Abstract
Mitochondrial morphology and function are intrinsically linked, indicating the opportunity to predict functions by analyzing morphological features in live-cell imaging. Herein, we introduce MoDL, a deep learning algorithm for mitochondrial image segmentation and function prediction. Trained on a dataset of 20,000 manually labeled mitochondria from super-resolution (SR) images, MoDL achieves superior segmentation accuracy, enabling comprehensive morphological analysis. Furthermore, MoDL predicts mitochondrial functions by employing an ensemble learning strategy, powered by an extended training dataset of over 100,000 SR images, each annotated with functional data from biochemical assays. By leveraging this large dataset alongside data fine-tuning and retraining, MoDL demonstrates the ability to precisely predict functions of heterogeneous mitochondria from unseen cell types through small sample size training. Our results highlight the MoDL’s potential to significantly impact mitochondrial research and drug discovery, illustrating its utility in exploring the complex relationship between mitochondrial form and function within a wide range of biological contexts.
Similar content being viewed by others
Introduction
Mitochondria, as essential organelles within cells, play crucial roles in energy production, substance metabolism, and cell apoptosis1,2. The interplay between mitochondrial morphology and functionality is a pivotal aspect of cellular bioenergetics3. The shape and structure of mitochondria are not static but change in response to the cell’s metabolic demands and stress signals4. These morphological changes are not merely consequences of cellular state but actively contribute to mitochondrial functions (i.e., morphofunction)5. For example, elongated mitochondrial networks are efficient at adenosine triphosphate (ATP) production and are associated with a high-energy state6,7, whereas fragmented mitochondria are often found in cells that are undergoing mitophagy8,9. Understanding this relationship is crucial for unraveling the complexities of cellular function and dysfunction. However, the identification of the intricate network structure of mitochondria and the quantification of morphological features have always presented challenges10.
Fluorescence imaging technology has proven effective for studying mitochondrial morphology, enabling real-time visualization of information such as shape, position, and quantity without the need of cell fixation11. Nevertheless, manual analysis of mitochondrial morphology remains labor-intensive and requires significant expertize12. Therefore, it is of great importance to provide an efficient and convenient method to assist researchers in quantifying and automating the analysis of mitochondrial morphology. Methods for efficient measurement of mitochondrial morphology have been established based on image segmentation10. Threshold-based methods have emerged as excellent and user-friendly options for the automated quantification of various morphological features13,14,15. However, these methods often struggle to achieve smooth and continuous segmentation of intricate mitochondrial contours13,14,15. The utilization of deep learning approaches for segmenting target images and defining morphological boundaries prior to analysis has shown promise16,17,18,19,20. Nevertheless, the challenge in collecting representative ground truth training data, compounded by the mitochondrial diameter nearing the optical diffraction resolution limit (~200 nm)21, often results in deep learning-based models struggling with effective feature extraction for segmentation.
In addition to mitochondrial image segmentation, several existing algorithms have realized mitochondrial image analysis. MitoMo leverages machine learning for categorizing mitochondria in healthy and damaged A549 cells22. MitoHacker predicts the DRP1 genotype based on morphology with an accuracy of 87%23. Additionally, Mitometer has employed random forest algorithm to classify mitochondria in triple-negative breast cancer cells and their receptor-positive counterparts15. Despite these technological strides, all these approaches are limited to cellular levels outcomes such as cell fates or classification. Therefore, a notable deficiency in comprehensive and precise prediction of mitochondrial functionality grounded in morphological attributes remains. This lacuna underscores a significant opportunity in the field, where the development of methodologies that intricately links mitochondrial morphology with functions could provide profound insights into cell biology and pathophysiology.
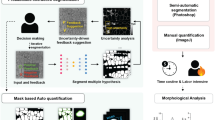
To achieve precise mitochondrial segmentation and function prediction in live-cell images, we have developed MoDL, a deep learning-based software package. MoDL possesses three key pipelines: (1) Trained on over 20,000 manually labeled mitochondria from SR images, MoDL achieves high-precision segmentation of mitochondrial contours from live-cell fluorescence images. This framework outperforms existing methods in delineating mitochondrial morphological features and is adaptable to diverse imaging platforms and cell types (Fig. 1a); (2) Based on high-quality segmentation and morphological features, MoDL can accurate prediction on various mitochondria functions by employing an ensemble learning algorithm. This pipeline is powered by an extended dataset contains over 100,000 SR images, each annotated with corresponding functional data obtained through biochemical assays (Fig. 1b); (3) MoDL demonstrates the ability to precisely predict functions of mitochondria from unseen cell types through small sample size training. The robustness and generalization of MoDL have been further verified by a drug-resistant cancer cell line that has an abnormal pattern of mitochondrial morphological features. (Fig. 1c).
a The first key pipeline of MoDL employs deep learning algorithms for precise and high-quality segmentation of mitochondrial fluorescence images. MoDL was trained on a set of over 20,000 independently manual annotations, derived from original fluorescence images obtained by SR microscopy. b The second key pipeline of MoDL centers on the regression analysis to accurately predict mitochondrial functions, including the mitochondrial membrane potential (MMP) polarization, respiration rate, reactive oxygen species (ROS) production, adenosine triphosphate (ATP) generation, and mitophagy level. This process was enhanced by a dataset for a training ensemble learning algorithm that comprising over 100,000 images and annotated functionalities. c The third key pipeline of MoDL, employing data fine-tuning and retraining alongside a substantial dataset, enables the precise prediction of mitochondrial functions in drug-resistant cancer cells, which exhibit abnormal morphology patterns, with small sample size training (created with BioRender.com).
By using MoDL, we have successfully predicted the different trends of mitochondrial damages induced by mitochondrial targeting or non-targeting cytotoxic reagents, including mitochondrial membrane potential (MMP) polarization, respiration rate, reactive oxygen species (ROS) production, ATP generation, and mitophagy level. Notably, by employing data fine-tuning and retraining, with a small training set, MoDL has revealed an abnormal pattern of mitochondrial morphology in drug-resistant cancer cells, and accurately predicted mitochondrial functions upon cisplatin intervention. Together, MoDL offers a powerful tool for mitochondrial morphology analysis, enabling precise investigation of the relationship between mitochondrial morphology and function.
Results
MoDL’s first pipeline: high-quality mitochondrial segmentation
Mitochondria structures are highly dynamic and have irregular shapes, with the diameter approaching the optical resolution limit (~200 nm)24,25. Accurate localization of mitochondria at this scale requires defining morphological boundaries with single-pixel precision. This process, known as segmentation, constitutes a critical first step in image analysis pipelines26,27. However, conventional optical imaging tools often fail to capture the precise shape of mitochondria, including intricate branching networks and ridges. This limitation makes acquiring high-quality mitochondrial segmentation a significant challenge. Therefore, structured illumination microscopy (SIM, resolution of 115 nm)28 was utilized to gather an extensive SR dataset for training. This approach enabling effectively augments model robustness and generalizability29. Next, we manually annotated approximately 20,000 individual mitochondria to obtain a high-quality training set with desirable characteristics (Supplementary Note 1 and Supplementary Fig. 1). The annotated cell lines include 143B (human osteosarcoma cells), HeLa (human cervix cancer cells), U87 (human astrocytoma cells), HepG2 and Hep3B (human carcinoma cells), L02 (human normal liver cells), A2780 (human ovarian cancer cells), and MCF7 (human breast cancer cells, Supplementary Table 1). As the gold standard in the field, this manual annotation of SR mitochondrial images ensures the accuracy of feature recognition and generates high-quality segmentation of mitochondrial fluorescence images in living cells (Fig. 1a).
The foundational framework of the MoDL image segmentation is based on U-Net30. To effectively extract deeper image features and alleviate issues of gradient vanishing and exploding, we have introduced residual networks (ResNets) after each convolutional layer in U-Net31. Specifically, the introduced residual units consist of an improved convolutional residual block and two identical residual blocks. In the convolutional residual block, both the main and the side paths are replaced with a combination of convolutional and max pooling layers with a stride of two (Supplementary Fig. 2a, b). Simultaneously, to prevent an increase in skip connections span caused by the growing depth of the architecture, we have introduced convolutional block attention modules (CBAM) after the outermost skip connection of the framework (Supplementary Fig. 2c). This enhancement enables the model to focus on crucial information from a widely array of feature maps in both spatial and channel dimensions. For spatial attention, feature maps from global average and max pooling are merged and processed through a convolutional layer to produce a spatial attention map. For channel attention, global average, and max pooling are applied to each channel, with the resulting descriptors transformed through fully connected layers to obtain attention weights. These mechanisms help enhance important features and suppress less important ones, thereby avoiding potential errors when concatenating features with the maximum spatial size in the outermost skip connection. The final model consists of a total of 104 convolutional layers and 16 max-pooling layers (Supplementary Note 2). In summary, the incorporation of improved ResNets and CBAM has significantly enhanced the model architecture (Supplementary Fig. 2d and Supplementary Table 2). To reduce the computational load, during the model training, we initially decomposed 1 original stack (2048 × 2048 pixel2 resolution) into 16 patches (512 × 512 pixel2 resolution) for training, later reassembling these segmentation patches to reconstruct a complete image. Due to the potential loss of edge information in the patches after being segmented by the model, we utilized stacking and overlapping method to reconstruct this missing edge information, generating a seamless and precise high-quality mitochondria segmentation image (Supplementary Note 3 and Supplementary Fig. 3).
