Abstract
Aniridia is a rare congenital vision-loss disorder that is caused primarily by heterozygous loss-of-function variants in the PAX6 gene. There is currently no curative treatment. Gene therapy has emerged as a powerful strategy for treating inherited retinal diseases. Here, we aim to establish a systemic PAX6 gene-augmentation therapy capable of addressing retinal pathologic phenotypes in a preclinical Sey mouse model of aniridia. We evaluated the intravenous delivery of FLAG-tagged PAX6 packaged in the AAV-PHP.eB capsid. Treatment was administered on postnatal day 21 to mimic the average age at diagnosis and the likely early therapeutic window in humans. The transcript levels of the virally-delivered PAX6 were 7.1% at 1 month, and 8.9% at 5 months, of Wt endogenous levels. These transcripts produced PAX6 protein in the ganglion (GCL) and inner nuclear cell layers. Importantly, in this pilot study, a significant increase in GCL thickness at 5 months post-injection was also observed. Furthermore, this structural change in the Sey retina was preceded by a significant increase in Notch1 transcription at 1-month post-injection. Thus, we demonstrated the first successful viral-mediated augmentation of PAX6 in the retina in a preclinical mouse model of aniridia, resulting in a phenotypic change at the structural and molecular levels.
Similar content being viewed by others
Introduction
Aniridia (OMIM: 106210) is a rare congenital disorder that can result in vision loss. It is caused primarily by heterozygous loss-of-function variants in the PAX6 gene [1, 2], which is essential for eye development [3], and its expression is tightly regulated [4,5,6]. While aniridia is named for its hallmark feature of iris hypoplasia, multiple additional ocular structures, including the cornea, lens, retina, and optic nerve, are affected. Patients often experience a range of ocular complications, such as foveal hypoplasia, nystagmus, cataracts, glaucoma, and vision loss. In addition to having ocular abnormalities, patients may also present with systemic complications, including obesity, diabetes, hypertension, cerebral malformations, olfactory dysfunction, and neurodevelopmental delay [7,8,9]. There is currently no curative treatment for aniridia. Management remains largely palliative, focusing on symptom relief through interventions such as cataract surgery, artificial iris and lens implants, or corneal transplantation. These procedures carry substantial risks and typically offer temporary benefits.
Gene therapy has emerged as a powerful strategy for treating inherited retinal diseases, with Luxturna, a treatment for RPE65-associated retinal dystrophy, becoming the first FDA-approved ocular gene therapy [10]. Numerous additional recombinant adeno-associated virus (rAAV)-based therapies are now in development for the treatment of retinal diseases, such as retinitis pigmentosa (ClinicalTrials.gov ID NCT05203939, NCT06292650, and NCT06388200), Stargardt disease (ClinicalTrials.gov ID NCT07063251 and NCT06467344), and X-linked retinoschisis (ClinicalTrials.gov ID NCT06289452). For aniridia, both CRISPR-based editing and gene augmentation are promising strategies [11, 12]. Compensating for the overall reduction in gene expression via gene augmentation may be a more broadly applicable approach than specifically targeting each of the 600 different PAX6 pathogenic variants via CRISPR [13, 14]. However, gene augmentation therapy may be limited by a narrow therapeutic index because of PAX6 dosage sensitivity [15, 16]. Typically, ocular gene therapy has been delivered via the subretinal route [17]. However, systemic gene therapy delivery may be safer for patients with aniridia, as they typically have fragile retinal tissues [18] and systemic complications [7,8,9]. Systemic rAAV-based gene therapies for other diseases, such as onasemnogene abeparvovec (Zolgensma) for spinal muscular atrophy [19] and etranacogene dezaparvovec (Hemgenix) for haemophilia B [20], have already been approved by the FDA. The former uses a ubiquitous promoter that expresses in most cell types of the body, and the latter uses a restricted promoter that can expresses in only a subset of cells. Clinically, both the cornea and the retina are important therapeutic targets in the treatment of aniridia; however, treatments that target the retina have remained largely unexplored [21]. Establishing treatments the promote retinal foveal development would most likely improve vision, and although foveal formation begins prenatally, it continues postnatally for months to years, offering a potential therapeutic window [22, 23].
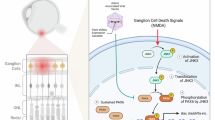
Studies of aniridia at the molecular level have implicated several highly conserved signalling pathways—Notch1, mTOR, and Wnt/β-catenin—in corneal disease [24]. Notch1, which is expressed in proliferating progenitor cells during development and retained in Müller glia in the adult retina [25,26,27], regulates cell fate, differentiation, proliferation, and survival, with transient activation in mature Müller glia capable of inducing retinal cell proliferation [28]. Downstream, Hes1 influences retinal ganglion cell proliferation, differentiation, and morphogenesis [29, 30]. Both canonical and noncanonical Wnt signalling is essential for eye development [31, 32], and the noncanonical ligand Wnt5a supports adult retinal ganglion cell survival, axon growth, and optic nerve regeneration [33]. Additionally, mTOR signalling is critical for ocular cell proliferation, and its loss in mice results in a corneal phenotype reminiscent of aniridia [34]. Within the context of aniridia, the role of these pathways in the postnatal retina is less understood.
Small eye (Sey) [35,36,37] is a heterozygous Pax6 variant in mice, and this variant has been reported in a patient with aniridia [38]; the Sey mouse model recapitulates many ocular features seen in patients with aniridia [39,40,41]. While mice lack a fovea, Sey mice exhibit multiple retinal abnormalities, including disorganized retinal layering and thinning of all layers, including the ganglion cell layer [39,40,41]. This makes the Sey mouse a robust and reproducible preclinical model for testing gene therapies for aniridia. In mice, the AAV-PHP.eB capsid, derived from AAV9, has been optimized using a ubiquitous promoter for widespread biodistribution and enhanced central nervous system tropism [42]. Transduction of multiple retinal cell types, including cell types that express PAX6 in adults, such as retinal ganglion cells, amacrine cells, horizontal cells, [43,44,45] and, less consistently, Müller glia, has been demonstrated [43,44,45]. Unlike its AAV9-derived predecessor PHP. B, which causes lethality in mice [46], PHP.eB has been shown to be well tolerated, with fewer off-target effects [47]. Although gene therapies with ubiquitous promoters are used clinically, gene therapies with restricted promoters are also used and may increase safety by delivering therapeutic agents specifically to target cells [48,49,50,51]. Efforts have been made to create restricted promoters from the PAX6 gene [52, 53]. One example of a human-DNA MiniPromoter tested in mice is Ple331, a synthetic promoter designed for increased expression in PAX6-positive cell types, including retinal ganglion cells, amacrine cells, horizontal cells, and Müller glia [53].
Here, we aim to establish a proof-of-concept for a systemic PAX6 gene augmentation therapy capable of addressing retinal pathologic phenotypes in a preclinical Sey mouse model. We evaluated the efficacy of an intravenously delivered FLAG-tagged PAX6 packaged in the AAV-PHP. eB capsid, whose expression was driven by either the ubiquitous smCBA promoter or the restricted Ple331 MiniPromoter. The FLAG tag has previously been shown not to interfere with PAX6 function [54,55,56], enabling the detection of viral expression without compromising therapeutic potential. Treatment was administered at postnatal day 21 (P21), after both eye opening (P13–14) and weaning (P18–21), but prior to sexual maturity (P35–42). This approximates the average age of humans at aniridia diagnosis (22.1 months) [57] and the likely therapeutic window for early intervention. Experiments were performed in defined hybrid B6129F1-Sey mice, whose genetic background provides a more naturalistic model of aniridia disease than the severe ocular phenotypes observed in pure C57BL/6 J (B6) mice [58]. Together, these design features aim to restore PAX6 expression in relevant retinal cell populations through a safe gene augmentation approach.
