Abstract
Mediator25 (MED25) has been ascribed as a signal-processing and -integrating center that controls jasmonate (JA)-induced and MYC2-dependent transcriptional output. A better understanding of the regulation of MED25 stability will undoubtedly advance our knowledge of the precise regulation of JA signaling-related transcriptional output. Here, we report that Arabidopsis MED16 activates JA-responsive gene expression by promoting MED25 stability. Conversely, two homologous E3 ubiquitin ligases, MED25-BINDING RING-H2 PROTEIN1 (MBR1) and MBR2, negatively regulate JA-responsive gene expression by promoting MED25 degradation. MED16 competes with MBR1&2 to bind to the von Willebrand Factor A (vWF-A) domain of MED25, thereby antagonizing the MBR1&2-mediated degradation of MED25 in vivo. In addition, we show that MED16 promotes hormone-induced interactions between MYC2 and MED25, leading to the activation of JA-responsive gene expression. Collectively, our findings reveal a multiprotein regulatory module that robustly and tightly maintains MED25 homeostasis, which determines the strength of the transcriptional output of JA signaling.
Similar content being viewed by others
Introduction
Jasmonate (JA) is a lipid-derived plant hormone that plays a pivotal role in the regulation of plant defense responses, development, and environmental acclimation1,2,3,4. To mediate the plant response to external stress or endogenous stimuli, JA triggers a genome-wide transcriptional reprogramming primarily governed by MYC2, a basic helix-loop-helix (bHLH) transcription factor (TF) that serves as the master regulator of JA signaling5,6,7,8,9,10. Recent elucidation of the core JA signaling pathway has significantly advanced our understanding of the molecular mechanisms underlying MYC2-regulated gene transcription. The core JA signaling pathway comprises interconnected regulatory modules that govern the transcriptional state of JA-responsive genes. Briefly, at the resting (i.e., repressed) stage of JA signaling, a group of plant-specific JASMONATE-ZIM DOMAIN (JAZ) proteins physically bind to and prevent MYC2 from activating the expression of JA-responsive genes11,12,13,14,15. In response to signals that trigger the accumulation of the bioactive JA ligand JA ( + )-7-iso-JA-Ile (JA-Ile), JAZ proteins act as co-receptors by forming a JA-Ile-dependent complex with the F-box protein CORONATINE-INSENSITIVE1 (COI1), a component of the SKP1–CUL1–F-box protein (SCF) E3 ubiquitin ligase complex (SCFCOI1)12,13,16,17, which leads to proteasome-dependent ubiquitination and degradation of JAZ repressors, thereby releasing MYC proteins from transcriptional repression11,13,14,15,16. Biochemical and structural analyzes indicate that JAZ degradation unmasks the JAZ-interaction domain (JID) and transcriptional activation domain (TAD) of MYC proteins to facilitate their binding to MED2518, a subunit of the Mediator transcriptional coactivator complex, and the recruitment of RNA polymerase II (Pol II) to the promoter of MYC2 target genes, thereby establishing an activated transcriptional state.
Our previous studies demonstrated that, at the activation stage of JA signaling, MED25 co-occupies a significant proportion (~25%) of MYC2 target genes together with MYC2 through occupancy pattern profiling19, suggesting that the function of MYC2 in activating JA-responsive gene expression is largely dependent on its interaction with MED25. MED25 physically and functionally interacts with MYC2 to form the MYC2–MED25 functional transcription complex (MMC), which in turn activates the transcription of JA-responsive genes20,21,22,23. Increasing evidence indicates that MED25 acts as a signal-processing and -integrating center during JA-induced transcriptional reprogramming19,20,21,22,23,24,25. In recent years, our understanding of JA signaling has greatly expanded owing to the elucidation of multiple regulators that interact with MED25 and thus coordinate their actions during the hormone-induced activation of MYC2. First, MED25 recruits the Pol II transcriptional machinery to MYC2 target gene promoters, thereby facilitating the assembly of the pre-initiation complex (PIC) for transcriptional initiation by bridging MYC2 and Pol II21,22. Second, MED25 recruits the hormone receptor COI1 to MYC2 target promoters at the resting stage and facilitates COI1-dependent degradation of JAZ repressors in the presence of JA-Ile20. Third, MED25 interacts with the epigenetic regulator HISTONE ACETYLTRANSFERASE OF THE CBP FAMILY1 (HAC1) to selectively regulate histone H3K9 acetylation of MYC2 target promoters20. Fourth, MED25 recruits the coactivator LEUNIG_ HOMOLOG (LUH), which in turn acts as a scaffold to enhance MMC assembly in a hormone-dependent manner23. Fifth, MED25 bridges the dynamic communication between remote JA ENHANCERS (JAEs) and MYC2 target promoters through chromatin looping19. Sixth, MED25 recruits the splicing factors PRE-mRNA-PROCESSING FACTOR 39a (PRP39a) and PRP40a to promote the complete splicing of JAZ genes, which plays a critical role in preventing excessive JAZ splice variant-mediated desensitization of JA signaling25. Collectively, these recent findings systematically establish MED25 as an integrative hub for the transcriptional activation of the JA signaling pathway.
Although these studies have enhanced our comprehension of MED25 activation during hormone perception and signaling19,20,21,22,23,24,25, the regulation of MED25 stability remains less well understood. MED25 was originally identified as a regulator of flowering time in response to changes in light quality and was named PHYTOCHROME AND FLOWERING TIME1 (PFT1)26. Since then, MED25 has become one of the most extensively studied plant Mediator subunits and is currently regarded as a crucial convergence point for various pathways encompassing hormone signaling, biotic and abiotic stress responses, and plant development4,27. Given that MED25 acts as a convergence node for multiple signaling networks, its stability is presumed to be under robust and tight regulation to enable the plant to respond rapidly to the ever-changing environment. Advancing our understanding of the regulation of MED25 stability would undoubtedly contribute to our knowledge of the regulation of Mediator-mediated transcriptional output in plants. A previous report found that two homologous E3 ubiquitin ligases, MBR1 and MBR2, target MED25 for proteasomal degradation, and that MED25 instability is required for the activation of its target gene, FLOWERING LOCUS T (FT)28. However, regulators capable of stabilizing MED25 are yet to be discovered. Considering the prominent role of MMC in JA signaling during hormone-induced activation of MYC2, it is reasonable to speculate that MED25 stability must be regulated by the components of MMC.
Using an unbiased proteomic analysis of MED25- and MYC2-associated proteins in Arabidopsis, we identified MED16 (also known as SENSITIVE TO FREEZING6 [SFR6])29,30,31,32,33,34,35,36,37,38 as an additional component of the MMC. In contrast to the classical transcriptional coactivator function of the Mediator subunit, MED16 plays a crucial role in stabilizing MED25 upon hormone elicitation.
Here, we report the mechanistic function of MED16 in the regulation of JA signaling. We demonstrate that MED16 regulates multiple JA responses by interacting with and stabilizing MED25. In addition, our findings indicate that MBR1 and MBR2 are homologous E3 ligases that target MED25 for degradation, while MED16 competes with MBR1 and MBR2 to bind to the vWF-A domain of MED25, thereby impeding MBR1&2-mediated degradation. Together, we demonstrate that MED16-MBR1&2 acts as a multiprotein regulatory module that tightly and robustly regulates MED25 homeostasis during JA signaling.
Results
MED16 interacts with and stabilizes MED25
MMC acts as an integrative hub that coordinates the activities of multiple regulators during JA-triggered activation of MYC220,23. To identify novel components of the MMC, we previously extracted total proteins from MED25-myc22 and MYC2-myc39 transgenic plants, immunoprecipitated MED25-myc and MYC2-myc using anti-myc agarose beads, and analyzed the composition of immunoprecipitated products by liquid chromatography-tandem mass spectrometry (LC-MS/MS). Previously, we identified SEU, SLK1, SLK2, LUG, and LUH as MMC components in the immunoprecipitated products23. Further analysis revealed that Mediator subunit MED16 was identified in both MED25-myc and MYC2-myc immunoprecipitated products23 (Supplementary Data 1), indicating that MED16 is a candidate component of MMC. We therefore focused on MED16 in subsequent studies.
To confirm the interaction of MED16 with MED25 and MYC2, we performed yeast two-hybrid (Y2H) assays using the fusions of full-length MED16 and MYC2 with the GAL4 DNA activation domain (AD) and those of full-length MED16 and MED25 with the GAL4 DNA binding domain (BD). The results showed that MED25 interacts with MYC2 and MED16 in yeast (Saccharomyces cerevisiae) (Fig. 1a); however, our Y2H results showed that MED16 does not interact with MYC2 (Fig. 1a). To confirm this observation, in vitro pull-down assays were performed using purified 6×His-tagged full-length MED16 (His-MED16) (Supplementary Fig. 1) and Flag epitope-tagged MED25 (MED25-Flag) or MYC2 (MYC2-Flag). The recombinant fusion protein His-MED16 was able to pull down MED25-Flag but not MYC2-Flag (Fig. 1b), indicating that MED16 interacts directly with MED25 but not with MYC2. The Y2H-based domain mapping assays showed that the MED25 vWF-A (MED251–242) domain26, but not the previously reported MED251–226 polypeptide22,36, was sufficient for the interaction of MED25 with MED16 (Fig. 1c, d), indicating that amino acid residues 227–242 of MED25 (Supplementary Fig. 2) are required for its interaction with MED16. Therefore, we hereafter refer to the MED251–242 polypeptide as the vWF-A domain of MED25 (Fig. 1c).