To assess MoDL’s performance, we compare several of these open-source algorithms in the same stack. For the quantitative evaluation, we employed a comprehensive metrics including the Dice coefficient, mean Intersection over Union (mIoU), and Pixel Accuracy (PA), along with the extracted morphological features (mean area, form factor, and branch length, Supplementary Note 4). The results showed that retraining with our SR dataset has improved performance of other deep learning-based algorithms: U-Net30 (Dice, mIoU, PA = 0.85, 0.80, 0.92, respectively), MitoSegNet20 (0.83, 0.71, 0.90), StarDist17 (0.80, 0.73, 0.91). However, MoDL (Dice, mIoU, PA = 0.92, 0.84, and 0.95, respectively) still significantly outperformed them (Fig. 2b and Supplementary Table 3). The threshold methods, including Mitometer15 (0.76, 0.63, 0.84), and other competitors also underperformed compared to MoDL (Fig. 2b and Supplementary Table 3). Meanwhile, the extracted morphological features from MoDL showed no statistical difference compared to the ground truth (P > 0.05). Similar performance was observed only with U-Net30 and MitoSegNet20 after retraining with SR dataset (Supplementary Table 3). Subsequently, for visualizing MoDL segmentation, we generated a pseudo-color image reflecting the fluorescence intensity of each mitochondrion by multiplying the original stack with the mask (Fig. 2c and Supplementary Note 5).
a Schematic diagram showing super resolution-structured illumination microscopy (SR-SIM) imaging and the key steps of MoDL in mitochondrial segmentation (created with BioRender.com). b Segmentation performance (Dice coefficient, mIoU, and PA) comparison of MoDL (red dot), open-source deep learning algorithms that are retrained with SR dataset, i.e., U-Net30, MitoSegNet20, Stardist17, and Otsu’s threshold-based Mitometer (green, cyan, blue, and orange dots, respectively)15. n = 200 images (512 × 512 pixel2 resolution). c The segmentation results by MoDL. The original images were first predicted into binary masks of segmentation images and then multiplied by the fluorescence intensity in the original to obtain a pseudo-color image that reflects individual mitochondrion fluorescence intensity. n = 30 images (2048 × 2048 pixel2 resolution), scale bar = 5 μm. Zoomed-in the original and pseudo-color images of mitochondria in two separate cells (i and ii), scale bar = 1 μm. d A comparison between the ground truth and the results of five segmentation algorithms (512 × 512 pixel2 resolution). e Pseudo-color images (1024 × 1024 pixel2 resolution) generated by MoDL for mitochondrial segmentation in cell lines that are included (HeLa, 143B, L02) and excluded (PC9, A549, PC12) in the dataset, scale bar = 2 μm. The Dice coefficient and relative fluorescence intensity of original image were assigned in the pseudo-color. n = 28 (HeLa), 20 (143B), 27 (L02), 13 (PC9), 22 (A549), 16 (PC12) images (512 × 512 pixel2 resolution). f Fluorescence imaging data obtained from SIM and confocal under the same viewing field, scale bar = 2 μm. g Quantitative assessment of mitochondrial morphological features (mean area, form factor, branch length) between SIM and confocal images employing MoDL (blue dot) and an adaptive threshold method (orange dot). n = 60 images (512 × 512 pixel2 resolution). Data are given as the mean ± SD (b, g). Statistical differences were calculated using a one-way ANOVA followed by Dunnett’s multiple comparison test, P < 0.0001 vs. MoDL (b), or a two-tailed Student’s t-test (g). No significant (ns, P > 0.05). Source data are provided as a Source Data file.
Next, we compared the segmentation outputs from MoDL and reported algorithms with the ground truth. MoDL provides smoother and more continuous segmentation of individual mitochondrion, effectively removing noise while preserving authentic details (Fig. 2d). Moreover, MoDL successfully disentangled two tightly juxtaposed, yet distinct, mitochondrial entities (Supplementary Fig. 2e). In contrast, Mitochondria Analyzer13 and MitoGraph14 failed to identify most of the mitochondria. This may be the thresholding method’s inability to adapt to the complex and diverse structural of mitochondria morphology (Supplementary Fig. 4). On the other hand, deep learning algorithms, such as U-Net30, MitoSegNet20, and StarDist17, that are retrained and fine-tuned using our SR dataset, could identify most of the mitochondria. However, the retention of true mitochondria contours and segmentation continuity of their outputs were weaker than that achieved by MoDL (Fig. 2d). We also evaluated the contribution of SR dataset on the performance of MoDL by using confocal images as the training dataset. The results demonstrated that SR dataset-trained MoDL performed superior performance than confocal dataset-trained MoDL in terms of Dice, mIoU, and PA (Supplementary Table 4). These results further support the rationale and necessity of the construction of the SR dataset for algorithm training. In general, MoDL represents the current state-of-the-art algorithm in terms of both performance and mitochondrial segmentation quality.
To validate the segmentation performance of MoDL, we conducted tests assessing its robustness and generalizability, along with evaluating its efficacy in mitochondrial morphology tracking in living cells. In the robustness test, we subjected the original images to various blur and noise models, quantifying the resulting morphological features and comparing them with ground truth (Supplementary Fig. 5a). Our results demonstrated that significant differences (P < 0.05) were observed only under conditions of exceptionally strong interference (Supplementary Fig. 5b).
Subsequently, to evaluate MoDL’s generalization capability across different cell lines, we tested cell lines that are included (HeLa, 143B, L02) and excluded (PC9, human lung cancer cells; A549, adenocarcinoma human alveolar basal epithelial cells; and PC12, rat adrenal pheochromocytoma cells) in the training data. The results demonstrated that MoDL maintained excellent segmentation performance on all the tested cell lines, exhibiting high-quality segmentation with smooth contours and clear shapes, as well as high Dice coefficients, mIoU, and PA (Fig. 2e and Supplementary Table 5). Furthermore, we evaluated the fluorescence intensity of each segmentation image through pseudo-color restoration, enabling assessment of the loading capacity of mitochondria trackers across diverse cell lines (Supplementary Fig. 6). Next, we compared MoDL’s generalization to process fluorescence data from different imaging platforms, including SIM and confocal microscopy (Fig. 2f and Supplementary Fig. 7). The results indicated that MoDL displayed very high robustness and compatibility, with no significance between the morphological features extracted from the two-microscope data (P > 0.05), along with high Dice, mIoU and PA (SIM vs. Confocal = 0.92 vs. 0.88, 0.84 vs. 0.80, 0.95 vs. 0.93, Fig. 2g, Supplementary Table 6). In contrast, an adaptive thresholding method showed performance discrepancies between the two groups (Fig. 2g, Supplementary Note 6, and Supplementary Table 6). Additionally, we conducted mean absolute percentage error (MAPE) comparisons of morphological features for individual mitochondrion instances obtained from SIM and confocal against the ground truth (Supplementary Table 6). The results provided strong evidence that MoDL can be effectively applied to precisely segment mitochondrial images taken from conventional resolution imaging systems. Moreover, to further evaluated MoDL’s generalization, we segmented images that were captured with various the microscopic resolutions (2048 × 2048 vs. 1024 × 1024 pixel2 resolution) and object lens magnifications (100× vs. 60×). The results showed that MoDL’s remained stable across different configuration parameters, with minimal fluctuations in Dice coefficient, mIoU, PA, and low MAPE in the morphological metrics of individual mitochondrion instances (Supplementary Table 6). Together, these results underscored the high robustness and generalization capabilities of MoDL in mitochondria segmentation, highlighting its potential for broad utility in mitochondria morphology analysis.
