Abstract
Mycoplasma pneumoniae causes atypical pneumonia in children and young adults. Its lack of a cell wall makes it resistant to beta-lactams, which are the first-line treatment for typical pneumonia. Current diagnostic tests are time-consuming and have low specificity, leading clinicians to administer empirical antibiotics. Using a LASSO regression simulation approach and blood microarray data from 107 children with pneumonia (including 30 M. pneumoniae) we identify eight different transcriptomic signatures, ranging from 3-10 transcripts, that differentiate mycoplasma pneumonia from other bacterial/viral pneumonias with high accuracy (AUC: 0.84–0.95). Additionally, we demonstrate that existing signatures for broadly distinguishing viral/bacterial infections and viral/bacterial pneumonias are ineffective in distinguishing M. pneumoniae from viral pneumonia. The new signatures are successfully validated in an independent RNAseq cohort of children with pneumonia, demonstrating their robustness. The high sensibility of these signatures presents a valuable opportunity to guide the treatment and management of M. pneumoniae pneumonia patients.
Similar content being viewed by others
Introduction
Mycoplasma pneumoniae commonly causes respiratory infections in children and young adults, presenting with clinical manifestations ranging from asymptomatic to pneumonia. M. pneumoniae is one of the most common causes of atypical pneumonia. The incidence of M. pneumoniae pneumonia is uncertain and is typically described as endemic, marked by cyclic epidemics occurring every 3 to 5 years. The COVID-19 pandemic, along with associated non-pharmaceutical interventions, had a significant impact on the circulation of respiratory pathogens, including M. pneumoniae1. A global surveillance network noted a decrease in M. pneumoniae detection from April 2020 to March 20212. Despite a resurgence in other pathogens, M. pneumoniae infections remained low until March 2022-23. Concerns about a potential upsurge in mycoplasma infections have arisen due to waning herd immunity3,4. Recent outbreaks of M. pneumoniae infections have been reported during the last months in different European countries, including Spain4, Denmark5, France6, and the Netherlands7 (see also https://edition.cnn.com/2023/12/01/health/mycoplasma-pneumonia/index.html). In November 2023, the World Health Organization (WHO) reported an increase in outpatient consultations and hospitalizations for M. pneumoniae pneumonia in China since May, along with rises in respiratory syncytial virus (RSV), adenovirus, and influenza virus cases since October1,3,8.
Establishing the microbiological etiology of pneumonia remains a challenge for clinicians within healthcare facilities, primarily due to the complexity of obtaining direct microbiological samples, which often require invasive procedures. Additionally, there is a delay in obtaining results from other indirect tests, such as pneumococcal antigen in urine, PCR in respiratory swabs, or serology, when available. In the particular case of M. pneumoniae, cultures are considered the gold-standard for bacterial diagnosis; however, due to the difficulty in obtaining representative samples, slow growth rates, and low sensitivity, culture methods are impractical for timely clinical and treatment decision-making9. Serology is another commonly used approach for diagnosing M. pneumoniae9, but tests show low specificity. IgM antibodies can remain elevated for months after infection, with frequent cross-reactivity with other pathogens but also potential for generating false positives10. As a result, serology has gradually been replaced by PCR11, a rapid method offering higher sensitivity and specificity compared to serology and culture. PCR is now widely used in developed countries for the diagnosis of M. pneumoniae. Specific diagnosis of M. pneumoniae is important since, due to its lack of a cell wall, it does not respond to beta-lactams antibiotics such as amoxicillin, which is the first-line treatment for typical community acquired pneumonia (CAP). Azithromycin is the first line of treatment instead12. Thus, clinicians often initiate empirical antibiotic treatment and adjust it later based on microbiological test results13.
Recent advances in transcriptomics have allowed for the investigation of specific signatures triggered by the host in response to infections by viruses, bacteria14,15 and different pathogens16. Given the outbreaks of M. pneumoniae and the lack of precise and clinically useful diagnostic tools, it is essential to explore the specificity of the gene expression response against M. pneumoniae and other microorganisms responsible for pneumonia. While there have been a few attempts to characterize the gene expression profile in pneumonia patients infected by different microorganisms, none of them have focused specifically on the atypical pneumonia caused by M. pneumoniae. However, we noted that a transcriptomic microarray study by Wallihan et al.17 included 30 children infected by M. pneumoniae, although a specific blood transcriptomic signature for this infection was not investigated.
Considering the urgent need prompted by recent outbreaks, it may be helpful to explore host gene expression mechanisms specifically triggered by M. pneumoniae and evaluate them in the context of those generated from pneumonia caused by other pathogens. Delivering a host-transcriptome signature specific for M. pneumoniae infection would be desirable to overcome the limitations of current clinical and non-clinical procedures.
In this study, we identify and validate eight distinct transcriptomic signatures, consisting of 3 to 10 genes each, that can accurately differentiate M. pneumoniae pneumonia from other bacterial and viral pneumonias in the pediatric population. These signatures represent a significant advancement over previous transcriptomic models, enhancing diagnostic accuracy and offering valuable insights for clinical decision-making.
Results
A robust multi-cohort study design incorporating discovery and validation phases across different platforms was implemented to derive optimal transcriptomic signatures for the specific diagnosis of M. pneumoniae. The overall design is summarized in Fig. 1. Initially, differential gene expression was analyzed between M. pneumoniae and viral datasets from17 (step 1 in Fig. 1). The most promising differentially expressed genes (DEGs) were then used in downstream analyses to identify the best transcriptomic signatures through a systematic resampling approach and penalized regression methods (steps 2–7 in Fig. 1). The top M. pneumoniae signatures were rigorously evaluated for their ability to discriminate between M. pneumoniae and viruses, as well as other pathogens, coinfections, and healthy controls17 (step 8 in Fig. 1). Additionally, the performance of these signatures was benchmarked against other published signatures to compare their discriminative power (step 9 in Fig. 1). Finally, the leading signatures were validated in an independent RNAseq cohort of pneumonia patients, confirming their robustness and clinical utility (step 10 in Fig. 1). Further methodological details are provided in Fig. 1 and the Materials and Methods section.
Differential expression was investigated between M. pneumoniae and virus datasets from17 (step 1). We employed a re-sampling approach (step 2) to generate 1000 transcript signatures of size n (step 3). These signatures were utilized to compute 1000 AUC values on both the training (TA) and test (TE) datasets (step 4), as well as on the non-mycoplasma bacteria and virus datasets from the EUCLIDS (discovery) cohort (step 4 and 5). Next, Signature Scores (SS) were obtained for the different n-transcript signatures (from n = 3 to 10) based on AUCTA, AUCTE, and AUCEU values (step 6), allowing to derive the best signature of size n (step 7); n-SSMAX. AUC values were then computed with the best n-transcript signatures for the comparison between M. pneumoniae vs. virus and M. pneumoniae vs. virus + other pathogens from ref. 17 (step 8). For comparative purposes, we also assessed the performance of other available signatures in the literature (step 9). Finally, the best signatures were subsequently validated in the EUCLIDS (validation cohort), DIAMONDS, and PERFORM cohorts (step 10).
