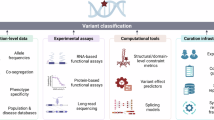
Abstract
Over the past three decades, Hereditary Cancer Testing (HCT) has evolved from single gene assays into multigene panel testing (MGPT), which allows for the screening of all known hereditary cancer genes in a single assay. MGPT is currently the standard approach for clinical HCT. However, with decreasing sequencing costs and increased instrument throughput, the scalability of exome sequencing (ES) and genome sequencing (GS) for HCT indications is becoming more viable. These methods provide broader insights into the coding exons and/or the entire genome, respectively. ES/GS data can also be reanalyzed to identify variants in novel genes that were not characterized at the time of initial testing, or to support research efforts aimed at uncovering additional associations between germline variants and cancer predisposition. Additionally, the emerging use of long-read sequencing (LRS) is noteworthy, enabling improved variant detection compared to short-read sequencing, especially for complex/structural variants and variation in difficult-to-sequence or paralogous regions in genes such as PMS2. This has the potential to increase the accuracy of HCT, reduce the turnaround time, find previously unidentifiable cancer risk variants, and ultimately increase the diagnostic yield. This article provides a comprehensive summary of the sequencing approaches used in HCT, discussing their strengths and limitations. We also highlight the added value of complementing DNA-only testing with RNA and tumor sequencing. Furthermore, we explore LRS-based approaches and discuss opportunities for their implementation in routine genetic testing for hereditary cancer.
Similar content being viewed by others
Introduction
The field of clinical cancer genetics has been evolving ever since the discovery and characterization of the first cancer predisposition genes (CPG) in the 80s–90s [1]. In the past, Hereditary Cancer Testing (HCT) consisted of single gene assays based primarily on Sanger sequencing to interrogate the protein coding regions and intron/exon boundaries of CPG such as APC, BRCA1/2 and the mismatch repair (MMR) genes MLH1 and MSH2 [2, 3].
With the emergence of Next Generation Sequencing (NGS, i.e., second generation sequencing) during the late 2000s, sequencing cost and scalability improved drastically, allowing the development of assays covering multiple CPG simultaneously. In parallel, researchers have continued uncovering novel gene-disease associations raising the number of bona fide CPG to >100 genes [4]. Starting from the mid-2010s, multigene panel testing (MGPT) became the standard HCT approach and is widely used by molecular diagnostic laboratories around the world, who continue to expand and refine their gene menu following novel gene discoveries and/or new evidence supporting or refuting previously established associations [5]. Moreover, some clinical laboratories are complementing DNA-only MGPT with RNA testing to provide additional insights into the splicing effects of germline variation, and its association with increased cancer risk [6]. Similarly, tumor-normal sequencing allows the robust identification of somatic alterations to guide treatment decisions and may contribute additional information in the evaluation of germline variants, particularly variants of uncertain significance (VUS) [7, 8].
The falling cost of sequencing has also impacted Exome Sequencing (ES) and Genome Sequencing (GS), with modern ultra-high-throughput instruments capable of generating >1000 exomes or >100 genomes per run, at a considerably lower cost compared to a decade ago. For HCT, this could potentially translate into ES/GS replacing MGPT, as this creates an opportunity for data reanalysis over time, and powers research studies evaluating non-coding variation, polygenic risk scores (PRS) and novel CPG discovery [9, 10].
Alongside the evolution of NGS, Long Read Sequencing (LRS, i.e., third generation sequencing) exhibited remarkable improvements over the past five years, resulting in lower error rates, cost, and higher instrument throughput. Compared to short-read sequencing (SRS), LRS detects over 50% more structural variants (SV) and can directly identify epigenetic marks like 5-methylcytosine that influence gene expression [11]. It also enables phasing of variants in recessive cancer syndromes. These capabilities position LRS as a valuable tool in HCT—initially as an orthogonal method for complex regions like PMS2, and potentially as a future first-tier sequencing approach.
DNA multigene panel testing
DNA MGPT were introduced commercially for hereditary cancer in 2012; however, widespread adoption did not begin until 2013 following the U.S. Supreme Court’s ruling to reverse the patent held on BRCA1/2. This decision enabled laboratories to offer comprehensive HCT via a single assay. In a study of individuals with breast and ovarian cancer, the proportion of HCT orders attributable to MGPT increased from ~40% at the end of 2013 to >90% by 2019 [12]. The average number of genes tested has also increased over time, with a trend toward larger pan-cancer panels over smaller, phenotype-specific panels [12, 13].
The clinical advantages of a DNA MGPT approach have been well established. Several large MGPT studies have demonstrated a significant increase in diagnostic yield over single gene/syndrome testing across cancer types. The diagnostic yield for MGPT is highly dependent upon the number of genes tested, indication for testing, and cohort selection. Initial results from commercial laboratories offering small-to-mid size panels (i.e., 25–34 genes) reported an overall diagnostic yield of 7–9% [13,14,15]. Subsequent studies of unselected cancer cohorts tested with larger panels (i.e., >75 genes) reported an overall diagnostic yield in the range of 13–17%, with the highest diagnostic yield observed among individuals with ovarian and pancreatic cancers [16, 17]. Other benefits of DNA MGPT include the identification of individuals with pathogenic/likely pathogenic variants (PV) in multiple CPG, in genes outside the initial testing indication, and refinement of penetrance estimates as a result of genotype-first characterization [18]. Based on these clinical advantages, expert guidelines recommend a MGPT approach to CPG testing [19].
Commercial laboratories generally perform MGPT using short-read targeted sequencing panels. The target enrichment of CPG is achieved by biotinylated probes that are designed to bind and capture regions of interest within CPG, primarily covering coding exons and intron-exon boundaries. Laboratories can also supplement their probe designs to capture regions of well-known deep intronic PV, including small variants and/or SV breakpoints [20]. Copy-number variants (CNV), mainly deletions and duplications, are inferred primarily from variations in expected read depth but may require orthogonal validation methods such as Multiplex Ligation-dependent Probe Amplification (MLPA) or microarrays. With the exception of exon-level deletions, CNVs can be reliably detected if read depth is sufficient. Of note, CNV calling algorithms are also evolving and display variable performance, with tools like GATK-gCNV showing high performance in MGPT data across CNV of different lengths [21].
The operational advantage of MGPT is cost effectiveness and scalability, as it allows laboratories to run hundreds-thousands of samples per day at a reduced cost compared to ES or GS (Table 1). MGPT also offers a higher sequencing depth for the desired targets, typically around 500x, which facilitates germline variant identification, as well as detection of mosaicism or clonal hematopoiesis of indeterminate potential [22].