Mitochondria morphology analysis based on MoDL segmentation
To validate the ability of MoDL in monitoring mitochondrial morphologies under different conditions, we first introduced mitophagy by carbonyl cyanide 3-chlorophenylhydrazone (CCCP, an oxidative phosphorylation uncoupler)32, followed by inhibiting mitochondrial fission via Mdivi-1 (a DRP1 inhibitor) treatment (Supplementary Fig. 8a)33. MoDL segmentation showed that with increasing incubation time of CCCP, the overall mitochondrial morphology gradually transformed from highly elongated structures to sparsely fused tubules and spheres (Supplementary Fig. 8b). After that, with the addition of Mdivi-1, mitochondria exhibited a lengthening trend, accompanied by an increase in branching and network complexity (Supplementary Fig. 8c). Simultaneously, we calculated the overlap of original and segmented images, indicating their high correlation (Pearson’s r = 0.84, Supplementary Fig 8d). This analysis affirms MoDL’s capability for detail assessment of mitochondria, including challenging features such as ridges.
Next, to further visually depict and quantitatively assess changes in mitochondrial morphology under different statuses32,34, varying concentrations of CCCP-treated cells were imaged and segmented by MoDL (Fig. 3a, b). However, due to the intricate and diverse nature of mitochondrial morphology, connecting mitochondrial functionalities solely on visual observation or manual analysis of each single mitochondrion is challenging. Based on the MoDL segmentation results, we extracted 31 morphological features for individual mitochondrion and over 100 quantitative metrics (comprising average, median, and standard deviation for each image, Supplementary Table 8), offering a broader set than current algorithms (Supplementary Table 9).
a A schematic diagram illustrating the establishment of cellular models with mitochondrial dysfunctional through carbonyl cyanide 3-chlorophenylhydrazone (CCCP) intervention. The study explored the relationship between mitochondrial functions (MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate) and morphological features (created with BioRender.com). b Pseudo-color images obtained through MoDL under CCCP treatment with different concentrations (0, 1, 5, 10, 20 µM, Time = 4 h) in HepG2 cells, showing intuitive changes in mitochondrial morphology. n = 16 images (2048 × 2048 pixel2 resolution), scale bars = 5 μm. c Comparisons of mitochondrial morphological features analyzed by MoDL. Statistical differences were calculated using a one-way ANOVA followed by Dunnett’s multiple comparison test, CCCP treatment (0, 1, 5, 10, and 20 µM) groups, respectively, vs. control. Pearson’s correlation coefficient (r) between feature changes and CCCP concentration (0, 1, 5, 10, and 20 µM). n = 2763 mitochondria. (1–10) area, eccentricity, equivalent diameter, Euler’s number, extent, major axis, minor axis, orientation, perimeter, solidity, (11–20) centroid x, centroid y, distance, weighted cent x, weighted cent y, weighted distance, form factor, roundness, branch count, total branch length, (21–31) mean branch length, median branch length, std branch length, mean branch angle, median branch angle, std branch angle, total density, average density, median density, aspect ratio, fluorescence intensity. d. Mitochondrial functions of HeLa, 143B, HepG2, L02, and U87 (green, blue, red, purple, and orange lines, respectively) cells under CCCP treatment (0, 1, 5, 10, 20 µM, Time = 4 h), n = 3 independent experiments, data are given as the mean ± SD. The original experimental biochemical results are provided in Supplementary Data 1. e Gene expression levels, including fission-related (DRP1, FIS1 and MIEF1), fusion-related (MFN1 and OPA1), mitophagy-related (PINK1 and BNIP3), and biosynthesis-related (SIRT1), in HeLa, 143B, L02, HepG2, and U87 cells under CCCP treatment (0, 1, 5, 10, 20 µM, Time = 4 h). n = 3. Source data are provided as a Source Data file.
Based on the comprehensive analysis, we observed a high correlation between the concentration of CCCP and various features, such as area (1), perimeter (9), and equivalent diameter (3, Fig. 3c), indicating a noticeable decrease in the size of individual mitochondrion. Positive correlation between CCCP concentration and other features like extent (5), solidity (10), and roundness (18) has also been observed, suggesting a shape shift from elongated to circular forms and toneless morphology. Furthermore, eccentricity (2), and branch count (19) length (20–23) showed a strong inverse correlation, implying a decline in mitochondrial network complexity and a trend towards uniformity. Additionally, fluorescence intensity (31) shows a negative correlation with CCCP. This may be due to mitochondrial dysfunction leading to the MMP depolarization, which reduces dye accumulation, and consequently lowers intensity35. However, several features such as orientation (8), centroid (11–13), and weighted distance (14–16) exhibited no significant connection to mitochondrial damage. This could be attributed to the mitochondrial layout and orientation changing with dysfunction, but still predominantly oriented around the nucleus. These analytical results uncovered distinct patterns of mitochondrial morphology upon the treatment of CCCP at different concentrations, indicating that changes in mitochondrial shapes represent key indicators for assessing the degree of dysfunction (Fig. 3c).
We next confirmed the mitochondrial dysfunction introduced by CCCP through a comprehensive range of biochemical assays, including MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate3 (Fig. 3d, Supplementary Note 7, and Supplementary Figs. 9, 10). Subsequently, we conducted analysis of gene expression changes associated with mitochondrial dynamics and biogenesis. Our qRT-PCR results showed that upon the treatment of CCCP, fusion and biogenesis-related genes were downregulated, while fission and mitophagy-related genes were upregulated (Fig. 3e and Supplementary Fig. 11). These results strongly suggested mitochondrial morphology as a hallmark of mitochondrial functions. More importantly, MoDL reveals intricate morphological features that strongly interconnect mitochondrial functions, empowering our next stage of mitochondrial function prediction via MoDL segmentation and analysis.
MoDL’s second pipeline: accurate prediction of mitochondrial functions
Our utilization of MoDL has facilitated the identification of a strong association between mitochondrial morphology and function. Next, we aimed to predict mitochondrial function based on various morphological features extracted by MoDL. Morphological feature-based mitochondrial function prediction, being a significant challenge, is a regression prediction issue for continuous target variables. For dataset preparation, we assembled a comprehensive collection of over 115,405 mitochondrial images (512 × 512 pixel2 resolution) from various cell lines, including HeLa (32,451), 143B (21,338), HepG2 (30,947), U87 (10,777) and L02 (19,892) cells. Each image in the dataset comes annotated with five indicators representative of mitochondrial functions (i.e., MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate, Supplementary Fig. 12). To ensure the precision and robustness of our algorithm, we proposed an ensemble learning approach that integrates image and numerical input (morphology data) in the analysis of mitochondrial morphology. By leveraging various data types instead of relying solely on single-modality information, we have gained a deeper understanding of the intricate relationship between diverse mitochondrial morphology and functions (Fig. 4a).
a A flowchart depicting the ensemble learning pipeline: the boosting algorithm processes the morphology data from numerical input, while the convolution neural network (CNN)+ analyzed mask features from the image input. Their integrated output facilitates precise mitochondrial function prediction (created with BioRender.com). b Correlation (orange dot) and residual (gray dot) analysis of MoDL’s prediction vs. ground truth for relative MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate. A comparison of Pearson’s correlation coefficient (r) is presented. The shaped band in the plot represents the 95% prediction (orange) and confidence (gray) interval. n = 100. c Comparative performance metrics, including mean squared error (MSE), mean absolute error (MAE), and R2 of CNN+, boosting, and MoDL (green, blue, and orange columns, respectively) in mitochondrial function prediction, across 5 cell lines: HeLa (9735), 143B (6401), HepG2 (9284), U87 (3233), and L02 (5967). n = 5 cell lines. d Statistical discrepancies between prediction and ground truth values using MoDL, XGBoost + LightGBM, XGBoost, LightGBM, and CNN in HepG2 cells, across varied training sizes of 30, 20, 10, 5, and 1 k, respectively. P values are displayed in blue for non-significant and in orange for significant differences. Data are given as the 95% prediction and confidence interval (b) or mean ± SD (c). Statistical differences were calculated using a Kruskal-Wallis test followed by Dunn’s multiple comparison test (c) and one-way ANOVA followed by Dunnett’s multiple comparison test (d). No significant (ns, P > 0.05). Source data are provided as a Source Data file.
