Abstract
Cell ion channels, cell proliferation and metastasis, and many other life activities are inseparable from the regulation of trace or even single copper ion (Cu+ and/or Cu2+). In this work, an electrochemical sensor for sensitive quantitative detection of 0.4−4 amol L−1 copper ions is developed by adopting: (1) copper ions catalyzing the click-chemistry reaction to capture numerous signal units; (2) special adsorption assembly method of signal units to ensure signal generation efficiency; and (3) fast scan voltammetry at 400 V s−1 to enhance signal intensity. And then, the single-atom detection of copper ions is realized by constructing a multi-layer deep convolutional neural network model FSVNet to extract hidden features and signal information of fast scan voltammograms for 0.2 amol L−1 of copper ions. Here, we show a multiple signal amplification strategy based on functionalized nanomaterials and fast scan voltammetry, together with a deep learning method, which realizes the sensitive detection and even single-atom detection of copper ions.
Similar content being viewed by others
Introduction
Single-atom detection (SAD) and single-molecule detection (SMD) usually refer to methods for detecting the properties and behaviors of individual atoms and molecules in a nanoscale1,2,3. Undoubtedly, they have reached the detection sensitivity limit of atoms and molecules, converting the statistical measurement of macroscopic objects into the discrete detection of microscopic individuals4,5,6,7, and then providing individual information in a microenvironment that cannot be obtained by traditional macroscopic measurement methods8,9. Involved in many research fields such as physics, chemistry, material science, life science, medicine, information science, electronics and so on, they have theoretical and practical application prospects in reaction kinetics mechanism, medical or forensic diagnosis, single-atom or single-molecule recognition, biomacromolecule structure and function, and microscopic performance measurement of nanomaterials, etc10,11,12,13.
Currently, there are two main types of SAD and SMD. The first type is spectroscopic methods, including surface-enhanced Raman scattering (SERS) spectroscopy14, resonance ionization spectroscopy (RIS)15, laser-induced fluorescence16, atom-trap trace analysis17, etc. The second type is scanning probe microscopic methods, including stimulated emission depletion (STED) microscopy18, scanning tunneling microscopy (STM)19, scanning near-field optical microscopy (SNOM)20, atomic force microscopy (AFM)21, etc.
In fact, since Bard et al. inaugurated observing the redox reaction of a single [(trimethylammonio)methyl]ferrocene (Cp2FeTMA+) molecule in the solution between the Pt-Ir microelectrode probe and the conductive substrate22, some advantageous electrochemical SAD and SMD methods based on laser-pulled nanoelectrodes and nanofabrication techniques23,24,25 have been attempted. Generally, to achieve electrochemical SAD and SMD, the following three capabilities should be available. (1) Being able to capture the properties of a single atom or molecule. Presently, two main strategies are usually adopted, one is to use nanoelectrode as the working electrode, performing electrochemical detection of countable atoms and molecules in the solution near the electrode26,27,28; the other is to use fast scan voltammetry (FSV) to implement fast potential perturbation, restricting the electrode reactions to a nanoscale space29,30. (2) Being able to amplify weak signals. Since the electrochemical signal from one atom or molecule is too weak, it should be amplified to a detectable level in various ways. Increasing the number of signal labels through target-driven catalytic coupling strategies could be employed31. The Cu+-catalyzed click chemistry reaction (copper-catalyzed azide-alkyne cycloaddition reaction, CuAAC), which is of mild reaction conditions, fast kinetics and high yield, has been applied in electrochemical sensing to achieve signal triggering and amplification32,33,34. In addition, the development of electrode designs such as nanopore electrode arrays can be adopted35, as well as the introduction of advanced acquisition components such as electrochemical transistors to enhance signal collection capabilities36, and the use of highly sensitive detection instruments such as lock-in amplifiers to suppress background interference37. (3) Being able to process complex signals. As easily masked by noise, complex signals should be processed to extract valuable information using data processing techniques such as wavelet transform38 and deep learning39,40,41,42, etc. Among them, deep learning is gradually demonstrating its application potential in analysis, which can extract vague information from high-noise and low-resolution sensing data, discover hidden relationships between sample parameters and sensing signals, and mine the correlation between signals and chemical events43,44,45.
In this study, based on the specific catalytic behavior of a single-atom Cu+ towards the CuAAC reaction46, SAD of copper ions (Cu+ and/or Cu2+) was achieved by using functionalized bionanomaterials to increase the number of signal labels, adopting FSV to enhance the signal intensity, and developing a deep learning method to identify and extract weak signals. In this case, electrochemical single-atom detection of conventional samples using conventional electrodes was realized, providing an alternative strategy for SAD and SMD.
Results
SAD principle of the sensor
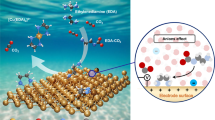
The combination of click chemistry and FSV was pioneered to develop an electrochemical sensor based on functionalized two-dimensional nanomaterials for the SAD of copper ions. As shown in Supplementary Fig. 1, [Ag(NH3)2]+ is reduced in situ by black phosphorus nanosheet (BPNS) to obtain BPNS@AgNPs with a large number of Ag nanoparticles (AgNPs) uniformly distributed on the surface of BPNS, and then propargylamine is bound to AgNPs via the Ag-N bond to obtain the BPNS@AgNPs-Alkyne signal unit. When Cu+ or Cu2+, which can be reduced to Cu+ by sodium ascorbate (SA), is present in the sample, a click chemistry reaction is triggered between the alkynyl group in the signal unit and the azide group in Azide-Fe3O4, resulting in the formation of a complex signal unit-Fe3O4 (Fig. 1a). Under the action of magnetic force, the complex could be quickly and firmly adsorbed onto the surface of the magnetic glassy carbon electrode (MGCE, 2 mm diameter), and then an electrochemical sensor could be prepared in one step (Fig. 1b). Thus, FSV signals could be produced by the redox reaction of AgNPs. Obviously, more copper ions correspond to more AgNPs and stronger electrochemical signal, so sensitive or even single-atom detection of copper ions can be achieved, mainly due to the multiple signal amplification mechanism as follows. (1) The number of labels is greatly increased owing to the special labeling mode. In this system, a Cu2+ ion is reduced to a Cu+ ion, which can capture multiple signal units by continuously catalyzing the click chemistry reaction between the BPNS@AgNPs-Alkyne signal unit and Azide-Fe3O4 as a catalyst. Each signal unit contains a large number of AgNPs with thousands of Ag atoms on the surface. In this case, a cascading signal amplification chain, that is, copper ions - signal unit - AgNPs - Ag atoms, is constructed. According to a simple estimation that each Cu+ could trigger the formation of 102 signal units by catalyzing the click chemistry reaction, each signal unit could bind 104 AgNPs, and each AgNP contains 105 Ag atoms on its surface, therefore the number of signal label Ag atoms n initiated by one copper ion is 1011. (2) The adsorption assembly method ensures the efficiency of signal generation. The electron transfer between the signal label and the electrode is greatly affected by the distance d between them. According to the basic theory of electrochemistry, the signal current i decays exponentially with the increase of d. In the construction of this sensor, the signal unit BPNS@AgNPs-Alkyne is directly adsorbed on the electrode surface, and BPNS with an electrical conductivity of about 300 S m−1 makes itself a new electrode surface. Thus, AgNPs on BPNS can be considered as being directly adsorbed and assembled on the electrode surface, resulting in d = 0. Therefore, all the Ag atoms on the surface of AgNPs can directly participate in the electrode reaction to ensure the signal generation efficiency close to 100%. (3) FSV amplifies the signal current. Once the sensor is constructed, the charge Q = ne- produced in one redox process is determined, as the number of AgNPs captured on the electrode surface is determined. The shorter the time t = E/v to complete the process, where E is the potential window and v is the scan rate, the larger the signal current i = Q/t = Qv/E = ne – v/E. Assuming E = 0.4 V and v = 400 V s−1, i is in the order of 10−5 A, which is an easily measured electrochemical signal (Fig. 1c). Furthermore, the SAD of copper ions could be realized by deep learning (Fig. 1d).
