Abstract
The detection and tracking of metastatic cancer over the lifetime of a patient remains a major challenge in clinical trials and real-world care. Advances in deep learning combined with massive datasets may enable the development of tools that can address this challenge. We present NYUMets-Brain, the world’s largest, longitudinal, real-world dataset of cancer consisting of the imaging, clinical follow-up, and medical management of 1,429 patients. Using this dataset we developed Segmentation-Through-Time, a deep neural network which explicitly utilizes the longitudinal structure of the data and obtained state-of-the-art results at small (<10 mm3) metastases detection and segmentation. We also demonstrate that the monthly rate of change of brain metastases over time are strongly predictive of overall survival (HR 1.27, 95%CI 1.18-1.38). We are releasing the dataset, codebase, and model weights for other cancer researchers to build upon these results and to serve as a public benchmark.
Similar content being viewed by others
Introduction
With the advancement of modern electronic health records (EHRs) and picture and archiving systems (PACs), it is possible to study the detailed, real-world outcomes of cancer patients and their tumors over the entire course of care1,2. The complexity of metastatic cancer makes this challenging, as tracking multiple metastatic lesions and diverse treatments spanning medications, radiation, and surgery is a formidable challenge that existing analytical systems struggle to perform. In the specific case of the imaging of brain metastases, MRI is the gold standard for tracking metastatic cancer3. Several scoring systems (RECIST4,5 and RANO-BM6) based on MRI have been created as a means of simplifying and standardizing radiological information, and have been shown to be reliable measures of treatment failure. However, these systems are not without their flaws5, and rely on time-consuming, expert assessments for their application utilizing only the patient’s imaging.
Recent advances in artificial intelligence (AI) and deep learning have led to a renewed interest in automated tools for tumor detection and tracking7,8 that can provide quantitative metrics for an arbitrary number of lesions. This interest has been dampened by a lack of large, real-world benchmarks to assess the utility of proposed solutions, as well as large-scale datasets to train upon. The development of DNNs for other use cases in computer vision typically involves large datasets and public benchmarks9,10,11. These open benchmarks and datasets have had a transformative impact on research and development, and the data-centric AI movement cites these as being the driving force behind the advance of modern AI systems12,13. Efforts to build large datasets, develop benchmarks, and publicly release them present substantial logistical challenges as well as concerns over the protection of patient health information (PHI). The few biomedical datasets and benchmarks that do exist are much smaller, hundreds rather than millions, of samples14,15,16 and frequently lack accompanying clinical, genomic, and therapeutic information. The impact of building even these smaller biomedical datasets and benchmarks can be enormous, with benchmarks such as the brain tumor segmentation (BraTS) dataset16 or the MIMIC-III17 dataset having a transformative impact on the development of medical AI models in a uniform and reproducible manner. Without these large, public benchmarks and datasets, medical DNNs run the risk of overfitting and lacking generalizability, thereby limiting their real-world potential18.
The data-centric AI movement12,13 emphasizes this transformative value of datasets to drive progress. Data-centric AI approaches emphasize that novel datasets drive algorithmic advances, and can lead to the development of entirely new classes of DNNs by incorporating new, diverse datatypes (genomic, imaging, or clinical) or longitudinal information1,2. For biomedical imaging, longitudinal information represents a unique departure due to the significant change in its extraction, transformation, and loading, which resembles sparsely sampled video data due to the inclusion of an additional dimension of time across irregular intervals19. First-order statistics derived from the dynamics of patients and tumors changing over time could provide a source of biomarkers for treatment progression, response, and survival. Just as cancer evolution has been increasingly recognized as a key driver of cancer on a molecular level20, dynamic changes in disease at the level of phenotype may signal key clinical events. Prostate-specific antigen (PSA) dynamics are definitive of biochemical treatment failure for prostate cancer21, tumor size change was noted to be predictive of progression-free survival (PFS) in the SELECT trial21,22, and other tumor growth metrics were noted as potential biomarkers for both metastatic colorectal cancer (CRC)23 and non-small cell lung cancer (NSCLC)24.
In this work, we built and publicly released the NYUMets-Brain dataset as the initial phase of a larger initiative to build public, multimodal, longitudinal benchmarks for cancer research and AI model development. Brain metastases are the most common type of brain cancer, and anywhere from 10–30% of cancer patients are expected to develop them, resulting in 100,000–240,000 new cases per year25,26. Successful treatment of brain metastases is critical, and if untreated, the average survival is short. We hypothesized that a large, multimodal, longitudinal dataset and benchmark combined with AI tools for handling it could improve upon existing solutions in brain metastasis detection and tracking, and investigated whether quantitative measurements of metastases over time could serve as a novel source of biomarkers.
Results
Dataset creation and preprocessing
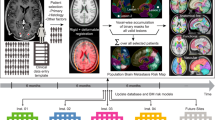
As part of this IRB-approved study, we worked in concert with NYU Langone’s Center for Advanced Radiosurgery to identify patients with metastatic cancer to the brain and to extract their clinical and radiographic data from disparate clinical records (Fig. 1A). The resulting dataset, NYUMets-Brain, consists of 1429 patients with a median of six imaging timepoints per patient (IQR 3–11) over a median of 8.4 months of care (IQR 2.8–22.8 months), resulting in a dataset of 8366 MRI studies and a total of 4860 timepoints of clinical follow-up (Extended Data Tables 1, 2). The imaging studies include 2367 expert segmentations, created for targeting radiosurgery treatment, in addition to T1, T1 post contrast, T2, and FLAIR MRI sequences. Out of these expert segmentations, 921 segmentations from 901 patients were verified a second time by an expert (neurosurgeon) after data preprocessing. The dataset was de-identified and standardized (see Methods for details on deidentification and standardization) to facilitate accession across timepoints and data modalities prior to sharing on Amazon S3.
