Abstract
The leucine-rich repeat kinase 2 (LRRK2) phosphorylates a subset of RAB GTPases, and their phosphorylation levels are elevated by Parkinson’s disease (PD)-linked mutations of LRRK2. However, the precise function of the LRRK2-regulated RAB GTPase in the brain remains to be elucidated. Here, we identify RAB12 as a robust LRRK2 substrate in the mouse brain through phosphoproteomics profiling and solve the structure of RAB12-LRRK2 protein complex through Cryo-EM analysis. Mechanistically, RAB12 cooperates with LRRK2 to inhibit primary ciliogenesis and regulate centrosome homeostasis in astrocytes through enhancing the phosphorylation of RAB10 and recruiting RILPL1, while the functions of RAB12 require a direct interaction with LRRK2 and LRRK2 activity. Furthermore, the ciliary and centrosome defects caused by the PD-linked LRRK2-G2019S mutation are prevented by Rab12 deletion in astrocytes. Thus, our study reveals a physiological function of the RAB12-LRRK2 complex in regulating ciliogenesis and centrosome homeostasis. The RAB12-LRRK2 structure offers a guidance in the therapeutic development of PD by targeting the RAB12-LRRK2 interaction.
Similar content being viewed by others
Introduction
The leucine-rich repeat kinase 2 (LRRK2) variants cause the most common inherited forms of Parkinson’s disease (PD)1,2. Lrrk2 encodes a large, complex protein containing both GTPase and kinase domains3. High-resolution cryo-EM structures of full-length LRRK2 protein were determined, and the kinase domain was shown to adopt an inactive conformation4,5, suggesting a highly regulated kinase activity. Multiple pathogenic variants of LRRK2 were identified and shown to enhance LRRK2 kinase activity in vitro6. Among those disease mutants, the LRRK2-G2019S is the most prevalent7,8,9. Available evidence has shown that LRRK2 regulates multiple cellular pathways, including immune response, vesicle trafficking, lysosome homeostasis, and synaptic transmission3,10,11.
Previous proteomics analysis reported that LRRK2 phosphorylates a subset of RAB GTPases at a conserved residue threonine or serine in their switch II domains, resulting in a potential inhibition of RAB activation and the effector protein binding12,13. The phospho-RAB has a profound impact on their functions as evidenced in impaired autophagosome motility in the axons14, primary ciliogenesis15,16,17,18,19,20,21,22, and centriolar cohesion19,23. For example, LRRK2 was shown to regulate primary ciliogenesis through RAB8 and RAB10 and their interactions with effectors Rab interacting lysosomal protein like 1/2 (RILPL1/2)16,18,24. Subsequent studies verified that LRRK2 pathogenic variants R1441C and G2019S interfere with primary cilia formation associated with an impairment of sonic hedgehog signaling15.
The identification of the selective RAB GTPases as the targets of LRRK2 kinase began to provide important insight into the molecular mechanism underlying LRRK2 pathogenic signaling pathways10,25,26. However, the exact mechanism for how multiple RAB proteins participate in LRRK2 pathogenic pathways remains to be clarified. Interestingly, the study of Lrrk2-G2019S knockin (KI) mouse tissues demonstrated an increase of phospho-RAB12 levels, while the change of phosphorylation levels of other RAB proteins is not understood27, implicating RAB12 as an important substrate of LRRK2 in the brain at basal condition. RAB12 can be recruited to LRRK2-associated lysosomes or endolysosomes and are phosphorylated by LRRK228,29,30. Two most recent reports showed that RAB12 is an activator of LRRK2 kinase activity that can promote lysosome recruitment of LRRK2 and facilitate LRRK2 phosphorylation of RAB10 in cell cultures from non-brain cell types22,31. Despite the notions, the physiological function of RAB12 and the precise functional relationship between LRRK2, RAB12, and other RAB GTPases remains to be elucidated particularly in neurons and glia.
Here we report RAB12 as a physiological substrate of LRRK2 in the brain, reveal the cryo-EM structure of RAB12-LRRK2 protein complex, and characterize the function of RAB12 in the glia. We show that RAB12 is an inhibitor of primary ciliogenesis, which requires a direct binding with LRRK2. Our study provides insight into the potential mechanism whereby the RAB12-LRRK2 complex controls the homeostasis of astrocytic ciliogenesis and centrosomes. Furthermore, our study shows the role of RAB12 in mediating the pathogenic function of LRRK2-G2019S in astrocytes. Our study thus establishes the physiological function of the RAB12-LRRK2 complex and provides a structural framework for targeting RAB12-LRRK2 interaction in therapeutic development.
Results
Structure of the RAB12-LRRK2 complex
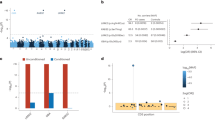
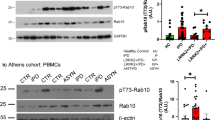
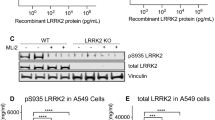
To identify the physiological substrates of LRRK2 kinase in the brain, we performed a deep proteome and phosphoproteomics analysis using brain lysates from Lrrk2 knockout (KO), Lrrk2 BAC transgenic mice overexpressing Lrrk2 wildtype (WTtg), PD-mutant G2019S (GStg)32, and control mice (WT). Through the TMT-LC/LC-MS/MS analysis, we identified a total of 10,195 proteins and 72,766 phosphopeptides (52,187 phosphorylation sites out of 7829 phosphoproteins) (Fig. S1a–c, Supplementary Data 1–3). The data revealed that the relative phosphorylation levels at multiple sites on LRRK2 (S910, S935, S955, and S973), OXR1 (S15, S308, and S396), and RAB12 (S105) were correlated with the LRRK2 levels/activities (Fig. S1c). We validated the RAB12 phosphorylation levels in the above mutant mice through immunoblot analysis (Fig. S1d). The above observation suggested that RAB12 is a robust substrate of LRRK2 in the brain under basal conditions.
To characterize the interaction between RAB12 and LRRK2 at atomic details, we solved the cryo-electron microscopy (cryo-EM) structures of the human RAB12-LRRK2 complex (Fig. S2 and Supplementary Table 1). We introduced the GTPase-deficient mutation (active form), Q101L33, to RAB12 to obtain homogeneous and stable RAB12-LRRK2 complexes for structural analysis. Like LRRK2-alone structures4, the RAB12-LRRK2 complex was also captured in both monomer and dimer states at overall resolutions of 4.1 Å and 3.9 Å, respectively (Fig. S2a), and LRRK2 protomers in both states adopt an inactive conformation as previously reported4,34 (Fig. S3a). Focused refinement was performed for the RAB12-LRRK2 interface to improve the local resolution (Fig. S2a). The GTP analog-bound RAB12 resembles RAB29 and RAB32 in their complexes with LRRK2 (Fig. 1b and Fig. S3b).
a The cryo-EM map and atomic model of the RAB12-LRRK2 complex. RAB12 and LRRK2 are colored in purple and blue, respectively. Side chains involved in interactions between RAB12 and LRRK2 are shown as sticks. b Interactions between the LRRK2 ARM domain and RAB12. The ARM repeats of LRRK2 are indicated by numbers. Key structural elements in RAB12 are labeled. The phosphorylation site (p-site) is shown as green sphere. Cryo-EM density of the GTP analog and Mg2+ bound to RAB12 is shown. c Structural comparison of the RAB12-LRRK2 interfaces between RAB12-LRRK2 monomer and dimer. d Superposition analysis of the RAB12-LRRK2 and RAB29–LRRK2 (PDB 8FO2) interfaces based on RAB proteins. e Superposition analysis of the RAB12-LRRK2 and RAB29–LRRK2 (PDB 8FO2) interfaces based on the LRRK2 ARM repeats. The three helices (H1, H2, and H3) of each ARM repeat are labeled.
The interaction interfaces between RAB12 and LRRK2 in both monomers and dimers are almost identical and mediated by CDR1 and Switch I-Interswitch-Switch II motifs of RAB12 and ARM6-ARM7 of LRRK2 (Fig. 1a–c). Key interface residues are annotated in Fig. 1a. Comparing the RAB12-LRRK2 and RAB29/RAB32–LRRK2 complexes, we found that all RABs use the CDR1 and Switch I-Interswitch-Switch II motifs for LRRK2 binding34, though the α2 helix in Rab12 seems to make closer contacts with ARM repeats (Fig. 1d and Fig. S3c, d). As a result, Tyr111 of RAB12 sits in the middle of the interface, while the side chain of corresponding residue Tyr77 points away from the ARM domain in RAB29 (Fig. S3c, d). On the other hand, RAB12 binds to the H1 helices of ARM6-ARM7 while RAB29 and RAB32 mainly interact with the H2 helices of ARM9-ARM10 of LRRK2 (Fig. 1e and FIg. S3f). Of note, each ARM repeat contains three helices: H1-H3. Sequence alignment revealed that only a few interface residues are not conserved among the three RAB GTPases, including Lys42, Ile81, and Ile107 of RAB12 and Leu76 of RAB29, resulting in a totally different binding surface on the ARM domain of LRRK2 (Fig. S3c–f). Glu240 of LRRK2 interacts with two of the three RAB12-specific residues (Lys42 and Ile81), forms salt bridges with Arg92 and Lys42 of RAB12 (Fig. S3c) and should be critical for RAB12-LRRK2 interactions.
