Abstract
Carbapenem-resistant Acinetobacter baumannii (CRAB) is a persistent nosocomial pathogen that poses a significant threat to global public health, particularly in intensive care units (ICUs). Here we report a three-month longitudinal genomic surveillance study conducted in a Hangzhou ICU in 2021. This followed a three-month study conducted in the same ICU in 2019, and infection prevention and control (IPC) interventions targeting patients, staff and the ICU environment. Most A. baumannii isolated in this ICU in 2021 were CRAB (80.9%; 419/518) with higher-level resistance to carbapenems. This was accompanied by the proportion of global clone 2 (GC2) isolates falling from 99.5% in 2019 to 50.8% (213/419) in 2021. The phylogenetic diversity of GC2 increased, apparently driven by regular introductions of distinct clusters in association with patients. The remaining CRAB (40.2%; 206/419) were a highly clonal population of ST164. Isolates of ST164 carried blaNDM-1 and blaOXA-23 carbapenemase genes, and exhibited higher carbapenem MIC50/MIC90 values than GC2. Comparative analysis of publicly available genomes from 26 countries (five continents) revealed that ST164 has evolved towards carbapenem resistance on multiple independent occasions. Its success in this ICU and global capacity for acquiring resistance determinants indicate that ST164 CRAB is an emerging high-risk lineage of global concern.
Similar content being viewed by others
Introduction
Acinetobacter baumannii is a Gram-negative bacterial pathogen that causes ventilator-associated pneumonia and bloodstream infections in critically ill hospital patients globally1,2. The morbidity and mortality associated with A. baumannii infections are exacerbated by its extensive resistance to antibiotics. Carbapenem-resistant A. baumannii (CRAB) are often only sensitive to tigecycline and polymyxins3, and can persist in hospital environments for extended periods1. Resolving CRAB outbreaks can require modifications to practice and infrastructure that imposes significant clinical, logistical, and financial costs4. In 2024, the World Health Organisation (https://www.who.int/publications/i/item/9789240093461) reiterated its declaration of CRAB as a priority organism (Critical group) for which novel therapeutics are urgently required5.
Given the limited options for antimicrobial chemotherapy, infection prevention and control (IPC) strategies must play a substantial role in limiting the impact of CRAB in hospitals6. Effective genomic surveillance has the capacity to inform IPC approaches by defining outbreaks and identifying sources and pathways for bacterial transmission. Recent hospital studies have demonstrated the utility of whole-genome sequencing (WGS) for high-resolution characterisation of A. baumannii outbreaks or persistent populations6,7,8,9. The crucial need to develop a thorough understanding of the longer-term persistence, transmission and evolution of CRAB populations in nosocomial settings provided the motivation for undertaking this study.
In August–October 2019, we conducted a 13-week genomic surveillance study of CRAB in a 28-bed intensive care unit (ICU) located in Hangzhou, China10. We found that the ICU CRAB population was dominated by representatives of global clone 2 (GC2), which is one of two clones that account for the majority of CRAB globally3. Consistent with global reports, carbapenem resistance in this GC2 population was conferred by the blaOXA-23 carbapenemase gene, which was present in composite transposons Tn2006 or Tn2009 inserted at several different chromosomal sites10. Comparative analyses revealed that the GC2 population in this ICU was phylogenetically diverse, and the diversity appeared to have been shaped by independent introductions of multiple distinct GC2 clusters to the ICU. We detected putative CRAB transmission events between bed units and rooms within the ICU, and the spatiotemporal co-location of diverse CRAB clusters facilitated three horizontal gene transfer events over the course of the study10.
Informed by that first study, we planned to implement a set of IPC interventions and then perform a follow-up study to evaluate their effectiveness in reducing the prevalence of CRAB in the ICU. However, ICU operations were interrupted by the outbreak of COVID-19 in early 2020, and access to the ICU was restricted. Consequently, the implementation of planned CRAB-focused interventions was delayed until September 2020. Once introduced, the suite of interventions targeted ICU patients, the ICU environment (including equipment and sinks), and ICU staff (Supplementary Table 1). Full access to the ICU was restored in late November 2020, enabling us to conduct a second high-resolution genomic surveillance study, with a methodology identical to that of our pre-COVID-19 pandemic study, in May–July 2021.
Here we describe the follow-up longitudinal study of A. baumannii in this Hangzhou ICU. Over a 13-week period, we applied a deep-sampling approach to ICU patients and the ICU environment. All A. baumannii isolates were whole-genome sequenced. The structure and dynamics of this population were compared to those from our original study to evaluate the population’s longer-term development, and its response to IPC interventions. Moreover, a global phylogenetic and geographic distribution analysis of the high-risk emerging ST164 CRAB lineage was performed to lay a foundation for targeted infection prevention and control (IPC) strategies to control the spread of this clone in ICUs.
Results
A. baumannii infections were rare despite a high abundance in patients and bed units
A total of 131 patients (79 male; 52 female; median age 68 years; interquartile range [IQR] = 55.5–77.5) were sampled over the 3-month study period (Table 1). The median length of patient stay, from admission to discharge or the end of the study, was 10 days (IQR: 5–21.5 days). Most patients (53.4%) were in the ICU for ≤10 days, but 17.1% were present for >31 days. Moreover, the median length of ICU stay of A. baumannii-infected patients (clinical isolative positive) was 28.5 days (IQR: 24–42.3 days). The median length of other ICU patients was 9 days (IQR: 5–16 days). The length of stay (LoS) of A. baumannii-infected patients was significantly longer than other patients in the ICU. Samples were taken each Tuesday, and patients were screened within the first 48 h after admission when possible. “Bed units”, defined as the environmental sites of each bed and its associated equipment, were sampled on 363/364 (99.7%) planned sampling occasions, as one patient was undergoing bedside surgery on a single planned sampling occasion. Patients were sampled on 303/346 (87.6%) planned occasions.