While MGPT has offered significant advantages over single-gene testing with respect to efficiency and identification of germline cancer predisposition, there are opportunities for improvement. One of the major limitations of targeted SR panels is the fixed design and subsequent need to regularly update the panel content. Professional testing guidelines are constantly evolving as novel CPG are discovered and established gene-disease relationships are refined. This, in turn, prompts laboratories to update their panels [19, 20]. While removing content could easily be achieved by bioinformatic masking, adding new content requires altering the probe design and necessitates clinical validation of the assay in each iteration, to ensure optimal performance in accordance with strict regulations of laboratory developed tests, such as the CLIA requirements in the US, and the IVDR in Europe.
There are also certain types of variants that are challenging to detect with targeted SR panels (Table 1). There are limitations in the ability to detect SV, especially when the breakpoints of these events are located outside the captured regions. For instance, copy-neutral events such as inversions and balanced translocations could go undetected if the breakpoints are not covered. Similarly, the inherent fluctuations of coverage uniformity in targeted sequencing might complicate the detection of smaller CNVs [23]. Another limitation of these assays is the accurate detection of variation in genes complicated by pseudogenes due to the shortcomings of SRS, prompting laboratories to validate suspicious findings with MLPA and long-range PCR [24].
RNA sequencing as a complement TO HCT
RNA testing was traditionally performed with RT-PCR using primers overlapping exons followed by Sanger sequencing. The emergence of a more scalable approach - targeted RNA sequencing (RNA-seq) with hybridization capture – allows for the evaluation of multiple CPG in parallel. The utility of RNA-seq in HCT resides in providing qualitative and quantitative evidence to identify and characterize abnormal splicing events and aberrant expression levels [25]. Unlike other putative loss-of-function variants, most splicing variants are classified as VUS without additional functional evidence [26].
As a complement to HCT, RNA-seq provides added clinical value by facilitating the interpretation of splicing VUS and the detection of deep intronic splicing PV in regions that are not typically included in MGPT. This ultimately results in improved diagnostic resolution, along with a higher molecular diagnostic rate [6]. The first diagnostic study in a large consecutive series included >43,000 individuals undergoing paired DNA and RNA testing of 1–18 genes. Horton et al. reported a variant classification impact based on RNA evidence in 1.3% individuals (n = 547), including medically significant upgrades (0.2%, n = 70), novel deep intronic PV (0.1%, n = 27) and VUS downgrades (0.7%, n = 305) [27]. As many VUS are recurrent in the tested populations, these updated classifications were subsequently applied to a larger series of >500,000 previously tested individuals, yielding an impact on 4.5% of individuals (n = 22,963) with DNA-only testing, and where reclassifications were made in 33.1% of individuals (n = 7602) resulting in revised clinical reports sent to ordering clinicians. In another large laboratory-based cohort study involving paired DNA/RNA testing of 63 genes in >20,000 patients, 6.3% of individuals (n = 1279) had a non-VUS variant that would have been classified as a VUS without RNA evidence, including 0.4% with PV (n = 75), and 5.9% with benign or likely benign variants (n = 1204). Additionally, 0.2% (n = 40) of individuals had splicing PV that otherwise would not have been detected, as the variants were located outside the reportable range for the targeted panel [28].
RNA analysis also highlights the importance of assessing naturally occurring alternative splicing for clinical evaluation of variants in disease-causing genes [29]. For example, an analysis performed by the ENIGMA (Evidence-based Network for the Interpretation of Germline Mutant Alleles) Consortium indicated that BRCA1 c.[594-2 A > C;641 A > G] (NM_007294), previously described to cause exon 10 skipping (a truncating alteration), displays characteristics inconsistent with those of a high risk pathogenic BRCA1 variant [30]. Their RNA analysis of human samples and splicing minigenes showed that exon 10 skipping results from c.641 A > G altering splicing regulation, not from c.594-2 A > C disrupting the acceptor site. RNA assays revealed that while most transcripts were truncating, 20–30% were in-frame (Δ9,10) and predicted to encode a functional BRCA1 protein, indicating it should not be considered a high-risk pathogenic variant.
Another way to leverage RNA-Seq data is by performing allele specific expression analyses. Allelic expression levels can be altered by different mechanisms such as gene deletions, epigenetic silencing, cis-regulatory elements or loss-of-function variants that trigger non-sense mediated decay. These alterations could result in monoallelic expression, resulting from the complete silencing of one allele, or in allelic imbalances, which clinical significance is more challenging to determine depending on dosage sensitivity and the abundance of rescue transcripts that maintain sufficient function [31, 32].
The Clinical Genome Resource (ClinGen) Variant Classification Working Group has developed updated guidelines to clarify how the American College of Medical Genetics and Genomics (ACMG)/Association for Molecular Pathology (AMP) codes should be applied to splicing evidence [26]. The continued refinement and integration of these recommendations aim to standardize the interpretation of RNA and in-silico data, improving consistency in variant classification and expanding the clinical utility of RNA sequencing.
Short read exome and genome sequencing
The cost to sequence a short read genome has dramatically declined over time, with sequencing costs expected to drop to $100–200/genome, consequently leading to a lower cost for ES. Similar to MGPT, the enrichment of all coding regions in ES is achieved by hybridization capture, whereas GS libraries are typically PCR-free and require less library preparation time. Required DNA amounts for ES and GS have also drastically decreased over time. These assays now require as low as 100–250 ng of gDNA, making them compatible with challenging specimen types that yield very limited amounts of gDNA, such as dried blood spots. Notably, there are now multi-omic GS assays that capture methylation data concurrently with genetic variation, leading to the detection of 5-methylcytosine and 5-hydroxymethylcytosine, such as Biomodal’s 6-base sequencing (Cambridge, UK), and Illumina’s 5-base sequencing (San Diego, CA, USA). These assays simplify the laboratory procedure to read the genome and epigenome without requiring multiple library preparation workflows, and without compromising the identification of C-to-T genetic variants [33].
As shown in Table 1, GS offers additional advantages over ES. The expanded coverage beyond exonic regions enables the identification of deep intronic variants and allows the delineation of SV breakpoints. For instance, there are two reported PALB2 (NM_024675) Ex13 duplications, 4.8 kb and 13.8 kb, where only the smaller leads to abnormal RNA splicing and is of clinical relevance [34, 35]. MGPT or ES requires additional orthogonal testing to disambiguate the breakpoints in the absence of RNA testing. Additionally, GS allows the identification of non-coding alterations that could have an effect on splicing or gene regulation. While these variants are challenging to classify, the emergence of powerful new AI tools such as SpliceAI, PromoterAI and AlphaGenome in conjunction with RNA testing is expected to facilitate their interpretation [36,37,38]. Moreover, GS offers more uniform coverage compared to ES, especially in GC-rich regions, which facilitates CNV identification [39]. GS data can also be leveraged for PRS analysis, especially in patients of non-European ancestry. Notably, existing PRS were based on genome-wide association studies primarily from the European population, and consequently not suitable for clinical use in different populations [9]. In this context, the adoption of GS will guide PRS recalculation in underrepresented populations leading to more equitable and accurate cancer risk predictions.
