Abstract
Our research focuses on Rhodovulum sulfidophilum, a marine, anoxygenic purple nonsulfur photosynthetic bacterium (PNSB), as a tool for the production of materials for a sustainable society. The most sustainable approach with PNSB is to culture it under autotrophic conditions using carbon dioxide and nitrogen fixation pathways. By introducing exogenous genes of interest, R. sulfidophilum, one of the most studied PNSB, can serve as a protein expression host under autotrophic conditions. Herein, through autotrophic culture, recombinant spider silk proteins were successfully produced. This research is expected to contribute to the development of sustainable material production using PNSB.
Similar content being viewed by others
Introduction
The production of useful substances using light energy by microorganisms with photosynthetic metabolic pathways has attracted attention as an environmentally friendly method1,2. Compared to traditional plant-based protein production, it offers advantages such as reduced area requirements, higher cellular protein content, and shorter growth times1,3. Furthermore, some of these microorganisms have metabolic pathways for nitrogen fixation in addition to photosynthesis, and unlike plants, they can supply their own nitrogen source, which is necessary for growth. Purple nonsulfur photosynthetic bacteria (PNSB) are attracting attention as such tools for environmentally friendly, resource-circulating societies because they can fix gaseous CO2 and N2 in the environment in their metabolic system3. PSNB perform anoxygenic photosynthesis but are facultative anaerobes; thus, they do not require strict oxygen-free conditions, making culturing methods simple3. Since PNSB are rich in amino acids, they are expected to be used as agricultural fertilizers4, as bait in fisheries3 or as hosts for the expression of heterologous proteins via genetic engineering5,6. PNSB also accumulates other useful substances, such as carotenoids, vitamins, and biodegradable plastics, depending on the culture conditions3,6; thus, they are expected to be useful production tools. By utilizing light energy, PNSB can grow rapidly in mineral medium containing simple organic carbon (C) sources such as butyrate, acetate, or propionate anaerobically7. Even in the absence of organic C compounds, by using light energy, PNSB can grow lithoautotrophically8: they can fix CO2 via electrons from reduced inorganic compounds such as H2S, Na2S2O38, and metals9. PNSB have also been investigated as bioremediation tools because they have a relatively high tolerance to metals and can detoxify toxic metals through oxidation and reduction9. Among PNSB, we focus on Rhodovulum sulfidophilum, a marine PNSB that exhibits superior reduced inorganic sulfur compound metabolism8,10,11. The sulfur concentration in seawater is higher than that in freshwater rivers and lakes12; hence, R. sulfidophilum possesses a sulfur-oxidizing gene cluster, the sox gene8,10. In our previous study, little growth of R. sulfidophilum was observed when N2 gas and NaHCO3 were added as inorganic N and C sources to artificial seawater5, but the addition of Na2S2O3 promoted bacterial growth by inducing not only the CO2 and nitrogen (N) fixation pathways but also possibly the lithotrophic sulfur oxidation metabolic pathway13. When light energy is provided for growth, PNSB exhibit a high growth rate under heterotrophic conditions; therefore, applied research on PNSB has been conducted mainly under photoheterotrophic conditions3,7, while the most sustainable approach would likely involve culturing bacteria under autotrophic conditions with pathways to fix CO2- and N, using gaseous CO2 and N2 present in large amounts in the air5. Furthermore, seawater is inexhaustible and has less impact on the environment than freshwater does, reducing costs and the risk of bacterial contamination due to its high sodium chloride content13. Exploiting these culture conditions, we aimed to produce useful heterologous R. sulfidophilum proteins for the purpose of sustainable material production (Fig. 1a).
a Schematic diagram of the experiment. b Plasmid map of the recombinant MaSp2 protein. c Amino acid sequence of the MaSp2 coding region.
As a recombinant protein produced by R. sulfidophilum, we focused on major ampullate spidroin 2 (MaSp2), which is produced by the spider Trichonephila clavipes14,15. The soluble form of the protein self-assembles into hierarchical structures, generating insoluble fibrous structural proteins by being processed in spider glands. The fibrous protein produced by spiders is commonly called spider silk and has attracted high interest: it is strong and flexible, making it promising for industrial applications in clothing, medical textiles, and other areas15,16,17,18. Spiders cannot be cultured artificially because of their cannibalism; thus, efforts are underway to use genetic engineering to overexpress soluble silk proteins in microorganisms such as yeast and Escherichia coli15 grown in medium rich in organic carbon and nitrogen sources, after which the collected silk proteins are converted into fibrous forms with original hierarchical structures using organic solvents or aqueous solutions such as phosphate buffer17,18. Here, we aimed to produce spider silk protein using an autotrophic process to address sustainability issues in biomanufacturing fields. In our previous paper, recombinant R. sulfidophilum harboring a plasmid encoding a different major ampullate spidroin 1 (MaSp1) was cultivated under photoautotrophic conditions5. When cultured for 7 days in artificial seawater supplemented with 12 mM NaHCO3 as an inorganic C source and 0.5 L/day of N2 gas as an N source, the cells showed little growth. In contrast, in this study, recombinant R. sulfidophilum harboring a MaSp2 expression plasmid was cultured in artificial seawater supplemented with CO2 gas as an inorganic C source, N2 gas as a N source, and a relatively high concentration of Na2S2O3 (approximately 9.7 mM) as an electron donor. This promoted cell growth (Fig. 2, Table 1) and resulted in the production of partially purified soluble MaSp2 protein at a yield of 3.2 ± 3.8 µg/L (Table 2).