A total of 5341 samples were collected from patients and environmental sites. A. baumannii was isolated from 505 samples: 382/4450 (8.6%) environmental samples and 123/891 (13.8%) patient samples. Bed units yielded more environmental isolates (374/4317, 8.7%) than communal areas outside bed units (8/133 samples, 6.0%). Within bed units, ventilators (56/281 samples, 19.9%) and ventilator shelves (40/275 samples, 14.6%) yielded the most A. baumannii. Of the 131 patients screened, 41.2% (54/131) were A. baumannii-positive on at least one sampling occasion, with most of the positive samples originating from nasogastric tube (33/174, 19.0%) or oral (40/220, 18.2%) swabs, followed by rectal swabs (38/273, 13.9%), nasojejunal tube (4/49, 8.2%), endotracheal tube (5/94, 5.3%) and tracheostomy tube (3/81, 3.7%). A set of 13 isolates obtained from diagnostic clinical specimens were collected from 10 patients over the study period; one from an abdominal excision exudate and 12 from sputum.
Most ICU A. baumannii were carbapenem-resistant
Over the study period, 19.1% (25/131) of ICU patients were treated with carbapenems imipenem, meropenem, or ertapenem. MIC testing of all 518 A. baumannii in this collection revealed that the rates of resistance to both imipenem and meropenem were 80.9% (419/518) (Fig. S4). A larger proportion of patient isolates (116/123; 94.3%) were CRAB than environmental isolates (303/382; 79.3%). However, the isolation rate of CRAB clinical strains of patients decreased from 8.6% (2019) to 7.6% (2021). The β-lactam/β-lactamase inhibitor combinations cefoperazone/sulbactam and piperacillin/tazobactam were administered to 14.5% (19/131) and 40.5% (53/131) of patients, respectively. The resistance rates to these combinations were 80.3% (416/518) for cefoperazone-sulbactam (1:1) and 81.1% (420/518) for cefoperazone-sulbactam (2:1). Resistance rates to sulbactam alone, ciprofloxacin, and amikacin were 80.9% (419/518), 80.9% (419/518), and 23.0% (119/518), respectively. All isolates were sensitive to colistin and tigecycline.
To quantify levels of carbapenem resistance in the CRAB populations from before and after the intervention period, we compared the imipenem MICs that would be sufficient to inhibit 50% (MIC50) and 90% (MIC90) of all isolates in each collection. For the 419 CRAB isolated in this study, the MIC50 and MIC90 to imipenem were 64 and 128 mg/L, respectively. The imipenem MIC50/MIC90 values for the 551 CRAB isolated in our first study were 32 and 64 mg/L, respectively. This data reflects an increase in carbapenem resistance levels in this ICU between 2019 and 2021. However, total sensitivity to colistin and tigecycline remained consistent across study periods.
Whole-genome sequencing revealed shifts in the circulating CRAB population
All (n = 518) A. baumannii isolates were whole-genome sequenced (Supplementary Data 1). Genome sequence data revealed that the population included representatives of 36 different sequence types according to the Pasteur MLST scheme (Fig. 1A). Most of the diversity was seen amongst the carbapenem-sensitive population, which included isolates from 34 different STs, 20 of which were represented by a single isolate each. Only three of these STs were represented by ten or more isolates: ST240, ST40, and ST221. The carbapenem-resistant population was comprised of just two sequence types: ST2/GC2 (213 isolates; 50.8%) and ST164 (206 isolates; 49.2%). The presence of ST164, accounting for approximately half of the CRAB isolates in this study, represents a major shift from 2019, where the CRAB population was dominated by GC2 (548/551; 99.5%) (Fig. 1A). ST164 isolates were obtained from clinical specimens in this ICU earlier in 2021, and this ST has recently been described as a new international clone (IC11), with reports of human infections in Europe, Asia, Africa and the Middle East11. Imipenem MIC50/MIC90 values for the 206 ST164 isolates were 128/128, and 32/64 mg/L for the 213 GC2 isolates, indicating that the appearance of ST164 was responsible for the overall increase in carbapenem resistance between studies.
A The upper panel shows the CRAB clone change model with their isolation rate in the ICU (Created in BioRender. Chen, Y. (2024) BioRender.com/q30i340). The below panel diaplays the comparison of the A. baumannii populations collected in this ICU over the initial (2019) and follow-up (2021) study periods. Horizontal bars are proportional to the number of isolates of given sequence types (STs), which are labelled and represented by different colours. Thinner horizontal lines and labels indicate whether isolates were carbapenem-resistant (CRAB; red) or carbapenem-sensitive (CSAB; blue). B Overview of the spatiotemporal distribution of GC2/ST2 (blue) and ST164 (orange) isolates over the course of the 2021 study. Bed unit and room numbers are indicated on the horizontal axis, and study week numbers are on the vertical axis. The sizes of the circles correspond to the number of isolates collected from each patient/bed unit at each sampling timepoint. Source data are provided as a Source Data file.
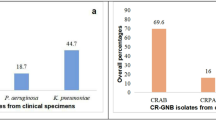
GC2 and ST164 were isolated throughout the study period and across the ICU (Fig. 1B). On 37/364 (10.2%) sampling occasions, GC2 and ST164 were isolated from the same bed unit. Despite their near-even prevalence in the ICU environment and patients, GC2 strikingly accounted for 12/13 (92.3%) clinical isolates obtained over the study period, with the remaining sputum isolates belonging to ST164. In contrast to GC2 and ST164, carbapenem-sensitive (CSAB) STs appeared sporadically over the course of the study and were generally localised to a single room, patient or time point (Fig. S5).
The GC2 population shifted significantly between studies through the introduction of multiple discrete sub-clusters
Consistent with our first study, carbapenem-resistance in GC2 CRAB isolates was conferred by the carbapenemase gene blaOXA-23. To determine the genomic contexts of these blaOXA-23 genes, we examined 19 hybrid-assembled complete GC2 genomes and screened the collection of 213 GC2 draft genomes with signature sequences indicative of the Tn2006 or Tn2009 insertions identified in our first study10. This revealed that the chromosomal positions encountered in the first study accounted for blaOXA-23 carriage in all GC2 isolates collected here (Fig. 2A; Supplementary Data 1). Additionally, one hybrid-assembled genome (DETAB-E2110) contained two copies of Tn2006, with the second copy found in the 13,545 bp plasmid pDETAB10/Tn2006 (Fig. S6). pDETAB10/Tn2006 was generated by the insertion of Tn2006 into the backbone of the 8731 bp R2-T1 type plasmid pDETAB10, which was present in 75/547 (13.7%) GC2 isolates in our first study and 175/213 (82.2%) GC2 isolates here. Screening this collection with the signatures of this insertion revealed that pDETAB10/Tn2006 was not present in any other isolates. DETAB-E2110 had MIC values of 128 and >128 mg/L for imipenem and meropenem, which were the equal-highest observed in this collection.