The extent of genomic regions analyzed by MGPT, ES, and GS leads to substantial differences in average sequencing depth, typically around 500x for MGPT, 100x for ES, and 30x for GS, though these values can vary between laboratories. This variability directly impacts analytical performance, especially for read-depth-based CNV detection, where identifying small events that span only one or a few exons remains challenging [23]. Using 50x GS data, De La Vega et al. demonstrated that genome-wide CNV detection performance varies considerably depending on the CNV caller used, with sensitivity for smaller events (1–5 kb) generally lower, and duplications being detected less reliably than deletions [40]. Notably, performance improved when analyses focused on a virtual panel covering exons from 184 clinically relevant genes, including 89 CPG, after applying custom CNV filters and removing common artifacts. Ultimately, further research is needed to enable a direct comparison of the analytical performance of MGPT versus ES and GS for HCT, which will help clarify the benefits and trade-offs associated with each approach.
Interestingly, some laboratories are now creating hybrid GS-ES assays, to combine the higher sequencing depth of ES with the genome wide coverage of GS in a cost-effective manner. Boltz TA et al. devised and benchmarked the performance of a blended exome (30–40x mean depth) and low-pass genome (1-4x mean depth) sequencing assay against matched conventional ES, GS, or array data. The authors reported high recall and positive predictive value when calling coding CNV affecting ≥3 exons and high concordance for common variant imputation [41].
One of the main clinical advantages of ES and GS over MGPT is the opportunity for data reanalysis over time. This minimizes the need for sequential testing, as a patient’s sequencing data can be reanalyzed as phenotype evolves and as novel CPG and gene-disease relationships are discovered. ES and GS are the recommended first-tier testing approaches for patients diagnosed with rare disease. In contrast, the use of ES/GS in HCT has, for the most part, been limited to the research setting. Several CPG have been identified in studies leveraging ES/GS, such as POLE, POLD1 and NTHL1 [42, 43]. More recently, MBD4 and RPS20 have also been added to guideline-based testing for colorectal cancer and polyposis [44]. As to guideline-based breast cancer testing, no novel CPG additions were noted, highlighting the variable success of gene discovery studies based on ES/GS depending on cancer type [45].
The clinical approach to HCT has traditionally prioritized clinical actionability over diagnostic resolution. SR MGPT currently offers a pragmatic solution to obtain high quality sequencing data for the coding regions of clinically actionable cancer predisposition genes. Comparable analytic sensitivity needs to be established before transitioning HCT to ES/GS-based assays to ensure quality is not compromised.
Other aspects that should also be considered include the larger data footprint and increased computational requirements for storage, reprocessing, and reanalysis of ES/GS data. Importantly, this also raises the possibility of uncovering incidental findings unrelated to oncology, such as variants associated with severe rare genetic disorders.
The utility of SOMATIC testing
Tumor-only testing is conducted on patients primarily to identify biomarkers to guide treatment selection and predict response. While it is possible to infer germline variants from somatic testing by relying on variant allele frequencies, confirmatory germline testing is recommended as allelic frequencies of true germline variants can be skewed outside the expected range (40–60%) due to several factors including tumor sample purity, somatic CNV, coverage, sequencing depth, and variant location [46]. Consequently, tumor-only sequencing is known to miss a significant number of germline variants [47].
To overcome these limitations, paired germline-somatic testing is conducted to unequivocally identify both germline and somatic variation, enabling the detection of somatic second hits, loss of heterozygosity, microsatellite instability, and tumor mutational burden [48]. In a study by Salvador MU et al., approximately 50% of patients referred for Lynch Syndrome (LS) testing, with abnormal immunohistochemistry results or microsatellite instability, were found to be double somatic mutation carriers or showing somatic MLH1 promoter hypermethylation [8]. The authors presented examples where loss of MMR staining was explained by double somatic MMR mutations while a germline PV was identified in a different MMR gene. They further proposed that somatic tumor data may serve as supporting evidence for variant classification; however, this approach has not yet been fully integrated into established testing guidelines, including those of the ACMG/AMP or the International Society for Gastrointestinal Hereditary Tumors (InSiGHT).
In this context, somatic mutational signatures could also prove useful, as they originate with high specificity from defects in certain CPG. For example, signature SBS36 and SBS30 emerge as a result of biallelic inactivation of MUTYH and NTHL1, respectively. Likewise, POLE/POLD1 defects result in SBS10 signatures and hypermutated tumors [49]. Identifying these signatures in patient tumors undergoing HCT could guide VUS reclassification or point to potentially undetected germline PV.
For research studies, large matched normal-tumor datasets are leveraged to uncover novel associations, or to better understand the contribution of germline PV to tumorigenesis. For instance, paired data from >34,000 patients recently suggested a potential role of NBN as a pan-cancer susceptibility gene [50]. Similarly, data from >17,000 patients revealed that 27% of cancers diagnosed in patients with high penetrance germline PV were not dependent on the contribution of the germline allele for tumorigenesis and occurred in tumor lineages outside the phenotypic spectrum of the corresponding CPG [51]. This aspect is particularly relevant when considering therapeutic implications, as not all cancers diagnosed in hereditary cancer patients will benefit from targeted therapies, highlighting the added clinical value of paired normal-tumor testing. In this setting, tumor retesting may reveal additional therapeutic opportunities, particularly when considering the accumulation of somatic alterations as the result of tumor evolution.
Tumor molecular testing is most commonly performed on formalin‑fixed, paraffin‑embedded tissue due to its cost‑effectiveness and suitability for long‑term preservation and storage. However, formalin fixation induces DNA fragmentation and chemical damage, which can introduce sequencing errors and artifacts [52]. Although mitigation strategies have been developed, such as S1 nuclease treatment, higher‑quality sequencing data are obtained when fresh or fresh‑frozen tissue is used as the input material. This is particularly important for LRS, as DNA integrity is a key determinant of average read length.