a Photograph of R. sulfidophilum transformants cultured in seawater supplemented with CO2 gas, N2 gas, and Na2S2O3. b The growth curve and the amount of inorganic C or N taken up by the cells on each culture day. These data are the averages from 3 independent experiments, and the error bars represent the SDs. c The expression level of MaSp2 protein in the cells. Whole cell lysates from equal numbers of cells harvested on days 2 to 6 of the culture were loaded onto SDS-PAGE. The left and right panels show Coomassie blue staining images and Western blot images, respectively. His-tag (6xHis-tag) antibodies were used for Western blotting. The black triangle indicates the 43 kDa band of the model MaSp2 protein.
Results
The addition of inorganic reduced sulfur compounds to artificial seawater increased cell proliferation
To facilitate R. sulfidophilum growth on a larger scale in a shorter culture period, in addition to inorganic and organic C/N dissolved in artificial seawater5, it is necessary to inject gaseous CO2 and N2 into seawater. The pH of seawater is slightly alkaline, approximately 8, and at this pH, most CO2 dissolved in seawater exists as HCO3- ions. By injecting gaseous CO2, the pH of seawater quickly decreases from approximately 8 to approximately 5, with CO2 dissolved in seawater existing in the form of gaseous CO2 rather than HCO3- ions, and the solubility of gaseous CO2 is low. Thus, during gas injection, 1 N NaOH was continuously added dropwise to dissolve CO2 in the form of HCO3- ions. In a 10 L-capacity incubator filled with 8.9 L of artificial seawater, CO2 and N2 gas were bubbled at rates of 0.15 L/min and 0.35 L/min, respectively, for 2 h at 30 °C. Before gas injection, the inorganic C concentration of artificial seawater was approximately 20–22 mg/L, and the 2 h gas injection resulted in approximately 340–420 mg/L inorganic C. After the gas supply was stopped, Na2S2O3 was added as an electron donor to a final concentration of 9.7 mM, followed by the addition of an antibiotic solution and a culture of R. sulfidophilum transformed with a plasmid encoding the MaSp2 gene (Fig. 1b). The coding region consisted of the three parts, N-terminal domain (NTD) and C-terminal domain (CTD), with a core 6 tandem repeat sequence between them, mimicking the structure found in nature (Fig. 1c). Because the core tandem repeat is directly related to the mechanical properties of spider silk, such as its strength, studies using recombinant proteins often focus exclusively on the core repeat domain but the NTD and CTD globular proteins are also important for the solubility of the protein and formation of hierarchical structures19,20, affecting the properties of the produced fiber19.
Cell division continued until day 4, after which the bacteria entered the stationary phase (Fig. 2a, b, Table 1). The specific growth rate was highest on days 0-2, at 0.56 ± 0.13 (Table 1). After the second day, the amount of inorganic C or N taken up into the cells was measured using total C (TC), inorganic C (IC), and total N (TN) analyses. There was a rough correlation between cell growth and the amount of C and N assimilated each day of culture (Fig. 2b, Table 1). This result is reasonable considering that amino acids are the main components of microorganisms and that the absorbed CO2/HCO3- ions and N2 gas are synthesized into amino acids21,22. The intracellular C:N molar ratio was roughly constant, ranging from 7.9 to 10.7 (Table 1), regardless of the specific growth rate (Table 1). The C:N ratio of marine bacteria has been reported to range from 5.0 to 8.323, suggesting that the cells in this culture system were prone to nitrogen deficiency stress.