A Core-gene phylogeny (left) and cluster characteristics (right). Reference genomes for clusters obtained in the 2019 study are indicated by coloured circles at branch tips. Cluster designations from this study are indicated by coloured boxes to the right of the phylogeny, along with boxes shaded as outlined in the key to the left of the Figure. The phylogeny featured three large clusters were labelled as x, y and z. B Temporal distribution of GC2 clusters over the course of this study. For each study week, columns are divided according to the number of isolates of each GC2 cluster that were obtained, with colours corresponding to the key in part A. C Spatiotemporal distribution of transient GC2 clusters introduced by patients over the course of this study. The meaning of shapes is outlined in the key below, and colours correspond to those in the key in part A. Source data are provided as a Source Data file.
Although the GC2 isolates collected here shared Tn2006/Tn2009 insertions with the GC2 clusters examined in our previous study, chromosomal segments that contain transposons can be exchanged by homologous recombination between distinct A. baumannii strains12, so we sought phylogenetic evidence to assess cluster persistence in this ICU. To determine whether the GC2 population observed in this study was derived from that present in 2019, we constructed a phylogeny from the GC2 genomes obtained here, along with representative genomes for each of the 17 GC2 clusters identified in our first study (Fig. 2A). The phylogeny featured three large clusters (labelled x, y and z in Fig. 2A) that included 49, 72 and 92 GC2 genomes from this study. Each of the large clusters was separated from the others by long branches in the phylogeny. Representatives of 16 GC2 clusters from the 2019 study were confined to the diverse central cluster (x in Fig. 2A), with a single representative present in one of the two more clonal clusters, cluster 24 (C24) (y in Fig. 2A). Within the diverse central cluster, representatives from the 2019 study were largely distinct from isolates collected here, with the exception of C16, which differed from isolate DETAB-P494 by just one core-gene SNP (cgSNP). Overall, the GC2 population was more diverse in 2021 than it was in 2019, which is consistent with frequent and ongoing introductions of distinct GC2 clusters to the ICU. Further supporting this, some 2021 GC2 clusters (C18–23) appeared briefly in the ICU, and were present for no more than three sampling weeks (Fig. 2B). These will be referred to as transient GC2 clusters.
To detect putative patient-associated introductions to the ICU over the course of this study, we identified instances where, apart from in week 1, the first isolates of transient GC2 clusters appeared in patient samples. We found six such instances, involving six different clusters (Fig. 2C). Within transient clusters, isolates differed by 0–4 cgSNPs. The numbers of patient samples within each transient cluster (C18–C23) were 1, 5, 1, 7, 1 and 2, respectively (Fig. 2C). C22 was represented by a single patient isolate, but isolates of the other five clusters were found in two or more different environments each. These included the bed units occupied by the patients that initially carried the clusters, and either bed units within the same rooms (C18, C19, C21), bed units in different rooms (C23), or a communal cleaning cart (C20) (Fig. 2C). C19 and C21 were carried by two patients each, and the second patient that each cluster was isolated from most parsimoniously acquired C19/C21 from their contaminated room environments. All six transient GC2 clusters that first appeared in patient samples disappeared from the ICU following the departure of the last patient that carried them (Fig. 2C).
The largest GC2 clusters, C24 and C25, were present in most (10/13 for C24) or all (13/13 for C25) study weeks (Fig. 2B). The largest cluster, C25, only contained genomes from this study, all of which differed from one another by 0–6 cgSNPs (Supplementary Data 2). Isolates in C24 differed from one another by 0-11 cgSNPs, and from the 2019 study representative C2 genome by 5–10 cgSNPs (Supplementary Data 2). As representatives of C24 and C25 were already present in the first week of this study (Fig. 2B), we did not capture their introductions to the ICU, so cannot determine whether they were introduced in single or multiple events. However, the presence of sub-clusters of C24 and C25, containing isolates that do not differ at the cgSNP level (0 cgSNPs) but were found in multiple bed units or rooms, is evidence for the ongoing transmission of C24 and C25 within the ICU.
ST164: A dominant emerging clone recently introduced but fully established in the ICU
ST164 was not detected in this ICU in our first study. Over the course of this study, we obtained 206 ST164 isolates from 26 different sample types from patients and the ICU environment. The three most common positive sample types were oral (17/206, 8.3%), nasogastric tube (12/206, 5.8%) and rectal (12/206, 5.8%) swabs for patients, and bedside table (25/206, 12.1%), ventilator (22/206, 10.7%) and bed rail (16/206, 7.8%) swabs for the environment.
It was, therefore, important to establish whether this lineage had been introduced recently or had persisted in the ICU for an extended period. To assess their structural diversity, eight ST164 isolates were hybrid-sequenced to generate complete genomes. All eight complete genomes consisted of a 3.9 Mbp chromosome and five plasmids, pDETEC17–pDETEC21, that ranged from 12,790 to 2309 bp (Fig. 3A). Screening the entire collection of 206 ST164 short-read assemblies revealed that the five plasmids were present in almost all ST164 isolates (pDETAB19 195/206; pDETAB17 201/206; others 206/206).
A Schematic overview of the genome of ST164 isolates DETAB-P462. The chromosome is shown as a large circle with the locations of antibiotic resistance genes, MLST alleles, capsule (K) and outer core (OC) loci indicated. To the right, plasmids pDETAB17-21 and their sizes are shown (not to scale). B Time-dated phylogeny of the ST164 population examined in this study. Prominent branch dates are labelled, and the colours to the right of the phylogeny reflect ST164 clusters or the presence of antibiotic-resistance genes and plasmids as outlined in the key. SNP distances between the four ST164 clusters are indicated in circles filled in various colours. The blue arrow indicates the position of DETAB-R21, the reference isolated that was isolated in this ICU in early 2021. Source data are provided as a Source Data file.