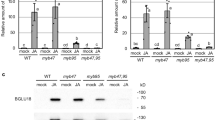
a Y2H assays examining interactions between MED16, MED25 and MYC2. b In vitro pull-down assays to verify interactions between MED16 and MED25, MYC2. Purified His-MED16 protein was incubated with MED25-Flag or MYC2-Flag for the His pull-down assays. MED25-Flag and MYC2-Flag were detected by immunoblotting using an anti-Flag antibody. The purified His-MED16 proteins were detected by immunoblotting using an anti-His antibody. c Schematic domain architecture of MED25. vWF-A, von Willebrand Factor A domain; MD, middle domain; ACID, conserved activator-interacting domain; GD, Gln-rich domain. d Y2H assays examining the interactions of MED25-BD and MED25 derivatives-BD with MED16-AD. e Co-IP assay to verify in vivo interactions between MED16 and MED25 by using MED16-GFP seedlings. Protein extracts from 10-d-old WT and MED16-GFP seedlings were immunoprecipitated with GFP antibody-bound agarose beads. f Co-IP assay to detect the association between MED16 and MYC2 in planta. Protein extracts from 10-d-old WT and MYC2-myc seedlings were immunoprecipitated with myc antibody-bound agarose beads. The white asterisk indicates the position of MED16. g Immunoblotting analyzes showing the gel-filtration patterns of MED25 protein in WT and med16-3 plants, and MED16 protein in WT plants. The red arrowheads indicate the most enriched fractions of MED25 and MED16. Molecular masses are indicated below the blot. Total: total unfractionated extracts. h Immunoblot analyzes of MED25 protein levels in the indicated genotypes. i Quantitative analyzes of the band intensity in (h). Statistical analysis was performed via ANOVA; bars with different letters are significantly different from each other (P < 0.05). j Immunoblot analyzes of MED25 protein levels in WT and med16-3 in response to CHX in the absence or presence of MeJA. To check MED25 protein level, seedlings were treated with CHX with or without 100 μM MeJA and protein levels were analyzed at indicated times. k Quantitative analyzes of the band intensity in (j). In (i) and (k), data shown are mean values of three biological repeats with standard deviations (SD). In (h) and (j), bands were quantified using Image J. In (a, b, d–h, j), n = 3 independent experiments. Source data are provided as a Source Data file.
To further determine whether MED16 interacts with MED25 in planta, we performed coimmunoprecipitation (co-IP) assays using the previously described MED16-GFP transgenic plants40 (overexpressing the GFP-fused MED16 coding sequence) and anti-MED25 antibody22. Results showed that MED16-GFP could coprecipitate endogenous MED25 (Fig. 1e), confirming the interaction between MED16 and MED25 in Arabidopsis. Since MED25 and MYC2 form a transcriptional complex through direct physical interaction20,22, we further investigated whether MED16 associates with MYC2 by performing co-IP assays using MYC2-myc transgenic plants39 and anti-MED16 antibody (Supplementary Fig. 3a). Co-IP results showed that the MYC2-myc protein was able to coprecipitate endogenous MED16 (Fig. 1f), indicating that MED16 associates with MYC2 in plants. Taken together, these results indicate that MED16 physically interacts with MED25 and associates with MYC2, confirming the reliability of our LC-MS/MS results.
Our results indicated that the vWF-A domain is required for the MED16–MED25 interaction. Interestingly, we found that vWF-A is the most conserved domain of MED25, which is evolutionarily conserved not only in plants but also in mammals (Supplementary Fig. 2). Notably, the vWF-A domain of human MED25 (hMED25) is responsible for its recruitment into the Mediator complex41. We therefore hypothesized that MED16 may be responsible for recruiting MED25 into the Mediator complex. To test this possibility, we performed gel-filtration analyzes using protein extracts prepared from wild-type (WT) and med16-3 (also known as sfr6-3) mutant plants. As expected, the majority of MED25 and MED16 proteins were present in similar fractions, with predicted sizes greater than 1 MDa (Fig. 1g, top and middle panels), which is consistent with the predicted size of the Mediator complex42 and indicates the integrity of the Mediator complex in WT plants. By contrast, in the absence of MED16, the majority of the MED25 complexes were shifted to smaller fractions (<500 kDa) (Fig. 1g, bottom panel), a significant deviation from the results obtained in WT plants, confirming that MED16 is required for the recruitment of MED25 into the Mediator complex.
The fact that MED16 physically interacts with MED25 led us to investigate the significance of the MED16–MED25 interaction. Interestingly, our gel-filtration assays consistently revealed a significant reduction in the endogenous MED25 protein level in med16-3 compared with WT plants (Fig. 1g). This result prompted us to investigate the role of MED16 in MED25 accumulation. We first examined MED25 gene expression in WT and med16-3 plants. The expression of MED25 was essentially comparable between WT and med16-3 plants (Supplementary Fig. 3b), suggesting that MED16 has little, if any, effect on the transcriptional regulation of MED25. Next, we compared MED25 protein accumulation between the WT, multiple med16 mutant and med25-4 mutant plants using an endogenous MED25 antibody22. We found a specific band at a molecular weight of approximately 100 kDa, which is consistent with predicted size of MED25, present in WT but absent in med25-4, thus validating the specificity of the MED25 antibody (Fig. 1h and Supplementary Fig. 3c). Furthermore, the level of MED25 protein was significantly reduced in multiple med16 mutant plants (Figs. 1h, i and Supplementary Fig. 3c), suggesting that MED16 is required for the protein accumulation of MED25.
To examine the functional specificity of MED16 on the accumulation of MED25, we examined the endogenous levels of several Mediator subunits including MED8, MED18, MED31, and MED35 in WT and med16-3 mutant plants using endogenous antibodies. Our protein gel blot assays showed that the levels of these Mediator subunit proteins were essentially comparable between WT and med16-3 plants, suggesting that MED16 has a negligible effect on the accumulation of Mediator subunits examined in this study (Supplementary Fig. 3d). Furthermore, considering that MED16 associates with MYC2, we also examined the accumulation of MYC2 in WT and med16-3 using endogenous MYC2-specific antibody20. The results showed that the protein level of MYC2 was similar in WT and med16-3 plants (Supplementary Fig. 3e), indicating that MED16 has little effect on MYC2 accumulation.
To investigate whether the regulation of MED25 stability by MED16 is a part of JA signaling, we treated WT and med16-3 plants with cycloheximide (CHX), an inhibitor of de novo protein synthesis, in the absence or presence of methyl jasmonate (MeJA). In WT plants, the MED25 protein level began to decrease gradually at 6 h (h) after the CHX treatment (Fig. 1j, k and Supplementary Fig. 3f) and was significantly reduced at 12 h (Fig. 1j, k and Supplementary Fig. 3f), indicating that MED25 is unstable after prolonged exposure to CHX. Interestingly, co-treatment with MeJA largely blocked the effect of CHX in WT plants (Fig. 1j, k and Supplementary Fig. 3f), suggesting that MeJA treatment promotes MED25 stability. In parallel experiments, MED25 protein levels were lower in the med16-3 mutant, compared with the WT, in the absence of CHX (Fig. 1j, k and Supplementary Fig. 3f) and were further reduced as early as 3 h after the CHX treatment, suggesting that MED25 is more unstable in med16-3 (Fig. 1j, k and Supplementary Fig. 3f). Importantly, in contrast to WT, co-treatment with MeJA no longer blocked the effect of CHX in med16-3 plants (Fig. 1j, k and Supplementary Fig. 3f), indicating that MED16 plays a critical role in stabilizing MED25 during JA signaling.
MED16 positively regulates MYC2/MED25-dependent JA signaling
To elucidate the biological significance of the MED16–MED25 interaction, we investigated whether MED16 is involved in JA-induced root growth inhibition and defense-related gene expression. We first compared the root length of WT (Col-0) and med16 mutant plants grown on half-strength Murashige and Skoog (1/2 MS) medium supplemented with or without JA. Consistent with previous observations39, exogenous JA application inhibited root growth in both WT and med16 seedlings; however, this inhibition was significantly less in med16 seedlings (Fig. 2a), suggesting that MED16 is a positive regulator of JA-induced root growth inhibition.
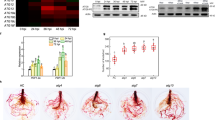
a Root growth inhibition assay of 10-d-old WT and med16 seedlings. Plants were grown on ½MS medium without or with 20 μM JA, and seedling root length was measured at 10 d after germination. Results shown are the mean ± SD, (n = 25 seedlings). b–e RT-qPCR showing the MeJA-induced expression of VSP1 (6 h treatment; [b]), VSP2 (6 h treatment; [c]), PDF1.2 (48 h treatment; [d]), and Thi2.1 (48 h treatment; [e]) in WT and med16 mutant plants. Plants were treated without or with 100 μM MeJA for the indicated times before RNA extraction. Data are presented as mean ± SD. f Root growth inhibition assay of Ten-d-old plants in the indicated genotypes. Plants were grown on ½MS medium without or with 20 μM of JA, and seedling root length was measured at 10 d after germination. Results shown are the mean ± SD, (n = 21 seedlings). g–j RT-qPCR showing the MeJA-induced expression of VSP1 (6 h treatment; [g]), VSP2 (6 h treatment; [h]), PDF1.2 (48 h treatment; [i]), and Thi2.1 (48 h treatment; [j]) in the indicated genotypes. Ten-day-old seedlings were treated without or with 100 μM MeJA for the indicated times before RNA extraction. Data in (b–e) and (g–j) are the mean values of three biological repeats with SD. Statistical analysis was performed via ANOVA; bars with different letters are significantly different from each other (P < 0.05). In (a) and (f), the central line of the box plot is the median. The box’s edges are the lower (25th percentile) and upper quartiles (75th percentile). Whiskers extend to data points within 1.5 times the interquartile range (IQR). Source data are provided as a Source Data file.
Next, we examined whether JA-induced expression of wound- and pathogen-responsive genes was affected in med16 plants. In Arabidopsis, JA-responsive genes form two distinct branches that were differentially regulate by MYC2. VEGETATIVE STORAGE PROTEIN1 (VSP1) and VSP2, which are widely used as marker genes for JA-regulated wound responses, were positively regulated by MYC2, whereas PLANT DEFENSIN1.2 (PDF1.2) and THIONIN2.1 (Thi2.1), which are widely used as marker genes for JA-regulated pathogen responses43,44, were negatively regulated by MYC2. Our RT-qPCR results showed that the MeJA-induced expression of VSP1 was significantly reduced in med16-2 and med16-3 than in the WT (Fig. 2b). Similarly, the MeJA-induced expression of JA-responsive genes VSP2, PDF1.2, and Thi2.1 was significantly reduced in med16-2 and med16-3 than in the WT (Fig. 2c–e), suggesting that MED16 is required for the JA-induced expression of these genes.