A The NYUMets dataset brought together multiple resources within a large academic medical center spread across the Cancer Center clinical database, Radiology picture and archiving system (PACS), Radiation Oncology treatment planning system, and the hospital electronic health records (EHR) systems. B We designed a deep learning model to explicitly incorporate the longitudinal segmentations and imaging data called segmentation-through time ›(STT). STT has a UNet backbone with a recurrent neural network (RNN) to carry learned representations forward across timepoints. While there are many possible ways of utilizing this unique, real-world longitudinal data, we present this as an initial benchmark attempt. In order to maximize the utility of the unlabeled data, we also experimented with self-supervised learning (SSL), where we first pretrained the STT and UNet models with a reconstruction task before finetuning it for the segmentation task.
Creation of longitudinal imaging API
Studies of tumor dynamics and evolution have been performed in a wide range of model systems, both in vitro and in vivo, to investigate these dynamics and how they interact with treatments20,27,28. Recently, with the benefit of advanced imaging, serial liquid biopsies, and initiatives like TRACERx, it has become feasible to track the radiographic and genomic dynamics of cancer in real-world clinical populations28,29,30. To facilitate the tracking of metastatic cancer on a per-metastasis basis, we developed an Application Programming Interface (API) for interacting with this dataset (Fig. 1B) and potentially other longitudinal cancer datasets as microscopic evolutionary changes can be associated with phenotypic correlates. In order to automatically segment and track metastases over longitudinal scans, we built a custom API built on top of the MONAI framework that includes a custom dataset class for feeding longitudinal scans into a neural network during training, evaluation, and inference, functions for image coregistration across timepoints, tumor identification, and matching algorithms, as well as novel clinically meaningful metrics that measure tumor burden and growth both by scan and by an individual tumor. We demonstrate the capabilities of this API through the inference of 4 sequential studies from a single patient in Fig. 2.
A The NYUMets API provides autosegmentation and tracking of metastases across timepoints, enabling more efficient investigation of patient-level and population-level cancer dynamics. Metastase segmentations were inferred using the SSL STT model and longitudinal T1 post-contrast inputs. The model-generated segmentations are shown in pink. Each individual tumor segmentation is assigned a color. B Lines corresponding to each tumor’s volume over time are color-matched to (A). The dotted lines represent the volumes from gold-standard segmentations. C The total tumor count, shown here, and other aggregative metrics, such as total intracranial volume, can be plotted over time as well.
Development of a novel model architecture for segmentation-through time
We developed a deep neural network for longitudinal semantic segmentation of metastatic cancer, segmentation-through time (STT), that explicitly incorporates the history of each metastases as it identifies existing and new lesions (Fig. 1B) incorporating LSTM layers between the embedding layers of the popular UNet architecture (see Methods for details regarding model architecture and training)31,32. This STT model was trained and evaluated on a longitudinal sequence of images of up to eight timepoints, and compared to the basic UNet architecture that serves as the STT model architecture’s backbone. In addition to supervised learning, we evaluated whether self-supervised learning (SSL) could improve the segmentation and detection of brain metastases. Full details on the methods for training and evaluation of the models, including the pretraining and finetuning of self-supervised learning networks, can be found in the Methods and Extended Fig. 4. In addition to investigating the effects of different model architectures and training paradigms, we investigated the impact of multisequence inputs (T1 post-contrast vs T1 post-contrast and T2 weighted images) as well as label quality during training (gold and silver labels vs gold only labels). Labels of “gold” quality were reviewed and adjusted as needed by an expert physician, while “silver” quality labels were not reviewed nor adjusted after preprocessing. These results can be found in the Extended Tables and Extended Fig. 7.
Benchmarking of longitudinal metastasis tracking using API
We created three clinically relevant metrics to evaluate our model’s performance on tracking metastases across longitudinal scans: change in total tumor volume, change in per-tumor volume, and change in total tumor count. Within a single timepoint, we also calculated the total tumor volume, the per-tumor volumes, and the total tumor count. We found that the UNet model trained with SSL had the lowest mean absolute error for total tumor volume (2,540.857 mm3), while the STT model trained with SSL and evaluated on longitudinal data had the lowest mean absolute error for change in per-tumor volume (210.026 mm3). The UNet model trained with SSL had the lowest mean absolute error for change in tumor count (6.464). Full results of both longitudinal and single-timepoint clinical metrics can be found in Extended Table 17 and Extended Fig. 7.
Benchmarking evaluation of metastasis segmentation
The UNet model pretrained using SSL had the highest median scan Dice similarity coefficient of 0.667 and the lowest median scan Hausdorff distance of 24.356. Stratified Dice similarity scores by volume showed that STT pretrained using SSL and finetuned on T1 post-contrast and T2 images with both Silver and Gold segmentations performed the best at segmenting metastases under 10 mm3 (mean Dice = 0.418) (Fig. 3A). The best mean Dice coefficients for tumors volumes 10–100 mm3, 100–1000 mm3, 1000–10,000 mm3, and over 10,000 m3 were 0.517, 0.680, 0.766, and 0.804, respectively. Similar to prior studies, STT segmentations demonstrated a linear relationship between Dice and individual tumor volume (R2 = 0.221, p < 0.001) as well as between predicted and target tumor volumes (R2 = 0.872, p < 0.001) (Fig. 3B). All benchmark results for segmentation can be found in Extended Table 6 and Extended Fig. 7.