Long read sequencing as an emerging approach to improve the accuracy of HCT
LRS: technology
LRS differs from SRS primarily on the length of reads, ranging from a few kilobases to 100 s of kb, compared to ~150 bp in SRS. There are two native LRS technologies developed by Pacific Biosciences of California (PacBio; Menlo Park, CA, USA) and Oxford Nanopore Technologies (ONT; Oxford, UK). PacBio sequencing relies on circularizing DNA fragments and real time amplification with light signal detection as bases are incorporated. The polymerase performs several passes over the inserts generating multiple subreads that are then collapsed in silico into a consensus high-fidelity read. ONT sequencing relies on passing DNA libraries through nanopores to directly measure electrical signal disruptions, called squiggles, that are specific to each base. In both technologies, the kinetics of base incorporation or squiggles, i.e., the time it takes to register consecutive signals, is leveraged to robustly identify epigenetic modifications. In short, PacBio sequencing offers higher accuracy, similar to SRS, and read lengths up to ~20 kb. ONT allows longer reads reaching >10–100 s kb but suffers from higher sequencing error rates. Both technologies are continually improving through changes of reagent chemistry, nanopore/polymerase bioengineering, and/or base calling algorithm updates.
Over 50% of the human genome consists of repetitive sequence, such as mobile elements (DNA/RNA transposons) and tandem repeats, and about 7% consists of segmental duplications [53, 54]. These regions are difficult to sequence and map accurately to the right location in the reference genome with ~150 bp reads, primarily due to the high sequence homology or the repetitive/GC-rich sequence context. Additionally, repetitive regions are particularly prone to damage during DNA replication and recombination, making them susceptible to drive structural aberrations [55]. The longer reads are superior in resolving these challenging regions while also allowing robust SV identification as more sequence identity is available to delineate the breakpoints compared to SRS (Table 1). In fact, studies leveraging long-read WGS (LR-WGS) have consistently identified ~25,000 unique SV/genome compared to ~11,000 in short-read WGS data [11]. Based on these observations, SRS-based assays could easily miss SV in CPG, providing an opportunity to increase diagnostic yield.
In parallel, specialized Bioinformatic algorithms specifically tailored to LRS are also emerging and evolving. Notably, the Paraphase tool has been developed to resolve medically challenging genes such as PMS2/PMS2CL, facilitating the disambiguation of variation with clinical relevance without relying on orthogonal confirmation with MLPA or long-range PCR sequencing [56]. Similarly, improvements to SV/CNV calling were made, with tools like Sawfish and Sniffles2 resulting in better accuracy and speed and enabling the detection of mosaic events [57, 58].
LRS: DNA-seq
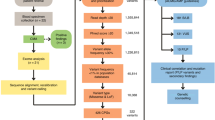
Over the past five years, >20 research studies have leveraged LR DNA sequencing in cases with known or suspected hereditary cancer conditions (Table 2). These studies utilized different sequencing protocols including LR-WGS and targeted LRS, the latter being more cost effective and scalable thanks to the higher multiplexing. Targeted LRS approaches varied from PCR amplicon sequencing, hybridization capture, CRISPR-Cas9 excision, or ONT adaptive sampling (ONT-AS), each having different strengths and limitations that we summarized in Fig. 1. Notably, ONT-AS consists in the real-time enrichment of pre-defined genomic regions of interest, and rejection of off-targets. As DNA strands pass through the nanopore, the first few hundred bases are read, and a decision is made to continue sequencing or rejecting the strand, resulting in increased sequencing depth in the desired targets (usually 20–60x with long reads), and low-pass genome-wide coverage ( ≤ 5x with shorter reads). Interestingly, the off-target reads can be leveraged for the imputation of single nucleotide polymorphisms and showed strong concordance (99.8%) with SRS data in a recent study by Nakamura et al., enabling PRS estimation [59].
Simplified workflow description, strengths and limitations. Abbreviations: BrdU: 5-bromo-2’-deoxyuridine, iDMRs: imprinted differentially methylated regions, lpWGS: low-pass WGS, m6A-MTase: N6-methyladenine methyl-transferase, ONT: Oxford Nanopore Technologies, ROI: region of interest, Seq: Sequencing.
Validation studies aimed at using LRS primarily to confirm and fully characterize SV calls that were previously identified using other approaches such as SRS, MLPA, or Optical Genome Mapping. By using LR-WGS, Dixon et al. were able to fully resolve the breakpoints of 13/14 SV from 19 patients with sizes varying from 510 bp to 108 kb, leading to the identification of allelic heterogeneity in BRCA1 Ex1-2 deletions (6.6-37 kb; NM_007294) and CHEK2 Ex9-10 deletions (5.4 kb or 6.2 kb; NM_007194) [60]. A common 1.26 Mb haplotype was identified among carriers of the 5.4 kb CHEK2 deletion, a founder in the Eastern European population, and a shared 1.08 Mb haplotype was found in three unrelated carriers of the British founder BRCA1 Ex6 duplication (6.1 kb). Similarly, in an unexplained familial adenomatous polyposis family, Baumann et al. first identified evidence of an intronic insertion in APC intron 7 with SRS, and leveraged LR-WGS to fully resolve the event as a 6.1 kb LINE-1 insertion that resulted in abnormal RNA splicing [61]. In another study, Watson et al. used long-range PCR and LRS on one patient to confirm a 4 kb RB1 Ex23 deletion (NM_000321), initially identified by MLPA, uncovering an additional 85 bp Ex24 tandem duplication that was previously missed [62].
Other studies leveraged LRS to explore missing heritability in unsolved cases with a clinical suspicion of hereditary cancer. Gulsuner et al. utilized ONT-AS in 120 unsolved kindreds affected by hereditary breast, ovarian, pancreatic, and/or prostate cancer to explore the frequency of rare deep intronic variants. The authors identified 7 PV in BRCA1, PALB2 and ATM in 8/120 (6%) families. LR RNA-seq revealed aberrant transcripts for all 7 variants and consisted of pseudoexon inclusion resulting in premature truncation [63]. Likewise, Paske et al. utilized long-range PCR and LRS to interrogate the entire coding and non-coding regions of the MMR genes in a series of 32 patients suspected to have LS. Their analysis revealed 6/32 (19%) deep intronic MLH1/MSH2 PV carriers, where evidence for abnormal splicing was also provided [64].
LRS: RNA-seq
Compared to SRS, LR RNA-Seq provides multiple advantages: (1) It allows the sequencing of entire transcripts and enables the discovery of novel ones that could be associated with disease; (2) It allows the quantification of transcript levels and the transcript-aware characterization of multiple splicing events, which could have implications on variant classification; (3) in the presence of heterozygous exonic markers, it enables allele specific quantification of transcripts. Taken altogether, this provides a higher resolution when analyzing RNA data and facilitates VUS interpretation.
