Abstract
Copy number variation (CNV) can drive rapid evolution in changing environments. In microbial pathogens, such adaptation is a key factor underpinning epidemics and colonization of new niches. However, the genomic determinants of such adaptation remain poorly understood. Here, we systematically investigate CNVs in a large genome sequencing dataset spanning a worldwide collection of 1104 genomes from the major wheat pathogen Zymoseptoria tritici. We found overall strong purifying selection acting on most CNVs. Genomic defense mechanisms likely accelerated gene loss over episodes of continental colonization. Local adaptation along climatic gradients was likely facilitated by CNVs affecting secondary metabolite production and gene loss in general. One of the strongest loci for climatic adaptation is a highly conserved gene of the NAD-dependent Sirtuin family. The Sirtuin CNV locus localizes to an ~68-kb Starship mobile element unique to the species carrying genes highly expressed during plant infection. The element has likely lost the ability to transpose, demonstrating how the ongoing domestication of cargo-carrying selfish elements can contribute to selectable variation within populations. Our work highlights how standing variation in gene copy numbers at the global scale can be a major factor driving climatic and metabolic adaptation in microbial species.
Similar content being viewed by others
Introduction
Populations occupying heterogeneous environments may evolve locally advantageous traits under divergent selection pressures1. How different forms of genetic variation contribute to such environmental adaptation remains unclear. Most broad-scale comparative evolutionary analyses are focused on single nucleotide variants (SNVs)2,3,4. However, large-effect structural variants have also been shown to play a role in species range adaptation5,6,7,8,9. Adaptative chromosomal inversions are well documented across populations of Drosophila and are linked to seasonal temperature fluctuations6,7,10 and cold tolerance8,9. Copy number variation (CNV) is a type of unbalanced structural variant defined by the loss or gain of sequence fragments ranging from ~50 bp in length to entire chromosomes. Analyzing CNVs systematically remains challenging due to limits in the detection and resolution of the exact sequence rearrangements11,12. CNVs can drive genome evolution13, contribute to domestication and speciation events14,15, and promote environmental adaptation16,17. Population-based studies revealed CNVs associated with environmental adaptation in seabirds, with a large 60 kb CNV likely contributing to plumage and thermal adaptation18. In wild lobster populations, CNVs but not SNVs are associated with sea surface temperature adaptation19. Hence, elucidating the population genetic context of widely distributed species is necessary to assess how CNVs and dynamic genome compartments contribute to adaptation. The impacts of gene gains and losses mediated by CNVs across the genome vary from local gene dosage effects20 to reshuffling gene structures21, global transcriptional changes, and chromatin reconfiguration22,23. CNVs mainly arise from inaccurate DNA repair and nonhomologous recombination24. Segmental duplications are often triggered by transposable element (TE) activity, and simple repeats are targets for nonallelic homologous recombination (NHR), leading to CNVs25,26. Overall, CNVs are linked to replicative or non-replicative nonhomologous processes based on weak homology24,27.
CNVs have been implicated in numerous phenotypic traits, including human disease28, life-history traits of crops29,30, and drug resistance31,32. For example, gene duplication of ACE-1, the target site for organophosphate and carbamate insecticides, confers resistance to the malaria vector Anopheles gambiae31. CNVs in fungal plant pathogens are a major concern because such genetic variation is linked to fungicide resistance33,34, pathogen virulence35,36, and nutrient absorption efficiency37. Rapid adaptation in plant pathogens is a threat to global food security38 and facilitates climate change39,40. Zymoseptoria tritici is one of the most destructive pathogens of wheat crops worldwide41. This haploid ascomycete underwent global population expansion concurrent with the introduction of wheat cultivation across continents42. With global spread, the pathogen accumulated mutations likely involved in adaptation to new climates42. The genome is organized in highly dynamic chromosomes, including eight accessory chromosomes and high degrees of structural variation43,44. The genome has expanded recently, most likely caused by TE activity and a weakening of genomic defense mediated by repeat-induced point mutations (RIPs)42,45. RIP is thought to be active during sexual reproduction and promotes mutations in any duplicated sequences46. Hence, the genomic defense mechanism is thought to constrain adaptation through CNVs. The species exhibits high gene set polymorphisms across populations47; however, how the global spread of the wheat host has shaped environmental adaptation remains unknown.
Here, we analyze a global panel of 1109 Z. tritici genomes covering all major regions linked to the domestication history of the wheat host. We validate a set of high-confidence CNVs to recapitulate the evolution of gene gain and loss across the global population genetic context of the species to assess the impact on gene functions. Finally, we show how CNVs contributed to chromosomal polymorphism and environmental range adaptation, including an ~68 kb cargo-carrying Starship element.
Results
Chromosomal and gene copy number variants in a 1000-genome panel
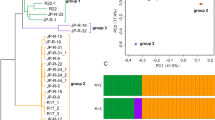
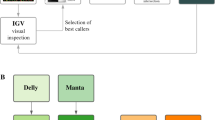
We used short-read sequencing data generated for a global panel of 1109 genomes covering the global distribution range of Z. tritici (Fig. 1A). The collection of genomes covers 42 countries, capturing the spread of the pathogen concurrently with the historic spread of wheat cultivation across continents42. The genomes were collected from a broad range of climates from hot and dry Middle Eastern regions to cooler and humid regions at high latitudes. We performed short-read mapping along the genome to assess segmental deletions and duplications. We implemented multiple filtering procedures and validation steps to ensure high-confidence CNV calls (Fig. 1B). To evaluate CNV call performance, we first compared the gene CNV calling of seven matching pair strains with replicated sequencing data. We found largely congruent calls for duplications and deletions (Fig. 1C). Next, we validated the CNV calling independently based on completely assembled genomes and synteny analyses48 (Supplementary Data 3). We found high consistency for variant calls between short-read CNV calling and the chromosome-level synteny approach (Fig. 1D). We used the empirically assessed confidence threshold (i.e., CNQ) to filter the global short-read dataset. Deletion and single-copy event calls were filtered to reduce false positives and retain high-confidence calls (Fig. 1D, E, Supplementary Fig. S1A, B. Finally, we evaluated CNV call quality for 14 core chromosome genes based on PCR assays conducted on 18 strains47. We compared against the CNV unfiltered dataset and the filtered dataset (Fig. 1F). The resulting dataset of high-confidence calls included 1104 strains and 8625 distinct gene loci affected by CNVs (Fig. 1B; Supplementary Data 4).
A Geographic distribution of the samples. N=1109 samples. The world map was created with the R package maps v. 3.4.2 based on Natural Earth project data (https://www.naturalearthdata.com). Source data can be found in Supplementary Data 1. B CNV call dataset after hard filtering across genetic clusters. Genetic clusters reflect the main genetic differentiation across the globe and were retrieved from Feurtey et al.42 Source data can be found in Supplementary Data 4. C Gene CNV call accuracy comparison between strains sequenced twice. Conflicting loci are defined as mismatches in CNV calls among the sequencing replicates. Source data can be found in Supplementary Data 4. D Gene CNV calls congruence comparison between GATK and chromosome-level assembly (SyRI) based CNV calling methods. The bar plot refers to the number of matching loci between methods. N = 43,517 loci. The boxplot refers to the quality call parameter CNQ score between matching and nonmatching calls. The box center line represents the median, and the limits represent the first and third quartiles. Whiskers indicate maximum and minimum values. Source data can be found in Supplementary Data 3. E CNQ score distribution for duplications and deletions per chromosome after filtering. Source data can be found in Supplementary Data 4. F Number of matches of gene CNV calls with PCR validation (14 core chromosome genes tested in 18 strains). The comparison shows both matches with the unfiltered and the filtered CNV callset. Circle size reflects the number of occurrences.
The species carries a set of eight highly polymorphic accessory chromosomes with unknown contributions to environmental adaptation. Accessory chromosomes are more polymorphic compared to core chromosomes44, creating challenges to define clear read depth thresholds for chromosome presence and absence (Fig. 2A). Shorter accessory chromosome variants of the canonical chromosome variant are expected to show reduced overall read depth due to missing segments. We applied a threshold of >60% genes per chromosome showing deletions to call the chromosome missing. Similarly, we required >60% of the genes to be called duplicated to call an accessory chromosome duplication (Fig. 2B, Supplementary Fig. S2A). Based on these thresholds, chromosomes with very substantial segmental deletions were considered in the same category as complete losses. Similarly, substantial partial duplications affecting were grouped with complete duplications (Supplementary Fig. S2B). To assess the reliability of chromosome CNV calling, we assessed the presence of accessory chromosomes in eight strains where a chromosome-level assembly was produced using PacBio long-read sequencing. We found a 100% match between chromosome CNV calling and chromosome sets present in the assemblies (Supplementary Fig. S2C). Furthermore, we confirmed the absence of particular chromosomes using transcriptomics data (Supplementary Fig. S3A)48. Finally, we verified accessory chromosome CNV calls using data from a previously conducted PCR assay covering 71 loci across two cores and all eight accessory chromosomes49. A total of 59 strains overlapped the global genome panel used in this study and the PCR assay. We defined missing chromosomes based on the PCR essay if 45% or more loci (a total mean of 7 loci tested per chromosome) were missing within the respective chromosome. We could validate 99.5% of chromosome presence calls and 97% of all chromosome absence calls (Supplementary Fig. S3B).