Materials and methods
Breeding of B6129F1-Sey Mice
The Sey variant (MMRRC: 050624-MU) was maintained on the genetic background 129S1/SvImJ (The Jackson Laboratory (JAX), RRID:IMSR_JAX:002448, Bar Harbor, ME) backcrossed for at least 10 generations. Experimental Wt and Sey mice were first-generation B6129F1 hybrids produced by crossing C57BL/6J wild-type dams (RRID:IMSR_JAX:000664) with 129S1/SvImJ-Sey heterozygous studs, which also carried the X-linked Hmgcr-lacZ transgene (RRID:IMSR_JAX:038651) (Table S1). B6-Pax6em3(3xFLAG)Ems mice (RRID:MMRRC_066967-MU), hereafter called Fax [54,55,56], were used for molecular assay validation. Genotyping PCR was performed on genomic DNA from ear notches of P18 mice with primers designed to bind to the Sey and Fax alleles, and lacZ transgene, as previously described [55, 58] (Table S2). Mice were on a 7 am–7 pm light cycle with food and water ad libitum. All procedures involving mice were in accordance with the guidelines of the Canadian Council on Animal Care and UBC Animal Care Committee (Protocols A21-0140 and A21-0184).
Cloning and virus production
Two promoters were used: ubiquitous chimeric CMV-chicken β-actin (smCBA) and restricted PAX6-derived Ple331 [53]. Two proteins were delivered: emerald-green fluorescent protein (EmGFP) [59] and three tandem copies of the FLAG peptide fused to the amino end of human PAX6 (3xFLAG/PAX6) [54,55,56]. This allowed the construction of three viral-genome plasmids for virus production: the published smCBA-driving EmGFP (pEMS2143) [53, 60,61,62,63] and the new smCBA-driving 3xFLAG/PAX6 (pEMS2244) and Ple331-driving 3xFLAG/PAX6 (pEMS2393).
The cloning of the pEMS2143 smCBA-EmGFP plasmid has been published previously [53, 62]. The new plasmid pEMS2244 smCBA-3xFLAG/PAX6 was assembled via a multistep cloning strategy using the same synthesized smCBA plasmid (pEMS2075) [53, 62] as that used for the published smCBA-EmGFP (pEMS2143). In this study the smCBA promoter was driving a synthesized 3xFLAG/PAX6 (pEMS1906) gene instead of EmGFP. The new plasmid pEMS2393 Ple331-3xFLAG/PAX6 was also assembled via a multistep cloning strategy from the same synthesized 3xFLAG/PAX6 plasmid (pEMS1906), but it was driven by Ple331, a synthesized promoter (pEMS2166). All three were designed with our “plug and play” ssAAV2 backbone plasmid version 1, which included the following additional elements: multiple cloning site for promoter insertion, chimeric intron, NotI for protein sequence insertion, AsiSI flanked woodchuck hepatitis virus posttranscriptional regulatory element (WPRE), and SV40 polyA [52, 64].
Viral genome plasmids were propagated in E. coli SURE cells (Agilent Technologies, Santa Clara, CA, USA). DNA was prepared with a Spin MiniPrep Kit (QIAGEN, Germantown, MD, USA), and the plasmids were confirmed to be free of rearrangements by AhdI digestion; the ITRs were verified by SmaI digestion, and the cloning sites were verified by sequencing. Confirmed plasmids were sent to the University of Pennsylvania Vector Core (Philadelphia, PA, USA) for large-scale DNA amplification using the EndoFree Plasmid Mega Kit (QIAGEN). Quality control of the plasmid preparation was performed by SmaI, PvuII, and SnaBI digestion, and confirmed plasmids were packaged into AAV-PHP.eB capsid, with the titre determined by droplet digital PCR (ddPCR). Three viruses were produced, namely AAV-PHP.eB smCBA-EmGFP-WPRE, AAV-PHP.eB smCBA-3xFLAG/PAX6-WPRE, and AAV-PHP.eB Ple331-3xFLAG/PAX6-WPRE, hereafter referred to as smCBA-EmGFP, smCBA-FLAG/PAX6, and Ple331-FLAG/PAX6, respectively.
Intravenous Injection and In Vivo Retinal Imaging
Mice were injected with 50 µl of phosphate-buffered saline (PBS) and 0.001% pluronic acid (PBS + P) into the tail vein using a 31-gauge needle on a 0.33 cc syringe (320440, BD, Franklin Lakes, NJ, USA) on P21. The mice were injected with either vehicle control (PBS + P), 1.50 × 1011 smCBA-FLAG/PAX6 genome copies (GC), or 3.09 × 1012 Ple331-FLAG/PAX6 GC alone or coinjected with 50 µl of 3.09 × 1012 Ple331-FLAG/PAX6 GCs plus 1.50 × 1011 smCBA-EmGFP GCs (Fig. 1, Table S1). Mice were assigned to an injection condition using constrained randomization (sex and genotype) by a noninjecting researcher. The injector was blinded to the genotype and solution injected.
A Timeline of tail vein injection, imaging, and mouse tissue harvesting. B Virus injections and numbers of mice that received each injection. The injections were vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 alone or in combination with smCBA-EmGFP. P, postnatal.
For retinal imaging, the mice were placed into an induction chamber and induced with isoflurane using a SomnoSuite (Kent Scientific, Torrington, CA, USA). After the induction phase, the mice were transferred to a nose cone, and the eyes were covered with tropicamide (0.5%) and phenylephrine (2.5%). The pupils were allowed to dilate for two minutes, after which lubricating eye drops were added. A Micron IV retinal imaging microscope (Phoenix Research Labs, Pleasanton, CA, USA) was used to image the eye.
Immunofluorescence and image processing
At 1 or 5 months mice were given a lethal dose of avertin (MilliporeSigma, Burlington, MA, USA), and either perfused for histology or tissues flash-frozen in liquid nitrogen for RNA. For histology, the mice were perfused transcardially with 4% paraformaldehyde; the eyes, brain, spinal cord, liver, heart, and pancreas were dissected; and the tissues were postfixed for 2 h at 4 °C. Eyes were submerged in 25% sucrose overnight at 4 °C for cryoprotection and then embedded in optimal cutting temperature compound (Tissue-Tek, Sakura FineTek, Torrance, CA, USA) on dry ice, after which 20 μm-thick sections were prepared and directly mounted onto slides. Antigen retrieval was performed by incubating the slides in Dako target retrieval solution (Agilent, Santa Clara, CA, USA) for 1 h at 55 °C prior to blocking for 1 h at room temperature. Primary antibody staining was performed using polyclonal chicken anti-GFP (Cat. GFP-1020; Aves Labs, Davis, CA, USA), monoclonal mouse anti-FLAG (Cat. F1804; Millipore Sigma), monoclonal rabbit anti-PAX6 produced with a synthetic peptide corresponding to residues near the carboxy terminus (Cat. 60433; Cell Signaling Technology, Danvers, MA), polyclonal rabbit anti-AP-2β (Cat. 2509; Cell Signaling Technology), or polyclonal rabbit anti-RBPMS (Cat. ABN1362; Millipore Sigma). Secondary antibody staining was performed with goat anti-chicken immunoglobulin (IgG) Alexa Fluor 488 (Cat. A11039; Thermo Fisher Scientific, Waltham, MA, USA), goat anti-mouse IgG Alexa Fluor 594 (Cat. A11032; Thermo Fisher Scientific), and goat anti-rabbit IgG Alexa Fluor 488 (Cat. A11034; Thermo Fisher Scientific). Images were taken at 10X using a Bx61 fluorescence microscope (Olympus Corporation, Tokyo, Japan) and processed using Fiji software version 1.54f [65].
FLAG-positive cells were quantified by calculating the proportion of FLAG-positive cells relative to the total number of Hoechst-stained nuclei in the ganglion cell layer (GCL). FLAG-positive and Hoechst-stained cells were manually counted using the Cell Counter plugin in Fiji software version 1.54 f [65]. For each mouse retina, 2 different sections were analysed.
