chamber. The lower chamber was filled with RPMI-1640 containing 10% FBS as a chemoattractant. After incubation for 16–19 h, non-invading cells on the upper surface were removed using cotton swabs. Invading cells attached to the lower surface were fixed with 95% ethanol and stained with 1% crystal violet. Stained cells were imaged and quantified using a digital microscope (Nikon, Tokyo, Japan).
Migration assays were performed following the same protocol but without Matrigel coating.
Collagen contraction assays
Following treatment with exosomes or plasmids for 48–72 h, fibroblasts were harvested and resuspended in 400 μL of a collagen mixture comprising 186.46 µL dH2O, 4.23 µL 1 N NaOH, 40 µL 10× PBS, and 169.31 µL Type I Rat Tail Collagen (354249, Corning). Afterwards, the mixture was seeded into a 24-well plate and incubated at 37 °C for 30 min to allow polymerization. Subsequently, 500 µL DMEM-F12 supplemented with 10% FBS was added on top of the gels. After incubation for 16 h, the gels were imaged, and the gel surface area was quantified using ImageJ software.
ELISA
For plasma analysis, heparinized blood samples were collected from ACC patients with (n = 9) or without (n = 5) lung metastasis. Plasma exosomes were isolated using the exoRNeasy Serum/Plasma Maxi Kit (Qiagen, Hilden, Germany). Exosomal S100A9 levels were quantified using a Human S100A9 ELISA Kit (Elabscience, Wuhan, China).
For fibroblast analysis, IL-17A concentrations in cell culture supernatants were measured using a Human IL-17A ELISA Kit (H014-2, Nanjing Jiancheng Bioengineering Institute, Jiangsu, China) following the manufacturer’s protocol. For neutralization experiments, fibroblast-conditioned medium was incubated with 10 μg/mL control IgG (R&D Systems, Minneapolis, MN, USA) or an IL-17A-neutralizing antibody (R&D Systems) prior to functional assays.
Mouse model of lung metastasis
All animal procedures were conducted in accordance with institutional regulations and the ARRIVE guidelines. Female NOD/SCID mice (4–6 weeks old, 20–22 g) were stratified by body weight and randomly assigned to control or treatment groups. Three independent in vivo experiments were performed:
Premetastatic niche model
To evaluate the effect of exosomes on lung stromal remodeling, mice (n = 8 per group) received daily tail vein injections of Vector-exo or S100A9-exo (70 μg) for 7 consecutive days. On day 8, mice were sacrificed, and lungs were harvested for immunohistochemical and immunofluorescence analyses.
Exosome-mediated metastasis model
To assess exosome-induced metastasis, mice (n = 7 per group) were pretreated with exosomes as described above. On day 8, 1 × 106 luciferase-expressing SACC-83 cells were injected via the tail vein. After 8 weeks, lung metastases were evaluated by bioluminescence imaging, followed by H&E staining.
CAF co-culture metastasis model
To assess the prometastatic potential of activated fibroblasts, luciferase-expressing SACC-83 cells were co-cultured with exosome-treated fibroblasts prior to injection. SACC-83 cells (1 × 106) were subsequently harvested and injected via the tail vein into NOD/SCID mice (n = 7 per group). After 8 weeks, lung metastases were assessed by bioluminescence imaging and H&E staining.
Immunohistochemistry (IHC) and multiplex immunohistochemistry (mIHC)
FFPE ACC tumor sections were deparaffinized, rehydrated, and subjected to heat-induced antigen retrieval in sodium citrate buffer (pH 6.0). Following blocking with 5% BSA for 1 h at room temperature, slides were processed. For standard IHC, slides were incubated overnight at 4 °C with primary antibodies, followed by incubation with secondary antibodies for 30 min at room temperature. Signals were visualized using a DAB substrate kit (ZLI-9017, ZSGB-Bio), and nuclei were counterstained with hematoxylin. Multiplex IHC (mIHC) was performed using a multi-labeling kit (Panovue, Beijing, China) following the manufacturer’s protocol. Targets were visualized using HRP conjugates and fluorophores at 520, 570, 620, 690, and 440 nm. Nuclei were counterstained with DAPI (ZLI-9556, ZSGB-Bio). Slides were scanned using the Aperio AT2 system (Leica, Wetzlar, Germany), and images were analyzed using ImageJ software. Detailed antibody information is provided in Supplementary Table 3.
