Abstract
Diazotrophic cyanobacteria can fix nitrogen gas (N2) but are usually scarce in nitrogen-limited coastal waters, which poses an apparent ecological paradox. One hypothesis is that high salinities (> 10 g/L NaCl) may inhibit cyanobacterial N2 fixation. However, here we show that N2 fixation in a unicellular coastal cyanobacterium exclusively depends on sodium ions and is inhibited at low NaCl concentrations (< 4 g/L). In the absence of Na+, cells of Cyanothece sp. ATCC 51142 (recently reclassified as Crocosphaera subtropica) upregulate the expression of nifHDK genes and synthesise a higher amount of nitrogenase, but do not fix N2 and do not grow. We find that the loss of N2-fixing ability in the absence of Na+ is due to insufficient ATP supply. Additional experiments suggest that N2 fixation in this organism is driven by sodium energetics and mixed-acid fermentation, rather than proton energetics and aerobic respiration, even though cells were cultured aerobically. Further work is needed to clarify the underlying mechanisms and whether our findings are relevant to other coastal cyanobacteria.
Similar content being viewed by others
Introduction
Organisms require combined nitrogen (N) for essential processes such as nucleic acid and protein synthesis. Despite the abundance of atmospheric nitrogen gas (N2), bioavailable N is often scarce in nature1. In aquatic ecosystems, cyanobacteria can partially meet the need for N by fixing N2, which impacts food chains, greenhouse gas sequestration, global climate change, and biogeochemical cycles1,2. This ability provides diazotrophic cyanobacteria with a competitive advantage over non-diazotrophic cyanobacteria and eukaryotic phytoplankton when environmental N is low. It enables them to exploit and dominate N-scarce environments worldwide, and can even lead to the development of dense cyanobacterial blooms when other nutrients are abundant3. Interestingly, in contrast to the frequently reported blooms of cyanobacterial N2-fixers (note that non-N2-fixers also bloom) in freshwater ecosystems, few blooms have been reported in N-limited saline coastal waters, despite their clear ecological advantage4,5. The usual scarcity and low abundance of coastal N2-fixing cyanobacteria in an N-deficient environment is striking and represents an ecological paradox.
One commonly proposed explanation is that high salinity levels (> 10 g/L NaCl), a fundamental difference between coastal water and freshwater ecosystems, severely inhibit N2 fixation and population development of resident N2-fixers4,5,6. However, most of these studies refer to multicellular N2-fixing cyanobacteria and therefore much remains unknown about the NaCl tolerance of coastal unicellular N2-fixers, which have only recently been recognised for their significant contribution to the oceanic N cycle1,7.
Here, we investigate the effect of NaCl on N2 fixation using Cyanothece sp. ATCC 51142 (hereafter Cyanothece sp., recently reclassified as Crocosphaera subtropica8), a coastal N2-fixing unicellular cyanobacterium9. First, we examine whether high salinity suppresses unicellular cyanobacterial N2 fixation, as is commonly proposed for multicellular cyanobacteria4. We then integrate physiological, transcriptomic, and enzymatic analyses to study the underlying mechanisms.
Contrary to the accepted view that high salinity negatively affects coastal N2 fixation4,5, here we demonstrate not only that N2 fixation of Cyanothece sp. is NaCl-dependent, but rather that Na+ energetics is essential to provide ATP for N2 fixation. Overall, we offer an alternative perspective on the paradox of the scarcity of N2-fixing cyanobacteria in N-limited coastal waters and provide insights into the role of Na+ energetics in cyanobacterial metabolism, specifically in N2 fixation.
Results
N2 fixation depends on the presence of NaCl
We first carried out growth experiments of Cyanothece sp. cultured in artificial seawater medium (ASP2, 18 g/L NaCl) and freshwater medium (BG11) in the presence or absence of N. Under N-rich conditions, cells from ASP2 and BG11 showed similar population growth, as expected for a coastal isolate. Under N deficiency, population growth was only observed in ASP2 without N (hereafter ASP2-N), whereas cells barely grew in BG11 without N (hereafter BG110) (Fig. 1a). Since the most prominent discrepancy between ASP2-N and BG110 is the presence of NaCl, we recultured these non-growing cells in fresh BG110 amended with 18 g/L NaCl (hereafter BG110 (NaCl)), and NaCl reactivated population growth (Fig. 1b). Here N2 fixation is the prerequisite for cell growth and division, it is therefore confirmed that N2 fixation depends on NaCl. Notably, although a slight population growth of cells grown in BG110 was observed in the first few days after inoculation, this was probably due to the consumption of the intracellular nitrogen storage product cyanophycin and the degradation of some proteins (such as light-harvesting complex phycobilisomes)10 rather than N2 fixation, which would otherwise have resulted in continuous population growth.
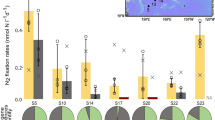
a Growth curves of cells grown in four different media over a period of 9 days. b Growth curves of 7-day-old non-growing cells in BG110 with the addition of NaCl over a period of 7 days. c Cell number increase of cells grown in BG11 with NaCl gradients. d Cell number increase of cells grown in BG110 with NaCl gradients. e N2 fixation activity of cells grown in BG110 with NaCl gradients. Nitrogenase activity is expressed as ethylene (C2H4) production by acetylene reduction. Data are presented as mean ± standard deviation (n = 3 biologically independent samples). Different letters on each bar (c, d, e) indicate statistical significance (p < 0.05) calculated by one-way ANOVA with Tukey’s HSD post-hoc analysis over all populations tested. In figure d, line segments and corresponding significance markers indicate statistical significance calculated by one-way ANOVA across four salinity treatments (2, 4, 9, 18 g/L). Significance: ns (no significance), *(p < 0.05), **(p < 0.01), ***(p < 0.001), ****(p < 0.0001). Source data are provided as a Source Data file.
The effect of NaCl gradients on N2 fixation and growth was investigated in more detail. When N was replete, except for the three highest NaCl concentrations (32, 34, 36 g/L) (ANOVA, F3,8 = 22.04, p < 0.001), cells in the other treatments did not differ in growth from cells grown at 18 g/L, which is equivalent to the salinity of the marine medium ASP2 (Fig. 1c). In contrast, when N was depleted, NaCl was required for N2 fixation and population growth. Cell proliferation was almost completely inhibited when NaCl was less than 4 g/L and maximised at 18 g/L NaCl (Fig. 1d, ANOVA, F3,8 = 137.1, p < 0.001). The requirement for NaCl on N2 fixation was further verified by the direct quantification of N2 fixation activities. In accordance with the growth pattern (Fig. 1d), N2 fixation activities were almost completely inhibited at low NaCl concentrations (0, 2 g/L) and maximised at 18 g/L NaCl (approximately 75 nmol ethylene produced per 108 cells per hour) (Fig. 1e, ANOVA, F3,8 = 318.3, p < 0.001). In addition, we found that NaCl is a general requirement for N2 fixation irrespective of cellular growth stage, as cells from the exponential phase (Fig. 1a), the stationary phase (Supplementary Fig. 1), and non-N2-fixing growth-arrested cells (Fig. 1b) all required NaCl to fix N2 under diazotrophy.
Cellular N starvation due to lack of N2 fixation
In the absence of NaCl, non-N2-fixing growth-arrested exhibited a yellow colouration (Supplementary Fig. 2a), a typical symptom of nitrogen-starvation-induced chlorosis10. To test whether these non-growing cells in BG110 are physiologically N-starved due to the non-functional N2 fixation, nitrogenous compounds and dry weight of cells cultured in either BG110 or BG110 (NaCl) were quantified. Consistent with our expectation, compared to normally growing cells (functioning N2 fixation) with NaCl, NaCl deprivation led to a significant decrease in chlorophyll (C55H72O5N4Mg) (t test, p < 0.05), total protein content (t test, p < 0.01), dry weight (t test, p < 0.01), and total protein ratio (of dry weight) (t test, p < 0.05) of non-N2-fixing cells grown in BG110 (Supplementary Figs. 2b-e), indicating that these cells were N-starved due to the lack of N2 fixation.