A We evaluated the models across three primary metrics: Dice similarity coefficient, recall, and false positives per scan (FP/scan). Our models show state-of-the-art performance on segmentation and detection of very small (<10 mm3) metastases on the NYUMets test dataset. N = 4 experiments with the four training data variations (T1 post vs T1 post + T2, gold labels vs. gold + silver labels). B We plotted the relationship between the Dice similarity coefficient and individual tumor volumes, as well as predicted tumor volumes and individual tumor volumes (one green dot = one NYUMets segmented metastasis) after inference using the STT model. These relationships were also demonstrated on our external test sets, Stanford BrainMetsShare and UCSF Brain Metastases Stereotactic Radiosurgery (BMSR) dataset, using the same model (one orange dot = one Stanford segmented metastasis). Pearson correlations were performed with the two-sided alternative hypothesis.
Benchmarking evaluation of metastasis detection
In order to identify whether a tumor was successfully detected (true positive), missed (false negative), or falsely present (false positive), as well as to determine whether a tumor persisted across two sequential timepoints, we developed a tumor matching procedure that uses two heuristics, centroid-to-centroid distance and intersection over reference (IoR), and based on the closest match above certain thresholds for each, determines a matched pair of tumors (see Methods for details and Extended Fig. 4). The UNet model pretrained using SSL and finetuned on T1 post-contrast images had the highest median precision (0.909) and F1 score (0.8) aggregated by study. Several models achieved a median recall of 0.667. The lowest mean false positive rate aggregated across all tumors was measured at 1.416. Precision, recall, and false positive rates all varied by tumor volume, with the highest precision and lowest false positive rate in tumors between 1000 and 10,000 mm3 (0.966 and 0.042, respectively) and the highest recall in tumors above 10,000 mm3 (1). In contrast, the best models’ precision, recall, and false positive rate on tumors below 10 mm3 was 0.095, 0.060, and 0.750, respectively. All benchmark results for detection can be found in Extended Table 7.
Evaluation of tumor segmentation and detection on external datasets
In order to evaluate the generalizability of our models, we evaluated our models on three external brain tumor datasets: the Stanford Brain Mets Share dataset33, the University of California San Francisco Brain Metastasis Stereotactic Radiosurgery (UCSF BMSR) Dataset34, the 2021 Brain Tumor Segmentation (BraTS) dataset14,16,35. All three datasets contain data collected from external institutions, and the BraTS dataset contains MRI images of glioblastoma multiforme, a primary brain tumor, thus evaluating the generalizability of the models across different diseases. Descriptions of both datasets can be found in Extended Fig. 2, individual tumor counts across volumes can be found in Extended Data Table 5 for the NYUMets, Stanford, and UCSF datasets, and full data processing and evaluation details can be found in Methods.
The UNet model pretrained using SSL and finetuned on gold-standard T1 post-contrast images had the best per-scan segmentation accuracy on Stanford Brain Mets Share with a Dice similarity coefficient of 0.284. This model also had the highest median precision (0.273), F1 score (0.513), and false positive rate (6) aggregated by scan. The STT model trained without SSL and finetuned on both gold- and silver-standard labels had the best overall per-tumor Dice (0.514), while the UNet model finetuned on both gold and silver labeled T1 post-contrast images had the highest Dice (0.262) for tumors less than 10 mm3. On the UCSF BMSR Dataset, the UNet model pretrained with SSL and T1 post-contrast gold and silver standard images performs the best with an overall per-tumor Dice of 0.645 and a per-tumor Dice of 0.467 on metastases less than 10 mm3. As the BraTS dataset contains segmentations of a single tumor, we evaluated our model’s segmentation quality for detecting tumor by combining the necrotic tumor core and GD-enhancing tumor classes. When using SSL for pretraining, the UNet model outperformed the STT model, with a Dice similarity coefficient of 0.597. Extended Data Tables 11 through 21 report the full results for the Stanford, UCSF, and BraTS datasets. Similar to the results on the NYUMets dataset, STT segmentations on the Stanford dataset demonstrated a linear relationship between Dice and individual tumor volume (R2 = 0.509, p < 0.001) as well as between predicted and target tumor volumes (R2 = 0.989, p < 0.001) (Fig. 3B).
Cancer dynamics strongly predict the overall survival of patients with metastatic cancer
To characterize the dynamic features of cancer, we evaluated how the burden of brain disease changed over time, specifically evaluating rates of new tumor influx and changes in tumor volume over time. We evaluated these changes for each patient over the shortest available time interval between expert segmented images (between initial and second GKS: median 5.4 months, IQR 2.5−10.8 months) and the longest available time interval between expert segmented images (between initial and last GKS: median 8.8 months, IQR 4.4−21.2) to create predictor variables for survival analysis. On a univariable proportional hazards regression of the shortest interval, survival was significantly associated with both the rate of additional tumors per month (HR 1.27 per doubling rate, 95% CI 1.21–1.33, p < 0.001, Fig. 4A, B) and change in volume per tumor per month (HR 1.17 per doubling rate, 95% CI 1.12–1.22, p < 0.001, Extended Fig. 9A, B). These effects became more pronounced over the longer interval, with HR of 1.53 per doubling rate of new tumors per month (95% CI 1.43–1.62, p < 0.001) and HR of 1.25 per doubling change of volume per tumor per month (95% CI 1.18–1.31, p < 0.001). See extended results for univariable analysis of traditional predictive factors, including age, functional status, and initial number of brain metastases (Extended Table 18). In a multivariable Cox proportional hazards model, including dynamic characteristics and traditional predictor variables, the rate of new tumors per month was the only variable associated with higher risk of death (HR 1.27 per doubling rate, 95% CI 1.18–1.38, p < 0.001) (Fig. 4C). For the longer time interval, there was an association with worse survival for both increase in new tumors per month (HR 1.47 per doubling rate, 95% CI 1.34–1.61, p < 0.001) and change in volume per tumor per month (HR 1.12 per doubling rate, 95% CI 1.03–1.20, p < 0.001) (Extended Fig. 9).
