Abstract
Dysregulated microglia activation, leading to neuroinflammation, is crucial in neurodegenerative disease development and progression. We constructed an atlas of human brain immune cells by integrating nineteen single-nucleus RNA-seq and single-cell RNA-seq datasets from multiple neurodegenerative conditions, comprising 241 samples from patients with Alzheimer’s disease, autism spectrum disorder, epilepsy, multiple sclerosis, Lewy body diseases, COVID-19, and healthy controls. The integrated Human Microglia Atlas (HuMicA) included 90,716 nuclei/cells and revealed nine populations distributed across all conditions. We identified four subtypes of disease-associated microglia and disease-inflammatory macrophages, recently described in mice, and shown here to be prevalent in human tissue. The high versatility of microglia is evident through changes in subset distribution across various pathologies, suggesting their contribution in shaping pathological phenotypes. A GPNMB-high subpopulation was expanded in AD and MS. In situ hybridization corroborated this increase in AD, opening the question on the relevance of this population in other pathologies.
Similar content being viewed by others
Introduction
Microglia are central immune mediators in the brain. They originate from myeloid precursors and migrate from the yolk sac to the brain parenchyma during the embryonic stage1,2. In the brain of healthy individuals, microglia orchestrate a wide network of processes essential for the development and maintenance of the normal functioning central nervous system (CNS), including modulation of neuronal connectivity through synaptic remodelling and structural organization, myelination, blood-brain barrier integrity and vasculogenesis. In pathological contexts, microglia generally acquire an activated state, transitioning from a resting elongated and ramified form to an amoeboid macrophage-like morphology, accompanied by an overall enhanced pro-inflammatory phenotype3. Despite being a normal physiological mechanism, unbalanced microglia activation has consistently been associated with neurological disease4,5.
The landscape of microglia activation was initially described by the M1/M2 polarization, wherein the M1 pro-inflammatory phenotype was considered neurotoxic and the M2 tolerogenic phenotype was thought to be neuroprotective6. The first description of a pathology-associated microglia gene expression signature was performed by Chiu and collaborators in a mice model of amyotrophic lateral sclerosis7. In 2017, Keren-Shaul et al. described what has been regarded as the consensus microglia pathology-associated phenotype, known as the “disease-associated microglia” (DAM). This milestone was achieved through the application of single-cell (sc) transcriptomics in a mice model of Alzheimer’s disease (AD). The authors identified phenotypically distinct subpopulations of microglia that were enriched in pathological conditions. These subpopulations exhibited downregulation of homeostatic microglia markers and activation of pathways involved in the immune response and phagocytosis8.
The implementation of single-nucleus (sn)RNA-seq methods has enabled the analysis of frozen tissue samples at the cellular level9, thereby maximizing the utilization of numerous previously frozen brain samples. This advancement has paved the way for human brain tissue studies across various neurological diseases, such as AD10,11,12,13,14,15, multiple sclerosis (MS)16,17, autism spectrum disorder (ASD)18 and Lewy Body Diseases (LBD)19. Individual attention has been devoted to microglia, revealing evidence of dysregulated gene expression in pathology and, in certain cases, an enrichment of DAM genes in subpopulations of microglia expanded in pathology. These findings suggest an inter-disease prevalence of this microglial phenotype. However, it is understandable that most of these studies place greater emphasis on the most abundant and traditionally considered more relevant cell types, particularly neuron populations.
It is becoming increasingly evident that the M1/M2 activation and the DAM phenotype are inadequate to fully comprehend the complexity of microglia behavior in both health and disease. It can be inferred that the plasticity of microglial function in homeostasis and injury/disease may follow a multi-subpopulation pattern influenced by the type of disease, sex, age and even spatial localization20,21.
In the human brain, the comprehensive characterization of microglia subpopulations is significantly limited by the low yield of microglia within individual datasets. To address this limitation, we performed an in silico integration of multiple snRNA-seq and scRNA-seq datasets from human brain tissue. These datasets included samples from patients diagnosed with AD, MS, ASD, LBD and epilepsy, as well as samples collected from neurologically healthy individuals. Recent studies demonstrated a pro-inflammatory DAM-like phenotype in severe COVID-19 brain tissue22,23. Considering these findings and the high incidence of long-term cerebral symptomatology associated with SARS-COV2 infection24,25,26, we deemed it relevant to include snRNA-seq data from severe COVID-19 patients.