We adopt the boosting algorithms, including XGBoost (eXtreme Gradient Boosting) and LightGBM (Light Gradient Boosting Machine) fusion, and CNN (Convolution neural network) to train the numerical and image data, respectively (Supplementary Fig. 13 and Supplementary Tables 10, 11). Inspired by the stacking approach36, our architecture involves a sequential learning process. The model comprises multiple layers, each contributes uniquely to the overall model performance. Since selecting initial learners is crucial, we chose XGBoost as the first-layer base learner for its superior handling of numerical features. Regularization prevents overfitting, enhancing base predictions and providing more reliable input for subsequent layers37,38,39. Firstly, we trained a feature set (Feature1-Featuren) using XGBoost-1 (XGB1) to produce predictions, which were then used as new features (Featuren + 1). This new feature, Featuren + 1, was combined with the original feature set and served as input features for LightGBM-1 (LGB1) and LightGBM-2 (LGB2) in the second layer. The predictions from LGB1 were further incorporated as new features (Featuren + 2), acting as input features for XGBoost-2 (XGB2). The XGB1 and LGB1 layers provide foundational feature processing and optimization, enabling the model to extract more representative inputs from the initial raw features. The XGB2 and LGB2 layers further enhance the model’s predictive capability and stability by integrating features generated from the previous layers. Concurrently, the third layer incorporated a CNN+ model, trained on all segment image data, improving the model’s performance in interpreting multi-dimensional data. The predictions from LGB2 in the second layer were weighted together with the predictions from XGB2 and the CNN+ model in the third layer to yield the final fused output (Supplementary Fig. 13). This stacking framework enhanced the performance of MoDL, enabling effective integration and fusion of diverse data types. Ablation experiments revealed that removing any components increased the model’s prediction mean squared error (MSE). Moreover, altering the module sequence and increasing the stacking depth beyond the optimal point led to diminished predictive performance (Supplementary Table 12).
Accounting for the mitochondria heterogeneity inherent to various cell types, we conducted separate trainings for HeLa, 143B, HepG2, U87, and L02 cells in MoDL. Subsequently, we performed a linear analysis between the prediction and ground truth obtained from biochemical assays. Our results showed that the data points are largely distributed around the fit line (y = x), with a Pearson correlation coefficient (r > 0.90) indicating a significant linear correlation between the predictions and ground truth (Fig. 4b). Additionally, the residual plot demonstrates minimal prediction bias, with residuals evenly distributed without any systematic pattern (Fig. 4b). This suggests robustness and reliability of MoDL in predicting various mitochondrial functions.
Through the comparison of various indicators, including error analysis and the coefficient of determination, MoDL showed notable advantages (Fig. 4c and Supplementary Fig. 14). Firstly, MoDL exhibited significantly lower MSE, root mean square error (RMSE), and relative root mean square error (R-RMSE) compared to the single-modality algorithms (P < 0.05), indicating smaller prediction errors in regression tasks. Subsequently, MoDL yielded lower MAE and MAPE, demonstrating accurate prediction of target variables and enhanced stability in prediction results. Additionally, the MoDL algorithm outperformed the key indicator of the coefficient of determination (P < 0.05), reflecting a higher degree of fitting between prediction and ground truth. This superior performance may stem from the ensemble learning’s adeptness in fully leveraging complementary information between image and numerical data, thereby enhancing model prediction accuracy (Fig. 4c and Supplementary Fig. 14).
To further validate the robustness of MoDL, ablation experiments were conducted. The obtained results demonstrated that even with a reduced training data size of 5000 images (512 × 512 pixel2 resolution), MoDL achieved accurate predictions of mitochondrial function, displaying no significant deviations from the ground truth values (Fig. 4d, P > 0.05). However, when the training sample size was further reduced, pronounced discrepancies emerged between the prediction and the ground truth (Fig. 4d, P < 0.05). Moreover, ablation experiments were performed to investigate the impact of individual modules within the fusion framework. Results showed that solely employing the XGBoost + LightGBM fusion, XGBoost, LightGBM, and CNN resulted in substantial disparity between the predicted and ground truth (Fig. 4d, P < 0.05). These findings highlighted the importance of ensemble learning algorithm in enhancing the prediction accuracy (Fig. 4d). Lastly, we evaluated the computational costs of the MoDL vs. single modules (Supplementary Table 13). Although ensemble learning increases computational costs, they remain manageable. Notably, the substantial improvement in prediction accuracy justifies this trade-off.
MoDL evaluates drug efficacy on mitochondria
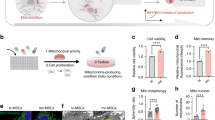
Encouraged by the capabilities of MoDL in mitochondrial function prediction, we extended our investigation to observe variations in mitochondrial function following treatment with different drugs in human liver cancer and normal cells. The mitochondrial functions of each batch of cells were imaged by SIM predicted by MoDL, and evaluated by wet bench biochemistry assays, respectively (Fig. 5a and Supplementary Fig. 12). To begin with, we evaluated the accuracy of MoDL’s predictions of mitochondrial function using different numbers of mitochondrial images (512 × 512 pixel2 resolution, Fig. 5b). As expected, the MAE between MoDL’s predictions and the ground truth descended with increased number of images contributing to the MoDL prediction process. The prediction stabilized near the ground truth with an ensemble of 80 images (i.e., 5 images in 2048 × 2048 pixel2 resolution), underscoring the method’s enhanced confidence and robustness. Therefore, in our following mitochondrial function prediction, we acquired 5 SIM images (2048 × 2048 pixel2 resolution) for each data point. MoDL also assessed the feature importance in mitochondrial function prediction (Supplementary Fig. 15), identifying the top 3 pivotal attributes for diverse mitochondrial function predictions (Fig. 5c).
a A scheme illustrating the MoDL predicts the impact of various drugs on mitochondrial functions in human liver cancer and normal cells (created with BioRender.com). b The MAE between the MoDL’s predictions and ground truth were calculated from different numbers of tested images (16, 32, 48, 64, 80, and 160), respectively. c Identification of the top 3 important features in MoDL’s predictions of various mitochondrial functions, along with Pearson’s correlation coefficient (r) for each feature relative to the ground truth. d MoDL prediction of changes in MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate in HepG2 cells treated with CCCP (orange and red lines) and Doxorubicin (DOX, 1 and 5 μM, green and blue lines) for 0 (control), 2, 4, 6, 12, and 24 h. The dash and solid lines represent prediction and ground truth (biochemical assays), respectively. n = 3 independent experiments. e MoDL’s prediction of mitochondrial protection in L02 cells treated with Polydatin (Pol, 50 μM) and Mitoquinone (MitoQ, 1 μM) for 24 h, followed by CCCP (5 μM) for 4 h. The orange and blue columns represent prediction and ground truth (biochemical assays), respectively, n = 3 independent experiments. Data are given as the mean ± SD (d, e). The original experimental biochemical results are provided in Supplementary Data 2 and 3 (d, e). Statistical differences were calculated using a two-sided Mann-Whitney test (d, e). No significant (ns, P > 0.05). Source data are provided as a Source Data file.
We next utilized MoDL to predict mitochondrial functions under the treatment of CCCP or Doxorubicin40 (DOX, a DNA synthesis inhibitor) at two concentrations (1 and 5 μM). Although CCCP and DOX are both cytotoxic agents, their mechanisms of action are mitochondria and nucleus-targeted, respectively (Fig. 5a). MoDL processed the corresponding images, generating predictions of mitochondrial functions in MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate (Fig. 5d, dash line). According to MoDL predictions, both CCCP and DOX significantly reduced ATP generation in HepG2 cells (Fig. 5d, P > 0.05), attributable to the cytotoxicity from both drugs lowering cell viability and ATP generation. However, MoDL analysis revealed that, compared to DOX, CCCP was more effective in inducing MMP depolarization, mitophagy, and respiration rate at both concentrations (Fig. 5d, P < 0.05). While only at higher concentration did CCCP introduce significantly higher ROS production compared to DOX. This disparity may arise from targeting action of CCCP on mitochondria, promoting an earlier and prominent effect on mitochondrial function. Notably, all these prediction results from MoDL have been further verified by biochemistry assays in parallel (Fig. 5d, solid line) which are consistently aligned with the predicted outcomes (P > 0.05).
In another experiment, L02 cells were first treated with Mitoquinone (MitoQ, a synthetically engineered antioxidant)41 and Polydatin (Pol, a natural small molecule antioxidant derivatized of resveratrol)42 followed by the introduction of CCCP (Fig. 5a). Compared to the more established mechanisms of MitoQ, Pol remains largely unexplored regarding its biological benefits and underlying mechanisms. Employing MoDL to investigate the potential pharmacological activity of Pol in protecting mitochondrial functionality would provide valuable insights. According to the analysis from MoDL, Pol exhibited more effective inhibition in MMP depolarization, preserved ATP synthesis, and respiration rate compared to MitoQ (Fig. 5e, P < 0.05). Furthermore, both drugs demonstrated efficacy in suppressing ROS production. However, MitoQ effectively restored mitophagy level to normal, Pol exhibited no influence in this regard. These findings highlighted the significance of delving deeper into the protective mechanisms of natural products like Pol on mitochondria. Similarly, the corresponding biochemistry assays (Fig. 5e, blue columns, P > 0.05) further substantiated MoDL’s precision in accurately predicting the dynamic changes in mitochondrial functions (Fig. 5e, orange columns).