A Line graph demonstrating rates of new tumors per patient with the X-axis representing time in months and the y-axis as the number of cumulative tumors treated. Each line is an individual patient connecting two points: the total number of tumors treated to date at t_0 (initial gamma knife treatment) and t_1 (second gamma knife treatment), allowing the slope to equal the rate of new tumor influx, i.e., the number of new tumors developed between t_0 and t_1 divided by time in months. The median time between t_0 and t_1 was 5.5 months (IQR 2.5–10.8 months). Patients were segmented into quartiles based on the rate of tumor influx, each represented by a color (Quartile 1 = Blue, Quartile 2 = Red, Quartile 3 = Green, Quartile 4 = Yellow). B Kaplan–Meier curve demonstrating these rates of tumor influx were highly predictive of overall survival (log-rank: p < 0.001). C Forest plot of multivariable cox-regression analysis. Controlling for tumor histology, performance status, age, and initial tumor burden demonstrates that the rate of monthly tumor changes as being strongly predictive of OS (HR 1.27 per doubling rate of new tumor development, p < 0.001). The measure of the center for error bars is the hazard ratio, with bars extending to 95% CI.
Discussion
NYUMets-brain is one of the world’s largest oncology datasets, and the only large-scale public benchmarks for real-world brain metastases segmentation upon which we developed a novel approach to segmenting longitudinal imaging, STT. NYUMets-brain represents the beginning of a larger data-centric AI initiative to advance the use of AI in oncology by constructing public datasets, models, and benchmarks for tasks in cancer research and clinical care. In confirmation of our hypothesis that this unique dataset could potentially lead to the discovery of novel biomarkers, we intuitively found that the monthly rate of change in intracranial tumors is highly predictive of survival. As noted in a recent editorial in Radiology by JN Acosta et al2, there is a need for medical artificial intelligence (AI) that incorporates prior imaging, which is analogous to how physicians perceive and utilize medical imaging. Thinking about longitudinal imaging also empowers research to develop metrics that are clinically significant (e.g., changes in the number of tumors or changes in tumor volume) that necessarily require longitudinal data to compute. Longitudinal thinking and modeling can allow us as a cancer AI community to move beyond optimizing for technical metrics like detection and segmentation, to clinical assessments like progression, detection of new lesions, and response.
There are other first-order statistics for cancer prognosis, most notably prostate serum antigen (PSA) dynamics21, and this study is the first to extend those concepts to statistics calculated from radiological imaging. The finding that a dynamic measure of intracranial disease is highly predictive of overall survival is surprising, given that prior studies have indicated only a minority of deaths in patients with brain metastases are attributable to neurologic causes36. One explanation is that intracranial disease is not causing death, but serving as a marker for systemic disease. With more active systemic disease, a higher metastatic potential would be expected, resulting in an increased influx of new intracranial tumors. Given the high sensitivity of brain imaging, this could serve as a highly sensitive measure of systemic disease control.
We demonstrate the use of this unique dataset and interface to serve as a benchmark by training several deep learning models that demonstrate results at the segmentation and detection of metastatic tumors comparable to existing works33,37,38. Most importantly, we release in full the datasets, code, and trained models to enable other researchers to directly compare against our published results—something which is not possible until now with prior work holding the combination of datasets and codebases as proprietary. In addition to comparable segmentation and detection results within studies and across all metastasis sizes, our models show state-of-the-art detection of metastases less than 10 mm3. While prior studies define small metastases with different thresholds and define true positive detection criteria less rigorously, for metastases with a diameter less than 2.5 mm (equivalent spherical volume = 14.1 mm3), our detection of small metastasis is comparable with prior studies on our NYUMets dataset (recall = 0.12) and exceeds prior studies on the Stanford Brain Mets Share dataset (recall = 0.536). For metastases of this size, ref. 38 reported a recall of 0.147, while ref. 37 did not report their results for this volume. We are also the first to report segmentation of metastasis less than 10 mm3, with best mean Dice similarity coefficients of 0.41 on the NYUMets dataset, 0.262 on the Stanford Brain Mets Share dataset, and 0.467 on the UCSF BMSR Dataset. Finally, we also define and report novel longitudinal clinical metrics, improving the clinical applicability of these methods.