Lack of Rab12 increases the number of ciliated astrocytes in vivo and in vitro
To elucidate the physiological function of Rab12 in the brain, we generated Rab12 KO mice by deleting the exon 3 of Rab12 from the genomic sequence using the CRISPR-Cas9 gene editing (Fig. S4a). The Rab12 KO mice are viable and show no apparent developmental defect. Given the evidence for the involvement of LRRK2 in primary ciliogenesis13,15,16,17,18,20,24, we tested the effects of Rab12 deletion on primary cilia formation in mouse brains. Staining of Rab12 KO brain slices with anti-Arl13b antibody showed a significant increase of the ciliated cell percentage in the striatum and cortex and a trend of increase of cilia length in the striatum. However, the average cilia volume of the mutant mice changed little in both regions (Fig. 2a). To ascertain the cell types affected in Rab12 KO brains, we performed co-staining with anti-Arl13b and the antibodies specific for astrocyte or neuron, two main ciliated cell types in the brains35. The results indicated that the astrocytes had higher ciliated cell percentage yet no significant change in the cilia length or volume in the mutant brains (striatum and cortex) compared to the controls (Fig. 2b). In contrast, the neurons from Rab12 KO brains showed no difference in the ciliated cell percentage but increased cilia length and volume in the striatum, not in the cortex, compared to the control mice (Fig. S4b). Consistently, primary astrocyte cultures derived from Rab12 KO brains also showed elevated percentage of ciliation but no change of the cilia length and volume (Fig. 2c). Therefore, our in vivo and in vitro data indicate that loss of Rab12 results in an increased number of ciliated astrocytes.
a Primary cilia were detected with anti-Arl13b antibody (white arrowheads) and imaged in the striatum and frontal cortex from wildtype (WT) and Rab12 KO mice. The ciliated cell percentage, cilia length, and cilia volume were quantified. Biological replicates: nWT = 9, and nRab12KO = 8 for the striatum; nWT = 4, and nRab12KO = 4 for the frontal cortex. b The primary cilia were detected in the astrocytes (anti-GFAP) in the striatum and frontal cortex of WT and Rab12 KO mice, and ciliated astrocyte percentage and cilium size were quantified (nWT = 9, and nRab12KO = 8 for both the striatum and frontal cortex). c The cilia in primary astrocytes from neonatal Rab12 KO mice and littermate controls were examined as in (b) (n = 5 for each group). The boxed images of cilia in (a–c) were enlarged at the upper left corners in each panel in (a, b), and the bottom left corners in each panel in (c). Each dot in the graphs represents an individual mouse (a, b) or >60 cells from each mouse (c). P-values were calculated using unpaired two-tailed t-tests; *P < 0.05, **P < 0.01, ***P < 0.001; error bars = SEM; scale bars are indicated. Source data are provided as a Source Data file.
Rab12 regulates the homeostasis of primary cilia and centrosomes
We next tested the idea that RAB12 suppresses primary ciliogenesis by focusing on astrocytes. We overexpressed Rab12 gene in astrocytes by AAV-mediated delivery of HA-tagged Rab12 into primary cultures. HA-RAB12 overexpression often resulted in large-size amorphic structures with short “spikes” protruding from the surface with anti-Arl13b staining in contrast to GFP-overexpressed astrocytes (Fig. 3a). The average intensity of Arl13b fluorescence of the spike-like structures in HA-RAB12-overexpressed astrocytes, however, is less than the cilia (Arl13b+) in GFP-overexpressed cells (Fig. 3a). Staining with anti-γ-tubulin, a marker for centrosome which forms basal bodies and sequentially templates cilia formation36, showed a significant overlap of γ-tubulin with Arl13b and HA-RAB12 in the large structures. The above staining pattern in HA-RAB12 overexpressed astrocytes is in stark contrast with non-infected or Gfp-infected astrocytes, which showed typical primary cilia linked to centrosomes with both γ-tubulin and Arl13b staining (Fig. 3a). Indeed, HA-RAB12 overexpression led to a significant increase of γ-tubulin staining volume, compared to GFP overexpression. Using another centrosome-specific marker anti-Pericentrin antibody, we validated the greater volume of centrosomes (Pericentrin + /HA-RAB12 + ) in the HA-RAB12 overexpressed astrocytes (Fig. 3b). The observation suggests an expansion of centrosomes and perhaps aberrant cilia formation resulted from the overexpression of RAB12.
Primary culture of astrocytes was infected with AAVs harboring GFAP promoters to express GFP, HA-RAB12 (a–c), or the active HA-RAB12 Q100L variant (HA-RAB12QL, d) conjugated with an HA tag. a, b The astrocytes overexpressing HA-RAB12 or GFP were marked with HA or GFP antibodies by IF, accordingly. Their primary cilia and centrosomes were detected using Arl13b antibody and γ-tubulin antibody, respectively. The regions of cells containing centrosomes and typical/failed cilia are marked by white-dash boxes and enlarged (a, b). The graphs at the bottom of (a, b) are the quantification of γ-tubulin volume (n = 4 for each group), γ-tubulin–GFP/HA Pearson coefficient (n = 4), and Arl13b average fluorescent intensity (n = 7) from the images as shown in (a), and Pericentrin volume, and Pericentrin–GFP/HA Pearson coefficient value from images as shown in (b) (n = 3). c Electron microscope images from WT, Rab12 KO, and HA-RAB12 overexpressed astrocytes. Centrosomes are indicated in blue, cilia are in pink, and ciliary pockets are in green. The arrowheads indicate the amplificated centrosomes, and the representatives of failed cilia in boxes 1 and 2 are enlarged and displayed in the right panels (n = 1 for each group). d The cilia and centrosomes in the astrocytes overexpressing GFP, or HA-RAB12QL were detected by IF, and the representative cilia and centrosomes from each group were enlarged. e–i The qualification of ciliated cell percentage, cilia length, cilia volume, γ-tubulin volume, and γ-tubulin–GFP/HA-RAB12 Pearson coefficient from images as shown in (d) (n = 5 for e–g, n = 4 for h, n = 6 for i). Each dot in the graphs represents >60 cells from each mouse. Unpaired two-tailed t-tests, *P < 0.05, **P < 0.01, ***P < 0.001; error bars = SEM; scale bars are indicated. Source data are provided as a Source Data file.
To investigate the details of the cilia alterations, we performed an electron microscopy (EM) analysis of the infected astrocytes. In control and Rab12 KO astrocytes, the normal primary cilia are associated with the centrosome at base (Fig. 3c). In contrast, we found frequent clustering of centrosomes/centrioles in HA-Rab12 infected cells, in which the “dwarf” cilia are associated with aberrant cilia membranes and enlarged lumen enwrapping the abnormal axoneme (Fig. 3c). The clustering of centrosomes/centrioles from the EM analysis is consistent with the large volume of γ-tubulin/Pericentrin labeled structures under the confocal microscopy (Fig. 3a). They are analogous to what was previously described as centrosome amplification or supernumerary centrioles associated with tumor cells37,38. Thus, our EM result suggested the failure of primary cilia formation in HA-RAB12 overexpressed astrocytes. Our observation agreed with a previous report, which suggested that centrosome amplification can inhibit cilia formation or cause clustered, diluted cilia (e.g. reduced Arl13b levels)37.
Moreover, overexpression of HA-RAB12-Q100L (HA-RAB12QL) mouse mutant, a constitutively active form of RAB1239,40, in astrocytes markedly reduced the ciliated cell percentage (Fig. 3d–g). However, in contrast to the wildtype, the active RAB12 had little effect on the γ-tubulin volume (Fig. 3h) and Pericentrin volume (Fig. S5a) and showed no significant co-localization with γ-tubulin (Fig. 3i) and Pericentrin (Fig. S5b) staining. The inhibitory effect of the active RAB12 variant on ciliated number of astrocytes corroborates the findings with RAB12 wildtype (WT) overexpression, which stunted cilia formation (Fig. 3a–c), while the active RAB12 variant appears not to cause centrosome amplification or associate with centrosomes.
We next asked if overexpression of LRRK2 or other LRRK2 substrates, such as RAB10 and RAB8a, affects the homeostasis of primary cilia and centrosomes in astrocytes. Interestingly, overexpression of RAB10, RAB8a, or LRRK2 influenced neither cilia morphology nor the volume of centrosomes (Fig. S5c–e). However, overexpression of RILPL1, an effector of RAB proteins, caused increased centrosome volume (Fig. S5d–e). Therefore, the effect of overexpression on primary cilia and centrosome homeostasis appears to be specific for RAB12, which may act through the effector RILPL1.
The RAB12-LRRK2 complex regulates primary cilia and centrosomes
Our structural analysis showed RAB12-LRRK2 complex formation and the binding interface between the two proteins (Fig. 1). Therefore, we next tested the role of RAB12-LRRK2 complex in regulating primary cilia and centrosomes in astrocytes, and we postulated that mutations of the critical residues of the interface in RAB12 disrupt the RAB12-LRRK2 interaction and consequently abolish the effects in suppressing ciliogenesis or promoting centrosome amplification. The analysis suggested that K42, F79, R92, and Y111 positioning at the interface of RAB12 are important for binding the ARM of LRRK2 (Fig. 1a). To this end, we engineered K42A and Y111A in the active RAB12 variant (HA-RAB12QL, Fig. S6a) as well as WT (HA-RAB12, Fig. S6a). Firstly, we verified that both active and WT RAB12 carrying K42A or Y111A lost the interactions with LRRK2 through co-immunoprecipitation (Co-IP) assay (Fig. 4a, b). Secondly, in contrast to HA-RAB12QL, overexpression of double mutants HA-RAB12QL-K42A or HA-RAB12QL-Y111A in astrocytes had no impact on the ciliated cell percentage. They did not influence cilia length or cilia volume (Fig. 4c). Thirdly, overexpression of HA-RAB12QL in Lrrk2 KO astrocytes appeared to alleviate the effect of reduced ciliated cell percentage as observed in WT astrocytes, while no influence was found in cilia length or volume (Fig. 4d, Fig. S6b). Thus, the role of RAB12 in the suppression of primary ciliogenesis in astrocytes depends on the presence of LRRK2 and requires RAB12-LRRK2 interaction.
a, b The human LRRK2WT plasmids were co-transfected with the human RAB12 variant plasmids, such as HA-RAB12QL-K42A, HA-RAB12QL-Y111A, HA-RAB12-K42A, or HA-RAB12-Y111A into HEK293T cells. The interactions between LRRK2 and RAB12 or its mutants were examined by co-immunoprecipitation (Co-IP) assay. a The Co-IP between LRRK2 and HA-RAB12QL or its binding deficient mutants (n = 3). b The Co-IP between LRRK2 and HA-RAB12 or its binding deficient mutants (n = 3). c Gfp, HA-RAB12QL, HA-RAB12QL-K42A, or HA-RAB12QL-Y111A were transduced into astrocytes by AAVs carrying GFAP promotors (n = 3). d WT and Lrrk2 KO astrocytes were infected with AAVs carrying Gfp, or HA-Rab12QL (n = 7). The cilia of the infected astrocytes from (c, d) were detected and their ciliated cell percentage, cilia length, and cilia volume were measured (c, d). e, f Gfp, HA-RAB12, HA-RAB12-K42A, or HA-RAB12-Y111A were transduced into astrocytes by AAVs carrying GFAP promotors. The infected astrocytes’ centrosome volume was determined (n = 3). g, h The centrosome volume in WT and Lrrk2 KO astrocytes infected with AAV-GFAP-Gfp, or AAV-GFAP-HA-Rab12 was measured (n = 4). The representative cilia from (c, d) and centrosomes from (e) and (g) were enlarged in each panel. Each dot in the graphs represents >60 cells from each mouse. P-values were calculated by one-way ANOVA (c, f) or unpaired two-tailed t-tests (d, h). Dunnett’s (c) and Turkey’s (f) multiple comparisons tests were performed when P-values from one-way ANOVA < 0.05. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; error bars = SEM; scale bars are indicated. Source data are provided as a Source Data file.