The intracellular protein expression of MaSp2 was not detected by Coomassie blue staining but rather by Western blotting (Fig. 2c). MaSp2 expression per cell was highest on day 2 (Fig. 2c) of the logarithmic growth phase, but the culture was continued until the stationary phase to obtain more cells (Fig. 3a). After 8–10 days of incubation, TC, IC and TN analyses using the medium collected on the final day of each culture (Table 2) revealed that approximately 65.5 ± 27.9 mg/L and 5.4 ± 1.1 mg/L inorganic C and N were taken up into the cells (Table 2) and 1.6 ± 0.7 g wet cells were collected (Table 2).
a Growth curve for R. sulfidophilum transformants in triplicate experiments. b Cell lysates disrupted by sonication were eluted through a Ni column and analyzed by SDS‒PAGE. Lane 1: cell lysate immediately before loading onto the Ni column. FT: flow-through. After imidazole passed through the column, the eluate was collected in 500 µL aliquots, and 10 µL of each aliquot was loaded into the wells from left to right in elution order. The three elution fractions marked with * were collected and subjected to ultracentrifugation. c Lane 1 represents the cell lysate before being applied to the Ni column, and lane 2 represents the protein mixture after being subjected to ultrafiltration (MWCO, 10,000). The left and right panels show Coomassie blue staining and Western blot results, respectively, for b and c. The black triangle indicates the 43 kDa band of the model MaSp2 protein.
Partial purification of the MaSp2 protein
After sonication, cell lysates were passed through a standard affinity Ni‒nitrilotriacetic acid (NTA) column24. The N-terminus of the coding region of MaSp2 contains six consecutive histidine residues (Fig. 1c). The affinity of these residues for Ni-NTA was exploited to bind the MaSp2 protein to the column, which was then eluted by competition with imidazole. The MaSp2 protein band was detected in 10 µL of the Ni‒NTA column eluate (approximately 1/50 of each elution fraction), indicating that the protein was concentrated (Fig. 3b). The three fractions with the highest MaSp2 elution were mixed and ultrafiltered, and the solvent was replaced with purified H2O, resulting in the purification of MaSp2 (Fig. 3c). The total soluble protein (TSP) content resulted in 21.4 ± 13.9 µg/L (141.8 ± 128.6 µg/g wet weight cell), with the MaSp2 protein accounting for 12.1 ± 9.0% of the total soluble protein (TSP) content, resulting in 3.2 ± 3.8 µg/L (24.6 ± 34.7 µg/g wet weight cell) MaSp2 (Table 2 and Supplementary Fig. 1).
Discussion
From a sustainability perspective, we focused on the use of inorganic resources in bacterial cultivation, using the reduced inorganic sulfur compound Na2S2O3 as the electron donor and gaseous CO2/N2 as the C/N source. Artificial seawater was used as the basal medium because the use of seawater is considered superior to the use of freshwater because it utilizes an unused resource. Under these sustainable and autotrophic conditions, a soluble form of the spider silk protein MaSp2, 3.2 ± 3.8 µg/L (24.6 ± 34.7 µg/g wet weight cell) (Table 2), was obtained from R. sulfidophilum. Previously, our research group cultured R. sulfidophilum carrying a plasmid encoding the gene for ampullate spidroin MaSp1 in seawater supplemented with 12 mM NaHCO3 (equivalent to an IC of 143 mg/L) and N2 gas, resulting in little cell growth that did not support further investigation5; under these experimental conditions, the yield of MaSp1 was 0.12 ± 0.10 mg/L, which was estimated to be derived from seed culture cells using organic N/C-rich MB medium. However, in the present study, R. sulfidophilum proliferated under similar autotrophic conditions when Na2S2O3 was added as an electron donor. Thus, sufficient and balanced electron donors and N and C sources are still important even for the autotrophic bacterial production of exogenous proteins. On the other hand, in the same paper, the addition of 0.4 g/L yeast extract in seawater supplemented with 12 mM NaHCO3 and N2 gas promoted cell growth and increased MaSp1 protein yield to 3.9 ± 2.8 mg/L, and when MB medium containing abundant organic N/C sources were used, the MaSp1 yield was 52.3 ± 11.2 mg/L5, which is comparable to that of heterologous protein expression systems in cyanobacteria25. These data indicate that under photoheterotrophic conditions, production of spider silk protein or possibly other heterologous proteins by R. sulfidophilum can increase from a few to tens of mg/L or mg/g cell levels. We speculate that such high levels of heterologous protein expression may be achievable even under autotrophic culture conditions if the efficiency of photosynthesis is improved and, consequently, the level of intracellular organic C is increased.
Data on the molar ratio of C and N incorporated into cells revealed that cells tend to be in an N-starved state when both CO2 and N fixation were simultaneously induced (Table 1). The C:N ratio of marine bacteria has been reported to be in the range of 5.0 to 8.323, but in this study, the C:N ratio obtained was in the range of 7.9 to 10.7 (Table 1), suggesting that the current method only achieves the minimum N fixation required for the cells, and N fixation is the rate-limiting factor in this system. Because 16 mol of ATP is required to fix 1 mol of N2, and ATP is also required for the subsequent amino acid synthesis, it is presumed that ATP is insufficient under the culture conditions. We speculate that restoring the intracellular C:N ratio to the standard levels (5.0 to 8.3) can be achieved by increasing the efficiency of photosynthesis, as mentioned above, to increase the amount of ATP available for N fixation and amino acid synthesis. When N-starved state is alleviated, an overall increase in amino acid synthesis is expected, along with increased production of MaSp2.