In this study, we have successfully generated a comprehensive cross-methodology Human Microglia Atlas (HuMicA) consisting of 90,716 cells/nuclei, derived from 241 individual samples across nineteen datasets and seven clinical settings. We observed an expansion of a microglia activated subpopulation with lipid-related transcriptomic traits in neuropathology, further validated by in situ RNA hybridization in postmortem human brain sections. Our results indicate that the HuMicA is a unique resource and toolset that will facilitate further studies in human microglia in both health and disease.
Results
Integration of the Human Microglia Atlas (HuMicA)
A total of nineteen publicly available snRNA-seq and scRNA-seq datasets on human brain tissue samples were utilized for this study (Table 1). These datasets encompassed samples from patients diagnosed with AD, ASD, epilepsy, MS, LBD and severe COVID-19, as well as samples from individuals without any neurodegenerative diagnosis. Detailed information of each individual subject included in this study is available in Supplementary Data 1. The integration pipeline (Fig. 1a) started by separately processing each dataset and annotating the main CNS cell types (neurons, oligodendrocytes, astrocytes, oligodendrocyte progenitor cells (OPCs), endothelial cells and immune cells) (Supplementary Fig. 1 and Supplementary Fig. 2) (see Methods). Subsequently, the immune cell clusters from each dataset were integrated, resulting in a final object comprising a total of 102,390 cells/nuclei. We used clustree27 to assess the evolution of hierarchical clustering among all cells as resolution increased, hinting at the ontogenic nature (Supplementary Fig. 3a). We observed two populations at the lowest resolution (0.01), a T cell cluster segregated from the bulk of the myeloid cells. Furthermore, at a clustering resolution of 0.025, an additional population emerged from the myeloid population, showing increased expression of markers from other CNS cell-types, which we considered doublets (Supplementary Fig. 3a–c). For the development of the HuMicA, only the main myeloid population at 0.025 resolution (cluster 0) was considered, whereas T cells and doublets were excluded.
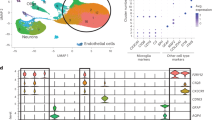
a Schematic overview of the pipeline used to obtain the integrated HuMicA object. b UMAP visualization of the integrated and clustered HuMicA object, annotated by the nine obtained clusters. c Barplot representation of the number of nuclei/cells and the mean genes detected per nucleus/cell (between parentheses) in each cluster. d Dot plot representation of the expression of canonical markers of microglia (P2RY12, CX3CR1) and macrophages (MRC1, CD163) across all clusters. e UMAPs showing the module score (MSc) expression of macrophage (MRC1, CD163) markers. The gene expression values correspond to the normalized “RNA” assay of the integrated Seurat object. f Heatmap representation of the average expression values by cluster of the significant markers of all HuMicA clusters. Gene expression is represented as the z-score of the averaged normalized “RNA” counts. The number of significant markers for each cluster is represented, as well as the most significant and biologically relevant genes from each list. The dot plot represents a gene set enrichment analysis, performed using fgsea with one-tailed test, depicting the significance of the overlap between the lists of genes from the literature and the expression patterns in each HuMicA cluster. Significance is represented by the negative of the log of the adjusted p value (Benjamini-Hochberg) and the normalized enrichment score. The dots highlighted in bold are considered statistically significant (FDR < 0.05). Source data are provided as a Source Data file.
The HuMicA was composed by 90,716 cells/nuclei clustered into 9 subpopulations, distributed across 241 samples (Fig. 1b, Supplementary Fig. 3d and Supplementary Data 1). After integration, we observed a uniform distribution of cells/nuclei per cluster in relation to the methodology used (scRNA-seq and snRNA-seq), sex and the state of the sample (postmortem or surgically resected tissue) (Supplementary Fig. 4a–o). In addition, we did observe significant differential distribution of some clusters in relation to age and brain tissue region (Supplementary Fig. 5a–j); however, one must consider the lack of uniformity in the distribution of each of these variables throughout the independent datasets, and the inability to differentiate between dataset/pathology-related and age- or tissue-related effects. Taking into account the incapacity to overcome this limitation with our in silico approach, we base our rationale on the conception that microglia, unlike other CNS cell types, exhibit consistent patterns across different brain sections28. We also assessed potential confounding effects and variance explained for multiple variables. We observed that inter-sample variability is the main contributor to variance, while the other variables demonstrated a residual contribution (Supplementary Fig. 5k). Nevertheless, we ruled out sample bias as the maximum contribution of an individual sample to each of the clusters did not exceed 16% (Supplementary Fig. 5l).