We next asked whether LRRK2-binding mutants of RAB12 are affected by the centrosome regulation. Overexpression of HA-RAB12WT-K42A (HA-RAB12-K42A) or HA-RAB12WT-Y111A (HA-RAB12-Y111A) was unable to induce centrosome expansion in astrocytes in contrast to the HA-RAB12 as shown in the staining with anti-Pericentrin antibody (Fig. 4e, f). In addition, overexpression of HA-RAB12 caused a smaller effect in the increasing volume of Pericentrin+ centrosomes in Lrrk2 KO astrocytes than that in control astrocytes (Fig. 4g, h, Fig. S6c), suggesting that the effect of RAB12 overexpression in centrosome amplification depends on LRRK2 and requires LRRK2 binding. Taken together, the above results demonstrated the requirement of a direct LRRK2 binding for the role of RAB12 in controlling the homeostasis of primary cilia and centrosomes.
RAB12-mediated recruitment of pRAB10/RILPL1 to centrosomes and elevation of pRAB10 levels requires LRRK2 binding in astrocytes
Previous studies suggested that RAB10 and RILPL1 regulate centrosomes and primary cilia16,18,19,24,41. We next asked whether RAB10 and RILPL1 participate in the action of RAB12 (or active RAB12) in the regulation of centrosomes and cilia in astrocytes. Overexpression of HA-RAB12 promoted co-localization and clustering of the endogenous LRRK2 (Fig. 5a), phospho-RAB10 (pRAB10) (Fig. 5b), and RILPL1 (Fig. 5c) with a marked increase of their staining intensity, compared to GFP-overexpressed astrocytes. The specificity of the endogenous LRRK2 staining was validated using Lrrk2 KO astrocytes (Fig. S7a). The clustering of the endogenous LRRK2, pRAB10, and RILPL1 colocalized with Pericentrin or γ-tubulin staining (Fig. 5a–c), suggesting that they are recruited to amplified centrosomes or centrioles. Interestingly, although the overexpression of HA-RAB12QL also increased the intensity of pRAB10 and RAB12-LRRK2 co-clustering, the active mutant did not show a significant co-localization with pRAB10, RILPL1, or centrosome markers (Fig. 5a–c).
a–c Primary culture of astrocytes was infected with AAV-GFAP-Gfp, AAV-GFAP-HA-Rab12, or AAV-GFAP-HA-Rab12QL. IF staining was performed to detect LRRK2 (a), pRAB10 (b), and RILPL1 (c) in the cells by corresponding antibodies. Images as shown in (a–c) were quantified to determine LRRK2-GFP/HA Pearson coefficient (n = 3 for each group), LRRK2 intensity colocalized with GFP or HA (n = 7), and LRRK2-γ-tubulin Pearson coefficient (n = 3) in (a), pRAB10-GFP/HA Pearson coefficient (n = 3), pRAB10 intensity colocalized with GFP or HA (n = 3), and pRAB10-Pericentrin Pearson coefficient (n = 4) in (b), and RILPL1-GFP/HA Pearson coefficient (n = 3) and RILPL1 intensity colocalized with GFP or HA (n = 3) in (c). d AAVs carrying Gfp, HA-RAB12, HA-RAB12QL, or their LRRK2-binding mutants, such as HA-RAB12-K42A, HA-RAB12-Y111A, HA-RAB12QL-K42A, or HA-RAB12QL-Y111A were used to infect primary culture of astrocytes. IF staining was performed to detect pRab10 in the infected cells, and its intensity was measured (n = 3 for each group). The magnified views of the dash boxes in the images in (a–d) were displayed in the last image column in each panel. e, f. Immunoblot analysis of primary cultured astrocytes overexpressing GFP, HA-RAB12, or HA-RAB12QL. The levels of pRAB10, total RAB10, RILPL1, GFP, HA, pRAB12, total RAB12, LRRK2 p-S1292 (n = 3 for each group), and total LRRK2 (n = 9 for each group) were detected. g, h Immunoblot analysis of WT and Rab12 KO primary cultured astrocytes. The levels of pRAB10, total RAB10, RILPL1, LRRK2 p-S1292, total LRRK2, pRAB12, and total RAB12 were quantified (n = 6 for each group). Each dot in the graphs represents >60 cells from each mouse (a–d) or cells from an individual mouse (f, h). P-values were calculated using one-way ANOVA followed by Dunnett’s (a–d) and Turkey’s (f) multiple comparisons tests, or unpaired two-tailed t-tests (h). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; error bars = SEM; scale bars are indicated. Arb. units: arbitrary units. Source data are provided as a Source Data file.
Furthermore, we found that overexpression of LRRK2-binding mutant HA-RAB12-K42A or HA-RAB12-Y111A failed to potentiate pRAB10 levels or recruit pRAB10 to the RAB12+ clusters (Fig. 5d). Lack of increase of pRAB10 intensity was also observed with the overexpression of active RAB12 mutants (HA-RAB12QL-K42A or HA-RAB12QL-Y111A) in astrocytes (Fig. 5d). In addition, the pRAB10 staining intensity was completely abolished in Lrrk2 KO astrocytes with the overexpression of HA-RAB12WT or active HA-RAB12QL (Fig. S7b, c). The levels of RILPL1 colocalizing with HA-RAB12 were also markedly reduced in Lrrk2 KO compared to the control astrocytes although the total fluorescent intensity of RILPL1 did not change (Fig. S7d–f), indicating RAB12 overexpression-induced RILPL1 recruitment is likely partially mediated by LRRK2.
We next performed immunoblot to analyze the impact of overexpression or loss-of-function of Rab12 in astrocytes. Both WT and active RAB12 overexpression led to a significant increase of pRAB10 and reduction of RILPL1 and LRRK2 p-S935 levels but with no impact on LRRK2 p-S1292 and total RAB10 levels. Only WT RAB12 overexpression, but not its active form, resulted in a reduction of endogenous total LRRK2 protein levels (Fig. 5e, f, Fig. S7g, h). In contrast, Rab12 KO astrocytes showed a decrease of pRAB10 and little change in total RAB10, RILPL1, LRRK2 p-S935, p-S1292, or total LRRK2 levels (Fig. 5g, h, Fig. S7i, j). Thus, our results collectively demonstrate the critical role of Rab12 in the control of RAB10 phosphorylation levels and the recruitment of pRAB10/RILP1 to centrosomes, which requires direct LRRK2 binding.
Block of LRRK2 kinase activity prevents RAB12-mediated centrosome amplification and suppression of primary ciliogenesis
We next asked if the block of LRRK2 activity influences the RAB12-regulated centrosome and primary cilia homeostasis. We treated GFP, HA-RAB12, or HA-RAB12QL overexpressed astrocytes with a well-characterized LRRK2 inhibitor MLi242 and found effective depletion of pRAB10 signals in all cases (Fig. 6a, b). The LRRK2 inhibitor significantly reduced the volume of γ-tubulin or Pericentrin labeled centrosomes in HA-RAB12 overexpressed astrocytes (Fig. 6c–e), while it had a negligible effect on the centrosomes in GFP-overexpressed astrocytes. Moreover, the MLi2 treatment rescued the decreased ciliated astrocyte percentage caused by the overexpression of active HA-RAB12QL and did not change the cilia length or volume (Fig. 6f–i). Thus, the data suggests that the block of LRRK2 activity abolishes the phosphorylation of RAB10 and consequently prevents the deregulated centrosome and primary ciliogenesis caused by RAB12 hyperactivity.
a, b Primary culture of astrocytes was infected with AAV-GFAP-Gfp, AAV-GFAP-HA-Rab12, or AAV-GFAP-HA-Rab12QL and pRAB10 average intensity was quantified in the infected cells treated with MLi2 or DMSO (n = 6 for each group). c–e The volume of γ-tubulin and Pericentrin was examined in astrocytes infected with AAV-GFAP-Gfp, or AAV-GFAP-HA-Rab12 and treated with MLi2 or DMSO (n = 6 for each group). f–i The ciliated cell percentage, cilia length, and cilia volume were quantified in astrocytes infected with AAV-GFAP-Gfp, or AAV-GFAP-HA-Rab12QL (n = 6 for each group). The boxed areas in (a), (c), and (f) were enlarged as shown. Each dot in the graphs represents >60 cells from each mouse. Unpaired two-tailed t-tests, *P < 0.05, ****P < 0.0001; error bars = SEM; scale bars are indicated. Arb. units: arbitrary units. Source data are provided as a Source Data file.