Compared with the growth of PNSB in common mineral medium26,27, the bacterial growth in the medium used herein was slightly lower, even though the seawater in the present study was presumably supplemented with sufficient C, N, and electron sources. We recently reported that when photosynthesis and N fixation by nitrogenase were simultaneously performed in mineral medium containing vitamins and trace metal elements instead of seawater, the growth of R. sulfidophilum was suppressed compared with that of R. sulfidophilum in heterotrophic medium22. We therefore consider that the low growth rate is not due to a lack of vitamins and/or trace metal elements in the culture medium but rather to a lack of other components that properly regulate the assimilation of C and N. Alternatively, the necessary components may be present in the seawater and the mineral medium above, but the concentrations of these components may need to be adjusted to efficiently induce both CO2 and N fixation simultaneously. Phosphorus and iron are well-known rate-limiting factors for the growth of marine diazotrophs28; hence, they could be candidates for controlling photosynthetic activity and cellular growth. In the future, we will attempt to precisely adjust the concentrations of these potential components.
We speculate that manipulating the promoter region of the plasmid may also increase the production of spider silk proteins. The plasmid used in this study constitutively expresses a heterologous protein that is thought to be a constant source of stress for cells; controlling the timing of heterologous protein expression would reduce the stress, promote cell growth, and ultimately increase protein production.
As shown in Fig. 1c, the 6xMaSp2 repeat sequences of the MaSp2 protein are particularly enriched for certain amino acids, such as alanine, glutamine, and glycine, which would lead to a shortage of intracellular tRNAs corresponding to these amino acids. Introduction of genes encoding these tRNAs is also expected to lead to increased protein production. Besides, codon optimization and 5’ untranslated region engineering may be promising options to increase spider silk production15,29.
It is well known that under photoheterotrophic conditions, the metabolism of PNSB varies significantly depending on the intensity, wavelength, and duration of light irradiation30,31. Nitrogen fixation in PNSB also depends on light32. Because our study was conducted under photoautotrophic conditions, the role of photosynthesis is likely to be more important than in previous studies. Light irradiation conditions may be a factor that determines the amount of heterologous protein produced, and further investigation of light irradiation conditions is essential.
Alterations to the culture method may also increase cell numbers and heterologous protein production. In this study, we performed batch culture, in which all components were added before starting the culture. However, perfusion culture, in which old medium is removed and new medium is added periodically, may yield better results33 because perfusion culture is expected to remove harmful waste products and add trace elements such as metals contained in seawater.
Like PNSB, cyanobacteria could fix CO2 and N2, and have been established early as tools for producing a wide range of useful substances1. Compared to cyanobacteria, PNSB has two major advantages as a host for recombinant proteins: first, it can utilize a wide range of substances as electron donors, and second, it does not require or produce oxygen during cultivation. While most cyanobacteria use H2O as the sole electron source during photosynthesis, PNSB cannot use H2O as an electron source but can utilize a variety of organic carbon compounds, including lignin, and reduced inorganic compounds as electron sources34. In particular, R. sulfidophilum can utilize sulfur compounds such as Na2S2O3 and H2S11, it is expected that various waste materials from food factories, paper factories, and factories that emit sulfur-containing wastewater and exhaust gases can be used as electron sources to grow bacteria. Nitrogen fixation is highly sensitive to oxygen and can even be inhibited by environmental oxygen levels, but cyanobacteria require oxygen supply to grow and release oxygen during photosynthesis35. Although cyanobacteria have several sophisticated mechanisms to mitigate the inhibitory effect of oxygen on nitrogen fixation, the high concentrations of oxygen still inhibit nitrogen fixation35. PNSB neither requires nor produces oxygen, which may make them superior to cyanobacteria in simultaneous photosynthesis and nitrogen fixation induction.
The Cartagena Protocol on Biosafety is an international agreement that mandates the prevention of the spread of genetically modified microorganisms into the environment. Each country has its own national regulations based on the Cartagena Protocol. For example, in Japan, recombinant R. sulfidophilum must be grown in a controlled environment, and the risk of the bacteria spreading into the environment is considered low; cultivation must be carried out in a laboratory with specific equipment, and the culture medium must be inactivated (e.g., autoclaved) at the end of the experiment. However, if the scale of aquaculture expands to several hundred liters in the future, the risk of spreading to the environment is expected to increase. In case they do spread into the environment, it would be necessary to establish a method to prevent genetically modified organisms from multiplying there. Auxotrophic strategies that require nutrients that are rarely present in the environment should be introduced in this system in future36.