A Read coverage ratio between accessory chromosomes and core chromosomes in the global collection (n = 1104). Individual chromosomes exceeding 1.5 times the mean core chromosome coverage were categorized as being either fully or at least partially duplicated. Chromosomes with coverage near the threshold (1.5 ± 0.25) were manually curated. B Distribution of gene presence/absence calls for the chromosomes 9 (core) and 19 (accessory). The 60% gene presence threshold in red was used to label the whole chromosome to be present or absent. C Frequency of chromosome CNVs in the global collection. Complex CNVs are defined as chromosomes exhibiting large deletions and duplications across chromosome arms. D Distribution of chromosome duplications (full and partial). E Total chromosome count including duplicates. Source data for Fig. 2A-E can be found in Supplementary Data 4.
As expected, core chromosomes 1–13 were fixed in the global collection (Fig. 2C). We found 17 (0.1%) cases of core chromosome duplications (full or partial), with chromosomes 5 and 12 accounting for 64% of all cases (Fig. 2D). We found that 19% of accessory chromosomes were missing from the global collection (n = 1698 out of 8872; Fig. 2C). Chromosome 18 was missing in 57% of strains, followed by chromosome 21, which was missing in 26% of samples (Fig. 2C). Overall, chromosome 18 accounts for 70% of all complex arrangements (i.e., high degree of duplications and deletions across the chromosome arm; Supplementary Fig. S3C), with most variation being associated with high repeat content. We found that the chromosome 14 structural variation in the population was associated with a large insertion44,49 of 351 kb encoding ca. 40 transcriptionally repressed genes (Supplementary Fig. S3D). The total chromosome number per genome varied up to 56% (13–23 chromosomes), with an average of 20 chromosomes underpinning substantial genetic diversity (Fig. 2E). Two strains (0.6% of total) carried only core chromosomes (Supplementary Fig. S3E). We found no evidence supporting yet undescribed accessory chromosomes analyzing scaffolds produced by de novo genome assembly. Higher chromosome numbers than found in the reference genome strain were caused by chromosomal duplications. We found that even strains with the highest chromosome numbers (n ≥ 21) were able to successfully infect wheat leaves and reproduce (Supplementary Fig. S3F).
To identify candidate CNVs associated with climatic adaptation, we analyzed gene CNV variation across the global distribution range of the pathogen. We focused on biallelic CNVs with single-copy, deletion, or duplication of genes (Supplementary Fig. S4A). For each strain, missing or duplicated chromosomes were removed to retain only single-copy chromosomes for further analyses. We found that 3%, 5.1%, and 18.3% of CNVs found in genes were common (>1% CNV frequency), rare (≤1% CNV frequency), and singletons, respectively, across the 1000-genome panel for a total of 8511 loci (Fig. 3A). Most genes (73.9%) showed no CNVs (i.e. fixed CNV frequency category). Most CNV genes (85.7%) share an ortholog in at least one sister species (Supplementary Fig. S4B). Using parsimony, we infer that most deletions of CNV events are likely gene losses. Most genes on accessory chromosomes (90%; n = 203) exhibited CNVs compared to 24% (n = 2021) of core chromosome genes (Fig. 3A). Among the core chromosomes, chromosome 5 exhibited the strongest skew toward low-frequency CNVs (i.e., singleton and rare category with CNV frequency ≤1%; Fig. 3A). CNV variation was also found across chromosome arms (Fig. 3B). We found overall gene duplications to be more abundant (n = 1400) but mostly at low frequency (1388 CNVs with ≤1% duplication frequency in panel) compared to gene deletions (n = 682, Fig. 3C; Supplementary Fig. S4C). The distinct frequency patterns of duplications and deletions were consistent across quality filtering stages (i.e. low and high-frequency CNVs; Supplementary Fig. S4D). Hence, we asked whether gene deletions and duplications would segregate differently among populations (Fig. 3D). Rare CNVs (global frequency <1%) showed similar proportions for gene deletions and duplications compared to the common frequency category (>1%). Gene duplications were four times less likely shared between populations compared to deletions (p-value < 0.00001, 95% CI 14–41%; Supplementary Fig. S5A). We then searched for CNV segment size variation in the global genome panel. We binned 1-kb CNV events into larger contiguous calls of presence or absence (Supplementary Data 5; Supplementary Fig. S5B). In the upper quartile (QA > 21), we found duplicated gene segments to be larger in size and encode more genes compared to deletions (Fig. 3E; Supplementary Fig. S5C). Taken together, our findings are consistent with CNVs being under purifying selection and that gene duplications show high population specificity.
A Pie chart of the overall percentage of gene CNV frequency in the global collection. The bar plot shows the gene CNV frequency per chromosome. Genes without evidence for any CNV event are grouped in the category CNV frequency fixed. The analysis was performed solely with euploid chromosomes to remove any bias from missing or duplicated chromosomes. B Density plot showing an overview of CNV frequency types across chromosome arms. Colors refer to gene CNV events. C Gene CNV event of each CNV frequency type. The density plot shows the CNV event distribution of low-frequency (singleton and rare frequency type) versus high-frequency (frequencies > 1%) CNV. D Number of gene CNVs shared between the 11 genetic clusters (i.e. populations). Colors refer to gene CNV events. Source data can be found in Supplementary Data 4. E CNV segment size across different CNV events and frequencies in the highest quartile CNV segment quality score (QA). The number within the boxplot refers to the sample size. N = 1137. Pairwise two-sided Wilcoxon test. Values on top refer to adjusted p value (Holm method). Groups with <3 samples lacked statistical power. Source data can be found in Supplementary Data 5. F CNV gene distance to the closest transposable element. The box center line represents the median, and the limits represent the first and third quartiles. Whiskers indicate maximum and minimum values. N = 8164. Pairwise two-side Wilcoxon test, adjusted p-value (Holm method). G Gene ontology enrichment analysis of CNV genes. Values in gray refer to two-sided Fisher’s test p values. BP refers to biological process and MF refers to molecular function. Source data can be found in Supplementary Data 6.
We investigated features shared by CNV-affected genes and found that such genes were, on average, closer to TEs than conserved genes (Supplementary Fig. S6A). Furthermore, gene deletions were closer to TEs than duplications (p-value < 0.01; Fig. 3F). We then asked whether coding sequences of CNV genes carry more high-impact SNVs (i.e., variants with disruptive effects) compared to conserved genes. Genes segregating deletions harbored higher impact variants compared to conserved genes and genes with duplications. Common duplicated genes exhibit the highest number of high-impact variants (median 0.0012; Supplementary Fig. S6B), both consistent with functional redundancy (i.e., relaxed selection) and genomic defenses affecting coding sequences. We also found that deleted or partially deleted genes were enriched for H3K27me3 repressive histone methylation marks. In contrast, conserved genes and genes with duplications were enriched for H3K4me2 euchromatin marks (Supplementary Fig. S7A). Consistent with these observations, CNV genes show higher transcriptional variation during host infection and possibly higher functional redundancy (Supplementary Fig. S7B). Overall, CNV genes were functionally enriched in metabolic processes, including toxic secondary metabolic processes and peptidase activity (Fig. 3G; Supplementary Data 6; Supplementary Fig. S7C). Gene dispensability and functional enrichment of metabolic processes suggest that gene CNVs facilitate metabolic diversification and local adaptation.
Effects of genome defenses and signatures of local adaptation
Population differentiation across the globe detected at 218 gene CNVs is broadly congruent with SNV-based assessments of population differentiation42 (Fig. 4A, Supplementary Fig. S8A). In Europe, the CNV-based population structure showed a pattern consistent with recent immigration events (Fig. 4A)42. In addition, the European population showed higher rates of gene flow with other regions (Supplementary Fig. S8B), corroborating the role the continent played in historic pathogen dispersal. Populations across the globe differed substantially in the rate of CNV events per strain (19–83 with a median of 44; Fig. 4B). Interestingly, we found a strong enrichment in duplications in the clusters assigned to Australia, NA—USA and Europe (Fig. 4B). An important factor shaping observed CNV rates among populations is the potential activity of RIP genomic defenses. Genomes with a functional copy of dim2 likely express functional RIP machinery42,50. Dim2-carrying populations tended to show higher rates of gene deletions (Fig. 4C). A mixed linear model accounting for population effects showed a weak but significant association of gene deletions and the strength of RIP (r2 = 0.023; Supplementary Fig. S8C, D).
A Principal component (PC) analyses based on 218 CNV genes (left panel) and 26,728 single nucleotide variants (SNVs; right panel). The color scheme identifies the genetic clusters (see Fig. 1). B Number of gene CNVs per strain and variant frequencies at CNV loci per population in core and accessory chromosomes. The sample number per population is shown in the last column. Different letters above the boxplots identify significantly (adjusted p-value < 0.05) different groups based on a one-way ANOVA followed by Tukey’s test. N = 1104 samples. C Relationship between mean gene CNV number per strain and RIP composite index42 in gene deletions and duplications. Genomes were labeled as carrying a functional RIP machinery if an intact copy of the dim2 methyltransferase gene underpinning activation of RIP was detected. Source data for A and B can be found in Supplementary Data 4.
We identified candidates for local adaptation across all genetic clusters. We assessed the 95th upper quantile fixation index (VST) score per gene CNV among populations (Fig. 5A). The top VST CNV gene (Zt09_13_00035) was predicted to be an effector gene with predicted functions in plant infection (Supplementary Data 7). The gene was rare and stable over time in the North African population but fixed across all other populations (Fig. 5B–D). Consistent with the predicted function, the gene is highly expressed during the initial stages of wheat infection and shares no homology outside of the species (Fig. 5E). Hence, the gene may play a role in adaptation to local host genotypes, favoring gene loss to avoid host recognition.