Loss of Rab12 rescues Lrrk2-G2019S - induced ciliary deficiency and centrosome alteration in astrocytes
A previous study showed that PD mutation LRRK2-G2019S led to cilium deficiency and enhanced rate of split centrosomes due to its increased kinase activity17. We thus asked if the pathogenic effect of LRRK2-G2019S occurs in astrocytes and depends on RAB12. Indeed, we observed that the primary astrocytes from BAC transgenic Lrrk2-G2019S mice32 (GStg) showed decreased ciliated astrocyte percentage (with no change in cilia volume or length) and increased incidence of split centrosomes compared to control astrocytes (Fig. 7), consistent with the previous observations24. Interestingly, mutant astrocytes derived from compound mice combining GStg and Rab12 KO showed no difference in the percentage of ciliation or split centrosomes when compared to control astrocytes (Fig. 7). The above observations demonstrated that the disruption of Rab12 abolished the impact of the Lrrk2 disease mutant on primary cilia and centrosome homeostasis in astrocytes.
The mice overexpressing Lrrk2-G2019S (GStg) were crossed with Rab12 KO mice to generate GStg + Rab12 KO double mutant mice. IF staining was performed to detect the cilia and the centrosomes in the primary cultures of astrocytes from WT, GStg, Rab12 KO, and GStg + Rab12 KO neonatal mice. The images from each group were analyzed to quantify the ciliated cell percentage, cilia length, cilia volume, and split centrosome percentage. In this study, split centrosomes are defined as the distance between the duplicated centrosomes in individual cells >0.8 μm24,64. The enlarged images show the representative cilia and centrosomes inside the box areas. Each dot in the graphs represents >60 cells from each mouse and n = 4 for each group. One-way ANOVA followed by Dunnett’s multiple comparisons test, *P < 0.05, **P < 0.01; error bars = SEM; scale bars are indicated. Source data are provided as a Source Data file.
Discussion
Our current study determines the structure of RAB12-LRRK2 complex and provides the atomic details for RAB12-LRRK2 interaction. We show that the RAB12-LRRK2 complex regulates primary ciliogenesis and centrosome homeostasis in astrocytes and that RAB12 is crucial for LRRK2 kinase activity acting upon the phosphorylation of RAB10. Furthermore, the pathogenic effects of LRRK2-G2019S on cilia and centrosome homeostasis in astrocytes depend on RAB12. Thus, our study identifies the interaction between LRRK2 and RAB12 as a therapeutic target.
Our structural analysis demonstrates that multiple small RAB GTPases, including RAB12, RAB29, and RAB32a, share the similar binding site of the CDR1 and Switch I-Interswitch-Switch II motifs in LRRK2 binding34. However, RAB12 differs from others by binding to the H1 helices of ARM6-ARM7, while RAB29 and RAB32 primarily interact with the H2 helices of ARM9-ARM10 of LRRK2 (Fig. 1). RAB12 and RAB29 were previously shown to activate LRRK2 when monitoring pRAB10 and LRRK2 Ser1292 autophosphorylation22,31,43. The RAB29-dependent activation of LRRK2 with Ser1292 autophosphorylation was proposed to be mediated by the formation of asymmetric LRRK2 tetramers34. However, the binding of RAB12 on ARM6-ARM7 is not comparable with the tetrameric configuration of LRRK2 (Fig. S3g), suggesting that RAB12-dependent activation should be mechanistically different from RAB29-dependent activation of LRRK2 at the molecular level. In fact, we observed that deletion or overexpression of Rab12 has little effect on Ser1292 autophosphorylation of LRRK2 in contrast to the phospho-Rab10 levels, indicating that the role of RAB12 in activating LRRK2 kinase activity could be substrate specific.
Through both loss-of-function and gain-of-function analysis, we establish the physiological function of RAB12 as an inhibitor of primary ciliogenesis in astrocytes. We show that RAB12 is essential for the maintenance of primary ciliary homeostasis in astrocytes of multiple brain regions including the cortex and the striatum. Our result indicates altered cilium size (unchanged ciliated cell percentage) in Rab12 KO neurons, implicating RAB12 in ciliary regulation also in neurons. A recent study suggested a potential function of RAB12 in ciliogenesis in RPE and A549 cell cultures with unclear mechanism22. Our study provides insight into the mechanism whereby RAB12 suppresses ciliogenesis. We observe that the expression of active RAB12 reduces ciliated astrocyte percentage, which requires a direct binding between RAB12 and LRRK2 and the kinase activity of LRRK2. Interestingly, we find that overexpression of wildtype RAB12 not only abolishes the primary ciliogenesis but also drives centrosome/centriole amplification. Such an effect of RAB12 also requires direct LRRK2 interaction and LRRK2 kinase activity. Furthermore, the centrosome/centriole amplification involves the recruitment of pRAB10 and RILPL1. The lack of centrosome phenotypes in astrocytes with active RAB12 overexpression may reflect an advanced stage, whereas wildtype RAB12 overexpression causes an intermediate phase that “arrests” amplified centrosomes to be resolved by RAB12 activation associated with the membrane. Several lines of evidence have demonstrated that centrosome/centriole amplification could lead to the failure of ciliogenesis37,44,45. Thus, our study suggests a model that the RAB12-LRRK2 protein complex suppresses primary ciliogenesis through altering centrosome/centriole homeostasis and recruiting centrosomal pRAB10 and RILPL1. Previous works proposed that LRRK2 signaling converged on a centriolar pRAB10-RILPL1 complex to cause the deregulation of centrosomes and that ciliogenesis and centrosomal defects are the two distinct consequences of LRRK2 downstream signaling24. Our study, however, reveals RAB12 as a critical link coupling the two processes regulated by LRRK2 signaling. Furthermore, we observe no effect of RAB8a, RAB10, or LRRK2WT overexpression in altering primary cilia morphology or centrosome amplification, suggesting that RAB12 is a rate-limiting factor.
Our study clearly demonstrates the physiological function of RAB12 in controlling homeostatic LRRK2 kinase activity towards the phosphorylation of small GTPase RAB10 in astrocytes or mouse brains, placing RAB12 upstream of LRRK2-RAB10 signaling. Our study also suggests the presence of RAB12-LRRK2 complex under basal conditions, which helps maintain the homeostatic levels of pRAB10. We hypothesize that the basal RAB12-LRRK2 complex plays a role in preventing excessive primary ciliogenesis through regulating centrosome homeostasis in astrocytes. Such a homeostatic function of the RAB12-LRRK2 complex, however, is disrupted by the LRRK2-G2019S mutation. This gain-of-function mutation exacerbates the suppression of primary ciliogenesis and increases the occurrence of split centrosomes, based on our observations and others19,24. It remains possible that the split centrosomes corroborate the stage of cell division in astrocytes, leading to the inhibition of primary ciliogenesis24,46. Our study also demonstrates that the LRRK2-G2019S-induced defects in the percentage of ciliation and split centrosomes were brought to the normal levels as in WT astrocytes after Rab12 deletion (Fig. 7). The result reveals the critical role of RAB12 in mediating the pathogenic functions of the LRRK2-G2019S in impairing ciliogenesis and causing excessive split centrosomes in astrocytes. Our findings, therefore, suggest that disruption of RAB12-LRRK2 interaction can be explored to block LRRK2 pathogenic signaling in addition to LRRK2 kinase inhibition in therapeutic development. For such a purpose, our RAB12-LRRK2 structure provides valuable information for designing the intervention to disrupt the RAB12-LRRK2 complex formation.
Our study has limitations and raises many questions. First, we showed that RAB12 is a robust substrate of LRRK2 in the brain and forms a complex with LRRK2. It is unclear, however, how phosphorylation of RAB12 may affect RAB12-LRRK2 interaction and RAB12 function. According to the structure of the RAB12-LRRK2 complex, RAB12’s phosphorylation site S106 does not directly involve binding to LRRK2 (Fig. 1b). We speculate that the p-S106 of Rab12 is unlikely to affect its binding to LRRK2. A report also showed that RAB12’s phospho-dead mutant is fully capable of recruiting and activating LRRK222. Future studies should elucidate the function of RAB12 phosphorylation. Second, our phosphoproteomics study did not find any significant changes in the phosphorylation sites of known LRRK2 substrates (RAB proteins) beyond RAB12, such as RAB8, RAB10, or RAB29. It remains unclear whether RAB12 is the most (patho)physiological substrate of LRRK2 in the brains. The result could be due to the low abundance of the above RAB proteins in mouse brain tissues and the sensitivity of the current methodology. Interestingly, a previous report found that LRRK2-R1441C mutation increases the levels of pRAB12, but not pRAB10, in mouse brain, lung, and kidney tissues47. Third, the detailed molecular mechanism for how the RAB12-LRRK2 complex inhibits ciliogenesis and regulates centrosome homeostasis is unknown.
Finally, although our study establishes that RAB12-LRRK2 complex controls ciliary and centrosomal homeostasis, it is unclear whether such a function is related to the previously reported role of RAB12 in regulating lysosome damage or stress response29,31. Furthermore, the significance of astrocytic ciliary defects and centrosome abnormality induced by the LRRK2 variant in the pathogenesis of PD remains to be clarified. Nonetheless, our structural analysis and functional characterization of the RAB12-LRRK2 complex provide insight into the pathogenic mechanism and pave the way for the development of novel therapeutics for PD.