Transcriptomic analysis
We conducted a principal component analysis (PCA) on a subset of the 500 most variable genes with the entire pneumonia cohort, which comprised four different groups: bacterial infection, viral infection, M. pneumoniae infection and co-infections. The first principal component (PC1), explaining 16.85% of the variation, clearly shows the segregation of the M. pneumoniae samples from pneumonias caused by other bacteria. As expected, co-infected samples are distributed evenly between the two groups. Viral profiles intermingle with other profiles in the plot, including M. pneumoniae and bacterial transcriptomes; Fig. 2A. PC2 (accounting for 8.16% of the variation) makes a subtle distinction between bacterial infections on one pole of the component, with most co-infections included in this cluster. M. pneumoniae samples are slightly displayed towards the center of the component, while viral samples are evenly distributed along the whole component, reproducing the same behavior as in PC1 (Fig. 2A).
A PCA of transcriptome profiles from blood samples of viral, M. pneumoniae, co-infections or other bacteria pneumonia infections. The two first principal components (PC1 and PC2) are shown. B Volcano plot showing the DEGs between conditions: M. pneumoniae pneumonia vs. viral pneumonia. The statistical test used to obtain P-values was a two-sided moderated t-test implemented in the limma package. Adjustments for multiple comparisons were performed using the Benjamini-Hochberg procedure to control the false discovery rate (FDR). The plot shows the log2 fold change (Log2FC) on the x-axis and the -log10 adjusted P-value on the y-axis. Genes with an adjusted P-value < 0.05 and a log2FC > |1| were colored in orange, genes with an adjusted P-value < 0.05 but a log2FC < |1| were colored in dark blue, genes with an adjusted P-value > 0.05 but a log2FC > |1| were colored in light blue and non-significant genes with log2 FC < |1| were colored in grey. Only genes included in the final signatures were labeled in the graphic.
In a comparative analysis of transcriptomes in children with pneumonia caused by M. pneumoniae vs. viral pneumonia, we identified 3783 DEGs, using a False Discovery Rate (FDR) significance threshold of 5% (Supplementary Data 1). Among these DEGs, 2288 were found to be upregulated, while 1495 were downregulated (Fig. 2B).
Diagnostic signature discovery
To identify the best minimal signatures for differentiating M. pneumoniae infection from viral infections, we constructed a LASSO model using the top 276 DEGs, applying the criteria of adjusted P-value < 0.01, Log2FC > |1 | , and Log2 Average Expression > 2 (Supplementary Data 1). Subsequently, we generated 1000 different transcriptomic signatures using the 1000 randomized training datasets (TA). We then calculated the area under the receiver operating characteristic curve (AUC) for each signature in the TA dataset, TE dataset and the EUCLIDS cohort (non-mycoplasma bacterial and viral pneumonias). Among the 1000 signatures, we computed the Signature Scores (SS) to obtain the best signatures of size 3 to 10. Eight transcriptomic signatures emerged as the best candidates for distinguishing pneumonia caused by M. pneumoniae from viral pneumonia (Table 1). These signatures include genes in common, totaling 18 different transcripts, with 15 over-expressed and 3 under-expressed in mycoplasma pneumonias compared to pneumonias from viral etiology (Supplementary Fig. 1). Notably, all these signatures demonstrated reliable performance when employed to compare M. pneumoniae samples against all non-mycoplasma samples, including cases of co-infections and other bacterial pneumonias (Supplementary Data 2).
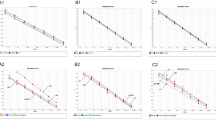
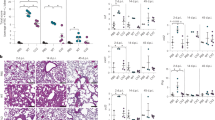
We assessed the performance for each of the eight selected signatures to differentiate between M. pneumoniae and viral pneumonia across all randomized datasets (TA), excluding those used to generate each signature (therefore, 999 subsets). The median AUC values ranged from 0.84 (for the 3-transcript signature) to 0.95 (for the 10-transcript signature) (Fig. 3; Supplementary Data 2), demonstrating the overall high-accuracy of the signatures to discriminate between both phenotypes. We set the optimal cut-off for each signature to optimize the discrimination of the two categories. Additionally, we evaluated each transcriptomic signature in the complete dataset to test their performance in differentiating mycoplasma pneumonias from all other pneumonias, represented with a receiver operating characteristic curve (ROC) and their respective AUC values. (Fig. 3; Supplementary Data 2). Equally, the AUC values from this analysis aligned with the AUC median values observed across all TA datasets.
AUCs and ROC curves from the density plots (left panels) of the AUC values computed on the 999 training resamples and using the optimal n-transcript signature; red dashed vertical lines in the left panels indicate the median values. ROC curves and AUC values (central panels) for the total cohort of i) M. pneumoniae (n = 30) vs. viral pneumonia (n = 77) (black line; AUCTO|M-V), ii) M. pneumoniae pneumonias (n = 30) vs. Bacterial pneumonias (n = 5) (red line; AUCTO|M-B) and iii) M. pneumoniae pneumonias (n = 30) vs. all non-mycoplasma pneumonia infections (including, virus, non-mycoplasma bacteria, and non-mycoplasma co-infections; n = 92) (violet line; AUCTO|M-VBC). Boxplots of the predicted values using each optimal model in the total cohort with two-sided Wilcoxon rank-sum test P-values (right panels). Red dashed line represents the optimal cutpoint. The boxes are defined by the upper and lower quartile (Q1 and Q3); the median is shown as a bold-colored horizontal line; whiskers extend to the most extreme data point which is no more than 1.5 times the interquartile range (IQR) from the box. VBC all non-mycoplasma pneumonias (including, virus, non-mycoplasma bacteria, and non-mycoplasma co-infections), B bacterial, M M. pneumoniae, TO total sample, V viral.
We further tested the accuracy of the signatures in discriminating between M. pneumoniae pneumonia from other clinically relevant groups, including: i) M. pneumoniae pneumonia vs. pneumonias without a detected pathogen, ii) M. pneumoniae pneumonia in coinfection with other pathogens vs. all other pneumonias; and iii) M. pneumoniae pneumonia vs. healthy controls. Notably, the signatures performed exceptionally well in distinguishing M. pneumoniae from healthy controls, with AUC values ranging from 0.87 to 0.98, and from pneumonias without a detected pathogen, with AUC values ranging from 0.81 to 0.90; Supplementary Data 2. As expected, the accuracy of the signatures decreases when tested against pneumonias caused by M. pneumoniae in co-infection with other pathogens, compared to viral and bacterial pneumonias; Supplementary Data 2. However, the 9-transcript signature still yielded AUC values of 0.77 or higher for all comparisons: M. pneumoniae in co-infection vs. viral + bacterial pneumonias (AUC = 0.77 [CI:0.65–0.89]), vs. bacterial pneumonias (AUC = 0.80 [CI:0.57–1.00]) and vs. viral pneumonias (AUC = 0.77 [CI:0.65–0.89]). Interestingly, the 3-transcript signature also showed AUC values above 0.70 in M. pneumoniae co-infection scenarios, achieving its highest performance when distinguishing from bacterial pneumonias (AUC = 0.83 [CI:0.63–1.00]).