This study demonstrated that under autotrophic conditions, both recombinant and exogenous proteins were reliably produced; however, the addition of high concentrations of electron donors and CO2/N2 gas was still insufficient to achieve sufficient protein production through photosynthesis and N fixation for industrial applications. To improve and enhance autotrophic productivity, some strategic modifications to culture conditions are needed to drive these metabolic processes efficiently. Since N fixation by nitrogenase appears to be rate-limiting and photosynthesis and N fixation are closely related22,37, improvement in the N fixation pathway could increase photosynthesis via reduced sulfur compounds, resulting in increased biomass production from the photosynthetic process.
Methods
Growth conditions
R. sulfidophilum (ATCC35886/DSM1374) was obtained from the American Type Culture Collection (ATCC). R. sulfidophilum containing the plasmid was precultured with mineral medium M6 containing 20 mM NaHCO3 as the inorganic C source, 7.6 mM (NH4)2SO4 as the fixed N source, 8 mM Na2S2O3 as the electron source, and 50 µg/mL kanamycin. The composition of the M6 medium per 1 L was as follows: K2HPO4, 0.78 g; KH2PO4, 0.75 g; CaCl2·2H2O, 0.029 g; MgSO4·7H2O, 0.247 g; FeSO4·7H2O, 0.011 g; NaCl, 20 g; vitamin solution, 10 mL; and trace element mixture, 10 µL. The pH was adjusted to 7.0 with NaOH, and the medium was sterilized by autoclaving at 121 °C for 15 min. The composition of the vitamin mixture per 1 L was as follows: nicotinamide acid, 1.0 g; thiamine, 1.0 g; biotin, 50.0 mg; PABA, 0.5 g; vitamin B12, 10.0 mg; Ca pantothenate, 0.5 g; folic acid, 0.5 g; pyridoxine HCl, 0.5 g; and EDTA (3Na), 2.0 g. The composition of the trace element mixture per 1 L was as follows: MnSO4·4H2O, 11.2 g; ZnSO4·7H2O, 2.9 g; Co(NO3)2·6H2O, 2.9 g; CuSO4·5H2O, 2.5 g; H3BO3, 3.1 g; Na2MoO4·2H2O, 2.4 g; and EDTA (3Na), 41.2 g. Precultured cells in the logarithmic growth phase in M6 medium were collected, and the cell pellets were washed and concentrated with artificial seawater (AIR WATER, Osaka, Japan). Cell density was assessed by measuring the absorbance of the culture medium at 660 nm using a spectrophotometer (AS ONE, Osaka, Japan). A total of 8.9 L of artificial seawater filtered through a 0.2 µm filter was placed in a 10 L capacity fermenter, which was equipped with a pH sensor to automatically inject acid or alkaline solutions to adjust the pH (Marubishi Bioengineering, Tokyo, Japan). Artificial seawater was used as the basal medium because the composition of seawater varies greatly depending on the season and collection location. CO2 and N2 gases were bubbled at rates of 0.15 L/min and 0.35 L/min, respectively, while stirring at 100 rpm for 2 h at 30 °C, and 1 N NaOH was added dropwise automatically. Approximately 250 mL of 1 N NaOH was used over the 2-h infusion, and the pH reached approximately 6.6. After the gas supply was stopped, 90 mL of 1 M Na2S2O3 was added as an electron donor to a concentration of 9.7 mM, followed by the addition of 4.5 mL of 100 mg/mL kanamycin solution to a concentration of 49 µg/mL, and approximately 10 mL of the concentrated culture of R. sulfidophilum transformant was added to obtain an optical density of approximately 0.02 at 660 nm. The inorganic C concentration reached 340–420 mg/L 2 h after injection, as determined using a TOC instrument (Shimadzu, Kyoto, Japan). All parts of the fermenter exposed to the outside air were sealed, and the fermenter was kept at 30 °C under 730 nm far-red light with gentle stirring at 50 rpm. Light irradiation panels (CCS, Kyoto, Japan) were installed on both sides of the fermenter, and the incident light intensity on each was set to 20 W/m2 and 170 µmol photons/m2/s (380 nm to 780 nm). The λmax and full width at half maximum of the LED emission spectrum were 733 nm and 717–748 nm, respectively (Supplementary Fig. 2). The 730 nm wavelength was used because its effect on the growth of R. sulfidophilum has been studied. After 8–10 days, the culture was stopped. After the cells were collected by centrifugation, they were washed once with artificial seawater, and the pellet was stored at -80 °C until cell disruption and protein purification began. A total of 1.0–2.4 g of wet cells was collected in 3 experiments.