Methods
Cloning, expression, and purification of human LRRK2 and RAB12
The human LRRK2 construct was expressed and purified as described4,34. Briefly, bacmids carrying LRRK2 construct were generated in E. coli DH10Bac cells (Invitrogen). Recombinant baculoviruses were produced and amplified in Sf9 cells. Then, the P3 virus of full-length LRRK2 was used to transfect HEK293F cells for protein expression. Briefly, for 1 L cultures of HEK293F cells (~2–3 × 106 cells/mL) in Freestyle 293 media (Gibco) supplemented with 2% FBS (Gibco), about 100 ml P3 virus was used. Infected cells were incubated at 37 °C overnight, and protein expression was induced by adding 10 mM sodium butyrate. Cells were cultured at 30 °C for another 48–60 h before harvest. The cell pellet from the 600 mL culture was resuspended in 30 mL lysis buffer (20 mM Tris pH 8.0, 200 mM NaCl, 5% glycerol, 2 mM DTT, and protease inhibitors), and then cells were lysed by brief sonication. LRRK2 was separated from the insoluble fraction by high-speed centrifugation (38,000 × g for 1 h), and incubated with 1 mL CNBr-activated sepharose beads (GE Healthcare) coupled with 1 mg high-affinity GFP nanobodies (GFP-NB)48. The GFP tag was cleaved by preScission protease at 4 °C, and LRRK2 was further purified by size-exclusion chromatography with a Superose 6 Increase 10/300 GL column (GE Healthcare) equilibrated with 20 mM Tris pH 8.0, 200 mM NaCl and 2 mM DTT. The purified protein was collected and concentrated to 12.5 mg/ml (OD280) using a 100-kDa MWCO centrifugal device (Ambion), flash-frozen in liquid N2, and stored at −80 °C.
DNA encoding human RAB12 (residues 36–211) was synthesized by Genewiz and subcloned into the modified pGEX plasmid (Sigma), which attaches a GST tag and a TEV protease cleavage site at the N-terminus. For structural and biochemical studies, we generated the RAB12Q101L mutant, which was constructed using the QuickChange Site-Directed Mutagenesis kit (Strategene). The recombinant proteins were overexpressed in E. coli strain BL21 (DE3) in LB media supplemented with 0.05 mg/ml ampicillin, and cells were grown at 37 °C until OD600 reached 0.8. Further expression was induced by adding 0.4 mM IPTG, and cells were allowed to grow at 16 °C for 20 h. Harvested cells were lysed by sonication in the lysis buffer (20 mM Tris-HCl pH 8.0, 200 mM NaCl, 5% glycerol, 5 mM MgCl2, and 1 mM PMSF), and lysates were cleared by centrifugation at 38,000 × g for 45 min. Subsequently, the proteins were purified by GST affinity chromatography on the glutathione Sepharose beads (GE Healthcare) in lysis buffer. The bound proteins were then eluted with lysis buffer containing 20 mM GSH and incubated with 1 mM GppNHp at 4 °C overnight, followed by gel filtration chromatography using a Superdex 200 Increase 10/300 GL column (GE Healthcare) in a storage buffer (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 1 mM MgCl2, and 2 mM DTT). For the structural study, the GST tag was removed by on-column cleavage with TEV protease at 4 °C overnight before gel filtration chromatography. The peak fractions were collected and concentrated to 4.3 mg/ml (OD280), flash-frozen in liquid N2, and stored at −80 °C.
Cryo-EM sample preparation
Cryo-EM grids were prepared with a Vitrobot Mark IV (FEI). Quantifoil R1.2/1.3 300 Au holey carbon grids (Quantifoil) were glow-discharged for 30 s. Purified full-length LRRK2 and RAB12Q101L were incubated together on the ice for 1 h with a final concentration of 30 μM and 60 μM in the presence of 2 mM AMP-PNP, 1 mM GppNHp, and 1 mM MgCl2. In addition, 2.3 mM fluorinated Fos-Choline-8 was added right before freezing the grids, and then 3.5 μL of protein sample were pipetted onto the grids, which were blotted for 3.5 s under blot force −3 at 16 °C and 95% relative humidity and plunge-frozen in liquid nitrogen-cooled liquid ethane.
Cryo-EM data acquisition and processing
The RAB12-LRRK2 dataset was collected on a Titan Krios (Thermo Fisher Scientific) transmission electron microscope equipped with a K3 direct electron detector and post-column GIF energy filter (Gatan). Data collection was performed in an automated manner using EPU (Thermo Fisher Scientific). Movies were recorded at defocus values from −1.2 to −2.6 μm at a magnification of 130k× in hardware binning mode, corresponding to a pixel size of 0.6485 Å at the specimen. During 2.0 s exposure, 70 frames were collected with a total electron dose of ~78 e-/Å−2 (at a dose rate of 1.1143 e-/frame/Å2). In total, 20,023 images were collected. Motion correction was performed on hardware-binned movie stacks and binned by 1 using MotionCor249. Contrast transfer function (CTF) estimation was performed using Gctf50. Prior to particle picking, micrographs were analyzed for good power spectrum, and the bad ones were discarded (with 19,637 good images remaining).
Particles were selected using the template picker with reported inactive full-length LRRK2 structures4 as a reference in cryoSPARC51 and extracted using a binning factor of 2. Several rounds of the 2D classification were performed to eliminate ice artifacts, carbon edges, and false-positive particles containing noise. During 2D classification, two groups of good classes were observed, corresponding to the RAB12-LRRK2 monomer and dimer states, respectively. Both groups were selected, and ab initio reconstruction was performed. In order to further separate the RAB12-LRRK2 monomer and dimer states, we performed Heterogeneous refinement in cryoSPARC. As a result, 95,596 particles were assigned to the monomer class and 71,784 particles to the dimer. Both 3D classes were further refined using cryoSPARC after extraction of unbinned particles corresponding to each identified subset. For the RAB12-LRRK2 monomer state, we performed a standard NU refinement without imposing symmetry, followed by a focused 3D refinement (with a soft mask around the RAB12-LRRK2 interface region) to improve the map quality of the ARM domain of the LRRK2 with RAB12 binding. All resolution estimates were calculated according to the gold-standard Fourier shell correlation (FSC) using the 0.143 criterion52. Local resolution was estimated in cryoSPARC. The density maps were B-factor sharpened in cryoSPARC and used to produce figures and build models.
Model building and refinement
The reported structures of LRRK2 (PDB 7LI4)4 and RAB12 (PDB 2IL1) were fitted and adjusted into the cryo-EM maps of RAB12-LRRK2 monomer state using Chimera53 and Coot54. The structural model was refined against the map using the real space refinement module with secondary structure and non-crystallographic symmetry restraints in the Phenix package55. Fourier shell correlation curves were calculated between the refined model and the full map. The geometries of the models were validated using MolProbity56. All the figures were prepared in PyMOL (Schrödinger, LLC.), UCSF Chimera53 and UCSF ChimeraX57.
Animals
All animal studies were carried out according to the guidance of the Icahn School of Medicine at Mount Sinai Animal Care and Use Committee (IACUC-2021-00031). All the mice in this study were adopted under a 12 h light/12 h dark cycle environment at 20–23 °C and 40–60% humidity with available food and water. Lrrk2 BAC transgenic mice overexpressing Lrrk2 wildtype (WTtg) and PD-mutant G2019S (GStg) were generated by our group32. Lrrk2 knockout (KO) mice (strain #: 012453 for phosphoproteomics profiling; strain #: 016121 for primary astrocyte culture) and C57BL/6J mice (WT, strain #: 000664) were purchased from The Jackson Laboratory. Rab12 KO mice were generated by deleting the exon 3 of Rab12 gene from the mouse genome of C57BL/6J background using CRISPR-Cas9 technology by our group. For the generation of GStg + Rab12 KO double mutant mice, GStg heterozygous mice were crossed with Rab12 KO homozygous mice to obtain GStg heterozygous + Rab12 KO homozygous mice. PCR was performed with genomic DNA isolated from mouse tail clips to confirm mice genotypes. The genotyping primer sequences for both WTtg and GStg are 5′- GACTACAAAGACGATGACGACAAG-3′ (forward) and 5′-CTACCACCACCCAGATAATGTC-3′ (reverse). For Lrrk2 KO strain #: 012453′s genotyping, the primers are 5′- AGGAGGAGGAGGCTCTGAAG-3′(forward) and 5′-GGCATTCGGGAAACATCACT-3′ (reverse). For Lrrk2 KO strain #: 016121′s genotyping, the primers are 5′- CTTGTTGGTCACTCACAGAGCAAGATAGTT-3′(common forward), 5′- CTACTTCATACTTTCCTTGATGTTTGGTGG-3′ (reverse of KO mutant), and 5′- CCACTCCTCTGTCTCCGTCT-3′ (reverse of WT). For Rab12 KO′s genotyping, the primers are 5′-GGTTGTTCATCTCGGTGGGT-3′ (forward) and 5′-GTATTTCCCCAGTGCCCCAT-3′ (reverse).
Mouse brain tissue collection and processing
Mice from both males and females at 3 months old were anesthetized with 10 mg/kg xylazine and 50 mg/kg ketamine by intraperitoneal injection. For the samples used for phosphoproteome profiling and western blot, the mice were sacrificed by cervical dislocation after anesthetization. Then, striatal tissues were dissected from their brains and stored at – 80 °C for further processing. For the samples used for immunofluorescent staining, the anesthetized mice were cardiac perfused with cold PBS followed by 4% paraformaldehyde-PBS solution. The mouse brains were dissected, kept in the 4% paraformaldehyde-PBS solution at 4 °C overnight, and sequentially dehydrated in 30% sucrose-PBS buffer for 72 h. Finally, the brain tissues were embedded in OTC (Fisher Scientific, #23-730-571), frozen on dry ice, and sectioned at 30 μms with a cryostat (Leica) for the following experiments.