Validation of M. pneumoniae signatures in an independent cohort
The diagnostic accuracy of the proposed RNA signatures was evaluated using additional gene expression data generated from a new pediatric cohort of pneumonia samples and a different technology (RNAseq); see details in Supplementary Data 3 and Supplementary Data 4 The results confirmed that all RNA signatures can discriminate between viral and M. pneumoniae pneumonias with AUCs higher than 0.68 (7-transcript), being the 9-transcript signature the most predictive (AUC = 1.00 [CI: 1.00–1.00]; sensitivity: 1.00; specificity: 1.00); Fig. 4, Supplementary Data 5. Unexpectedly, the third most predictive model was the 3-transcript signature (AUC = 0.86 [CI: 0.65–1.00]; sensitivity: 0.89; specificity: 0.90) (Fig. 4, Supplementary Data 5). Remarkably, all signatures can also accurately differentiate M. pneumoniae pneumonia from other bacterial pneumonias, with AUCs ranging from 0.74 (CI: 0.55–0.92) for the 4-transcript signature to 0.85 (CI: 0.73–0.99) for the 10-transcript signature. Considering all viral and non-mycoplasma bacterial pneumonias against M. pneumoniae pneumonias, the AUC values were similar to those obtained from the comparison M. pneumoniae vs. bacterial pneumonia. In this case, the 9-transcript signature yielded the best performance of all the signatures tested (AUC = 0.89 [CI: 0.79–0.99]); Fig. 4, Supplementary Data 5.
ROC curves with AUC values for the three tested validation subsets: M. pneumoniae (n = 9) vs. viral pneumonias (n = 10) (black line; AUCM-V), M. pneumoniae (n = 9) vs. non-mycoplasma pneumonias including viral and other bacterial pneumonias (n = 32) (violet line; AUCM-VB) and M. pneumoniae vs. non-mycoplasma bacterial pneumonias (n = 22) (red line; AUCM-B) (left panels). Boxplots of the predicted values using each model in the three validation subsets (right panels). Two-sided Wilcoxon P-values are also displayed. The red dashed line represents the optimal cut off. The boxes represent the distribution of scores for M. pneumoniae, viral and bacterial pneumonia samples. They are defined by the upper and lower quartiles (Q1 and Q3), with the median displayed as a bold-colored horizontal line. The whiskers extend to the most extreme data points within 1.5 times the interquartile range (IQR) from the box. Abbreviations as in legend of Fig. 3. VB all non-mycoplasma pneumonias including viral and other bacterial pneumonias, B bacterial, M M. pneumoniae, V viral.
As in the case of the discovery dataset, we determined the optimal cut-off for each signature in the validation that establishes the threshold to discriminate between pneumonias from different etiologies in the three comparisons groups (Fig. 4; Supplementary Data 5).
Additionally, we also validated the eight signatures in other groups, some of which are clinically relevant, including pneumonias diagnosed without a detected pathogen, controls with asthma and healthy controls. The AUC values obtained were: i) from 0.76 (4-transcripts signature) to 0.85 (9-transcripts signature) for the contrast M. pneumoniae pneumonia vs. pneumonias diagnosed without a detected pathogen, ii) from 0.68 (7-transcript signature) to 0.86 (6-transcripts signature) for the contrast M. pneumoniae pneumonia vs. controls with asthma, and iii) from 0.83 (4-transcripts signature and 9-transcripts signature) to 0.96 (8-transcripts signature) for the contrast M. pneumoniae pneumonia vs. healthy controls (Supplementary Data 5).
Performance of other available signatures in M. pneumoniae samples
As M. pneumoniae pediatric pneumonias appear to induce a different alteration in the transcriptome compared with other bacterial infections causing pneumonia in children, we opted to examine the performance of two available signatures designed to differentiate viral and bacterial pediatric infections in samples from children with viral pneumonia and pneumonia caused by M. pneumoniae: the two-transcript signature (IFI44L/FAM89A), capable of differentiating between pediatric viral from bacterial infections14; and the novel 5-transcript signature18, designed to specifically distinguish between viral and bacterial CAP in children. We observed that neither of these two tested signatures could effectively diagnose M. pneumoniae as a bacterial infection or a bacterial pneumonia, yielding AUC values of 0.56 and 0.52, respectively (Supplementary Fig. 2).
Differentially regulated pathways between M. pneumoniae and viral pneumonia
To detect biological pathways responsible for the different response to M. pneumoniae and viral pneumonia in our study cohort, we performed a GSVA analysis directly on gene expression data. We identify 525 significantly (adjusted P-value < 0.05) differentially regulated pathways (DRPs) between both categories, with over half upregulated in M. pneumoniae pneumonia (358/525; 68%); Supplementary Data 6; Fig. 5A. Among the top 20 pathways, the most notable changes in pathways activity were mainly represented by up-regulated processes in atypical pneumonia (18/20; 90%); Supplementary Fig. 3.
A Volcano plot showing the genes in DRPs between conditions: M. pneumoniae pneumonia (n = 30) vs. viral pneumonia (n = 77). The statistical test used to obtain P-values was a two-sided moderated t-test implemented in the limma package. Adjustments for multiple comparisons were performed using the Benjamini-Hochberg procedure to control the false discovery rate (FDR). The plot shows the log2 fold change (Log2FC) on the x-axis and the -log10 adjusted P-value on the y-axis. Pathways meeting the criteria of an adjusted P-value < 0.05 and a log2FC > |0.1| were colored in yellow, pathways with an adjusted P-value < 0.05 but a log2FC < |0.1| were colored in dark blue, genes with an adjusted P-value > 0.05 but a log2FC > |0.1| were colored in light blue and non-significant genes were colored in grey. B Boxplots representing the pathway activity of the DRPs for the M. pneumoniae (n = 30) and viral pneumonia (n = 77) groups in which genes from the transcriptomic signatures are involved (left panel). Two-sided Wilcoxon rank-sum test P-values are displayed. The boxes are defined by the upper and lower quartiles (Q1 and Q3), with the median displayed as a bold-colored horizontal line. The whiskers extend to the most extreme data points within 1.5 times the interquartile range (IQR) from the box. Bubble plot of the DRPs in which genes included in the transcriptomic signatures are involved (right panel). Numeric identifies indicate DRPs in which genes from the transcriptomic signatures are involved. The statistical test used to obtain P-values was a two-sided moderated t-test implemented in the limma package. Adjustments for multiple comparisons were performed using the Benjamini-Hochberg procedure to control the false discovery rate (FDR). Size of the points is proportional to the -log10 adjusted P-value and color bar indicates the log2FC values.