Plasmid construction and conjugation
A model MaSp2 coding region consisting of the N-terminal domain, six repeats of the tandem domain, and the C-terminus was transferred from the plasmid used in Escherichia coli14 into a modified pBBR1MCS-2 broad-host-range vector5. The NTD and 6 repeats were derived from MaSp2 of Trichonephila clavipes, and the CTD was derived from MaSp2 of Latrodectus hesperus. The coding region was located downstream of the trc promoter, and there was a histidine tag at the N-terminus5. This plasmid was introduced into R. sulfidophilum via the RP4/RK2 conjugation system, as described previously5. Plasmid maps were generated using SnapGene® software (Dotmatics, MA, USA).
Analysis of MaSp2 protein expression levels
Based on the optical density values at 660 nm, equal numbers of cultured cells were collected by centrifugation. The cell pellet was suspended in Laemmli sample buffer containing 2-mercaptoethanol, heated at 100 °C for 5 min, and centrifuged at 13,000 rpm for 5 min. The supernatant was analyzed by SDS‒PAGE on a 4–15% gradient gel (Bio-Rad, CA, USA), and protein bands were analyzed by Coomassie blue staining (Tokyo Chemical Industry, Tokyo, Japan) or semidry Western blotting using a PVDF membrane (Bio-Rad, CA, USA) and an anti-His-tag (6xHis-tag) antibody (MBL, Tokyo, Japan).
Protein purification
Fifty milliliters of purified water was added per 1 g of wet cells, and the suspended cells were subjected to sonication (TOMY SEIKO, Tokyo, Japan). The homogenate was centrifuged, and the supernatant was collected. The supernatant was adjusted to 20 mM sodium phosphate, 500 mM NaCl, and 20 mM imidazole and run over a 1 mL histidine-tagged column (HisTrap FF, Cytiva, Tokyo, Japan). The bound proteins were eluted via isocratic elution with 500 mM imidazole. Approximately 1.5 mL of the eluate was subjected to ultrafiltration (Cytiva, Tokyo, Japan, molecular weight cutoff of 10,000), replaced with H2O by adding 5 mL of H2O three times, and concentrated to a final volume of 300–500 µL. The protein concentration was determined using the BCA protein assay (Thermo Fisher Scientific, MA, USA). Soluble protein was subjected to SDS‒PAGE on a 4–15% gradient gel (Bio-Rad, CA, USA) and detected by Coomassie blue staining (Tokyo Chemical Industry, Tokyo, Japan) or semidry Western blotting using a PVDF membrane (Bio-Rad, CA, USA) and an anti-His-tag (6xHis-tag) antibody (MBL, Tokyo, Japan). The amount of MaSp2 in TSP was calculated using Coomassie-stained gels by image analysis software (ATTO, Tokyo, Japan). The validity of this calculation method was confirmed by comparing the obtained protein solution with a purified His-tagged standard protein solution using Coomassie staining and Western blotting (Supplementary Fig. 1).
Analysis of the amount of carbon and nitrogen taken up into cells
The culture mixture and the supernatant obtained by centrifugation (9000 g, 10 min, 4 °C) were subjected to total organic carbon (TOC) and total nitrogen (TN) analyses (Shimadzu, Kyoto, Japan), respectively, to determine the total carbon (TC, mg/L), inorganic carbon (IC, mg/L), and total N (TN, mg/L) in each sample. The total organic carbon (TOC, mg/L) content was calculated by subtracting the IC value from the TC value. The amounts of carbon and nitrogen taken up into the cells were calculated using the following formulas:
Carbon taken up into cells (mg/L) = TC (mg/L) of the culture – IC (mg/L) of the supernatant.
Nitrogen taken up into cells (mg/L) = TN (mg/L) of the culture – TN (mg/L) of the supernatant.
TOC analyzers use heat and a platinum catalyst to decompose TC into CO2, and measure the amount of CO2 gas as the peak area in the detector38,39. By calculating the ratio between this value and the area value of the TC standard solution, the TC concentration in the sample can be calculated. The amount of IC is determined by acid degradation of the sample (pH below 3), which releases CO2 from all carbonates in the sample, which is then measured in the same way as TC38,39. The amount of TN is measured by heating the sample to 720 °C, which causes the TN in the sample to decompose into nitric oxide, which is then analyzed in the gas analyzer and detected as a peak area39. However, this method cannot measure N2 gas because N2 gas in the sample is not converted to nitric oxide even when heated to 720°C. Therefore, fixed nitrogen was measured indirectly by subtracting the TN value of the supernatant from the TN value of the culture medium.
Data availability
The raw data were generated at Kyoto University and the RIKEN Center for Sustainable Resource Science. Derived data supporting the findings of this study are available from the corresponding author (K.N.) upon request.