For the quantification of retinal cell layer thickness, the ImageJ (http://rsbweb.nih.gov/ij/, version 1.52) straight tool was used in the DAPI channel in 2 × 500 µm windows [40]. The following measurements were obtained on both the left and right sides of the optic nerve of the left eye of each mouse: total retinal thickness, total retinal thickness with the retinal pigment epithelium (RPE), the GCL, the inner nuclear layer (INL), and the outer nuclear layer (ONL).
RT‒ddPCR
RNA was isolated using the PureLink RNA Micro Kit (Cat. 12183-016; Thermo Fisher Scientific), and quality was assessed using a NanoDrop Spectrophotometer. cDNA was produced via reverse transcription using Superscript VILO (Cat. 11754-050; Thermo Fisher Scientific) and diluted to 0.5 ng/μm. Reverse transcriptase Droplet Digital PCR (RT-ddPCR) was performed on the QX200 Droplet Digital System (Bio-Rad, Hercules, CA, USA) using ddPCR EvaGreen Supermix (Cat. #1864034, Bio-Rad). The RT‒ddPCR assays and primer sequences are shown in Table S2. Four ubiquitous genes, Rpl13a, Rplp0, Sdha, and Tbp, were quantified using published assays [66] (Table S2). Normalization was subsequently performed to obtain the geometric mean of Rplp0 and Rpl13a. Viral FLAG-tagged PAX6 was quantified with the FLAG-PAX6 assay, which was specific to the viral transcript and did not amplify genome transcripts, as determined by RT‒ddPCR. Endogenous Pax6 (Endo-Pax6) transcripts were quantified using a commercially available assay, which was specific to genomic mouse transcripts and did not amplify viral transcripts as determined by RT‒ddPCR. Notch1, Hes1, Mtor, and Wnt5a transcripts were quantified using commercially available assays.
Statistical analysis
Graphs were generated using GraphPad Prism software (Boston, MA, USA). Each data point corresponds to a single retina from an individual mouse. The data shown are the mean ± standard error of the mean (SEM). Given the sample size, normality was assessed by visual inspections of quantile‒quantile plots. The data had no major deviation from the diagonal line and were therefore judged to be normally distributed. Assumptions of equal variance were verified visually (distribution plots) and supported by consistent results when analysed using nonparametric tests that do not assume equal variance. Differences between the means of the two conditions were analysed using an unpaired two-tailed t-test, with p ≤ 0.05 considered to indicate statistical significance. For comparisons between three or more conditions, ordinary one-way ANOVA was performed, followed by Tukey’s multiple comparisons test if the difference was significant (p ≤ 0.05; GraphPad Prism).
Results
A cohort of ninety-four mice received intravenous injections of control and PAX6-augmented viruses
For this study, we bred a cohort of 94 aged-matched mice that were genotyped for the Sey variant at P18, injected them at P21, and harvested their tissues at either 1 or 5 months post-injection (Fig. 1, Table S1). This timeline is presented to scale in Fig. 1A, and the number of mice of each genotype (Wt or Sey) receiving either single-virus injection or coinjection is shown in Fig. 1B. Single-virus injections involved either a vehicle control or one of two augmentation rAAVs encoding 3xFLAG/PAX6 driven by either a ubiquitous smCBA promoter (smCBA-FLAG/PAX6) or a restricted PAX6-derived Ple331 MiniPromoter (Ple331-FLAG/PAX6). Coinjections involved the same three conditions as single-virus injections, but added to each condition was a fluorescent virus with the smCBA promoter driving EmGFP. All the mice survived with no detriment, and all the Sey/+ (heterozygous) mice qualitatively showed the expected overall ocular phenotype associated with this mouse model of aniridia [46, 54, 58, 67].
Widespread viral transduction in the retina revealed by fundus in vivo imaging of GFP expression
Retinal fundus imaging of mice injected with the fluorescent EmGFP virus enabled longitudinal in vivo assessment of viral expression (Fig. 2). In vivo imaging was not possible in Sey mice because the keratopathy observed in these model mice renders the cornea opaque and prevents the retina from being viewed clearly. In Wt mice, no fluorescence was detected in single-injection vehicle-only controls (Fig. 2A–C), whereas GFP expression was observed throughout the retina in the mice that received a coinjection of smCBA-EmGFP with vehicle (Fig. 2D–F) or one of the augmentation viruses (Fig. 2G–L) at 1, 3, and 5 months post-injection. Coinjection of FLAG/PAX6 augmentation viruses with smCBA-EmGFP did not affect transduction patterns, with all 3 coinjection conditions revealing an even and panretinal distribution of GFP expression (Fig. 2D–L). The intensity of GFP fluorescence appeared brightest at 1 month post-injection (Fig. 2D, G, J), with a slight reduction in the GFP signal at 3 and 5 months (Fig. 1E, F, H, I, K, L). Although some mice had overall lower intensity of GFP fluorescence, the transduction was nonetheless uniform across the entire retina (Fig. 2J–L).
Fundus retinal images of mice injected at P21 with vehicle control, vehicle control coinjected with smCBA-EmGFP, smCBA-FLAG/PAX6 coinjected with smCBA-EmGFP, or Ple331-FLAG/PAX6 coinjected with smCBA-EmGFP. Retinal imaging of wild-type mice was performed at 1, 3, and 5 months post-injection. Brightfield (BF) and GFP epifluorescence images are shown of a mouse that received A–C vehicle control injection, D–F vehicle control coinjected with smCBA-EmGFP, G–I smCBA-3xFLAG/PAX6 coinjected with smCBA-EmGFP, and J–L Ple331-3xFLAG/PAX6 coinjected with smCBA-EmGFP. The numbers above the panels denote the animal IDs.
Transduction of therapeutically relevant cells in the retina revealed by GFP expression
It was important to demonstrate that the AAV-PHP.eB virus was able to transduce adult retinal cells that endogenously express Pax6, including ganglion cells, amacrine cells, horizontal cells, and Müller glia. Thus, GFP expression in cryosections of the retinas of mice that received smCBA-EmGFP was analysed by immunofluorescence at 1 month (Fig. 3) and 5 months (Fig. S1) post-injection. smCBA-GFP was coinjected with vehicle, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6. Retinal cell transduction was evaluated using immunofluorescence and cell morphological analysis, with RBPMS as a marker for RGCs and AP-2β as a marker for amacrine and horizontal cells. Strong GFP expression was consistently observed in the GCL and INL of all the retinas of the mice. Colabelling with RBPMS revealed robust transduction of retinal ganglion cells with smCBA-EmGFP in both Wt (Fig. 3A, E, I, Fig. S1A) and Sey mice (Fig. 3C, G, K, Fig. S1C). Colabelling with AP-2β revealed that amacrine cells were also transduced by smCBA-EmGFP under all coinjection conditions, along with horizontal cells, which were identified by their localization in the distal INL and their dendritic branching into the IPL (Fig. 3B, D, F, G, H, L, Fig. S1B, D). Transduction of Müller glia was observed in both Wt (Fig. 3E, J, Fig. S1A) and Sey (Fig. 3K, Fig. S1C, D) mice, as determined by their distinct vertical morphology. Coinjection of FLAG/PAX6 augmentation viruses with smCBA-EmGFP did not affect transduction patterns (Fig. 3E–L).