Whole proteome profiling and phosphoproteome profiling by TMT-LC/LC-MS/MS
Proteome and phosphoproteome profiling were performed as described previously58. Mouse brain samples (n = 3 for Lrrk2 KO, n = 2 for WT, n = 2 for WTtg, n = 3 for GStg) were lysed in fresh lysis buffer (50 mM HEPES, pH 8.5, 8 M urea, 0.5% sodium deoxycholate and phosphatase inhibitor cocktail) (PhosphoSTOP, Roche). The protein concentration of the lysate was quantified by BCA protein assay (Thermo Fisher Scientific). One mg of proteins from each sample were digested with Lys-C (Wako, 1:100 w/w) at room temperature for 2 h, diluted 4 times with 50 mM HEPES, pH 8.5, and further digested with trypsin (Promega, 1:50 w/w) overnight at room temperature followed by Cys reduction and alkylation. The peptides were acidified by 1% trifluoroacetic acid, desalted with Sep-Pak C18 cartridge (Waters), and then labeled with 10-plex TMT reagents (Thermo Fisher Scientific) followed by mixing equally. The mixture was desalted, dried, and solubilized in 600 µL of buffer A (10 mM ammonium formate, pH 8) and separated on two concatenated XBridge C18 columns (3.5 μm particle size, 4.6 mm × 25 cm, Waters) into 42 fractions with a 60 min gradient from 13% to 45% buffer B (95% acetonitrile, 10 mM ammonium formate, pH 8) with flow rate of 0.4 mL/min. For each fraction, phosphopeptide enrichments were carried out by TiO2 beads (GL sciences) as previously reported59. Briefly, the dried peptides were dissolved in 100 μL of binding buffer (65% acetonitrile, 2% TFA, and 1 mM KH2PO4). TiO2 beads (1 mg) were washed twice with washing buffer (65% acetonitrile, 0.1% TFA), mixed with the peptide solution, and incubated with end-over-end rotation at room temperature for 20 min. The phosphopeptide-bound beads were collected by brief centrifugation, washed twice with 150 μL washing buffer, and transferred to a C18 StageTip (Thermo Fisher Scientific) sitting on the top of a 2-mL centrifuge tube. The StageTip was centrifuged to remove the wash buffer completely and phosphopeptides were eluted under basic pH conditions (15 μL, 15% NH4OH, 40% acetonitrile). The eluents were dried, dissolved in 5% formic acid, and loaded on a reverse phase column (50 µm × 40 cm, 1.9 µm C18 resin) (Dr. Maisch GmbH, Germany) interfaced with a Q-Exactive HF mass spectrometer (Thermo Fisher Scientific). Peptides were eluted at 65 °C by 9–35% buffer B gradient in 4 h (buffer A: 0.2% formic acid, 3% DMSO; buffer B: buffer A plus 67% acetonitrile, flow rate of 0.15 µL/min). The mass spectrometer was operated in data-dependent mode with a survey scan in Orbitrap (60,000 resolution, 1 × 106 AGC target and 50 ms maximal ion time) and 20 MS/MS high-resolution scans (60,000 resolution, 1 × 105 AGC target, 105 ms maximal ion time, HCD, 35 normalized collision energy, 1.5 m/z isolation window, and 20 s dynamic exclusion). The whole proteome profiling was performed in a similar protocol but excluding the step of phosphopeptide enrichment.
The data analysis was performed by our JUMP suite60. Briefly, acquired MS/MS raw files were converted into mzXML format and searched by the JUMP algorithm against a composite target/decoy database to estimate FDR. The target protein database was downloaded from the Uniprot mouse database and the decoy protein database was generated by reversing all target protein sequences. Searches were performed with 10 ppm mass tolerance for precursor ions and 15 ppm for fragment ions, fully tryptic restriction, two maximal missed cleavages and the assignment of a, b, and y ions. TMT tags on lysine residues and peptide N termini (+229.16293 Da) and carbamidomethylation of Cys residues (+57.02146 Da) were used for static modifications and the dynamic modifications included the oxidation of methionine residues (+15.99492 Da) and Ser/Thr/Tyr phosphorylation (+79.96633). The assigned peptides were filtered by mass accuracy, minimal peptide length (7), matching scores, charge state and trypticity to reduce phosphopeptide FDR to 1%. Phosphosite reliability was evaluated by the localization score (Lscore) from JUMP suite. TMT reporter ion intensities of each PSM were extracted. The raw intensities were then corrected based on isotopic distribution of each labeling reagent and loading bias. The mean-centered intensities across samples were calculated and phosphopeptide intensities were derived by averaging related PSMs. Finally, phosphopeptide absolute intensities were determined by multiplying the relative intensities by the grand-mean of three most highly abundant PSMs. The whole proteome analysis was carried out similarly but excluding the mass shift of phosphorylation.
Proteins or phosphopeptides showing changes were identified using P-values calculated through the limma package61, which employs moderated one-way ANOVA for the four genotypes’ comparison. These P-values were subsequently adjusted using the Benjamini–Hochberg procedure to control the false discovery rate. Phosphopeptides that passed the adjusted P-value threshold of 0.05 were then examined manually. In addition, the phosphopeptide levels were also normalized by their related protein levels to investigate the impact of kinase activity62.
Plasmid construction
Total RNA was purified from the mouse brain tissue using RNeasy Mini Kit (QIAN, #74106) and was reverse-transcribed to cDNA. The mouse WT Rab12 (mRab12) coding sequence was amplified from the cDNA obtained. An HA tag was added to the N terminal of the mRab12 coding sequence and subcloned into pAAV-GFAP-EGfp (addgene, #50473) using SalI and EcoRI restriction enzyme sites and LV-hGFAP-Gfp (addgene, #183906) using AgeI and XhoI restriction enzyme sites, respectively. The original DNA fragments between the restriction enzyme sites in the two plasmids were removed. Mouse Rilpl1, Rab8a, and Rab10 were also amplified from the cDNA above. Rilpl1 was conjugated with a His tag at its C terminal, and a V5 tag and a Flag tag were conjugated at the N terminal of Rab8a and Rab10, respectively. All three genes were also inserted into LV-hGFAP-Gfp using AgeI and XhoI sites. Human WT RAB12 (hRAB12) and its QL variant (hRAB12QL) DNA fragments were amplified from pcDNA3.1-hRAB12 and pcDNA3.1-hRAB12QL generated in our lab before. NEBuilder® HiFi DNA Assembly Kit (NEB, # E2621L) was used to construct the mRab12Q100L variant, hRAB12 and hRAB12Q101L binding mutations (K42A and Y111A). All these mutant variants were subcloned into the same location of pAAV-hGFAP-GFP as done with mRab12.
Virus production
For AAV particles generation for in vitro studies, the AAV plasmids harboring the genes of interest were co-transfected with helper plasmids pAAV2/1 (addgene, #112862) and pAdDeltaF6 (addgene, #112867) into HEK293T cells (ATCC, #CRL-11268) using lipofectamine 3000 (Thermo Fisher Scientific, #L3000008). The medium was changed 6 h after transfection and replaced with fresh culture medium (DMEM with 10% fetal bovine serum). The medium was collected 48 h after transfection and centrifuged at 1000 × g for 10 min to spin down the cell debris. The supernatant was transferred to a new tube followed by addition of Polyethylene glycol 8000 (PEG000)-NaCl buffer (40% PEG8000, 2.5 M NaCl) to precipitate the AAV particles at 4 °C overnight. Then, the AAV particles were spun down at 4000 × g at 4 °C for 1 h. The supernatant was removed, and the remaining pellets were resuspended in glia medium (DMEM with 10% fetal bovine serum, 1× GlutaMax, 1× sodium pyruvate, and 1% penicillin/streptomycin). The resuspended AAV particles were aliquoted and frozen at −80 °C.
For lentivirus production, HEK293T cells were co-transfected with lentiviral expression plasmids containing the gene of interest and helper plasmids pMDLg/pRRE (addgene, #12251), pRSV-Rev (addgene, #12253), and pCAG-VSVG (addgene, #64084). The culture was replaced with fresh culture medium 6 h post-transfection. The culture medium containing the lentivirus particles was collected at 24 h, 48 h, and 60 h post-transfection. The medium from the indicated time points was combined and centrifuged at 1000 × g for 10 min to remove the cell debris, and then transferred to a sterile ultracentrifuge tube. The lentivirus particles were precipitated by ultracentrifugation at 75,600 × g at 4 °C for 2 h. The supernatant was removed, and the lentivirus pellets were resuspended with 1× lenti-freezing medium (0.5 M sucrose in DMEM), aliquoted, and frozen at −80 °C.
Primary astrocyte culture
Briefly, P0-P2 neonatal mice were anesthetized on ice. The whole brains were dissected, and the olfactory bulbs, meninges, cerebellums, and brain stems were removed. The remaining tissues were kept in tubes containing chilled glia medium composed of DMEM with 10% fetal bovine serum (HyClone, #SH30396.03HI), 1× GlutaMax (GIBCO, #35050-061), 1× sodium pyruvate (GIBCO, #11360-070), and 1% penicillin/streptomycin (GIBCO, #15140-122). Brain tissue from different neonatal mice were placed in separate tubes. Cells were then dissociated using 1 mL pipette tips followed by a fire-polished glass pipette and cultured in T75 flasks in a humidified 5% CO2 incubator at 37 °C for 14 days. On DIV14, the astrocytes were detached from the flasks’ bottoms by 0.05% trypsin (GIBCO, #25200056) and replaced on culture plates for either immunoblot analysis or immunofluorescent staining.
Immunofluorescent staining
For immunofluorescent staining of the mouse brain, brain slices were mounted on a superfrost plus microscope slide (Thermo Fisher Scientific, # 12-550-15) and washed with PBS once for 5 min. Slices were blocked with blocking buffer (5% goat serum in PBS containing 0.3% Triton X-100) for 1 h at room temperature and incubated with primary antibodies at 4 °C for about 16 h. Then, slices were rinsed with PBST (PBS containing 0.05% Triton X-100) twice for 5 min each and once with PBS for 5 min, followed by incubation with corresponding fluorescent dye-conjugated secondary antibodies for 1 h at room temperature shielded from light. The secondary antibody was rinsed from the slices as previously indicated and incubated with 2 µM Hoechst solution (Thermo Fisher Scientific, #62249) for 5 min at room temperature. Finally, slices were rinsed with PBS twice for 5 min and mounted with ProLong™ Diamond Antifade Mountant (Thermo Fisher Scientific, # P36961). For the immunofluorescent staining for primary cultured astrocytes, the cells were fixed with 4% paraformaldehyde-PBS solution for 15 min at room temperature and sequentially washed three times with PBST. The cells were blocked and stained in a similar manner as described above, except that the concentration of Triton X-100 in the blocking buffer was reduced to 0.1%. For the centrosome staining, the cells were additionally fixed with methanol at −20 °C for 15 min after paraformaldehyde fixation for enhanced visualization. The information on the antibodies used is listed in Supplementary Table 2.