The clustering analysis of the final HuMicA object resulted in the identification of nine clusters (Fig. 1b and Supplementary Fig. 3d), annotated 0 to 8, from the largest to the smallest in terms of cell number (Fig. 1c). As expected, microglia were naturally the predominant cell type, with the exception of cluster 7 which represented border-associated macrophages (MAC, described as located in meninges, choroid plexus and perivascular spaces of the CNS29,30) as shown by the upregulation of known markers (CD163 and MRC1) (Fig. 1d, e, and Supplementary Data 2). Upregulated gene markers were calculated for each cluster in relation to all other, generating distinct cluster-specific signatures (Fig. 1f and Supplementary Data 2). Furthermore, we crossed the lists of markers obtained within the HuMicA with genesets collected from the literature (Supplementary Data 3). These include standard human microglia transcriptomic signatures obtained comparing sorted microglia and bulk cortical tissue (Gosselin 2017 and Galatro 2017)31,32; the original DAM signatures (Keren-Shaul 2017)8; and markers of three populations within the DAM-verse, including the bona-fide DAM, youth-associated microglia (YAM) (prevalent in embryonic and neonatal stages), and disease-inflammatory macrophages (DIMs), microglia-like cells that differentiate from infiltrating monocytes (Silvin 2022)33. Moreover, we incorporated lists of microglia population markers from previous single-cell transcriptomic studies, some whose datasets are included in the HuMicA (Mathys 2019, Olah 2020, Zhou 2020 and Schirmer 2019)10,15,17,34 and others fully independent (Gerrits 2021 and Sun 2023)35,36. In general, we observed a distinctive enrichment of specific transcriptomic patterns within the markers of each HuMicA cluster (explored in detail in the following sections). This demonstrates that the populations observed in our integrated analysis are prevalent in external and independent data, reinforcing the biological replicability of our findings.
Homeostatic microglia clusters
The homeostatic microglia signature is primarily characterized by high expression of the so-called homeostatic genes, particularly P2RY12 and CX3CR1. We observed this pattern predominantly in the largest microglia population, cluster 0, which we designated as Homeos1 (Fig. 1f and Supplementary Data 2). The cross analysis with signatures from other studies (Fig. 1f) showed that the expression profile of cluster 0 related to the literature signatures corresponding to the largest populations also deemed “homeostatic” (Mic0 – Mathys 2019; MG2 – Olah 2020; d – Schirmer 2019; 0 – Gerrits 2021; MG0 – Sun 2023). To further characterize each of the HuMicA clusters, we performed a hierarchical clustering based on the significant markers of each cluster. We plotted a dendrogram from a matrix of count values consisting of the average expression of all cells/nuclei within each cluster for all the unique cluster markers. Clusters 4 and 8 grouped together with Homeos1 (Fig. 2a) and were also considered homeostatic populations (annotated as Homeos2 and Homeos3) (Fig. 2b). In addition, these two clusters also show a similar pattern of geneset enrichment to Homeos1 (cluster 0) (Fig. 1f). The lists of markers for the three homeostatic clusters show a high degree of shared patterns. However, each population also showed exclusive upregulation of specific genes (Fig. 2c), with GRID2 and SERPINE1 being prominent among the genes characteristic of Homeos2 and Homeos3, respectively (Fig. 2c and Supplementary Fig. 6a). Gene ontology (GO) analysis revealed a striking enrichment of GTPase related terms for the three homeostatic clusters. In addition, Homeos3 presents a particular enrichment of chemotaxis and regulation of B cell differentiation (Fig. 2d, Supplementary Fig. 6b and Supplementary Data 4).