a Target-triggered click chemistry reaction. The reduction of Cu2+ to Cu+ in the sample triggers a click chemistry reaction between the signal unit and the Azide-Fe3O4, resulting in the formation of a complex signal unit-Fe3O4. b Electrochemical sensor. The complex signal unit-Fe3O4 is adsorbed onto the magnetic glassy carbon electrode (MGCE) surface under the action of magnetic force. c FSV detection. At a scan rate of 400 V s−1, the in-situ redox reaction of AgNPs generates electrochemical signals. d Deep learning method for SAD of copper ions. Deep learning enables the distinction between positive and negative signals. BPNS: black phosphorus nanosheet.
Characterization of nanomaterials
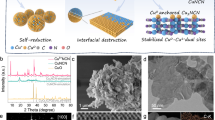
The morphology of the nanomaterials was characterized by scanning electron microscopy (SEM). The SEM and inset dynamic light scattering (DLS) analysis images in Fig. 2a show that the azide-Fe3O4 particles are spherical with a diameter of about 250 nm. BPNS exhibits a typical two-dimensional sheet structure with large surface area and good dispersibility (Fig. 2b). Due to the high affinity between the phosphorus scaffold and the metal ions/atoms, BPNS can efficiently enrich and reduce [Ag(NH3)2]+ in situ to obtain BPNS@AgNPs (Fig. 2c). A large number of AgNPs with a uniform size of about 30 nm are evenly and densely distributed on BPNS, which is also confirmed by energy dispersive X-ray spectroscopy (EDS) elemental analysis (Supplementary Fig. 2) and mapping (Supplementary Fig. 3). BPNS@AgNPs are functionalized by alkynyl groups to form the signal unit BPNS@AgNPs-Alkyne (Supplementary Figs. 4 and 5), which can combine with Azide-Fe3O4 through a click chemistry reaction to form a complex signal unit-Fe3O4 (Fig. 2d and the inset). The complex can be easily adsorbed on the electrode surface under magnetic force through minimal loading of ≈5–10 magnetic Fe3O4 nanoparticles, thus to construct the sensor and carry out electrochemical detection (Supplementary Fig. 6).
SEM images of a Azide-Fe3O4 (inset: DLS analysis of azide-Fe3O4 particles), b BPNS, c BPNS@AgNPs, and d complex signal unit-Fe3O4.
The preparation of BPNS@AgNPs-Alkyne was also characterized by Fourier transform infrared spectroscopy (FTIR) and zeta potential. As shown in Supplementary Fig. 7, compared to BPNS@AgNPs, BPNS@AgNPs-Alkyne exhibits a new peak at 2115 cm−1 corresponding to the stretching vibration of the alkynyl group, suggesting that propargylamine is bound to BPNS@AgNPs. Zeta potentials of BPNS, BPNS@AgNPs, propargylamine, and BPNS@AgNPs-Alkyne are −22.2, −24.3, 23.5 and 3.55 mV, respectively (Supplementary Fig. 8), also demonstrating the preparation of BPNS@AgNPs-Alkyne.
Characterization of the sensor
The click chemistry reaction between BPNS@AgNPs-Alkyne and Azide-Fe3O4 was characterized by FTIR and EDS. As shown in Fig. 3a, the peak of BPNS@AgNPs-Alkyne at 2115 cm−1 corresponds to the stretching vibration of the alkynyl group (green line), and the peak of Azide-Fe3O4 at 2100 cm−1 corresponds to the stretching vibration of the azide group (orange line). In the presence of Cu2+ and SA, the peaks at 2115 and 2100 cm−1 disappear, while the other peak appears at 1457 cm−1, which suggests that the alkynyl group reacts with the azide group to form a triazole ring (Fig. 3a, blue line), and the CuAAC reaction is completed. EDS elemental analysis (Supplementary Fig. 9) and mapping (Supplementary Fig. 10) show that the elemental composition of the complex is Ag, P, Fe, C, N and O, also indicating that it is obtained by CuAAC reaction between signal unit BPNS@AgNPs-Alkyne and Azide-Fe3O4.
a FTIR spectra to verify the CuAAC reaction between signal unit BPNS@AgNPs-Alkyne and Azide-Fe3O4. The curves are BPNS@AgNPs-Alkyne (green line), Azide-Fe3O4 (orange line), and complex signal unit-Fe3O4 (blue line). b CV and c EIS to characterize the sensor assembly process, where the curves are bare MGCE (blue line), and MGCE + complex signal unit-Fe3O4 (orange line). Experimental conditions: 0.1 mol L−1 KCl containing 5.0 mmol L−1 [Fe(CN)6]3−/4−; potential scan range: −0.2 to 0.6 V vs. Ag/AgCl (3 mol L−1 KCl); scan rate of CV: 50 mV s−1; frequency range and AC amplitude of EIS: 0.01 to 10 kHz, 5 mV. d Feasibility of electrochemical sensor characterized by FSV. Curves shown here are a subset of FSV voltammograms of bare MGCE (orange line), Azide-Fe3O4 (blue line), BPNS@AgNPs-Alkyne (pink line), Azide-Fe3O4 + BPNS@AgNPs-Alkyne (gray line), Azide-Fe3O4 + BPNS@AgNPs-Alkyne + Cu2+ (brown line), Azide-Fe3O4 + BPNS@AgNPs-Alkyne + SA (green line) and Azide-Fe3O4 + BPNS@AgNPs-Alkyne + Cu2+ + SA (violet line), all of which are background subtracted and baselines are all set to 0 as templated as Supplementary Fig. 11. e Identical FSV voltammograms in continuous-cycle mode. The curves are the 1st cycle at 400 V s−1 (orange line), the 800th cycle at 400 V s−1 (gray line). Curves shown here are FSV voltammograms without background subtraction. Experimental conditions of d and e: 0.5 mol L−1 KNO3 containing 1 amol L−1 Cu2+; potential window: −1.0 −1.5 V vs. Ag/AgCl (3 mol L−1 KCl); scan rate: 400 V s−1. Source data are provided as a Source Data file.