The observations indicated that MoDL has the potential to predict the effects of drugs on various mitochondrial function indicators, thereby distinguishing their respective action pathways and offering an initial assessment of drug efficacy. Consequently, MoDL emerges as a powerful tool for facilitating high-throughput drug screening and providing insights into drug efficacy and mechanisms.
MoDL’s third pipeline: prediction in unseen cell types with small sample size training
Drug resistance is the primary factor contributing to the failure of tumor chemotherapy, often stemming from the mutations that occur in tumor cells under prolonged drug pressure43. A variety of anticancer drug-resistance mechanisms have been reported, including significant disparities in mitochondrial function and morphology, representing a novel avenue for exploring the intricacies of tumor drug-resistance44,45. Leveraging MoDL, we uncovered aberrations in mitochondrial morphological changes in cisplatin-treated drug-resistant A2780 cells (Fig. 6a). Basal A2780 S cells (cisplatin-sensitive) exhibited a highly fused mitochondrial morphology characterized by MoDL. In contrast, A2780 CP cells (cisplatin-resistant) displayed a propensity towards spherical and short rod-shaped mitochondria with enhanced fission and mitophagy events46 (Fig. 6a). Subsequently, we conducted cisplatin intervention experiments and MoDL indicated that as the dosage of cisplatin increased, the mitochondrial morphology of A2780 S cells transitioned gradually from the initial highly fused state to a fragmented state (Fig. 6b, c and Supplementary Fig. 17a), showing the similar trend with other cancer cell lines (Fig. 3b).
a An illustrative scheme presents the distinct mitochondrial morphological trends of drug-sensitive and -resistant human ovarian cancer cells upon cisplatin treatment (created with BioRender.com). b MoDL-generated pseudo-color images (1024 × 1024 pixel2 resolution) display the mitochondrial morphology under cisplatin treatment (0, 1, 5, 10 μM) for 24 h, and the corresponding morphological quantitative features (c), including mean area, form factor, branch length, and solidity. The blue and orange dot represent A2780 S and A2780 CP cells, respectively. n = 85 images (512 × 512 pixel2 resolution), scale bars = 2 μm. d The impact of collecting diverse sample sizes from a pre-existing dataset on the MSE in MoDL’s predictions (initial training set = 1120, test set = 320, i.e., 70 and 20 images in 2048 × 2048 pixel2 resolution, respectively). e MoDL’s predictions of changes in MMP polarization, ATP generation, ROS production, mitophagy level, and respiration rate in A2780 S (blue line) and A2780 CP (orange line) cells treated with cisplatin (0, 1, 5, 10 μM) for 24 h. The dash and solid lines represent prediction and ground truth (biochemical assays), respectively. n = 3 independent experiments. Data are given as the mean ± SD (c, e). The original experimental biochemical results are provided in Supplementary Data 4 (e) Statistical differences were calculated using a two-way ANOVA, A2780 S vs. A2780 CP cells and cisplatin treatment 10 μM groups vs. control (c) or a two-sided Mann–Whitney test, prediction vs. ground truth (e). No significant (ns, P > 0.05). Source data are provided as a Source Data file.
Intriguingly, our segmentation of cisplatin-resistant A2780 CP cells by MoDL revealed a phenomenon contrary to that in cisplatin-sensitive A2780 S. As cisplatin dosage increased, mitochondrial morphology underwent a gradual transition from an initially highly fragmented state towards a more fused configuration (Fig. 6b and Supplementary Fig. 17a). Moreover, our detailed quantitative analysis of mitochondrial morphology of A2780 CP cells via MoDL highlighted a notable trend in response to increasing cisplatin concentrations, which is characterized by a consistent increase in mitochondrial area, form factor, and branch length, accompanied by a decrease in solidity (Fig. 6c). This anomalous phenomenon underscores the complex and divergent adaptations in drug-resistant cell lines. These findings not only contribute to a better understanding of the biological mechanisms underlying tumor drug resistance but also inspire novel therapeutic strategies targeting drug-resistant tumor cells. It should be noted that the observation of these “abnormal” changes in mitochondrial morphology was realized under consistent live-cell imaging with the aid of precise segmentation and analysis from MoDL.
In our subsequent work, we endeavored to use MoDL to predict mitochondrial functions in A2780 cells. Given the unique characteristics of this cell line, retraining of the model was necessary. Fortunately, this did not require a vast new collection of mitochondrial images. By strategically selecting highly relevant features from our extensive pre-existing dataset, we employed Z-score calculations to extract data from the large dataset that was highly compatible with the A2780 dataset (Supplementary Note 8). We found that a subset of 20,000 samples significantly minimized the MSE between predicted mitochondrial functions and ground truth for the A2780 task (Fig. 6d). However, beyond this point, increasing subset size introduced high Z-score samples, leading to more irrelevant or noisy data (Supplementary Table 14). This degraded the training process and decreased prediction accuracy, resulting in a higher MSE (Fig. 6d). Notably, the newly assembled dataset of just 1440 images (equivalent to 90 images at 2048 × 2048 pixel2 resolution, which could be completed within 2 h using NIS-Elements) was adequate to re-train MoDL to predict mitochondrial functions in A2780 cells with high accuracy (Fig. 6e). Predicted outcomes from MoDL (Fig. 6e, dash line) are consistently aligned with the biochemistry assays (Fig. 6e, solid line, P > 0.05). These findings imply a distinct mitochondrial adaptation to cisplatin in drug-resistant cells. More importantly, our results have demonstrated the practical potential of MoDL in predicting mitochondrial functions in unseen cell types, despite the limitations of sample size and distinct morphology patterns.
Discussion
MoDL is a deep learning algorithm that performs high-accurate segmentation of mitochondria and predicts its function in live-cell imaging. The first key pipeline of MoDL is mitochondrial segmentation. In training set preparation, we annotated over 20,000 mitochondria to optimize learning features and achieve accurate segmentation. Furthermore, we incorporated ResNets, CBAM, and U-Net to deepen the model architecture. Compared to published methods, MoDL not only exhibited higher performance but also demonstrated very high robustness through blur/noise models. Next, we verified the ability of MoDL to be deployed across SIM and confocal imaging platforms. Meanwhile, MoDL could also precisely track the mitochondrial morphological changes under different conditions. Based on the high-quality segmentation outputs from MoDL, we comprehensively and quantitatively analyzed mitochondrial morphologies, generating 31 morphological features for individual mitochondrions and over 100 quantitative metrics. We then conducted a wide range of mitochondrial function analyses both by MoDL and biochemistry assays, demonstrating a high correlation between mitochondrial morphology and functions.
We further established an ensemble learning algorithm, the second key pipeline of MoDL, which realizes regression and prediction of mitochondrial functions. Ensemble learning integrates numerical input of morphological data and image of mitochondrial features. Its robustness was enhanced by training on a dataset contains over 100,000 SR images, each annotated with corresponding functional data obtained through biochemical assays. Our findings indicated that the predictive outcomes of the ensemble learning algorithm are not significantly different from the ground truth values obtained through biochemical methods. Furthermore, we utilized MoDL to assess drug impact patterns on mitochondrial function in liver cancer and normal cells. We verified that a natural product, Pol, can effectively protect mitochondrial functions.
Our current study implementing five key metrics, however, mitochondrial function is inherently diverse, and other indicators such as changes in fatty acid oxidation (FAO) are also closely linked to mitochondrial morphology47,48,49. Using MoDL, we have successfully predicted FAO level in the CCCP-treated HepG2 model (Supplementary Note 9, Supplementary Fig. 18, and Supplementary Table 15). Looking forward, we anticipate that incorporating a wider array of indicators in the future will enhance the generalization capability and application scope of MoDL. This expanded framework aims to provide a more comprehensive depiction of mitochondrial physiological states, potentially deepening our understanding of mitochondrial morphology and its implications in health and dysfunction. This will pave the way for more accurate diagnostics, improved therapeutic strategies, and novel insights into cellular metabolisms.