a Cluster dendrogram calculated considering the average expression values of all cells/nuclei for each cluster and the unique gene markers for all clusters (distance: “spearman”; clustering: “ward.D”). The box highlights the three grouped clusters that show homeostatic microglia profiles. b UMAP representation of the three clusters annotated as homeostatic microglia. Cluster 0, Homeos1; Cluster 4, Homeos2; Cluster 8, Homeos3. c Dot plot representation of the expression of the top 10 most significant upregulated markers (ordered by avg_log2FC) for the three homeostatic cluster, within the full HuMicA object. The red dashed boxes highlight the top 10 markers for each cluster. The gene expression values correspond to the normalized “RNA” assay of the Seurat object. d Net plot representation of the three most significant gene ontology (GO) terms enriched for the respective markers of the three homeostatic clusters. GO was calculated using enrichGO package followed by the simplify function. The significance of the enrichment is represented in function of the negative of the log of the adjusted p-value (Benjamini-Horchberg). The range of expression of each gene associated with the GO terms is represented by the avg_log2FC. e Representation of the enrichment of the statistically significant (FDR < 0.05) transcription factors (TFs) relative to the comparison of differential expression of each HuMicA homeostatic clusters vs all other. Significance is represented by the normalized enrichment score and the negative of log of the adjusted p value (Benjamini-Horchberg). All presented TFs showed statistically significant enrichment, and the red dashed boxes highlight the TFs with an FDR < 0.05 for each of the homeostatic clusters. Source data are provided as a Source Data file.
To gain insights into the potential transcription factor (TF) involvement, we employed the CollecTRI regulon-target repository37, to predict the activity of regulatory TFs, based on the specific differential expression of each homeostatic cluster. Decreased activity of MYC, previously described as an important factor in microglia activation38, appears to be a general homeostatic feature, shared by the three homeostatic populations. In addition, down activation of TF involved in inflammatory response (NFKB1, RELA) and other factors previously associated with microglia activation, such as SP139, STAT140, TP5341 and AR42, was observed for Homeos1 and Homeos2 (Fig. 2e). The latter also demonstrated activation of BCL6, previously shown to act as a suppressor of pro-inflammatory macrophage activation43 (Fig. 2e).
The disease-associated microglia (DAM) phenotype
The occurrence of a DAM gene signature has been the gold standard for characterizing microglia activation8. Additionally, in a recent study, Silvin and colleagues, described in mice two DAM-related populations with distinct ontogeny. One corresponded to legitimate DAM, while the other consisted of disease-inflammatory macrophages (DIMs), which represent a microglia-like population derived from infiltrating monocytes33. The gene set enrichment analysis from Fig. 1f demonstrated that the DAM signatures were enriched for microglia clusters 1, 3, 5 and 6; whereas DIM signature was explicitly enriched for cluster 2. The evaluation of the module score (MSc) expression of the full list of human genes in the DAM signature (n = 210), based on Keren-Shaul et al.8, and as previously reported for human data by Thrupp et al.44, and the summarized human gene signatures for DAM, YAM and DIMs provided by Silvin et al.33 reinforces these patterns (Fig. 3a). A higher level of YAM signature expression is observed in clusters associated with the DAM signature, highlighting the similarities between these two signatures, which form the basis of the hypothesis regarding the acquisition of an embryonic-related signature in a pathological environment. Nonetheless, our integrated object consists solely of adult tissue, limiting our insights into embryogenic microglia. Furthermore, clusters 1, 2, 3, 5 and 6 (highlighted in Fig. 3b) present characteristic gene expression signatures (Fig. 3a, Supplementary Fig. 7a and Supplementary Data 2), corresponding GO (Supplementary Fig. 7b and Supplementary Data 4) and pathway enrichment calculated using gene set enrichment analysis (GSEA) (Fig. 3c), which guided our annotation. Cluster 1 or Inflam.DAM presents TMEM163 as the main marker and SPP1, a categorical DAM gene, within the list of top markers (Fig. 3a, Supplementary