Immunofluorescence staining was performed on eye cryosections from mice at 1 month post-injection using anti-GFP (green), anti-RBPMS (ganglion cell marker, red), and anti-AP-2β (amacrine/horizontal cell marker, red) antibodies and Hoechst nuclear counterstain (blue). Two separate cryosections are presented for each Wt and Sey mouse that received injections of A–D vehicle control coinjected with smCBA-EmGFP, E–H smCBA-FLAG/PAX6 coinjected with smCBA-EmGFP, and I–L Ple331-FLAG/PAX6 coinjected with smCBA-EmGFP. For data at 5 months post-injection, see Fig. S1. The arrows indicate EmGFP-positive cells colabelled with RBPMS (ganglion cells), the arrowheads indicate EmGFP-positive cells colabelled with AP-2β (amacrine/horizontal cells), and the J and K asterisks (*) denote EmGFP-positive Müller glia. The numbers above the panels denote the animal IDs. Scale bar = 50 μm. AP-2β, transcription factor AP-2 beta; GCL, ganglion cell layer; IPL, inner plexiform layer; INL, inner nuclear layer; OPL, outer plexiform layer; ONL, outer nuclear layer; RBPMS, RNA binding protein with multiple splicing.
Cells in the ganglion cell layer and inner nuclear layer expressed the PAX6 protein after transduction with the augmentation viruses
Viral PAX6 expression was analysed in cryosectioned retinas using an anti-FLAG antibody to distinguish FLAG/PAX6 from the endogenously expressed Pax6 protein (Fig. 4). As expected, retinas from vehicle-injected Wt and Sey mice showed no FLAG-positive cells, whereas endogenous Pax6 protein expression was clearly observed (Fig. 4A, B, G, H). Both Wt and Sey mice injected with smCBA-FLAG/PAX6 virus displayed FLAG-positive cells at 1 month post-injection (Fig. 4C, D) and sustained this expression at 5 months post-injection (Fig. 4I, J). Importantly, FLAG/PAX6 was predominantly localized to the GCL, in which ganglion cells express endogenous Pax6, while FLAG/PAX6 expression was also observed in the INL, in which amacrine and horizontal cells express endogenous Pax6 (Fig. 4J, L). Similar patterns of FLAG-expressing cells were observed in Wt and Sey mice injected with the Ple331-FLAG/PAX6 virus at both 1 and 5 months post-injection (Fig. 4E, F, K, L).
Immunofluorescence staining was performed on eye cryosections from mice at 1 month and 5 months post-injection using anti-FLAG (red) and anti-Pax6 (green) antibodies and Hoechst nuclear counterstain (blue). Wt and Sey mice received injections of vehicle control at A, B 1 month post-injection and G, H 5 months post-injection; smCBA-FLAG/PAX6 at C, D 1 month post-injection and I, J 5 months post-injection; or Ple331-FLAG/PAX6 at E, F 1 month post-injection and K, L 5 months post-injection. The arrows indicate FLAG-positive cells also stained with Pax6. The numbers above the panels denote the animal IDs. Scale bar = 50 μm. GCL ganglion cell layer, IPL inner plexiform layer, INL inner nuclear layer, OPL outer plexiform layer, ONL outer nuclear layer.
Both augmentation viruses resulted in significant viral PAX6 protein expression in the Wt and Sey ganglion cell layers
To quantify the percentage of cells expressing the viral-delivered PAX6 protein in the GCL, immunofluorescence images were scored for the number of FLAG-positive cells and the total number of Hoechst-stained cells (Fig. 5, Fig. S2). As expected, GCLs from vehicle-injected Wt and Sey GCLs at 1 month and 5 months post-injection showed no FLAG-positive cells (Fig. 5). In contrast, FLAG-positive cells were detected in both smCBA-FLAG/PAX6- and Ple331-FLAG/PAX6-injected Wt and Sey GCLs at 1 month and 5 months post-injection. The percentage of FLAG-positive cells was lower at 1 month, ranging from 1.5% to 5.3%, and higher at 5 months, ranging from 1.9% to 11.1%. Considering the significant differences for each genotype and harvest time point separately, the differences at 5 months post-injection for Wt were very to highly significant, and those for Sey were highly significant. For all significance values, see Fig. S2.
Immunofluorescence staining was performed on eye cryosections from mice harvested at 1 month and 5 months post-injection using an anti-FLAG antibody and Hoechst nuclear counterstain. FLAG-positive (FLAG +) cells were counted as a percentage of total Hoechst counterstained cells in the GCL of Wt and Sey mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6. The data shown are the mean ± SEM, with each dot indicating a single retina analysed from each mouse. Not all significant differences are shown; only those within each genotype and harvest time-point group are shown. For all significance values, see Fig. S2. Statistical significance was determined using ANOVA followed by a post hoc Tukey’s comparison test. A p value < 0.05 was considered to indicate statistical significance. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001.
The retinas of mice injected with SmCBA-FLAG/PAX6 had significantly increased ganglion cell layer thickness
To assess the effects of smCBA-FLAG/PAX6 and Ple331-FLAG/PAX6 virus injection, retinal cell layer thickness was quantified in immunofluorescence images of Wt and Sey mice at 1 month and 5 months post-injection (Fig. 6, Fig. S3). First, considering only the vehicle-injected mice, we were disappointed that the retinal layers and total retinal thickness were not significantly thicker in the Wt mice than in the Sey mice (Fig. 6, Fig. S3, S4). Assuming no impact of the injection, we would have expected the Sey retinas to be thinner [40, 41, 68]. Additionally, in the retinas of mice injected with the ubiquitous smCBA-FLAG/PAX6 augmentation virus, no significant differences in retinal cell layers or total retinal thickness were observed at 1 month post-injection (Fig. 6A, Fig. S3A, C, E, G). Interestingly, at 5 months post-injection, compared with vehicle-treated Sey mice, smCBA-FLAG/PAX6-injected Sey mice exhibited a significantly thicker GCL (Fig. 6B). This increase in thickness was also significantly different than that in the eyes of Ple331-FLAG/PAX6 injected Sey mice and smCBA-FLAG/PAX6 injected Wt mice. This increase was specific to the GCL, since injection of smCBA-FLAG/PAX6 in Wt or Sey mice had no effect on the thickness of other retinal layers or total retinal thickness at 5 months post-injection (Fig. S3B, D, F, H). Furthermore, it was specific to the smCBA-FLAG/PAX6 virus, since injection of the restricted Ple331-FLAG/PAX6 augmentation virus resulted in no significant differences in the thickness of either the GCL (Fig. 6) or other retinal layers or total retinal thickness (Fig. S3) at 1 and 5 months post-injection.
Immunofluorescence images were examined for Wt and Sey mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6. The ganglion cell layer thickness was quantified at A 1 month post-injection and B 5 months post-injection. The data shown are the mean ± SEM, with each dot indicating a single retina analysed from each mouse. Statistical significance was determined using ANOVA and, if significant, a post hoc Tukey’s comparison test. A p value < 0.05 was considered to indicate statistical significance. *p ≤ 0.05; **p ≤ 0.01.
Both augmentation viruses significantly increased the transcript levels of viral PAX6 in the retinas of Wt and Sey mice
Previously, others have shown the potential of Rpl13a, Rplp0, Sdha, and Tbp as ubiquitous genes for RT‒ddPCR normalization in the adult mouse retina [66]. Here, we measured their transcript levels across all treatment groups and at the 1- and 5-month harvest time points (Fig. S5). We found that only Rpl13a and Rplp0 exhibited consistent transcript levels, with no significant difference between genotypes, injection conditions, or time points; thus, these genes were used for normalization. To validate the results of the Endo-Pax6 RT‒ddPCR assay, endogenous Pax6 transcript levels were quantified in the retinas of Wt and Sey mice injected with vehicle only (Fig. S6). As expected for B6129F1 adult retinas [58], Sey mice presented approximately 50% of the Pax6 transcript levels observed in Wt mice, which is consistent with the presence of a single functional Pax6 allele. To validate the results of the FLAG-PAX6 RT‒ddPCR assay, we analysed retinas from uninjected Wt, Fax/+ (heterozygous for a FLAG-tagged Pax6 allele in the genome) [54,55,56], and Fax/Fax (homozygous for the same allele) mice. The sequence of this genomic FLAG/PAX6 gene is the same as that in the augmentation viruses. As expected, all three genotypes exhibited similar levels of Endo-Pax6 transcripts (Fig. S7A), whereas FLAG-PAX6 transcript levels varied with genotype (Fig. S7B). Specifically, no transcripts were detected in the Wt mice; the FLAG-PAX6 transcript levels in the Fax/+ mice were approximately 50% of those in the Fax/Fax mice (Fig. S7C), and the FLAG-PAX6 transcript levels were the same as Endo-Pax6 transcript levels in the Fax/Fax mice (Fig. S7D).