In considering features for model training, fluorescence intensity also should be taken into account. However, the intensity of mitochondria is influenced by various factors during imaging50. For instance, different dyes exhibit varying photobleaching resistance, leading to discrepancies in signal intensity. Additionally, imaging conditions, such as focus plane adjustments, laser power, and exposure time, significantly impact observed intensity. Therefore, standardized imaging protocols are crucial. In our study, due to the prolonged data collection period and inherent variability, we have not yet incorporated fluorescence intensity as a feature for further functional insights. Moving forward, we aim to standardize imaging protocols to include fluorescence intensity in our model training. We anticipate that this will enhance the robustness of MoDL, offering more comprehensive and reliable insights into mitochondrial health and disease.
Lastly, through the application of MoDL, we have identified distinct morphological changes in mitochondria within drug-resistant A2780 CP cells. Upon cisplatin treatment, these cells demonstrated morphological trends that were contrary to those observed in normal cells. Furthermore, MoDL’s third key pipeline, bolstered by the substantial dataset alongside data fine-tuning and retraining, has achieved precise predictions of mitochondrial functions in A2780 CP cells, despite the limitations of small sample size training and abnormal mitochondrial morphology patterns. In practice, MoDL requires just a modest collection of approximately 90 images in 2048 × 2048 pixel2 resolution from users, which are corroborated with experimentally tested functions. By capitalizing on data fine-tuning and retraining with our comprehensive dataset, MoDL demonstrated robustness and generalization capabilities in accurately predicting mitochondrial functions across a spectrum of cell types. Looking forward, our aim is to develop MoDL into an online, freely accessible service, available on a public platform for the benefit of the wider scientific community.
We believe that MoDL serves as a robust tool for accurate and effective analysis of mitochondria in live-cell images. Furthermore, we envision MoDL fostering the widespread dissemination of feature data related to mitochondrial morphology and function. This, in turn, could enable the utilization of deep learning to enhance generalization and conduct morphological analysis under various circumstances. Additionally, we anticipate MoDL’s contribution to diverse mitochondria-related scientific research, such as drug discovery51, mitochondrial transplantation52, sensing53, disease diagnosis54, and targeted drug delivery55, thereby opening avenues for the study of mitochondria.
Methods
Cell lines and cell culture
The cell lines 143B (human osteosarcoma cells, CRL-8303), HeLa (human cervix cancer cell, CCL-2), U87 (human astrocytoma cell, HTB-14), HepG2 (HB-8065) and Hep3B (human carcinoma cells, HB-8064), and MCF7 (human breast cancer cell, HTB-22) were obtained from American Type Culture Collection (ATCC). The cell lines L02 (human normal liver cell, CL-0111) and A2780 (human ovarian cancer cells, CL-0013) were purchased from Procell Life Science & Technology Co., Ltd. Cells were cultured under standard conditions (95% humidity, 5% CO2, 37 °C) and harvested by 0.5% (w/v) trypsin-EDTA solution. Before undergoing further experiments, cells were seeded in 96- or 6-well plates in DMEM (Dulbecco’s modified eagle medium, Gibco) supplemented with 10% fetal bovine serum (Gibco), 100 U/ml penicillin, and 100 μg/mL streptomycin (HyClone).
Live-cell imaging
The Nikon C2plus&N-SIM E is an integrated microscope capable of compatibility with both SIM and confocal. The training dataset for the MoDL segmentation pipeline was collected using SIM, while the test and validation datasets were obtained using SIM and confocal microscopy. Prior to imaging, the wild-type HeLa, 143B, U87, HepG2, Hep3B, L02, A2780, and MCF7 cells were seeded in glass-bottomed dishes and grown to 60–70% confluence in DMEM. After staining the cells with MitoTracker-Green, MitoTracker-Red, and MitoTracker-DeepRed (100 nM) for 30 min, the cells were washed with PBS buffer three times and supplemented with phenol red-free DMEM. The glass-bottomed dishes were then transferred to the imaging platform of the microscope for imaging. HepG2 cells were transfected with GFP (green fluorescent protein) or RFP (red fluorescent protein)-expressing lentiviral vectors (Xi’an Orbitalgene Co, China) to express high amounts of endogenous GFP or RFP targeted to the mitochondria. Fluorescence imaging was performed with Nikon N-SIM E with the CFI Apochromat TIRF 100XC Oil (N.A. ≥1.49, WD. ≥ 0.12 mm).
Live-cell mitochondrial morphological tracking
Wild-type HepG2 cells were seeded in glass-bottomed dishes and allowed to reach 60–70% confluency. After incubation with MitoTracker-Green for 30 min, the medium was replaced with fresh DMEM containing CCCP (10 μM). The dishes were then placed in a live-cell imaging workstation maintained at 37 °C with 5% CO2. Imaging coordinates of fixed viewing region were recorded using Nikon N-SIM E to capture images at 0, 1, and 2 h points. Subsequently, the medium was removed again and replaced with fresh DMEM containing Mdivi-1 (10 μM). Additional images were acquired at 2, 3, and 4 h time points using the same imaging coordinates established earlier. This experimental setup enabled the observation of mitochondrial dynamics in HepG2 cells, allowing for the investigation of the effects of CCCP and Mdivi-1 on mitochondrial morphology.
Detection of mitochondrial ROS production
MitoSOXTM, a commercial fluorogenic dye specifically targeted to mitochondria in living cells. Oxidation of MitoSOX™ by ROS produces fluorescence. The cells were seeded in glass-bottomed dishes and grown to 50–60% confluence in DMEM. The above cells were incubated with drugs for different time points, followed by replacing with fresh DMEM containing MitoSOXTM (5 μM) for 20 min. The cells were then washed with PBS buffer three times and imaged with the EVOS fluorescent inverted microscope immediately with a band path of 550–600 nm upon excitation at 488 nm in same exposure and excitation settings.
Seahorse XFe-24 metabolic flux analysis
A total of 3 × 105 cells/well were seeded into XFe 24-well cell culture plates and incubated overnight to allow attachment. The cells were then treated with CCCP (0, 1, 5, 10, 20 µM) for 4 h. The vehicle-alone (DMSO) control cells were processed in parallel. After treatment, the cells were washed in pre-warmed XF assay media (XF assay media supplemented with 10 mM of glucose, 1 mM of Pyruvate, and 2 mM of L-glutamine, and the pH adjusted to 7.4). The cells were then maintained in 500 µL/well of XF assay media at 37 °C in a non-CO2 incubator for 1 h. For OCR measurement, 1 µM of oligomycin, 0.5 µM of carbonyl cyanide-4-(trifluoromethoxy) phenylhydrazone (FCCP), 0.5 µM of rotenone/antimycin A were subsequently added in XF assay media, and loaded in the XFe-24 sensor cartridge. The measurements were normalized by protein content (SRB assay). All experiments were performed in triplicate. The Cell Mito Stress Test was generated by following the manufacturer’s guidelines and instructions (Seahorse, Agilent Technologies).
Detection of MMP polarization
The JC-10 commercial assay kit (Abcam) was used to detect MMP polarization. In the healthy mitochondria, JC-10 aggregates within the mitochondrial matrix, forming polymers that emit red fluorescence. Conversely, under conditions of mitochondrial damage and decreased MMP, JC-10 cannot accumulate within the matrix and exists as monomers, generating green fluorescence. The transition from red to green fluorescence provides a quantifiable measure of mitochondrial depolarization. Cells were seeded in 6-well plates and allowed to grow to 60–70% confluence before treatment with drugs. Subsequently, the culture medium was replaced with fresh DMEM supplemented with JC-10 (10 μM) and incubated for 30 min. After the desired treatment duration, cells were washed with PBS buffer and detached using trypsin-EDTA. Then, the cells were collected and centrifuged to obtain a cell pellet, which was then resuspended in 200 μL of PBS buffer in 1.5 mL tubes. Flow cytometry (CytoFLEX S) was then employed to analyze MMP polarization. Forward scatter (FSC) and side scatter (SSC) gates were applied to exclude low-FSC/low-SSC debris. In the healthy mitochondria, JC-10 aggregates within the mitochondrial matrix, forming polymers that emit red fluorescence (λex = 561 nm, λem = 585 nm, detected in the PE channel). Under conditions of mitochondrial damage and decreased MMP, JC-10 cannot accumulate within the matrix and exists as monomers, generating green fluorescence (λex = 488 nm, λem = 525 nm, detected in the FITC channel). The PE/FITC ratio provides a quantifiable measure of mitochondrial depolarization.
Detection of ATP generation
The commercial CellTiter-Glo® assay kit (Promega) was utilized to quantify ATP generation, a key indicator of mitochondrial function. The assay relies on a homogeneous quantification of ATP through a luminescence-based readout. CellTiter-Glo® Reagent was prepared by mixing CellTiter-Glo® Buffer and CellTiter-Glo® solution. The cells were seeded in a 96-well opaque plate at a density of 70–80% confluency and treated with 100 μL of DMEM followed by the addition of 100 μL of CellTiter-Glo® Reagent per well. The plate was then placed on an orbital shaker for 2 min to induce cell lysis. Following shaking, the plate was incubated at room temperature for 10 min to stabilize the luminescent signal before recording the phosphorescence signal using a microplate reader (TECAN).