The assembly process of the electrochemical sensor was characterized by cyclic voltammetry (CV) and electrochemical impedance spectroscopy (EIS). As shown in Fig. 3b, c, compared to those of the bare MGCE (blue line), the peak currents of the probe [Fe(CN)6]3-/4- on the electrode modified by the complex signal unit-Fe3O4 increase and the charge transfer resistance (Rct) decreases (orange line), due to the large surface area and good conductivity of the complex.
The feasibility of the electrochemical sensor was also characterized by FSV. As shown in Fig. 3d, bare MGCE (orange line), Azide-Fe3O4 (blue line), BPNS@AgNPs-Alkyne (pink line), Azide-Fe3O4 + BPNS@AgNPs-Alkyne (gray line), Azide-Fe3O4 + BPNS@AgNPs-Alkyne + Cu2+ (brown line), and Azide-Fe3O4 + BPNS@AgNPs-Alkyne + SA (green line) could not produce faradaic current signals. Only in the case of Azide-Fe3O4 + BPNS@AgNPs-Alkyne + Cu2+ + SA (violet line), a significant faradaic current signal is observed, indicating that it is feasible to detect copper ions with this sensor based on the CuAAC reaction.
To eliminate some factors degrading the precision of single-cycle triggered signals, voltammograms were carried out in the continuous-cycle mode with no delay between repetitive scans. In ref. 47. we found that Ag undergoes in-situ redox reaction when the scan rate is higher than 200 V s−1, that is, Ag atoms on the surface of AgNPs are oxidized to Ag+ ions in the anodic scan and Ag+ ions are quickly reduced to Ag atoms in the cathodic scan without diffusing away from their established location into the bulk solution beyond the outer Helmholtz plane (OHP). In this case, the voltammograms are almost identical even after successive 800 scan cycles at 400 V s−1 (Fig. 3e), so we can take any one of these identical voltammograms for analysis.
Performance of the sensor
To achieve the best performance of the electrochemical sensor, the click chemistry reaction time and the scan rate have been systematically optimized. As shown in Fig. 4a, the FSV response increases gradually with the increment of the click chemistry reaction time, and the increase slows down after 40 min. It is not difficult to understand that the occurrence of the click chemistry reaction requires the simultaneous existence of BPNS@AgNPs-Alkyne, Azide-Fe3O4, and copper ions in a limited space. As the reaction progresses, the available concentrations of BPNS@AgNPs-Alkyne and Azide-Fe3O4 around copper ions will decrease, resulting in a decrease in the rate of the click chemistry reaction and no further obvious increase in the signal current. As for the scan rate, the FSV response increases with it and reaches a plateau at 400 V s−1 (Fig. 4b). It is because the diameter of the electrode used in this experiment was 2 mm. When the electrode size is at the level of mm, the maximum usable scan rate is about 100–1000 V s−1 30. Else, the charging current will be coupled with the faradaic current, causing severe distortion to the electrochemical signal. In addition, the redox reaction kinetics of AgNPs may also be a potential limiting factor. Therefore, the optimum click chemistry reaction time and scan rate for the FSV response were selected as 40 min and 400 V s−1, respectively.
The optimization of the click chemistry reaction time a and the scan rate b for FSV response. The concentration of Cu2+ used was 1.0 amol L−1. c Selectivity and anti-interference of the sensor. The concentration of Cu2+ used was 1.0 amol L−1. The concentration of some common interfering metal ions including Al3+, Ca2+, Cd2+, Co2+, Fe3+, Hg2+, Mg2+, Mn2+, and Pb2+ was 1000 amol L−1. Mixture 1 was the mixture of 1.0 amol L−1 Cu2+ and common interfering metal ions including Al3+, Ca2+, Cd2+, Co2+, Fe3+, Hg2+, Mg2+, Mn2+, and Pb2+ at the concentration of 1000 amol L−1. Mixture 2, 3, 4 and 5 were the mixture of 1.0 amol L−1 Cu2+ and 10 amol L−1 of cysteine, serine, chitosan and Nafion, respectively. d Storage stability and reproducibility. Batches of functional materials and sensors were stored at 4 °C, and used to determine 1 amol L−1 Cu2+ for 10 consecutive days. e A screenshot of real-time FSV voltammogram of 1 amol L−1 Cu2+. The output potential is converted into the current in the voltammogram according to the specific circuit design. f FSV signals of the electrochemical sensor at different Cu2+ concentrations from 0.4 to 4 amol L−1. The curves shown here are a subset of FSV voltammograms with background subtracted and baselines all set to 0 as in Supplementary Fig. 11. g Linear relationship between the peak current and the Cu2+ concentration. Error bars represent standard deviations, n = 5. Source data are provided as a Source Data file.
a Negative and positive FSV signals. The curves shown here are a subset of FSV voltammograms without averaging by oscilloscope and background subtraction. The theoretical number of Cu2+ contained in the sample is 1.2. b The framework of our image classifier FSVNet based on DCNN. FSV voltammogram images are captured by the home-built electrochemical system and inputted to the computer for deep learning, including feature extraction and classification predictions. c Accuracy and d loss rates of FSVNet during the learning process. e AUC diagram of FSVNet model on testing dataset. The receiver operating characteristic (ROC) curve illustrates an area under the curve (AUC) of 1.00. Source data are provided as a Source Data file.
The selectivity of the sensor was investigated by comparing the FSV response currents of 1 amol L−1 Cu2+ and some common interfering metal ions including Al3+, Ca2+, Cd2+, Co2+, Fe3+, Hg2+, Mg2+, Mn2+, and Pb2+ at the concentration of 1000 amol L−1. No obvious FSV response of interfering ions is observed (Fig. 4c). In addition, when 1 amol L−1 Cu2+ coexists with all interfering ions at the concentration of 1000 amol L−1 (mixture 1), and 10 amol L−1 of cysteine (mixture 2), serine (mixture 3), chitosan (mixture 4) and Nafion (mixture 5), respectively, the FSV response currents of these mixtures are almost the same as that in the presence of Cu2+ alone. The above results show that the electrochemical sensor has good selectivity for copper ion detection without significant interference from common metal ions, amino acids and polyelectrolytes, due to the high specificity of the CuAAC reaction. It is worth mentioning that there is a perceptible decrease in the signal at high concentrations of cysteine, serine, chitosan and Nafion up to 100 amol L−1 (Supplementary Fig. 12), mainly due to their coordination with copper ions.