The most DRP was “SRP dependent co-translational protein targeting to membrane signal sequence recognition” (adjusted P-value = 1 × 10−06) followed by ”interleukin-12 mediated signaling” pathways (adjusted P-value = 2 × 10−08), and a group of biological routes related to polysaccharides and glycolipids synthesis, including “nucleotide sugar biosynthetic process” (adjusted P-value = 9 × 10−05), “UDP N-Acetylglucosamine metabolic process” (adjusted P-value = 9 × 10−05), “amino sugar biosynthetic process” (adjusted P-value = 2 × 10−05), “UDP N-Acetylglucosamine biosynthetic process” (adjusted P-value = 4 × 10−05) and “GDP-mannose metabolic process” (adjusted P-value = 9 × 10−05); Supplementary Fig. 3; Supplementary Data 6.
Afterwards, we investigated the functional involvement of the genes included in the transcriptomic signatures by examining the DRPs related to these genes. We found that 10 out of the 18 genes (UAP1, PTGDR2, CAV1, HYOU1, GALM, GAS6, GINS4, CDCA2, MAPRE3, DIO3; Supplementary Data 7, Fig. 5B) participate in 9 significantly DRPs. Notably, among these pathways, “the nucleotide sugar biosynthetic process” emerged as one of the most significant pathways in the overall analysis (adjusted P-value = 9 × 10−05), Fig. 5B. Additionally, the involvement of two of the predictive genes in the same DRP (CAV1 and HYOU1) was only detected for the “response to endoplasmic reticulum stress process” (adjusted P-value = 0.002).
Discussion
Pneumonia etiology in children is highly diverse and typically challenging to effectively assess in the clinical practice. The increase of cases of M. pneumoniae from May 2023 raised concerns within the clinical community. Infections caused by this pathogen require specific treatment protocols, as they do not respond to standard antibiotic prescriptions typically employed when bacterial pneumonia is suspected12,13. Current diagnostic tests can suffer from long turnaround times and may have limited sensitivity and specificity. PCR-based diagnostic tests for mycoplasma detection, while highly sensitive and specific, have become more widely used in developed countries, especially following the SARS-CoV-2 pandemic. However, PCR tests are ineffective to discern between a M. pneumoniae active infection and asymptomatic carriage, which is common in children’s upper respiratory tract19. They also struggle in cases of coinfections, where M. pneumoniae may not be the primary pathogen. Given that M. pneumoniae pneumonia requires tailored treatment, misdiagnosis can have serious consequences for patient care. A host-based diagnosis using specific transcriptomic signatures can help to address the limitations of the pathogen-based tests by improving the ability to distinguish between carriage and active infection in M. pneumoniae pneumonia cases. Our analysis shows that transcriptomic-based signatures available in the literature, previously proven to be effective to broadly differentiate between viral and bacterial infections or specifically pneumonia from viral and bacterial etiology, do not function properly for discriminating M. pneumoniae pneumonia cases from viral pneumonias in children. Our findings also indicate that M. pneumoniae pneumonia alters the expression of >3000 genes in blood of affected children, enabling the inference of eight new specific signatures capable of specifically differentiating pneumonia caused by M. pneumoniae from viral pneumonia. Notably, the performance of these signatures does not decline when used to discriminate between M. pneumoniae infections from other bacterial pneumonias or non-mycoplasma co-infections, ensuring high specificity in detecting M. pneumoniae pneumonias. The identification of these specific transcriptomic signatures capable of diagnosing M. pneumoniae pneumonias in children holds promise for enhancing the diagnosis and treatment for these patients, who cannot be identified using the currently available signatures. Most notably, these signatures could guide the decision to administer a specific macrolide instead of a beta-lactam antibiotic. As all the n-transcript signatures exhibit high sensitivities, the test would ensure the detection of most M. pneumoniae infected patients and improve their management.
Employing an innovative simulation-driven theoretical framework, we uncovered eight distinct transcriptomic signatures, each representing the optimal combinations of 3 to 10 transcripts, capable of differentiating M. pneumoniae pneumonia from other bacterial and viral pneumonias with high accuracy (AUC: 0.84–0.95). These also signatures demonstrated strong discriminatory power in other clinically relevant groups, such as coinfections (with or without M. pneumoniae), pneumonias without a detected pathogen, and healthy controls. This broad applicability makes the signatures valuable not only for distinguishing M. pneumoniae pneumonia from viral pneumonia but also for a variety of common clinical scenarios.
The robustness of these signatures was confirmed in an independent validation cohort, which included viral and bacterial pneumonias, pneumonias without a detected pathogen, controls with asthma and healthy controls. While our theoretical model (Fig. 3) predicted that signature performance would improve with a greater number of predictors, in the validation cohort, the 9-transcript signature exhibited the best performance, but the 3-transcript signature also displayed unexpectedly high accuracy.
The choice and usefulness of these signatures would completely depend on different factors of the clinical context: available equipment, budget, number of samples, urgency for results, etc. Few technologies can accurately measure that number of transcripts simultaneously at a reasonable cost and in a timely fashion. For instance, RNA sequencing is not a suitable choice because, while it is time- and cost-effective for high-throughput gene expression, it is impractical when studying only a limited number of target genes. However, NanoString (www.NanoString.com/) offers high precision and is much more convenient when targeting a limited number of genes. Unfortunately, these platforms are often reserved for research purposes and are not typically found in the daily laboratory routine of medical centers. Conventional RT-qPCR is the gold standard for these cases, but can be time-consuming, laborious, and less cost-effective, especially when the number of target genes and the sample size increase. New PCR approaches, such as 1-step digital PCR (dPCR), allow for the combination of up to 5 genes, with a high sensibility and bypassing certain steps20, and would be an option to study the biomarkers of some of the proposed signatures. On the other hand, reducing the number of predictors increases the likelihood of transferability to a point-of-care (POC) device, and the signatures with the lower number of transcripts are the most promising candidates for implementation in this type of tests, specially the 3-transcript signature, which shows remarkably accurate performance compared to other signatures with a larger number of predictors. The use of compact signatures would facilitate the translation to a rapid diagnostic POC test enhancing its widespread adoption and leading to reduced costs and time.