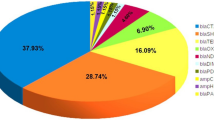
All ST164 genomes contained five acquired antibiotic resistance genes (ARGs). Four of these encoded beta-lactamases: blaNDM-1, blaOXA-23, blaCARB-16 and ampC (also called blaADC). In the complete ST164 genomes, all ARGs were located in the chromosome (Fig. 3A). The blaNDM-1 gene was in the complex transposon Tn692413 inserted downstream of glmS, along with the bleomycin resistance gene bleMBL and the aminoglycoside resistance gene aphA6. The remaining beta-lactamase genes were in four ISAba1 composite transposons: blaOXA-23 in two copies of Tn2006, ampC in Tn6168, and blaCARB-16 in a novel transposon named Tn7735 (Fig. 3A). Screening draft genomes with the junction sequences associated with all five ARG-containing insertions revealed that they were totally conserved across this ST164 population.
The near-total conservation of mobile genetic elements and chromosomal insertion sites suggested that the ST164 population in this ICU was highly clonal. To confirm this, we determined cgSNPs between all isolates in this collection and found that they ranged from 0 to 21. We did not find evidence for the introduction of ST164 clusters with patients over the course of this study, and we were aware that the same ST had been isolated from a sputum specimen collected in this ICU in January 202114, four months prior to this study. We therefore used the genome of that clinical isolate, DETAB-R21, as a reference for a time-dated phylogeny for the genomes generated here (Fig. 3B). The time-dated phylogeny was consistent with this ST164 population having diversified from an initial introduction to the ICU in mid-2020. The two deepest branches in this phylogeny had estimated dates of divergence from a common ancestor in April 2020, giving rise to clusters 164-A and 164-B (Fig. 3B). DETAB-R21 clustered with 164-B. Both 164-A and 164-B were present in the first and last weeks of this study, but were not present in every intervening week, with no 164-A isolates in week 11 and no 164-B in weeks 3 and 12 (Fig. 4). Clusters 164-C (present in weeks 5-13) and 164-D (weeks 2, 5-10), emerged from 164-A and 164-B, respectively (Fig. 3B), and were not present in the ICU at the outset of the study (Fig. 4). Notably, the first isolates of 164-C and 164-D were derived from environmental samples rather than patient samples, consistent with their emergence from a pre-existing ICU population rather than from patient-associated introductions over the course of this study.
A Temporal distribution of ST164 clusters over the course of this study. For each study week, columns are divided according to the number of isolates of each ST164 cluster that were obtained, with colours corresponding to the key. Spatiotemporal distribution of ST164 clusters isolated from patients (B) and the ICU environment (C). Colours correspond to the key in part A, and the sizes of bubbles to the numbers of isolates indicated by the key in part B. Source data are provided as a Source Data file.
Over the course of the study, we observed 10 clear instances where patients were admitted to the ICU, were A. baumannii-negative at one or more sampling points, but then yielded ST164 from subsequent oral, rectal or tube swabs. All of these patient-derived ST164 isolates were identical (0 cgSNPs; Supplementary Data 3) to one or more isolates that had previously been obtained from the ICU environment or from other ICU patients. We conclude that these events represent acquisition of ST164 from the population persisting in the ICU. Acquisition events involved three of the four ST164 phylogenetic clusters: ST164-A in 3/10 cases, ST164-B in 4/10, and ST164-C in 3/10. In one further case, a patient who produced GC2 from their first screening samples appeared to acquire ST164-D, which was isolated from a subsequent oral swab and a clinical sputum sample.
Persistence of CRAB in the ICU results in clinically relevant CRAB acquisition by patients
Using the phylogenetic evidence described above, we sought to determine the derivation of each of the 13 CRAB clinical isolates collected over the course of this study and, therefore, to identify the routes through which they might have been acquired by these 10 patients over the study period. Most (12/13) of the clinical isolates were GC2, and amongst these, 10 were C25, one was C24, and one was C21. The remaining clinical isolate was ST164-D. Representatives of all four of these clusters were present in multiple patient and environmental samples prior to their appearance in clinical samples, but to avoid any ambiguity associated with small numbers of cgSNPs (≤4), we focused on comparing identical (0 cgSNP) isolates when describing clinically-relevant CRAB acquisition in the ICU.
The GC2 C21 cluster appears to have been introduced to the ICU by patient 200 (P200) (see above), who was admitted to the ICU from either the community or a different hospital (data not available). The presence of C21 in a P200 clinical sample is, therefore, indicative of acquisition outside this hospital, before the development of clinically relevant symptoms requiring sputum sample collection in this ICU. The GC2 C24 isolates obtained from P172 were identical to isolates obtained from P172’s first screening samples on arrival to the ICU, which might suggest that they carried C24 on arrival. However, because C24 was circulating in the ICU (see above), and identical isolates had been obtained from other patients and the ICU environment prior to the admission of P172, it is also possible that P172 acquired C24 in the ICU before their first screening samples were taken.
In the remaining eight cases, patients had been CRAB-negative at one or more sampling points prior to the development of symptoms that necessitated clinical sample collection. In three of these cases, isolates identical to the clinical isolates (0 cgSNPs) had previously been isolated from the ICU environment. It, therefore, seems most parsimonious that these patients acquired CRAB in the ICU. The five remaining clinical isolates were not identical to any other isolates in the collection but differed from multiple isolates by ≤4 cgSNPs. We conclude that acquisition in the ICU was responsible for the majority (9 or 10/10) of CRAB found in clinically relevant patient samples over the course of this study.