The storage stability and preparation reproducibility were further checked. Batches of functional materials and electrochemical sensors were prepared, stored at 4 °C, and used to determine 1 amol L−1 Cu2+ for 10 consecutive days. As shown in Fig. 4d, the average response value is 97.5% of the initial value with a relative standard deviation (RSD) of 5.0%, indicating high storage stability, which could be attributed to the fact that it is completely prepared from chemical materials. Reproducibility is also good, mainly because the simple preparation can be performed in a single step.
Under the optimal experimental conditions (Fig. 4a, b), the linear range and the sensitivity of the electrochemical sensor were studied. As shown in Fig. 4e, the digital oscilloscope can capture the FSV voltammogram in real time. After data processing according to the steps described in Supplementary Fig. 11, a series of peak current curves were obtained (Fig. 4f), and the peak current of FSV increases with the increment of Cu2+ concentration. In the concentration range of 0.4–4 amol L−1, there is a good linear relationship between the peak current and the Cu2+ concentration. The linear regression equation is y = 0.325 mA L amol−1 × x − 0.032 mA with a coefficient of determination of R2 = 0.996, where y is the peak current in mA and x is the Cu2+ concentration in amol L−1. When BPNS@AgNPs-Alkyne and Azide-Fe3O4 are in large excess, copper ion becomes the rate-limiting agent in the CuAAC reaction. The higher the copper ion concentration, the higher the CuAAC reaction rate, and the more signal units adsorbed on the electrode surface, the stronger the signal. That is to say, the signal current is proportional to the Cu2+ concentration. However, when the Cu2+ concentration reaches 10 amol L−1 or higher, the linear relationship between the peak current and the Cu2+ concentration no longer exists, and the peak current is a little lower than expected, mainly due to the emerging stacking of the complex signal unit-Fe3O4 (Supplementary Fig. 13). In this case, some AgNPs assembled are too far away from the electrode surface, thus making electron tunneling and the subsequent electrochemical reaction difficult. And more Fe3O4 nanoparticles were immobilized onto each BPNS@AgNPs by CuAAC reaction is another reason. Compared to the main methods reported for sensitive detection of copper ions (Table 1), this strategy exhibits much higher sensitivity owing to the combination of specially designed functionalized bionanomaterials, elaborately arranged electrode assembly and FSV with signal enhancement capability. Among them, a target-induced click conjugation for monitoring of Cu2+ using horseradish peroxidase as the indicator and nanogold particles as the enhancer was reported with a dynamic working range of 1.0 amol L−1 - 10 μmol L−1 and a detection limit of 0.38 amol L−1, but effective sensitivity comparison with this work48 cannot be achieved because the volume of the reaction solution was not mentioned.
It is worth noting that, the RSDs were large at lower concentrations. For instance, five current responses of 0.4 amol L−1 Cu2+ solution were 0.059, 0.122, 0.059, 0.121 and 0.097 mA, respectively, with a mean of 0.0916 mA, a standard deviation of 0.0314 mA and an RSD of 34%. This is mainly because, when measuring at or close to single-atom level, there will be inherent variability. In this case, the notion of concentration is no longer so well defined, but instead solutions containing 0, 1, 2, … atoms are expected with statistics following a Poisson distribution49,50. In fact, it is not possible for the concentration to be exactly 0.4 amol L−1, since this implies 2.4 copper ions in the 10 μL sample solution used. According to the prediction of a Poisson distribution,
it is possible to have 0, 1, 2, 3, 4, or ≥5 ions with a probability of 9%, 22%, 26%, 21%, 12%, and 10%, respectively. Therefore, the RSD of 0.4 amol L−1 sample measured 10 times is about 60% theoretically, which is not significantly different from our results of 34% considering the small sample size, and uncertainties and the stochastic nature of measurements at or close to single-atom level.
Accuracy and precision were investigated by the standard addition method using Cu2+ solutions of 0.5, 1.5 and 3.5 amol L−1, each of which was added with an appropriate amount of high-concentration standard solution to achieve a final concentration of ≈4 amol L−1. As shown in Table 2, the recoveries are 90.0%-96.6% and the RSDs are 4.7%-6.7%, showing good accuracy and precision. The accuracy and precision of the 0.5 amol L−1 sample detected using the standard addition method are much better than those of 0.4 amol L−1 sample detected directly, indicating that increasing the sample concentration within the allowable linear range by the standard addition method can effectively improve the accuracy and precision of measurements at or close to single-atom level.
Deep learning-assisted SAD of copper ions
Since the sample volume used is 10 μL, the theoretical average number of Cu2+ ions contained in the 0.2 amol L−1 sample is 1.2. As shown in Fig. 5a, the signal intensity of the 0.2 amol L−1 sample (positive) cannot reach three times that of the noise, and its FSV images are difficult to be directly distinguished from those of the blank sample (negative) according to the traditional signal identification criterion of signal/noise ratio of 3, resulting in the inability to apply traditional quantitative strategies (Supplementary Fig. 14). Therefore, a deep learning strategy based on image classification, as shown in Fig. 5b, was adopted to achieve SAD.
The accuracy and the loss rate of typical deep convolutional neural network (DCNN) models including VGGNet, InceptionNet, ResNet, DenseNet and our FSVNet, were investigated using training and validation sets. As shown in Fig. 5c, d, a steady increase in accuracy and a steady decrease in loss rate during the learning process are exhibited in the case of our FSVNet, better than those of VGGNet, InceptionNet, ResNet and DenseNet (Supplementary Fig. 15). In addition, about 100% accuracy is achieved at nearly 1/3 of the epoch, indicating that FSVNet generalizes well to the current signal. And, the accuracy and the loss rate in the training set and the test set of FSVNet are 99.07% and 0.91%, 99.96% and 1.69%, respectively (Table 3), which is the best in detecting hidden features in FSV voltammograms. Interestingly, according to a Poisson distribution, 30% of the 0.2 amol L−1 sample (positive) should contain 0 copper ion, that is, the signals should be negative. However, the accuracy of the deep learning is high up to >99%, implying that the distribution of ions in solution at or close to single-atom level does not completely follow a Poisson distribution, which describes the probability of completely random events. In fact, the distribution of ions in a solution is not completely random but is strongly influenced by the movements of their own and surrounding solvent molecules. In this case, the randomness is greatly reduced, and the distribution is greatly averaged.