A CNV variant differentiation based on the fixation index VST across core chromosomes. N = 218 genes. The red dotted line marks the 95th quantile used as a threshold. Source data can be found in Supplementary Data 7. B CNV frequencies of the top VST CNV locus across populations. C Frequency variation at the effector locus in MEA-North Africa genomes and D changes over time in MEA genomes. E Expression profile155 of the effector gene during host infection in four strains (dpi: days post-infection). Source data for panels C and D can be found in Supplementary Data 4.
Structural variation underpinning climate adaptation
The species underwent climatic adaptation over the course of continental colonization42. We investigated the contributions of chromosome and gene CNVs to overall climatic adaptation using genome-environment association (GEA) analyses. We analyzed a total of 1099 samples and examined 19 bioclimatic factors (Supplementary Data 8; Supplementary Fig. S9A, B) based on two mixed-model association approaches for adaptive CNV discovery (Bonferroni α = 0.05). Several climatic factors showed strong positive correlations (r > 0.8, p-value < 0.000; Supplementary Fig. S9C). We identified significant associations for the chromosome CNVs of accessory chromosomes 15, 17, and 20 (Supplementary Fig. S10A; Supplementary Data 9). Chromosome 20 was the most consistently retrieved by both GEA methods, revealing the mean temperature to the coldest and driest quarters (Supplementary Fig. S10A). Next, we analyzed phenotypic trait variation for a subset of the global collection of strains (n = 145; Supplementary Data 10)51, and no trait was significantly associated with CNVs (FDR 5% threshold). We identified 21 gene-level CNV associations with climatic factors spanning 14 different loci (Fig. 6A, Supplementary Data 11). Associated loci encoded functions, including epigenetic regulation, metabolism, and cell signaling functions (Supplementary Data 11). We found the strongest correlations with the climatic factors of the maximum temperature of the warmest month and the mean temperature of the warmest quarter (Fig. 6A; r = 0.84; p-value < 0.001, Supplementary Fig. S9C). The associated CNVs were segregating variable gene deletion frequencies (Fig. 6B).
A Manhattan plot of the genome-environment association (GEA) analyses using two mixed model methods (Tassel and GEMMA). The red dotted line refers to the Bonferroni (alpha = 0.05) threshold. The Venn diagram shows the GEA hit overlap between mixed model methods (Tassel and GEMMA). The bar plot shows the number of significant hits per climatic factor. Colors refer to climatic factors. Source data can be found in Supplementary S11. B Distribution of gene CNV frequency and significant GEA hits in the global collection. The red dotted line refers to the mean frequency. Source data can be found in Supplementary Data 8. C Biosynthetic gene cluster (BGC) 19 frequency in the global collection. Circles at the top refer to mean transcriptional activity under in vitro conditions in a subset of the global collection (n = 74). D Presence of BGC19 across geography. N = 1104. The world map was created with the R package maps v. 3.4.2 based on Natural Earth project data (https://www.naturalearthdata.com). E Correlation of BGC presence with the associated phenotype (annual mean temperature). Two-sided Pearson correlation test. Source data can be found in Supplementary Data 4. F On-host reproduction rate in a subset (n = 145) of global collection grouped by presence/absence of BGC19 and three different wheat cultivars. Data displayed the overlapping hits using Tassel and GEMMA. The box center line represents the median, and the limits represent the first and third quartiles. Whiskers indicate maximum and minimum values. N = 130 samples per cultivar. Pairwise two-sided Wilcoxon test, adjusted p value (Holm method). Source data can be found in Supplementary Data 10.
We integrated CNV and SNV-based GEA analyses and identified three genes with shared evidence from both marker types (Supplementary Data 11)42. In general, gene functions identified by each variant type were only moderately overlapping (r2 < 0.6, Supplementary Fig. S9C; Supplementary Data 11). This suggests largely independent contributions by CNVs and SNVs to climate adaptation. We identified a significant association in the gene CNV of Zt09_9_00561 encoding interferon 6 (Fig. 6A), which was supported by both association mapping methods. Gene presence is associated with higher mean temperatures of the wettest quarter (Supplementary Data 11). We also found congruent mean annual temperature associations for the CNVs for the gene pair Zt09_2_00058/60 (Fig. 6A) and the SNV association at Zt09_2_00069 (Supplementary Data 11). The associated CNVs flank a biosynthetic gene cluster (BGC19) on chromosome 2 with the second highest fixation index values (VST) of all gene CNVs. (Figs. 5A, 6C; Supplementary Fig. S10B). We investigated the nature of BGC19 and found that it is a 63-kb cluster of unknown function (Fig. 6C). The BGC is present in the sister species Z. brevis, and the loss of the core biosynthetic gene (Zt09_2_00067) was confirmed by PCR47. The presence of BGC19 was negatively correlated with mean annual temperatures (r = −0.61, p-value < 2.2e−16; Fig. 6D and E). The loss of the BGC was also associated with higher reproduction on wheat leaves of 12 wheat cultivars (Fig. 6F; Supplementary Data 10). Taken together, the deletion of the BGC19 gene cluster is likely under antagonistic pleiotropy for high-temperature adaptation and host colonization potential.
The climate-associated Sir2 locus is occupied by a massive Starship mobile element
Causal factors contributing to environmental adaptation remain poorly understood. We took advantage of the high-quality pangenome resources for the species to investigate the significant association of the annual temperature range with a CNV on chromosome 7 (Figs. 6A, 7A). The gene Zt09_7_00034 is predicted to encode a homolog of Sir2, an NAD-dependent deacetylase major protein family associated with lifespan and mating type silencing in yeast, as well as aging in humans52,53,54. A phylogenetic analysis of the Sirtuin family showed that Z. tritici Sir2 is an ancient duplication of Sir5 in Dothideomycetes followed by multiple independent gene losses (Supplementary Fig. S11). To investigate the chromosomal context of the Sir2 deletion, we analyzed both unfiltered CNV call data and chromosome-level assembled genomes (n = 19). CNV frequency analyses of the global collection showed that the region segregates a large insertion encoding transcriptionally active genes as well as TEs (Fig. 7A). The Sir2 locus was partially present in the center of origin populations (i.e., Middle East, Iran) and fixed in Oceania and the USA (Supplementary Fig. S12A). The gene Zt09_7_00033 adjacent to Sir2 encodes a DUF3435 domain characteristic of a newly described family of tyrosine recombinases55. This fungal-specific tyrosine domain is required for the mobilization of massive TEs identified as Starships55,56. Genes unrelated to DUF3435 encoded inside the Starship are called cargo genes. The massive mobile element resides close to the major global regulator Velvet57, which is linked to sexual reproduction and growth58. We found a single insertion in the Starship (hereafter identified as Swordfish) present mostly as a single copy (global frequency of 64.5%) or entirely missing (14.3% globally), and additional haplotypes showed cargo gene variation (Supplementary Fig. S12B). Swordfish boundaries contained no detectable direct repeats flanking the element, suggesting that the element lost the ability to transpose. The Swordfish region is rich in TEs (Fig. 7A), and chromosome-level assembled genomes revealed syntenic Swordfish flanks among sister species (Supplementary Fig. S13A). Strains harboring Starship carry ~68k additional sequences (SD = 13.5 kb; Supplementary Fig. S13B).
A Temperature variation associated with the CNV of Sir2. Starship gene order in the reference genome IPO323. The black circle represents the median. N = 920 samples. Source data can be found in Supplementary Data 8. B Synteny plot between chromosome-level assemblies of Z. tritici strains and sister species within the Starship region. Diagonal bars refer to different genomic contexts, and dotted arrows refer to transposition events between regions. C Predicted impact on protein sequences (normalized by transcript size), transcriptional activity during the infection host cycle, and post-translational modifications for Starship and flanking region genes. Dpi: days post-infection. Expression data126 includes three biological replicates per condition (i.e. dpi). The light gray refers to the Starship cargo genes.
We used gene synteny and phylogenetic analysis to retrace the evolution of Swordfish. Phylogenies of the flanking genes match the evolutionary history of the genus (Supplementary Fig. S14). Swordfish gene cargo underwent a complex sequence of duplications, transposition, and multiple, independent gene losses (Fig. 7B, Supplementary Fig. S15A). For example, the genes Zt09_7_00039/38 have paralogs in different genetic contexts, suggesting ancestral duplications (Supplementary Fig. S15A). Chromosome-level assemblies lacking Swordfish (strains IR01_48b, CNR93, UR95) show the flanking gene Zt09_7_00030 and cargo genes Zt09_7_00037/38 in a different location on chromosome 7. Although the Swordfish is never present in more than one copy, a close homolog of the Starship tyrosine recombinase was found in strains lacking Starship (i.e., strain Zt269). Distant duplicates of the tyrosine recombinase (identity < 60%) were found on chromosomes 1 and 12 (Supplementary Fig. S15B). Strain 3D1 carried a tandem duplication of the cargo genes Zt09_7_00033/34/35 (Supplementary Fig. S13A). Swordfish is rich in TE sequences, including specific retrotransposons (RLX_LARD_Gridr and RII_Philae; Supplementary Fig. S16A, B). Some strains lacking Swordfish carry cargo genes (Zt09_7_00037-39) in a different genomic location, suggesting that Swordfish cargo turnover was recent and possibly mediated by TEs (Supplementary Fig. S16C). Epigenetic analyses revealed a bi-repressive pattern regulated by H3K27me3 and H3k9me2 modifications (Fig. 7C). Flanking regions are marked by H3K4me2 euchromatin. The predicted secreted protein Zt09_07_00037 and the flanking gene Zt09_7_00040 are highly expressed during infection (Fig. 7C). We found a high degree of sequence conservation among the triplet Zt09_7_00034/35 lacking coding sequence variants, suggesting strong purifying selection and conservation of synteny (Fig. 7C, Supplementary Fig. S17).