Differential pathway analysis also reveals different processes that are over-activated in M. pneumoniae compared to viral pneumonias causing pneumonia. Many of these significantly altered pathways are related to nucleotide sugar metabolism, which is closely associated with human infections due to its involvement in surface carbohydrate biosynthesis, protein glycosylation, and glycosylated lipid biosynthesis; all essential processes for pathogens’ ability to infect and survive within the human host21,22,23. Nucleotide sugars function as glycosyl donors for glycosylation reactions in proteins catalyzed by the glycosyltransferases. Mycoplasmas, with their small genome, cannot synthesize some essential biomolecules de novo, relying on obtaining them from the host with which they live in close association. UDP-N-acetylglucosamine (GlcNAc), a nucleotide sugar, is one of the major carbon sources utilized by several human pathogens, including M. pneumoniae, and its catabolism is also involved in the regulation of virulence and pathogenesis24. M. pneumoniae has been reported to produce biofilm on the epithelial surface, contributing to evasion of host immune system and providing resistance to antibiotics25,26. Biofilms, complex communities of microorganisms, typically bacteria, encased within a self-produced extracellular matrix, can contribute to persistent infections27. Polysaccharides containing GlcNAc contribute to the structure and virulence of these biofilms in different microorganisms28 including M. pneumoniae.
Another significantly altered pathway is the “Interleukin-12 (IL-12) mediated signalling pathway”, found to be over-regulated in M. pneumoniae compared with viral pneumonia infection. IL-12 plays an immune regulatory role in innate and adaptive responses during infection, triggering the production of interferon-γ and promoting the differentiation of type 1 T helper cells29. Elevated levels of IL-12 have been correlated with the severity of pneumonia in M. pneumoniae infections and slower microbiologic and histological recovery in mice, suspected to be related to a boost in the innate immune host response30.
Remarkably, some of the pathways involving genes included in our signatures, are among or related to the top 20 most significant pathways from the global analysis of M. pneumoniae vs. viral pneumonias (Supplementary Fig. 3, Supplementary Data 7). This suggests an important role of these genes in the host-response to M. pneumoniae infection. Nucleotide sugar biosynthesis is one of the most significantly up-regulated pathways in the global analysis and, as described above, is closely related to pathogens’ metabolism. The endoplasmic reticulum stress or the endoplasmic reticulum unfolded protein response is known to be involved in the innate immune signalling response to different pathogens31. Pathogens hijack endoplasmic reticulum functions for their own benefit leading to a stress state in host cells. The unfolded protein response mitigates the endoplasmic reticulum stress, but if it cannot counterbalance this stress, apoptotic and autophagic processes are activated, contributing to the disease pathogenesis. Other pathways related to the regulation of cell cycle, specifically around the mitotic phase, are largely describe to be altered in viral processes32.
Although our results show evidence of biological processes differentially activated in M. pneumoniae pneumonia, the specific contribution of these pathways to the M. pneumoniae pathogenesis needs further exploration.
There are a few limitations in the present study. First, while we analyze the largest cohorts of M. pneumoniae infections to date, the sample size remain limited, highlighting the need to explore additional cohorts and sub-phenotypes, such as patients with respiratory symptoms of non-infectious etiology. Second, our signatures were derived from microarray data, leaving room for the discovery of potentially superior signatures once RNAseq data become available. Third, technical validations of these signatures using gold standard methods such as dPCR or qPCR would be valuable, as well as exploring the possibility of translating these signatures into a POC diagnostic tool. Fourth, since gene expression patterns can vary across populations33, it would be advantageous to examine the performance of the signatures in diverse populations from different geographic regions.
This study emphasizes the urgent need for expanded research on M. pneumoniae, providing compelling evidence of this microorganism’s unique ability to elicit a distinct gene expression response in infected pneumonia patients. By leveraging this pathogen-specific reaction, we successfully derived flexible gene expression signatures, comprising 3 to 10 transcripts. Validation in an independent cohort of patients provided further evidence indicating their specificity and sensitivity in distinguishing M. pneumoniae from other viral, bacterial and undiagnosed pneumonias. This breakthrough enables accurate differentiation between pneumonia caused by M. pneumoniae and other pneumonias of different etiologies, providing a framework for appropriate treatment and offering promising avenues for further exploration and potential clinical applications.
Methods
Samples and study design
We selected microarray blood transcriptomic profiles from 151 children hospitalized with pneumonia and 20 healthy controls as reported by Wallihan et al. 17. The pneumonia cases included infections caused by M. pneumoniae (n = 30), various viral infections (n = 77), pyogenic bacteria (n = 5), co-infections not involving M. pneumoniae (n = 10), co-infections involving M. pneumoniae (n = 11), and pneumonia infections without detected pathogen (n = 18); Fig. 1. The data were downloaded from the Gene Expression Onmibus (GEO) database with accession number GSE103119. Inclusion criteria required evidence of acute infection, signs or symptoms of respiratory illness, and radiologic confirmation of lower respiratory tract disease. For more detailed information on the inclusion and exclusion criteria, see Supplemental Table 1 in Wallihan et al. 17.
Additionally, we have used RNAseq data from blood of pediatric patients with viral and bacterial pneumonia not caused by M. pneumoniae recruited, recruited through the European Union Childhood Life-threatening Infectious Diseases Study (EUCLIDS- https://www.diamonds2020.eu/our-research-history/euclids/34,35,36 consortium available in GEO and ArrayExpress under accession numbers GSE261482/E-MTAB-11671); Fig. 1. Specifically, the EUCLIDS discovery cohort contains RNAseq data for 39 definitive non-mycoplasma bacterial and 9 definitive viral pneumonias, all of them used and reported by Viz-Lasheras et al.18(clinical data available in Table 118); see definitions in ref. 14.
For validation, we used new RNAseq data from an additional pediatric cohort comprising blood samples obtained from children infected with M. pneumoniae (n = 9), samples from viral (n = 10) and bacterial (n = 22) pneumonias, samples from pneumonias with no causal pathogen detected (n = 57) along with healthy controls (n = 35) and controls with asthma (n = 8); Fig. 1. Clinical features and demographic information of this cohort are provided in Supplementary Data 3. These samples were recruited under the umbrella of i) the EUCLIDS Study (EUCLIDS validation), ii) the PErsonalised Risk assessment in Febrile illness to Optimise Real-life Management across the European Union (PERFORM - https://www.perform2020.org/) and iii) the Diagnosis and Management of Febrile Illness using RNA Personalised Molecular Signature Diagnosis (DIAMONDS - https://www.diamonds2020.eu) consortiums16,37,38 and are available in ArrayExpress under accession numbers E-MTAB-12793, E-MTAB-14564 and E-MTAB-14588. Pneumonia phenotype in the non-mycoplasma pneumonia cohort (EUCLIDS discovery), and in the validation cohort (EUCLIDS validation-PERFORM-DIAMONDS) was defined as follows: i) Inflammation of one or both lungs (lobar, segmental or multilobar collapse/consolidation) on chest X-ray (excluding perihilar consolidation or patchy consolidation); ii) Clinical symptoms compatible with acute respiratory infection and radiological evidence of consolidation or pleural effusion: alveolar consolidation (dense or fluffy opacity occupying part or all of a lobe or lung, with or without air-bronchograms) or pleural effusion (fluid in the lateral pleural space, excluding minor or oblique fissure), spatially associated with a pulmonary parenchymal infiltrates (including other infiltrate) or obliterating enough of the hemithorax to obscure opacity. All samples were collected within 24 h of admission. M. pneumoniae pneumonia was diagnosed using PCR and/or serology in throat swab samples along with clinical symptoms compatible with atypical pneumonia (Supplementary Data 8).