Global epidemiology and carbapenemase gene distribution in ST164
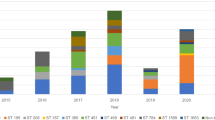
To explore the global distribution of ST164 isolates, we chose four representative strains from our study to represent the total diversity observed in this ICU and further incorporated 131 ST164 publicly-available genomes that were collected in 26 countries from five continents between 2017 and 2023 (Supplementary Data 4). Overall, ST164 strains are primarily distributed in Asian countries (Fig. 5C). Most publicly-available ST164 genomes (95.6%; 129/135) were isolated from patients, and the highest numbers of isolates were collected between 2016 and 2019 (Fig. 5A and B). The most common KL type in global ST164 isolates was KL47 (63.4%, 83/131), which differs from the GC2 isolates where KL30 (43.2%, 92/213) and KL93 (33.8%, 72/213) were most common. ST164 genomes could be further divided into ST234 (20%, 27/135), ST1418 (54.8%, 74/135) and novel types (25.2%, 34/135) using the Oxford MLST scheme (Fig. 5A). ST164 genomes differed by between 3 and 6,037 cgSNPs (Supplementary Data 5), and could be divided into seven main clades via evolutionary branches (Fig. 5A). All isolates assigned to Clade 1 were from the USA and carried the blaNDM-1 cabapenemase gene. Isolates from Clade 6 exhibited the widest geographical distribution, with representatives isolated on five different continents. The largest cluster, Clade 7, is dominated by strains isolated from Thailand. Notably, strains collected in China formed a monophyletic cluster, Clade 2. Importantly, Clade 2 strains harboured the highest number of carbapenemase genes, which were carried in multiple transposons, including Tn6924, Tn7735, and up to three copies of Tn2006. Isolates of other clades (clade 1, clade 3–7) carry at most one carbapenem resistance gene. Moreover, all strains from clade 1 carry blaNDM-1, and the majority of clade 3–7 isolates only harboured blaOXA-23. Co-carriage of blaNDM-1 and blaOXA-23 has so far only been seen in China and Malaysia.
A A tree of 131 global ST164 strains and 4 representative isolates collected from our ICU was built and further visualised using iTOL v5. The information of ST_Oxford, location, continent, host and cabapenemases distribution was shown. Clades 1–7 were labelled as different colours. The reference strain DETAB-R21 was labelled as a blue name. Four representative strains from this study were indicated with red names. B Temporal distribution of strain numbers and carbapenemase genes. C Geographical distribution of major ST164 strains in the world and characteristics of carbapenemase carried by the strains. OXA-23, OXA-420, OXA-23+OXA-58, OXA-23+NDM-1, NDM-1 and none were shown as various colours. The black circle with three arrows means that ST164 CRAB may be widely spread in Asia. The map file was downloaded from ChiPlot v1 (https://www.chiplot.online/). Source data are provided as a Source Data file.
Discussion
This follow-up observational study has allowed us to assess the influence of bundled IPC interventions on an endemic ICU CRAB population, in a region where CRAB prevalence is highest globally15. The nature of bundled intervention strategies complicates assessments of the impacts of individual IPC measures, and this was further complicated here by the time that passed between studies, as well as by the changes to ICU access and practice associated with responses to COVID-19. However, the unique timing of our studies on either side of the emergence of COVID-19 also afforded us an opportunity to determine the joint impacts of the intervention strategies as well as the COVID-19 pandemic on antimicrobial resistance in A. baumannii.
The early and middle periods of the COVID-19 pandemic were associated with increased rates of antibiotic prescribing in China16, which may have augmented resistance levels in nosocomial bacterial populations. Increased prescriptions of ß-lactam antibiotics (including carbapenems) and tigecycline, which have been reported across multiple studies16, are particularly concerning in the context of A. baumannii. The phenotype-agnostic approach to sampling in this study allowed us to determine that the majority (80.8%) of A. baumannii in this ICU were CRAB. We observed an increase in the magnitude of carbapenem resistance in this ICU, with MIC50/MIC90 values doubling between our 2019 and 2021 studies. The increase in carbapenem resistance levels was driven by the emergence of ST164, which co-produced NDM-1 and OXA-23 carbapenemases. Despite the increase in carbapenem resistance, all isolates obtained in 2021 remained sensitive to colistin and tigecycline, though monitoring CRAB populations for resistance to these last-resort antibiotics should remain a high priority.
Following IPC interventions, we observed a decrease in the numbers of CRAB isolated from patients and from the ICU environment and saw a reduction in both the total number of clinical isolates (from 19 in 2019 to 13 in 2021) and the number of patients from which clinical isolates were obtained: 12/140 patients (8.6%) in 2019, and 10/131 patients (7.6%) in 2021. However, IPC practices remained imperfect, with multiple examples of GC2 CRAB clusters being imported to the ICU before spreading to different bed units or rooms and being acquired by patients, highlighting the significant challenges associated with prevention of CRAB transmission in hospital wards once it has colonised the hospital environment. More importantly, A. baumannii could persist in the oral cavity based on the positive samples of oral swabs in ICU patients. Our previous research showed A. baumannii DETAB-R21 was isolated from an oral swab in the ICU14, which was considered as one colonisation isolate. Moreover, a study by A M Richards et al. showed subgingival colonisation by A. baumannii could increase the risk of refractory periodontitis17. Apart from oral colonisation, a study by Patrick M. Ketter et al. revealed the gastrointestinal (GI) tract has been found to be a major reservoir for A. baumannii, as well as to potentially contribute to the development of multidrug resistance18. Our study also showed that nasogastric tubes have a high CRAB isolation rate. These persistent carriage strains in patients require further attention in the ICU. We used a highly-stringent 0-cgSNP cut-off for assessing putative transmission events over the course of this study. This allowed us to unambiguously implicate environmental persistence in the development of hospital-acquired infections. New data presented here adds to our previous findings that suggested GC2 clusters were being imported to the ICU regularly from external sources. We hypothesise that other wards within this hospital, as well as patients transferred from other hospitals, serve as a source for incoming CRAB strains. If this is the case, hospital-wide IPC measures or highly stringent screening and isolation of CRAB-positive patients prior to ICU admission will be required to ultimately prevent CRAB spread in the ICU. To further prevent its dissemination in this ICU, the patients can be admitted directly to the isolation ward and cared for by only one nurse if the screening result is positive before the patients prior to the ICU admission.