This study has several limitations worth noting. First, the underlying imaging dataset and population is limited to patients with metastatic cancer to the brain treated at a single institution and lack imaging data regarding systemic disease, including extracranial metastases There is also inherent heterogeneity in this patient sample which is not currently captured, such as molecular data, comorbidities, and interval open surgery. Second, several patients received their oncological care at other institutions where we did not have access to comprehensive records of their pharmacotherapy. Third, to de-identify the images we opted for skull stripping vs. more sophisticated forms of brain MRI deidentification (e.g., defacing), which induces a degree of noise to the de-identified dataset. Fourth, we did not exhaustively benchmark existing solutions for metastatic brain tumor segmentation and detection. Many existing works lack publicly available codebases, model weights, and/or datasets, and we viewed re-implementation of all of them as being potentially error-prone and out of scope due to the inherent non-determinism of deep learning approaches. Finally, while we report state-of-the-art results on small metastases, our overall results on metastasis segmentation and detection do not perform above other prior work. This is potentially due to inherent differences in our datasets, including the proportions of metastases of different sizes as well as the origin of our segmentations (which are derived from radiation planning software). There could also be differences in image quality, scan parameters, and/or approaches to image processing (such as other datasets’ lack of skull stripping, the process of which introduces noise but is necessary for deidentification). The lack of improvement when including T2 scans could be due to noise introduced during processing steps such as coregistration. Some of the deep learning approaches, which, as noted previously, are infeasible to reproduce without openly released codebases.
We hope that NYUMets encourages rethinking how we practice machine learning in the biomedical space to be more data-centric and more open. We note that for biomedical projects, the de facto standard is to not release data, models, and frequently even code39. Open datasets, open codebases, open models (weights), and most of all, open benchmarks can ensure scientific reproducibility and can accelerate the use of AI in the biomedical space similar to other domains. Massive real-world datasets and public benchmarks such as NYUMets-Brain may enable the tracking and detection of metastatic brain cancer, and be broadly applicable to advancing the development of AI models in other types of metastatic cancer as well.
Methods
Prospective radiosurgery clinical registry
Since November 2012, the Center for Advanced Radiosurgery at NYU Langone Health has maintained a prospective data collection system. This registry is based on a SQL server platform with manual prospective entry of every patient treated with stereotactic radiosurgery at the time of treatment and at each clinical follow up. Major elements entered include demographic details, clinical and disease-based characteristics (i.e., tumor histology, symptoms, functional status, prior treatments), results of clinical grading systems (i.e., EQ-5D, RPA), treatment details, and follow-up clinical characteristics. Details of this registry’s development can be found in this contemporary report40. The present study protocol was approved by the NYU Langone Health Institutional Review Board (IRB# s20-01881) under a waiver of informed consent as the study involves de-identified data and meets all five criteria under the Common Rule (45 CFR 46.116(f)).
Clinical data processing
Clinical data drawn from the prospective radiosurgery clinical registry, radiosurgery treatment software (Leksell GammaPlan), medication orders from electronic medical records (Epic Clarity), and imaging records were standardized and de-identified across three datasets: (1) time series dataset, (2) individual dataset, and (3) radiosurgery details dataset. For all datasets, the patient, radiosurgery treatment, and MR images were indexed based on randomly generated numbers for patient identifier (eight-digit), gamma knife identifier (nine-digit), and image identifier (ten-digit).
See API at nyumets.org for details of variables and legends for each variable in all datasets. For the time series dataset, 100 variables were indexed on time of occurrence or data collection in units of days from initial brain radiosurgery. The individual dataset was indexed based on patient identifiers with 95 variables, including clinical variables that did not change with time. The “Radiosurgery Details” dataset was indexed on a radiosurgery identifier to provide 19 variables with further treatment details for available radiosurgery treatments.
Image preprocessing
For studies without radiosurgery planning segmentations, pre-contrast T1, post-contrast T1, T2, and FLAIR brain MRI images were extracted from each timepoint’s DICOM files and combined into a single 3D array using the pydicom and dicom_numpy packages. All images were resampled to isotropic 1 mm3 dimensions using linear interpolation and then saved in NIFTI format using the Nibabel package. For studies with Gamma Knife segmentations, tumor segmentations were extracted from the RTSTRUCT file and combined into one binary array using the rt_utils package. Post-contrast T1 gradient-echo MRI associated with each RTSTRUCT were also extracted from their DICOM files into a 3D array. Both arrays were resampled to 1 mm3 dimensions and saved in NIFTI format. Studies with radiosurgical planning and without radiosurgical planning were combined into one study timepoint if the non-radiosurgical planning study was acquired 30 days or less before radiosurgical treatment and was the closest in date (if multiple non-radiosurgical studies were acquired within the 30-day window).
All studies, whether matched or unmatched, were then reoriented to the same orientation, and all NIFTI files within each study were coregistered using symmetric normalization with the ANTsPy package. Coregistered images were then skull stripped using the AFNI 3dSkullStrip function. Segmentations were either manually adjusted by a board-certified neurosurgeon using ITK-SNAP or run through a low pass filter to remove aliasing effects from the processing pipeline.
The dataset was randomly split into predetermined train, validation, and testing splits by patient ID such that images from the same patient were only present in one split. A description and comparison of the training, validation, and testing splits can be found in the Extended Data Table 3. Immediately prior to training and evaluation, image intensity was normalized, images of multiple MRI modalities (if used) were combined in the channel dimension, and the images were cropped or padded to (192, 192, 192) arrays using the MONAI package. Any labeled pixels falling outside of the brain were removed. Several spatial and intensity augmentations were optionally added prior to model training; a detailed list can be found in Extended Data Table 4. Finally, the full-size image was cropped into 96 × 96 × 96 patches to be fed into the network. If multiple MRI sequences were used, they were processed as multi-channel images (e.g. T1C in the first channel, T2 in the second channel).