Since MED16 interacts with and stabilizes MED25, we proceeded to investigate whether the involvement of MED16 in JA signaling is dependent on MMC. Towards this goal, we generated med16-3 med25-4, med25-4 myc2-2, and med16-3 myc2-2 double mutant lines by crossing the respective single mutants, and compared JA-induced root growth inhibition, wound-responsive gene expression and pathogen-responsive gene expression between the double mutants and the corresponding parental lines. Consistent with previous observations, our results showed that JA-induced root growth inhibition and wound-responsive gene expression was significantly reduced in med16-3, med25-4, and myc2-2 plants compared with the WT (Fig. 2f–h). Importantly, JA-induced root growth inhibition and wound-responsive gene expression were essentially comparable between the med16-3 med25-4 double mutant and its parental lines (Fig. 2f–h), suggesting that there is no additive effect between MED16 and MED25 in regulating these JA responses. This was consistent with the proposed role of MED16 in promoting MED25 stability during JA signaling. Additionally, regarding the JA-induced root growth inhibition assays, the phenotypes of the med16-3 myc2-2 and med25-4 myc2-2 double mutants were comparable to that of myc2-2 (Fig. 2f), suggesting that myc2-2 is epistatic to both med16-3 and med25-4 in the inhibition of JA-induced root growth. In terms of wound-responsive gene expression, med16-3 myc2-2 and med25-4 myc2-2 double mutants did not exhibit either significant additive effects or clear epistatic effects (Fig. 2g, h). These observations suggest that MED16 and MED25 act in the same pathway as MYC2 and positively regulate the expression of JA-induced wound-responsive genes.
Next, we compared the JA-induced expression levels of PDF1.2 and Thi2.1 between the double mutants and their parental lines. Consistent with previous observations, PDF1.2 and Thi2.1 expression levels were significantly decreased in med16-3 and med25-4 mutants and significantly increased in myc2-2 compared with the WT (Fig. 2i, j). Importantly, JA-induced expression levels of PDF1.2 and Thi2.1 were essentially comparable between the med16-3 med25-4 double mutant and its parental lines (Fig. 2i, j), suggesting that there is no additive effect between MED16 and MED25 in regulating pathogen-responsive gene expression. Interestingly, consistent with previous observations22, in the med25-4 myc2-2 double mutant, JA-induced expression levels of PDF1.2 and Thi2.1 were essentially comparable to those in med25-4, suggesting that med25-4 is epistatic to myc2-2 in terms of JA-induced PDF1.2 and Thi.2.1 expression (Fig. 2i, j). Similarly, in the med16-3 myc2-2 double mutant, JA-induced expression levels of PDF1.2 and Thi2.1 were essentially comparable to those in med16-3, indicating that med16-3 is also epistatic to myc2-2 in terms of JA-induced PDF1.2 and Thi2.1 expression (Fig. 2i, j). In sum, these results suggest that MED16 and MED25 are required for the inhibitory function of MYC2 in the regulation of pathogen-responsive gene expression, and that MED16 and MED25 acts genetically downstream of MYC2 to regulate pathogen-responsive gene expression.
Collectively, our results support the conclusion that MED16 positively regulates MYC2/MED25-dependent JA signaling via promoting MED25 stability.
MBR1&2 facilitate MED25 poly-ubiquitination
The key role of MED16 in promoting MED25 stability led us to focus on the factors that regulate MED25 stability. A previous study showed that two putative Ring-type E3 ligases, MBR1 and MBR2, interact with MED25 and promote its degradation via the ubiquitin-26S proteasome pathway28; however, whether MBR1 and MBR2 exhibit true E3 ubiquitin ligase activities in vitro remained unclear. To address this query, we conducted in vitro ubiquitination assays to examine if MBR1 and MBR2 are functional E3 ubiquitin ligases mediating poly-ubiquitination of MED25 in vitro. First, Y2H, co-IP, and in vitro pull-down assays were performed to determine the physical interaction between MED25 and MBR2. In agreement with previous studies28, Y2H assays confirmed the interaction of MED25 with both MBR1 and MBR2 (Fig. 3a). In in vitro pull-down assays, full-length MBR2 expressed in Escherichia coli remained insoluble despite various efforts. To circumvent this problem, we expressed a truncated MBR2 (MBR2C, MBR2316–666) containing the intact Ring-H2 domain (Fig. 3b) and successfully obtained soluble glutathione S-transferase (GST)-tagged MBR2C. Y2H results showed interaction between MBR2C and MED25 in yeast cells (Supplementary Fig. 4a). Subsequently, we performed pull-down assays using purified GST-MBR2C and in vitro translated MED25-Flag, and found that GST-MBR2C, but not GST, was able to pull down MED25 (Fig. 3c), indicating that MED25 physically interacts with MBR2. Consequently, we investigated whether MBR2 is a functional E3 ubiquitin ligase using the MBR2C protein in subsequent experiments. Furthermore, in co-IP assays performed using 35Spro:MBR2-GFP (MBR2-GFP) transgenic plants (Supplementary Fig. 4b) and anti-MED25 antibody, MBR2 could co-immunoprecipitate endogenous MED25, suggesting that MBR2 interacts with MED25 in plants (Fig. 3d). Taken together, these results indicate that MBR2 physically interacts with MED25 both in vitro and in vivo.
a Y2H assays examining the interactions of MED25-BD and MED25 derivatives-BD with MBR1-AD or MBR2-AD. b Schematic domain architecture of MBR2 and MBR2C. Ring, Ring finger domain. c In vitro pull-down assays to verify interactions between MBR2C and MED25. MED25-Flag and pulled down GST-MBR2C protein were detected by immunoblotting using anti-Flag and anti-GST antibodies, respectively. CBB, Coomassie Brilliant Blue staining. The white asterisk indicates the position of GST-MBR2C. d Co-IP assay to verify in vivo interactions between MBR2 and MED25 by using MBR2-GFP seedlings. Ten-d-old WT and MBR2-GFP seedlings were treated with 50 μM MG132, and then the protein extracts from WT and MBR2-GFP plants were immunoprecipitated with GFP antibody-bound agarose beads. e In vitro E3 ubiquitin ligase activity of MBR2. GST-MBR2C fusion protein was assayed for E3 ubiquitin ligase activity in the presence of E1 (from wheat), E2 (from human), and 6xHis-tagged ubiquitin (His-Ub). GST was used as a negative control. The anti-ubiquitin antibody was used to detect the poly-ubiquitination of His-Ub by GST-MBR2C. f MBR2 ubiquitinates MED25. In vitro ubiquitination assays were carried out with the indicated recombinant proteins, and poly-ubiquitination of MED25-Flag was detected by immunoblot using anti-Flag antibody. In (e) and (f), the white asterisk indicates the position of GST-MBR2C; and the presence of E1 (His-UBA1) and E2 (UBCh5b) were detected by using anti-His and anti-UBCh5b antibodies, respectively. g MED25 is poly-ubiquitinated in plants. Ten-day-old seedlings were pretreated with 50 mM MG132 for 6 h before immunoprecipitation. MED25-myc proteins were enriched from myc-trap beads. Ubiquitinated proteins and input were detected using anti-ubiquitin and myc antibodies, respectively. h MED25 is accumulated in the mbr mutant plants. Bands were quantified using Image J. i Quantitative analyzes of the band intensity in (h). Data shown are mean values of four biological repeats with SD. Statistical analysis was performed via ANOVA; bars with different letters are significantly different from each other (P < 0.05). In (a, c–g), all experiments were repeated three times, and similar results were obtained. Source data are provided as a Source Data file.
Next, to test whether MBR2 possesses E3 ubiquitin ligase activity, we performed in vitro ubiquitination assays45. In the presence of wheat (Triticum aestivum) His-E1 (ubiquitin-activating enzyme), human E2 (UBCh5b, ubiquitin-conjugating enzyme), and His-ubiquitin, ubiquitination activity was observed only upon the addition of purified GST-MBR2C (Fig. 3e), indicating that MBR2 is a functional E3 ubiquitin ligase. We further used this assay to investigate whether MED25 serves as the substrate of MBR2. Incubation of MED25-Flag, GST-MBR2C, ubiquitin, E1, and E2 resulted in the formation of ladder-like bands (Fig. 3f), indicating that MBR2 promotes the poly-ubiquitination of MED25-Flag in vitro. Furthermore, to evaluate whether MED25 can be ubiquitinated in plant cells, we performed standard in vivo ubiquitination experiments10,46 using MED25-myc transgenic plants. We enriched the MED25-myc proteins from transgenic seedlings using myc-trap beads, with wild-type plants serving as a negative control. The bound proteins were analyzed by immunoblotting with an anti-ubiquitin antibody. As shown in Fig. 3g, a ladder-like protein pattern was detected in the enriched proteins from MED25-myc transgenic plants, but not in proteins from wild-type plants (Fig. 3g), indicating that MED25 can be ubiquitinated in plant cells. Taken together, these results indicate that MBR2 is a functional E3 ubiquitin ligase that promotes the poly-ubiquitination of MED25.
To determine the role of MBR1 and MBR2 in the regulation of MED25 proteolysis in plants, we created mbr1 and mbr2 single mutants using the CRISPR/Cas9 gene editing system47 and subsequently crossed these single mutants to generate the mbr1-1 mbr2-1 double mutant. Sequence analysis of the MBR1 open reading frame (ORF) revealed the insertion of an adenine at nucleotide 24 in mbr1-1 and that of a thymine at nucleotide 66 in mbr2-1, both of which resulted in a frame shift and the generation of a premature stop codon (TGA, mbr1-1; TAG, mbr2-1; Supplementary Fig. 4c). We then examined the MED25 protein level in WT, mbr1-1, mbr2-1, and mbr1-1 mbr2-1 plants. Compared with the WT, the protein level of MED25 was similar in mbr1-1, slightly higher in mbr2-1, and significantly higher in mbr1-1 mbr2-1 (Fig. 3h, i and Supplementary Fig. 5), indicating that MBR1&2 act redundantly to promote the proteolysis of MED25.