Statistical analysis
Bioinformatics and statistical analyses were performed using R 4.4.2, SPSS 27, and GraphPad Prism 8.0. All in vitro experiments were conducted in at least three independent replicates, and representative data are shown. Numerical data are presented as the mean ± standard deviation (SD) or mean ± standard error of the mean (SEM), as specified in the figure legends. Statistical comparisons were performed using the two-tailed Student’s t test, Mann–Whitney U test, one-way or two-way ANOVA, Kruskal–Wallis test, Pearson correlation analysis, the log-rank test, or the chi-squared (χ²) test. Statistical significance was defined as p < 0.05 (ns, not significant; *p < 0.05; **p < 0.01; ***p < 0.001).
Ethics approval and consent to participate
This study was conducted in compliance with the principles of the Declaration of Helsinki. All methods were performed in accordance with the relevant guidelines and regulations. Human tissues and primary human cells were provided by the participants with written informed consent. Ethics approval for human subjects (number KQEC-2024-140-01) was provided by the Ethics Committee of the Hospital of Stomatology, Sun Yat-sen University. All animal procedures were approved by Permit number JENNIO-IACUC-2023-A068.
Data availability
The data or code supporting the findings of this study are available from the corresponding author upon reasonable request. The human scRNA-seq data have been deposited in the Genome Sequence Archive (GSA) with Accession Number HRA014947.
References
Drier Y, Cotton MJ, Williamson KE, Gillespie SM, Ryan RJ, Kluk MJ, et al. An oncogenic MYB feedback loop drives alternate cell fates in adenoid cystic carcinoma. Nat Genet. 2016;48:265–72.
de Sousa LG, Neto FL, Lin J, Ferrarotto R. Treatment of recurrent or metastatic adenoid cystic carcinoma. Curr Oncol Rep. 2022;24:621–31.
Ouyang DQ, Liang LZ, Zheng GS, Ke ZF, Weng DS, Yang WF, et al. Risk factors and prognosis for salivary gland adenoid cystic carcinoma in southern china: a 25-year retrospective study. Medicine. 2017;96:e5964.
Lorini L, Ardighieri L, Bozzola A, Romani C, Bignotti E, Buglione M, et al. Prognosis and management of recurrent and/or metastatic head and neck adenoid cystic carcinoma. Oral Oncol. 2021;115:105213.
Liu Y, Sinjab A, Min J, Han G, Paradiso F, Zhang Y, et al. Conserved spatial subtypes and cellular neighborhoods of cancer-associated fibroblasts revealed by single-cell spatial multi-omics. Cancer Cell. 2025;43:905–24.e6.
Li RF, Liu S, Gao Q, Fu M, Sun XY, Xiao M, et al. Inhibition of CDH11 activates cGAS-STING by stimulating branched chain amino acid catabolism and mitigates lung metastasis of adenoid cystic carcinoma. Adv Sci. 2025;12:e2408751.
Lu Y, Zheng J, Lin P, Lin Y, Zheng Y, Mai Z, et al. Tumor microenvironment-derived exosomes: a double-edged sword for advanced T cell-based immunotherapy. ACS Nano. 2024;18:27230–60.
Chen CW, Fu M, Du ZH, Zhao F, Yang WW, Xu LH, et al. Long noncoding RNA MRPL23-AS1 promotes adenoid cystic carcinoma lung metastasis. Cancer Res. 2020;80:2273–85.
Fu M, Gao Q, Xiao M, Li RF, Sun XY, Li SL, et al. Extracellular vesicles containing circMYBL1 induce CD44 in adenoid cystic carcinoma cells and pulmonary endothelial cells to promote lung metastasis. Cancer Res. 2024;84:2484–500.