Transcriptomic profiles in the presence and absence of NaCl
Transcriptomic analysis was performed to reveal the molecular background and transcriptomic features of the NaCl requirement for N2 fixation. Compared to N2-fixing cells cultured with NaCl (18 g/L), 5226 (1909 genes upregulated, 3317 genes downregulated) genes of non-N2-fixing cells grown without NaCl were identified as differentially expressed genes (DEGs) (adjusted P value < 0.05) (Supplementary Fig. 3a). According to KEGG enrichment analysis, all DEGs could be classified into several functional categories (Supplementary Fig. 3b), showing striking metabolic differences. Detailed heatmap visualisation of DEGs enriched in KEGG ribosome biosynthesis (map03010, Fig. 2a), photosynthesis (map00195, Fig. 2b), and porphyrin metabolism (participating in chlorophyll synthesis) (map00860, Supplementary Fig. 3c) indicated that almost all DEGs enriched in non-N2-fixing cells showed a downregulated pattern at the transcript level compared to N2-fixing cells. Given the importance of ribosomes and photosynthesis for cell growth and division, these distinct gene expression patterns provide clues at the molecular level as to why diazotrophic cells cultured without NaCl fail to grow or divide.
a Heatmap analysis showing the DEGs enriched for ribosome biosynthesis (map03010) and (b) for photosynthesis (map00195). Two components of the ribosomal proteins (small subunit, large subunit) and six photosynthetic subunits (photosystem II (PSII), cytochrome, photosystem I (PSI), ATP synthase, allophycocyanin, phycocyanin) were analysed. KEGG annotations were assigned from the genome annotation. Column dendrograms indicate similarity based on Euclidean distance and hierarchical clustering. Gene clusters were determined by k-means clustering with Euclidean distance. The heatmap colour gradient shows low gene expression (blue) and high gene expression (red). c Expression trend of nitrogenase structural genes (nifHDK). Data (TPM, transcripts per million) are presented as box plots (lower bound at 25th percentile, centre line at the median, upper bound at 75th percentile) with whiskers at minimum and maximum values. Shaded-in plots indicate significantly different expression levels of genes and arrows indicate the directionality of statistically significant trends between these two treatments (p < 0.05, right-tailed Fisher’s exact t test followed by Benjamin-Hochberg (BH) adjustment). Statistical significance of selected genes was calculated from pairwise TPM comparisons of triplicate samples. “NaCl” denotes the addition of extra NaCl (18 g/L) to BG110, while “None” indicates that no additional substances were added to BG110. Source data are provided as a Source Data file.
It is reasonable to expect that non-N2-fixing (N-starved) cells would reduce their primary metabolism to a minimal level in order to survive. Indeed, these cells exhibited the stringent response11 (a persistent state in which cells reduce protein synthesis and stop dividing), as evidenced by non-N2-fixing cells in the absence of Na+: 1) Cells were nitrogen-starved (Supplementary Fig. 2) and growth-arrested (Figs. 1a, d), which represents the most typical cause and consequence of the stringent response11,12; 2) They exhibited typical stringent response transcriptomic patterns, which involved the downregulation of ribosome biosynthesis genes (Fig. 2a) and the upregulation of nutrient transporter genes12 (Supplementary Fig. 4a, 32 out of 45 annotated genes); 3) They upregulated the expression of the relA gene (Supplementary Fig. 4b, gene location: NC_010546.1 (21413−22405), NCBI), encoding ppGpp synthase/hydrolase, a key enzyme that regulates levels of ppGpp (the signalling molecule for the stringent response)11. Intriguingly, although Cyanothece sp. is capable of N2 fixation, the morphological, physiological and transcriptomic performance of the non-N2-fixing growth-arrested cells in the absence of Na+ also align with the nitrogen chlorosis theory, which typically describes the processes involved in the acclimation of non-diazotrophic cyanobacteria to N shortage10. It is important to note, however, that the stringent response (or nitrogen chlorosis) described above is the consequence (resulting from cellular N starvation) rather than the cause of N2 fixation dysfunction in the absence of NaCl. The reason for the N2-fixers’ N-starvation in the absence of NaCl has yet to be determined.
With regard to the nitrogenase (the key enzyme responsible for N2 fixation) structural genes (nifHDK)13, we found a higher level of expression of nifHDK in the non-N2-fixing cells, i.e., cultured without NaCl, than in the N2-fixing cells, cultured with NaCl (Fig. 2c). This expression pattern suggests that non-N2-fixing growth-arrested cells in the absence of Na+ were N-starved, which is in accordance with the result that these cells were physiologically N-starved (Supplementary Fig. 2).
N2 fixation depends exclusively on Na+
To determine whether the need for NaCl is specific, we tested whether four other metal chlorides, including KCl, LiCl, MgCl2 and CaCl2, at doses equivalent to NaCl (310 mM, following the medium recipe for ASP29), could stimulate the growth of cells grown in BG110 in the same way as NaCl. As N2 fixation is a must for population growth under N deprivation, population growth is used as a proxy for N2 fixation activity in the following contexts, unless otherwise specified. We found that population growth was exclusively observed in the NaCl treatment (Fig. 3a, ANOVA, F4,10 = 65.74, p < 0.001). Given that high concentrations of LiCl are toxic to cells and the observed growth arrest of the LiCl (310 mM) treatment could have been caused by its toxicity, we further tested whether LiCl at low concentrations can activate population growth. The results show that LiCl at low concentrations cannot stimulate population growth either (Supplementary Fig. 5), indicating that LiCl cannot function in the same manner as NaCl to activate N2 fixation. Overall, these results demonstrate that the requirement for NaCl for N2 fixation is exclusive and cannot be substituted by other metal chlorides. Moreover, since all the tested metal chlorides contain a chloride moiety, this result also indicates that Na+ and not Cl- plays a role in N2 fixation. Notably, BG110 medium alone contains 0.082 g/L Na+ from its constitutive ingredients (Na2CO3, Na2EDTA·2H2O and Na2MoO4·2H2O), nevertheless, this Na+ concentration is far below the effective concentration (4 g/L) for activating N2 fixation as shown earlier (Figs. 1d, e).
a Increase in cell number of cells grown in BG110 with the addition of five different metal chlorides: NaCl, KCl, LiCl, MgCl2 and CaCl2. b Schematic representation of the pathway of unicellular cyanobacterial N2 fixation. Created in BioRender. Tang, S. (2023) BioRender.com/e77x697. N2 is first reduced to NH4+ by nitrogenase (Step 1) and then is incorporated into carbon skeletons via the glutamine (Gln) synthetase-glutamate (Glu) synthase pathway, where further N-containing organic metabolites are formed (Step 2). NifHDK, nitrogenase complex; PEP carboxylase, phosphoenolpyruvate carboxylase; 2-OG, 2-oxoglutarate; GS, glutamine synthetase; GOGAT, glutamate synthase. c Increase in cell number of cells grown in BG110 amended with NaCl, NH4Cl and Gln. d Nitrogenase content of cells grown in BG110, BG110 (NaCl) and BG110 (NaNO3). e Cell number increase of cells grown in BG110 and BG110 (NaCl) with and without N2 sparging. f Nitrogenase activity of cells grown in BG110, BG110 (NaCl) and BG110 (NaNO3). One unit (U) is the amount of enzyme that catalyses the reaction of 1 µmol of substrate per minute. The graphs show the mean ± standard deviation, n = 3 biologically independent samples for a, c and e, n = 6 biologically independent samples for (d and f). Different letters on each bar represent statistical significance (p < 0.05) calculated by one-way ANOVA with Tukey’s HSD post-hoc analysis over all populations tested. Source data are provided as a Source Data file.
Active nitrogenase despite the lack of Na+
Unicellular cyanobacterial N2 fixation can generally be divided into two steps, i.e., N2 fixation (from gaseous N2 to ammonium) and N assimilation (from ammonium to organic nitrogenous compounds)14 (Fig. 3b). We have shown that N2 fixation is severely inhibited (Fig. 1e), but it remains to be determined whether N assimilation is also suppressed under NaCl deprivation. Therefore, we performed a nutrient addition experiment in which the product of N2 fixation (ammonium (NH4Cl)) and an intermediate of N assimilation (glutamine (Gln)) was added to cells grown in BG110. Compared to cells grown in BG110, the addition of NH4Cl (2 mM) and Gln (2 mM) mimicked the Na+ effect and resulted in normal growth (Fig. 3c, ANOVA, F3,8 = 58.83, p < 0.001), confirming that N2 fixation only was inhibited by Na+ deprivation.
We then tested: 1)whether Na+ deficiency inhibits the biosynthesis of nitrogenase; 2) whether Na+ plays a role in protecting the enzyme from oxygen (O2) inactivation, as nitrogenase is known to be extremely sensitive to O215; 3) whether Na+ participates in the conformational activation of nitrogenase. Although almost no N2 fixation activity was detected in cells grown in BG110 (Fig. 1e), these cells expressed a significantly higher amount of nitrogenase (Fig. 3d, ANOVA, F2,15 = 337.6, p < 0.001), which is consistent with our previous transcriptomic analysis showing that non-N2-fixing cells expressed higher levels of nifHDK (Fig. 2c). The results of the sparging experiment show that even under anaerobic conditions, Na+ was still required for N2 fixation and population growth (Fig. 3e, ANOVA, F3,8 = 40.52, p < 0.001), suggesting that the dependence of nitrogenase activity on Na+ is not due to a role of the cation in providing oxygen protection to the enzyme. Nitrogenase from non-N2-fixing cells was tested to be conformationally active with a significantly higher catalytic rate compared to that of N2-fixing cells (Fig. 3f, ANOVA, F2,15 = 289.2, p < 0.001), which makes sense since the catalytic capacity of an enzyme is generally proportional to its amount (Fig. 3d). Taken together, these results indicate that: 1) Na+ is not involved in the biosynthesis, protection against O2 inactivation or conformational activation of nitrogenase; 2) non-N2-fixing cells grown without Na+ synthesised an unusually high amount of nitrogenase but could not fix N2.