Negative regulation of JA responses by MBRs depends on MED25
The subsequent investigation aimed to determine the involvement of MBR1&2 in JA-induced root growth inhibition and defense-related gene expression. We first examined the root length of WT and mbr1-1 mbr2-1 plants grown on 1/2 MS medium supplemented with or without JA. Consistent with previous observations, exogenous JA application led to root growth inhibition in both genotypes, and this inhibition was significantly increased in mbr1-1 mbr2-1 than in the WT (Fig. 4a), suggesting that MBR1&2 negatively regulate JA-induced root growth inhibition. Consistently, MeJA-induced expression of VSP1, VSP2, PDF1.2 and Thi2.1 was significantly higher in mbr1-1 mbr2-1 than in the WT (Fig. 4b–e). These results indicate that MBR1&2 negatively regulate multiple JA responses.
a Root growth inhibition assay of Ten-d-old WT, med25-4, mbr1-1 mbr2-1 and med25-4 mbr1-1 mbr2-1 plants. Plants were grown on ½MS medium without or with 20 μM of JA, and seedling root length was measured at 10 d after germination. (n = 25) seedlings. The central line of the box plot is the median. The box’s edges are the lower (25th percentile) and upper quartiles (75th percentile). Whiskers extend to data points within 1.5 times the IQR. b–e RT-qPCR showing the MeJA-induced expression of VSP1 (6 h treatment; [b]), VSP2 (6 h treatment; [c]), PDF1.2 (48 h treatment; [d]), and Thi2.1 (48 h treatment; [e]) in WT, med25-4, mbr1-1 mbr2-1 and med25-4 mbr1-1 mbr2-1 plants. Plants were treated without or with 100 μM MeJA for the indicated times before RNA extraction. Data are presented as mean ± SD. Data in (b–e) are the mean values of three biological repeats with SD. Statistical analysis was performed via ANOVA; bars with different letters are significantly different from each other (P < 0.05). In (a–e), all experiments were repeated three times, and similar results were obtained. Source data are provided as a Source Data file.
We then tested whether MBR1&2 are involved in regulating diverse aspects of the JA response by promoting MED25 degradation. We first generated the mbr1-1 mbr2-1 med25-4 triple mutant by crossing mbr1-1 mbr2-1 with med25-4, and then compared JA-induced root growth inhibition and JA-responsive gene expression between the triple mutant and their parents. The mbr1-1 mbr2-1 double mutant was highly sensitive to JA-induced root growth inhibition, and this phenotype was suppressed by the MED25 mutation (Fig. 4a), suggesting that med25-4 is epistatic to mbr1-1 mbr2-1 in regulating JA-induced root growth inhibition. Similarly, RT-qPCR results showed that the expression levels of pathogen-responsive genes VSP1, VSP2, PDF1.2 and Thi2.1 were significantly increased in mbr1-1 mbr2-1 mutant plants, and this increase was largely suppressed in the med25-4 background (Fig. 4b–e), suggesting that med25-4 is epistatic to mbr1-1 mbr2-1 in regulating the expression of VSP1, VSP2, PDF1.2 and Thi2.1. Taken together, these results led to the conclusion that MBR1&2 negatively regulate multiple JA responses by promoting the degradation of MED25.
MED16 competes with MBR2 to bind to MED25
MED16 plays a key role in promoting MED25 stability, exhibiting an unconventional function that distinguishes it from the conventional transcriptional coactivator activity of Mediator subunits. We aimed to determine how MED16 regulates MED25 stability. Since MED16 and MBRs play opposite roles in regulating MED25 stability, we hypothesized that MED16 either directly inhibits the E3 ubiquitin ligase activity of MBRs or competes with MBRs to bind to MED25. The results of protein interaction analysis showed no interaction between MED16 and MBR1&2 (Supplementary Fig. 6), thus ruling out the possibility that MED16 directly inhibits the E3 ubiquitin ligase activity of MBRs through physical interaction. To test whether MED16 competes with MBRs to bind to MED25, we performed Y2H-based domain mapping assays. The results showed that the vWF-A domain of MED25 is responsible for the MED25–MBRs interaction (Fig. 3a). Considering this result together with the requirement of the vWF-A domain for the MED16–MED25 interaction (Fig. 1d), we concluded that MED16 and MBRs bind to the same domain of MED25.
Next, to determine whether MED16 competes with MBR2 to bind to MED25, we performed yeast three-hybrid (Y3H) assays. MED16–MED25 interaction was detected on the synthetic defined (SD) medium SD/-Ade/-His/-Trp/-Leu (SD/-4) but was abolished on the SD/-Ade/-His/-Trp/-Leu/-Met (SD/-5) selection medium upon the induction of MBR1 or MBR2 expression (Fig. 5a), suggesting that MBR1 and MBR2 competitively inhibit the MED16–MED25 interaction in yeast cells. In parallel experiments, MED25 interacted with MBR1 and MBR2 on SD/-4; however, the strength of MBRs–MED25 interaction decreased on SD/-5 upon the induction of MED16 expression (Fig. 5a), suggesting that MED16 also interferes with the MBRs–MED25 interaction in yeast cells. Taken together, these results suggest that MED16 and MBRs compete with each other to bind to MED25 in yeast cells.
a Yeast three-hybrid (Y3H) assays showing that addition of MBR1, MBR2 dramatically reduced the MED16–MED25 interaction (Top three panels) and addition of MED16 reduced the MED25–MBR1/2 interactions (Bottom four panels). (Top three panels) Yeast cells cotransformed with pGADT7-MED16 and pBridge-MED25-MBR1,2 were dropped onto SD/-Trp/-Leu (SD/-2) and SD/-Trp/-Leu/-Ade/-His (SD/-4) media to assess the MED25–MED16 interaction. The cotransformed yeast cells were dropped onto SD/-Trp/-Leu/-Ade/-His/-Met (SD/-5) medium to induce expression of MBR1 or MBR2. (Bottom four panels) Yeast cells cotransformed with pGADT7-MBR1, 2 and pBridge-MED25-MED16 were dropped onto SD/-2 and SD/-4 media to assess the MED25/MBR1,2 interactions. The cotransformed yeast cells were dropped onto SD/-5 medium to induce expression of MED16. AD, activation domain fusion; BD, binding domain fusion. b In vitro quantitative pull-down assays showing that MED16 competes with MBR2 for binding to MED25. For each sample, the amounts of MED25-Flag and GST-MBR2C were equal, and His-MED16 was added according to the indicated gradient. GST-MBR2C was pulled down by MED25-Flag immobilized on an anti-Flag resin. Proteins were eluted and analyzed using an anti-Flag antibody. The white asterisk indicates the position of His-MED16 (upper) and His-TF (lower). c MED16 prevents MED25 from degradation in planta. MED25-myc was infiltrated with MBR2-Flag and/or MED16-GFP co-expression. d Co-IP assays of MED25 and MYC2 in the MYC2-myc/WT and MYC2-myc/med16-3 plants. Ten-day-old MYC2-myc/WT and MYC2-myc/med16-3 seedlings were treated with 100 mM MeJA for the indicated times. Proteins extracted from MYC2-myc/WT and MYC2-myc/med16-3 plants were immunoprecipitated using an anti-myc antibody and immunoblotted using an anti-MED25. In (a–d), all experiments were repeated three times, and similar results were obtained. e Proposed working model for the mechanistic roles of MED16 coordinates with MED25 in regulating JA-induced activation of MYC2. MED16 competes with MBRs for binding to vWF-A domain of MED25, thereby recruiting MED25 into Mediator complex, which impairs the interaction ability of MBRs with MED25 and stabilizes MED25. In addition, MED16 also promotes hormone-dependent enhancement of protein interactions between MYC2 and MED25, thereby activating JA-responsive gene expression. Source data are provided as a Source Data file.
To further substantiate the above observations, we performed in vitro pull-down assays using a constant protein concentration of GST-MBR2C and an increasing protein concentration of His-MED16. The results showed that MED25-Flag pulled down less GST-MBR2C as the amount of His-MED16 increased (Fig. 5b, lanes 2–5). However, the ability of MED25-Flag to pull down GST-MBR2C was not apparently affected by the increase in the amount of His-Trigger Factor (His-TF; negative control) (Fig. 5b, lanes 2, 6–8). These results indicate that MED16 competes with MBR2 to bind to MED25 in vitro.
Next, to confirm that MED16 and MBR2 play antagonistic roles in MED25 degradation in vivo, we co-expressed MED25-myc and MBR2-Flag, together with or without MED16-GFP, in Nicotiana benthamiana leaves and examined MED25-myc protein levels. Consistent with previous observations, MED25-myc abundance was significantly reduced in leaves co-expressing MED25-myc and MBR2-Flag (Fig. 5c), suggesting that MBR2 promotes MED25 degradation. Notably, the reduction in MED25-myc abundance was significantly inhibited in leaves co-expressing MED25-myc with MBR2-Flag and MED16-GFP (Fig. 5c), suggesting that MED16 prevents MED25 from MBR2-mediated degradation in plants. Taken together, these results suggest that MED16 and MBRs act antagonistically by engaging in competitive interactions with MED25 to regulate MED25 homeostasis.
MED16 facilitates MYC2–MED25 interaction during JA signaling
Previous studies showed that hormone elicitation enhances the interaction between MYC2 and MED2520,23, and the results described above showed that MED16 directly interacts with MED25 and associates with MYC2 (Fig. 1). Therefore, we hypothesized that MED16 is involved in the hormone-induced enhancement of MYC2–MED25 interaction. To test this hypothesis, we performed co-IP experiments and evaluated the ability of MYC2-myc to pull down native MED25 in med16 mutant and WT backgrounds. Consistent with previous observations20,23, our results showed that MeJA treatment significantly increased the ability of MYC2-myc to pull down MED25 in the WT; however, this increase was completely abolished in the med16 mutant (Fig. 5d), suggesting that MED16 plays a critical role in promoting the MYC2–MED25 interaction during the activation of JA signaling.
In summary, we propose a working model elucidating the role of MED16 in establishing MMC-dependent JA-responsive gene expression. We provide evidence showing that MED16 genetically and physically interacts with MED25 and positively regulates JA signaling. Additionally, our study reveals that MED16 competes with MBRs to bind to the vWF-A domain of MED25, leading to the recruitment of MED25 into the Mediator complex. This competition weakens the ability of MBRs to interact with MED25 and ultimately stabilizes MED25 (Fig. 5e). Furthermore, we showed that MED16 plays a crucial role in facilitating the hormone-dependent enhancement of MYC2–MED25 interaction. These findings illustrate that a multiprotein regulatory module regulates JA-induced transcriptional reprogramming by controlling the stability of MED25, and provide mechanistic insights into the interplay between different Mediator subunits.