Considering that this image classification is to distinguish nominally positive signal images for single copper ion and negative ones for the blank, ablation experiments were carried out to compare the effectiveness of the model. Specifically, different activation functions such as tanh, ReLU, and Leaky ReLU were chosen for nonlinear transformation processing, convolutional bases with different numbers and convolutional kernels with different sizes were selected to optimize the model and training parameters, max pooling and average pooling were adopted to achieve feature optimization, and batch normalization and dropout were used to enhance the generalization ability and robustness of the model. As shown in Table 4, our FSVNet exhibits the best overall performance in terms of four indicators such as F1-Score, Accuracy, Precision and Recall. The receiver operating characteristic (ROC) curve illustrates that the area under the curve (AUC) is 1.00 (Fig. 5e), also demonstrating that FSVNet effectively discriminates between positive and negative signals.
To further investigate whether this sensor has truly achieved SAD of copper ions, 100 samples at the concentration of 0.1 amol L−1 were tested. It was found that, 56 samples are positive (≥1 ions) while 44 samples are negative (0 ions). Considering that the theoretical average number of Cu2+ in 10 μL of 0.1 amol L−1 sample is 0.6, the observed probability is approximately equal to the theoretical one, demonstrating that SAD of copper ions has indeed been achieved.
Discussion
In summary, a sensitive deep learning-assisted electrochemical sensor was developed, benefitted from the multi-level signal amplification strategy, which is a delicate integration of functionalized two-dimensional nanomaterials, click chemistry, and FSV. Additionally, single-atom detection of copper ions was achieved, based on a custom-built deep learning model FSVNet. It is worth mentioning that Cu+ is the substantial catalyst for the click reaction, which is unstable in water due to disproportionation. In the presence of Cu2+, sodium ascorbate should be introduced to reduce Cu2+ to Cu+ in situ to meet the detection protocol. Therefore, the proposed device is eventually an electrochemical sensor for copper ions, including Cu+ and/or Cu2+. It is also expected that the introduction of deep learning methods will greatly improve the ability to extract weak signals from complex backgrounds, changing the traditional signal identification criterion in analytical chemistry, that is, signal/noise ratio of 3.
Obviously, the current method still needs to increase the number of signal units by click chemistry to improve detection sensitivity, which requires a certain reaction time. Therefore, real-time SAD could not be realized up to now. However, it is optimistic that the targets of this SAD method can be extended to single molecule enzymes or single atom/particle catalysts with catalytic amplification capability without the need for real-time monitoring. For example, single Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-associated protein or DNA ligase protein could be detected by the aid of their cleavage or ligation of DNA/RNA sequences, and single atom/particle catalysts could be traced with their help towards breaking and forming special chemical bonds.
Methods
Materials and reagents
Black phosphorus nanosheet (BPNS, 99.998%) dispersion was purchased from XFNANO Materials Tech. Co., Ltd. (Nanjing, China). Ascorbic acid (AA, United States Pharmacopeia Reference Standard), copper powder (99.999% trace metals basis), silver nitrate (AgNO3, 99.9999% trace metals basis), polyvinylpyrrolidone (PVP, Mw 10,000 g mol−1), 3-aminopropyltriethoxysilane (APTES, ≥98%), and 3-azidopropylamine (≥95%) were purchased from Merck (Darmstadt, Germany). Propargyl amine (98%) was purchased from Shanghai Macklin Biochemical Technology Co., Ltd. (Shanghai, China). Diethylene glycol (DEG, analytical standard, ≥99%), ethylene glycol (EG, analytical standard, ≥99.9%), glutaraldehyde (50% in water, specialized for electron microscopy), ammonia (NH3·H2O, guaranteed reagent, 25 wt.%) and FeCl3·6H2O (puriss. p.a., reag. Ph. Eur., ≥99%) were purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China). All other chemicals used were of at least analytical grade. Millipore water (18.2 MΩ cm) was used throughout the experiment.
Pretreatment of glassware
To prevent any external contamination, all glassware was pre-cleaned with aqua regia (a chemical mixture of 38 wt.% hydrochloric acid and 69 wt.% nitric acid in a 3:1 volume ratio), thoroughly rinsed with water and distilled ethanol, and well dried prior to use. Since aqua regia is highly corrosive and hazardous, extreme caution should be exercised with appropriate laboratory precautions.
Apparatus
The morphology, element distribution and energy dispersive X-ray spectroscopy (EDS) of various materials were completed by scanning electron microscopy (SEM, Nova NanoSEM450, FEI, USA). Fourier transform infrared (FTIR) spectra were obtained by Nicolet AVATAR FTIR 6700 spectrometer (Woburn, Massachusetts, USA). Electrochemical impedance spectroscopy (EIS) and cyclic voltammetry (CV) were measured at conventional scan rates using CHI 760E electrochemical workstation (CH instrument, Shanghai, China). Zeta potential analysis was performed using Malvern Zana ZS 90 Mastersizer (Malvern, UK). Fast scan voltammetry (FSV) was performed using a home-built electrochemical system, which includes an AFG3021B arbitrary/function generator (Tektronix, Oregon, USA) to generate potential waveforms, a TBS1102 digitizing oscilloscope (Tektronix, Oregon, USA) to acquire voltametric data, and a home-built circuit to provide perfect on-line positive feedback compensation for the ohmic drop in the solution according to our previous works with minor modifications51,52. A traditional three-electrode system was used, including a magnetic glassy carbon electrode (MGCE, 2 mm diameter, GaossUnion, Wuhan, China) as the working electrode, an Ag/AgCl (3 mol L−1 KCl) electrode as the reference electrode, and a platinum wire as the counter electrode.
Preparation of Fe3O4 nanoparticles dispersion
First, 540 mg of FeCl3·6H2O was added into 20 mL of a mixture of DEG and EG (VDEG/VEG = 7/13) and stirred for 30 min. Then, 2 g of PVP was added and stirred in an oil bath at 120 °C until PVP was completely dissolved. Next, 2 g of anhydrous sodium acetate powder was added and stirred at room temperature for 30 min, followed by transferring the mixture into a 100 mL Teflon-lined stainless-steel autoclave and heated at 200 °C for 8 h. After the obtained black precipitate was washed and dispersed in 20 mL of water, the dispersion of magnetic Fe3O4 nanoparticles was obtained.
Preparation of Cu2+ standard solutions
A 0.3177 g of copper powder was accurately weighed, placed in a 250-mL beaker, and covered with a watch glass. Then, 20 mL of HNO3 solution (1:1) was carefully added to the beaker for dissolution. Next, 100 mL of 2.5 mol L−1 H2SO4 solution was added, and the mixture was carefully boiled to remove nitrogen oxides. After cooling, it was transferred to a 500 mL volumetric flask and diluted with water to obtain a 1.000 × 10−2 mol L−1 Cu2+ standard stock solution. Through stepwise dilution, Cu2+ standard solutions with different concentrations were obtained. The blank was prepared in the same manner but in the absence of Cu2+.