To quantify the levels of viral FLAG-tagged PAX6 and endogenous Pax6, RT‒ddPCR was performed on the retinas of mice injected with vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 viruses at 1 month and 5 months post-injection using virus-specific FLAG-PAX6 and mouse genome-specific Endo-Pax6 assays (Fig. 7, Fig. S8). Here, we consider significant differences for each genotype and harvest time point separately (Fig. 7A). Significant FLAG-PAX6 transcript levels were detected in both Wt and Sey smCBA-FLAG/PAX6-injected mice at 1 month post-injection. Additionally, for mice injected with Ple331-FLAG/PAX6, significant FLAG-PAX6 transcript levels were detected in Wt mice at 1 month post-injection and in Sey mice at 5 months post-injection. For all significance values, see Fig. S8. The transcription of endogenous Pax6 was not affected by the transcription of viral PAX6 in Wt (Fig. 7B) or Sey mice (Fig. 7C). When genotypes were analysed together, the only significant differences were between Wt and Sey mice, with the latter having lower endogenous Pax6 transcript levels, as expected (Fig. S9). The level of viral PAX6 transcripts delivered, as a percentage relative to Endo-Pax6 levels in vehicle-injected Wt mice, was 5.6–10.1% at 1 month post-injection (Fig. 7D) and 5.7–10.1% at 5 months post-injection (Fig. 7E).
The transcript levels of virally delivered FLAG-tagged PAX6 (FLAG-PAX6) or endogenously transcribed Pax6 (Endo-Pax6) were quantified by RT‒ddPCR. A FLAG-PAX6 transcript levels in the retinas of Wt and Sey mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 harvested at 1 month or 5 months post-injection. Not all significant differences are shown; only those within each genotype and harvest time-point group are shown. For all significance values, see Fig. S8. B Endo-Pax6 transcript levels in the retinas of Wt mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 harvested at 1 month and 5 months post-injection. C Endo-Pax6 mRNA transcript levels in the retinas of Sey mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 harvested at 1 month and 5 months post-injection. B and C were reanalyzed by single-way ANOVA, as shown in Fig. S9. D, E Using data in panels A, B, and C, the relative transcription levels of Endo-Pax6 or FLAG-PAX6 were presented as a percentage of the Endo-Pax6 transcript levels of the vehicle-injected Wt samples. D Harvested at 1 month postinjection. E Harvested at 5 months postinjection. A subset of the data in B and C is presented in Fig. S6. The data shown are the mean ± SEM, with each dot indicating a single retina analysed from each mouse. Statistical significance was determined using ANOVA and, if significant, a post hoc Tukey’s comparison test. A p value < 0.05 was considered to indicate statistical significance. *p ≤ 0.05; **p ≤ 0.01; ****p ≤ 0.0001. RT‒ddPCR, reverse transcription–droplet digital PCR.
The smCBA-FLAG/PAX6 virus significantly increased Notch1 transcript levels in the retina
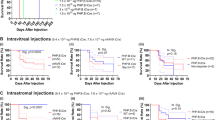
To assess the impact of viral PAX6 delivery on related signalling pathways, we quantified Notch 1, Hes1, Mtor, and Wnt5a transcript levels in retinas using RT‒ddPCR at 1 and 5 months post-injection (Fig. 8, Fig. S10). At 1 month post-injection, compared with that in Sey vehicle control mice, Notch1 expression in Sey mice treated with ubiquitous smCBA-FLAG/PAX6 significantly increased (Fig. 8A). This increase in transcription was also significantly different than Ple331-FLAG/PAX6 and smCBA-FLAG/PAX6 injected Wt eyes, and vehicle injected Wt eyes. At 5 months post-injection, the Notch1 levels in smCBA-FLAG/PAX6-injected Sey mice were still elevated but not significantly elevated (Fig. 8B). In contrast, restricted Ple331-FLAG/PAX6 injection at either 1 or 5 months post-injection did not affect Notch1 transcription in either Wt or Sey mice (Fig. 8A, B). Viral delivery of FLAG/PAX6 with either augmentation virus at 1 or 5 months post-injection had no significant effect on the transcription of the other signalling pathway genes Hes1, Mtor, and Wnt5a (Fig. S10).
Transcript levels of endogenous Notch1 were quantified by RT‒ddPCR in the retinas of Wt and Sey mice that received vehicle control, smCBA-FLAG/PAX6, or Ple331-FLAG/PAX6 at A 1 month and B 5 months post-injection. The data shown are the mean ± SEM, with each dot indicating a single retina analysed from each mouse. Statistical significance was determined using ANOVA and, if significant, a post hoc Tukey’s comparison test. A p value < 0.05 was considered to indicate statistical significance. *p ≤ 0.05; **p ≤ 0.01. RT‒ddPCR, reverse transcription–droplet digital PCR.
Discussion
Treatment with the ubiquitous smCBA-FLAG/PAX6 augmentation virus in Sey mice at P21 resulted in PAX6 transcript levels being 7.1% and 8.9% of Wt endogenous levels at 1 month and 5 months post-injection, respectively (Fig. 7D, E). These viral transcripts produced PAX6 protein in the therapeutically relevant retinal cell types (Fig. 4), which was present in up to 8.4% of the GCL cells by 5 months post-injection (Fig. 5). We also observed a significant increase in GCL thickness at 5 months post-injection (Fig. 6B). Given the modest increase in PAX6 levels and the limited sample size, attributing this change in GCL thickness to the gene therapy should be done with caution and warrants further experimental validation. Furthermore, this change was not observed in the Wt mice (Fig. 6), perhaps suggesting that the modest viral-induced protein increase was not biologically impactful, or at least not when the Pax6 levels were normal. In the Sey/+ mice, the increase in GCL thickness was not present at 1 month post-injection (Fig. 6A), which is consistent with the percentage of GCL cells expressing virally derived PAX6 being lower at 1 month and increasing at 5 months post-injection (Fig. 5). The increased thickness was specific to the GCL, potentially because RGCs were the predominant cell type to be transduced by the virus (Figs. 3, 4). The increased GCL was also specific to the ubiquitous smCBA promoter, leading to the conclusion that the restricted PAX6-derived Ple331 promoter, which delivered similar overall levels of PAX6 transcripts and protein, had an unknown difference in spatial and/or temporal expression. Although the vehicle-injected controls showed that on average the GCL in Wt mice was thicker than Sey mice, this did not reach statistical significance (Fig.S4). We do not attribute this lack of significance to the vehicle injections but rather to a lack of power. Using the data now available in Kaad et al. (2025) [40], we calculated that rather than 8 Wt and 6 Sey mice, we should have used 34 Wt and 34 Sey mice for a 100% effect and 80% power. Thus, we cannot rigorously claim that the smCBA-FLAG/PAX6 augmentation virus had a therapeutic effect; however, we are encouraged by this first, exploratory evidence of the ability of a PAX6-augmentation virus to increase GCL thickness in Sey mice.