Fig. 7a and Supplementary Data 2); as well as a marked enrichment of pathways related to antigen presentation, pro-inflammatory signaling, namely interferon gamma, leukocyte activation and cytokine production, and phagocytosis (Fig. 3c, Supplementary Fig. 7b and Supplementary Data 4). In addition, cluster 1 markers are enriched for the MG2 inflammatory population from the Sun et al.36 study (Fig. 1f). Cluster 2 or DIMs showed high expression of SLC2A3 and CD83 (Fig. 3a, Supplementary Fig. 7a and Supplementary Data 2) and solely carries the DIM signature, which is supported by the enrichment of multiple pro-inflammatory pathways associated with interleukin signalling (Fig. 3c, Supplementary Fig. 7b and Supplementary Data 4). Notably, cluster 8 (Homeos3) shows a relatively increased expression of the DIM signature, particularly in CD83, DUSP1, and FOS (Fig. 3a and Supplementary Data 2). This finding may support the possibility of a pre-activation state in this population, as indicated by the enrichment of activation-related GO terms (Fig. 2d). Clusters 3 and 5, Ribo.DAM1 and Ribo.DAM2 respectively, demonstrated similar enrichment profiles with a striking overrepresentation of ribosome-related terms (Fig. 3c, Supplementary Fig. 7b and Supplementary Data 4). In addition, both relate with MG3 from the Sun study, denominated Ribosome Biogenesis (Fig. 1f). Nevertheless, each present its own transcriptomic signature, with SYT1 and PCDH9 heading the upregulated markers of Ribo.DAM1, whilst Ribo.DAM2 specifically overexpresses PLEKHA7 and MECOM (Fig. 3a, Supplementary Fig. 7a and Supplementary Data 2). Lastly, cluster 6 was characterized by elevated expression of GPNMB and PTPRG (Fig. 3a, Supplementary Fig. 7a and Supplementary Data 2), and showed enrichment of pathways associated with lysosome, lipoproteins and lipid metabolism (Fig. 3c, Supplementary Fig. 7b and Supplementary Data 4). In accordance, this cluster’s signature was specifically enriched for Sun’s MG4: Lipid Processing (Fig. 1f). The TF enrichment analysis was not as precise in uncovering specific drivers for each microglia population, as it rather demonstrated a dual pattern distinguishing DIMs and DAM-related clusters. The abovementioned MYC, down activated in homeostatic microglia, was enriched only for Inflam.DAM (cluster 1), Ribo.DAM1 (cluster 3) and Ribo.DAM2 (cluster 5); whereas the DIMs (cluster 2) exhibited enhanced involvement of, JUN, SP1, NF-kB subunits, STAT1, FOS, HIF1A supporting their overall pro-inflammatory profile. Of note, MITF, recently shown to drive a disease-associated transcriptional signature in human stem-cell-differentiated microglia45, is also activated in the DIMs. Finally, Lipo.DAM (cluster 6) showed the singular activation of PPARA (Fig. 3d).
a Dot plot representation of the expression of the module scores (MSc) of the original upregulated DAM signature from Keren-Shaul et al.8 and the refined human signatures for DAM, youth-associated microglia (YAM) and disease-inflammatory macrophages (DIMs) from Silvin et al.33, as well as the expression of top 10 most significant upregulated markers (ordered by avg_log2FC) for the five clusters considered within the “DAM-verse”. Cluster 1: Inflam.DAM, Cluster 2: DIMs, Cluster 3: Ribo.DAM1, Cluster 5: Ribo.DAM2 and Cluster 6: Lipo.DAM. The red dashed boxes highlight the expression of the MSc of the DAM, YAM and DIM signatures and the top 10 markers for each cluster. The expression values correspond to the normalized “RNA” assay of the Seurat object. b UMAP representation of the five DAM-related and DIM clusters and border-associated macrophages, MAC. c Barplot representing of selected gene set enrichment analysis (GSEA) terms from the REACTOME, WIKIPATHWAYS and KEGG repositories for the markers of the DAM-related and DIM clusters. Enrichment is represented as a function of the negative of the log of the adjusted p-value and the normalized enrichment score. d Representation of the enrichment of the statistically significant (adjusted p-value (Benjamini-Horchberg) < 0.05) transcription factors (TFs) relative to the comparison of differential expression of each HuMicA DAM and DIM clusters vs all other. Significance is represented by the normalized enrichment score and the negative of the log of the adjusted p-value (FDR). All presented TFs showed statistically significant enrichment, and the red dashed boxes highlight the TFs with an FDR < 0.05 for each of the DAM and DIM clusters. Source data are provided as a Source Data file.