References
Nawaz, T. et al. Sustainable protein production through genetic engineering of cyanobacteria and use of atmospheric N2 gas. Food Energy Secur. 13, e536 (2024).
Cheng, J. et al. Cyanobacteria-mediated light-driven biotransformation: the current status and perspectives. ACS Omega 8, 42062–42071 (2023).
Rashid, N., Onwusogh, U. & Mackey, H. R. Exploring the metabolic features of purple non-sulfur bacteria for waste carbon utilization and single-cell protein synthesis. Biomass-.-. Convers. Biorefin. 14, 12653–12672 (2024).
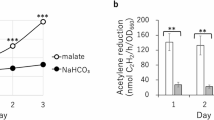
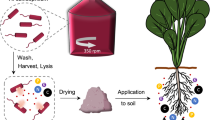
Morey-Yagi, S. R. et al. Utilization of lysed and dried bacterial biomass from the marine purple photosynthetic bacterium Rhodovulum sulfidophilum as a sustainable nitrogen fertilizer for plant production. npj Sustain. Agric. 2, 10 (2024).
Foong, C. P. et al. A marine photosynthetic microbial cell factory as a platform for spider silk production. Commun. Biol. 3, 357 (2020).
Srisawat, P., Higuchi-Takeuchi, M. & Numata, K. Microbial autotrophic biorefineries: Perspectives for biopolymer production. Polym. J. 54, 1139–1151 (2022).
Bayon-Vicente, G. et al. Metabolic pathways to sustainability: review of purple non-sulfur bacteria potential in agri-food waste valorization. Front. Bioeng. Biotechnol. 13, 1529032 (2025).
Ghosh, W. & Dam, B. Biochemistry and molecular biology of lithotrophic sulfur oxidation by taxonomically and ecologically diverse bacteria and archaea. FEMS Microbiol. Rev. 33, 999–1043 (2009).
Grattieri, M., Labarile, R., Buscemi, G. & Trotta, M. The periodic table of photosynthetic purple non-sulfur bacteria: intact cell-metal ions interactions. Photochem. Photobiol. Sci. 21, 101–111 (2022).
Appia-Ayme, C., Little, P. J., Matsumoto, Y., Leech, A. P. & Berks, B. C. Cytochrome complex essential for photosynthetic oxidation of both thiosulfate and Sulfide in Rhodovulum sulfidophilum. J. Bacteriol. 183, 6107–6118 (2001).
Hansen, T. A. & Veldkamp, H. Rhodopseudomonas sulfidophila, nov. spec., a new species of the purple nonsulfur bacteria. Arch. f. ür. Mikrobiol. 92, 45–58 (1973).
Jørgensen, B. B., Findlay, A. J. & Pellerin, A. The Biogeochemical Sulfur Cycle of Marine Sediments. Front. Microbiol. 10, 849 (2019).
Morey-Yagi, S. R. et al. Low-salinity medium for large-scale biomass production of the marine purple photosynthetic bacterium Rhodovulum sulfidophilum. bioRxiv, 2025.2003.2014.643199 (2025).
Malay, A. D. et al. Spider silk self-assembly via modular liquid-liquid phase separation and nanofibrillation. Sci. Adv. 6, eabb6030 (2020).
Ramezaniaghdam, M., Nahdi, N. D. & Reski, R. Recombinant spider silk: promises and bottlenecks. Front. Bioeng. Biotechnol. 10, 835637 (2022).
Numata, K. & Kaplan, D. L. Silk proteins: designs from nature with multipurpose utility and infinite future possibilities. Adv. Mater. 37, 2411256 (2025).
Malay, A. D., Craig, H. C., Chen, J., Oktaviani, N. A. & Numata, K. Complexity of spider dragline silk. Biomacromolecules 23, 1827–1840 (2022).
Numata, K. The biology of natural polymers accelerates and expands the science of biomacromolecules: a focus on structural proteins. Biomacromolecules 26, 1393–1403 (2025).
Finnigan, W. et al. The effect of terminal globular domains on the response of recombinant mini-spidroins to fiber spinning triggers. Sci. Rep. 10, 10671 (2020).
Chen, J. et al. Replicating shear-mediated self-assembly of spider silk through microfluidics. Nat. Commun. 15, 527 (2024).
Srisawat, P. et al. Engineered nanogel particles enhance the photoautotrophic biosynthesis of polyhydroxyalkanoate in marine photosynthetic bacteria. ACS Sustain. Chem. Eng. 10, 4133–4142 (2022).
Suzuki, M., Shirai, T., Morey-Yagi, S. R., Kondo, A. & Numata, K. Evaluation of nitrogen fixation in the marine purple photosynthetic bacterium Rhodovulum sulfidophilum under autotrophic and heterotrophic conditions. Sci. Rep. 15, 18344 (2025).