Synthesis of signal unit BPNS@AgNPs-Alkyne
Briefly, 0.05 mL of 0.08 mmol L−1 AgNO3 was added into 10 mL of 4 mol L−1 NH3·H2O, followed by adding 10 mL of 0.2 mg mL−1 BPNS and stirring in dark at room temperature for 12 h. After centrifugation at 5249× g and dispersion in 10 mL of water, BPNS@AgNPs dispersion was obtained. Then, 3 mL of 1 mg mL−1 propargyl amine was added and incubated for 12 h. After centrifugation at 5249× g and centrifugation in 1 mL of water, the signal unit BPNS@AgNPs-Alkyne dispersion was finally obtained by immobilizing propargylamine on BPNS@AgNPs through Ag-N bond.
Preparation of Azide-Fe3O4
After a 10 mL of the as-prepared Fe3O4 nanoparticle dispersion and 200 μL of 3-aminopropyltriethoxysilane (APTES) were mixed in 10 mL of ethanol, sonicated for 1 h, stirred for 12 h at room temperature, magnetically separated, washed and dispersed in 10 mL of water, amino-functionalized Fe3O4 nanoparticle dispersion was obtained. Then, 600 μL of 2.5 wt.% glutaraldehyde aqueous solution was added and incubated for 6 h at room temperature, followed by magnetically separating, washing and dispersing in 2 mL of water. Subsequently, 2 mL of 2.5 mg mL−1 3-azidopropylamine was added and stirred for 6 h at room temperature. Finally, Azide-Fe3O4 dispersion was obtained after being magnetically separated, rinsed and dispersed in 2 mL of water.
Construction of electrochemical sensor
A 50 µL of signal unit BPNS@AgNPs-Alkyne dispersion, 50 µL of Azide-Fe3O4 dispersion and 10 μL of sample solution containing known or unknown amount of Cu2+ were mixed. Then, 10 µL of 0.02 mol L−1 ascorbic acid was added and shaken in dark for 40 min to perform the click chemistry reaction. After being magnetically separated, rinsed and dropped, the precipitate was firmly adsorbed onto the surface of MGCE, and the electrochemical sensor was constructed in one step.
Zeta potential measurement
A 0.75 mL of each sample (BPNS, BPNS@AgNPs, propargylamine and BPNS@AgNPs-Alkyne) was added to a clear disposable zeta cell DTS1060C and analyzed using the dynamic light scattering particle size analyzer. The parameters for zeta potential detection were set as follows: refractive index was 2.42 and absorption was 0.01, dispersant was water, temperature was 25 °C, and equilibration time was 90 s.
EIS and CV measurement
EIS and CV were recorded using an electrochemical workstation at room temperature. The parameters for EIS measurement were set as follows: the electrolyte solution was 0.1 mol L−1 KCl containing 5.0 mmol L−1 [Fe(CN)6]3-/4-, frequency range was 0.01 to 10 kHz, and AC amplitude was 5 mV. The parameters for CV measurement were set as follows: the potential scan range was −0.2 to 0.6 V, and the scan rate was 50 mV s−1.
FSV measurement
Parameters of FSV were as follows. Supporting electrolyte solution: 0.5 mol L−1 KNO3; potential window: −1.0−1.5 V; scan rate: 400 V s−1.
Data processing of FSV
The FSV data processing includes three steps: peak current interception, data shifting, and Gaussian fitting (Supplementary Fig. 11). First, the upper part (Supplementary Fig. 11b) of the complete voltammogram of FSV (Supplementary Fig. 11a) was intercepted. Next, the baseline was subtracted from these data points, which is the data shifting operation (Supplementary Fig. 11c). Finally, Gaussian fitting was performed on the translated waveform data to obtain the final peak current curve (Supplementary Fig. 11d).
Experimental temperature
All experiments were performed at 25 ± 1 °C.
Deep learning for SAD of copper ions
(1) Construction of the dataset. A source dataset was created by acquiring 1280 FSV voltammograms of 0.2 amol L−1 Cu2+ and 1632 FSV voltammograms of blank solutions, each voltammogram containing 512 data points and being 5184 ×4575 pixels in size. It was randomly partitioned into training, testing, and validation sets in a ratio of 6:3:1, with the training set to train classification models to obtain efficient image classifiers, the validation set to adjust the hyperparameters of the model and preliminarily evaluate its ability, and the testing set to further evaluate the generalization ability of the final model. (2) Development of image classifier. Our end-to-end image classifier FSVNet based on deep convolutional neural network for detecting hidden signals in voltammograms was established utilizing Pytorch 2.2.1+cu118. Through multi-layer convolution and pooling operations, potential features and signal patterns of voltammograms were revealed, and then automatic image classification was achieved by a fully connected network. It mainly composed of three parts (Fig. 5b): (i) 8 convolutional layers and ReLU non-linear activation function, (ii) a pooling layer, and (iii) 5 fully connected layers. To standardize the input data and expedite the network training process, batch normalization was performed after each convolution layer and fully connected layer. Additionally, max-pooling and dropout operations were carried out between convolution layers and fully connected layers, thus to enlarge the receptive field, suppress noise, reduce model parameters, enhance the model robustness, and reduce the overfitting risk. (3) Training and optimization of image classifier. Adam optimizer with a learning rate of 0.00001 was chosen for model training and optimization. VGGNet, InceptionNet, ResNet, and DenseNet were selected as the baseline models, and their performance was compared to that of our FSVNet (Supplementary Note 1). They were all trained for 20 epochs with a batch size of 10. Losses were evaluated by the CrossEntropy function L(p,q), which is defined as
where the true probability distribution p(xi) is the label of the training set, and the predicted probability distribution q(xi) is the result predicted by the model. And an early stopping technique was utilized to prevent model overfitting. In addition, the training was terminated if the loss did not decrease in five consecutive epochs. The classification accuracy was defined as
where n* represents the count of accurately classified samples, while Ntotal denotes the count of overall samples. All training was operated using NVIDIA RTX 2060 Super GPU.
Data availability
The data that support the findings of this study are available from Zenodo66 and from the corresponding authors upon request. Source data are provided with this paper.
Code availability
Code and raw data relevant to data analysis in this study is available at GitHub (https://github.com/NBUGUOzy/FSVNet) and Zenodo66.
References
Alkemade, C. & Th, J. Single-atom detection. Appl. Spectrosc. 35, 1–14 (1981).
Elmar, H. et al. Single-atom imaging of fermions in a quantum-gas microscope. Nat. Phys. 11, 738–742 (2015).
Li, H. Trapping atoms of krypton-81 to date groundwater and ice cores. Nat. Rev. Earth Environ. 5, 231 (2024).
Legrand, T. et al. Three-dimensional imaging of single atoms in an optical lattice via helical point-spread-function engineering. Phys. Rev. A 109, 033304 (2024).
Ahmad, M. et al. A generalizable nanopore sensor for highly specific protein detection at single-molecule precision. Nat. Commun. 14, 1374 (2023).