External datasets were processed to match the NYUMets dataset. If needed, all images were converted to Nifti format and resampled to isotropic 1 × 1 × 1 mm resolution. All other processing steps were completed identically to the NYUMets dataset before inference.
Model architecture
We use the Basic UNet architecture from the MONAI library as the control network, and modify it with a single unidirectional convolutional LSTM layer to process imaging data through time, which we call “segmentation-through time” (STT) (Fig. 1B). For the STT network, data was combined into batches that contained the studies of one patient, with the batch dimension representing the sorted sequential images. The default layer dimensions were used for the UNet backbone and standalone UNet (feature encoding sizes = (32, 32, 64, 128, 256), last feature decoding size = 32).
The STT model was compared to a UNet with the same architecture other than a convolutional LSTM layer. Extended Data Table 3 presents the characteristics of the dataset used for model training and evaluation. In order to measure the relative value of additional MRI sequences, we ran experiments training on only T1 post-contrast images as well as T1 post-contrast and T2 images. We also evaluated the relative value of training on both non-expert-adjusted and expert-adjusted segmentations compared to training on only expert-adjusted segmentations. In order to evaluate the performance of our models on metastases of different volumes, we stratified our evaluation metrics by metastasis volume. Extended Table 4 shows the distribution of metastasis volume in the dataset used for modeling. Finally, we also compared our modeling results with a standard training regimen to models with the same architecture that were pretrained using self-supervised learning (SSL) (see next section for details).
Model training
The models for this study are implemented using PyTorch41, MONAI42, and PyTorch lightning43. The models are trained using 1x A100 GPU, and all are trained with an equivalent batch size of 64 (adjusted with gradient accumulation). We first ran a hyperparameter search for each configuration of the dataset. The best model of each configuration, determined by validation metrics, is then trained with three different random seeds to obtain the final model and performance. Hyperparameter tuning for the model is done using Optuna44 with the TPE sampler45, which is a tree-based algorithm for finding optimal hyperparameters. We use a search budget of 100 trials for each study, and run eight jobs concurrently on 16 A100 GPUs. Hyperparameters tuned using this procedure include the learning rate, loss function, and the hidden size of the decoder. The best hyperparameters were determined by the mean Dice similarity coefficient on the validation dataset. Using these hyperparameters, each model was then trained for 150 epochs to obtain the final trained model that would be used for model evaluation.
Dice similarity coefficients were calculated per-scan as well as on an individual metastasis level to evaluate metastasis segmentation quality. Hausdorff Distances at the 95th percentile were also calculated per-scan to measure segmentation shape fidelity.
Self-supervised learning (SSL) training
We experiment with a self-supervised learning approach similar to Tang et al46. to pretrain the model before finetuning it with the annotated segmentations. Our pretraining method involves the use of reconstruction loss and contrastive loss given by the following equations.
For the reconstruction loss (Eq. (4)), the network is given perturbed version of the original image \({P}_{1}({x}_{n})\), and is required to output an exact reconstruction of the unperturbed version of the input. The perturbations used in this step include local pixel shuffling and random cutouts. For the contrastive loss (Eq. (3)), we perturb each sample with two different random transformations \({P}_{1},\,{P}_{2}\) (Eq. (1)) and use those as positive samples (Eq. (2)), with the other images in the same batch as negative samples. We then combine the reconstruction loss and the contrastive loss using a weight of 1.0 and 0.5 for pretraining (Eq. (5)). The network is pretrained to convergence as marked by the plateauing of validation set reconstruction loss.
Model evaluation
Models were evaluated across several segmentation and detection metrics on the test dataset. Data inputted into the model for evaluation underwent identical processing steps as training. Model predictions were compared with gold (expert-adjusted) labels only. In order to separate individual metastasis, the connected components algorithm was run over the binarized model outputs and target labels. In order to match connected components between the target segmentation and predicted segmentation, as well as across sequential timepoints, a tumor matching algorithm was used to standardize the connected component IDs between two segmentations (Extended Fig. 4). For every pair of tumor IDs between two connected component segmentation, the Euclidean distance between metastasis centroids and the intersection over reference (IoR), where the reference segmentation is either the target segmentation or the first timepoint’s segmentation in a sequence, was calculated. All pairs with centroid distances of less than 1000 mm and IoR of over 0.001 were then selected, and the pair of metastasis segmentations with the greatest IoR was chosen as the matched pair in order to avoid a single metastasis being assigned to more than one pair.
Models were evaluated on single-timepoint segmentation and detection, as well as segmentation and detection across two sequential timepoints. In addition to being aggregated by scan, several metrics were calculated and aggregated across all metastases. When the Dice similarity coefficient was calculated on a per-lesion basis, only true positive lesions were included, as done in ref. 37. When the Dice similarity coefficient was calculated on a per-scan basis, it was calculated excluding the background channel and all segmentations were included (including false positive and false negative segmentations). A full list and description of the metrics used to evaluate models can be found in Extended Fig. 5.
In addition to the evaluation of our internal test set of NYUMets data, we also conducted external tests with the Stanford BrainMetsShare dataset and the BraTS 2021 dataset. We performed no preprocessing on these datasets except for converting the provided Stanford BrainMetShare files to NIFTI format.
Clinical outcomes statistics
Kaplan–Meier survival analysis was used to assess survival with equality of survivor functions tested with log-rank tests. Univariable and multivariable Cox proportional hazards models were fitted to assess factors contributing to overall survival and individual variable significance was assessed with likelihood ratio tests. Statistical analyses were performed using Stata version 17.0 (StataCorp LP). Alpha was set at 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data is publicly available at https://nyumets.org.