The Swordfish cargo gene Sir2 CNV is significantly associated with the annual range of temperature, and the neighboring highly expressed secreted gene Zt09_7_00037 is associated with the mean diurnal range. Both climatic factors are moderately correlated (r = 0.55, p-value < 0.001). Overall, we identified a possible new role for Sir2 as a factor in climate adaptation. Hence, the massive swordfish mobile element likely contributes to thermal adaptation and shapes the species range across highly variable environmental conditions.
Discussion
Climatic factors are strong determinants of pathogen spread and disease severity39,59. Genetic variability provides the substrate for rapid adaptation to a changing environment60 and to cope with environmental stressors61,62,63. How copy number variation shapes climate gradients spanned by individual species remains poorly understood. We show that multiple gene deletions contribute to pathogen adaptation across continental climatic gradients. CNVs are also drivers of metabolic diversity, including contributions by Starship mobile elements reshuffling genes carried as cargo.
Approximately a quarter of all genes in the pathogen species were affected by CNV events, likely reflecting an equilibrium between new CNVs being generated and purifying selection acting against CNVs64. Genes segregating CNVs were located closer to TEs and were functionally enriched for catalytic activities and secondary metabolic processes compared to more conserved genes. Furthermore, gene duplications were the most abundant CNV events yet remained at low frequency in populations and were rarely shared among populations. The skew in CNV events may stem from a duplication detection bias; however, a more stringent control for call quality and CNV frequencies produced similar outcomes. Gene duplications are a powerful source for gene neofunctionalization65,66,67 and promote non-allelic homologous recombination due to homology among duplicates68,69. Z. tritici is a highly recombinant species70,71, suggesting that segmental duplications could impact the likelihood of nonallelic homologous recombination69. The evolutionary history of the pathogen has likely impacted the rates of duplications on chromosomes, as observed in the more recently colonized Oceanian and American continents. More recent populations exhibited signatures of bottlenecks with reduced genetic diversity42. Concurrently, the bottlenecked populations experienced an increase in TE activity, most likely caused by a loss of defense mechanisms against TEs42,50. RIP triggers rapid mutation accumulation after duplication, leading to loss-of-function or, more rarely, diversifying selection in populations72,73. RIP activity likely acted as a driver of gene loss by rapidly mutating genes and facilitating the purging of nonfunctional copies through excision. Such nonrandom processes underlying the creation and elimination of structural variation illuminate how species gene pools can evolve over short evolutionary time periods.
The role of gene duplications in environmental adaptation is well documented across kingdoms66,74,75. How gene loss can contribute to adaptation is less well understood. Our analyses are supporting previous work showing that gene deletions are under strong purifying selection in Z. tritici populations47. Furthermore, gene losses can largely be explained CNV-driven environmental adaptation. We found signatures for adaptive gene loss ranging from single genes to chromosome copy number variation. Gene loss typically arises from the abrupt rearrangement of coding sequences by repetitive elements, unequal crossing-over events, or by the accumulation of mutations leading to a loss of function76,77. For example, loss of the Desat2 gene in cosmopolitan D. melanogaster was linked to resistance to cold78. Hence, adaptive loss to changing environments shows convergence across kingdoms77,79,80,81,82.
We found strong associations indicating that climate adaptation across continents was likely facilitated by the presence/absence of variation in a biosynthetic gene cluster. The presence of the BGC19 cluster is positively associated with colder climates, and cluster deletion is associated with an increased reproduction rate in specific wheat cultivars. Fungal BGCs are a major source of chemical diversity, and tight physical organization improves coregulation efficiency and reduces cytotoxicity caused by intermediary products83. Secondary metabolites are involved in numerous cellular functions, including virulence, defense, and growth84. BGCs can be hotspots generating structural variation85,86,87 and favor adaptation88,89. Adaptive loss of BGCs is thought to be explained by the Black Queen hypothesis, which argues that communities sharing “leaky” resources such as metabolites will favor genome reduction and loss of gene sets necessary to make such metabolites accessible90. As we have identified linked antagonistic pleiotropy governing the presence of BGC19, the gene cluster may be under a complex selection regime across populations.
In addition to climate adaptation, virulence factors (i.e., effectors) can also undergo adaptive gene loss to facilitate pathogen adaptation52,91. In an arms race between host and pathogens, such virulence factors can be important triggers for host defense mechanisms or serve to manipulate the immune response91,92. The sampling in North Africa covered a 30-year timespan and showed consistently low frequencies of the virulence factor in populations. The region predominantly produces durum wheat (Triticum durum) in contrast to the more widely cultivated bread wheat (T. aestivum)93. Strong selection in North African populations was likely driven by pathogen-specific recognition mechanisms94,95. Overall, we found that gene loss at different scales likely played an important role in environmental adaptation across the historical spread of the pathogen population.
We identified a Starship mobile element associated with climate adaptation in fungal populations. Starships are widespread among fungi and were recently characterized as tyrosine-recombinase-mobilized DNA transposons amassing multiple host genes and TEs56,96. Elements can reach up to approximately half a megabase in size and carry species-specific cargo genes. Individual Starships were associated with adaptive functions such as heavy metal tolerance in a strain-specific manner97 spore killing-mediated meiotic drive96, and formaldehyde resistance98. Starships are powerful agents that reshuffle the core functions of gene clusters98. The cargo carried by swordfish expands our understanding of adaptive functions associated with massive selfish elements. The cargo includes gene CNVs associated with thermal range climatic factors that may contribute to the environmental range of pathogen populations19,99,100. Signatures of thermal adaptation were previously documented in Z. tritici without identifying a molecular mechanism101,102,103. Surprisingly, the Swordfish element resides within a highly conserved region across sister species and neighbors the gene VeltB, which is a key regulator of reproduction, metabolism, and growth processes in fungi57. The strong epigenetic repression across the entire Swordfish element is consistent with the strong repression of TE activity immediately adjacent to a transcriptionally open and conserved region of the chromosome104. We hypothesize that this Starship likely inserted itself at this locus after the emergence of the species, followed by a complex set of duplications and transposition events facilitated by high TE activity in the region.
Swordfish carries an ancient duplication of the Sir5 gene. The sirtuin protein family is a group of evolutionarily conserved NAD+-dependent histone deacetylases involved in regulating epigenetic processes and105 and through their enzymatic activity, sirtuins modulate the acetylation status of histones and other proteins, which have been implicated in a diverse range of biological processes in eukaryotes, including cellular aging, diseases, genome stability, metabolism, and stress response52. The Zt09_7_00034 sirtuin resides exclusively next to the tyrosine recombinase gene within the Swordfish. The lack of SNV variants in the coding sequence, coupled with transcription activity, suggests a local regulatory role within the epigenetic landscape of the element, partially explaining the successful insertion of the element in a highly conserved genomic region. In fungi, sirtuins are involved in mating-type loci silencing, rDNA stabilization, host defense suppression, and secondary metabolism54,106,107. Here, we found evidence of a sirtuin protein linked to climate adaptation in fungi.
We retraced the Starship element formation and hypothesize that Swordfish was formed after speciation and spread across the globe following duplication events, diversification, and retrotransposon-mediated gene transposition. Some pathogen populations nearly fixed or lost Swordfish. The absence of direct repeats at the flanks of the element is consistent with full integration into the fungal genome. The integration followed by mobility impairment suggests that Swordfish was domesticated after its insertion at this locus through selection for adaptative gene cargo and selection against deleterious effects of mobile element activity. Overall, we show that CNVs can be drivers of metabolic diversity and contribute to the global climate adaptation of a crop pathogen. We found that rates of gene loss were strongly associated with the efficiency of genomic defenses, influence adaptive loss-of-functions and prevent maladaptation to a changing environment108. Large genome panels enable the retracing of pathogens spread across continents and disentangle random effects from the most likely effects of local adaptation, highlighting the role CNVs play in the evolution of microbial species.
Methods
Sampling
We performed copy number variation analysis on a global collection of Zymoseptoria tritici comprising 1109 Illumina short-read genomes (Fig. 1A). The collection covers strains originating from 42 countries representative of the history of wheat domestication and historical expansion of wheat cultivation (Supplementary Data 1)42. Additionally, we used high-quality full chromosome assemblies based on PacBio long reads of 19 strains of Z. tritici representative of the global genetic diversity of the species and genomes of four sister species (Z. pseudotritici, Z. ardabilae, Z. brevis and Z. passerinii)44,109 for CNV call validation. Information about sample origin, sequence quality, and accession numbers is provided in Supplementary Data 1.
CNV calling
Illumina raw reads were trimmed with Trimmomatic v. 032110 and mapped to the Z. tritici (IPO323) reference genome using Bowtie2 v.2.4.0111 very-sensitive-local parameter. We used GATK CNV caller v.4.1.9.0112 with recommended parameters on alignment BAM files (n = 1109). The software scans read coverage and models sequencing biases based on negative binomials, taking copy number states and genomic regions of CNV activity into account for a hierarchical hidden Markov model (HHMM). We set the CNV interval to 1000 bp windows with no overlap. Such intervals are recommended to account for variation in sequence coverage (Supplementary Data 1). We filtered for GC content in windows (min = 0.1 and max = 0.9), as well as extremely low and high read counts (--low-count-filter-count-threshold = 5, --extreme-count-filter-minimum-percentile = 1, --extreme-count-filter-maximum-percentile = 99). We then built a prior table for chromosomal ploidy to assign prior probabilities for each ploidy state. Finally, we called CNV genotypes using the germlineCNVCaller and PostprocessGermlineCNVCalls functions. After genome-wide CNV calling, we filtered and validated gene CNVs. We used bedtools v2.31113 annotate to overlap gene elements with the CNV calling.