A striking finding here was the reduction in the prevalence of GC2 amongst CRAB isolates, falling from 99.5% in 2019 to 50.8% in 2021. The remainder of the 2021 CRAB population was comprised of a highly clonal set of ST164 isolates that appear to have been diversifying in the ICU since mid-2020. This study has, therefore, added further context to the recent recognition of the emergence of ST164 as an international A. baumannii clone11, with our data specifically highlighting this clone’s capacity to establish itself and persist in an ICU environment, even in direct competition with clusters of the highly successful GC2. Based on the global view of diversity amongst ST164 genomes, we speculate that this clone has spread widely in Asia, particularly in China and Thailand, and has acquired carbapenemase genes on multiple independent occasions. Strikingly, it appears that the convergence of blaOXA-23 and blaNDM-1 has occurred twice in ST164, highlighting the potential this clone has for accumulating carbapenem resistance determinants. All ST164 strains isolated in Hangzhou carry blaNDM-1 and blaOXA-23 simultaneously, suggesting that this particular sub-clone has been highly successful in our hospital settings. While ST164 caused fewer infections than GC2 in this ICU over the 2021 study period, its presence in one clinical specimen, and in clinical specimens collected previously in this ICU and elsewhere11,14, along with its high carbapenem resistance levels, suggest that it is a lineage that urgently requires surveillance and further characterisation. In addition, the most common KL type was KL47 in the 131 global ST164 isolates. One recent study from the United Kingdom showed the majority of the KL47 isolates belonged to GC2 and ST164 clone1. KL47 type was also identified in ST16, ST215 and ST374 isolates (Pasteur MLST scheme). These different ST types, which carry KL47, would potentially reveal the capsular loci interaction of their common ancestors.
The increase in CRAB diversity between 2019 and 2021, including the appearance of ST164 and the diversification of GC2, was likely a product of regular imports to the ICU over an approximately 20-month period. However, it would be interesting to determine whether IPC interventions have introduced new selective pressures and contributed to population turbulence. If so, these environmental conditions might be selected for fitter lineages that can better establish persistence in hospital environments. Under such conditions, IPC measures might have differential effects at the strain level, and effective responses will require informed, dynamic IPC strategies. Experimental characterisation of diverse CRAB lineages will be needed to assess their variable ability to persist in hospital environments and ultimately infect patients. The GC2 cluster C25, which was persistent in this ICU and prominent amongst clinical isolates over the course of this study (10/13; 76.9%), is an intriguing target for future laboratory studies that might address these questions.
A limitation of this study is that the IPC interventions did not occur immediately after the first study in 2019 due to the COVID-19 pandemic. COVID-19 may have played a role in CRAB population changes in the ICU, considering the surge in cases, increased antibiotic use, and associated IPC measures. Hence, it is difficult to accurately measure the impact of IPC interventions and COVID-19 on the ICU CRAB population. We expect that the CRAB population was influenced by the combined effects of both our IPC interventions and COVID-19-related changes to clinical practice.
In conclusion, we report on the emergence of a new CRAB clone, ST164, in this Hangzhou ICU. Our data show that ST164 was as successful as the globally dominant clone GC2 in this ICU setting, but exhibited higher carbapenem resistance levels due to dual production of NDM-1 and OXA-23 carbapenemases. Through comparative analyses, we found that ST164 has been isolated in 26 countries from five continents, and has evolved towards carbapenem resistance on multiple independent occasions. We conclude that ST164 is an emerging high-risk lineage which requires urgent global surveillance, and our findings will lay a foundation for the formulation of more reasonable and effective targeted IPC strategies for ICU staffs, patients and the environment.
Methods
Consent and research ethics
Ethical approval and informed consent were obtained by the Sir Run Run Shaw Hospital (SRRSH) Zhejiang University local ethics committee (approval number 20190802-1). This work was part of a study registered as a clinical trial with ClinicalTrials.gov (NCT04310722).
IPC interventions implemented before this study
A bundled set of interventions (from September 2020 to July 2021) that aimed to address the abundance of CRAB in this ICU was implemented 8 months prior to the outset of this study. The interventions included changes to daily practice in the ICU, alongside raising staff awareness around the issue. The interventions are listed in Supplementary Table 1. Briefly, patient care was modified by introducing “water-free” cleaning methods and by installing disinfection devices for liquid waste. ICU sinks ceased operation. Environmental cleaning was performed more regularly, with particular attention paid to end-of-stay cleaning procedures and communal surfaces such as keyboards used routinely by ICU staff. Education for staff was enhanced, and enforcement of hand hygiene practices was targeted through compliance monitoring and the introduction of a compliance management system.
Study design and sample collection
This longitudinal study was conducted in a 28-bed ICU in SRRSH, Hangzhou, China, from May to July 2021. A map of the ICU is presented in Fig. S1. Unless sampling was perceived to be medically detrimental, ICU patient sampling was planned for the beginning of the study or on admission to the ward and weekly thereafter. Patient samples were routinely collected from oral and rectal swabs, and from nasogastric, nasojejunal, endotracheal, or tracheostomy tube swabs when present. Clinical samples were obtained from patients as part of normal medical practice (the specific process of microbiological diagnostics was shown in Fig. S2), and all A. baumannii obtained from clinical samples over the study period were retained. Equipment, sinks and other surfaces within patient rooms were swabbed weekly. Equipment, surfaces, and sinks in communal areas outside patient rooms were sampled monthly. The overall flow chart is shown in Fig. S3.
Sample processing and species identification
Environmental surfaces (approximately 10 × 10 cm) were swabbed with sterile swabs (COPAN, Italy) pre-moistened with 2 mL TSB supplemented with 0.1% sodium thiosulfate (Sangon Biotech, China). The swabs were immediately placed into 14-mL sterile tubes containing 2 mL TSB with 0.1% sodium thiosulfate and then incubated for 24 h at 37 °C. After culture, 20 µL overnight bacteria were streaked onto Acinetobacter CHROMagar plates (CHROMagar, Paris, France), followed by overnight incubation at 37 °C.
For patient samples, rectal swabs containing Cary-Blair Transport Medium (Gongdong, Taizhou, China) were used and other swabs were moistened using 0.9% saline before sampling. These swabs were plated directly onto Acinetobacter CHROMagar plates and incubated at 37 °C for 24 h. Based on red colour and morphology, a single, isolated colony of presumptive A. baumannii was selected and streaked onto a Mueller–Hinton (MH) agar plate (Oxoid, Hampshire, UK), then incubated again at 37 °C for 24 h. A single, isolated colony from the MH plate was selected and stored for downstream analyses.