Discussion
Mediator, an evolutionarily-conserved multisubunit protein complex, is required for gene transcription by RNA Pol II48. Extensive studies conducted to date have established the functional versatility of the Mediator complex, demonstrating its role in sensing, integrating, and processing diverse signaling pathways by physically interacting with a range of TFs48,49,50,51,52 and assembling at promoters as a PIC to control transcription initiation. Although experimentally purified from a number of fungal and metazoan organisms, the Mediator complex was successfully isolated from plants only in 2007, 13 years after its initial discovery53. To date, 29 conserved and 6 plant-specific subunits have been identified in the plant Mediator complex3,53. Similar to metazoans, most research in plants focuses on how Mediator subunits interact with signaling-specific TFs to activate or repress gene expression. For example, in Arabidopsis, MED25 interacts with the master regulator MYC2 and positively regulates multiple JA responses22. MED8 regulates plant defense against Botrytis cinerea infection by interacting with FAMA to activate OCTADECANOID-RESPONSIVE ARABIDOPSIS59 (ORA59) expression54. MED18 interacts with the YIN YANG1 (YY1) TF to bind to the promoter regions of disease-related genes and repress their expression55. CDK8 negatively regulates B. cinerea-induced plant immunity by interacting with WAX INDUCER1 (WIN1), an ETHYLENE RESPONSIVE FACTOR (ERF) family protein involved in cuticular wax biosynthesis56. In addition to interacting with TFs and assembling into the PIC, Mediator subunits also regulate various steps of transcriptional regulation, including transcription elongation and alternative mRNA processing57,58. Recently, novel roles of Mediator in gene expression regulation have been revealed by showing its connection to the nuclear pore and linking Mediator to the regulation of gene positioning in the nuclear space, extending its function from gene expression regulation to gene positioning59. Nonetheless, as a multisubunit complex, our understanding of the function of Mediator subunits in any signaling pathway is still in its infancy.
Arabidopsis med16 mutants were first identified in a screen for mutants that failed to acclimate to freezing temperatures60. Over a decade later, the MED16 subunit of the Mediator complex was identified as the causal factor responsible for the mutant phenotype29. Subsequent studies have shown that MED16 regulates many biological processes, including plant defense and abiotic stress responses, iron/phosphate homeostasis, and endoreduplication30,31,33,34,35,36,37,40,49,61. Previous reports showed that MED16 acts as a transcriptional activator that associates with the WRKY33 TF and is required for defense against S. sclerotiorum infection30. In this study, we extend the function of MED16 and show systemically that MED16 directly interacts with and stabilizes MED25 to regulate multiple JA responses. Specifically within the JA signaling pathway, our findings suggest that, rather than acting as a classical transcriptional coactivator, MED16 acts as a guardian of MED25. The following lines of evidence support our claim: (1) MED16 directly interacted with MED25 but not with the master regulator MYC2 (Fig. 1a–e); (2) MED16 exhibited genetic interaction with MED25 and MYC2 (Fig. 2f–j); (3) MED25 protein levels were significantly and specifically reduced in med16 mutant plants, with negligible impact on the accumulation of other Mediator subunits (Fig. 1h, i and Supplementary Fig. 3c, d); (4) the MeJA-enhanced stability of MED25 was largely blocked in med16 plants (Fig. 1j, k), and (5) MED16 is required for recruiting MED25 into the Mediator complex (Fig. 1g). Taken together, these findings strongly support that MED16 functions either as a typical transcriptional activator or as an atypical guardian in a context-specific manner.
Upon perception by plant cells, JA triggers a genome-wide transcriptional reprogramming, in which a significant portion (~39%) of JA-regulated genes were also regulated by MYC27, suggesting MYC2 acts as a master regulator during JA signaling. Previous study investigated the genome-wide binding profiles of MYC2 and MED25 in response to MeJA and identified 11,446 and 3411 binding peaks for MYC2 and MED25, respectively19. Further comparison of these datasets revealed that MYC2 and MED25 shared 2863 binding peaks, representing approximately 25% of MYC2 binding peaks and 84% of MED25 binding peaks19. These results suggest that the function of MYC2 in regulating JA signaling depends largely on MED25. Considering the fact that MED16 promotes MED25 stability during JA signaling, MED16 and MED25 undoubtedly co-regulate a large number of MYC2 target genes. Consistent with these findings, our studies showed that the phenotypes of med16-3, med25-4 and myc2-2 mutant plants were all exhibited a significantly less sensitive phenotype to JA than that of wild-type plants in terms of JA-induced root growth and wound-responsive gene expression (Fig. 2f–h). In addition, the phenotypes of the med16-3 myc2-2 and med25-4 myc2-2 double mutants were comparable to that of myc2-2 (Fig. 2f–h). These results suggested that MED16 and MED25 act in the same pathway as MYC2 and regulate JA-induced root growth inhibition and wound-responsive gene expression as a unitary module with MYC2. However, for a certain phenotype (i.e., JA-induce growth inhibition), myc2-2 showed a more severe phenotype compared to med25-4 and med16-3 (Fig. 2f), implying that MYC2 interacts with other regulatory elements in addition to the MED25–MED16 module in regulation of JA-induced root growth inhibition.
Notably, albeit MMC components co-regulate numerous JA-responsive gene expression, the phenotypic defects caused by MED16 or MED25 mutation were not always identical to those caused by MYC2 mutation, suggesting a more complex genetic relationship between these genes. For example, in terms of pathogen-responsive gene expression, JA treatment significantly increased the expression of pathogen-responsive genes in Arabidopsis thaliana, but this increase did not appear to be regulated by MYC2, as it exerted a negative regulatory effect on these genes (Fig. 2i, j). Previous studies have shown that MYC2 inhibits pathogen-responsive gene expression by repressing the expression of ORA59 and ERF1. Both transcription factors are essential for activating the expression of pathogen-responsive genes through direct binding to the GCC-box in their promoters10,21. By downregulating ORA59 and ERF1, MYC2 effectively suppresses expression of these genes10,21. By contrast, the levels of pathogen-responsive gene transcripts decreased significantly in both MeJA-treated med16 and med25 mutants (Fig. 2i, j), highlighting the critical role of MED16 and MED25 in JA-induced pathogen defense. Specifically, MED25 interacts not only with MYC2, but also with ORA59 and ERF1 in Arabidopsis21. This network of interactions explains the significant reduction in pathogen-responsive gene expression observed in MeJA-treated med25 mutants. Indeed, since a limited number of Mediator subunits can transcribe all protein-coding genes, it is unsurprising that individual Mediator subunits interact with a variety of transcription factors, some of which may play overlapping roles within a given pathway.
Particularly, as shown in Fig. 2i, the expression of PDF1.2 was extremely low in both MeJA-treated med16 and med25 mutants (Fig. 2i). For MED25, in addition to its interactions with ORA59 and ERF1 mentioned above, this may also be due to its interaction with CDK8, which collectively enhances PDF1.2 expression56. Regarding MED16, in addition to stabilizing MED25, previous studies have shown that MED16 physically associates with WRKY33 to promote transcription of PDF1.2 and ORA5930, this would further explain the extremely low expression of PDF1.2 in MeJA-treated med16 mutant.
Interestingly, in addition to MED16, several Mediator subunits, including MED8, MED18, and CDK8, interact with MED25 and perform functions overlapping those of MED25 to regulate JA-induced gene expression in Arabidopsis54,55,56. Like MED16, these Mediator subunits directly interact with MED25 but not with MYC2. Therefore, it is not surprising that these Mediator subunits function in the JA signaling pathway by interacting and cooperating with MED25. However, because of the limited studies conducted on the architecture of the plant Mediator complex, the molecular details of functional interactions between these Mediator subunits and MED25 remain unclear. A deeper understanding of the nature, in particular the structural and biochemical bases, of these protein–protein interactions may be crucial for the systematic elucidation of the role of MMC in the regulation of JA-mediated transcriptional output.
MED25 has emerged as one of the most extensively studied plant Mediator subunits, and its role in a variety of plant developmental processes and stress responses has been well-studied; however, the regulation of MED25 stability has received relatively little attention. MBR1 and MBR2 were identified through Y2H assays using the vWF-A domain of MED25 as bait. Previous research demonstrated that MBR1 and MBR2 can facilitate RING-H2-dependent degradation of MED25 in vivo28. Interestingly, it was also reported that MED25 degradation is coupled with its role in stimulating FT gene transcription during the induction of flowering. This degradation-coupled function has been termed “activation by destruction”28. It is evident that distinct requirements for MED25 degradation may be necessitated by different signaling events. For example, although the med16 mutant exhibits a constitutive reduction in MED25 accumulation (Fig. 1h, i and Supplementary·Fig. 3c), MED16 and MED25 have been reported to play opposing roles in ABSCISIC ACID (ABA) responses32. By contrast, both MED16 and MED25 positively regulate the expression of iron homeostasis-related genes in Arabidopsis36. In this study, MED16 stabilized MED25 and performed functions similar to those of MED25 in the regulation of multiple JA responses. This suggests that a stable MED25 is critical for the activation of JA responses. Notably, the vWF-A domain of MED25 has been implicated in the recruitment of MED25 into the Mediator complex in mammals41.
In the current study, we found that the vWF-A domain is evolutionarily conserved and is indispensable for the interaction of MED25 with MED16 (Fig. 1d and Supplementary Fig. 2). MED16 competes with MBRs to bind to the vWF-A domain of MED25 and stabilizes MED25 by recruiting it into the Mediator complex. Thus, MED16 and MBRs form a regulatory module that robustly and tightly regulates MED25 homeostasis, which determines the strength of the transcriptional output of JA signaling.
Methods
Plant materials and growth conditions
Arabidopsis thaliana ecotype Columbia (Col-0) was used as the WT. The following materials were used in this study and have been previously described: MYC2-myc39, med16-1 (sfr6-1)29,32, med16-2 (SALK_048091)29,32, med16-3 (CS859103)29, MED16-GFP40, med25-422, med25-4 myc2-222. The MYC2-myc/med16-3 transgenic line was generated by introducing the MYC2-myc transgene into the med16-3 background via crossing. The mbr1-1 and mbr2-1 mutants were generated in the WT background using the CRISPR/Cas9 technology, and homozygous plants were identified by DNA sequencing. The mbr1-1 mbr2-1 double mutant was generated by crossing mbr1-1 and mbr2-1. The med16-3 med25-4 and med16-3 myc2-2 double mutants and med25-4 mbr1-1 mbr2-1 triple mutant were generated via crossing, and homozygous plants were selected by genotyping. Other Arabidopsis plants were grown at 22 °C under long-day photoperiod (16 h light/8 h dark) and 120 μmol photons m-2 s-1 light intensity. For MeJA treatment, 8 mL liquid 1/2 MS medium containing MeJA at a final concentration of 100 μM was added to the plate for the indicated durations. JA-induced root growth inhibition assays were performed as described previously39 using seedlings grown on 1/2 MS medium supplemented with or without 20 μM JA. Nicotiana benthamiana plants were grown under long-day conditions (16 h light [28 °C]/8 h dark [22 °C]).