CNV filtering and validation
CNV coherence validation
To assess CNV calling quality, we analyzed seven pairs of independently sequenced strain (i.e., the same strain with at least two independent library preparation and sequencing efforts; Supplementary Data 2). After validation, we retained the strain with the higher mean coverage of each replicate pair in the dataset for further analysis.
Gene CNV events
CNV calling was performed in 1-kb windows across the genome as described above (see “CNV calling”), allowing for ambiguous calls in polymorphic gene elements. Deletion, duplication and single-copy CNV events were attributed to a gene if the event coverage was >80% of the gene. We defined partial deletions if the deletion covered 50–80% of the gene. Additional combinations were defined as single-copy events.
CNV filtering and validation
To reduce false positive calls in the dataset, we used quality scores (CNQ), which are defined as the difference between the two best genotypes, Phred-scaled log posteriors. We set the CNQ threshold based on the structural variant calling verified using the fully assembled genomes of Z. tritici44. We contrasted CNV calls by the GATK calling pipeline to pairwise whole-genome comparisons of chromosome-level assemblies based on the software SyRI v1.3114 using IPO323 as the reference genome. For a direct comparison of variant genotyping, we analyzed four strains present in both the global dataset (Illumina reads) and the chromosome-level assembly dataset (PacBio reads; samples 3D1, 3D7, 1E4, and 1A5). We first subset gene sets to unambiguous CNV calls (i.e., with more than 60% of event coverage) in both datasets. We then subset the SyRI structural variant calls to single-copy regions, translocation, and DNA gain and loss, removing SNV calls (Supplementary Data 3). We compared the call quality based on CNQ to the gCNV GATK output. The levels of matching and discordant calls between tools were used to define thresholds for CNV GATK calls. We filtered for missing data in the global dataset by removing calls with <50% coverage or <80% call frequency. To define CNV segments (i.e., larger regions with consistent CN calls), we binned GATK CNV Caller segment output per strain and calculated the CNV segment quality call QA defined as complementary Phred-scaled probability at all points (i.e., bins) in the segment, which agree with the segment copy-number call. We then subset for CNV segments based on the filtered dataset and kept CNV segments within the upper quartile quality. We retrieved 14 loci to cross-check the validation with a PCR screen performed earlier for deletion polymorphisms47.
Chromosome number variation
Z. tritici is haploid and carries 13 core chromosomes and up to eight accessory chromosomes (not shared by all members of the species). We used chromosome-wide copy number estimates using the median coverage (MAPQ > 20) of core chromosomes for each strain to define accessory chromosome presence/absence among all strains. Core chromosomes with more than 1.5 times the core coverage were defined as likely duplicated. Chromosome coverage close to the cutoff threshold (1.5 ± 0.3) was manually curated. The high polymorphism of accessory chromosomes44 makes the implementation of thresholds challenging. We defined accessory chromosome absence with gene presence falling below 60% and chromosome duplication with >60% gene duplications. To validate the utility of these thresholds, we analyzed eight strains included in this study for which chromosome-level assemblies are available (3D1, 3D7, 1E4, 1A5, 08STF040, 08STCZ015, 08STCH015 and 09STD078)44.
Single nucleotide variant call and de novo genome assembly
To analyze the genomic context of CNVs in the population, we retrieved SNV calls and de novo genome assembly analyses previously performed for the same global collection (n = 1109)42. Briefly, reads were mapped to the reference genome (IPO323) using Bowtie2 v.2.4.139111. SNVs and short indels were assessed using the short-variant pipeline performed with GATK v4.1.4. HaplotypeCaller115. Ploidy was set to 1, and hard filtering was performed. The per-site filters included FS > 10, 444 MQ < 20, QD < 20, ReadPosRankSum between −2 and 2, MQRankSum between −2 and 2, and BaseQRankSum between −2 and 2. We filtered the dataset for biallelic SNV genotypes, a minor allele frequency of 0.05, and max missing genotype data of 20%. We further used the dataset to predict the effect of SNVs on encoded proteins (categories high, moderate, low, and modifier) using SnpEff version 4.3116. We filtered for CNV genes with a ≥80% single copy genotype in the global panel. We built a database based on the reference genome IPO323 annotation and filtered for the top impact effect per variant. De novo assemblies were generated using SPADES v3.14.1 software117 with the “--careful” parameter to reduce mismatches.
Population structure analysis
The global collection included 1003 genomes grouped into 11 genetic clusters and 106 admixed samples defined as showing <75% assignment to any specific cluster42. CNV-based population structure and VST outlier analyses were performed on the filtered, biallelic gene CNV dataset with a single copy defined as the reference allele and the most frequently observed CNV event at the locus as the alternative allele. We filtered core chromosome gene CNVs for a minor allele frequency ≥1% and 20% maximum missing genotypes per locus. The SNV-based PCA was generated using the SNV data subset with the BCFtools118 "M2" option to keep only biallelic SNV and "-q 0.05:minor" for a minor allele frequency filter of 5%. We used vcftools119 "--thin 1000" to keep only 1 SNV for every 1 kb interval. The PCA was performed with the ade4 R package v1.7-22120 and visualized with the ggplot2 v3.4.2 R package121. CNV fixation indices (VST) were calculated using the hierfstat v0.5-11 R package122.
Environmental and life-history trait adaptation analysis
We performed whole-chromosome CNV and gene CNV genotype-environment association (GEA) analyses using bioclimatic factors from the WorldClim database v.2.1 at 10’ resolution. The bioclimatic data comprise historic averages (between 1970-2000) of 19 climatic variables related to temperature and precipitation. Geographic coordinates of strain collection sites were used to approximate climatic dataset gridpoints. We used two widely used mixed linear model association mapping tools: GEMMA v.0.98.3123 and Tassel v5124 using the Rtassel R package v0.9.29125. We used the thinned SNV dataset (see the section “Population structure”) to estimate the kinship matrix to account for non-random relatedness among genotypes and reduce spurious associations. We contrasted our CNV-based GEA with the SNP-based GEA performed by Feurty et al.42 using identical climatic datasets and tools. We used Bonferroni corrections (alpha = 0.05) to account for multiple testing in GEA analyses. We performed phenotype-genotype GWAS analyses on a subset of the global collection (n = 145 strains) using 24 life-history phenotypes51. The phenotypic data comprised virulence (i.e., lesion size) and reproduction (i.e., pycnidia production) of individual strains during host infection on 12 different wheat cultivars. Tests were conducted with GEMMA and Tassel software using the same parameters as for the GEA analyses.
Transcriptome profiling
We analyzed transcriptional profiles based on gene expression in minimal medium conditions of 19 Z. tritici chromosome-level assembly strains44 and a collection of strains from a Swiss field population (Eschikon, Switzerland; n = 74)48. In summary, 10e5 cells were inoculated using liquid minimum media (MM) with a limited carbon source and grown for 7-10 days to reach the hyphal growth stage. RNA extraction was performed using a NucleoSpin RNA Plant Kit following the manufacturer’s instructions44. We analyzed the publicly available transcriptome dataset (NCBI SRA accession SRP077418) of four strains also included in this study (3D1, 3D7, 1E4, and 1A5) inoculated on wheat plants and analyzed 7, 12, 14, and 28 days after infection126. Illumina raw sequencing reads were trimmed and filtered for adapter contamination using Trimmomatic v.0.32 (parameters: ILLUMINACLIP:Trueseq3_PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36)110. Filtered reads were aligned using HISAT2 v. 2.0.4 with default parameters127 to the Z. tritici reference genome (IPO323). Mapped transcripts were quantified using HTSeq-count v.2.0.2128. Read counts were normalized by calculating trimmed means of M-values (TMM) with the calcNormFactors option. To account for gene length, we calculated reads per kilobase per million mapped reads (RPKM) values using the R package edgeR v.3.42.2129.
Orthology analyses and characterization of gene functions
Orthologs were searched among the chromosome-level assemblies (n = 19) and four sister species of Z. tritici with Orthofinder v.2.2130. Orthologs shared between Z. tritici and at least one sister species were defined as conserved and used to infer gene losses versus gains in Z. tritici. TE annotations of the chromosome-level assemblies were retrieved from refs. 44,45. Chromosome-level assemblies of Z. tritici were also analyzed to predict biosynthesis-related gene clusters (BGCs) using antiSMASH v.5.0131. Identified gene clusters were further annotated using InterProScan v.5.54132. GO term enrichment analyses were performed using Fisher’s exact tests based on gene counts with the topGO R package133. The GO term treemap was plotted using the treemapify R package134. We retrieved publicly available ChIP-seq datasets of histone modifications H3K4me2, H3K9m2, and H3K27me3 from the NCBI SRA (SRP059394) of the Z. tritici IPO323 reference genome isolate grown in rich medium135. ChIP-seq reads were trimmed with Trimmomatic v.0.32110 and mapped to the IPO323 reference genome with Bowtie2 v.2.4111. Alignment BAM files were converted using BEDtools v.2.30113, and peak calling was performed using Homer v.4.11136. Gene coverage was analyzed with the BEDtools intersect command.