Zhong C, Niu Y, Liu W, Yuan Y, Li K, Shi Y, et al. S100A9 derived from chemoembolization-induced hypoxia governs mitochondrial function in hepatocellular carcinoma progression. Adv Sci. 2022;9:e2202206.
Chung YH, Park J, Cai H, Steinmetz NF. S100A9-targeted cowpea mosaic virus as a prophylactic and therapeutic immunotherapy against metastatic breast cancer and melanoma. Adv Sci. 2021;8:e2101796.
Shen S, Zhang M, Wang X, Liu Q, Su H, Sun B, et al. Single-cell RNA sequencing reveals S100a9(hi) macrophages promote the transition from acute inflammation to fibrotic remodeling after myocardial ischemia‒reperfusion. Theranostics. 2024;14:1241–59.
Mehta AR, Armstrong AJ. Tasquinimod in the treatment of castrate-resistant prostate cancer—current status and future prospects. Ther Adv Urol. 2016;8:9–18.
Prieto D, Sotelo N, Seija N, Sernbo S, Abreu C, Durán R, et al. S100-A9 protein in exosomes from chronic lymphocytic leukemia cells promotes NF-κB activity during disease progression. Blood. 2017;130:777–88.
Wang F, Li Y, Yang Z, Cao W, Liu Y, Zhao L, et al. Targeting IL-17A enhances imatinib efficacy in Philadelphia chromosome-positive B-cell acute lymphoblastic leukemia. Nat Commun. 2024;15:203.
Straus DS. TNFα and IL-17 cooperatively stimulate glucose metabolism and growth factor production in human colorectal cancer cells. Mol Cancer. 2013;12:78.
Dal Lago EA, Sousa LG, Yang Z, Hoff CO, Bonini F, Sawyer M, et al. Prognostic value of tumor volume doubling time in lung-metastatic adenoid cystic carcinoma. Oral Oncol. 2024;151:106759.
de Visser KE, Joyce JA. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell. 2023;41:374–403.
Mashouri L, Yousefi H, Aref AR, Ahadi AM, Molaei F, Alahari SK. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer. 2019;18:75.
He G, Peng X, Wei S, Yang S, Li X, Huang M, et al. Exosomes in the hypoxic TME: from release, uptake and biofunctions to clinical applications. Mol Cancer. 2022;21:19.
Yang WW, Yang LQ, Zhao F, Chen CW, Xu LH, Fu J, et al. Epiregulin promotes lung metastasis of salivary adenoid cystic carcinoma. Theranostics. 2017;7:3700–14.
Gysler SM, Drapkin R. Tumor innervation: peripheral nerves take control of the tumor microenvironment. J Clin Invest. 2021;131:e147276.
Chen Y, Ouyang Y, Li Z, Wang X, Ma J. S100A8 and S100A9 in cancer. Biochim Biophys Acta Rev Cancer. 2023;1878:188891.
Jukic A, Bakiri L, Wagner EF, Tilg H, Adolph TE. Calprotectin: from biomarker to biological function. Gut. 2021;70:1978–88.
Wang Y, Yin K, Tian J, Xia X, Ma J, Tang X, et al. Granulocytic myeloid-derived suppressor cells promote the stemness of colorectal cancer cells through exosomal S100A9. Adv Sci. 2019;6:1901278.
Fan R, Satilmis H, Vandewalle N, Verheye E, De Bruyne E, Menu E, et al. Targeting S100A9 protein affects mTOR-ER stress signaling and increases venetoclax sensitivity in acute myeloid leukemia. Blood Cancer J. 2023;13:188.
Sousa LG, McGrail DJ, Lazar Neto F, Li K, Marques-Piubelli ML, Ferri-Borgogno S, et al. Spatial immunoprofiling of adenoid cystic carcinoma reveals B7-H4 is a therapeutic target for aggressive tumors. Clin Cancer Res. 2023;29:3162–71.
Parikh AS, Wizel A, Davis D, Lefranc-Torres A, Rodarte-Rascon AI, Miller LE, et al. Single-cell RNA sequencing identifies a paracrine interaction that may drive oncogenic notch signaling in human adenoid cystic carcinoma. Cell Rep. 2022;41:111743.