Fukuda, R., Ogawa, H., Nagata, T. & Koike, I. Direct determination of carbon and nitrogen contents of natural bacterial assemblages in marine environments. Appl. Environ. Microbiol. 64, 3352–3358 (1998).
Hochuli, E., Bannwarth, W., Döbeli, H., Gentz, R. & Stüber, D. Genetic approach to facilitate purification of recombinant proteins with a novel metal chelate adsorbent. Bio/Technol. 6, 1321–1325 (1988).
Betterle, N., Hidalgo Martinez, D. & Melis, A. Cyanobacterial production of biopharmaceutical and biotherapeutic proteins. Front. Plant Sci. 11, 237 (2020).
Huang, J. J., Heiniger, E. K., McKinlay, J. B. & Harwood, C. S. Production of hydrogen gas from light and the inorganic electron donor thiosulfate by Rhodopseudomonas palustris. Appl Environ. Microbiol 76, 7717–7722 (2010).
Gupta, D. et al. Photoferrotrophy and phototrophic extracellular electron uptake is common in the marine anoxygenic phototroph Rhodovulum sulfidophilum. ISME J. 15, 3384–3398 (2021).
Tanita, I. et al. Regionally Variable Responses of Nitrogen Fixation to Iron and Phosphorus Enrichment in the Pacific Ocean. J. Geophys. Res.: Biogeosci. 126, e2021JG006542 (2021).
Ren, J., Oh, S. H. & Na, D. Untranslated region engineering strategies for gene overexpression, fine-tuning, and dynamic regulation. J. Microbiol. 63, e2501033 (2025).
Chen, J. et al. Photosynthetic bacteria-based technology is a potential alternative to meet sustainable wastewater treatment requirement? Environ. Int. 137, 105417 (2020).
Yu, S., Xu, Y., Liang, C., Lou, W. & Peng, L. Spectral bands of incandescent lamp leading to variable productivity of purple bacteria biomass and microbial protein: Full is better than segmented. Sci. Total Environ. 823, 153736 (2022).
Maeda, I. Potential of phototrophic purple nonsulfur bacteria to fix nitrogen in rice fields. Microorganisms 10, 28 (2021).
Cerruti, M. et al. Enrichment and aggregation of purple non-sulfur bacteria in a mixed-culture sequencing-batch photobioreactor for biological nutrient removal from wastewater. Front. Bioeng. Biotechnol. 8, 557234 (2020).
Stephens, S., Mahadevan, R. & Allen, D. G. Engineering photosynthetic bioprocesses for sustainable chemical production: a review. Front. Bioeng. Biotechnol. 8, 610723 (2021).
Paerl, H. The cyanobacterial nitrogen fixation paradox in natural waters. F1000Res 6, 244 (2017).
Hirota, R. et al. A novel biocontainment strategy makes bacterial growth and survival dependent on phosphite. Sci. Rep. 7, 44748 (2017).
Zhang, C.-C., Zhou, C.-Z., Burnap, R. L. & Peng, L. Carbon/nitrogen metabolic balance: lessons from cyanobacteria. Trends Plant Sci. 23, 1116–1130 (2018).
Bucher, G. et al. Total organic carbon (TOC): a simple tool for assessing micro(nano)plastics and nanocellulose recovery during size-based fractionation. Anal. Bioanal. Chem. 417, 2983–2996 (2025).
Halewood, E. et al. Determination of dissolved organic carbon and total dissolved nitrogen in seawater using High Temperature Combustion Analysis. Front. Mar. Sci. 9, 1061646 (2022).
Acknowledgements
The authors thank Dr. Choon Pin Foong for technical support and expertise, as well as guidance in experimental design. This work was supported by Japan Science and Technology, COI-Next (Grant number JPMJPF2114) and ERATO (Grant number JPMJER1602). The funding agencies played no role in the study design; data collection, analysis, and interpretation; or writing of this manuscript.
Author information
Authors and Affiliations
Contributions
K.N. conceived the original research idea. M.S. and K.N. designed the experiments. M.S. conducted the experiments. M.S. and K.N. analyzed the results. M.S. wrote the manuscript. All the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
Keiji Numata is the CTO of Symbiobe Inc., a start-up company affiliated with Kyoto University.
Ethics approval and consent to participate
All genetic recombination experiments were performed in accordance with the guidelines and regulations of Kyoto University and were approved by the Kyoto University Recombinant DNA Experiment Safety Committee (Approval no. 250049).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suzuki, M., Numata, K. Production of the recombinant spider silk MaSp2 protein using the marine purple photosynthetic nonsulfur bacterium Rhodovulum sulfidophilum under autotrophic conditions. NPG Asia Mater 18, 13 (2026). https://doi.org/10.1038/s41427-026-00638-7
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41427-026-00638-7