Plasmid construction and plant transformation
The 35Spro:MBR2-GFP construct (MBR2-GFP) was generated in two steps; first, the coding sequence (CDS) of GFP was amplified and cloned into pCAMBIA1300 to obtain pCAMBIA1300-GFP, and then the CDS of MBR2 was amplified and cloned into pCAMBIA1300-GFP. To construct 35Spro:MED16-GFP (MED16-GFP), the MED16 CDS was amplified and cloned into pCAMBIA1300-GFP. Primers used for plasmid construction are listed in Supplementary Data 2.
The constructs were transformed into Agrobacterium tumefaciens strain GV3101, which was used to transform Arabidopsis plants via the floral dip method. Transformants were selected based on their resistance to hygromycin, and homozygous T3 lines were used for subsequent experiments.
Generation of the mbr mutant using CRISPR/Cas9 technology
Twenty-base pair (bp) fragments of the MBR1 CDS (22–41 bp) and MBR2 CDS (64–83 bp) were used as the targeting sequences for genome editing of MBR1 and MBR2, respectively. The designed targeting sequences were cloned into the BsaI site of the AtU6-26-sgRNA-SK vector to generate AtU6-26-MBR1-targetsgRNA and AtU6-26-MBR2-targetsgRNA. The AtU6-26-MBR1-targets-gRNA was digested with SpeI and NheI, and the cassette was cloned into the SpeI position of the pYAO:hSpCas9 vector to generate pYAO:hSpCas9-MBR1-targetsgRNA. Similarly, the AtU6-26-MBR2-targetsgRNA was digested with SpeI and NheI, and the cassette was cloned into the SpeI position of the pYAO:hSpCas9-MBR1-targetsgRNA vector to generate pYAO:hSpCas9-MBR1-MBR2-targetsgRNA. The resultant constructs were individually transformed into A. tumefaciens strain GV3101, which was used to transform WT Arabidopsis plants via the floral dip method. Further selection was based on their hygromycin resistance, and target gene mutation was verified by DNA sequencing. Cas9-free T2 plants harboring mutations in MBR1 or MBR2 were identified for further experiments.
Purification and MS analysis of MYC2 protein complexes
Samples (5 g) of 10-d-old MYC2-myc seedlings were harvested, ground in liquid nitrogen, and lysed with 10 mL of ice-cold extraction buffer (50 mM Tris-HCl [pH 7.5], 150 mM NaCl, 5 mM EDTA, 0.1% [v/v] Triton X-100, 0.2% [v/v] Nonidet P-40, 0.6 mM phenylmethylsulfonyl fluoride [PMSF], and 20 μM MG132 with Roche protease inhibitor cocktail). After vortexing for 30 s, the samples were centrifuged at 13,000 × g for 10 min at 4 °C. For each sample, 30 μL of the supernatant was subjected to immunoblot analysis, and the remainder was incubated with 50 μL of anti-myc antibody-bound agarose beads (Proteintech, yta-100) for 4 h at 4 °C with gentle shaking. The agarose beads were collected and washed four times with extraction buffer and once with 50 mM Tris-HCl (pH 7.5). The precipitate was eluted using 1× SDS protein loading buffer and separated using 10% SDS-PAGE. The gel was stained with Thermo GelCode Blue Safe Protein Stain and washed with double-distilled water.
The gel was cut into small pieces and destained in buffer containing 25 mM ammonium bicarbonate and 50% (v/v) acetonitrile (pH 8.0). Proteins were reduced with 10 mM DTT at 37 °C for 1 h and then alkylated with 25 mM iodoacetamide at room temperature for 1 h in the dark. In-solution trypsin digestion was performed overnight at 37 °C using a trypsin: substrate ratio of 1:50. Peptides were extracted from the gel with buffers containing 5% (v/v) trifluoroacetic acid and 50% (v/v) acetonitrile (pH 0.5) after two rounds of ultrasonication. Liquids were freeze-dried in a SpeedVac, and peptides were resolubilized in 0.1% (v/v) formic acid and filtered through a 0.45-µm centrifugal filter.
Peptides were analyzed using a TripleTOF 5600 mass spectrometer (AB SCIEX) coupled to an online Eksigent nanoLC Ultra HPLC system in the information-dependent mode. The LC gradient (solvent A = 0.1% formic acid in water; solvent B = 0.1% formic acid in acetonitrile) used for sample elution was 5% to 90% solvent B for 90 min at a flow rate of 300 nL min−1.
Peptides were identified by examining the tandem mass spectrometric spectra using ProteinPilot v4.2 and by searching against the Arabidopsis International Protein Index database. Carbamidomethylation of cysteine (Cys) residues was used as the fixed modification. Trypsin was specified as the proteolytic enzyme, allowing two missed cleavage sites. Mass tolerance was set to 0.05 D, and the maximum FDR for proteins and peptides was set to 1%.
Purification of ubiquitinated proteins
Purification of ubiquitinated proteins was performed as described with modifications46. Wild-type and MED25-myc transgenic plants were grown in 1/2 MS medium for 10 days and were then treated with 50 mM MG132 for 6 h. Total proteins were extracted with 1 ml of buffer I (50 mM Tris-Cl, (pH 7.5), 20 mM NaCl, 0.1% NP-40 and 5 mM ATP) in a prechilled mortar. The following were added to the protein homogenates: 1 mM PMSF, 50 mM MG132, 10 nM Ub aldehyde (Sigma-Aldrich, cat#662056), and 10 mM N-ethylmaleimide (Thermo, cat#23030). After proteins were quantified, 2 mg of total proteins in a total volume of 2 ml was used for the assay. A 100 μl volume of protein supernatants was reserved as input. Other protein supernatants were incubated with 30 μl of prewashed myc-trap beads (Proteintech, cat#yta-100) in 2 ml of buffer I at 4 °C. After 4 h, the agaroses were washed twice with buffer I and twice with buffer II (supplemented with 200 mM NaCl in buffer I). Samples were boiled in 50 ml of 1× SDS loading buffer for 5 min. The ubiquitinated proteins were separated by 10% SDS–PAGE gel, and anti-ubiquitin antibody (Thermo fisher, cat#14-6078-82) was used to detect ubiquitinated-MED25-myc protein.
Y2H assays
The CDSs of MED16, MYC2, MBR1, MBR2, MED25, and MED25-derivatives were cloned into pGADT7 and/or pGBKT7 using sequence-specific primers (Supplementary Data 2) to generate fusions with the GAL4 AD and/or GAL4 BD, respectively. The resultant constructs were cotransformed into yeast (Saccharomyces cerevisiae) strain AH109. The presence of transgenes was confirmed by growth on SD/-Leu/-Trp (SD/-2) medium (Clontech). To assess protein–protein interactions, the transformed yeast cells were suspended in liquid SD/-Leu/-Trp to OD = 1.0. Five-microliter samples of suspended yeast cells were plated on the SD/-Ade/-His/-Leu/-Trp (SD/-4) medium (Clontech). The plates were incubated at 30 °C, and protein–protein interactions were examined after 3 days.
Y3H assays
The full-length CDS of MED16 was cloned into the pGADT7 vector. To construct pBridge-MED25-MBR1 or pBridge-MED25-MBR2, the MED25 CDS was cloned into the multiple cloning site (MCS) I of the pBridge vector (Clontech) to generate a fusion with the GAL4 BD domain, and the CDS of MBR1 or MBR2 was cloned into the MCS II of the pBridge vector to express as the “bridge” protein only in the absence of methionine. Y3H assays were based on the MATCHMAKER GAL4 Two-Hybrid System (Clontech). The constructs used to test protein–protein interactions were cotransformed into S. cerevisiae strain AH109. The presence of transgenes was confirmed by growing the yeast cells on SD/-2 plates. The transformed yeast cells were plated on SD/-4 medium to assess the MED25–MED16 interaction without the expression of MBR1 or MBR2. Plates containing the SD/-Ade/-His/-Leu/-Trp/-Met (SD/-5) medium were used to induce MBR1 or MBR2 expression. Interactions were observed after 3 days of incubation at 30 °C. To construct pBridge-MED25-MED16, the CDSs of MED25 and MED16 were cloned into the MCS I and MCS II of the pBridge vector, respectively. MBR16 was expressed as the “bridge” protein to test its effects on MED25–MBR1 and MED25–MBR2 interactions. The experimental procedures were the same as those described above.
In vitro ubiquitination assays
MBR2C was cloned into the pGEX-4T-3 vector and expressed in Escherichia coli. Fusion proteins were prepared according to the manufacturer’s instructions. In vitro E3 ligase assays were performed as described previously45. Briefly, 30-μL reactions, each containing 1.5 μL of 20× reaction buffer (1 M Tris-HCl [pH 7.4], 200 mM MgCl2, 100 mM ATP, and 40 mM DTT), recombinant His-tagged wheat (Triticum aestivum) E1 (50 ng), human E2 (UBCh5b; ∼200 ng), and purified GST-tagged E3 (∼1 μg), and purified His-tagged Arabidopsis ubiquitin (UBQ14, AT4G02890; ∼2 μg), were prepared. The reactions were incubated at 37 °C for 90 min, and proteins were separated by SDS-PAGE. Anti-ubiquitin antibody was used to detect His-tagged ubiquitin, and anti-Flag antibody was used to detect Flag-tagged MED25.