Schwenk et al. developed the capture and ultradeep long-read RNA seq approach (CAPLRseq), covering 123 CPG. Focusing on the diagnosis of LS, the authors validated the assay in 17 cases with PV in the MMR genes [65]. Eight patients were carriers of variants with previously available RNA data from PCR and Sanger sequencing, while nine patients were carriers of PV without previously available RNA data. CAPLRseq data was concordant with abnormal splicing and allowed the accurate identification and quantification of transcript levels. Interestingly, one of the variants was a MLH1-DCLK3 inversion that resulted in fusion transcripts that are challenging to resolve with SRS, and another consisted in constitutional methylation (hereinafter, epimutation) affecting MLH1, where CAPLRseq expression levels were consistent with MLH1 monoallelic expression. The authors also applied this method to resolve two VUS in PMS2 and MSH6, successfully leading to reclassifications.
Similarly, Aucouturier et al. developed a LR-RNA target enrichment approach capturing 28 CPG compatible with both PacBio and ONT platforms. They also developed a computational framework named SOSTAR (iSofOrmS annoTAtoR) allowing the processing of raw data and the automatic annotation of transcripts in a human readable nomenclature [66]. The authors validated this approach in eight samples including four negative controls from healthy donors, two positive controls with BRCA1 spliceogenic variants, and two relatives with abnormal BRCA1 splice junctions suspected to have a deep intronic cryptic event. The negative controls recapitulated known BRCA1/2 alternative splicing events and allowed a comprehensive look into transcript architecture, including the identification of novel transcripts. In positive controls, both LRS technologies correctly identified the expected aberrant BRCA1 transcripts. For the missing heritability patients, it led to the identification of a 900 bp pseudoexon inclusion caused by a 2.7 kb SVA (SINE-VNTR-Alu) insertion leading to its classification as a PV. Noteworthy, the authors described differences in the ability of the two LRS technologies in capturing longer transcripts, with ONT being superior to PacBio. This is informative to future studies focusing on CPG with large cDNAs such as ATM and BRCA2, where selecting one technology over the other is more appropriate, and where RNA sample integrity is critical, underscoring the importance of specimen type and RNA isolation methods.
LRS: Epigenetics
When used on native DNA molecules, LRS allows the direct detection of epigenetic modifications such as 5-methylcytosines, creating an opportunity to directly identify these marks on PCR-free LRS data. Epimutations are a well-known mechanism for cancer predisposition, albeit understudied and infrequently tested for in the clinical setting [67]. Notably, MLH1 epimutations are known to be associated with LS. LRS can directly identify allele specific MLH1 epimutation carriers, however, certain individuals exhibit mosaic levels of methylation ( ≤ 10%), making them harder to detect without sensitive molecular assays [59, 68]. Such events might not be detected by current PCR-free LRS approaches (LR-WGS, ONT Adaptive Sampling or CRISPR/Cas9 excision) due to the limited sequencing depth. Beyond promoter methylation in CPG, LRS could be leveraged to study epimutation signatures, such as the known episignature in Beckwith-Wiedemann syndrome, consisting in abnormal methylation patterns in the imprinted 11p15.5 region, and predisposing to embryonal tumors [69].
Additionally, methylation data from LRS can be leveraged to assign a parent-of-origin in carriers of PV without parental testing, positively impacting clinical management by simplifying cascade testing, improving variant curation and penetrance estimation. Akbari et al. described an approach based on short-read single-cell template strand sequencing (Strand-seq) to create sparse chromosome haplotypes serving as scaffolds for LR-WGS to identify imprinted differentially methylated regions, resulting in chromosome-length haplotypes and phased variant calls assigned to each parent [70] (Fig. 1).
Finally, Fiber-Seq is a recently developed LRS-based approach to study chromatin accessibility, transcription factor and nucleosome occupancy on multi kilobase chromatin fibers, enabled by the ability of LRS to detect additional epigenetic marks such as m6A (N2-methyladenine) [71] (Fig. 1). It provides synchronous genome and epigenome profiling, shedding light on the contribution of rare non-coding variation to human disease.
Conclusion
Multigene panels have been the standard sequencing approach for clinical HCT for the past decade, representing a cost-effective strategy to interrogate multiple CPG simultaneously, without compromising on analytical performance [13, 19]. Despite its convenience compared to Sanger sequencing, molecular diagnostic laboratories often rely on orthogonal confirmation assays to validate observations, especially when it comes to deletions/duplications or variants in regions complicated by pseudogenes [24]. This translates into longer turnaround times, complex laboratory workflows relying on multiple complementary assays, the need to maintain personnel competency and validated equipment, and in cases of complex alterations, uncertainty in fully characterizing DNA alterations. Additionally, requirements for assay validation limit the ability to rapidly update panels as new disease genes are discovered. While these hurdles only affect a small percentage of cases that receive clinical testing, they have a high impact on clinical lab operations resulting in an increased investment of resources.
The addition of RNA-seq to DNA-only MGPT provides direct functional evidence that aids in the interpretation of DNA variants, resulting in the reduction of VUS rates. Additionally, the cascade effect of VUS reclassification was shown to impact a larger number of patients who received DNA-only testing [27]. Similarly, the addition of tumor testing enables the identification of somatic second hits, double somatic mutation carriers, and hallmarks associated with defects in CPG, such as mutational signatures or microsatellite instability [8, 49].
The dramatic decrease of ES/GS cost over time, and the development of ultra-high-throughput sequencers could trigger a paradigm shift in the future, creating an opportunity for data reanalysis and fueling research studies. However, there are important considerations prior to this potential transition. First, the lower sequencing depth of these approaches could reduce sensitivity to detect certain alterations such as CNV. Second, the cost and time to develop and validate these assays and insurance reimbursement. And lastly, the underlying short-read sequencing technology has well-known shortcomings in the detection of structural variation and variation in complex regions [11, 24].
In this context, the emergence of LRS approaches could be transformative both in the clinical and translational research settings. On one hand, it will allow clinical laboratories to consolidate and simplify orthogonal confirmation workflows, improving testing accuracy and potentially resulting in shorter reporting times for ordering clinicians. On the other hand, it opens several research fronts that could improve our understanding of the association of germline variants and increased cancer susceptibility, especially when it comes to non-coding variation and epigenetic modifications.
It is worth noting that PRS studies will benefit from the growing adoption of GS, blended ES/GS assays, and adaptive LRS, enabling the formulation of population specific PRS for individuals of non-European ancestry and ultimately advancing equity in genetic research for populations that have been historically underrepresented [9].