Starship characterization and annotation
To characterize Starship mobile elements in the Z. tritici global collection, we searched for genes with CNVs for Starship-associated functional domains56 using the hmmsearch function of HMMER v3.3.2 (E-value ≤ 0.001)137. We focused on genes belonging to a newly described family of tyrosine recombinases with DUF3435 domains (Protein Family accession: PF11917) that are both necessary and sufficient for the movement of entire elements98. We defined the boundaries of candidate elements by annotating their putative empty insertion sites. We aligned 25 kb upstream of the candidate tyrosine recombinases to the corresponding homologous region in isolates that were missing the gene to determine the upstream element boundary and then aligned 25 kb downstream of the homologous region back to the contig containing the tyrosine recombinase to determine the downstream element boundary. All alignments and quality filtering were performed with MUMmer4138 (nucmer settings: -mum; delta-filter settings: -m -l 2000 -i 90) and manually inspected.
To establish phylogenies for Starship cargo genes, we performed pairwise alignments of predicted proteins for each genome using blastP v.2.12139. To ensure that phylogenies are not biased by missed gene annotations, we performed pairwise alignments of the predicted protein sequence against the chromosome-level assemblies and draft assemblies of the global collection using Exonerate v.2.70140 with the parameter --model protein2genome -minintron 20 --maxintron 3000. We aligned sequences with MAFFT v. 7.310141 using the --maxiterate 1000 –localpair options. Phylogenetic trees were built using RAxML v.8142 with the parameters -m GTRGAMMA for nucleotide sequences and -m PROTGAMMAAUTO for protein sequences with 1000 bootstrap replicates.
Visualizations and statistical analyses
All described statistical tests were performed using R143. Analyses of differences among genetic clusters were performed using ANOVAs with the multcomp R package v.1.4-25144. Heatmaps were generated with the pheatmap R package v.1.0.12145. Phylogenetic trees were plotted with the ggtree R package v3.8.0146. Synteny plots of the region were plotted with the genoplotR v.0.8.11147 and gggenomes v.0.9.9148 R packages. The correlation plot was generated using the corrplot R package v.0.92149. Additional graphics were produced using the ggplot2 R package121. To analyze associations of the RIP composite index and gene CNV events, we used a mixed-effect linear regression model with the R package nlme v.3.1.164150. We first used a baseline model of the explanatory variable (i.e., RIP mean per strain) and response variable (i.e., CNV event per strain) and compared it to more complex models adding population and RIP index as random effects. We used the ANOVA function from the package car v.3.1-2151 to assess model fit. We used the function in r.squaredGLMM152 to calculate the conditional (R2c) and marginal coefficients (R2m) of the generalized mixed-effect models. RIP composite data was retrieved from42.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data are available from the NCBI Sequence Read Archive. Individual accession numbers are reported in Supplementary Data 1. Supplementary Data 1, 2 and 5–11 are available in the Supplementary_Data_1-2_5-11.xlsx file. Supplementary Data 3 and 4 are available from https://zenodo.org/records/11616291153.
Code availability
Code used for analyses can be found in Zenodo (https://zenodo.org/records/8344848)154: scripts for plotting Figs. 1–5 can be found in the GATK_CNV_caller.zip and General.zip files, scripts for Fig. 6 can be found in the GEA.zip file, and scripts for Fig. 7 can be found in the swordfish.zip file within the repository154.
References
Savolainen, O., Lascoux, M. & Merilä, J. Ecological genomics of local adaptation. Nat. Rev. Genet. 14, 807–820 (2013).
Martínez-Berdeja, A. et al. Functional variants of DOG1 control seed chilling responses and variation in seasonal life-history strategies in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 117, 2526–2534 (2020).
Fournier-Level, A. et al. A map of local adaptation in Arabidopsis thaliana. Science (1979) 334, 86–89 (2011).
Exposito-Alonso, M. et al. Genomic basis and evolutionary potential for extreme drought adaptation in Arabidopsis thaliana. Nat. Ecol. Evol. 2, 352–358 (2017).
Wellband, K. et al. Chromosomal fusion and life history-associated genomic variation contribute to within-river local adaptation of Atlantic salmon. Mol. Ecol. 28, 1439–1459 (2019).
Bergland, A. O., Behrman, E. L., O’Brien, K. R., Schmidt, P. S. & Petrov, D. A. Genomic evidence of rapid and stable adaptive oscillations over seasonal time scales in Drosophila. PLoS Genet. 10, e1004775 (2014).
Machado, H. E. et al. Broad geographic sampling reveals the shared basis and environmental correlates of seasonal adaptation in Drosophila. Elife 10, e67577 (2021).
Collinge, J. E., Anderson, A. R., Weeks, A. R., Johnson, T. K. & McKechnie, S. W. Latitudinal and cold-tolerance variation associate with DNA repeat-number variation in the hsr-omega RNA gene of Drosophila melanogaster. Heredity 101, 260–270 (2008).
Pool, J. E., Braun, D. T. & Lack, J. B. Parallel evolution of cold tolerance within Drosophila melanogaster. Mol. Biol. Evol. 34, 349–360 (2017).
Durmaz, E., Benson, C., Kapun, M., Schmidt, P. & Flatt, T. An inversion supergene in Drosophila underpins latitudinal clines in survival traits. J. Evol. Biol. 31, 1354–1364 (2018).
Zare, F., Dow, M., Monteleone, N., Hosny, A. & Nabavi, S. An evaluation of copy number variation detection tools for cancer using whole exome sequencing data. BMC Bioinform. 18, 1–13 (2017).
Gabrielaite, M. et al. A comparison of tools for copy-number variation detection in germline whole exome and whole genome sequencing data. Cancers 13, 6283 (2021).
Mérot, C., Oomen, R. A., Tigano, A. & Wellenreuther, M. A roadmap for understanding the evolutionary significance of structural genomic variation. Trends Ecol. Evol. 35, 561–572 (2020).
Steenwyk, J. & Rokas, A. Extensive copy number variation in fermentation-related genes among Saccharomyces cerevisiae wine strains. G3: Genes Genomes Genet. 7, 1475–1485 (2017).
O’Neill, M. J. & O’Neill, R. J. Sex chromosome repeats tip the balance towards speciation. Mol. Ecol. 27, 3783–3798 (2018).
Hull, R. M., Cruz, C., Jack, C.V., & Houseley, J. Environmental change drives accelerated adaptation through stimulated copy number variation. PLoS Biol. 15, e2001333 (2017).
Whale, A. J., King, M., Hull, R. M., Krueger, F. & Houseley, J. Stimulation of adaptive gene amplification by origin firing under replication fork constraint. Nucleic Acids Res. 50, 915–936 (2022).
Tigano, A., Reiertsen, T. K., Walters, J. R. & Friesen, V. L. A complex copy number variant underlies differences in both colour plumage and cold adaptation in a dimorphic seabird. Preprint at bioRxiv https://doi.org/10.1101/507384 (2018).
Dorant, Y. et al. Copy number variants outperform SNPs to reveal genotype–temperature association in a marine species. Mol. Ecol. 29, 4765–4782 (2020).
Iantorno, S. A. et al. Gene expression in Leishmania is regulated predominantly by gene dosage. mBio 8, e01393–17 (2017).
Wang, Y., Tan, X. & Paterson, A. H. Different patterns of gene structure divergence following gene duplication in Arabidopsis. BMC Genom. 14, 1–9 (2013).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Ghavi-Helm, Y. et al. Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression. Nat. Genet. 51, 1272–1282 (2019).
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nat. Rev. Genet. 10, 551–564 (2009).
Lu, P. et al. Analysis of Arabidopsis genome-wide variations before and after meiosis and meiotic recombination by resequencing Landsberg erecta and all four products of a single meiosis. Genome Res. 22, 508–518 (2012).
Klein, S. J., O’neill, R. J., Klein, S. J. & O’neill, R. J. Transposable elements: genome innovation, chromosome diversity, and centromere conflict. Chromosome Res. 26, 5–23 (2018).
Verdin, H. et al. Microhomology-mediated mechanisms underlie non-recurrent disease-causing microdeletions of the FOXL2 gene or its regulatory domain. PLoS Genet. 9, e1003358 (2013).
Zhang, F., Gu, W., Hurles, M. E. & Lupski, J. R. Copy number variation in human health, disease, and evolution. Annu. Rev. Genom. Hum. Genet. 10, 451–481 (2009).
Nitcher, R., Distelfeld, A., Tan, C., Yan, L. & Dubcovsky, J. Increased copy number at the HvFT1 locus is associated with accelerated flowering time in barley. Mol. Genet. Genom. 288, 261–275 (2013).
Díaz, A., Zikhali, M., Turner, A. S., Isaac, P. & Laurie, D. A. Copy number variation affecting the photoperiod-B1 and vernalization-A1 genes is associated with altered flowering time in wheat (Triticum aestivum). PLoS ONE 7, e33234 (2012).
Assogba, B. S. et al. The ace-1 locus is amplified in all resistant Anopheles gambiae mosquitoes: fitness consequences of homogeneous and heterogeneous duplications. PLoS Biol. 14, e2000618 (2016).
Gimenez, S. et al. Adaptation by copy number variation increases insecticide resistance in the fall armyworm. Commun. Biol. 3, 1–10 (2020).
Todd, R. T. & Selmecki, A. Expandable and reversible copy number amplification drives rapid adaptation to antifungal drugs. Elife 9, 1–33 (2020).
Stalder, L., Oggenfuss, U., Mohd-Assaad, N. & Croll, D. The population genetics of adaptation through copy number variation in a fungal plant pathogen. Mol. Ecol. https://doi.org/10.1111/MEC.16435 (2022).
Farrer, R. A. et al. Chromosomal copy number variation, selection and uneven rates of recombination reveal cryptic genome diversity linked to pathogenicity. PLoS Genet. 9, e1003703 (2013).
Steenwyk, J. L., Soghigian, J. S., Perfect, J. R. & Gibbons, J. G. Copy number variation contributes to cryptic genetic variation in outbreak lineages of Cryptococcus gattii from the North American Pacific Northwest. BMC Genom. 17, 1–13 (2016).