Inclusion and ethics statement
Harmonized procedures for patient recruitment, classification, clinical data and sample collection, processing and storage were followed across the participating centers. Written, informed consent was obtained from parents or guardians at all sites using locally approved permissions. See study protocol EUCLIDS: see DOI: 10.1016/S2352-4642(18)30113-5; study protocol PERFORM: see DOI: 10.1093/cid/ciad615; study protocol DIAMONDS: see https://doi.org/10.1186/ISRCTN12394803. Each participating country’s Ethics Committee (EC) approval is available upon request. The study conformed to the ethical principles for medical research involving human participants outlined in the Declaration of Helsinki. This study was designed and implemented by clinicians and researchers embedded within treating clinical teams and this is reflected in the opportunities to contribute meaningfully to the data analysis, interpretation, and writing. This is reflected in the authorship. Members of the wider EUCLIDS, PERFORM and DIAMONDS Search Study consortia are identified in the Supplementary Information.
RNAseq analysis
Whole blood was collected into PAXgene blood RNA tubes (BD Biosciences; catalog number: 762165) and stored at − 80 °C. For the EUCLIDS cohort, total RNA was isolated using PAXgene blood isolation kit according to the manufacturer’s instructions (Qiagen; catalog number: 762174). RNA sequencing was conducted on a HiSeq 4000 (Illumina) platform, with library preparation and sequencing of 30 million 75 or 100 bp paired-end reads. The Illumina’s TruSeq RNA Sample Preparation Kit was used for library preparation, and ribosomal and globin RNA depletion was performed using the Illumina® Ribo-Zero Gold kit (Illumina; catalog number: MRZG12324). For the PERFORM/DIAMONDS cohort, RNA was isolated using PAXgene blood miRNA isolation kit according to the manufacturer’s instructions (Qiagen; catalog number: 763134). An additional DNAse treatment was carried out with the RNA clean & concentrator kit (Zymo Research; catalog number: R1013) prior to sequencing. RNA was quantified using RiboGreen (Invitrogen; catalog number: R11490) on the FLUOstar OPTIMA plate reader (BMG Labtech) and the integrity analyzed on the TapeStation 2200 (Agilent, catalog number: 5067–5576; 5067–5578; 5067–5577). After a normalization step, a strand specific library preparation was completed using NEBNext® Ultra™ II mRNA kit (New England Biolabs; catalog number: E7770) and NEB rRNA/globin depletion probes following manufacturer’s recommendations (New England Biolabs; catalog number: E7750). Individual libraries were normalized using Qubit, pooled together and diluted. The sequencing was performed using a 150 paired-end configuration in a Novaseq6000 platform (Illumina). Quality control of raw data was carried out using FastQC39, alignment and read counting were performed using STAR40, alignment filtering was done with SAMtools41 and read counting was carried out using FeatureCounts42.
Statistical analysis
Microarray data pre-process and normalization was performed using the illuminaHumanv4.db43 and limma44 packages. RNAseq data was processed for batch correction using control samples and COMBAT-Seq45 package. Data was subsequently normalized with DESeq246 package. Principal component analysis (PCA) was conducted to explore the different groups in the data and check for potential outliers. A differential expression (DE) analysis was carried out using limma44 package and accounting for differences in age, in order to compare the blood transcriptome of pneumonia patients with M. pneumoniae infection (n = 30) vs. viral pneumonia infections (n = 77); step 1 in Fig. 1.
In order to identify subsets of genes that could serve as predictive transcriptomic signatures differentiating M. pneumoniae pneumonia from other pneumonias of viral etiology, we randomly split the dataset (n = 10717); into 1000 independent subsets comprising 70% of the samples (TA datasets) and other 1000 subsets containing the remaining 30% of the samples (TE datasets); step 2 in Fig. 1. A predictive transcriptomic signature was computed using the R package glmnet47 for each of the 1000 TA. To do that, a logistic LASSO regression model was fitted with the alpha parameter set to 1 and a 10-fold cross validation (step 3 in Fig. 1); 276 differentially expressed genes (DEGs) were included as input for the logistic regression based on |Log2FC | > 1, adjusted P-value < 0.01, and a Log2 average expression > 2 (Supplementary Data 1).
The accuracy of the predictive transcriptomic signatures was measured by calculating the area under the receiver operating characteristic curve (AUC) with 95% confidence intervals (CI) using the pROC package48. The optimal cut-point value (cut-off) that maximize sensitivity and specificity, was calculated using the OptimalCutPoints R package49. AUC values were computed for the 1000 TA and the corresponding TE datasets (step 4 in Fig. 1), and also for the non-M. pneumoniae viral and bacterial pneumonias in the EUCLIDS cohort; step 5 in Fig. 1. Among the 1000 signatures of size n transcripts, we selected those i signatures of size n = 3 to 10 transcripts. Next, on these selected i signatures, we computed the score n-SSi = (0.4 × AUCTA) + (0.4 × AUCTE) – (0.2 × AUCEU) (step 6 in Fig. 1); this scoring method considers the AUC value for both training and test samples, while penalizing performance on non-M. pneumoniae pneumonia infections in the EUCLIDS cohort. We selected the best transcriptomic signature of size n = 3 to 10 (n-SSMAX) among the different signatures with the same number of transcripts. The AUC values for the signatures with the n-SSMAX values, were then obtained in the 999 complete training resamples (1000 minus the one used to generate the n-SSMAX); step 7 in Fig. 1. In addition, the robustness of the best signatures to differentiate M. pneumoniae pneumonia from all sub-groups included in the original pneumonia dataset17 was evaluated by calculating the AUC values in the following contrasts: i) M. pneumoniae pneumonia vs. viral pneumonia, ii) vs. bacterial pneumonia, iii) vs. viral pneumonia + non-mycoplasma bacterial pneumonia + non-mycoplasma co-infected pneumonias, iv) vs. M. pneumoniae pneumonia co-infections, v) vs. pneumonia without detected pathogen, and vi) vs. healthy controls; step 8 in Fig. 1.
The performance of other signatures available in the literature to differentiate M. pneumoniae from viral pneumonias were also investigated. In particular, we tested the 5-transcript signature recently developed to differentiate non-Mycoplasma pneumonia from viral pneumonia18, and the 2-transcript signature developed to differentiate viral from bacterial infections14; step 9 in Fig. 1.