The clinical utility of LRS for HCT remains to be fully delineated, as the number of studies remains scarce and focused on small, highly ascertained cohorts of cancer patients (Table 2). As LRS adoption continues to increase, future studies in larger series, especially in unselected cohorts, will be pivotal in estimating its real added clinical value. Encouraging data from rare disease research has already demonstrated an increase in diagnostic rate and shorter turnaround time enabled by this technology [72]. Importantly, compared to SRS, the cost of LRS remains high and instrument throughput is considerably lower. Optimal DNA and RNA integrity and quantity requirements are also more stringent for LRS, prompting laboratories to devise alternative protocols for DNA/RNA isolation and quality control. Additionally, downstream bioinformatic workflows and tertiary analysis solutions are still maturing, and currently available catalogs of SV, tandem repeats and epigenetic modifications are emerging but remain limited to provide accurate frequency estimates of these events in the general population to inform filtering, prioritization and interpretation.
Ultimately, integrating emerging technologies into hereditary cancer testing will require careful validation and collaboration across clinical laboratories, researchers, and professional organizations. As sequencing technologies and analytical tools continue to evolve, the boundary between research and clinical testing will become increasingly seamless. Strategic investment in LRS development, data integration, and evidence generation will determine how quickly these innovations translate into tangible improvements in diagnostic precision and the future of hereditary cancer testing.
References
Rahman N. Realising the promise of cancer predisposition genes. Nature. 2014;505:302.
Gima L, Solomon I, Hampel H. The evolution of genetic testing from focused testing to panel testing and from patient focused to population testing: are we there yet? Clin Colon Rectal Surg. 2023;37:133.
Kolata G. Breaking ranks, lab offers test to assess risk of breast cancer. N Y Times Web 1996.
Garutti M, Foffano L, Mazzeo R, Michelotti A, Da Ros L, Viel A, et al. Hereditary cancer syndromes: a comprehensive review with a visual tool. Genes (Basel). 2023;14:1025.
Herrera-Mullar J, Horton C, Weaver A, Towne M, Huang JM, VanNoy GE, et al. Understanding how gene-disease relationships can impact clinical utility: adaptations and challenges in hereditary cancer testing. Genome Med. 2025;17:1–12.
Karam R, Conner B, Laduca H, McGoldrick K, Krempely K, Richardson ME et al. Assessment of diagnostic outcomes of RNA genetic testing for hereditary cancer. JAMA Netw Open. 2019;2. https://doi.org/10.1001/JAMANETWORKOPEN.2019.13900.
Zehir A, Benayed R, Shah RH, Syed A, Middha S, Kim HR, et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat Med. 2017;23:703.
Salvador MU, Truelson MRF, Mason C, Souders B, LaDuca H, Dougall B, et al. Comprehensive paired tumor/germline testing for lynch syndrome: bringing resolution to the diagnostic process. J Clin Oncol. 2019;37:647–57.
Roberts E, Howell S, Evans DG. Polygenic risk scores and breast cancer risk prediction. Breast Off J Eur Soc Mastol. 2023;67:71.
Palles C, West HD, Chew E, Galavotti S, Flensburg C, Grolleman JE, et al. Germline MBD4 deficiency causes a multi-tumor predisposition syndrome. Am J Hum Genet. 2022;109:953–60.
Zhao X, Collins RL, Lee WP, Weber AM, Jun Y, Zhu Q, et al. Expectations and blind spots for structural variation detection from long-read assemblies and short-read genome sequencing technologies. Am J Hum Genet. 2021;108:919–28.
Kurian AW, Kurian AW, Abrahamse P, Furgal A, Ward KC, Hamilton AS, et al. Germline genetic testing after cancer diagnosis. JAMA. 2023;330:43.
LaDuca H, Polley EC, Yussuf A, Hoang L, Gutierrez S, Hart SN, et al. A clinical guide to hereditary cancer panel testing: evaluation of gene-specific cancer associations and sensitivity of genetic testing criteria in a cohort of 165,000 high-risk patients. Genet Med. 2020;22:407–15.
Neben CL, Zimmer AD, Stedden W, van den Akker J, O’Connor R, Chan RC, et al. Multi-gene panel testing of 23,179 individuals for hereditary cancer risk identifies pathogenic variant carriers missed by current genetic testing guidelines. J Mol Diagnost. 2019;21:646–57.
Rosenthal ET, Bernhisel R, Brown K, Kidd J, Manley S. Clinical testing with a panel of 25 genes associated with increased cancer risk results in a significant increase in clinically significant findings across a broad range of cancer histories. Cancer Genet. 2017;218–219:58–68.
Samadder NJ, Riegert-Johnson D, Boardman L, Rhodes D, Wick M, Okuno S, et al. Comparison of universal genetic testing vs guideline-directed targeted testing for patients with hereditary cancer syndrome. JAMA Oncol. 2021;7:230–7.
Ceyhan-Birsoy O, Jayakumaran G, Kemel Y, Misyura M, Aypar U, Jairam S, et al. Diagnostic yield and clinical relevance of expanded genetic testing for cancer patients. Genome Med. 2022;14:92.
Roberts ME, Ranola JMO, Marshall ML, Susswein LR, Graceffo S, Bohnert K, et al. Comparison of CDH1 penetrance estimates in clinically ascertained families vs families ascertained for multiple gastric cancers. JAMA Oncol. 2019;5:1325–31.
Tung N, Ricker C, Messersmith H, Balmaña J, Domchek S, Stoffel EM, et al. Selection of germline genetic testing panels in patients with cancer: ASCO guideline. J Clin Oncol. 2024;42:2599–615.
van den Akker J, Hon L, Ondov A, Mahkovec Z, O’Connor R, Chan RC, et al. Intronic breakpoint signatures enhance detection and characterization of clinically relevant germline structural variants. J Mol Diagnost. 2021;23:612–29.
Munté E, Roca C, Valle J, Feliubadaló L, Pineda M, Gel B, et al. Detection of germline CNVs from gene panel data: benchmarking the state of the art. Brief Bioinform. 2024;26:bbae645.
Rofes P, Castillo-Manzano C, Menéndez M, Teulé Á, Iglesias S, Munté E, et al. TP53 germline testing and hereditary cancer: how somatic events and clinical criteria affect variant detection rate. Genome Med. 2025;17:3.
Babadi M, Fu JM, Lee SK, Smirnov AN, Gauthier LD, Walker M, et al. GATK-gCNV enables discovery of rare copy number variants from exome sequencing data. Nat Genet. 2023;55:1589.
Herman DS, Smith C, Liu C, Vaughn CP, Palaniappan S, Pritchard CC, et al. Efficient detection of copy number mutations in PMS2 exons with a close homolog. J Mol Diagn. 2018;20:512.