A shortage of ATP prevents N2 fixation
Due to the incompatibility of oxygen-producing photosynthesis and O2-sensitive N2 fixation, N2 fixation of Cyanothece sp. occurs in the dark16,17. It is well established that N2-fixing cells synthesise and store large quantities of carbohydrate in the form of glycogen via photosynthesis in the light, which is used to generate adenosine triphosphate (ATP) via glycolysis to power N2 fixation in the dark18. Given that biological N2 fixation, including both N2 fixation itself (the theoretical minimum energy requirement) and the associated apparatuses supporting N2 fixation, is highly energy-intensive16, we reason that a limitation of ATP caused by Na+ deficiency might lead to N2 fixation dysfunction.
It has been reported that Na+ plays a role in bicarbonate uptake, which is subsequently catalysed to release CO2 for photosynthesis19,20. Therefore, Na+ deprivation may adversely affect photosynthesis and glycogen biosynthesis, resulting in glycogen shortage (the energy substance for N2 fixation) and insufficient ATP supply for N2 fixation. Accordingly, we determined whether Na+ was involved in glycogen synthesis and indirectly inhibited N2 fixation. To this end, we quantified and contrasted the intracellular glycogen levels of cells grown without N, either in the presence or absence of Na+, under ambient CO2 levels without forced aeration, in a diurnal cycle (a light-dark cycle of 12 h/12 h). As expected, the cellular glycogen content of N2-fixing cells grown in BG110 (NaCl) exhibited a diurnal pattern, with glycogen synthesis occurring during the light period and consumption to power N2 fixation in the dark period. It is noteworthy that, in comparison to N2-fixing cells, non-N2-fixing growth-arrested cells in BG110 exhibited elevated levels of glycogen, yet were unable to utilise it in the dark (Fig. 4a, ANOVA, F3,8 = 37.36, p < 0.001). Indeed, the accumulation of glycogen in cyanobacteria under N starvation represents a classic physiological response observed during nitrogen chlorosis10. This result, coupled with the fact that cells can grow normally in BG11 (Fig. 1a), suggests that Na+ is not involved in bicarbonate uptake and glycogen synthesis and that the energy source (glycogen) shortage is not the cause of N2 fixation failure in the absence of Na+.
a Cellular glycogen contents of non-N2-fixing cells grown in BG110 or N2-fixing cells grown in BG110 with 18 g/L NaCl (BG110 (NaCl)) in a diurnal cycle. Cells were sampled at the end of the light phase (L12) or at the end of the dark phase (D12). b Increase in cell number of cells grown in BG110 or supplemented with either NaCl or ATP. c Nitrogenase content of cells grown in BG110 (NaCl), BG110 (Gln), BG110 (ATP) and BG110 (NaNO3). d Increase in cell number of cells grown in BG110 supplemented with equivalent doses of ATP, ADP or AMP. The graphs show the mean ± standard deviation, n = 3 biologically independent samples for a, b and d, n = 6 biologically independent samples for (c). Different letters on each bar represent statistical significance (p < 0.05) calculated by ANOVA with Tukey’s HSD post-hoc analysis across all populations tested. Source data are provided as a Source Data file.
To further test the hypothesis that an ATP shortage in the absence of Na+ leads to N2 fixation dysfunction, an additional supply of ATP was provided to cells grown in BG110. The results show that additional ATP (0.1 mM) stimulated population growth (Fig. 4a, ANOVA, F2,6 = 49.95, p < 0.001), suggesting that: 1) The observed failure of the nitrogenase to function may be attributed to the lack of ATP; 2) Under diazotrophic conditions, Na+ may play a role in ATP biosynthesis. However, the presence of the nitrogen element in ATP (C10H16N5O13P3) promoted us to reconsider the role of ATP in activating N2 fixation, either by providing chemical energy or by providing N. To distinguish between these two possibilities, we first examined the nitrogenase content of cells grown in BG110 supplemented with additional ATP. Almost no nitrogenase was observed in cells supplemented with N (Gln (0.1 mM) and NaNO3 (0.1 mM)), whereas cells supplemented with ATP (0.1 mM) or Na+ (310 mM) synthesised the two highest levels of nitrogenase (Fig. 4b, ANOVA, F3,20 = 2269, p < 0.001). It can be concluded that in this case ATP does not act as a N supplier, which would otherwise not lead to the biosynthesis of nitrogenase. Furthermore, we also assessed population growth in BG110 supplemented with an equivalent dose (0.1 mM) of ATP, ADP (adenosine diphosphate, C10H15N5O10P2), or AMP (adenosine monophosphate, C10H14N5O7P). As shown in Fig. 4c, cells supplemented with ATP had the highest growth among all treatments (ANOVA, F3,8 = 52.06, p < 0.001), indicating that ATP serves as additional chemical energy rather than as an N source. Otherwise, this would have resulted in comparable growth patterns across all treatments, as all supplements provided an equivalent dose of N. From an energetic point of view, this result makes sense, as ATP possesses a higher energy content than ADP and AMP, thereby enabling the nitrogenase to fix more N2 for growth. Notably, although not statistically significant, there was a slight population growth in the AMP treatment when compared to the no additive control. This is probably due to the scavenging of the very low energy that AMP contains. Although it has been reported that many bacterial species are capable of releasing and actively depleting extracellular ATP21, further investigation is required to ascertain how Cyanothece sp. cells utilise extracellular ATP. Possibilities include that ATP may be transported into cells via water-filled porins (at least into the periplasm)22,23 or via transporters/carriers24 or other uptake systems, either independently or in combination, and then utilised by cells. In conclusion, the results collectively indicate that insufficient bioavailable ATP (energy) impairs N2 fixation in the absence of Na+. Nevertheless, the mechanism by which Na+ contributes to the energy metabolism of N2-fixing cells remains elusive.
Na+ energetics enables N2 fixation
Regarding energy metabolism in bacteria, it is well established that ATP synthesis is driven by the energy stored in a transmembrane electrochemical gradient of protons coupled to the light-driven or the respiration-driven proton pumps (H+ energetics)25. Nevertheless, there is evidence that Na+ can replace H+ as the coupling ion to drive ATP synthesis in several anaerobic bacteria25,26, as well as in cyanobacteria, as recently demonstrated20. This type of machinery for synthesising ATP by Na+-coupling (Na+ energetics) fits well with our findings that Na+ correlates with the supply of ATP for N2 fixation. Also, there is a striking resemblance between N2-fixing Cyanothece sp. and anaerobic bacteria with Na+ energetics, as both require non-oxic microenvironments. Therefore, we propose that a mechanism for Na+-coupled ATP synthesis exists in N2-fixing Cyanothece sp. To test this proposal, we first quantified the cellular ATP content and the ATP/ADP ratio of non-N2-fixing cells grown in BG110 (for seven days) following the addition of NaCl (18 g/L) during N2 fixation in the dark. The addition of NaCl resulted in a significantly higher amount of ATP (Fig. 5a, ANOVA, F3,16 = 21.28, p < 0.001) and higher ATP/ADP ratio (Fig. 5b, ANOVA, F3,16 = 37.02, p < 0.001) relative to the no NaCl addition control, indicating that Na+ plays a critical role in driving ATP synthesis. To further corroborate the involvement of Na+ in ATP synthesis during N2 fixation, dark-phase N2-fixing cells grown with NaCl were treated with monensin20,27 (a Na+-specific ionophore) or ethyl-isopropyl amiloride20 (EIPA, an inhibitor of Na+ channels and Na+/H+ antiporters), respectively. The ionophore monensin renders membranes permeable to Na+ and thus destroys transmembrane Na+ gradients, whereas EIPA suppresses transmembrane Na+ transportation. If Na+ fluxes were coupled to ATP synthesis, the collapse of the Na+ transmembrane gradient caused by monensin or EIPA should result in a reduction in cellular ATP levels and ATP/ADP ratios due to the inhibition of ATP synthesis. As shown in Figs. 5c, d, the addition of monensin (14 µM) resulted in significantly lower ATP levels (ANOVA, F3,16 = 64.21, p < 0.001) and ATP/ADP ratios (ANOVA, F3,16 = 31.87, p < 0.001) in comparison to the untreated control. Similarly, treatment with EIPA (100 µM) markedly suppressed ATP synthesis (Fig. 5e, ANOVA, F3,16 = 45.72, p < 0.001), leading to lower ATP/ADP ratios (Fig. 5f, ANOVA, F3,16 = 40.3, p < 0.001). These results substantiate the pivotal role of Na+ energetics in N2 fixation.
Cellular ATP content (a) and ATP/ADP ratio (b) of N-starved cells grown in BG110 after the addition of NaCl in the dark. Prior to NaCl addition, cells were grown in BG110 for 7 days. Cellular ATP content (c) and ATP/ADP ratio (d) of dark phase N2-fixing cells grown in BG110 with 18 g/L NaCl after the addition of monensin (14 µM). Cellular ATP content (e) and ATP/ADP ratio (f) of dark phase N2-fixing cells grown in BG110 with 18 g/L NaCl after the addition of EIPA (100 µM). The graphs show the mean ± standard deviation (n = 3 biologically independent samples). Different letters on each bar represent statistical significance (p < 0.05) calculated by ANOVA with Tukey’s HSD post-hoc analysis across all populations tested. Source data are provided as a Source Data file.