Pathology-associated alterations in microglia subpopulations
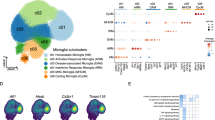
All HuMicA subpopulations were distributed throughout the majority of the subjects and were prevalent in all pathological settings (Supplementary Fig. 8 and Supplementary Data 5). To gain further insights into potential associations between the HuMicA subpopulations and neurodegenerative pathologies, we interrogated the enrichment of each cluster across each pathological settings in comparison to the healthy population, using the sccomp tool46. Our analysis revealed varying patterns of behaviour across different conditions. Clusters 0 (Homeos1), 3 (Ribo.DAM1) and 8 (Homeos3) are specifically depleted in MS, AD and COVID-19, respectively; whilst cluster 6 (Lipo.DAM) is expanded in AD and MS, and cluster 7 (MAC) is expanded only in COVID-19 (Fig. 4a and b, Supplementary Fig. 8 and Supplementary Data 5). It is worth noticing the increase of border-associated macrophages exclusive to COVID-19, the only non-neurodegenerative condition, which could be explained by the aggressive and systemic acute pro-inflammatory response in these patients that deceased at intensive care units due to causes associated with severe manifestation of the pathology. To experimentally validate the expansion of cluster 6: Lipo.DAM in AD and MS, we analysed human brain sections through immunohistochemistry and RNAscope in situ hybridization. Using the pan-macrophage marker IBA1 to identify brain resident macrophages, we performed RNAscope for P2RY12 and GPNMB across 6 AD, 8 MS and 6 PD patients, and 7 control subjects (Fig. 4c, Supplementary Fig. 9a and Supplementary Table 1). Our analysis is based on the number of P2RY12 or GPNMB-positive dots within IBA1+ cells/microglia (Supplementary Fig. 9b and f). Ten images were taken per sample, and we included a total of 1784 cells. We first evaluated the mean number of P2RY12 and GPNMB dots per cell, which were similar across the studied groups, although there was a slight tendency towards an increase in GPNMB+ dots in AD (Supplementary Fig. 9c and g). In addition, the fraction of P2RY12+ and GPNMB+ cells (number of dots higher than zero) also did not show significant differences (Supplementary Fig. 9d and h). We consider that our Homeos1 and Lipo.DAM populations are not merely positive for the respective markers but are instead enriched in them. To explore this, we focused our analysis on cells with a clear phenotype, defining P2RY12- and GPNMB-high cells based on the real proportions of Homeos1 (28%) and Lipo.DAM (2.88%) in the HuMicA. Thus, P2RY12- and GPNMB-high cells contain a number of dots equal or higher than 3 (72nd percentile) and 15 (97.5th percentile), respectively (Supplementary Fig. 9b and f). We found a significant increase for GPNMB-high cells in AD (Fig. 4d), while no differences in the fraction of P2RY12-high cells were observed (Fig. 4e). Looking at these fractions per sample/tissue section instead of group, it is clear that GPNMB-high cells are increased in AD and suggests a similar tendency in the other pathologies (Supplementary Fig. 9e and i). We also performed a morphometric analysis of IBA1+ cells, reflected in the number and the mean length of branches/ramifications, which were similar between all groups when considering all identified cells (Supplementary Fig. 9j and k). In addition, we observed no differences in the number of ramifications between P2RY12-high and P2RY12-low cells; whereas, GPNMB-high cells showed lower number of ramifications compared to GPNMB-low cells (Fig. 4f). This indicates that GPNMB-high cells go through a structural change towards an activated morphology. The images presented in Fig. 4c support these conclusions from a qualitative standpoint. The examples in AD samples show the colocalization of GPNMB-high cells with potential amyloid senile plaques; and P2RY12-high cells show higher number of ramifications while GPNMB-high appear to be less ramified and more rounded (Fig. 4c). These findings from in situ hybridization corroborate our HuMicA analysis and indicate the expansion of lipid metabolism-related microglia, which are structurally distinct from homeostatic cells, specifically in AD, and suggest a tendency towards an increase in other neuropathologies.