Antibody production
The CDSs of MED8 and MED351-350 were PCR-amplified from WT cDNA using gene-specific primers (Supplementary Data 2), and the resultant PCR products were cloned into the pGEX-4T-1 vector using 2×MultiF Seamless Assembly Mix (Abclonal, cat#RK21020) to express the GST-MED8 and GST-MED351-350 fusion proteins in E. coli BL21 (DE3). Similarly, the CDS of MED16 was PCR-amplified from WT cDNA using gene-specific primers (Supplementary Data 2), and the resultant PCR products were cloned into the pET28a vector to express the His-MED16 fusion protein in E. coli Rossetta (DE3). The E. coli cells were induced with 0.4 mM IPTG at 16°C for 16 h. GST Bind Resin (Millipore, cat#70541-4) or Ni-NTA resin (Novagen, cat#70666) was used to purify the recombinant fusion proteins. Antibody production was performed as described with modifications62,63. Briefly, purified recombinant proteins were concentrated and boiled. The samples were then separated using SDS-PAGE and the specific bands at approximately 160 kDa were excised from the gels and ground into powder under liquid nitrogen. The powdered samples were sent to the Antibody Center at the Institute of Genetics and Developmental Biology to immunize mice for antibody generation. Anti-MED8, anti-MED16, and anti-MED35 antibodies were used in protein gel blotting assays at a final dilution of 1:1,000.
Identification of full-length His-MED16 by MS
The recombinant protein excised from SDS-PAGE gel was digested by in-gel digestion. Briefly, samples were reduced with 10 mM DTT in 50 mM ammonium bicarbonate at 56 °C for 1 h and alkylated with 55 mM iodoacetamide in 50 mM ammonium bicarbonate in the dark for 45 min, and digested with trypsin at 37 °C overnight. Peptides were extracted from gel with buffers containing 5% trifluoroacetic acid and 50% acetonitirile by ultrasonic twice. The liquid were freeze dried by SpeedVac, and peptides were desalted by StageTip.
For MS analyzes, peptides were resuspended in 0.1% FA and analyzed by LTQ Orbitrap Elite mass spectrometer (ThermoFisher Scientific) coupled online to an Easy-nLC 1000 (Thermo Fisher Scientific) in the data-dependent mode. The peptides were separated by reverse phase LC with an 150 μm (ID) × 250 mm (length) analytical column packed with C18 particles of 1.9 µm diameter. The mobile phases for the LC contains buffer A (0.1 % FA) and buffer B (100% ACN, 0.1 % FA), and a non-linear gradient of buffer B from 3%–30% for 90 min was used for the separation. Precursor ions were measured in the Orbitrap analyzer at 240,000 resolution (at 400 m/z) and a target value of 106 ions. The twenty most intense ions from each MS scan were isolated, fragmented, and measured in the linear ion trap. The CID normalized collision energy was set to 35.
The data was analyzed using a pre-release version of Thermo Scientific Proteome DiscovererTM software version 1.4. The proteome sequences for Arabidopsis thaliana from TAIR were used for the database searching and the mass tolerance was set to 0.05 Da. The protease used for protein digestion was trypsin. The maximum number of missed cleavages was set at two, the minimum peptide length was set at six amino acids and the maximum peptide length was set at 144 amino acids. The false discovery rate was set at 0.01 for peptide and protein identifications. Cysteine carbamidomethylation and methionine oxidation were included in the search as the static modification and variable modification, respectively.
Protein expression and in vitro pull-down assays
To produce GST-MBR2C, the CDS of MBR2(316-666) was amplified and cloned into the pGEX-4T-1 vector. Protein expressed in E. coli BL21 (DE3) was purified using GST Bind Resin (Novagen, cat#70541-4). To produce MED25-Flag, the CDS of MED25 was amplified and cloned into the pF3K WG (BYDV) Flexi® Vector (Promega). Proteins were in vitro translated using the TnT® SP6 High-Yield Wheat Germ Protein Expression System. To perform pull-down assays, 5 μL of in vitro translated MED25-Flag protein was incubated with 1 μg immobilized GST-MBR2C or GST at 4 °C in binding buffer (25 mM Tris-HCl [pH 7.5], 100 mM NaCl, 1 mM DTT, and Roche protease inhibitor cocktail) for 2 h. Proteins retained on the beads were analyzed by immunoblotting using anti-Flag antibody (Abmart, cat#M20008L).
To conduct competitive pull-down assays, 1 μg of GST- MBR2C with 1, 2 or 5 μg of His-MED16 were incubated with immobilized MED25-Flag at 4 °C in binding buffer (25 mM Tris-HCl [pH 7.5], 100 mM NaCl, 1 mM DTT, and Roche protease inhibitor cocktail) for 2 h. Proteins retained on the beads were analyzed by immunoblotting with anti-His (Easybio, cat#BE2019-100), anti-GST (Abmart, cat#M20007M), and anti-Flag (Abmart, cat#M20008L) antibodies.
Co-IP assays
Ten-day-old MED16-GFP or MBR2-GFP transgenic seedlings were homogenized in extraction buffer (50 mM Tris-HCl [pH 7.5], 150 mM NaCl, 0.1% Triton X-100, 0.2% Nonidet P-40, 0.6 mM PMSF, and 20 μΜ MG132 with Roche protease inhibitor cocktail). WT seedlings were used as a negative control. After protein extraction, 20 μL of protein A/G plus agarose (Santa Cruz Biotechnology, cat#sc-2003) was added to 2 mg of the extracted proteins to reduce non-specific immunoglobulin binding. After 1 h of incubation, the supernatant was transferred to a new tube. GFP antibody-bound agarose beads (Proteintech, cat#gta-100) were added to the samples, which were then incubated for 4 h at 4 °C with gentle rocking. The precipitated samples were washed at least four times with protein extraction buffer, and the bound proteins were eluted by heating the beads in 1× SDS protein loading buffer at 95 °C for 6 min. MED25 was detected by immunoblotting using anti-MED25 antibody22. To test the MYC2–MED16 interaction in vivo, co-IP assays were performed (as described above) using MYC2-myc transgenic plants and anti-MED16 antibody.
To investigate the effect of MED16 on the MYC2–MED25 interaction, co-IP assays were performed using MYC2-myc and MYC2-myc/med16-3 transgenic plants, followed by immunoblotting using anti-MED25 antibody22 and anti-myc antibody (Abmart, 1:2000, cat#M20002L).
Immunoblot and gel-filtration assays
To analyze the MED25 protein level, protein extraction was performed by homogenizing 10-day-old seedlings in extraction buffer (50 mM Tris-HCl [pH 7.5], 150 mM NaCl, 1% Nonidet P-40, 1 μM DTT, 10 μM MG132, and Roche protease inhibitor cocktail). SDS sample buffer was added to the protein extracts, following which the protein samples were boiled for 5 min, separated on SDS-PAGE gels, and transferred to polyvinylidene fluoride membranes. Immunoblots were probed with anti-MED25 antibody. ACTIN protein was used as a control. Gel-filtration analysis was performed as described previously64, with a minor modification in the buffer composition (50 mM Tris [pH 7.5], 150 mM NaCl, 1 mM EDTA, 0.3% Triton X-100, 1 mM DTT, 1 mM PMSF, and 1× complete protease inhibitor cocktail [Roche]) used for lysis and gel filtration. A Superose 6 (Amersham) column was used for gel filtration.
RNA extraction, reverse transcription (RT), and RT-qPCR
To analyze gene expression, total RNA was extracted using TRIzol (Invitrogen) reagent from 10-day-old seedlings treated with or without 100 μM MeJA for the indicated durations. Then, cDNA was prepared from 2 μg of total RNA using PrimeScriptTM RT Reagent Kit (TaKaRa, cat#RR0447A), and quantified on Roche 480 cycler using KAPA SYBR FAST qPCR Master Mix (KAPA, cat#KK4601), according to the manufacturer’s instructions. The expression levels of target genes were determined by RT-qPCR using gene-specific primers (Supplementary Data 2) and normalized relative to the expression level of ACTIN7. Three independent biological replicates were performed. Data were expressed as mean ± standard deviation and statistically analyzed using one-way analysis of variance (ANOVA).
Statistics and reproducibility
Significant differences were analyzed using two-sample Student’s t tests or one-way ANOVA with GraphPad Prism 8.0. All experiments were repeated independently three times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information files. Nucleotide sequences of genes examined in this study can be found in The Arabidopsis Information Resource (TAIR) website (https://www.arabidopsis.org) under the following accession numbers: MED8, AT2G03070; MED16, AT4G04920; MED18, AT2G22370; MED25, AT1G25540; MED35, AT1G44910; MYC2, AT1G32640; MBR1, AT2G15530; MBR2, AT4G34040; VSP1, AT5G24780; VSP2, AT5G24770; PDF1.2, AT5G44420; Thi2.1, AT1G72260; ACTIN7, AT5G09810. The MS data of full-length His-MED16 protein identification was deposited to the ProteomeXchange via the PRIDE database under the identifier PXD056008. Any additional information is available from the corresponding author upon request. Source data are provided with this paper.
References
Browse, J. Jasmonate passes muster: a receptor and targets for the defense hormone. Annu Rev. Plant Biol. 60, 183–205 (2009).
Wasternack, C. & Hause, B. Jasmonates: biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 111, 1021–1058 (2013).
Zhai, Q., Deng, L. & Li, C. Mediator subunit MED25: at the nexus of jasmonate signaling. Curr. Opin. Plant Biol. 57, 78–86 (2020).
Zhai, Q. & Li, C. The plant Mediator complex and its role in jasmonate signaling. J. Exp. Bot. 70, 3415–3424 (2019).
Boter, M., Ruíz-Rivero, O., Abdeen, A. & Prat, S. Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev. 18, 1577–1591 (2004).
Dombrecht, B. et al. MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 19, 2225–2245 (2007).
Du, M. et al. MYC2 orchestrates a hierarchical transcriptional cascade that regulates jasmonate-mediated plant immunity in tomato. Plant Cell 29, 1883–1906 (2017).
Kazan, K. & Manners, J. M. MYC2: the master in action. Mol. Plant 6, 686–703 (2013).
Lorenzo, O., Chico, J. M., Sánchez-Serrano, J. J. & Solano, R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16, 1938–1950 (2004).
Zhai, Q. et al. Phosphorylation-coupled proteolysis of the transcription factor MYC2 is important for jasmonate-signaled plant immunity. PLoS Genet 9, e1003422 (2013).
Chini, A. et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671 (2007).
Pauwels, L. et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464, 788–791 (2010).
Sheard, L. B. et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 468, 400–405 (2010).
Thines, B. et al. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature 448, 661–665 (2007).
Yan, Y. et al. A downstream mediator in the growth repression limb of the jasmonate pathway. Plant Cell 19, 2470–2483 (2007).
Fonseca, S. et al. (+)-7-iso-Jasmonoyl-L-isoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 5, 344–350 (2009).