Hong, J. & Gresham, D. Molecular specificity, convergence and constraint shape adaptive evolution in nutrient-poor environments. PLoS Genet. 10, e1004041 (2014).
Madden, L. V. & Wheelis, M. The threat of plant pathogens as weapons against U.S. crops. Annu. Rev. Phytopathol. 41, 155–176 (2003).
Chaloner, T. M., Gurr, S. J. & Bebber, D. P. Plant pathogen infection risk tracks global crop yields under climate change. Nat. Clim. Chang. 11, 710–715 (2021).
Mora, C. et al. Over half of known human pathogenic diseases can be aggravated by climate change. Nat. Clim. Chang. https://doi.org/10.1038/s41558-022-01426-1 (2022).
Torriani, S. F. F. et al. Zymoseptoria tritici: a major threat to wheat production, integrated approaches to control. Fungal Genet. Biol. 79, 8–12 (2015).
Feurtey, A. et al. A thousand-genome panel retraces the global spread and adaptation of a major fungal crop pathogen. Nat. Commun. 14, 1–15 (2023).
Badet, T., Fouché, S., Hartmann, F. E., Zala, M. & Croll, D. Machine-learning predicts genomic determinants of meiosis-driven structural variation in a eukaryotic pathogen. Nat. Commun. 12, 1–14 (2021).
Badet, T., Oggenfuss, U., Abraham, L., McDonald, B. A. & Croll, D. A 19-isolate reference-quality global pangenome for the fungal wheat pathogen Zymoseptoria tritici. BMC Biol. 18, 1–18 (2020).
Oggenfuss, U. et al. A population-level invasion by transposable elements triggers genome expansion in a fungal pathogen. Elife 10, e69249 (2021).
Galagan, J. E. & Selker, E. U. RIP: The evolutionary cost of genome defense. Trends Genet. 20, 417–423 (2004).
Hartmann, F. E. & Croll, D. Distinct trajectories of massive recent gene gains and losses in populations of a microbial eukaryotic pathogen. Mol. Biol. Evol. 34, 2808–2822 (2017).
Abraham, L. N., Oggenfuss, U. & Croll, D. Population-level transposable element expression dynamics influence trait evolution in a fungal crop pathogen. MBio 15, e02840-23 (2024).
Croll, D., Zala, M., & McDonald, B. A. Breakage-fusion-bridge cycles and large insertions contribute to the rapid evolution of accessory chromosomes in a fungal pathogen. PLoS Genet. 9, e1003567 (2013).
Moller, M. et al. Recent loss of the Dim2 DNA methyltransferase decreases mutation rate in repeats and changes evolutionary trajectory in a fungal pathogen. PLoS Genet. 17, e1009448 (2021).
Dutta, A., Hartmann, F. E., Francisco, C. S., McDonald, B. A. & Croll, D. Mapping the adaptive landscape of a major agricultural pathogen reveals evolutionary constraints across heterogeneous environments. ISME J. 15, 1402–1419 (2021).
Jing, H. & Lin, H. Sirtuins in epigenetic regulation. Chem. Rev. 115, 2350–2375 (2015).
North, B. J. & Verdin, E. Protein family review Sirtuins: Sir2-related NAD-dependent protein deacetylases. Genome Biol. 5, 224 (2004).
Smith, K. M. et al. The fungus Neurospora crassa displays telomeric silencing mediated by multiple sirtuins and by methylation of histone H3 lysine 9. Epigenet. Chromatin 1, 1–20 (2008).
Vogan, A. A. et al. The Enterprise, a massive transposon carrying Spok meiotic drive genes. Genome Res. 31, 789–798 (2021).
Gluck-Thaler, E. et al. Giant starship elements mobilize accessory genes in fungal genomes. Mol. Biol. Evol. 39, msac109 (2022).
Calvo, A. M., Lohmar, J. M., Ibarra, B. & Satterlee, T. 18 Velvet regulation of fungal development. In Growth, Differentiation and Sexuality (ed Wendland, J.) 475–497, The Mycota, vol 1 (Springer, Cham, 2016). https://doi.org/10.1007/978-3-319-25844-7_18.
Tiley, A. M. M., White, H. J., Foster, G. D. & Bailey, A. M. The ZtvelB gene is required for vegetative growth and sporulation in the wheat pathogen Zymoseptoria tritici. Front. Microbiol. 10, 2210 (2019).
Fones, H. & Gurr, S. The impact of Septoria tritici blotch disease on wheat: an EU perspective. Fungal Genet. Biol. 79, 3–7 (2015).
Lande, R. & Shannon, S. The role of genetic variation in adaptation and population persistence in a changing environment. Evolution 50, 434 (1996).
Kutz, S. J., Hoberg, E. P., Polley, L. & Jenkins, E. J. Global warming is changing the dynamics of Arctic host–parasite systems. Proc. R. Soc. B: Biol. Sci. 272, 2571–2576 (2005).
Hueffer, K., O’Hara, T. M. & Follmann, E. H. Adaptation of mammalian host–pathogen interactions in a changing arctic environment. Acta Vet. Scand. 53, https://doi.org/10.1186/1751-0147-53-17 (2011).
Laaksonen, S. et al. Climate change promotes the emergence of serious disease outbreaks of filarioid nematodes. Ecohealth 7, 7–13 (2010).
Conrad, D. F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2009).
Sandve, S. R., Rohlfs, R. V. & Hvidsten, T. R. Subfunctionalization versus neofunctionalization after whole-genome duplication. Nat. Genet. 50, 908–909 (2018).
Ames, R. M. et al. Gene duplication and environmental adaptation within yeast populations. Genome Biol. Evol. 2, 591–601 (2010).
Kondrashov, F. A. Gene duplication as a mechanism of genomic adaptation to a changing environment. Proc. R. Soc. B: Biol. Sci. 279, 5048–5057 (2012).
Linardopoulou, E. V. et al. Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 437, 94–100 (2005).
Liu, P. et al. Frequency of nonallelic homologous recombination is correlated with length of homology: evidence that ectopic synapsis precedes ectopic crossing-over. Am. J. Hum. Genet. 89, 580–588 (2011).
Stukenbrock, E. H. & Dutheil, J. Y. Fine-scale recombination maps of fungal plant pathogens reveal dynamic recombination landscapes and intragenic hotspots. Genetics 208, 1209–1229 (2018).
Croll, D., Lendenmann, M. H., Stewart, E. & McDonald, B. A. The impact of recombination hotspots on genome evolution of a fungal plant pathogen. Genetics 201, 1213–1228 (2015).
Fudal, I. et al. Repeat-Induced Point Mutation (RIP) as an alternative mechanism of evolution toward virulence in Leptosphaeria maculans. Molecular Plant-Microbe Interactions. 22, 932–941 (2009).
Rouxel, T. et al. Effector diversification within compartments of the Leptosphaeria maculans genome affected by Repeat-Induced Point mutations. Nat. Commun. 2, 1–10 (2011).
Bratlie, M. S. et al. Gene duplications in prokaryotes can be associated with environmental adaptation. BMC Genom. 11, 1–17 (2010).
Xu, S. et al. Where whole-genome duplication is most beneficial: adaptation of mangroves to a wide salinity range between land and sea. Mol. Ecol. https://doi.org/10.1111/MEC.16320 (2021).
Xu, Y. C. & Guo, Y. L. Less is more, natural loss-of-function mutation is a strategy for adaptation. Plant Commun. 1, 100103 (2020).
Albalat, R. & Cañestro, C. Evolution by gene loss. Nat. Rev. Genet. 17, 379–391 (2016).
Greenberg, A. J., Moran, J. R., Coyne, J. A. & Wu, C. I. Ecological adaptation during incipient speciation revealed by precise gene replacement. Science (1979) 302, 1754–1757 (2003).
Prunier, J. et al. Gene copy number variations in adaptive evolution: the genomic distribution of gene copy number variations revealed by genetic mapping and their adaptive role in an undomesticated species, white spruce (Picea glauca). Mol. Ecol. 26, 5989–6001 (2017).
Monroe, J. G. et al. Drought adaptation in Arabidopsis thaliana by extensive genetic loss-of-function. Elife 7, e41038 (2018).
Castagnone-Sereno, P. et al. Gene copy number variations as signatures of adaptive evolution in the parthenogenetic, plant–parasitic nematode Meloidogyne incognita. Mol. Ecol. 28, 2559–2572 (2019).
Huelsmann, M. et al. Genes lost during the transition from land to water in cetaceans highlight genomic changes associated with aquatic adaptations. Sci. Adv. 5, 6671–6696 (2019).
Rokas, A., Mead, M. E., Steenwyk, J. L., Raja, H. A. & Oberlies, N. H. Biosynthetic gene clusters and the evolution of fungal chemodiversity. Nat. Prod. Rep. 37, 868–878 (2020).
Osbourn, A. Secondary metabolic gene clusters: evolutionary toolkits for chemical innovation. Trends Genet. 26, 449–457 (2010).
Hartmann, F. E., Vonlanthen, T., Singh, N. K., Mcdonald, M. & Milgate, A. The complex genomic basis of rapid convergent adaptation to pesticides across continents in a fungal plant pathogen. Mol. Ecol. https://doi.org/10.1111/mec.15737 (2020).
Tralamazza, S. M. et al. Complex evolutionary origins of specialized metabolite gene cluster diversity among the plant pathogenic fungi of the Fusarium graminearum species complex. Genome Biol. Evol. 11, 3106–3122 (2019).