To further understand the role of Na+ energetics in energy metabolism, we investigated cell growth after adding either monensin, a protonophore 3,5-di-tert-butyl4-hydroxybenzaldehyde (DTHB)28, EIPA, or a typical F-type ATP synthase inhibitor dicyclohexylcarbodiimide (DCCD)29 to the respective culture medium. Similar to monensin that destroys Na+ energetics, the protonophore DTHB makes membranes permeable to protons, abolishing proton gradients and thus inhibiting H+ energetics and ATP generation. Compared to normally growing cells from the control, 14 µM monensin completely inhibited the growth of cells in BG110 (NaCl) (Fig. 6a, ANOVA, F3,16 = 135.2, p < 0.001). Conversely, cells cultured in BG11 with monensin divided normally, indicating that monensin is not toxic to cells. Furthermore, cells were cultured in BG110 supplemented with ATP in the presence or absence of monensin, to examine whether monensin inhibits the performance of nitrogenase. The result shows that the addition of monensin does not impede the growth of ATP-supplemented cells, indicating that monensin does not disrupt the nitrogenase (Supplementary Fig. 6). With regard to DTHB exposure, only the cells that had been grown in BG11 were significantly inhibited by DTHB (200 µM) (Fig. 6b, ANOVA, F3,16 = 169.83, p < 0.001). No growth was observed in any of the treatments that were exposed to EIPA (100 µM) (Fig. 6c, ANOVA, F3,16 = 99.11, p < 0.001) or DCCD (100 µM) (Fig. 6d, ANOVA, F3,16 = 249.3, p < 0.001). These results suggest: 1) The presence of a Na+-coupled ATP synthesis machinery. This was demonstrated by non-toxicity of monensin to cells and nitrogenase, as well as by the abolition of transmembrane Na+ gradients, which resulted in the inhibition of ATP synthesis and the subsequent growth arrest of cells in BG110 (NaCl); 2) The existence of an H+-coupled ATP synthesis machinery was evidenced by the normal growth of cells in BG11, which was unaffected by monensin, but was severely impaired by DTHB. Furthermore, we found that cells in BG110 (NaCl) with DTHB grew normally, suggesting that H+ energetics is not involved in N2 fixation; 3) The inhibition of the Na+/H+ antiporter and suppression of other essential cellular functions through EIPA, as evidenced by the complete suppression of cell growth in all treatments. Indeed, EIPA has been reported to severely inhibit cyanobacterial PSII system20. This suggests that lower cellular ATP contents and ATP/ADP ratios observed following EIPA treatment (Fig. 5e, f) may be attributed not only to the inhibition of Na+-dependent ATP synthesis but also to the suppression of other key metabolic processes; and 4) The utilization of the same type of F0F1 ATP synthase by Na+ and H+ energetics, as DCCD suppressed the growth of cells in all treatments. Interestingly, we found that cells in BG11 (NaCl) with monensin were unable to grow. This suggests that Na+ availability may play a critical role in determining the coupling ion for ATP synthase. In this case, Na+-coupled ATP synthesis would be favoured by the presence of Na+, otherwise, the cells in BG11 (NaCl) with monensin should have grown due to H+ energetics. Overall, these results indicate that Na+ energetics plays an essential role in the activation of N2 fixation. Nevertheless, future experimental efforts are needed to provide more direct evidence to determine whether and to what extent Na+-dependent ATP synthesis takes place.
Increase in number of cells grown in the medium supplemented with monensin (a), DTHB (b), EIPA (c) or DCCD (d) as compared to non-supplemented cells. The graphs show the mean ± standard deviation (n = 3 biologically independent samples). Different letters on each bar represent statistical significance (p < 0.05) calculated by ANOVA with Tukey’s HSD post-hoc analysis across all populations tested. Source data are provided as a Source Data file.
Potential Na+ gradient generator
Similar to the H+ cycle, a Na+ cycle is required for continuous energy supply25,26. In this case, Na+ pumps (Na+ gradient generators) must work together with ATP synthase (Na+ gradient consumer) to re-energise the membrane in N2-fixing cells. Since it has been reported that the Na+/H+ antiporter acts as a Na+ pump to generate a transmembrane Na+ gradient20, we examined the Na+/H+ antiporter of the respective cells. Using transcriptomic analysis, we found that in the absence of Na+, non-N2-fixing cells expressed significantly higher levels of four genes encoding subunits of the Na+/H+ antiporter, namely mnhB (K05566, KEGG), mnhD (K05568, KEGG), mnhE (K05569, KEGG), and mnhG (K05571, KEGG), than N2-fixing cells (Supplementary Fig. 7), implying the key role of the Na+/H+ antiporter in generating the Na+ gradient. Here, these cells were unable to establish a Na+ gradient to drive ATP synthesis to meet their energy needs. Moreover, the result that N2-fixing cells were extremely sensitive to EIPA (an inhibitor of the Na+/H+ antiporter, Fig. 6c), provides additional evidence that the Na+/H+ antiporter is involved in the regeneration of Na+ gradients.
In summary, based on all the evidence provided in this study, we propose that Cyanothece sp. performs typical H+ energetics across the thylakoid membrane during the day (Fig. 7a). In contrast, during the night under N2-fixing conditions, we propose that the primary Na+ gradient consumer (Na+-coupled ATP synthase) works together with active Na+ gradient generators (Na+/H+ antiporter) across the plasma membrane to drive ATP synthesis for N2 fixation (Fig. 7b).
a Schematic representation of the typical H+ energetics across the thylakoid membrane coupled to photosynthesis in cyanobacteria (including Cyanothece sp.). Created in BioRender. Tang, S. (2023) BioRender.com/n32k956. b Schematic representation of the proposed Na+ energetics across the plasma membrane with Na+ gradient generator and consumer in N2-fixing Cyanothece sp. cells. Created in BioRender. Tang, S. (2023) BioRender.com/n32k956. Abbreviations: RTO, respiratory terminal oxidase; Cyt b6f, cytochrome b6f; PC/Cyt c6, plastocyanin/cytochrome c6; PQ, plastoquinone; Fd, flavodiiron; NDH, NADH-dehydrogenase; Fum, fumarate; Suc, succinate; SDH, succinate dehydrogenase; Glu, glutamic acid; Gln, glutamine; Ser, Serine; Val, valine; Tyr, tyrosine; Ala, alanine; ATP, Adenosine triphosphate; ADP, Adenosine diphosphate; Pie, inorganic phosphate; Δψ, membrane potential.
Anaerobic fermentation possibly powers N2 fixation
To understand why Cyanothece sp. cells capable of H+ energetics must perform Na+ energetics under N2-fixing conditions, it is important to note that N2 fixation in this species occurs in the dark and under anaerobic conditions in the cell16. Moreover, although H+ energetics are generally believed to be more energetically advantageous than Na+ energetics, H+ energetics in our case is no better than Na+ energetics due to the lack of a high potential electron acceptor, i.e., O226. Therefore, we propose that anaerobic energy metabolism coupled with Na+ energetics powers N2 fixation in the case of Cyanothece sp.
Cellular O2 dynamics during N2 fixation of Cyanothece sp. under aerobic conditions show that although there is a sharp decrease in the O2 level of the culture medium suggesting active aerobic respiration, this only occurs at the beginning of the dark phase30,31. In contrast, it has been repeatedly reported that the O2 level remains almost unchanged, albeit with some perturbations (meaning that almost no O2 is consumed by the cells), while at the same time active N2 fixation has been detected during the remaining hours (indicating functioning energy metabolisms)30,31. This suggests that anaerobic energy metabolism, rather than aerobic respiration, is functioning to power N2 fixation. Indeed, it has been shown that diazotrophic cyanobacteria (filamentous Oscillatoria limosa strain 23 and unicellular Synechococcus OS-A and OS-B’) rely on anaerobic fermentation to power N2 fixation under dark and anaerobic conditions32,33. Notably, Cyanothece sp. contains all the fermentation-related genes necessary to produce ethanol, lactate, and acetate in its genome17, indicating the capacity for anaerobic fermentation. Taken together, this evidence suggests that it is highly likely that anaerobic fermentation plays a crucial role in powering N2 fixation in Cyanothece sp.
To investigate whether fermentation powers N2 fixation in Cyanothece sp., we first examined the transcriptomic level of key fermentation-related genes. Although our transcriptomic analysis was designed to discriminate non-N2-fixing growth-arrested cells from normally growing N2-fixing cells at the transcript level, we identified transcriptomic patterns of fermentation-related genes suggesting that fermentation is involved in N2 fixation. Specifically, we found that three key genes involved in microbial fermentation34, namely ack encoding acetate kinase (K00925, KEGG), adh encoding alcohol dehydrogenase (K00001, KEGG) and ldh encoding lactate dehydrogenase (K00016, KEGG), were upregulated in non-N2-fixing cells compared to N2-fixing cells (Supplementary Fig. 8a). This suggests that non-N2-fixing cells may be challenged to meet their energy requirements in the absence of Na+ and therefore express higher levels of key fermentative energy metabolism genes. The presence of fermentative energy production in N2-fixing cells is further supported by the significantly higher activities of all three key fermentative enzymes in N2-fixing cells than in N-grown cells (Supplementary Fig. 8b). Furthermore, based on the activities of three different fermentative enzymes, it is reasonable to conclude the existence of a mixed acid type fermentation, a typical fermentation mode in bacteria34,35. In conclusion, in N2-fixing Cyanothece sp. cells under dark and anaerobic conditions, it is highly probable that aerobic respiration plays a role only in the initial period of the dark phase, probably to power the synthesis of nitrogenase and partial N2 fixation and to create an anoxic interior, while mixed-acid fermentation, which requires further validation, coupled with Na+ energetics, functions to power N2 fixation in the rest of the time.