Shimizu, K. et al. De novo design of a nanopore for single-molecule detection that incorporates a β-hairpin peptide. Nat. Nanotechnol. 17, 67–75 (2021).
Zevenbergen, M. A. G., Singh, P. S., Goluch, E. D., Wolfrum, B. L. & Lemay, S. G. Stochastic sensing of single molecules in a nanofluidic electrochemical device. Nano Lett. 11, 2881–2886 (2011).
Kang, S., Nieuwenhuis, A. F., Mathwig, K., Mampallil, D. & Lemay, S. G. Electrochemical single-molecule detection in aqueous solution using self-aligned nanogap transducers. ACS Nano 7, 10931–10937 (2013).
Farka, Z. et al. Advances in optical single-molecule detection: En route to supersensitive bioaffinity assays. Angew. Chem. Int. Ed. 59, 10746–10773 (2020).
Mitchell, S. et al. Automated image analysis for single-atom detection in catalytic materials by transmission electron microscopy. J. Am. Chem. Soc. 144, 8018–8029 (2022).
Gormal, R. S. et al. Modular transient nanoclustering of activated β2-adrenergic receptors revealed by single-molecule tracking of conformation-specific nanobodies. Proc. Natl. Acad. Sci. USA. 117, 30476–30487 (2020).
Deslignière, E., Rolland, A., Ebberink, E. H. T. M., Yin, V. & Heck, A. J. R. Orbitrap-based mass and charge analysis of single molecules. Acc. Chem. Res. 56, 1458–1468 (2023).
Su, Z., Li, T., Wu, D., Wu, Y. & Li, G. Recent progress on single-molecule detection technologies for food safety. J. Agric. Food Chem. 70, 458–469 (2022).
Choi, H. K. et al. Single-molecule surface-enhanced Raman scattering as a probe of single-molecule surface reactions: Promises and current challenges. Acc. Chem. Res. 52, 3008–3017 (2019).
Hurst, G. S., Payne, M. G., Kramer, S. D. & Young, J. P. Resonance ionization spectroscopy and one-atom detection. Rev. Mod. Phys. 51, 767–819 (1979).
Ruchkina, M., Raveesh, M., Dominguez, A., Bood, J. & Brackmann, C. Simultaneous single-shot imaging of H and O atoms in premixed turbulent flames using femtosecond two-photon laser-induced fluorescence. Opt. Express 31, 12932–12943 (2023).
Xia, T. Y. et al. Atom-trap trace analysis of 41Ca/Ca down to the 10–17 level. Nat. Phys. 19, 904–908 (2023).
Inavalli, V. V. G. K. et al. A super-resolution platform for correlative live single-molecule imaging and STED microscopy. Nat. Methods 16, 1263–1268 (2019).
Wu, H., Li, G., Hou, J. & Sotthewes, K. Probing surface properties of organic molecular layers by scanning tunneling microscopy. Adv. Colloid Interface Sci. 318, 102956 (2023).
Chen, X. et al. Modern scattering-type scanning near-field optical microscopy for advanced material research. Adv. Mater. 31, 1804774 (2019).
Fang, S. & Hu, Y. H. Open the door to the atomic world by single-molecule atomic force microscopy. Matter-US 4, 1189–1223 (2021).
Fan, F. R. F., Kwak, J. & Bard, A. J. Single molecule electrochemistry. J. Am. Chem. Soc. 118, 9669–9675 (1996).
Hassenkam, T. et al. Self-assembly and conductive properties of molecularly linked gold nanowires. Nano Lett. 4, 19–22 (2004).
Zhu, J. et al. Solution-processable carbon nanoelectrodes for single-molecule investigations. J. Am. Chem. Soc. 138, 2905–2908 (2016).
Li, G. X. & Lin, X. Q. A glass nanopore electrode for single molecule detection. Chin. Chem. Lett. 21, 1115–1118 (2010).
Morikawa, T., Yokota, K., Tsutsui, M. & Taniguchi, M. Fast and low-noise tunnelling current measurements for single-molecule detection in an electrolyte solution using insulator-protected nanoelectrodes. Nanoscale 9, 4076–4081 (2017).
He, Q. & Tang, L. Sub-5 nm nanogap electrodes towards single-molecular biosensing. Biosens. Bioelectron. 213, 114486 (2022).
Ghimire, G. et al. Potentiometric and SERS detection of single nanoparticle collision events on a surface functionalized gold nanoelectrode. J. Electrochem. Soc. 169, 047511 (2022).
Puthongkham, P. & Venton, B. J. Recent advances in fast-scan cyclic voltammetry. Analyst 145, 1087–1102 (2020).
Amatore, C., Bouret, Y., Maisonhaute, E., Abruña, H. D. & Goldsmith, J. I. Electrochemistry within molecules using ultrafast cyclic voltammetry. C. R. Chim. 6, 99–115 (2003).
Liu, Y., Liu, Y., Qiao, L., Liu, Y. & Liu, B. Advances in signal amplification strategies for electrochemical biosensing. Curr. Opin. Electrochem. 12, 5–12 (2018).
Meldal, M. & Diness, F. Recent fascinating aspects of the CuAAC click reaction. Trends Chem. 2, 569–584 (2020).
Edward Sekhosana, K., Majeed, S. A. & Feleni, U. Click chemistry in the electrochemical systems: Toward the architecture of electrochemical (bio)sensors. Coord. Chem. Rev. 491, 215232 (2023).
Lu, J. et al. Functionalization of covalent organic frameworks with DNA via covalent modification and the application to exosomes detection. Anal. Chem. 94, 5055–5061 (2022).
Fu, K., Kwon, S. R., Han, D. & Bohn, P. W. Single entity electrochemistry in nanopore electrode arrays: Ion transport meets electron transfer in confined geometries. Acc. Chem. Res. 53, 719–728 (2020).
Guo, K. et al. Rapid single-molecule detection of COVID-19 and MERS antigens via nanobody-functionalized organic electrochemical transistors. Nat. Biomed. Eng. 5, 666–677 (2021).
Wu, M. & Lu, H. P. Ultra-sensitive lock-in amplifier coupled oscillatory magnetic tweezers for piconewton force manipulation applications. J. Appl. Phys. 130, 014504 (2021).
Ganjalizadeh, V. et al. Fast custom wavelet analysis technique for single molecule detection and identification. Nat. Commun. 13, 1035 (2022).
Cho, S. Y. et al. Finding hidden signals in chemical sensors using deep learning. Anal. Chem. 92, 6529–6537 (2020).
Speiser, A. et al. Deep learning enables fast and dense single-molecule localization with high accuracy. Nat. Methods 18, 1082–1090 (2021).
Shao, S., Yan, R., Lu, Y., Wang, P. & Gao, R. X. DCNN-based multi-signal induction motor fault diagnosis. IEEE Trans. Instrum. Meas. 69, 2658–2669 (2020).