Species identification was confirmed using Matrix-assisted laser desorption ionisation-time of flight mass spectrometry (MALDI-TOF MS) (bioMérieux, France) and 16S rRNA gene sequencing.
Antimicrobial susceptibility profiling
Minimum inhibitory concentrations (MICs) of meropenem, imipenem, polymixin B, tigecycline, cefoperazone/sulbactam (1:1), cefoperazone/sulbactam (2:1), sulbactam, amikacin and ciprofloxacin were measured using the broth microdilution method of plate for “Susceptibility testing of non-fastidious Gram Negative isolates”. Results were interpreted according to the Clinical and Laboratory Standards Institute 2021 guidelines and European Committee on Antimicrobial Susceptibility Testing (EUCAST) 2023 breakpoint tables (http://www.eucast.org/clinical_breakpoints/) for polymyxin B (S: ≤2 mg/L, R: >2 mg/L). Tigecycline results were interpreted based on Food and Drug Administration (FDA) guidelines. Escherichia coli ATCC 25922 was used as the quality control strain.
Illumina sequencing
Collected isolates were all subjected to Illumina sequencing (Hangzhou Digital-Micro Biotech Co., Ltd., China). Briefly, DNA was isolated using the Qiagen Mini Kit (Qiagen, Germany). Libraries were prepared using the TruePrepTM DNA Library Prep Kit V2 (Vazyme) according to the manufacturer’s protocol. Paired-end sequencing (2×150-bp reads) was performed on the Illumina HiSeq X Ten platform (Illumina Inc., San Diego, CA, USA).
Oxford nanopore sequencing
Long-read sequencing (Hangzhou Digital-Micro Biotech Co., Ltd., China) was performed for 60 isolates chosen after Illumina sequencing on the basis of phylogenetic, resistance gene and plasmid diversity. Genomic DNA was extracted using the Gentra® Puregene® Yeast/Bact. Kit (Qiagen, Germany) according to the manufacturer’s protocol with minor modifications. In brief, libraries were prepared using the SQU-LSK109 Ligation Sequencing Kit (Oxford Nanopore Technologies, UK) in conjunction with the PCR-Free ONT EXP-NBD104 Native Barcode Expansion Kit (Oxford Nanopore Technologies, UK) according to the native barcoding genomic DNA protocol. DNA was processed without the optional shearing steps to select for long reads. After quantification of the individual libraries by Qubit and normalisation of library concentrations, the library was sequenced on the GridION X5 platform (Oxford Nanopore Technologies, UK).
Bioinformatic analyses
Sequence reads were trimmed, assembled using Unicycler v0.4.8 and assessed for quality. MLST was used to determine multi-locus sequence types with the Pasteur and Oxford typing schemes19,20. Typing of capsular polysaccharide (KL) was conducted with Kaptive 2.0 and Bautype 1.021. AMRFinder v1 was used to identify antimicrobial resistance genes22. Plasmid replicons were typed using the Hamidian lab typing scheme23.
Snippy v4.4.5 (https://github.com/tseemann/snippy) was used to align Illumina reads against a corresponding reference hybrid assembly (A. baumannii DETAB-R21, GenBank accession GCA_024205285.1) and generate a core genome alignment (3,709,099 bp). Polymorphic sites (120 polymorphic sites) were extracted with Gubbins v2.4.0, excluding those that were predicted to occur via recombination24. Phylogenies were constructed from these polymorphic sites using IQtree v2.0.3. Divergence dating was undertaken with the least-squares method implemented by IQTree v2.0.3, using the previously generated tree, Gubbins fasta file, and a GTR + G model25,26. SNP-distances were calculated from the Gubbins-filtered polymorphic sites file using SNP-dists 0.6.3 (https://github.com/tseemann/snp-dists).
Global analyses for ST164
To construct a global phylogeny for the ST164 CRAB clone, we downloaded 24,084 A. baumannii genome sequences from the National Center for Biotechnology Information (NCBI) database (search conducted on 11.12.2023). All A. baumannii genome sequences were analysed by mlst v2.19.0 (https://github.com/tseemann/mlst) and ABRicate v1.0.1 with the Resfinder 4.0 database27, aiming to identify sequence types (STs) and antibiotic resistance genes. Quality control was performed using FastANI v1.33 and QUAST v5.0.228. Assemblies with a genome size greater than five million base pairs, an N50 smaller than 15,000 or a genome average nucleotide identification (ANI) of <95% were excluded from further analysis. Following this, 135 genomes (four representative strains from this study and 131 global genomes, SNPs difference: 3–6037) were used to generate a full-length whole-genome alignment with the complete genome of A. baumannii DETAB-R21 (GenBank accession GCA_024205285.1) as the reference using the methods described above. The resulting tree was annotated in iTOL v529. The global distribution of ST164 CRAB carrying different carbapenemase genes was visualised using GraphPad Prism 8.0, ChiPlot v1 (https://www.chiplot.online/) and Adobe Illustrator CC 2023.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequence data and genome assemblies generated in this study have been submitted to GenBank under the BioProject accession PRJNA1034534. The individual Illumina sequence read accession numbers of all 518 A. baumannii isolates are listed in Supplementary Data 6. The raw data of patients’ clinical data and MICs of all 518 A. baumannii isolates are provided in the Source Data excel. Source data are provided with this paper.
References
Bergogne-Berezin, E. & Towner, K. J. Acinetobacter spp. as nosocomial pathogens: microbiological, clinical, and epidemiological features. Clin. Microbiol. Rev. 9, 148–165 (1996).
Muller, C. et al. A global view on carbapenem-resistant Acinetobacter baumannii. mBio 14, e0226023 (2023).
Hamidian, M. & Nigro, S. J. Emergence, molecular mechanisms and global spread of carbapenem-resistant Acinetobacter baumannii. Microb. Genom. 5, e000306 (2019).