Discussion
Contrary to the traditional view that coastal N2 fixation is sensitive to high levels of NaCl4,5, this study reports that N2 fixation by a coastal unicellular N2-fixer is exclusively dependent on the presence of Na+. In the absence of Na+, the cells failed to fix N2, were N-starved and showed a stringent response (or chlorosis), despite producing substantially higher levels of active nitrogenase. Further experiments show that the presence of Na+ is a prerequisite for the activation of Na+ energetics for the generation of ATP, which is subsequently used for the energy-intensive N2 fixation. Thus, this study demonstrates the exclusive and essential role of Na+ energetics in powering N2 fixation in unicellular cyanobacteria.
Conventionally, H+ energetics are thought to be chemically more advantageous than Na+ energetics26. So why should Cyanothece sp. cells capable of H+ energetics (Fig. 1a and Fig. 6b) perform Na+ energetics? In Cyanothece sp., as in many unicellular N2-fixing cyanobacteria and some multicellular species, N2 fixation occurs in the dark16. The necessary ATP is believed to be produced by H+ energetics, coupled with the aerobic respiratory breakdown of endogenous glycogen granules, which not only energizes this process but also consumes intracellular O2, creating an anoxic interior (so-called respiratory protection)15,36. Although an anoxic environment is required for nitrogenase to function, this process is very energy-intensive. Therefore, if this energy were produced solely by aerobic respiration of the glycogen granules, the cells would have to increase their respiration and hence their respiratory O2 uptake. Simultaneously, the O2 level in the cell should be kept low enough to avoid inactivation of the nitrogenase. Consequently, the demand for more O2 for increased aerobic respiratory activity is in conflict with the demand for an anaerobic interior. These two processes, which should work in tandem, seem to be incompatible. Therefore, it is reasonable to assume that under N2 fixation conditions (dark and anoxic), H+ energetics (coupled to aerobic respiration of glycogen) is not favoured and instead Na+ energetics (coupled to anaerobic fermentation of glycogen), which can function independently of aerobic respiration25, appears to be an alternative. Indeed, by integrating published data17,30,31,32,33 and our results (Supplementary Fig. 8), it is reasonable to infer that although the cells were grown aerobically, mixed-acid fermentation rather than aerobic respiration mainly powers N2 fixation, whereas active aerobic respiration only functions at the beginning of the dark phase.
Furthermore, under dark and anaerobic conditions, Na+ energetics is not inferior to H+ energetics from an energetic point of view. Only when there is a direct mechanistic link between redox reactions (oxidation of external resources) or photosynthesis in photosynthetic microbes and the translocation of H+ across the membrane is H+ energetics chemically more advantageous than Na+ energetics26. Thus, an energetic advantage only exists when high potential electron acceptors, such as O2, are available. This is probably the reason why H+ is mainly used as a coupling ion in the aerobic microbial world. In Cyanothece sp. under conditions where N is abundant, O2 does not interfere with cellular metabolism during the day or night, allowing a more efficient use of H+ energetics. However, when N is depleted, an obligate anaerobic environment is required for oxygen-sensitive N2 fixation. In this case, Na+ energetics coupled to fermentation are engaged, and organic intermediates of the glycolysis37, such as pyruvate, serve as electron acceptors.
From an energetic perspective, the advantage of using both H+ and Na+ energetics in Cyanothece sp. can be rationalised by considering the energy challenges faced by cells in their complex natural habitats. To meet the energy demands of dynamic coastal seas, in particular the energy-intensive anaerobic N2 fixation, organisms require a versatile energy metabolism, i.e., H+ energetics coupled to photosynthesis and aerobic respiration during the day, and Na+ energetics for anaerobic N2 fixation in the dark. This metabolic flexibility likely contributes to the evolutionary resilience of cyanobacteria in a wide range of environments.
In the study of Na+ energetics in cyanobacterial N2 fixation, although three species of filamentous N2-fixing cyanobacteria were reported to be dependent on NaCl for N2 fixation about forty years ago38, it was then concluded that inhibition of combined phosphorus (P) uptake in the absence of Na+ was responsible for the loss of nitrogenase activity. However, this cannot explain the Na+ dependence of N2 fixation in Cyanothece sp. as we show here that P uptake is independent of the presence of Na+ (Fig. 1a, Fig. 3c). Recently, although not directly related to N2 fixation, a study using Synechocystis sp. PCC 6803 reported that N-starved cells engage in Na+ energetics for ATP synthesis to maintain viability and to awaken from dormancy20,39. These reports suggest that Na+ energetics may be more widespread in cyanobacteria than commonly thought. Indeed, Na+ energetics is considered to be the most ancient form of bioenergetics, and genomic analyses suggest that it is still prevalent throughout the tree of life today26,40.
Overall, the reported Na+ energetics provides experimental evidence for their existence in microbial phototrophs in general, and the previously undescribed evidence for their role in cyanobacterial N2 fixation in particular, providing a different perspective to understand the low abundance of cyanobacterial N2-fixers in N-limited coastal waters. In nature, low coastal salinities caused by freshwater input (fluviatile and terrestrial freshwater)41,42 and other environmental factors (such as temperature variations, the presence of predators) may together explain the low abundance of Cyanothece sp. Furthermore, the results presented also suggest that sodium energetics may be more widespread than expected, given its probable evolutionary origin in LUCA, and that its metabolic importance in other organisms may still be underestimated.
Methods
Strain and culture conditions
Cyanothece sp. ATCC 51142 was obtained from the ATCC culture collection. Stock cells were grown photoautotrophically at continuous light with a light intensity of 30 μmol m−2 s−1 in artificial seawater medium ASP29 at 30 °C. All experiments were carried out in a light-dark cycle of 12 h/12 h to accommodate the temporal segregation of photosynthesis and N2 fixation, unless stated otherwise. The cells were cultivated without forced aeration, with the sole source of CO2 being diffusion from the atmosphere into the flasks.
N-dependent effects of NaCl on population growth
To evaluate the effect of NaCl on the growth and N2 fixation activity of Cyanothece sp., we performed growth experiments in which we grew cells in artificial seawater medium ASP2 and artificial freshwater medium BG1143, supplemented with or without N, respectively. In total, we had four media, i.e., ASP2, ASP2 without N (ASP2-N), BG11 and BG11 without N (BG110). Prior to growth experiments, exponentially growing stock cells were harvested by centrifugation, washed twice to remove NaCl and resuspended in these four sterile media. The resuspended cells were subsequently inoculated into tissue culture flasks (Thermo Scientific, USA) containing 10 mL of the respective medium with a starting density of approximately 4 × 106 cells/mL. Cultures were grown aerobically at a light intensity of 30 μmol m−2 s−1 in a light-dark cycle of 12 h/12 h on a shaker at 150 rpm (stroke: 26 mm, MQD-S3R, Shanghai Minquan Instruments Co., Ltd) at 30 °C (unless otherwise specified, the same culture parameters were used in all experiments). Growth experiments lasted for nine days, and we recorded cell density daily using a cell counting chamber (Neubauer haemocytometry) under a light microscope (Axio observer Z1, Carl Zeiss, Germany). It should be noted that the counting of cells does not take into account the potential for variation in cell size across different experimental conditions.
To investigate the effects of NaCl deficiency on growth in the absence of N, cells previously grown in BG110 for seven days were subcultured in BG110 supplemented with either NaCl (18 g/L) or NaNO3 (1.5 g/L or 17.6 mM). BG110 without extra additives was used as a blank control. Each culture was inoculated with a starting density of ~ 5 × 105 cells/mL. Growth experiments were conducted in triplicate for seven days. Cell density was quantified as described above.
To investigate the effect of NaCl on the growth of Cyanothece sp. in the presence of N, we conducted experiments with three replicates (with an initial density of approximately 4 × 106 cells/mL) for each of the fourteen NaCl concentrations (0, 2, 4, 9, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36 g/L) in BG11 medium. The experiments lasted seven days, after which cell density was quantified and the increase in cell number was determined. Similar experiments were conducted using BG110 medium instead of BG11 to evaluate the impact of NaCl on the growth of Cyanothece sp. in the absence of N. These experiments lasted seven days, during which cell density and population growth were recorded and calculated. Population growth (the increase in cell number) was determined by subtracting the cell density on day 0 from the cell density on day 7.
To investigate whether the NaCl requirement for N2 fixation depends on cellular growth stage, triplicate populations of cells (with an initial density of approximately 4 × 106 cells/mL) from exponential phase (7 days after inoculation) or stationary phase (20 days after inoculation) were grown in BG110 medium with or without NaCl addition. This experiment lasted for seven days, during which cell density and population growth were recorded and calculated as described above.