Gao, Z. et al. Complex networks and deep learning for EEG signal analysis. Cogn. Neurodynamics 15, 369–388 (2021).
Yang, Y. et al. Rapid detection of SARS-CoV‑2 RNA in human nasopharyngeal specimens using surface-enhanced Raman spectroscopy and deep learning algorithms. ACS Sens. 8, 297–307 (2023).
Noreldeen, H. A. A. et al. Deep learning-based sensor array: 3D fluorescence spectra of gold nanoclusters for qualitative and quantitative analysis of vitamin B6 derivatives. Anal. Chem. 94, 9287–9296 (2022).
Ma, Y. P., Li, Q., Luo, J. B., Huang, C. Z. & Zhou, J. Weak reaction scatterometry of plasmonic resonance light scattering with machine learning. Anal. Chem. 93, 12131–12138 (2021).
Wang, G. et al. Mechanistic investigation into single-electron oxidative addition of single-atom Cu(I)-N4 site: Revealing the Cu(I)-Cu(II)-Cu(I) catalytic cycle in photochemical hydrophosphinylation. J. Am. Chem. Soc. 146, 8668–8676 (2024).
Zhou, H. et al. Tunneling or Hopping? A direct electrochemical observation of electron transfer in DNA. Anal. Chem. 94, 15324–15331 (2022).
Lv, J., Tang, Y., Tang, D., Zhang, J. & Tang, D. Target-induced click conjugation for attomolar electronic monitoring of Cu(II) using horseradish peroxidase as indicator and nanogold particles as enhancer. Anal. Methods 9, 117–123 (2017).
Zhou, M., Dick, J. E. & Bard, A. J. Electrodeposition of isolated platinum atoms and clusters on bismuth-characterization and electrocatalysis. J. Am. Chem. Soc. 139, 17677–17682 (2017).
Ren, H. & Edwards, M. A. Stochasticity in single-entity electrochemistry. Curr. Opin. Electrochem. 25, 100632 (2021).
Chen, G. et al. A portable digital-control electrochemical system with automatic ohmic drop compensation for fast scan voltammetry and its application to ultrasensitive detection of chromium(III). Sens. Actuators B 301, 127135 (2019).
Xiao, F. et al. A fast scan cyclic voltammetric digital circuit with precise ohmic drop compensation by online measuring solution resistance and its biosensing application. Anal. Chim. Acta 1175, 338744 (2021).
Aydin, D., Karuk Elmas, S. N. & Arslan, F. N. Ultrasensitive detection of Cu2+: A cyanobiphenyl-based colorimetric and fluorescence chemosensor and its smartphone and food sample applications. Food Chem. 402, 134439 (2023).
Hou, J. et al. Fluorescence and colorimetric dual-mode ratiometric sensor based on Zr-tetraphenylporphyrin tetrasulfonic acid hydrate metal-organic frameworks for visual detection of copper ions. ACS Appl. Mater. Interfaces 14, 13848–13857 (2022).
Gao, H. et al. In situ growth visualization nanochannel membrane for ultrasensitive copper ion detection under the electric field enrichment. ACS Appl. Mater. Interfaces 12, 4849–4858 (2020).
Xie, S. et al. AIE-active metal-organic frameworks: Facile preparation, tunable light emission, ultrasensitive sensing of copper(II) and visual fluorescence detection of glucose. J. Mater. Chem. C 8, 10408–10415 (2020).
He, L. et al. Hydrophobic plasmonic silver membrane as SERS-active catcher for rapid and ultrasensitive Cu(II) detection. J. Hazard. Mater. 440, 129731 (2022).
Liu, Y., Wu, Y., Guo, X., Wen, Y. & Yang, H. Rapid and selective detection of trace Cu2+ by accumulation-reaction-based Raman spectroscopy. Sens. Actuators B 283, 278–283 (2019).
Ma, Y., Li, M., Li, Z. & Zhao, M. Light regulated heterojunctions with tunable interfacial energy barriers for sensitive and specific detection of copper ions. Chem. Eng. J. 431, 133880 (2022).
Li, J. et al. Ligand reduction and cation exchange on nanostructures for an elegant design of copper ions photoelectrochemical sensing. Sens. Actuators B 328, 129032 (2021).
Chen, J., Zhao, G. C., Wei, Y., Feng, D. & Zhang, H. Construction of a novel photoelectrochemical sensor for detecting trace amount of copper (II) ion. Electrochim. Acta 370, 137736 (2021).
Han, Q. et al. Multifunctional zinc oxide promotes electrochemiluminescence of porphyrin aggregates for ultrasensitive detection of copper ion. Anal. Chem. 92, 3324–3331 (2020).
Zhou, Y., Shi, X., Ma, L., Chai, Y. & Yuan, R. Thiol stabilized Au nanoclusters with efficient electrochemiluminescence for the simultaneous and ultrasensitive detection of GSH and Cu2+. Sens. Actuators B 382, 133518 (2023).
Qing, M. et al. Click chemistry reaction-triggered 3D DNA walking machine for sensitive electrochemical detection of copper ion. Anal. Chem. 90, 11439–11445 (2018).
Hui, X. et al. High-performance flexible electrochemical heavy metal sensor based on layer-by-layer assembly of Ti3C2Tx/MWNTs nanocomposites for noninvasive detection of copper and zinc ions in human biofluids. ACS Appl. Mater. Interfaces 12, 48928–48937 (2020).
Hao, T. FSVNet [Data set], Zenodo, https://doi.org/10.5281/zenodo.14027319 (2024).
Acknowledgements
We gratefully acknowledge the financial support from National Natural Science Foundation of China (82273681 and 42076193 to Z. Guo; 22374087 to P. Gai).
Author information
Authors and Affiliations
Contributions
T.H. and H.Z. conducted the FSV tests, analyzed the data and wrote the draft. Z.W. and Y.G. planned and performed deep learning and analyzed the relative data. H.L. and W.W. conducted material characterization. P.G. and Z.G. conceived the project, designed the experiments, supervised the study and revised the manuscript. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Manoj Kunbhakar and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hao, T., Zhou, H., Gai, P. et al. Deep learning-assisted single-atom detection of copper ions by combining click chemistry and fast scan voltammetry. Nat Commun 15, 10292 (2024). https://doi.org/10.1038/s41467-024-54743-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-54743-8
This article is cited by
-
Rational Green Synthesis of Eu MOF for Ratiometric Fluorescence Sensing Cu2+
Journal of Fluorescence (2025)
-
The roadmap of carbon-based single-atom catalysts: rational design and electrochemical applications
Rare Metals (2025)
-
15 Years of Progress on Transition Metal-Based Electrocatalysts for Microbial Electrochemical Hydrogen Production: From Nanoscale Design to Macroscale Application
Nano-Micro Letters (2025)