Ketkar S, Burrage LC, Lee B. RNA sequencing as a diagnostic tool. JAMA. 2023;329:85–86.
Walker LC, Hoya M, de la, Wiggins GAR, Lindy A, Vincent LM, Parsons MT, et al. Using the ACMG/AMP framework to capture evidence related to predicted and observed impact on splicing: recommendations from the ClinGen SVI Splicing Subgroup. Am J Hum Genet. 2023;110:1046–67.
Horton C, Hoang L, Zimmermann H, Young C, Grzybowski J, Durda K, et al. Diagnostic outcomes of concurrent DNA and RNA sequencing in individuals undergoing hereditary cancer testing. JAMA Oncol. 2024;10:212–9.
Kamps-Hughes N, Carlton VEH, Fresard L, Osazuwa S, Starks E, Vincent JJ, et al. A systematic method for detecting abnormal mRNA splicing and assessing its clinical impact in individuals undergoing genetic testing for hereditary cancer syndromes. J Mol Diagnost. 2023;25:156–67.
Landrith T, Li B, Cass AA, Conner BR, LaDuca H, McKenna DB, et al. Splicing profile by capture RNA-seq identifies pathogenic germline variants in tumor suppressor genes. NPJ Precis Oncol. 2020;4. https://doi.org/10.1038/S41698-020-0109-Y.
de la Hoya M, Soukarieh O, López-Perolio I, Vega A, Walker LC, van Ierland Y, et al. Combined genetic and splicing analysis of BRCA1 c.[594-2 A > C; 641 A > G] highlights the relevance of naturally occurring in-frame transcripts for developing disease gene variant classification algorithms. Hum Mol Genet. 2016;25:2256–68.
Robles-Espinoza CD, Mohammadi P, Bonilla X, Gutierrez-Arcelus M. Allele-specific expression: applications in cancer and technical considerations. Curr Opin Genet Dev. 2021;66:10–19.
Tubeuf H, Caputo SM, Sullivan T, Rondeaux J, Krieger S, Caux-Moncoutier V, et al. Calibration of pathogenicity due to variant-induced leaky splicing defects by using BRCA2 exon 3 as a model system. Cancer Res. 2020;80:3593.
Füllgrabe J, Gosal WS, Creed P, Liu S, Lumby CK, Morley DJ, et al. Simultaneous sequencing of genetic and epigenetic bases in DNA. Nat Biotechnol. 2023;41:1457–64.
Yang C, Arnold AG, Trottier M, Sonoda Y, Abu-Rustum NR, Zivanovic O, et al. Characterization of a novel germline PALB2 duplication in a hereditary breast and ovarian cancer family. Breast Cancer Res Treat. 2016;160:447.
Kwong A, Au CH, Shin VY, Ho DN, Wong EYL, Ho CYS, et al. Rapid breakpoint mapping of a novel germline PALB2 duplication by PCR-free long-read sequencing for interpretation of its pathogenicity. JCO Precis Oncol. 2021;5:1044–7.
Jaganathan K, Ersaro N, Novakovsky G, Wang Y, James T, Schwartzentruber J, et al. Predicting expression-altering promoter mutations with deep learning. Science (1979). 2025. https://doi.org/10.1126/SCIENCE.ADS7373.
Avsec Ž, Latysheva N, Cheng J, Novati G, Taylor KR, Ward T, et al. AlphaGenome: advancing regulatory variant effect prediction with a unified DNA sequence model. bioRxiv. 2025:2025.06.25.661532.
Jaganathan K, Kyriazopoulou Panagiotopoulou S, McRae JF, Darbandi SF, Knowles D, Li YI, et al. Predicting splicing from primary sequence with deep learning. Cell. 2019;176:535–548.e24.
Meienberg J, Bruggmann R, Oexle K, Matyas G. Clinical sequencing: is WGS the better WES? Hum Genet. 2016;135:359.
De La Vega FM, Irvine SA, Anur P, Potts K, Kraft L, Torres R et al. Benchmarking of germline copy number variant callers from whole genome sequencing data for clinical applications. Bioinforma Adv. 2025;5. https://doi.org/10.1093/BIOADV/VBAF071.
Boltz TA, Chu BB, Liao C, Sealock JM, Ye R, Majara L, et al. A blended genome and exome sequencing method captures genetic variation in an unbiased, high-quality, and cost-effective manner. bioRxiv. 2024:2024.09.06.611689.
Palles C, Cazier JB, Howarth KM, Domingo E, Jones AM, Broderick P, et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat Genet. 2013;45:136–43.
Weren RDA, Ligtenberg MJL, Kets CM, De Voer RM, Verwiel ETP, Spruijt L, et al. A germline homozygous mutation in the base-excision repair gene NTHL1 causes adenomatous polyposis and colorectal cancer. Nat Genet. 2015;47:668–71.
NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Genetic/Familial High-Risk Assessment: Colorectal, Endometrial, and Gastric V.1.2025. © National Comprehensive Cancer Network, Inc. 2025. All rights reserved. Accessed [February 16th, 2026]. To view the most recent and complete version of the guideline, go online to NCCN.org.
NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Genetic/Familial High-Risk Assessment: Breast, Ovarian, Pancreatic, and Prostate V.2.2026. © National Comprehensive Cancer Network, Inc. 2025. All rights reserved. Accessed [February 16th, 2026]. To view the most recent and complete version of the guideline, go online to NCCN.org.
Li MM, Chao E, Esplin ED, Miller DT, Nathanson KL, Plon SE, et al. Points to consider for reporting of germline variation in patients undergoing tumor testing: a statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2020;22:1142–8.
Jaber D, Zhang J, Godley LA. Detecting likely germline variants during tumor-based molecular profiling. J Clin Invest. 2025;135. https://doi.org/10.1172/JCI190264.
Schwartz A, Manning DK, Koeller DR, Chittenden A, Isidro RA, Hayes CP, et al. An integrated somatic and germline approach to aid interpretation of germline variants of uncertain significance in cancer susceptibility genes. Front Oncol. 2022;12:942741.
Joo JE, Viana-Errasti J, Buchanan DD, Valle L. Genetics, genomics and clinical features of adenomatous polyposis. Fam Cancer. 2025;24:38.
Belhadj S, Khurram A, Bandlamudi C, Palou-Márquez G, Ravichandran V, Steinsnyder Z, et al. NBN pathogenic germline variants are associated with pan-cancer susceptibility and in vitro DNA damage response defects. Clin Cancer Res. 2023;29:422–31.
Srinivasan P, Bandlamudi C, Jonsson P, Kemel Y, Chavan SS, Richards AL, et al. The context-specific role of germline pathogenicity in tumorigenesis. Nat Genet. 2021;53:1577.