Measurement of N2 fixation activity
N2 fixation activity of cells cultured in BG110 with seven NaCl concentrations (0, 2, 4, 9, 18, 20, 22 g/L) was measured by a modified acetylene reduction method. Cultures were grown for five days under the conditions described above and cell density was quantified on the last day. Assays were conducted in Vacutainer tubes containing 2 mL of the corresponding cell samples and 15% acetylene in the gas phase. Each sample was incubated for 6 hours under the same conditions of light, temperature, and shaking as in the growth experiments. 0.2 mL of triplicate samples were analysed for ethylene concentration by high-performance liquid chromatography (HPLC). The rates of nitrogenase activity are reported in nanomoles of C2H4 produced per 108 cells per hour.
Cellular chlorophyll, total protein, dry weight
To quantify cellular chlorophyll, 1 mL of cells from 7-day-old Cyanothece sp. cell cultures grown in either BG110 or BG110 with 18 g/L NaCl medium (hereafter BG110 (NaCl)), were centrifuged and then extracted twice with 5 mL of 80% aqueous acetone. The extracts were pooled and the spectra of this extract and a sample of whole cells were measured using a DW2000 spectrophotometer (Olis, GA, USA), both against 80% acetone or BG11 media as reference. Chlorophyll a and b levels from the acetone extracts were calculated as outlined in a previous study44. Specifically, the concentration of chlorophyll was determined using the equations:
where An is the absorbance spectrophotometrically measured at the specific wavelength of 663 and 647.
Total protein content was determined using a Bradford Protein Assay kit (Solarbio, China) following the manufacturer’s instructions. Cells grown in either BG110 or BG110 (NaCl) for 7 days were collected and cell density was recorded. Then, 2 mL of culture from each treatment was used to extract total protein using a Plant Protein Extraction Kit (Solarbio, China). Specifically, 2 mL of sample was centrifuged and the supernatant was discarded. Glass beads were used to grind the samples and thoroughly break the cell wall under liquid nitrogen. 1 mL of lysis solution was added at 4 °C for 20 minutes, samples were shaken every 5 minutes, followed by centrifugation at 4 °C at 20000 × g for 30 minutes. The supernatant was harvested for quantification. All experiments were done in triplicate.
To quantify cell dry weight, 10 mL of cells grown in either BG110 or BG110 (NaCl) were concentrated in pre-weighed Eppendorf tubes and cell density was quantified. Then, each treatment was centrifuged at 20000 × g for 5 minutes. The resulting pellet was washed twice with ddH2O and the supernatant was discarded. The tubes were dried in an oven at 65 °C for 24 hours and the total dry weight (cells + tubes) was subsequently determined gravimetrically. Cellular dry weight was calculated with the following equation: Cellular dry weight = (Total dry weight – Weight of tube) / cell number.
Metal chlorides, NH4Cl, Glutamine addition
To explore the mechanisms of growth inhibition in the absence of NaCl under diazotrophy, we grew cells in BG110 and provided them with additional substances to determine which part of the metabolism is affected by NaCl deficiency. To this end, we assessed whether NaCl could be replaced by other metal chlorides by adding an equivalent amount (310 mM) of KCl, LiCl, MgCl2 and CaCl2. Moreover, to investigate whether low concentrations of LiCl could stimulate population growth under diazotrophy, a gradient of LiCl (0, 5, 10, 20, 50,100, 200, 300 mM) was added to cells grown in BG110 medium. Furthermore, to investigate whether N2 fixation products can lead to population growth, we supplemented the BG110 medium with additional NH4Cl (2 mM) or glutamine (2 mM, Aladdin, China). The concentration (2 mM) of NH4Cl or glutamine was chosen based on the result of a pre-experiment, which was neither too low to support population growth nor too high to potentially inhibit population growth. We conducted 7-day-long growth experiments using three replicates (with an initial density of approximately 4 × 106 cells/mL) in these experimental setups, cell densities were recorded and the increases in cell number were calculated.
Sparging experiment
For anaerobic incubation, cells with a starting density of approximately 4 × 106 cells/mL were transferred into tissue culture flasks with either BG110 or BG110 supplemented with 18 g/L NaCl. A total of four treatments were included, i.e., sparging (BG110, BG110 (NaCl)) and no sparging (BG110, BG110 (NaCl)). For the two sparging treatments, tissue culture flasks were flushed with N2 for 10 minutes and then sealed with parafilm to prevent diffusive influx of O2. Triplicate cultures were cultivated for seven days under the conditions above. Afterwards, the cell density of each treatment was quantified.
Quantification of nitrogenase content and estimation of nitrogenase activity
The enzyme activities and contents of nitrogenase of Cyanothece sp. collected in the dark period were measured using commercial nitrogenase activity ELISA (enzyme-linked immunosorbent assay) kit (RF12981, Shanghai Ruifan Biological Technology Co., Ltd, Shanghai, China) and nitrogenase content ELISA kit (MM-246202, Jiangsu Meimian Industrial Co., Ltd, Yancheng, China), respectively. The NaNO3 treatment is used to assess the accuracy of the ELISA approach, as cells should not synthesise nitrogenase when N is replete. Enzymes were extracted from cells grown in the respective media (approximately 1 × 108 cells) in an Eppendorf tube containing 1 mL of ice-cold PBS buffer (pH = 7.4). The homogenate was centrifuged at 10000 × g for 5 minutes at 4 °C and the resulting supernatant was used for further assays. 50 µL of standard, sample and control (nothing added) were added to a well of the nitrogenase antibody pre-coated 96-well plate. The HRP (horseradish peroxidase)-conjugated reagent was then added and the whole plate was incubated at 37 °C for 30 minutes. Detection reagents A (50 µL) and B (100 µL) were added sequentially to each well, incubated for 30 minutes at 37 °C and washed with wash buffer for 5 times. 90 µL of TMB (tetramethylbenzidine) substrate was added and incubated for 15 minutes at 37 °C, after which 50 µL of stop solution was added. Finally, the OD (optical density) at 450 nm was measured and the nitrogenase concentration and activity were calculated from the standard curve. For each condition, six biological replicates were measured.
Transcriptomics
20 mL of cells (seven-day-old since inoculation) grown in either BG110 or BG110 with NaCl (18 g/L) were harvested in the dark phase and then centrifuged at 20000 × g for 10 minutes at 4 ˚C to remove the supernatant. Cell pellets were resuspended in ddH2O and centrifuged again. The supernatants were discarded to remove residual medium. This procedure was repeated twice. The collected cell pellets were transferred to RNase-free cryogenic vials and fast frozen in liquid nitrogen for 30 minutes until extraction. In total, we had both cell types in triplicate.
Total RNA was extracted using Plant RNA Purification Reagent for plant tissue (Invitrogen, USA), and genomic DNA was removed using DNase I (TaKara, Dalian). Only high quality RNA samples (OD260/280 = 1.8 ~ 2.2; OD260/230 ≥ 2.0; RIN ≥ 6.5; 28S:18S ≥ 1.0; and > 1 μg) were used for sequencing library construction. Total RNA samples were sent to Shanghai Majorbio Bio-pharm Technology Co., Ltd. (Shanghai, China). Paired-end sequencing was performed on the Illumina sequencing platform (San Diego, CA, USA) according to the manufacturer’s instructions (Illumina).
Data were analysed using the online platform Majorbio Cloud [www.majorbio.com]45. Raw paired-end reads were trimmed and quality controlled for adaptor contamination using SeqPrep [https://github.com/jstjohn/SeqPrep] and Sickle [https://github.com/najoshi/sickle] with default parameters. Clean reads were then aligned to the Cyanothece sp. ATCC 51142 genome using HISAT2 software [http://ccb.jhu.edu/software/hisat2/index.shtml]. Mapped reads from each sample were assembled using StringTie [https://ccb.jhu.edu/software/stringtie/index.shtml?%20t=example] in a reference-based approach.
To identify the differentially expressed genes (DEGs) between the two different treatments, the expression level of each transcript was calculated using the transcripts per million reads (TPM) method. RSEM was used to quantify gene abundance. Essentially, differential expression analysis was performed using DESeq2 with a Q-value ≤ 0.05, and significant DEGs were confirmed only if |log2FC | > 1 and a Q-value ≤ 0.05. Finally, KEGG pathway analyses were performed using Goatools [https://github.com/tanghaibao/Goatools] and KOBAS, respectively. TPM was used for data visualisation. Heatmaps were generated using ComplexHeatmap v2.8.0. Heatmap Z-scores were calculated for each gene by subtracting the gene expression from the row mean and then dividing by the row standard deviation. All other plots were generated using ggplot2 v3.3.5.
Glycogen quantification
Glycogen content was quantified using a hexokinase activity assay kit (BC0745, Solarbio, Beijing, China). Triplicate populations were cultured in BG110 or BG110 (NaCl) in a 12 h/12 h light/dark cycle for 3 days. 10 mL samples of each treatment were collected at the end of the light phase and at the end of the dark phase, respectively. The collected cells were pelleted and washed with distilled water. 30% (weight/volume) KOH was added to lyse the cells, followed by incubation at 95 °C for 90 minutes. Glycogen was precipitated by addition of cold absolute ethanol for 2 hours on ice and collected by centrifugation (15000 g for 10 minutes). The pellets were washed with absolute ethanol and dried at 60 °C. 300 µL of 100 mM sodium acetate buffer (pH 4.75) was added to resuspend the pellets, and 25 µL of the resuspended samples were used to measure the background glucose level. The remaining glycogen samples were digested with amyloglucosidase at 55 °C for 25 minutes. Then 25 µl of sample was mixed with the assay reagent in a light-proof microtitre plate followed by incubation at room temperature (15 minutes). NADPH levels were measured at 340 nm using a DW2000 spectrophotometer (Olis, GA, USA) and the corresponding glycogen levels were calculated.