a Differential cluster composition in each group in relation to “No Neuropathology” individuals using sccomp represented by the mean of the posterior distribution for the composition parameter, and the 2.5 and 97.5% quantiles. (FDR < 0.05 is highlighted in red). b Proportion of the clusters showing altered distribution (Cluster 0: Homeos1; Cluster 3: Ribo.DAM1; Cluster 6: Lipo.DAM; Cluster 7: MAC), individualized by sample, per group. The blue boxes represent the posterior predictive check and the red triangles outliers (*FDR < 0.05). In panels (a, b) the groups include 60 Alzheimer’s Disease (AD), 7 autism spectrum disorder (ASD), 15 COVID-19, 17 epilepsy, 23 Lewy Body Diseases (LBD), 10 multiple sclerosis (MS) and 109 control samples. c Maximum projection of an immunostaining of the pan-macrophage marker IBA1 in green combined with RNAscope in situ hybridization for P2RY12 (red) and GPNMB (magenta), counterstained with DAPI. Images correspond to human postmortem temporal cortex tissue sections from controls (n = 7), AD (n = 6), MS (n = 8) and Parkinson’s disease (PD) (n = 6). The yellow arrows indicate the blue autofluorescence of amyloid senile plaques in AD samples. The red and magenta arrows indicate P2RY12-high and GPNMB-high cells, respectively. Scale 15 µm. d Fraction of IBA1 + GPNMB-high cells, considered for a number of GPNMB dots equal or superior to 15. Each dot represents a donor sample. e Fraction of IBA1 + P2RY12-high cells, considered for a number of P2RY12 dots equal or superior to 3. Each dot represents a donor sample. The significance in panels (d, e) was calculated using the non-parametric Kruskal-Wallis test followed by the Dunn’s test [*adjusted p (Bonferroni) < 0.05]. f Comparison of the mean number of branches between P2RY12-high (n = 509) and P2RY12-low IBA1+ cells (n = 1275) and between GPNMB-high (n = 49) and GPNMB-low IBA1+ cells (n = 1735). Each dot represents a cell. The significance was calculated using the non-parametric Mann-Whitney test (*p < 0.05). For box plots, the hinges correspond to the first and third quartiles, the whiskers denote minimum and maximum values (excluding outliers) and the horizontal line to the median. All tests used were two-sided. Source data are provided as a Source Data file.
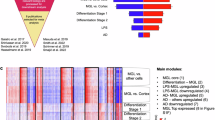
Pathology-associated transcriptomic changes in microglia were also investigated through the calculation of pseudobulk differentially expressed genes (DEGs) in pathology vs the healthy population using the complete HuMicA object, and each of the four main subpopulations; namely, Homeos (clusters 0, 4 and 8), DAMs (clusters 1, 3, 5, 6), DIMs (cluster 2), and MAC (cluster 7) (Supplementary Data 6). The total number of upregulated DEGs varied between pathologies, and was overall higher when calculated using the HuMicA object (Fig. 5a). This can relate to the positive association between the number of cells and the number of DEGs in pseudobulk analysis described elsewhere47 and should also be considered when interpreting the lower number of DEGs for the smaller DIM and MAC subpopulations. By performing a Fisher’s exact test for the overlap between the multiple lists of DEGs, separated by pathology and cell population, we demonstrated a high level of similarity between pathologies (Fig. 5b). A consistent overlap was equally observed between the analysed cellular subsets within each pathology (Supplementary Fig. 9l). These results suggest consensual pathology-associated pattern of differential gene expression manifested non-specifically in neuropathology. However, the potential bias introduced by using the same control population in all comparisons should be acknowledged. In addition, we performed GO enrichment of the lists of upregulated pseudo-bulk DEGs in each pathology vs controls considering the full HuMicA object, the Homeos (clusters 0, 4 and 8), DAMs (clusters 1, 3, 5, 6), and DIMs (cluster 2) populations (Fig. 5c, Supplementary Data 7). For a group of GO terms, we observe lack of pathology specificity, being commonly enriched across different diseases. From those we highlight pathways involved in cellular cytoskeleton and intracellular transportation (actin binding, dynein intermediate chain binding, minus−end−directed microtubule motor activity), extracellular environment (collagen-containing extracellular matrix). Contrarily, some terms demonstrate a pathology-specific enrichment, such as ones related to ATP synthesis (proton−transporting ATP synthase complex, ATPase complex), and lipoproteins (triglyceride−rich plasma lipoprotein particle) in LBD; mechanisms associated with ribosome (structural constituent of ribosome, ribosome assembly), myelin (myelination, myelin sheath), antigen presentation (MHC protein complex) and migration (leukocyte migration) in MS; immune response terms (cytokine receptor activity, immune receptor activity) in COVID-19; and WASH complex (Wiskott Aldrich Syndrome protein and scar homologue complex), involved in endosome and actin filament regulation, in epilepsy (Fig. 5c). In essence, these results demonstrate that there are pathology-related mechanisms that are shared between pathologies, whereas others may be particular to a pathology or a small group of pathologies. However, it is paramount to acknowledge the potential of statistical artifacts in this pseudobulk analysis, which by its own nature is more prone to batch effects in a heterogeneous integrated single-cell object like the HuMicA.