Code availability
All code and trained model weights will be made publicly available at https://github.com/nyumets/nyumets upon publication of the manuscript.
References
Acosta, J. N., Falcone, G. J., Rajpurkar, P. & Topol, E. J. Multimodal biomedical AI. Nat. Med. 28, 1773–1784 (2022).
Acosta, J. N., Falcone, G. J. & Rajpurkar, P. The need for medical artificial intelligence that incorporates prior images. Radiology https://doi.org/10.1148/radiol.212830 (2022).
Suh, C. H., Jung, S. C., Kim, K. W. & Pyo, J. The detectability of brain metastases using contrast-enhanced spin-echo or gradient-echo images: a systematic review and meta-analysis. J. Neurooncol. 129, 363–371 (2016).
Litière, S., Collette, S., de Vries, E. G. E., Seymour, L. & Bogaerts, J. RECIST - learning from the past to build the future. Nat. Rev. Clin. Oncol. 14, 187–192 (2017).
Abramson, R. G., McGhee, C. R., Lakomkin, N. & Arteaga, C. L. Pitfalls in RECIST data extraction for clinical trials: beyond the basics. Acad. Radiol. 22, 779–786 (2015).
Lin, N. U. et al. Response assessment criteria for brain metastases: proposal from the RANO group. Lancet Oncol. 16, e270–e278 (2015).
Chang, K. et al. Automatic assessment of glioma burden: a deep learning algorithm for fully automated volumetric and bidimensional measurement. Neuro. Oncol. 21, 1412–1422 (2019).
Tang, Y. et al. Semi-automatic RECIST labeling on CT scans with cascaded convolutional neural networks. In 21st International Conference Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2018 405–413 (Springer International Publishing, 2018).
Russakovsky, O. et al. Imagenet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252 (2015).
Lin, T. -Y. et al. Microsoft coco: common objects in context. In 13th European Conference Proceedings of the Computer Vision–ECCV 2014 740–755 (Springer International Publishing, 2014).
Everingham, M., Van Gool, L., Williams, C. K. I., Winn, J. & Zisserman, A. The pascal visual object classes (VOC) challenge. Int. J. Comput. Vis. 88, 303–338 (2010).
Strickland, E. Andrew Ng: unbiggen A. I. IEEE Spectrum https://spectrum.ieee.org/andrew-ng-data-centric-ai (2022).
Mazumder, M. et al. Dataperf: Benchmarks for data-centric ai development. Advances in Neural Information Processing Systems 36 (2024).
Bakas, S. et al. Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 4, 170117 (2017).
Freymann, J. B., Kirby, J. S., Perry, J. H., Clunie, D. A. & Jaffe, C. C. Image data sharing for biomedical research-meeting HIPAA requirements for de-identification. J. Digit. Imaging 25, 14–24 (2012).
Menze, B. H. et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 34, 1993–2024 (2015).
Johnson, A. E. W. et al. MIMIC-III, a freely accessible critical care database. Sci. Data 3, 160035 (2016).
Wynants, L. et al. Prediction models for diagnosis and prognosis of covid-19 infection: systematic review and critical appraisal. BMJ 369, m1328 (2020).
MONAI Consortium. MONAI: medical open network for AI. https://doi.org/10.5281/zenodo.7459814 (2022).
Swanton, C. Take lessons from cancer evolution to the clinic. Nature 581, 382–383 (2020).
Ma, T. M. et al. Refining the definition of biochemical failure in the era of stereotactic body radiation therapy for prostate cancer: the Phoenix definition and beyond. Radiother. Oncol. 166, 1–7 (2022).
Robinson, B. et al. Characterization of tumor size changes over time from the phase 3 study of lenvatinib in thyroid cancer. J. Clin. Endocrinol. Metab. 101, 4103–4109 (2016).
Claret, L. et al. Evaluation of tumor-size response metrics to predict overall survival in Western and Chinese patients with first-line metastatic colorectal cancer. J. Clin. Oncol. 31, 2110–2114 (2013).
Wang, Y. et al. Elucidation of relationship between tumor size and survival in non-small-cell lung cancer patients can aid early decision making in clinical drug development. Clin. Pharmacol. Ther. 86, 167–174 (2009).
Cagney, D. N. et al. Incidence and prognosis of patients with brain metastases at diagnosis of systemic malignancy: a population-based study. Neuro. Oncol. 19, 1511–1521 (2017).
Nayak, L., Lee, E. Q. & Wen, P. Y. Epidemiology of brain metastases. Curr. Oncol. Rep. 14, 48–54 (2012).
Burrell, R. A., McGranahan, N., Bartek, J. & Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013).
Bailey, C. et al. Tracking cancer evolution through the disease course. Cancer Discov. 11, 916–932 (2021).
Welter, L. et al. Treatment response and tumor evolution: lessons from an extended series of multianalyte liquid biopsies in a metastatic breast cancer patient. Cold Spring Harb. Mol. Case Stud. 6, a005819 (2020).
Xu, Y. et al. Deep learning predicts lung cancer treatment response from serial medical imaging. Clin. Cancer Res. 25, 3266–3275 (2019).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In 18th International Conference Proceedings of the Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015. 234–241 (Springer International Publishing, 2015).
Falk, T. et al. U-Net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2019).