ATP addition experiment
To examine whether the inhibition of nitrogenase activity was caused by the lack of ATP, additional ATP (0.1 mM, Solarbo, China) was added to BG110. As the commercially available aqueous ATP solution has a concentration of 10 mM and the volume of BG110 medium used in the experiment is 10 mL, a final concentration of 0.1 mM ATP was chosen to alleviate the dilution of the culture medium by the addition of the ATP mother solution and to ensure a sufficiently high ATP concentration to support population growth. Here, 100 µL of ATP mother solution was added to 9900 µL of each culture medium. We tested the growth of three replicates (with an initial density of approximately 4 × 106 cells/mL) in the modified BG110 media. After seven days, the cell density was quantified and the increase in cell number was calculated.
To determine the actual role of ATP in activating population growth, cells (approximately 4 × 106 cells/mL) were cultured in BG110 modified with NaCl (310 mM), glutamine (0.1 mM), NaNO3 (0.1 mM) or ATP (0.1 mM). To ensure that all treatments had the same concentration of additives, 0.1 mM glutamine and NaNO3 were used in accordance with the concentration of ATP, as outlined above. Each treatment had six replicates. Cells were harvested in the dark period on day 3 for quantification of nitrogenase content by ELISA according to the manufacturer’s protocol (Shanghai Ruifan Biological Technology Co., Ltd, Shanghai, China). If the role of ATP is to provide chemical energy, then nitrogenase biosynthesis would be expected; if ATP acts as an N resource, then nitrogenase synthesis is unnecessary. To further examine whether supplemented ATP functions as N or energy source, 0.1 mM (as outlined above) of ADP (Macklin, China) and AMP (Macklin, China) and ATP were added to the cells (a starting density of approximately 4 × 106 cells/mL) grown in BG110. Each treatment had three replicates. After seven days, the increase in cell number was calculated and contrasted.
ATP and ADP quantification
To test whether Na+ directly mediates the intracellular ATP content, we quantified the cellular ATP and ADP content of cells in different NaCl gradients. Cells with a starting density of 4 × 106 cells/mL were first cultured in BG110 for seven days. Since N2 fixation occurs in the dark, NaCl (18 g/L) was added to these non-growing cultures in the dark phase on day 7. Cells were sampled at four time points after the NaCl addition (0 minutes, 10 minutes, 30 minutes, 60 minutes). 1 mL of cells (enumerated under the microscope) were snap-frozen in liquid nitrogen for 20 minutes and stored at −80 °C until ATP and ADP quantification. For ATP extraction, samples were subjected to three consecutive cycles of boiling at 100 °C and freezing in liquid nitrogen, followed by centrifugation at 4 °C at 20,000 × g for 2 minutes. The resulting ATP supernatant was quantified using an established protocol of the ATP content assay kit (AKOP004M, Boxbio, Beijing, China). 200 μL of ATP supernatant and a standard sample (0.625 μmol/mL ATP in ddH2O) were incubated with a working buffer containing hexokinase, glucose, glucose dehydrogenase, and NADP, respectively. The absorbance of the synthesised NADPH was measured at 340 nm using a spectrophotometer at two time points: 10 s and 190 s after incubation. The ATP standard curve was generated and used to calculate the ATP content in the collected samples. For every condition, three biological replicates were measured.
For ADP quantification, cellular ATP and ADP were first extracted as described above, and ADP was subsequently quantified using an ADP Assay Kit (MAK133, Sigma-Aldrich, Missouri, USA) according to the manufacturer’s protocol. Briefly, 10 μL of the samples together with 10 μL of standard (24 μM ADP) were mixed with 90 μL of a working mix containing reaction buffer, luciferin and firefly luciferase, followed by incubation for 10 minutes at room temperature in a 96-well plate. The luminescence was quantified with a spectrophotometer (Olis, GA, USA) to determine the read luminescence for ATP (RLUATP). 5 μL ADP reagent was then added to each well immediately after reading (RLUATP) and incubated for 2 minutes at room temperature to determine the read luminescence for ADP (RLUADP). The ADP standard curve was generated. The luminescence of ADP present in the samples was calculated (RLUADP - RLUATP), and the corresponding ADP content was determined using the standard curve. Three biological replicates were measured for each condition.
Exposure to monensin, DTHB, EIPA and DCCD
The following assays were performed to understand the role of Na+ or H+-coupled ATP synthesis in driving ATP generation in the presence or absence of Na+ under diazotrophy. The Na+-specific ionophore monensin (Solarbo, Beijing, China), an inhibitor of Na+ channels and the Na+/H+ antiporter ethyl-isopropyl amiloride (EIPA, GLPBIO, USA) and the protonophore 3,5-di-tert-butyl4-hydroxybenzaldehyde (DTHB, Thermo Fisher Scientific, USA) were used to test which pathway drives ATP synthesis. A typical F0F1 ATP synthase inhibitor, dicyclohexylcarbodiimide (DCCD, Macklin, China), was used to test whether the H+-motive and Na+-motive machinery share a common ATP synthase. Firstly, to quantify changes in cellular ATP levels and ATP/ADP ratios, monensin (14 µM) or EIPA (100 µM) was added to N2-fixing cells (with a starting density of approximately 4 × 106 cells/mL) grown in BG110 with 18 g/L NaCl in the dark phase on day 7. 1 mL of cells were sampled at four time points after the addition of monensin or EIPA (0 minutes, 10 minutes, 30 minutes, 60 minutes). Cellular ATP and ADP levels were quantified as described above. Secondly, to understand the energetics of cells under different conditions, we tested the growth performance of cells in different media with the addition of monensin, DTHB, EIPA or DCCD. We used four media, including BG11, BG11 with NaCl (18 g/L), BG110, BG110 with NaCl (18 g/L), and cells (with a starting density of approximately 4 × 106 cells/mL) were cultured in each treatment with either monensin (14 µM), DTHB (200 µM), EIPA (100 µM), DCCD (100 µM) or no additive (as blank control). After seven days of growth, the increase in cell number was recorded and contrasted. To test whether monensin interferes with the performance of nitrogenase, cells (with a starting density of approximately 2.5 × 106 cells/mL) were cultured in BG110, BG110 (0.1 mM ATP) and BG110 (0.1 mM ATP & 14 µM monensin), respectively. After three days of growth, the increase in cell number was recorded and compared.
Estimation of the activity of fermentation-related enzymes
To determine the activities of fermentation-related enzymes, cells (approximately 4 × 106 cells/mL) were cultured in BG11 with NaCl (18 g/L) and in BG110 with NaCl (18 g/L), respectively. Each treatment had nine replicates. The experiment was carried out in a 12 h/12 h light-dark cycle, and cells were harvested twice on day 7 at midnight to estimate the activities of three fermentation-related enzymes, i.e., acetate kinase, alcohol dehydrogenase, and lactate dehydrogenase, using ELISA, according to the manufacturer’s protocol (MM-1181O2, MM-91569O2, MM-2023O2, Jiangsu Meimian Industrial Co., Ltd, Yancheng, China).
Statistical analysis
Physiological parameters were analysed using two-tailed Welch’s t test. Growth data, nitrogenase activity data (acetylene reduction method) and ELISA data were analysed by one-way ANOVA. Monensin, DTHB, EIPA and DCCD data were analysed by two-way ANOVA. Significance: ns (no significance), *(p < 0.05), **(p < 0.01), ***(p < 0.001), ****(p < 0.0001). All statistical analyses were performed in R v4.1.146.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw RNAseq reads for the analysis of differential gene expression have been submitted to NCBI’s SRA database (http://www.ncbi.nlm.nih.gov) under BioProject PRJNA1014498. The KEGG database [https://www.kegg.jp/] was used for functional enrichment analyses. The data generated in this study are provided in the Source Data file. Source data are provided with this paper.
Code availability
All scripts for data visualisation are available at https://github.com/SiTANG1990/N2-fixation-coastal-unicellular-cyanobacteria.git [https://doi.org/10.5281/zenodo.1392310947].
References
Zehr, J. P. & Capone, D. G. Changing perspectives in marine nitrogen fixation. Science 368, eaay9514 (2020).
Zehr, J. P. Nitrogen fixation by marine cyanobacteria. Trends Microbiol 19, 162–173 (2011).
Huisman, J. et al. Cyanobacterial blooms. Nat. Rev. Microbiol. 16, 471–483 (2018).
Howarth, R. W., Marino, R., Howarth, R. W. & Marino, R. Nitrogen as the limiting nutrient for eutrophication in coastal marine ecosystems: evolving views over three decades. Limnol. Oceanogr. 51, 364–376 (2006).
Conley, D. J. et al. Controlling eutrophication: nitrogen and phosphorus. Science 323, 1014–1015 (2009).