Yan, J. et al. The Arabidopsis CORONATINE INSENSITIVE1 protein is a jasmonate receptor. Plant Cell 21, 2220–2236 (2009).
Zhang, F. et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 525, 269–273 (2015).
Wang, H. et al. MED25 connects enhancer-promoter looping and MYC2-dependent activation of jasmonate signalling. Nat. Plants 5, 616–625 (2019).
An, C. et al. Mediator subunit MED25 links the jasmonate receptor to transcriptionally active chromatin. Proc. Natl Acad. Sci. USA 114, E8930–E8939 (2017).
Çevik, V. et al. MEDIATOR25 acts as an integrative hub for the regulation of jasmonate-responsive gene expression in Arabidopsis. Plant Physiol. 160, 541–555 (2012).
Chen, R. et al. The Arabidopsis mediator subunit MED25 differentially regulates jasmonate and abscisic acid signaling through interacting with the MYC2 and ABI5 transcription factors. Plant Cell 24, 2898–2916 (2012).
You, Y., Zhai, Q., An, C. & Li, C. LEUNIG_HOMOLOG mediates MYC2-dependent transcriptional activation in cooperation with the coactivators HAC1 and MED25. Plant Cell 31, 2187–2205 (2019).
Liu, Y. et al. MYC2 regulates the termination of jasmonate signaling via an autoregulatory negative feedback loop. Plant Cell 31, 106–127 (2019).
Wu, F. et al. Mediator subunit MED25 couples alternative splicing of JAZ genes with fine-tuning of jasmonate signaling. Plant Cell 32, 429–448 (2020).
Cerdán, P. D. & Chory, J. Regulation of flowering time by light quality. Nature 423, 881–885 (2003).
Kazan, K. The multitalented MEDIATOR25. Front Plant Sci. 8, 999 (2017).
Iñigo, S., Giráldez, A. N., Chory, J. & Cerdán, P. D. Proteasome-mediated turnover of Arabidopsis MED25 is coupled to the activation of FLOWERING LOCUS T transcription. Plant Physiol. 160, 1662–1673 (2012).
Knight, H. et al. Identification of SFR6, a key component in cold acclimation acting post-translationally on CBF function. Plant J. 58, 97–108 (2009).
Wang, C. et al. The Arabidopsis Mediator complex subunit16 is a key component of basal resistance against the necrotrophic fungal pathogen Sclerotinia sclerotiorum. Plant Physiol. 169, 856–872 (2015).
Wathugala, D. L. et al. The Mediator subunit SFR6/MED16 controls defence gene expression mediated by salicylic acid and jasmonate responsive pathways. New Phytol. 195, 217–230 (2012).
Guo, P. et al. Mediator tail module subunits MED16 and MED25 differentially regulate abscisic acid signaling in Arabidopsis. J. Integr. Plant Biol. 63, 802–815 (2021).
Hemsley, P. A. et al. The Arabidopsis mediator complex subunits MED16, MED14, and MED2 regulate mediator and RNA polymerase II recruitment to CBF-responsive cold-regulated genes. Plant Cell 26, 465–484 (2014).
Liu, Z. et al. Transcriptional repression of the APC/C activator genes CCS52A1/A2 by the Mediator complex subunit MED16 controls endoreduplication and cell growth in Arabidopsis. Plant Cell 31, 1899–1912 (2019).
Wang, C., Du, X. & Mou, Z. The Mediator complex subunits MED14, MED15, and MED16 are involved in defense signaling crosstalk in Arabidopsis. Front Plant Sci. 7, 1947 (2016).
Yang, Y. et al. The Arabidopsis Mediator subunit MED16 regulates iron homeostasis by associating with EIN3/EIL1 through subunit MED25. Plant J. 77, 838–851 (2014).
Zhang, X., Wang, C., Zhang, Y., Sun, Y. & Mou, Z. The Arabidopsis mediator complex subunit16 positively regulates salicylate-mediated systemic acquired resistance and jasmonate/ethylene-induced defense pathways. Plant Cell 24, 4294–4309 (2012).
Zhang, X., Yao, J., Zhang, Y., Sun, Y. & Mou, Z. The Arabidopsis Mediator complex subunits MED14/SWP and MED16/SFR6/IEN1 differentially regulate defense gene expression in plant immune responses. Plant J. 75, 484–497 (2013).
Chen, Q. et al. The basic helix-loop-helix transcription factor MYC2 directly represses PLETHORA expression during jasmonate-mediated modulation of the root stem cell niche in Arabidopsis. Plant Cell 23, 3335–3352 (2011).
Zhang, Y. et al. Mediator subunit 16 functions in the regulation of iron uptake gene expression in Arabidopsis. New Phytol. 203, 770–783 (2014).
Mittler, G. et al. A novel docking site on Mediator is critical for activation by VP16 in mammalian cells. EMBO J. 22, 6494–6504 (2003).
Richter, W. F., Nayak, S., Iwasa, J. & Taatjes, D. J. The Mediator complex as a master regulator of transcription by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 23, 732–749 (2022).
Berger, S., Bell, E., Sadka, A. & Mullet, J. E. Arabidopsis thaliana Atvsp is homologous to soybean VspA and VspB, genes encoding vegetative storage protein acid phosphatases, and is regulated similarly by methyl jasmonate, wounding, sugars, light and phosphate. Plant Mol. Biol. 27, 933–942 (1995).
Penninckx, I. A. et al. Pathogen-induced systemic activation of a plant defensin gene in Arabidopsis follows a salicylic acid-independent pathway. Plant Cell 8, 2309–2323 (1996).
Zhao, Q., Liu, L. & Xie, Q. In vitro protein ubiquitination assay. Methods Mol. Biol. 876, 163–172 (2012).
Kong, L. et al. Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases. Nat. Commun. 6, 8630 (2015).
Yan, L. et al. High-efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol. Plant 8, 1820–1823 (2015).
Soutourina, J. Transcription regulation by the Mediator complex. Nat. Rev. Mol. Cell Biol. 19, 262–274 (2018).
Allen, B. L. & Taatjes, D. J. The Mediator complex: a central integrator of transcription. Nat. Rev. Mol. Cell Biol. 16, 155–166 (2015).
Conaway, R. C. & Conaway, J. W. The Mediator complex and transcription elongation. Biochim Biophys. Acta 1829, 69–75 (2013).
Jeronimo, C. & Robert, F. The Mediator complex: at the nexus of RNA polymerase II transcription. Trends Cell Biol. 27, 765–783 (2017).
Malik, S. & Roeder, R. G. The metazoan Mediator co-activator complex as an integrative hub for transcriptional regulation. Nat. Rev. Genet 11, 761–772 (2010).
Bäckström, S., Elfving, N., Nilsson, R., Wingsle, G. & Björklund, S. Purification of a plant mediator from Arabidopsis thaliana identifies PFT1 as the Med25 subunit. Mol. Cell 26, 717–729 (2007).
Li, X., Yang, R. & Chen, H. The Arabidopsis thaliana Mediator subunit MED8 regulates plant immunity to Botrytis Cinerea through interacting with the basic helix-loop-helix (bHLH) transcription factor FAMA. PLoS One 13, e0193458 (2018).
Lai, Z. et al. MED18 interaction with distinct transcription factors regulates multiple plant functions. Nat. Commun. 5, 3064 (2014).
Zhu, Y. et al. CYCLIN-DEPENDENT KINASE8 differentially regulates plant immunity to fungal pathogens through kinase-dependent and -independent functions in Arabidopsis. Plant Cell 26, 4149–4170 (2014).
Huang, Y. et al. Mediator complex regulates alternative mRNA processing via the MED23 subunit. Mol. Cell 45, 459–469 (2012).
Takahashi, H. et al. Human mediator subunit MED26 functions as a docking site for transcription elongation factors. Cell 146, 92–104 (2011).
Schneider, M. et al. The nuclear pore-associated TREX-2 complex employs Mediator to regulate gene expression. Cell 162, 1016–1028 (2015).
Warren, G., McKown, R., Marin, A. L. & Teutonico, R. Isolation of mutations affecting the development of freezing tolerance in Arabidopsis thaliana (L.) Heynh. Plant Physiol. 111, 1011–1019 (1996).
Raya-González, J. et al. MEDIATOR16 orchestrates local and systemic responses to phosphate scarcity in Arabidopsis roots. New Phytol. 229, 1278–1288 (2021).
Amero, S. A., James, T. C. & Elgin, S. C. Production of antibodies using proteins in gel bands. Methods Mol. Biol. 32, 401–406 (1994).
Tjian, R., Stinchcomb, D. & Losick, R. Antibody directed against Bacillus subtilis rho factor purified by sodium dodecyl sulfate slab gel electrophoresis. effect on transcription by RNA polymerase in crude extracts of vegetative and sporulating cells. J. Biol. Chem. 250, 8824–8828 (1975).
Saijo, Y. et al. The COP1-SPA1 interaction defines a critical step in phytochrome A-mediated regulation of HY5 activity. Genes Dev. 17, 2642–2647 (2003).
Acknowledgements
We thank the Animal Center of Institute of Genetics and Developmental Biology, Chinese Academy of Sciences (CAS) for generating the antibodies. We also thank Dr. Yingfang Zhu for providing the seeds of med16-1 and med16-2; and Dr. Hong-Qing Ling for providing the seeds of MED16-GFP. This work was supported by National Natural Science Foundation of China (32161133018, 32370332, 31991183) and National Key R&D Program of China (2022YFD1400800).
Author information
Authors and Affiliations
Contributions
C.Y.L. and F.M.W. conceived and designed the study. C.Y.L. supervised the project. F.M.W., C.L.S., and Z.Y.Z. conducted the experiments and analyzed the data. F.F.Y. and Q.X. contribute to ubiquitination assays. C.Y.L., F.M.W., C.L.S., and L.D. drafted the manuscript. All authors approved the final version of the manuscript. The corresponding author attests that all the listed authors meet the authorship criteria and that no other individual meeting the criteria has been omitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, F., Sun, C., Zhu, Z. et al. A multiprotein regulatory module, MED16–MBR1&2, controls MED25 homeostasis during jasmonate signaling. Nat Commun 16, 772 (2025). https://doi.org/10.1038/s41467-025-56041-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56041-3
This article is cited by
-
Unlocking tropical adaptation: novel flowering time genes implicate JA and ROS signaling in maize
BMC Plant Biology (2026)