We are further encouraged by the structural changes in the Sey mouse retina being accompanied by molecular change. We detected a significant upregulation of Notch1 transcription in the retinas of smCBA-FLAG/PAX6-treated Sey mice at 1 month post-injection (Fig. 8). Thus, this increase preceded the increase in thickness of the GCL, which was not detected until 5 months (Fig. 6). Overall, the increase in Notch1 transcription correlated with increased GCL thickness in smCBA-FLAG/PAX6 virus-injected Wt mice but not in Ple331-FLAG/PAX6-injected mice. Finally, this transcriptional upregulation was not a general phenomenon, since the expression of Hes1, Mtor, and Wnt5a, which are representative members of signalling pathways critical in eye development [32, 34, 69] and altered in the aniridic cornea [24], remained stable (Fig. S10). Typically, Notch1 expression is high during development in proliferating and undifferentiated cells, and in the postnatal mouse retina, it is limited to Müller glial cells [25, 26]. A transient increase in Notch1 signalling has been shown to promote the proliferation of Müller glia in the postnatal retina [70], suggesting that Notch1-mediated signalling pathways may contribute to the observed increase in GCL thickness following treatment.
Rather surprisingly, given the known autoregulation of PAX6 [4,5,6] and the negative impact of overexpression [15, 16], we provide evidence that intravenous delivery of viral PAX6 is well tolerated. We detected no systemic changes, such as illness or death (Fig. S1), or retinal toxicity (Figs. 2, 3, 4), despite intravenous delivery, presumably resulting in widespread virus delivery [47]. In addition, RT‒ddPCR analysis confirmed that the transcription of endogenous Pax6 was unaffected by viral delivery (Fig. 7B, C, Fig. S9), indicating that no compensatory or autoregulatory mechanisms were triggered in response to supraphysiologic levels of Pax6 expression. While this tolerance may be secondary to the modest increase in total Pax6/PAX6 transcription (Fig. 7), the change was apparently sufficient to increase Notch1 transcript levels (Fig. 8) and GCL thickness (Fig. 6) in Sey mice.
As predicted by the histological studies of others, systemic delivery via the bloodstream of the AAV-PHP.eB capsid resulted in widespread and uniform expression of GFP throughout the mouse retina (Figs. 2, 3, Fig. S1) [43,44,45, 71, 72]. We have added to the knowledge about this delivery system in several ways. First, the ability of AAV-PHP.eB to cross the blood‒retinal barrier in the B6129F1 hybrid mouse strain has not previously been tested. However, its component strains, B6 and 129, are known to be in a subgroup of strains permissive for AAV-PHP.eB transport across the blood‒brain barrier [73], and AAV-PHP.eB transport across the blood‒retina barrier in the B6 mouse strain and a related 129 strain has been reported [43, 44, 71, 72]. Next, the mice were injected with the augmentation viruses at P21 to mimic the early treatment of aniridia patients and studied as late as 5 months post-injection to allow time for therapeutic impact. To date, this protocol is both the earliest treatment time and the longest study time used with AAV-PHP.eB in the retina [43,44,45, 71, 72]. Thus, we were delighted to discover that the protein expression pattern revealed at 1 month was sustained and even increased at 5 months post-injection (Figs. 2, 3, 5, Fig. S1, S2). Finally, for therapeutic impact, it is desirable to achieve transduction of adult retinal cells that endogenously express Pax6: ganglion cells, amacrine cells, horizontal cells, and Müller glia. We observed, as anticipated from the marker identification of retinal cells performed by others [43,44,45], the transduction of ganglion, amacrine, and horizontal cells (Fig. 3, Fig. S1). We also observed Müller glial transduction at both 1 and 5 months post-injection, consistent with recent work [44] but not earlier studies [43, 45]. We hypothesize that this difference may reflect the age at injection, since we injected mice at P21, and the former study used mice at 6–8 weeks; however, the latter two studies used mice at 20–22 weeks and 7 months.
In this study, we demonstrated the first successful viral-mediated augmentation of PAX6 expression in the retina in a preclinical mouse model of aniridia, apparently resulting in phenotypic, structural, and molecular changes. Using minimally invasive intravenous delivery in young mice, we achieved widespread retinal transduction, including that of therapeutically relevant cell types. PAX6 delivery correlated with a significant increase in Notch1 transcript levels and subsequent significant increase in retinal GCL thickness in Sey mice that received smCBA-FLAG/PAX6 virus. This occurred without detectable adverse effects: endogenous Pax6 expression remained stable following viral delivery, and no broad dysregulation of key signalling pathways was observed, supporting the specificity of the approach. We observed significantly increased FLAG-PAX6 transcript and protein levels in both Wt and Sey mice using RT‒ddPCR and immunohistochemistry, respectively. RT‒ddPCR yields an average across all retinal cells; thus, the use of single-cell RNA sequencing and spatial transcriptomics in future work may be beneficial. Immunohistochemistry can detect the type and number of viral protein-expressing cells but is not sufficiently quantitative to determine the amount of protein per cell; thus, the use of epifluorescently labelled viral PAX6 in future work may be beneficial. Furthermore, it would be beneficial to carry out studies designed to achieve a mechanistic understanding of the role of PAX6 and Notch1 in increasing retinal thickness in the eyes of Sey mice. In moving this work towards clinical application, it will be important to carry out a replicate study employing a larger cohort of mice, with the aim of detecting significant improvement in all the layers of the retina.
Data availability
The plasmids pEMS2075, pEMS2143, pEMS2166, pEMS2244, and pEMS2393 used in this study are available to the research community through the nonprofit distributor Addgene (www.addgene.org). All the mouse strains used in this study are available through the nonprofit distributor The Jackson Laboratory (www.jax.org).
References
Jordan T, Hanson I, Zaletayev D, Hodgson S, Prosser J, Seawright A, et al. The human PAX6 gene is mutated in two patients with aniridia. Nat Genet. 1992;1:328–32.
Kit V, Cunha DL, Hagag AM, Moosajee M. Longitudinal genotype-phenotype analysis in 86 patients with PAX6-related aniridia. JCI Insight. 2021;6:e148406.
Ochi S, Manabe S, Kikkawa T, Osumi N. Thirty Years’ History since the Discovery of Pax6: From Central Nervous System Development to Neurodevelopmental Disorders. Int J Mol Sci. 2022;23:6115.
Aota S, Nakajima N, Sakamoto R, Watanabe S, Ibaraki N, Okazaki K. Pax6 autoregulation mediated by direct interaction of Pax6 protein with the head surface ectoderm-specific enhancer of the mouse Pax6 gene. Dev Biol. 2003;257:1–13.
Bhatia S, Bengani H, Fish M, Brown A, Divizia MT, de Marco R, et al. Disruption of Autoregulatory Feedback by a Mutation in a Remote, Ultraconserved PAX6 Enhancer Causes Aniridia. Am J Hum Genet. 2013;93:1126–34.
Shukla S, Mishra R. Autoregulation of Pax6 in neuronal cells is mediated by Pax6(5a), Pax6(DeltaPD), SPARC, and p53. Mol Biol Rep. 2022;49:3271–9.
Dansault A, David G, Schwartz C, Jaliffa C, Vieira V, de la Houssaye G, et al. Three new PAX6 mutations including one causing an unusual ophthalmic phenotype associated with neurodevelopmental abnormalities. Mol Vis. 2007;13:511–23.
Obst J, Fries FN, Amini M, Naray A, Munteanu C, Stachon T, et al. Systemic Diseases in Patients with Congenital Aniridia: A Report from the Homburg Registry for Congenital Aniridia. Ophthalmol Ther. 2025;14:433–45.
Sisodiya SM, Free SL, Williamson KA, Mitchell TN, Willis C, Stevens JM, et al. PAX6 haploinsufficiency causes cerebral malformation and olfactory dysfunction in humans. Nat Genet. 2001;28:214–6.
Russell S, Bennett J, Wellman JA, Chung DC, Yu ZF, Tillman A, et al. Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet. 2017;390:849–60.
Campochiaro PA, Avery R, Brown DM, Heier JS, Ho AC, Huddleston SM, et al. Gene therapy for neovascular age-related macular degeneration by subretinal delivery of RGX-314: a phase 1/2a dose-escalation study. Lancet. 2024;403:1563–73.