Co-immunoprecipitation
HEK293T cell line was maintained in DMEM supplemented with penicillin/streptomycin and 10% FBS (R&D Systems, #S11550). One day prior to transfection, HEK293T cells were seeded onto a 6-well plate at 6 × 105 cells per well. Cells were transfected using Lipofectamine® 3000 (Invitrogen, #L3000015). Plasmid DNA concentrations of 0.5 µg were used for RAB12, RAB12-Y111A, RAB12-K42A, RAB12QL, RAB12QL-Y111A, RAB12QL-K42A plasmids and co-transfected with 4.5 µg of Flag-tagged LRRK2WT plasmids for double transfected conditions. Plasmid DNA was added to 125 µL Opti-MEM containing 10 µL of P3000 followed by the addition of 125 µL of Opti-MEM containing 3.75 µL Lipofectamine 3000. The plate was incubated at 37 °C for 24 h in a 5% CO2 incubator. After incubation, transfected cells were then used for co-immunoprecipitation (Co-IP). Cells were lysed on ice with lysis buffer consisting of 1% Triton X-100 TBS buffer containing 1× Halt™ protease and phosphatase inhibitor cocktail (#78440, Thermo Fisher Scientific) and 5 mM EDTA (Thermo Fisher Scientific #1861274). The concentration of cell lysates was examined by BCA assay (Thermo Fisher Scientific, #23227), and 1000 µg of proteins from each sample were incubated in 30 µL Anti-FLAG® M2 Magnetic Beads (Millipore Sigma, #M8823) overnight at 4 °C. After incubation, beads were washed three times with 1% Triton X-100 TBS. Proteins bound to the M2 Magnetic Beads were eluted by boiling in 1× loading buffers composed of NuPAGE™ LDS Sample Buffer (Thermo Fisher Scientific, #NP0007) and 0.05 M DTT (Sigma, #11583786001) at 95 °C for 10 min. The supernatants were transferred into new Eppendorf tubes for immunoblot analysis after brief centrifugation.
Immunoblot analysis
Brain tissue samples were homogenized through bead disruption using the Bullet Blender (Next Advance Inc.) technology. Samples were put in 1.5 mL RINO tubes (Next Advance Inc.) with zirconium oxide beads (ZrOB05 + ZrOB10 mixture, Next Advance Inc.) in the lysis buffer described in the Co-IP procedure but without Triton X-100 and homogenized for 2 min at speed 12 at 4 °C. The tissue homogenates were mixed with an equal volume of lysis buffer with 2% Triton X-100 and then lysed on ice for 20 min before centrifugation. For cell-culture samples, cells were harvested and lysed in the lysis buffer on ice for 15 min, and vortexed briefly before centrifugation. The centrifugation conditions for tissue samples and cell samples were both at 16,000 × g for 10 min at 4 °C. The supernatants were transferred to new tubes and BCA assay was performed to determine the samples’ protein concentrations. The samples were mixed with 3× loading buffers to a final concentration of 1×, and incubated at 70 °C for 10 min. After being cooled down on ice, the samples were loaded onto an SDS-PAGE gel. Proteins were separated by electrophoresis at 120 V for 1.5 h and then electroeluted onto a PVDF membrane (BIO-RAD, #10026933) at 25 V, 1 A for 0.5 h using Trans-Blot® Turbo™ Transfer System (BIO-RAD). PVDF membranes were then washed with TBS for 5 min and blocked with Intercept Blocking Buffers (LI-COR, #927-60001) for 1 h at room temperature. Membranes were incubated in primary antibodies overnight at 4 °C, followed by three times wash in TBST (0.1% Tween-20 in TBS) for 5 min each. The appropriate secondary fluorescent or HRP-linked antibodies were applied for 1 h at room temperature and then washed off three times for 5 min each. The signals were visualized by the ChemiDoc Imaging system (BIO-RAD). Signal intensities were then measured by ImageJ (https://imagej.nih.gov/ij/index.html). The uncropped and unprocessed blots are provided in the Source Data file.
Electron microscope
Primary astrocytes were grown on Permanox® Slide (Electron Microscopy Sciences, #70400). Samples were prepared by washing cells with 0.1 M sodium cacodylate solution (Electron Microscopy Sciences, #15960-01) three times and fixed with sodium cacodylate solution containing 2% paraformaldehyde and 2.5% glutaraldehyde for 2 h at 4 °C. They were then post-fixed with sodium cacodylate containing 2% osmium tetroxide and 1.5% potassium ferricyanide for 1 h at room temperature. After being briefly washed by sodium cacodylate buffer, the cells were stained with 2% uranyl acetate in distilled H2O for 1 h at room temperature followed by dehydration in an ascending ethanol series. The cells were then embedded in resin through an increasing ethanol/resin series for 16–18 h (Electron Microscopy Sciences Embed 812 Kit, #14120), and incubated in a vacuum oven for 72 h at 60 °C. Ultra-thin sections were obtained on copper 300 mesh grids (Electron Microscopy Sciences, G300H-Cu) and counterstained with 1% uranyl acetate and lead citrate. Images were taken by an HT7500 transmission electron microscope (Hitachi High-Technologies, Japan). Sample processing and imaging were performed by the Microscopy and Advanced Bioimaging CoRE at the Icahn School of Medicine at Mount Sinai.
Image acquisition and analysis
The fluorescent image acquisition in this study was performed using z-stack with a 63× objective at 1024 × 1024 resolution with the Zeiss LSM 780 confocal microscope (Carl Zeiss). The z-step interval was 0.5 µm for astrocytes, and 0.75 µm for neurons63. Sixteen z-stack images were taken for cultured astrocytes and 50 images for mouse brain slices. The images from astrocytes and neurons were obtained from the dorsal striatum and frontal cortex areas in the mouse brain slices.
For ciliated cell15,63 and split centrosome24,64 percentage quantitation, the z-stack images were collapsed by the maximum intensity projection function in ImageJ (https://imagej.nih.gov/ij/index.html) and manually counted as described previously. CiliaQ, an open-source ImageJ plugin65, was used to measure the cilia length and volume of primary cultured astrocytes, whereas the surface module in Imaris software (Bitplane) was utilized to determine the cilia length and volume of astrocytes and neurons in the mouse brain slices, and the centrosome volume of primary cultured astrocytes. The fluorescent intensity and Pearson coefficient between proteins were quantified by the particle analysis plugin and the JaCoP plugin66 in ImageJ, respectively.
Statistics & reproducibility
GraphPad Prism 9 was used to execute the statistical analyses in this study. All the results were presented as mean ± standard error (SEM). As for comparisons between the two groups, Student’s t-test (unpaired, two-tailed) was applied to determine the significance. As for the comparisons with more than two groups, one-way ANOVA was performed. When one-way ANOVA indicated statistical significance, a post hoc pairwise comparison test was used for multiple group comparisons. P-value < 0.05 was defined as statistical significance. The number of biological replicates for each experiment is indicated in the corresponding figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD048732. The cryo-EM maps of the RAB12-LRRK2 complex have been deposited in the Electron Microscopy Data Bank under the accession codes EMD-43234 (RAB12-LRRK2 monomer) and EMD-43235 (RAB12-LRRK2 dimer). The corresponding coordinates have been deposited in the Protein Data Bank under the accession codes 8VH4 (RAB12-LRRK2 monomer) and 8VH5 (RAB12-LRRK2 dimer). Previously published structures used in model building and structural comparison are available in PDB with accession codes: 7LHW, 7LHT, 8FO2, 8FO9, 4CYM, 2IL1. Source data are provided with this paper.
Code availability
JUMPSuite [https://github.com/JUMPSuite/JUMP] was used to analyze the proteome and phosphoproteomics data.
References
Paisán-Ruíz, C. et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 44, 595–600 (2004).
Zimprich, A. et al. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44, 601–607 (2004).
Bonet-Ponce, L. & Cookson, M. R. LRRK2 recruitment, activity, and function in organelles. FEBS J. 289, 6871–6890 (2022).
Myasnikov, A. et al. Structural analysis of the full-length human LRRK2. Cell 184, 3519–3527.e10 (2021).
Deniston, C. K. et al. Structure of LRRK2 in Parkinson’s disease and model for microtubule interaction. Nature 588, 344–349 (2020).
Kalogeropulou, A. F. et al. Impact of 100 LRRK2 variants linked to Parkinson’s disease on kinase activity and microtubule binding. Biochem J. 479, 1759–1783 (2022).
Bardien, S., Lesage, S., Brice, A. & Carr, J. Genetic characteristics of leucine-rich repeat kinase 2 (LRRK2) associated Parkinson’s disease. Parkinsonism Relat. Disord. 17, 501–508 (2011).
Deng, H. et al. Genetic analysis of LRRK2 mutations in patients with Parkinson disease. J. Neurol. Sci. 251, 102–106 (2006).
Correia Guedes, L. et al. Worldwide frequency of G2019S LRRK2 mutation in Parkinson’s disease: a systematic review. Parkinsonism Relat. Disord. 16, 237–242 (2010).
Taylor, M. & Alessi, D. R. Advances in elucidating the function of leucine-rich repeat protein kinase-2 in normal cells and Parkinson’s disease. Curr. Opin. Cell Biol. 63, 102–113 (2020).
Pan, P. Y. et al. Parkinson’s disease-associated LRRK2 hyperactive kinase mutant disrupts synaptic vesicle trafficking in ventral midbrain neurons. J. Neurosci. 37, 11366–11376 (2017).
Steger, M. et al. Phosphoproteomics reveals that Parkinson’s disease kinase LRRK2 regulates a subset of Rab GTPases. eLife 5, e12813 (2016).
Steger, M. et al. Systematic proteomic analysis of LRRK2-mediated Rab GTPase phosphorylation establishes a connection to ciliogenesis. eLife 6, e31012 (2017).