Valero-Jiménez, C. A. et al. Dynamics in secondary metabolite gene clusters in otherwise highly syntenic and stable genomes in the fungal genus Botrytis. Genome Biol. Evol. 12, 2491–2507 (2020).
Krishnan, P. et al. Transposable element insertions shape gene regulation and melanin production in a fungal pathogen of wheat. BMC Biol. 16, 1–18 (2018).
Wong, S. & Wolfe, K. H. Birth of a metabolic gene cluster in yeast by adaptive gene relocation. Nat. Genet. 37, 777–782 (2005).
Morris, J. J., Lenski, R. E. & Zinser, E. R. The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. mBio. 3, e00036-12 (2012).
Fouché, S., Plissonneau, C. & Croll, D. The birth and death of effectors in rapidly evolving filamentous pathogen genomes. Curr. Opin. Microbiol. 46, 34–42 (2018).
Sánchez-Vallet, A. et al. The genome biology of effector gene evolution in filamentous plant pathogens. Annu. Rev. Phytopathol. 56, 21–40 (2018).
Yacoubi, I. et al. New insight into the North-African durum wheat biodiversity: phenotypic variations for adaptive and agronomic traits. Genet. Resour. Crop Evol. 67, 445–455 (2020).
Hartmann, F. E., Sánchez-Vallet, A., McDonald, B. A. & Croll, D. A fungal wheat pathogen evolved host specialization by extensive chromosomal rearrangements. ISME J. 11, 1189–1204 (2017).
Zess Id, E. K. et al. Regressive evolution of an effector following a host jump in the Irish potato famine pathogen lineage. PLoS Pathog. 18, e1010918 (2022).
Vogan, A. A. et al. The Enterprise, a massive transposon carrying Spok meiotic drive genes. Genome Research 31, 789–798 (2021).
Urquhart, A. S., Chong, N. F., Yang, Y. & Idnurm, A. A large transposable element mediates metal resistance in the fungus Paecilomyces variotii. Curr. Biol. 32, 937–950.e5 (2022).
Urquhart, A. S., Vogan, A. A., Gardiner, D. M. & Idnurm, A. Starships are active eukaryotic transposable elements mobilized by a new family of tyrosine recombinases. Proc. Natl Acad. Sci. USA 120, e2214521120 (2023).
Cayuela, H. et al. Thermal adaptation rather than demographic history drives genetic structure inferred by copy number variants in a marine fish. Mol. Ecol. 30, 1624–1641 (2021).
Benestan, L. et al. Seascape genomics provides evidence for thermal adaptation and current-mediated population structure in American lobster (Homarus americanus). Mol. Ecol. 25, 5073–5092 (2016).
Zhan, J. & McDonald, B. A. Thermal adaptation in the fungal pathogen Mycosphaerella graminicola. Mol. Ecol. 20, 1689–1701 (2011).
Lendenmann, M. H., Croll, D., Palma-Guerrero, J., Stewart, E. L. & Mcdonald, B. A. QTL mapping of temperature sensitivity reveals candidate genes for thermal adaptation and growth morphology in the plant pathogenic fungus Zymoseptoria tritici. Heredity 116, 384–394 (2016).
Boixel, A. L., Gélisse, S., Marcel, T. C. & Suffert, F. Differential tolerance of Zymoseptoria tritici to altered optimal moisture conditions during the early stages of wheat infection. J. Plant Pathol. 104, 495–507 (2022).
Ohtani, H. & Iwasaki, Y. W. Rewiring of chromatin state and gene expression by transposable elements. Dev. Growth Differ. 63, 262–273 (2021).
Sauve, A. A. Sirtuin chemical mechanisms. Biochim. Biophys. Acta (BBA)—Proteins Proteom. 1804, 1591–1603 (2010).
Kawauchi, M., Nishiura, M. & Iwashita, K. Fungus-specific sirtuin HstD coordinates secondary metabolism and development through control of LaeA. Eukaryot. Cell 12, 1087–1096 (2013).
Itoh, E. et al. Sirtuin e is a fungal global transcriptional regulator that determines the transition from the primary growth to the stationary phase. J. Biol. Chem. 292, 11043–11054 (2017).
Campbell-Staton, S. C. et al. Parallel selection on thermal physiology facilitates repeated adaptation of city lizards to urban heat islands. Nat. Ecol. Evol. 4, 652–658 (2020).
Feurtey, A. et al. Genome compartmentalization predates species divergence in the plant pathogen genus Zymoseptoria. BMC Genom. 21, 1–15 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Babadi, M. et al. Abstract 2287: precise common and rare germline CNV calling with GATK. Cancer Res. 78, 2287–2287 (2018).
Quinlan, A. R. BEDTools: the Swiss‐army tool for genome feature analysis. Curr. Protoc. Bioinform. 47, 11.12.1–11.12.34 (2014).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 1–13 (2019).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, 1–4 (2021).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Dray, S. & Dufour, A. B. The ade4 Package: implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Wickham, H. ggplot2. Wiley Interdiscip. Rev. Comput. Stat. 3, 180–185 (2011).
Goudet, J. hierfstat, a package for r to compute and test hierarchical F-statistics. Mol. Ecol. Notes 5, 184–186 (2005).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Monier, B., Casstevens, T. M., Bradbury, P. J. & Buckler, E. S. rTASSEL: an R interface to TASSEL for analyzing genomic diversity. J. Open Source Softw. 7, 4530 (2022).
Palma-Guerrero, J. et al. Comparative transcriptomic analyses of Zymoseptoria tritici strains show complex lifestyle transitions and intraspecific variability in transcription profiles. Mol. Plant Pathol. 17, 845–859 (2016).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–14 (2019).
Blin, K. et al. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 47, W81–W87 (2019).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Alexa, A. R. J. Gene set enrichment analysis with topGO. 1–26 http://www.mpi-sb.mpg.de/∼alexa (2009).
Bruls, M., Huizing, K. & van Wijk, J. J. Squarified Treemaps 33–42 (2000) https://doi.org/10.1007/978-3-7091-6783-0_4.
Schotanus, K. et al. Histone modifications rather than the novel regional centromeres of Zymoseptoria tritici distinguish core and accessory chromosomes. Epigenet. Chromatin 8, 1–18 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Marçais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinform. 6, 1–11 (2005).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform. https://doi.org/10.1093/bib/bbx108 (2017).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
R. Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2023).
Bretz, F., Hothorn, T. & Westfall, P. Multiple Comparisons Using R (Chapman and Hall/CRC, 2016).
Maintainer, K. & Kolde, R. Package ‘pheatmap’. https://cran.r-project.org/web/packages/pheatmap/pheatmap.pdf (2018).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Guy, L., Roat Kultima, J. & Andersson, S. G. E. genoPlotR: comparative gene and genome visualization in R. Bioinformatics 26, 2334–2335 (2010).
Hackl, T. et al. gggenomes: A Grammar of Graphics for Comparative Genomics. https://thackl.github.io/gggenomes/ (2023)
Taiyun Wei, M. et al. Package ‘corrplot’ Title Visualization of a Correlation Matrix https://cran.r-project.org/web/packages/corrplot/corrplot.pdf (2017).
Pinheiro, J. & Bates, D. Package ‘Nlme’. https://bugs.r-project.org (2023).
Bates, D. et al. The Car Package https://rdrr.io/rforge/car/ (2007).
Bartón, K. Package ‘MuMIn’ Title Multi-Model Inference https://cran.r-project.org/web/packages/MuMIn/MuMIn.pdf (2023).
Tralamazza, S. & Croll, D. Copy number variation introduced by a massive mobile element facilitates global thermal adaptation in a fungal wheat pathogen—Supplementary Data files. Zenodo https://doi.org/10.5281/zenodo.11616290 (2024).
Tralamazza, S. & Croll, D. Copy number variation introduced by a massive mobile element underpins global thermal adaptation in a fungal wheat pathogen—scripts. Zenodo https://doi.org/10.5281/zenodo.8344847 (2024).
Palma-Guerrero, J. et al. Comparative transcriptome analyses in Zymoseptoria tritici reveal significant differences in gene expression among strains during plant infection. Mol. Plant–Microbe Interact. 30, 231–244 (2017).
Acknowledgements
We would like to thank Thomas Badet for facilitating access to genome assembly data. We thank lab members for their thoughtful discussions and input. E.G.-T. was supported by funding from the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement (grant number 890630). D.C. was supported by the Swiss National Science Foundation grants 177052 and 205401.
Author information
Authors and Affiliations
Contributions
S.M.T. and D.C. conceived the study; S.M.T. and E.G.-T. performed analyses; A.F. provided datasets; D.C. provided funding and supervised the work; S.M.T. and D.C. wrote the manuscript with input from E.G.-T. and A.F.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jesper Svedberg, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tralamazza, S.M., Gluck-Thaler, E., Feurtey, A. et al. Copy number variation introduced by a massive mobile element facilitates global thermal adaptation in a fungal wheat pathogen. Nat Commun 15, 5728 (2024). https://doi.org/10.1038/s41467-024-49913-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-49913-7
This article is cited by
-
Transposable elements hitchhike on Starships across fungal genomes
Nature Communications (2026)
-
Transposable elements: a key piece in the genomic evolution and adaptation of Myrtaceae species
Mobile DNA (2025)
-
The role of gene copy number variation in antimicrobial resistance in human fungal pathogens
npj Antimicrobials and Resistance (2025)
-
Starship giant transposons dominate plastic genomic regions in a fungal plant pathogen and drive virulence evolution
Nature Communications (2025)
-
Historic transposon mobilisation waves create distinct pools of adaptive variants in a major crop pathogen
Nature Communications (2025)