Detection of mitophagy level
Recently, we reported a development of a pyrimidine-based small-molecule fluorescent probe, Z2, for monitoring mitophagy through pH changes within the mitochondria56. The core structure of Z2 was designed with a push-pull electronic configuration, and the terminal hydroxy groups of the pyrimidine moiety were capped to effectively suppress the intramolecular charge transfer (ICT) effect, thereby minimizing the fluorescence quantum yield. Under acidic conditions, the dimethylaniline moiety undergoes protonation, leading to restoration of the ICT effect and subsequent fluorescence signal generation. To achieve mitochondrial targeting, Z2 was modified with a triphenylphosphonium cation (TPP+) moiety, resulting in a Pearson’s correlation coefficient (r = 0.90) comparable to commercially available mitochondrial dyes. This innovative design of Z2 offers a promising tool for investigating mitophagy by facilitating real-time monitoring of pH changes within the organelle. The cells were seeded in glass-bottomed dishes and grown to 50–60% confluence in DMEM. The above cells were incubated with drugs, the medium was removed, fresh DMEM containing Z2 (5 μM) was added and incubated for 20 min. After the desired time, the cells were washed with PBS buffer three times. Then the cells were imaged with the Nikon C2plus confocal microscope immediately with a band path of 550–600 nm upon excitation at 488 nm.
Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted by using TRIzol reagent (Invitrogen) after the indicated treatments. In total, 500 ng of isolated RNA was reverse transcribed into complementary DNA using a Prime Script RT Reagent Kit (Takara). qRT-PCR was performed with a CFX Connected real-time system (Bio-Rad) using Ultra SYBR Green Mixture (Takara) and specific primers. The amplification conditions were initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s. Each sample was run in triplicate. The specificity of PCR products was confirmed by melting curve analysis. Relative expression was quantified by the 2-DDCt method. GAPDH was used as the housekeeping gene to normalize gene expression levels. The primer sequences are listed in Supplementary Data 5.
Western blot analysis
Total cell protein lysates were separated by 10% SDS-PAGE electrophoresis and then transferred to polyvinylidene difluoride membranes (0.2 mm pore size) (Millipore, USA). After blockade with 5% skim milk powder TBST for 1 h at room temperature (RT), the membranes were then incubated with antibodies against DRP1 (EPR19274, #ab184247, abcam, 1:1000), OPA1 (D6U6N, #80471, Cell Signaling Technology, 1:1000), BNIP3 (EPR4034, #ab109362, abcam, 1:1000), SIRT1 (D1D7, #9475, Cell Signaling Technology, 1:1000), GAPDH (1E6D9, #60004-1-Ig, proteintech, 1:5000) overnight at 4 °C. The secondary antibodies were HRP-coupled goat anti-rabbit IgG (H + L) antibody, peroxidase-labeled (#074-1506, Kirkegaard & perry laboratories, 1:5000), anti-mouse IgA + IgG + IgM (H + L) antibody, human serum adsorbed and peroxidase-labeled (#074-1807, Kirkegaard & perry laboratories, 1:5000). The membranes were visualized with chemiluminescent peroxidase reagents (BioCytoSci, USA) and the expression of target proteins was analyzed with Quantity One software.
Statistics and reproducibility
The experiment was independently repeated and depicted in figure legends, and the results were similar. All representative images presented were reproduced with similar results. All the experimental data were analyzed with GraphPad Prism 9 (GraphPad Software, San Diego, CA). The results in all the experiments were presented as mean ± standard deviation (s.d.). The experiments were independently repeated at least 3 times. Normality and homogeneity of variances were assessed using the Shapiro-Wilk test and Brown-Forsythe test, respectively. For non-normally distributed data, statistical comparisons were conducted using the Kruskal-Wallis test followed by Dunn’s multiple comparison test (more than two groups) or Mann–Whitney test (two groups). For normally distributed data, statistical comparisons were conducted using one-way or two-way ANOVA followed by Dunnett’s multiple comparison test (more than two groups) or two-tailed Student’s t-test (two groups). A P value (<0.05) was considered statistically significant: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data needed to evaluate the conclusions in the paper are present in the Source Data file, Supplementary Data files, and in the Supplementary Information. The datasets generated and/or analyzed during the study are available on GitHub (https://github.com/OBPNPW2024/MoDL). Source data are provided with this paper.
Code availability
The MoDL algorithm is written in Python 3.8.0. The corresponding source codes and scripts are available through GitHub at (https://github.com/OBPNPW2024/MoDL). The executable files and codes are also deposited on (https://doi.org/10.5281/zenodo.10889133)57.
References
Amorim, J. A. et al. Mitochondrial and metabolic dysfunction in ageing and age-related diseases. Nat. Rev. Endocrinol. 18, 243–258 (2022).
Chen, L. et al. Mitochondrial heterogeneity in diseases. Signal. Transduct. Target Ther. 8, 311 (2023).
Monzel, A. S., Enríquez, J. A. & Picard, M. Multifaceted mitochondria: moving mitochondrial science beyond function and dysfunction. Nat. Metab. 5, 546–562 (2023).
Picard, M. & Shirihai, O. S. Mitochondrial signal transduction. Cell Metab. 34, 1620–1653 (2022).
Bulthuis, E. P., Adjobo-Hermans, M. J. W., Willems, P. H. G. M. & Koopman, W. J. H. Mitochondrial morphofunction in mammalian cells. Antioxid. Redox Signal. 30, 2066–2109 (2019).
Giacomello, M., Pyakurel, A., Glytsou, C. & Scorrano, L. The cell biology of mitochondrial membrane dynamics. Nat. Rev. Mol. Cell Biol. 21, 204–224 (2020).
Dong, W. T. et al. Mitochondrial fission drives neuronal metabolic burden to promote stress susceptibility in male mice. Nat. Metab. 5, 2220–2236 (2023).
Murakawa, T. et al. Bcl-2-like protein 13 is a mammalian Atg32 homologue that mediates mitophagy and mitochondrial fragmentation. Nat. Commun. 6, 7527 (2015).
Gomes, L. C., Di Benedetto, G. & Scorrano, L. During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat. Cell Biol. 13, 589–598 (2011).
Chu, C. H., Tseng, W. W., Hsu, C. M. & Wei, A. C. Image analysis of the mitochondrial network morphology with applications in cancer research. Front. Phys. 10, 855775 (2022).
Zhai, R. et al. Small-molecule fluorogenic probes for mitochondrial nanoscale imaging. Chem. Soc. Rev. 52, 942–972 (2023).
Lin, C. H., Lin, T. Y., Hsu, S. C. & Hsu, H. J. Expansion microscopy-based imaging for visualization of mitochondria in drosophila ovarian germline stem cells. FEBS Open Bio 12, 2102–2110 (2022).
Chaudhry, A., Shi, R. & Luciani, D. S. A pipeline for multidimensional confocal analysis of mitochondrial morphology, function, and dynamics in pancreatic β-cells. Am. J. Physiol. Endoc. Metab. 318, E87–E101 (2020).
Viana, M. P., Lim, S. & Rafelski, S. M. Quantifying mitochondrial content in living cells. Methods Cell Biol. 125, 77–93 (2015).
Lefebvre, A. E. Y. T., Ma, D., Kessenbrock, K., Lawson, D. A. & Digman, M. A. Automated segmentation and tracking of mitochondria in live-cell time-lapse images. Nat. Methods 18, 1091–1102 (2021).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. In Medical Image Computing and Computer Assisted Intervention-MICCAI (eds Frangi, A., Schnabel, J., Davatzikos, C., Alberola-López, C. & Fichtinger, G.) Lecture Notes in Computer Science, vol. 11071, 265–273 (Springer, Cham, 2018).
Sekh, A. A. et al. Physics-based machine learning for subcellular segmentation in living cells. Nat. Mach. Intell. 3, 1071–1080 (2021).
Cutler, K. J. et al. Omnipose: a high-precision morphology-independent solution for bacterial cell segmentation. Nat. Methods 19, 1438–1448 (2022).
Fischer, C. A. et al. MitoSegNet: easy-to-use deep learning segmentation for analyzing mitochondrial morphology. iScience 23, 101601 (2020).
Jakobs, S. & Wurm, C. A. Super-resolution microscopy of mitochondria. Curr. Opin. Chem. Biol. 20, 9–15 (2014).