Dou, D., Smith, E. M., Evans, C. S., Boecker, C. A. & Holzbaur, E. L. F. Regulatory imbalance between LRRK2 kinase, PPM1H phosphatase, and ARF6 GTPase disrupts the axonal transport of autophagosomes. Cell Rep. 42, 112448 (2023).
Dhekne, H. S. et al. A pathway for Parkinson’s disease LRRK2 kinase to block primary cilia and Sonic hedgehog signaling in the brain. eLife 7, e40202 (2018).
Dhekne, H. S. et al. LRRK2-phosphorylated Rab10 sequesters Myosin Va with RILPL2 during ciliogenesis blockade. Life Sci. Alliance 4, e202101050 (2021).
Khan, S. S. et al. Pathogenic LRRK2 control of primary cilia and Hedgehog signaling in neurons and astrocytes of mouse brain. eLife 10, e67900 (2021).
Sobu, Y., Wawro, P. S., Dhekne, H. S., Yeshaw, W. M. & Pfeffer, S. R. Pathogenic LRRK2 regulates ciliation probability upstream of tau tubulin kinase 2 via Rab10 and RILPL1 proteins. Proc. Natl Acad. Sci. USA 118, e2005894118 (2021).
Lara Ordóñez, A. J. et al. The LRRK2 signaling network converges on a centriolar phospho-Rab10/RILPL1 complex to cause deficits in centrosome cohesion and cell polarization. Biol. Open 11, bio059468 (2022).
Schmidt, S. et al. Primary cilia and SHH signaling impairments in human and mouse models of Parkinson’s disease. Nat. Commun. 13, 4819 (2022).
Yeshaw, W. M. et al. Localization of PPM1H phosphatase tunes Parkinson’s disease-linked LRRK2 kinase-mediated Rab GTPase phosphorylation and ciliogenesis. Proc. Natl Acad. Sci. USA 120, e2315171120 (2023).
Dhekne, H. S. et al. Genome-wide screen reveals Rab12 GTPase as a critical activator of Parkinson’s disease-linked LRRK2 kinase. eLife 12, e87098 (2023).
Fdez, E. et al. Pathogenic LRRK2 regulates centrosome cohesion via Rab10/RILPL1-mediated CDK5RAP2 displacement. iScience 25, 104476 (2022).
Lara Ordónez, A. J. et al. RAB8, RAB10 and RILPL1 contribute to both LRRK2 kinase-mediated centrosomal cohesion and ciliogenesis deficits. Hum. Mol. Genet. 28, 3552–3568 (2019).
Pfeffer, S. R. LRRK2 and Rab GTPases. Biochem. Soc. Trans. 46, 1707–1712 (2018).
Pfeffer, S. R. LRRK2 phosphorylation of Rab GTPases in Parkinson’s disease. FEBS Lett. 597, 811–818 (2023).
Kluss, J. H. et al. Preclinical modeling of chronic inhibition of the Parkinson’s disease associated kinase LRRK2 reveals altered function of the endolysosomal system in vivo. Mol. Neurodegener. 16, 17 (2021).
Kluss, J. H. et al. Lysosomal positioning regulates Rab10 phosphorylation at LRRK2(+) lysosomes. Proc. Natl Acad. Sci. USA 119, e2205492119 (2022).
Ito, K. et al. Pathogenic LRRK2 compromises the subcellular distribution of lysosomes in a Rab12-RILPL1-dependent manner. FASEB J. 37, e22930 (2023).
Kluss, J. H., Bonet-Ponce, L., Lewis, P. A. & Cookson, M. R. Directing LRRK2 to membranes of the endolysosomal pathway triggers RAB phosphorylation and JIP4 recruitment. Neurobiol. Dis. 170, 105769 (2022).
Wang, X. et al. Rab12 is a regulator of LRRK2 and its activation by damaged lysosomes. eLife 12, e87255 (2023).
Li, X. et al. Enhanced striatal dopamine transmission and motor performance with LRRK2 overexpression in mice is eliminated by familial Parkinson’s disease mutation G2019S. J. Neurosci. 30, 1788–1797 (2010).
Xu, J., Fotouhi, M. & McPherson, P. S. Phosphorylation of the exchange factor DENND3 by ULK in response to starvation activates Rab12 and induces autophagy. EMBO Rep. 16, 709–718 (2015).
Zhu, H. et al. Rab29-dependent asymmetrical activation of leucine-rich repeat kinase 2. Science 382, 1404–1411 (2023).
Sterpka, A. & Chen, X. Neuronal and astrocytic primary cilia in the mature brain. Pharmacol. Res. 137, 114–121 (2018).
Breslow, D. K. & Holland, A. J. Mechanism and regulation of centriole and cilium biogenesis. Annu Rev. Biochem. 88, 691–724 (2019).
Mahjoub, M. R. & Stearns, T. Supernumerary centrosomes nucleate extra cilia and compromise primary cilium signaling. Curr. Biol. 22, 1628–1634 (2012).
D’Assoro, A. B., Lingle, W. L. & Salisbury, J. L. Centrosome amplification and the development of cancer. Oncogene 21, 6146–6153 (2002).
Itoh, T., Satoh, M., Kanno, E. & Fukuda, M. Screening for target Rabs of TBC (Tre-2/Bub2/Cdc16) domain-containing proteins based on their Rab-binding activity. Genes Cells 11, 1023–1037 (2006).
Azouz, N. P., Matsui, T., Fukuda, M. & Sagi-Eisenberg, R. Decoding the regulation of mast cell exocytosis by networks of Rab GTPases. J. Immunol. 189, 2169–2180 (2012).
Schaub, J. R. & Stearns, T. The Rilp-like proteins Rilpl1 and Rilpl2 regulate ciliary membrane content. Mol. Biol. Cell 24, 453–464 (2013).
Fell, M. J. et al. MLi-2, a potent, selective, and centrally active compound for exploring the therapeutic potential and safety of LRRK2 kinase inhibition. J. Pharmacol. Exp. Ther. 355, 397–409 (2015).
Purlyte, E. et al. Rab29 activation of the Parkinson’s disease-associated LRRK2 kinase. EMBO J. 37, 1–18 (2018).
Dionne, L. K. et al. Centrosome amplification disrupts renal development and causes cystogenesis. J. Cell Biol. 217, 2485–2501 (2018).
Coelho, P. A. et al. Over-expression of Plk4 induces centrosome amplification, loss of primary cilia and associated tissue hyperplasia in the mouse. Open Biol. 5, 150209 (2015).
Goranci-Buzhala, G. et al. Cilium induction triggers differentiation of glioma stem cells. Cell Rep. 36, 109656 (2021).
Kalogeropulou, A. F. et al. Endogenous Rab29 does not impact basal or stimulated LRRK2 pathway activity. Biochem J. 477, 4397–4423 (2020).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Henderson, R. et al. Outcome of the first electron microscopy validation task force meeting. Structure 20, 205–214 (2012).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Tan, H. et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46, 488–503 (2017).
Tan, H. et al. Refined phosphopeptide enrichment by phosphate additive and the analysis of human brain phosphoproteome. Proteomics 15, 500–507 (2015).
Wang, X. et al. JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy. Mol. Cell. Proteom. 13, 3663–3673 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wang, H. et al. Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes. Nat. Commun. 10, 3718 (2019).
Shahzad, S. K., Herschel, S. D., Francesca, T., Suzanne, R. P. Analysis of primary cilia in rodent brain by immunofluorescence microscopy. protocolsio, https://doi.org/10.17504/protocols.io.bnwimfce (2020).
Fdez, E. et al. Protocol to measure centrosome cohesion deficits mediated by pathogenic LRRK2 in cultured cells using confocal microscopy. STAR Protoc. 4, 102024 (2023).
Hansen, J. N., Rassmann, S., Stüven, B., Jurisch-Yaksi, N. & Wachten, D. CiliaQ: a simple, open-source software for automated quantification of ciliary morphology and fluorescence in 2D, 3D, and 4D images. Eur. Phys. J. E Soft Matter 44, 18 (2021).
Bolte, S. & Cordelières, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Acknowledgements
We thank the support of Electron Microscopy core facility at Mount Sinai and the Cryo-Electron Microscopy Center at St. Jude Children’s Research Hospital for help with cryo-EM data collection. We thank members of the Yue lab and members of the Sun lab for critical reading and helpful discussion and P. Hixson for cell-culture support. This work was funded by NIH (P20NS123320), The Michael J. Fox Foundation, and Parkinson’s Foundation to Z.Y. This work was also funded by the American Lebanese Syrian Associated Charities (ALSAC) and NIH (R01NS129795) to J.S.
Author information
Authors and Affiliations
Contributions
Z.Y. conceived, designed, supervised the study and wrote the manuscript; J.S. conceived and supervised the structure research and wrote the manuscript; X-J.L. performed the cellular, molecular, and mouse experiments, analyzed the experimental data, and wrote the manuscript; H.Z. conducted the structure experiments and related data analysis, and wrote the manuscript; B.T.H performed the Co-IP experiments, data analysis and wrote the manuscript; H.K. sectioned the mouse brains for the IF experiments and analyzed the experimental data; X-T.L. participated in establishing the mouse models; H.T. and J.P. carried out the proteomics, phosphoproteomics and bioinformatics analysis and wrote the manuscript; I.C. carried out some of the culture experiments and analyzed the experimental data; Y.Z. performed some of the mouse genotyping experiments and analyzed the data. P.X. contributed financial support to X-J.L’s training and participated in the discussion in the manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stefan Knapp, John Salogiannis, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, X., Zhu, H., Huang, B.T. et al. RAB12-LRRK2 complex suppresses primary ciliogenesis and regulates centrosome homeostasis in astrocytes. Nat Commun 15, 8434 (2024). https://doi.org/10.1038/s41467-024-52723-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52723-6
This article is cited by
-
Epigenetic Insights into Vascular Cognitive Impairment: DNA Methylation in the Human Brain Tissue
Molecular Neurobiology (2025)
-
LRRK2 kinase-mediated accumulation of lysosome-associated phospho-Rabs in tauopathies and synucleinopathies
Acta Neuropathologica (2025)