Khanani AM, Boyer DS, Wykoff CC, Regillo CD, Busbee BG, Pieramici D, et al. Safety and efficacy of ixoberogene soroparvovec in neovascular age-related macular degeneration in the United States (OPTIC): a prospective, two-year, multicentre phase 1 study. EClinicalMedicine. 2024;67:102394.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2016;44:D862–8.
Lima Cunha D, Arno G, Corton M, Moosajee M. The Spectrum of PAX6 Mutations and Genotype-Phenotype Correlations in the Eye. Genes (Basel). 2019;10:1050.
Manuel M, Georgala PA, Carr CB, Chanas S, Kleinjan DA, Martynoga B, et al. Controlled overexpression of Pax6 in vivo negatively autoregulates the Pax6 locus, causing cell-autonomous defects of late cortical progenitor proliferation with little effect on cortical arealization. Development. 2007;134:545–55.
Schedl A, Ross A, Lee M, Engelkamp D, Rashbass P, van Heyningen V, et al. Influence of PAX6 gene dosage on development: overexpression causes severe eye abnormalities. Cell. 1996;86:71–82.
Irigoyen C, Amenabar Alonso A, Sanchez-Molina J, Rodriguez-Hidalgo M, Lara-Lopez A, Ruiz-Ederra J. Subretinal Injection Techniques for Retinal Disease: A Review. J Clin Med. 2022;11:4717.
Lee H, Khan R, O’Keefe M. Aniridia: current pathology and management. Acta Ophthalmol. 2008;86:708–15.
Ogbonmide T, Rathore R, Rangrej SB, Hutchinson S, Lewis M, Ojilere S, et al. Gene Therapy for Spinal Muscular Atrophy (SMA): A Review of Current Challenges and Safety Considerations for Onasemnogene Abeparvovec (Zolgensma). Cureus. 2023;15:e36197.
Anguela XM, High KA. Hemophilia B and gene therapy: a new chapter with etranacogene dezaparvovec. Blood Adv. 2024;8:1796–803.
Grainger RM, Lauderdale JD, Collins JL, Trout KL, McCullen Krantz S, Wolfe SS, et al. Report on the 2021 Aniridia North America symposium on PAX6, aniridia, and beyond. Ocul Surf. 2023;29:423–31.
Hendrickson A, Possin D, Vajzovic L, Toth CA. Histologic development of the human fovea from midgestation to maturity. Am J Ophthalmol. 2012;154:767–78.e2.
Provis JM, Diaz CM, Dreher B. Ontogeny of the primate fovea: a central issue in retinal development. Prog Neurobiol. 1998;54:549–80.
Vicente A, Bystrom B, Pedrosa Domellof F. Altered Signaling Pathways in Aniridia-Related Keratopathy. Invest Ophthalmol Vis Sci. 2018;59:5531–41.
Dorsky RI, Rapaport DH, Harris WA. Xotch inhibits cell differentiation in the Xenopus retina. Neuron. 1995;14:487–96.
Furukawa T, Mukherjee S, Bao ZZ, Morrow EM, Cepko CL. rax, Hes1, and notch1 promote the formation of Muller glia by postnatal retinal progenitor cells. Neuron. 2000;26:383–94.
Lindsell CE, Boulter J, diSibio G, Gossler A, Weinmaster G. Expression patterns of Jagged, Delta1, Notch1, Notch2, and Notch3 genes identify ligand-receptor pairs that may function in neural development. Mol Cell Neurosci. 1996;8:14–27.
Mills EA, Goldman D. The Regulation of Notch Signaling in Retinal Development and Regeneration. Curr Pathobiol Rep. 2017;5:323–31.
Bosze B, Moon MS, Kageyama R, Brown NL. Simultaneous Requirements for Hes1 in Retinal Neurogenesis and Optic Cup-Stalk Boundary Maintenance. J Neurosci. 2020;40:1501–13.
Takatsuka K, Hatakeyama J, Bessho Y, Kageyama R. Roles of the bHLH gene Hes1 in retinal morphogenesis. Brain Res. 2004;1004:148–55.
Shah R, Amador C, Chun ST, Ghiam S, Saghizadeh M, Kramerov AA, et al. Non-canonical Wnt signaling in the eye. Prog Retin Eye Res. 2023;95:101149.
Wang Y, Mahesh P, Wang Y, Novo SG, Shihan MH, Hayward-Piatkovskyi B, et al. Spatiotemporal dynamics of canonical Wnt signaling during embryonic eye development and posterior capsular opacification (PCO). Exp Eye Res. 2018;175:148–58.
Musada GR, Carmy-Bennun T, Hackam AS. Identification of a Novel Axon Regeneration Role for Noncanonical Wnt Signaling in the Adult Retina after Injury. eNeuro. 2022;9:ENEURO.0182-22.2022.
Hagglund AC, Jones I, Carlsson L. A novel mouse model of anterior segment dysgenesis (ASD): conditional deletion of Tsc1 disrupts ciliary body and iris development. Dis Model Mech. 2017;10:245–57.
Hill RE, Favor J, Hogan BL, Ton CC, Saunders GF, Hanson IM, et al. Mouse small eye results from mutations in a paired-like homeobox-containing gene. Nature. 1991;354:522–5.
Hogan BL, Horsburgh G, Cohen J, Hetherington CM, Fisher G, Lyon MF. Small eyes (Sey): a homozygous lethal mutation on chromosome 2 which affects the differentiation of both lens and nasal placodes in the mouse. J Embryol Exp Morphol. 1986;97:95–110.
Roberts RC. Small-eyes, a new dominant mutant in the mouse. Genet Res. 1967;9:121–2.
Hever AM, Williamson KA, van Heyningen V. Developmental malformations of the eye: the role of PAX6, SOX2 and OTX2. Clin Genet. 2006;69:459–70.
Cole JD, McDaniel JA, Nilak J, Ban A, Rodriguez C, Hameed Z, et al. Characterization of neural damage and neuroinflammation in Pax6 small-eye mice. Exp Eye Res. 2024;238:109723.
Kaad SG, Korecki AJ, Simpson EM. Establishing Preclinical Quantitative Parameters for Future Assessment of Corneal and Retinal Therapeutics for Aniridia. Invest Ophthalmol Vis Sci. 2025;66:60.
Wang X, Gregory-Evans K, Wasan KM, Sivak O, Shan X, Gregory-Evans CY. Efficacy of Postnatal In Vivo Nonsense Suppression Therapy in a Pax6 Mouse Model of Aniridia. Mol Ther Nucleic Acids. 2017;7:417–28.
Chan KY, Jang MJ, Yoo BB, Greenbaum A, Ravi N, Wu WL, et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat Neurosci. 2017;20:1172–9.
Palfi A, Chadderton N, Millington-Ward S, Post I, Humphries P, Kenna PF, et al. AAV-PHP.eB transduces both the inner and outer retina with high efficacy in mice. Mol Ther Methods Clin Dev. 2022;25:236–49.
Tang M, Zhong L, Rong H, Li K, Ye M, Peng J, et al. Efficient retinal ganglion cells transduction by retro-orbital venous sinus injection of AAV-PHP.eB in mature mice. Exp Eye Res. 2024;244:109931.
Touahri Y, Dixit R, Kofoed RH, Miloska K, Park E, Raeisossadati R, et al. Focused ultrasound as a novel strategy for noninvasive gene delivery to retinal Muller glia. Theranostics. 2020;10:2982–99.
Mirjalili Mohanna SZ, Korecki AJ, Simpson EM. rAAV-PHP.B escapes the mouse eye and causes lethality whereas rAAV9 can transduce aniridic corneal limbal stem cells without lethality. Gene Ther. 2023;30:670–84.
Mathiesen SN, Lock JL, Schoderboeck L, Abraham WC, Hughes SM. CNS Transduction Benefits of AAV-PHP.eB over AAV9 Are Dependent on Administration Route and Mouse Strain. Mol Ther Methods Clin Dev. 2020;19:447–58.