Ouellet, M., Guillebaud, G., Gervais, V., Lupien St-Pierre, D. & Germain, M. A novel algorithm identifies stress-induced alterations in mitochondrial connectivity and inner membrane structure from confocal images. PLoS. Comput. Biol. 13, e1005612 (2017).
Rohani, A., Kashatus, J. A., Sessions, D. T., Sharmin, S. & Kashatus, D. F. MitoHacker: a set of tools to enable high-throughput analysis of mitochondrial network morphology. Sci. Rep. 10, 18941 (2020).
Shirihai, O. S., Song, M. & Dorn, G. W. 2nd How mitochondrial dynamism orchestrates mitophagy. Circ. Res. 116, 1835–1849 (2015).
Chan, D. C. Mitochondrial dynamics and its involvement in disease. Annu. Rev. Pathol. 15, 235–259 (2020).
Wang, Z., Wang, E. & Zhu, Y. Image segmentation evaluation: a survey of methods. Artif. Intell. Rev. 53, 5637–5674 (2020).
Lucas, A. M. et al. Open-source deep-learning software for bioimage segmentation. Mol. Biol. Cell 32, 823–829 (2021).
Heintzmann, R. & Huser, T. Super-resolution structured illumination microscopy. Chem. Rev. 117, 13890–13908 (2017).
Li, Q. & Chen, C. A robust and high-precision edge segmentation and refinement method for high-resolution images. Math. Biosci. Eng. 20, 1058–1082 (2023).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI (eds Navab, N., Hornegger, J., Wells, W. & Frangi, A.) Lecture Notes in Computer Science, vol. 9351, 234–241 (Springer, Cham, 2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition, IEEE Conf. Comput. Vis. Pattern Recognit. 770–778 (2016).
Padman, B. S., Bach, M., Lucarelli, G., Prescott, M. & Ramm, G. The protonophore CCCP interferes with lysosomal degradation of autophagic cargo in yeast and mammalian cells. Autophagy 9, 1862–1875 (2013).
Aishwarya, R. et al. Pleiotropic effects of mdivi-1 in altering mitochondrial dynamics, respiration, and autophagy in cardiomyocytes. Redox Biol. 36, 101660 (2020).
Kane, M. S. et al. Current mechanistic insights into the CCCP-induced cell survival response. Biochem. Pharmacol. 148, 100–110 (2018).
Li, X., Zhao, Y., Yin, J. & Lin, W. Organic fluorescent probes for detecting mitochondrial membrane potential. Coord. Chem. Rev. 420, 213419 (2020).
Zhang, L. & Song, Q. Multimodel integrated enterprise credit evaluation method based on attention mechanism. Comput. Intell. Neurosci. 2022, 8612759 (2022).
Chen, T. & Guestrin, C. XGBoost: A scalable tree boosting system. ACM SIGKDD International Conference on Knowledge Discovery & Data Mining 785-794 (2016).
Kumar, D., Sood, S. K. & Rawat, K. S. Early health prediction framework using XGBoost ensemble algorithm in intelligent environment. Artif. Intell. Rev. 56, 1591–1615 (2023).
Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 1, 206–215 (2018).
Kciuk, M. et al. Doxorubicin-an agent with multiple mechanisms of anticancer activity. Cells 12, 659 (2023).
Xiao, L. et al. The mitochondria-targeted antioxidant MitoQ ameliorated tubular injury mediated by mitophagy in diabetic kidney disease via Nrf2/PINK1. Redox Biol. 11, 297–311 (2017).
Wu, M. et al. Polydatin for treating atherosclerotic diseases: a functional and mechanistic overview. Biomed. Pharmacother. 128, 110308 (2020).
Nallasamy, P. et al. Tumor microenvironment enriches the stemness features: the architectural event of therapy resistance and metastasis. Mol. Cancer 21, 225 (2022).
Jin, P. et al. Mitochondrial adaptation in cancer drug resistance: prevalence, mechanisms, and management. J. Hematol. Oncol. 15, 97 (2022).
Zampieri, L. X. et al. Mitochondria participate in chemo resistance to cisplatin in human ovarian cancer cells. Mol. Cancer Res. 18, 1379–1391 (2020).
Xie, L. et al. Drp1-dependent remodeling of mitochondrial morphology triggered by EBV-LMP1 increases cisplatin resistance. Signal. Transduct. Target Ther. 5, 56 (2020).
Carracedo, A., Cantley, L. C. & Pandolfi, P. P. Cancer metabolism: fatty acid oxidation in the limelight. Nat. Rev. Cancer 13, 227–232 (2013).
Castro-Sepulveda, M., Fernández-Verdejo, R., Zbinden-Foncea, H. & Rieusset, J. Mitochondria-SR interaction and mitochondrial fusion/fission in the regulation of skeletal muscle metabolism. Metabolism 144, 155578 (2023).
Ngo, J. et al. Mitochondrial morphology controls fatty acid utilization by changing CPT1 sensitivity to malonyl-CoA. EMBO J. 42, e111901 (2023).
Liu, X., Yang, L., Long, Q., Weaver, D. & Hajnóczky, G. Choosing proper fluorescent dyes, proteins, and imaging techniques to study mitochondrial dynamics in mammalian cells. Biophys. Rep. 3, 64–72 (2017).
Crunkhorn, S. Mitochondrial disease: eliminating mutant mitochondrial DNA. Nat. Rev. Drug Discov. 17, 788 (2018).
Zhang, T. G. & Miao, C. Y. Mitochondrial transplantation as a promising therapy for mitochondrial diseases. Acta Pharm. Sin. B. 13, 1028–1035 (2023).
Ji, W. et al. Optical/electrochemical methods for detecting mitochondrial energy metabolism. Chem. Soc. Rev. 51, 71–127 (2022).
Gorman, G. S. et al. Mitochondrial diseases. Nat. Rev. Dis. Prim. 2, 16080 (2016).
Guo, X. et al. Mito-Bomb: targeting mitochondria for cancer therapy. Adv. Mater. 33, e2007778 (2021).
Zhou, J. et al. Pyrimidine-based fluorescent probe for monitoring mitophagy via detection of mitochondrial pH variation. Chembiochem 23, e202200217 (2022).
Ding, Y. et al. Mitochondrial segmentation and function prediction in live-cell images with deep learning. Zenodo https://doi.org/10.5281/zenodo.10889133 (2024).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (62288102 to W.H., 62475216 to B.P., 22077101 to L.L.), Key Research and Development Program of Shaanxi (2024GH-ZDXM-37 to B.P.), Fujian Provincial Natural Science Foundation of China (2024J01060 to L.L.), Natural Science Basic Research Program of Shaanxi (2024JC-YBQN-0919 to J.Z.), Innovation Foundation for Doctorate Dissertation of Northwestern Polytechnical University (CX2023098 to Y.D.), Postdoctoral Fellowship Program of CPSF (GZB20230379 to H.F., GZC20240889 to B.F.), China Postdoctoral Science Foundation (2023M742919 to H.F.), the Open Project Program of the Analytical & Testing Center of Northwestern Polytechnical University (2020T018 to L.L.), Startup Program of XMU (L.L.), and Fundamental Research Funds for the Central Universities (W.H. and L.L.). Australian Research Council (FL220100185 to N.H.V.).
Author information
Authors and Affiliations
Contributions
B.P., L.L., and W.H. conceptualized the study. Y.D. and J.L. developed the code for the MoDL algorithm. Y.D., J.L., J.Z., P.L., H.B., B.F., and H.F. performed experiments and formal analyses. Y.D., J.L., and B.P. wrote the manuscript. K.H., G.W., C.J.N., N.H.V., B.P., L.L., and W.H. reviewed and edited the manuscript. B.P., L.L., N.H.V., and W.H. secured funding for this work. B.P., L.L., and W.H. supervised different parts of the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Carsten Marr, who co-reviewed with Abhinanda Punnakkal; Md. Shenuarin Bhuiyan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, Y., Li, J., Zhang, J. et al. Mitochondrial segmentation and function prediction in live-cell images with deep learning. Nat Commun 16, 743 (2025). https://doi.org/10.1038/s41467-025-55825-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-55825-x
This article is cited by
-
PROTAC-based degradation of catechol-O-methyltransferase for potential treatment of Parkinson’s disease
Science China Chemistry (2026)
-
AI-assisted SERS imaging method for label-free and rapid discrimination of clinical lymphoma
Journal of Nanobiotechnology (2025)
-
STK-former: an efficient transformer with adaptive TopK sparse attention for image inpainting
The Visual Computer (2025)