Haile S, Corbett RD, Bilobram S, Bye MH, Kirk H, Pandoh P et al. Sources of erroneous sequences and artifact chimeric reads in next generation sequencing of genomic DNA from formalin-fixed paraffin-embedded samples. Nucleic Acids Res. 2019;47. https://doi.org/10.1093/nar/gky1142.
Liao X, Zhu W, Zhou J, Li H, Xu X, Zhang B et al. Repetitive DNA sequence detection and its role in the human genome. Commun Biol. 2023;6. https://doi.org/10.1038/S42003-023-05322-Y.
Vollger MR, Guitart X, Dishuck PC, Mercuri L, Harvey WT, Gershman A, et al. Segmental duplications and their variation in a complete human genome. Science. 2022;376:eabj6965.
Pande S, Dawood M, Grochowski CM. Structural variants: mechanisms, mapping, and interpretation in human genetics. Genes (Basel). 2025;16:905.
Chen X, Baker D, Dolzhenko E, Devaney JM, Noya J, Berlyoung AS et al. Genome-wide profiling of highly similar paralogous genes using HiFi sequencing. Nat Commun. 2025;16. https://doi.org/10.1038/S41467-025-57505-2.
Smolka M, Paulin LF, Grochowski CM, Horner DW, Mahmoud M, Behera S, et al. Detection of mosaic and population-level structural variants with Sniffles2. Nat Biotechnol. 2024;42:1571–80.
Saunders CT, Holt JM, Baker DN, Lake JA, Belyeu JR, Kronenberg Z et al. Sawfish: improving long-read structural variant discovery and genotyping with local haplotype modeling. Bioinformatics. 2025;41. https://doi.org/10.1093/BIOINFORMATICS/BTAF136.
Nakamura W, Hirata M, Oda S, Chiba K, Okada A, Mateos RN, et al. Assessing the efficacy of target adaptive sampling long-read sequencing through hereditary cancer patient genomes. NPJ Genom Med. 2024;9. https://doi.org/10.1038/S41525-024-00394-Z.
Dixon K, Shen Y, O’Neill K, Mungall KL, Chan S, Bilobram S, et al. Defining the heterogeneity of unbalanced structural variation underlying breast cancer susceptibility by nanopore genome sequencing. Eur J Hum Genet. 2023;31:602–6.
Baumann AA, Knol LI, Arlt M, Hutschenreiter T, Richter A, Widmann TJ, et al. Long-read genome and RNA sequencing resolve a pathogenic intronic germline LINE-1 insertion in APC. NPJ Genom Med. 2025;10. https://doi.org/10.1038/s41525-025-00485-5.
Watson CM, Holliday DL, Crinnion LA, Bonthron DT. Long-read nanopore DNA sequencing can resolve complex intragenic duplication/deletion variants, providing information to enable preimplantation genetic diagnosis. Prenat Diagn. 2022;42:226–32.
Gulsuner S, AbuRayyan A, Mandell JB, Lee MK, Bernier GV, Norquist BM, et al. Long-read DNA and cDNA sequencing identify cancer-predisposing deep intronic variation in tumor-suppressor genes. Genome Res. 2024;34:1825–31.
Te Paske IBAW, Mensenkamp AR, Neveling K, Baert-Desurmont S, Claes KBM, de Leeneer K, et al. Noncoding aberrations in mismatch repair genes underlie a substantial part of the missing heritability in lynch syndrome. Gastroenterology. 2022;163:1691–1694.e7.
Schwenk V, Leal Silva RM, Scharf F, Knaust K, Wendlandt M, Häusser T, et al. Transcript capture and ultradeep long-read RNA sequencing (CAPLRseq) to diagnose HNPCC/Lynch syndrome. J Med Genet. 2023;60:747.
Aucouturier C, Soirat N, Castéra L, Bertrand D, Atkinson A, Lavolé T, et al. Fine mapping of RNA isoform diversity using an innovative targeted long-read RNA sequencing protocol with novel dedicated bioinformatics pipeline. BMC Genom. 2024;25:1–13.
Lønning PE, Nikolaienko O, Knappskog S. Constitutional epimutations: from rare events toward major cancer risk factors? JCO Precis Oncol. 2025;9:e2400746.
Dámaso E, Canet-Hermida J, Vargas-Parra G, Velasco À, Marín F, Darder E, et al. Highly sensitive MLH1 methylation analysis in blood identifies a cancer patient with low-level mosaic MLH1 epimutation. Clin Epigenet. 2019;11. https://doi.org/10.1186/S13148-019-0762-6.
Shuman C, Kalish JM, Weksberg R. Beckwith-Wiedemann Syndrome. GeneReviews® 2023. https://www.ncbi.nlm.nih.gov/books/NBK1394/ (accessed 6 Oct2025).
Akbari V, Hanlon VCT, O’Neill K, Lefebvre L, Schrader KA, Lansdorp PM, et al. Parent-of-origin detection and chromosome-scale haplotyping using long-read DNA methylation sequencing and Strand-seq. Cell Genom. 2022;3:100233.
Stergachis AB, Debo BM, Haugen E, Churchman LS, Stamatoyannopoulos JA. Single-molecule regulatory architectures captured by chromatin fiber sequencing. Science. 2020;368. https://doi.org/10.1126/SCIENCE.AAZ1646.
Thiffault I, Farrow E, Barrett C, Scott M, Ross A, Means JC, et al. Clinical long-read sequencing test for genetic disease diagnosis. JAMA Pediatr. 2025. https://doi.org/10.1001/JAMAPEDIATRICS.2025.3320.
Acknowledgements
We would like to thank our colleagues at Ambry Genetics, especially Melissa Pronold, Rebecca Mar-Heyming, Matthew Schultz, Katherine Shortt, Stuti Joshi, Nicole Lambert, Tiantian Geier and Medhat Mahmoud, for their valuable feedback and contributions to various topics addressed in this review.
Funding
No specific funding to declare.
Author information
Authors and Affiliations
Contributions
SB drafted the original manuscript. All authors participated in reviewing, editing, amending and approving the final text. SB compiled all the recommendations into the final submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors are full-time employees of Ambry Genetics Corporation (Aliso Viejo, CA, USA), A Tempus AI company (Chicago, IL, USA), at the time of writing this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Belhadj, S., Hatch, C.J., LaDuca, H. et al. Sequencing approaches in hereditary cancer testing: strengths, limitations and future directions. Eur J Hum Genet (2026). https://doi.org/10.1038/s41431-026-02078-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41431-026-02078-x