Grøvik, E. et al. Deep learning enables automatic detection and segmentation of brain metastases on multisequence MRI. J. Magn. Reson. Imaging 51, 175–182 (2020).
Rudie, J. D. et al. The University of California San Francisco Brain Metastases Stereotactic Radiosurgery (UCSF-BMSR) MRI Dataset. Radiol. Artif. Intell. 6, e230126 (2024).
Baid, U. et al. The RSNA-ASNR-MICCAI BraTS 2021 benchmark on brain tumor segmentation and radiogenomic classification. Preprint at arXiv [cs.CV] (2021).
Aoyama, H. et al. Stereotactic radiosurgery plus whole-brain radiation therapy vs stereotactic radiosurgery alone for treatment of brain metastases: a randomized controlled trial. JAMA 295, 2483–2491 (2006).
Chartrand, G. et al. Automated detection of brain metastases on T1-weighted MRI using a convolutional neural network: impact of volume aware loss and sampling strategy. J. Magn. Reson. Imaging https://doi.org/10.1002/jmri.28274 (2022).
Rudie, J. D. et al. 3D U-net convolutional neural network for detection and segmentation of intracranial metastases. Radiology https://doi.org/10.1148/ryai.2021200204 (2021).
Harish, K. B., Price, W. N. & Aphinyanaphongs, Y. Open-source clinical machine learning models: critical appraisal of feasibility, advantages, and challenges. JMIR Form. Res. 6, e33970 (2022).
Kondziolka, D., Cooper, B. T., Lunsford, L. D. & Silverman, J. Development, implementation, and use of a local and global clinical registry for neurosurgery. Big Data 3, 80–89 (2015).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. Preprint at arXiv [cs.LG] (2019).
The MONAI Consortium. Project MONAI. https://monai.io/ (2020).
Falcon, W. The PyTorch lightning team. Pytorch Lightning 3, 6 (2019).
Akiba, T., Sano, S., Yanase, T., Ohta, T. & Koyama, M. Optuna: a next-generation hyperparameter optimization framework. In Proc. 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining 2623–2631 (Association for Computing Machinery, 2019).
Bergstra, J., Bardenet, R., Bengio, Y. & Kégl, B. Algorithms for hyper-parameter optimization. Adv. Neural Inf. Process. Syst. 24, 2546–2554 (2011).
Tang, Y. et al. Self-supervised pre-training of swin transformers for 3d medical image analysis. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. (2022).
Acknowledgements
Erin Chu, Ph.D., and the Amazon Web Services Open Data team for their generous support of this project’s data hosting. Anthony Costa, Ph.D., Mona Flores, M.D., and the NVIDIA MONAI team. Michael Liu at Bearwave for his help with internal infrastructure construction. Dean Dafna Bar-Sagi, Ph.D. Michael Recht, M.D., Yvonne Lui, M.D., Daniel Sodickson, M.D., Ph.D., Erik Sulman, M.D., Ph.D., and everyone else at NYU Langone who supports the OLAB. Dr. Benjamin Neal and the NYU Langone Perlmutter Comprehensive Cancer Center (P30 CA0160087). E.K.O. is funded by the NCI’s Early Surgeon Scientist Program (ESSP) (3P30CA016087-41S1). K.E.L is funded by the NREF & Aaron Cohen-Gadol Medical Student Summer Research Fellowship and RSNA Medical Student Research Grant.
Author information
Authors and Affiliations
Contributions
E.K.O. and D.K. conceptualized and supervised the project. K.E.L., Y.J.K., Z.S., J.D.A., K.B., T.Q., and L.Y.J. collected data, cleaned data, and built the dataset. K.E.L., C.L., M.N.-M., and Y.J.K. performed modeling experiments. C.L. performed the self-supervised learning experiments. Z.S., S.N., performed the analysis of cancer dynamics. K.E.L., M.N.-M., Z.S., C.L., E.Y., and S.N. prepared figures. K.E.L. and C.L. debugged and tested the models and the pretraining and finetuning software. E.K.O. designed the NYUMets website and cloud infrastructure. J.G.G., D.O., V.C., and D.K. provided guidance and feedback throughout the project. K.E.L., Z.S., and C.L. wrote the initial draft. All authors edited and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests. E.K.O. reports consulting with Sofinnova Inc., Google Inc., income from Merck & Co., and Mirati Therapeutics, and equity in Artisight Inc. D.O. is a consultant and equity holder of Invenio Inc. D.K. reports consulting with Elekta Inc. K.E.L. is currently employed by NVIDIA. L.Y.J., M.N., S.N., J.D.A., K.B., T.Q., V.C., E.Y., and J.G.G. declare no competing interests. The work presented herein was performed exclusively within the NYU Langone Health System.
Peer review
Peer review information
Nature Communications thanks Antonio Meola, Jon Ottesen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Link, K.E., Schnurman, Z., Liu, C. et al. Longitudinal deep neural networks for assessing metastatic brain cancer on a large open benchmark. Nat Commun 15, 8170 (2024). https://doi.org/10.1038/s41467-024-52414-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52414-2
This article is cited by
-
Learning neuroimaging models from health system-scale data
Nature Biomedical Engineering (2026)
-
Applications of artificial intelligence in non–small cell lung cancer: from precision diagnosis to personalized prognosis and therapy
Journal of Translational Medicine (2025)
-
Outcomes of concurrent versus non-concurrent immune checkpoint inhibition with stereotactic radiosurgery for melanoma brain metastases
Journal of Neuro-Oncology (2025)