Dijkshoorn, L., Nemec, A. & Seifert, H. An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat. Rev. Microbiol. 5, 939–951 (2007).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Roberts, L. W. et al. Genomic surveillance, characterization and intervention of a polymicrobial multidrug-resistant outbreak in critical care. Microb. Genom. 7, mgen000530 (2021).
Lewis, T. et al. High-throughput whole-genome sequencing to dissect the epidemiology of Acinetobacter baumannii isolates from a hospital outbreak. J. Hosp. Infect. 75, 37–41 (2010).
Shenoy, E. S. et al. Community-acquired in name only: a cluster of carbapenem-resistant Acinetobacter baumannii in a burn intensive care unit and beyond. Infect. Control Hosp. Epidemiol. 41, 531–538 (2020).
Schultz, M. B. et al. Repeated local emergence of carbapenem-resistant Acinetobacter baumannii in a single hospital ward. Microb. Genom. 2, e000050 (2016).
Doughty, E. L. et al. Endemicity and diversification of carbapenem-resistant Acinetobacter baumannii in an intensive care unit. Lancet Reg. Health West Pac. 37, 100780 (2023).
Hansen, F. et al. Characterisation of carbapenemase-producing Acinetobacter baumannii isolates from danish patients 2014-2021: detection of a new international clone— IC11. Int. J. Antimicrob. Agents 62, 106866 (2023).
Hua, X. et al. Acquisition of a genomic resistance island (AbGRI5) from global clone 2 through homologous recombination in a clinical Acinetobacter baumannii isolate. J. Antimicrob. Chemother. 76, 65–69 (2021).
Mann, R., Rafei, R., Gunawan, C., Harmer, C. J. & Hamidian, M. Variants of Tn6924, a novel Tn7 family transposon carrying the bla(NDM) metallo-beta-lactamase and 14 copies of the aphA6 amikacin resistance genes found in Acinetobacter baumannii. Microbiol. Spectr. 10, e0174521 (2022).
Zhao, F. et al. Description of a rare pyomelanin-producing carbapenem-resistant Acinetobacter baumannii strain coharboring chromosomal OXA-23 and NDM-1. Microbiol. Spectr. 10, e0214422 (2022).
Oldenkamp, R., Schultsz, C., Mancini, E. & Cappuccio, A. Filling the gaps in the global prevalence map of clinical antimicrobial resistance. Proc. Natl Acad. Sci. USA 118, e2013515118 (2021).
Cong, W. et al. Prevalence of antibiotic prescribing in COVID-19 patients in China and other low- and middle-income countries during the pandemic (December 2019–March 2021): a systematic review and meta-analysis. J. Antimicrob. Chemother. 78, 2787–2794 (2023).
Richards, A. M., Abu Kwaik, Y. & Lamont, R. J. Code blue: Acinetobacter baumannii, a nosocomial pathogen with a role in the oral cavity. Mol. Oral Microbiol. 30, 2–15 (2015).
Ketter, P. M. et al. Acinetobacter baumannii gastrointestinal colonization is facilitated by secretory IgA which is reductively dissociated by bacterial thioredoxin A. mBio 9, e01298–18 (2018).
Diancourt, L., Passet, V., Nemec, A., Dijkshoorn, L. & Brisse, S. The population structure of Acinetobacter baumannii: expanding multiresistant clones from an ancestral susceptible genetic pool. PLoS ONE 5, e10034 (2010).
Bartual, S. G. et al. Development of a multilocus sequence typing scheme for characterization of clinical isolates of Acinetobacter baumannii. J. Clin. Microbiol. 43, 4382–4390 (2005).
Wyres, K. L., Cahill, S. M., Holt, K. E., Hall, R. M. & Kenyon, J. J. Identification of Acinetobacter baumannii loci for capsular polysaccharide (KL) and lipooligosaccharide outer core (OCL) synthesis in genome assemblies using curated reference databases compatible with Kaptive. Microb. Genom. 6, e000339 (2020).
Feldgarden, M. et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype–phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, e00483–19 (2019).
Lam, M. M. C., Koong, J., Holt, K. E., Hall, R. M. & Hamidian, M. Detection and typing of plasmids in Acinetobacter baumannii using rep genes encoding replication initiation proteins. Microbiol. Spectr. 11, e0247822 (2023).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
To, T. H., Jung, M., Lycett, S. & Gascuel, O. Fast dating using least-squares criteria and algorithms. Syst. Biol. 65, 82–97 (2016).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Mikheenko, A., Prjibelski, A., Saveliev, V., Antipov, D. & Gurevich, A. Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 34, i142–i150 (2018).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Acknowledgements
This work was undertaken as part of the DETECTIVE research project funded by the Medical Research Council (MR/S013660/1 (A.M.)), the National Natural Science Foundation of China (U22A20338 (Y.Y.), 82402677 (H.L.), 32270183 (X.H.), 81861138054 (Y.Y.), 32011530116 (X.H.)), Zhejiang Provincial Natural Science Foundation of China under Grant No. LQ24H200005 (H.L.) and a Wolfson Research Merit Award (WM160092 (W.v.S.)). We thank the doctors and nurses in the ICU for performing patient sampling. We appreciate the use of servers provided by MRC CLIMB BIG DATA grant MR/T030062/1.
Author information
Authors and Affiliations
Contributions
H.L.: funding acquisition, sample collection, sample processing, data curation, formal analysis, visualisation and writing—original draft. R.A.M.: data curation, formal analysis, visualisation, writing—original draft, writing—review and editing. E.L.D.: data curation, formal analysis, visualisation, writing—original draft. X.H.: supervision, formal analysis, project administration. A.E.S.: data curation, formal analysis. L.Z., X.C., and F.G.: sample collection, review and editing. W.v.S.: funding acquisition, conceptualisation, writing—review and editing. A.M.: funding acquisition, conceptualisation, formal analysis, writing—review and editing. Y.Y.: resources, funding acquisition, conceptualisation, writing—review and editing. All the authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, H., Moran, R.A., Doughty, E.L. et al. Longitudinal genomics reveals carbapenem-resistant Acinetobacter baumannii population changes with emergence of highly resistant ST164 clone. Nat Commun 15, 9483 (2024). https://doi.org/10.1038/s41467-024-53817-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53817-x