Paerl, H. W. Physiological ecology and regulation of N2 fixation in natural waters. Adv. Microb. Ecol. 8, 305–344 (1990).
Moisander, P. H. et al. Unicellular cyanobacterial distributions broaden the oceanic N2 fixation domain. Science 327, 1512–1514 (2010).
Mareš, J. et al. Taxonomic resolution of the genus Cyanothece (Chroococcales, Cyanobacteria), with a treatment on Gloeothece and three new genera, Crocosphaera, Rippkaea, and Zehria. J. Phycol. 55, 578–610 (2019).
Reddy, K. J., Haskell, J. B., Sherman, D. M. & Sherman, L. A. Unicellular, aerobic nitrogen-fixing cyanobacteria of the genus Cyanothece. J. Bacteriol. 175, 1284–1292 (1993).
Forchhammer, K. & Schwarz, R. Nitrogen chlorosis in unicellular cyanobacteria-a developmental program for surviving nitrogen deprivation. Environ. Microbiol. 21, 1173–1184 (2019).
Hauryliuk, V., Atkinson, G. C., Murakami, K. S., Tenson, T. & Gerdes, K. Recent functional insights into the role of (p)ppGpp in bacterial physiology. Nat. Rev. Microbiol. 13, 298–309 (2015).
Irving, S. E., Choudhury, N. R. & Corrigan, R. M. The stringent response and physiological roles of (pp)pGpp in bacteria. Nat. Rev. Microbiol. 19, 256–271 (2021).
Flores, E. & Herrero, A. Compartmentalised function through cell differentiation in filamentous cyanobacteria. Nat. Rev. Microbiol. 8, 39–50 (2010).
Flores, E. & Herrero, A. Nitrogen assimilation and nitrogen control in cyanobacteria. Biochem. Soc. Trans. 33, 164–7 (2005).
Berman-Frank, I., Lundgren, P. & Falkowski, P. Nitrogen fixation and photosynthetic oxygen evolution in cyanobacteria. Res. Microbiol. 154, 157–164 (2003).
Stal, L. J. Nitrogen Fixation in Cyanobacteria. In eLS, (John Wiley & Sons, Ltd, 2015) https://doi.org/10.1002/9780470015902.a0021159.pub2.
Welsh, E. A. et al. The genome of Cyanothece 51142, a unicellular diazotrophic cyanobacterium important in the marine nitrogen cycle. Proc. Natl Acad. Sci. USA 105, 15094–15099 (2008).
Schneegurt, M. A., Sherman, D. M., Nayar, S. & Sherman, L. A. Oscillating behavior of carbohydrate granule formation and dinitrogen fixation in the cyanobacterium Cyanothece sp. strain ATCC 51142. J. Bacteriol. 176, 1586–1597 (1994).
Burnap, R. L., Hagemann, M. & Kaplan, A. Regulation of CO2 concentrating mechanism in cyanobacteria. Life 5, 348–371 (2015).
Doello, S., Burkhardt, M. & Forchhammer, K. The essential role of sodium bioenergetics and ATP homeostasis in the developmental transitions of a cyanobacterium. Curr. Biol. 31, 1606–1615 (2021).
Mempin, R. et al. Release of extracellular ATP by bacteria during growth. BMC Microbiol 13, 301 (2013).
Alvarez, C. L. et al. Dynamic regulation of extracellular ATP in Escherichia coli. Biochem. J. 474, 1395–1416 (2017).
Spari, D. & Beldi, G. Extracellular ATP as an inter-kingdom signaling molecule: release mechanisms by bacteria and its implication on the host. Int. J. Mol. Sci. 21, 5590 (2020).
Winkler, H. H. & Neuhaus, H. E. Non-mitochondrial ATP transport. Trends Biochem. Sci. 24, 64–68 (1999).
Von Ballmoos, C., Wiedenmann, A. & Dimroth, P. Essentials for ATP synthesis by F1F0 ATP synthases. Annu. Rev. Biochem. 78, 649–672 (2009).
Mulkidjanian, A. Y., Dibrov, P. & Galperin, M. Y. The past and present of sodium energetics: may the sodium-motive force be with you. Biochim. Biophys. Acta 1777, 985–992 (2008).
Huczyński, A., Janczak, J., Łowicki, D. & Brzezinski, B. Monensin a acid complexes as a model of electrogenic transport of sodium cation. Biochim. Biophys. Acta 1818, 2108–2119 (2012).
Efiok, B. J. S. & Webster, D. A. Sodium-coupled ATP synthesis in the bacterium Vitreoscilla. Arch. Biochem. Biophys. 292, 102–106 (1992).
Toei, M. & Noji, H. Single-molecule analysis of F0F1-ATP synthase inhibited by N,N-Dicyclohexylcarbodiimide. J. Biol. Chem. 288, 25717–25726 (2013).
Bandyopadhyay, A., Elvitigala, T., Liberton, M. & Pakrasi, H. B. Variations in the rhythms of respiration and nitrogen fixation in members of the unicellular diazotrophic cyanobacterial genus Cyanothece. Plant Physiol. 161, 1334–1346 (2013).
Bandyopadhyay, A., Sengupta, A., Elvitigala, T. & Pakrasi, H. B. Endogenous clock-mediated regulation of intracellular oxygen dynamics is essential for diazotrophic growth of unicellular cyanobacteria. Nat. Commun. 15, 3712 (2024).
Heyer, H., Stal, L. & Krumbein, W. E. Simultaneous heterolactic and acetate fermentation in the marine cyanobacterium Oscillatoria limosa incubated anaerobically in the dark. Arch. Microbiol. 151, 558–564 (1989).
Steunou, A.-S. et al. In situ analysis of nitrogen fixation and metabolic switching in unicellular thermophilic cyanobacteria inhabiting hot spring microbial mats. Proc. Natl Acad. Sci. USA 103, 2398–2403 (2006).
Stal, L. J. & Moezelaar, R. Fermentation in cyanobacteria. FEMS Microbiol. Rev. 21, 179–211 (1997).
Trchounian, A. Mechanisms for hydrogen production by different bacteria during mixed-acid and photo-fermentation and perspectives of hydrogen production biotechnology. Crit. Rev. Biotechnol. 35, 103–113 (2015).
Fay, P. Oxygen relations of nitrogen fixation in cyanobacteria. Microbiol. Rev. 56, 340–373 (1992).
Müller, V. et al. Bacterial Fermentation. in eLS (John Wiley & Sons, Ltd). https://doi.org/10.1002/9780470015902.a0001415.pub2 (2008).
Apte, S. K. & Thomas, J. Effect of sodium on nitrogen fixation in Anabaena torulosa and Plectonema boryanum. Microbiology 130, 1161–1168 (1984).
Burkhardt, M., Rapp, J., Menzel, C., Link, H. & Forchhammer, K. The global influence of sodium on cyanobacteria in resuscitation from nitrogen starvation. Biology 12, 159 (2023).
Kozlova, M. I. et al. Expansion of the “Sodium World” through evolutionary time and taxonomic space. Biochemistry 85, 1518–1542 (2020).
Du, J. & Park, K. Estuarine salinity recovery from an extreme precipitation event: Hurricane Harvey in Galveston Bay. Sci. Total Environ. 670, 1049–1059 (2019).
Konapala, G., Mishra, A. K., Wada, Y. & Mann, M. E. Climate change will affect global water availability through compounding changes in seasonal precipitation and evaporation. Nat. Commun. 11, 3044 (2020).
Rippka, R., Deruelles, J., Waterbury, J. B., Herdman, M. & Stanier, R. Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111, 1–61 (1979).
Arnon, D. I., McSwain, B. D., Tsujimoto, H. Y. & Wada, K. Photochemical activity and components of membrane preparations from blue-green algae. I. Coexistence of two photosystems in relation to chlorophyll a and removal of phycocyanin. Biochim. Biophys. Acta 357, 231–245 (1974).
Ren, Y. et al. Majorbio Cloud: A one-stop, comprehensive bioinformatic platform for multiomics analyses. iMeta 1, e12 (2022).
Ihaka, R. & Gentleman, R. R. A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Tang, S. et al. A unicellular cyanobacterium relies on sodium energetics to fix N2. Zenodo https://doi.org/10.5281/zenodo.13923109 (2024).
Acknowledgements
We thank Yuelu Jiang and Peter Deines for their helpful comments on this paper. This work was supported by the S&T Projects of Shenzhen Science and Technology Innovation Committee (KCXFZ20211020163557022) to ZH.C. K.H. gratefully acknowledges the financial support of the John Templeton Foundation (#62220). The opinions expressed in this paper are those of the authors and not those of the John Templeton Foundation.
Author information
Authors and Affiliations
Contributions
S.T., K.H., and ZH.C. conceptualised and designed the experiments. S.T., X.Y.C., YQ.L., L.L., Q.Y., D.L., and JM.Z. (Jianming Zhu) performed the experiments. S.T., K.H., J.Z. (Jin Zhou), and YY.J. analysed and visualised the data. All authors interpreted the results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Karl Forchhammer, Hakuto Kageyama and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, S., Cheng, X., Liu, Y. et al. A unicellular cyanobacterium relies on sodium energetics to fix N2. Nat Commun 15, 9716 (2024). https://doi.org/10.1038/s41467-024-53978-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53978-9