Validation of the best signatures
The performance and accuracy of the predictive transcriptomic signatures were validated using new blood RNAseq data generated from an additional pediatric cohort, comprising new samples from EUCLIDS and PERFORM-DIAMONDS cohorts. Coefficients and intercepts from each LASSO model were applied to the new dataset to perform ROC analysis and calculate the AUC, sensitivity, and specificity of the signatures. Six different comparisons were tested using the validation cohort: i) M. pneumoniae pneumonia vs. viral pneumonia, ii) vs. bacterial pneumonia, iii) vs. both viral and non-mycoplasma bacterial pneumonias, iv) vs. pneumonias without detected pathogen; v) vs. controls with asthma, and vi) vs. healthy controls, step 10 in Fig. 1.
GSVA pathway analysis
Biological pathways differentially involved in viral and M. pneumoniae pneumonia were inferred from gene expression data using the GSVA algorithm included in the Gene Set Variation Analysis (GSVA) R package50. Gene Ontology (GO) biological pathways gene set collection from the Molecular Signatures Database (MSigDB)51 was used as reference database. Significantly differentially regulated pathways (DRPs) were determined using the limma package44 with the viral groups as a reference and a threshold of adjusted P-value < 0.05.
All graphics were created using R software v.4.3.2 (www-r-project.org).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
New RNAseq data generated in this study and used for validation are available in ArrayExpress repository (https://www.ebi.ac.uk/biostudies/arrayexpress) under accession number E-MTAB-14588. RNASeq data previously analyzed in other studies are available in ArrayExpress under accession numbers E-MTAB-12793 and E-MTAB-14564 and in Gene Expression Omnibus NCBI (GEO; http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE261482. Details are provided in Supplementary Data 4.
Code availability
Original R code has been deposited at github (https://github.com/Sandraviz/Mycoplasma-pneumoniae_-Transcriptomic-Signatures.git), https://doi.org/10.5281/zenodo.1425874852.
References
Larcher, R., Boudet, A., Roger, C., Villa, F. & Loubet, P. Mycoplasma pneumoniae is back! Is it the next pandemic? Anaesth Crit Care. Pain. Med. 43, 101338 (2024).
Meyer Sauteur, P. M. et al. Mycoplasma pneumoniae detections before and during the COVID-19 pandemic: results of a global survey, 2017 to 2021. Euro Surveill. 27, 2100746 (2022).
Meyer Sauteur, P. M. et al. Mycoplasma pneumoniae: delayed re-emergence after COVID-19 pandemic restrictions. Lancet Microbe 5, e100–e101 (2024).
Urbieta, A. D. et al. Mycoplasma pneumoniae at the rise not only in China: rapid increase of Mycoplasma pneumoniae cases also in Spain. Emerg. Microbes Infect. 13, 2332680 (2024).
Nordholm, A. C. et al. Mycoplasma pneumoniae epidemic in Denmark, October to December, 2023. Euro Surveill. 29, 2300707 (2024).
Wreghitt, T. Mycoplasma pneumoniae: current outbreak. Epidemiol. Infect. 152, e47 (2024).
Bolluyt, D. C. et al. Increased incidence of Mycoplasma pneumoniae infections and hospital admissions in the Netherlands, November to December 2023. Euro Surveill. 29, 2300724 (2024).
Parums, D. V. Editorial: Outbreaks of post-pandemic childhood pneumonia and the re-emergence of endemic respiratory infections. Med. Sci. Monit. 29, e943312 (2023).
Daxboeck, F., Krause, R. & Wenisch, C. Laboratory diagnosis of Mycoplasma pneumoniae infection. Clin. Microbiol. Infect.: Off. Publ. Eur. Soc. Clin Microbiol Infect Dis. 9, 263–273 (2003).
Landry, M. L. Immunoglobulin M for acute infection: True or false? Vaccin. Immunol. 23, 40–45 (2016).
Zhang, L., Zong, Z. Y., Liu, Y. B., Ye, H. & Lv, X. J. PCR versus serology for diagnosing Mycoplasma pneumoniae infection: a systematic review & meta-analysis. Indian J. Med. Res. 134, 270–280 (2011).
Atkinson, T. P., Balish, M. F. & Waites, K. B. Epidemiology, clinical manifestations, pathogenesis and laboratory detection of Mycoplasma pneumoniae infections. FEMS Microbiol. Rev. 32, 956–973 (2008).
Rueda, Z. V. et al. Etiology and the challenge of diagnostic testing of community-acquired pneumonia in children and adolescents. BMC Pediatr. 22, 169 (2022).
Herberg, J. A. et al. Diagnostic test accuracy of a 2-transcript host RNA signature for discriminating bacterial vs viral infection in febrile children. JAMA 316, 835–845 (2016).
Barral-Arca, R., Pardo-Seco, J., Martinón-Torres, F. & Salas, A. A 2-transcript host cell signature distinguishes viral from bacterial diarrhea and it is influenced by the severity of symptoms. Sci. Rep. 8, 8043 (2018).
Habgood-Coote, D. et al. Diagnosis of childhood febrile illness using a multi-class blood RNA molecular signature. Med 4, 635–654.e635 (2023).
Wallihan, R. G. et al. Molecular distance to health transcriptional score and disease severity in children hospitalized with community-acquired pneumonia. Front. Cell Infect. Microbiol. 8, 382 (2018).
Viz-Lasheras, S. et al. A 5-transcript signature for discriminating viral and bacterial etiology in pediatric pneumonia. iScience https://doi.org/10.1016/j.isci.2025.111747 (2025).
Spuesens, E. B. et al. Carriage of Mycoplasma pneumoniae in the upper respiratory tract of symptomatic and asymptomatic children: an observational study. PLoS Med. 10, e1001444 (2013).
Wainman, L. M., Sathyanarayana, S. H. & Lefferts, J. A. Applications of digital polymerase chain reaction (dPCR) in molecular and clinical testing. J. Appl. Lab. Med. 9, 124–137 (2024).
Daubenspeck, J. M., Jordan, D. S., Simmons, W., Renfrow, M. B. & Dybvig, K. General N-and O-Linked glycosylation of lipoproteins in mycoplasmas and role of exogenous oligosaccharide. PLoS One 10, e0143362 (2015).
Klement, M. L., Ojemyr, L., Tagscherer, K. E., Widmalm, G. & Wieslander, A. A processive lipid glycosyltransferase in the small human pathogen Mycoplasma pneumoniae: involvement in host immune response. Mol. Microbiol. 65, 1444–1457 (2007).
Jordan, D. S., Daubenspeck, J. M., Laube, A. H., Renfrow, M. B. & Dybvig, K. O-linked protein glycosylation in Mycoplasma. Mol. Microbiol. 90, 1046–1053 (2013).
Ansari, S., Kumar, V., Bhatt, D. N., Irfan, M. & Datta, A. N-Acetylglucosamine sensing and metabolic engineering for attenuating human and plant pathogens. Bioengineering 9, 64 (2022).
Simmons, W. L. et al. Type 1 and type 2 strains of Mycoplasma pneumoniae form different biofilms. Microbiol 159, 737–747 (2013).