Chhabra Y, Weeraratna AT. Fibroblasts in cancer: unity in heterogeneity. Cell. 2023;186:1580–609.
Sun X, He X, Zhang Y, Hosaka K, Andersson P, Wu J, et al. Inflammatory cell-derived CXCL3 promotes pancreatic cancer metastasis through a novel myofibroblast-hijacked cancer escape mechanism. Gut. 2022;71:129–47.
Özbay Kurt FG, Cicortas BA, Balzasch BM, De la Torre C, Ast V, Tavukcuoglu E, et al. S100A9 and HMGB1 orchestrate MDSC-mediated immunosuppression in melanoma through TLR4 signaling. J Immunother Cancer. 2024;12:e009552.
Wu F, Yang J, Liu J, Wang Y, Mu J, Zeng Q, et al. Signaling pathways in cancer-associated fibroblasts and targeted therapy for cancer. Signal Transduct Target Ther. 2021;6:218.
Li X, Bechara R, Zhao J, McGeachy MJ, Gaffen SL. IL-17 receptor-based signaling and implications for disease. Nat Immunol. 2019;20:1594–602.
Picard FSR, Lutz V, Brichkina A, Neuhaus F, Ruckenbrod T, Hupfer A, et al. IL-17A-producing CD8(+) T cells promote PDAC via induction of inflammatory cancer-associated fibroblasts. Gut. 2023;72:1510–22.
Amatya N, Garg AV, Gaffen SL. IL-17 signaling: the Yin and the Yang. Trends Immunol. 2017;38:310–22.
Noack M, Beringer A, Miossec P. Additive or synergistic interactions between IL-17A or IL-17F and TNF or IL-1β depend on the cell type. Front Immunol. 2019;10:1726.
Sun D, Novotny M, Bulek K, Liu C, Li X, Hamilton T. Treatment with IL-17 prolongs the half-life of chemokine CXCL1 mRNA via the adaptor TRAF5 and the splicing-regulatory factor SF2 (ASF). Nat Immunol. 2011;12:853–60.
Karlsen JR, Borregaard N, Cowland JB. Induction of neutrophil gelatinase-associated lipocalin expression by co-stimulation with interleukin-17 and tumor necrosis factor-alpha is controlled by IkappaB-zeta but neither by C/EBP-beta nor C/EBP-delta. J Biol Chem. 2010;285:14088–100.
Gao F, Chen X, Li X, Deng C, Luo P. The pro-migratory and pro-invasive roles of cancer-associated fibroblasts secreted IL-17A in prostate cancer. J Biochem Mol Toxicol. 2025;39:e70047.
Liu C, Liu R, Wang B, Lian J, Yao Y, Sun H, et al. Blocking IL-17A enhances tumor response to anti-PD-1 immunotherapy in microsatellite stable colorectal cancer. J Immunother Cancer. 2021;9:e001895.
Li SL. Establishment of a human cancer cell line from adenoid cystic carcinoma of the minor salivary gland. Zhonghua Kou Qiang Yi Xue Za Zhi. 1990;25:29–31.
Wheeler SE, Shi H, Lin F, Dasari S, Bednash J, Thorne S, et al. Enhancement of head and neck squamous cell carcinoma proliferation, invasion, and metastasis by tumor-associated fibroblasts in preclinical models. Head Neck. 2014;36:385–92.
Hao Y, Stuart T, Kowalski MH, Choudhary S, Hoffman P, Hartman A, et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. 2024;42:293–304.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–37.e4.
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16:1289–96.
Jin S, Plikus MV, Nie Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat Protoc. 2025;20:180–219.
Acknowledgements
We thank X.-Y. Ren for the time and support.
Funding
This study was funded by the National Natural Science Foundation of China (82103555) and the China Postdoctoral Science Foundation (2021M703688, 2022T150753).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Cw., Zhang, Sr., Yan, Ym. et al. Exosomal S100A9 promotes lung metastasis of adenoid cystic carcinoma via activating cancer-associated fibroblasts. Cell Death Discov. 12, 120 (2026). https://doi.org/10.1038/s41420-026-02991-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-026-02991-6