Fujita K, Nishiguchi KM, Shiga Y, Nakazawa T. Spatially and Temporally Regulated NRF2 Gene Therapy Using Mcp-1 Promoter in Retinal Ganglion Cell Injury. Mol Ther Methods Clin Dev. 2017;5:130–41.
Xiong W, Wu DM, Xue Y, Wang SK, Chung MJ, Ji X, et al. AAV cis-regulatory sequences are correlated with ocular toxicity. Proc Natl Acad Sci USA. 2019;116:5785–94.
Khabou H, Cordeau C, Pacot L, Fisson S, Dalkara D. Dosage Thresholds and Influence of Transgene Cassette in Adeno-Associated Virus-Related Toxicity. Hum Gene Ther. 2018;29:1235–41.
Petit L, Punzo C. Gene therapy approaches for the treatment of retinal disorders. Discov Med. 2016;22:221–9.
Hickmott JW, Chen CY, Arenillas DJ, Korecki AJ, Lam SL, Molday LL, et al. PAX6 MiniPromoters drive restricted expression from rAAV in the adult mouse retina. Mol Ther Methods Clin Dev. 2016;3:16051.
Korecki AJ, Cueva-Vargas JL, Fornes O, Agostinone J, Farkas RA, Hickmott JW, et al. Human MiniPromoters for ocular-rAAV expression in ON bipolar, cone, corneal, endothelial, Müller glial, and PAX6 cells. Gene Ther. 2021;28:351–72.
Adair BA, Korecki AJ, Djaksigulova D, Wagner PK, Chiu NY, Lam SL, et al. ABE8e Corrects Pax6-Aniridic Variant in Humanized Mouse ESCs and via LNPs in Ex Vivo Cortical Neurons. Ophthalmol Ther. 2023;12:2049–68.
Mirjalili Mohanna SZ, Hickmott JW, Lam SL, Chiu NY, Lengyell TC, Tam BM, et al. Germline CRISPR/Cas9-Mediated Gene Editing Prevents Vision Loss in a Novel Mouse Model of Aniridia. Mol Ther Methods Clin Dev. 2020;17:478–90.
Mirjalili Mohanna SZM, Djaksigulova D, Hill AM, Wagner PK, Simpson EM, Leavitt BR. LNP-mediated delivery of CRISPR RNP for wide-spread in vivo genome editing in mouse cornea. J Control Release. 2022;350:401–13.
Netland PA, Scott ML, Boyle JW, Lauderdale JD. Ocular and Systemic Findings in a Survey of Aniridia Subjects. J Am Assoc Pediatr Ophthalmol Strabismus. 2011;15:562–6.
Hickmott JW, Gunawardane U, Jensen K, Korecki AJ, Simpson EM. Epistasis between Pax6(Sey) and genetic background reinforces the value of defined hybrid mouse models for therapeutic trials. Gene Ther. 2018;25:524–37.
Teerawanichpan P, Hoffman T, Ashe P, Datla R, Selvaraj G. Investigations of combinations of mutations in the jellyfish green fluorescent protein (GFP) that afford brighter fluorescence, and use of a version (VisGreen) in plant, bacterial, and animal cells. Biochim Biophys Acta. 2007;1770:1360–8.
Kofoed RH, Heinen S, Silburt J, Dubey S, Dibia CL, Maes M, et al. Transgene distribution and immune response after ultrasound delivery of rAAV9 and PHP.B to the brain in a mouse model of amyloidosis. Mol Ther Methods Clin Dev. 2021;23:390–405.
Kofoed RH, Simpson EM, Hynynen K, Aubert I. Sonoselective delivery using ultrasound and microbubbles combined with intravenous rAAV9 CLDN5-GFP does not increase endothelial gene expression. Gene Ther. 2023;30:807–11.
de Moura Gomes A, Petkau TL, Korecki AJ, Fornes O, Galvan A, Lu G, et al. New MiniPromoter Ple389 (ADORA2A) drives selective expression in medium spiny neurons in mice and non-human primates. Sci Rep. 2024;14:28194.
Simpson E, Korecki AJ, Fornes O, McGill TJ, Cueva-Vargas JL, Agostinone J, et al. New MiniPromoter Ple345 (NEFL) Drives Strong and Specific Expression in Retinal Ganglion Cells of Mouse and Primate Retina. Hum Gene Ther. 2019;30:257–72.
de Leeuw CN, Korecki AJ, Berry GE, Hickmott JW, Lam SL, Lengyell TC, et al. rAAV-Compatible MiniPromoters for Restricted Expression in the Brain and Eye. Mol Brain. 2016;9:52.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Adachi H, Tominaga H, Maruyama Y, Yoneda K, Maruyama K, Yoshii K, et al. Stage-specific reference genes significant for quantitative PCR during mouse retinal development. Genes Cells. 2015;20:625–35.
Javidjam D, Moustardas P, Abbasi M, Dashti A, Rautavaara Y, Lagali N. A human-like model of aniridia-associated keratopathy for mechanistic and therapeutic studies. JCI Insight. 2024;10:e183965.
Pedersen HR, Baraas RC, Landsend ECS, Utheim OA, Utheim TP, Gilson SJ, et al. PAX6 Genotypic and Retinal Phenotypic Characterization in Congenital Aniridia. Invest Ophthalmol Vis Sci. 2020;61:14.
Brennan D, Giles S. Sonic hedgehog expression is disrupted following in ovo ethanol exposure during early chick eye development. Reprod Toxicol. 2013;41:49–56.
Jadhav AP, Cho SH, Cepko CL. Notch activity permits retinal cells to progress through multiple progenitor states and acquire a stem cell property. Proc Natl Acad Sci USA. 2006;103:18998–9003.
Basavarajappa D, Gupta V, Chitranshi N, Wall RV, Rajput R, Pushpitha K, et al. Siponimod exerts neuroprotective effects on the retina and higher visual pathway through neuronal S1PR1 in experimental glaucoma. Neural Regen Res. 2023;18:840–8.
Simpson CP, Bolch SN, Zhu P, Weidert F, Dinculescu A, Lobanova ES. Systemic Delivery of Genes to Retina Using Adeno-Associated Viruses. Adv Exp Med Biol. 2019;1185:109–12.
Hordeaux J, Yuan Y, Clark PM, Wang Q, Martino RA, Sims JJ, et al. The GPI-Linked Protein LY6A Drives AAV-PHP.B Transport across the Blood-Brain Barrier. Mol Ther. 2019;27:912–21.
Acknowledgements
Catherine Chan, Ph.D., from the UBC-Gut4Health Microbiome Core (RRID:SCR_023673) was very helpful during the establishment of the RT‒ddPCR assays in this study.
Funding
This work was supported by Fighting Blindness Canada, a charitable organization (GR015664).
Author information
Authors and Affiliations
Contributions
Conceptualization, Funding acquisition, Resources, Supervision: EMS. Formal analysis, Visualization: DD. Investigation, Validation: DD, SGK, AJK, SLL, TCL. Writing - original draft: DD, AJK, EMS. Writing - review & editing: DD, EMS.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All mice were bred and maintained in the pathogen-free mouse core facility of the Center for Molecular Medicine and Therapeutics at The University of British Columbia. All procedures involving animal work were in accordance with the guidelines of the Canadian Council on Animal Care and UBC Animal Care Committee (Protocols A21-0140 and A21-0184).
Consent for publication
All the authors have read and approved the manuscript and consent to its publication.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Djaksigulova, D., Kaad, S.G., Korecki, A.J. et al. First pilot study of intravenous rAAV-PAX6 gene therapy increases retinal-ganglion-cell-layer thickness and Notch1 transcription in a mouse model of aniridia. Gene Ther (2026). https://doi.org/10.1038/s41434-026-00605-5
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41434-026-00605-5