Lluch-Senar, M. et al. Comparative “-omics” in Mycoplasma pneumoniae Clinical Isolates Reveals Key Virulence Factors. PLoS One 10, e0137354 (2015).
Branda, S. S., Vik, S., Friedman, L. & Kolter R. Biofilms: the matrix revisited. Trends Microbiol. 13, 20–26 (2005).
Chen, M., Yu, Q. & Sun, H. Novel strategies for the prevention and treatment of biofilm related infections. Int. J. Mol. Sci. 14, 18488–18501 (2013).
Brombacher, F., Kastelein, R. A. & Alber, G. Novel IL-12 family members shed light on the orchestration of Th1 responses. Trends Immunol. 24, 207–212 (2003).
Salvatore, C. M., Fonseca-Aten, M., Katz-Gaynor, K., Gomez, A. M. & Hardy, R. D. Intranasal interleukin−12 therapy inhibits Mycoplasma pneumoniae clearance and sustains airway obstruction in murine pneumonia. Infect. Immun. 76, 732–738 (2008).
Choi, J. A. & Song, C. H. Insights into the role of endoplasmic reticulum stress in infectious diseases. Front. Immunol. 10, 3147 (2019).
Bagga, S. & Bouchard, M. J. Cell cycle regulation during viral infection. Methods Mol. Biol. 1170, 165–227 (2014).
Barral-Arca, R., Pardo-Seco, J., Bello, X., Martinón-Torres, F. & Salas, A. Ancestry patterns inferred from massive RNAseq data. RNA 25, 857–868 (2019).
Borghini, L. et al. Identification of regulatory variants associated with genetic susceptibility to meningococcal disease. Sci. Rep. 9, 6966 (2019).
Martinon-Torres, F. et al. Life-threatening infections in children in Europe (the EUCLIDS Project): a prospective cohort study. Lancet Child. Adolesc. Health 2, 404–414 (2018).
Kumar, V. et al. Variation in CFHR3 determines susceptibility to meningococcal disease by controlling factor H concentrations. Am. J. Hum. Genet. 109, 1680–1691 (2022).
Moradi Marjaneh, M. et al. Analysis of blood and nasal epithelial transcriptomes to identify mechanisms associated with control of SARS-CoV-2 viral load in the upper respiratory tract. J. Infect. 87, 538–550 (2023).
Jackson, H. et al. Characterisation of the blood RNA host response underpinning severity in COVID-19 patients. Sci. Rep. 12, 12216 (2022).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Available online. Retrieved May 17, 2018 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Dunning, M., Lynch, A. & Eldridge, M. illuminaHumanv4.db: Illumina HumanHT12v4 annotation data (chip illuminaHumanv4). R package version 1.26.0. (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Zhang, Y., Parmigiani, G. & Johnson, W. E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR. Genom. Bioinform 2, lqaa078 (2020).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with. DESeq2. Genome Biol. 15, 550 (2014).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform 12, 77 (2011).
López-Ratón, M., Rodríguez-Álvarez, M. X., Cadarso-Suárez, C. & Gude-Sampedro, F. OptimalCutpoints: An R package for selecting optimal cutpoints in diagnostic tests. J. Stat. Softw. 61, 1–36 (2014).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 14, 7 (2013).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Viz-Lasheras, S., Gómez-Carballa, A., Bello, X., F., M.-T. & Salas, A. A diagnostic host-specific transcriptome response for Mycoplasma pneumoniae pneumonia to guide pediatric patient treatment. https://doi.org/10.5281/zenodo.14258748 (2024).
Acknowledgements
This research project was made possible through the access granted by the Galician Supercomputing Center (CESGA) to its supercomputing infrastructure. The supercomputer FinisTerrae III and its permanent data storage system have been funded by the Spanish Ministry of Science and Innovation, the Galician Government, and the European Regional Development Fund (ERDF). This study also received support by i) ISCIII: TRINEO: PI22/00162; DIAVIR: DTS19/00049; Resvi-Omics: PI19/01039 (to A.S.), ReSVinext: PI16/01569, Enterogen: PI19/01090, OMI-COVI-VAC: PI22/00406 (to F.M.-T.), cofinanciados FEDER, ii) GAIN: IN607B 2020/08 and IN607A 2023/02 (to A.S.), GEN-COVID (IN845D 2020/23 (to F.M.-T.), IIN607A2021/05 (to F.M.-T.) and IN677D 2024/06 (to A.G.-C.); iii) ACIS: BI-BACVIR (PRIS-3, to A.S.), CovidPhy (SA 304 C, to A.S.), PneumoTrack (PRIST-VAL, to A.S.), and Respisal (PRIST-VAL, to F.M.-T.); v) consorcio Centro de Investigación Biomédica en Red de Enfermedades Respiratorias (CB21/06/00103; to A.S. and F.M.-T.) and vi) Spanish Ministry of Science and Innovation (MCIN)/Spanish Research Agency (AEI) (KAWA-TesT: PID2022-142156OB-I00, to A.G-.C.). This study has been funded by Instituto de Salud Carlos III (ISCIII) through the project “CP23/00080” and co-funded by the European Union (A.G-.C.). L.J.S. was supported by the NOMIS foundation. This work was supported by the European Union’s Horizon 2020 research and innovation programme under Grant Agreement Nos. 668303 (PERFORM) and 848196 (DIAMONDS). The funders were not involved in the study design, collection, analysis, interpretation of data, the writing of this article or the decision to submit it for publication.
Author information
Authors and Affiliations
Consortia
Contributions
A.S., F.M-.T., A.G-.C. and S.V-.L. conceptualized and designed the study. A.S., F.M-.T., M.L., M.vF., C.F., M.E., W.Z., A.J.P., T.T.K., S.A., M.P., H.A.M. and R.dG. provided key resources. I.R-.C., A.I.D., J.A.H., A.J.C., E.D.C., were responsible for clinical classification of the patients and collection of clinical metadata. M.T., D.Z., C.V., S.C.P., L.J.S., D.S.K., U.V.B. and P.A. contributed to patients recruitment. VJW and NS coordinated sample processing and biobanking. S.V-.L., X.B., A.G-.C., M.K. and D.H-.C. conducted the bioinformatics and computational biology analysis. Computational models and machine learning approaches used to analyze the data were developed by A.S., S.V-.L. and A.G-.C. A.S., S.V-.L. and A.G-.C. wrote the first draft of the manuscript. All authors contributed to discussions regarding the interpretation of results and reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Charles Langelier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Viz-Lasheras, S., Gómez-Carballa, A., Bello, X. et al. A diagnostic host-specific transcriptome response for Mycoplasma pneumoniae pneumonia to guide pediatric patient treatment. Nat Commun 16, 673 (2025). https://doi.org/10.1038/s41467-025-55932-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-55932-9
This article is cited by
-
Predicting the protein interaction landscape of a free-living bacterium with pooled-AlphaFold3
Molecular Systems Biology (2026)