a Barplot representation of the number of pseudobulk DEGs calculated for each pathology in comparison to the control population. Upregulated DEGs were calculated using the complete HuMicA object and subsets consisting of Homeos subpopulations (clusters 0, 1, 4), DAMs (clusters 1, 3, 5, 6) and DIMs (cluster 2). Significant DEGs are considered for an adjusted p value (FDR) inferior to 0.05 and log2FoldChange superior to 1. b Heatmap representation of the significance of the overlap between the lists of pseudobulk DEGS, calculated with the GeneOverlap function. Overlaps were calculated between each pathology within each subset (HuMicA, Homeos, DAMs, DIMS). The degree of significance is depicted by the Jaccard index and a p-value inferior to 0.05 is highlighted by a white asterisk. c Dot plot representation of the gene ontology (GO) enrichment of all lists of pseudobulk DEGs between each pathology and controls accounting for the complete HuMicA object and subsets consisting of Homeos subpopulations (clusters 0, 1, 4), DAMs (clusters 1, 3, 5, 6), DIMs (cluster 2) and MAC (cluster 7). Selected statistically significant GO terms are represented (adjusted p-value (Benjamini-Horchberg) < 0.05). Significance is represented by the log of the fold change and the negative of the log of the adjusted p-value (FDR). Source data are provided as a Source Data file.
Discussion
Our integrated analysis of snRNA-seq and scRNA-seq datasets, comprising brain biopsies from various neurological pathologies and healthy subjects, has resulted in the creation of a comprehensive multi-pathology human atlas of microglia called HuMicA, which includes 90,716 single cells. The clustering analysis of the integrated object confirmed that microglia encompass a range of distinct cell subpopulations rather than a one-directional transition from homeostasis to activation. In mice, several studies have already developed atlases covering the spectrum of microglia throughout embryonic development, aging, and in pathological conditions33,48,49. The translation of these findings into human biology has become feasible due to the availability of snRNA-seq studies, which have significantly contributed to the more precise characterization of the human cerebral environment, by identifying specific cellular subpopulations. However, microglia often take on secondary attention in human studies primarily due to their low abundance within bulk brain samples. In our analysis, the average proportion of the immune cell population relative to the entire dataset was approximately 5%, which aligns with the reported amounts in the original studies12,14. This limitation has previously impacted the scalability of the studies, that require larger number of samples, and hinders the identification of smaller microglia subpopulations, which remain underrepresented in smaller datasets. In this study, we have successfully overcome this significant limitation by employing a bioinformatic strategy that maximizes the utilization of already published data.
The HuMicA comprises nine subpopulations, eight microglial and one border-associated macrophages, with a substantial number of cells per cluster, allowing for detailed characterization. For instance, we successfully described the classical homeostatic microglia phenotype, which accounted for the most abundant population, Homeos1, that shared a strong resemblance with the main homeostatic microglia clusters from previous literature10,35,36. In addition, we identified two additional populations with a homeostatic gene profile, one of which, Homeos3, was smaller in size and potentially represented a pre-activation state considering the expression of inflammatory chemotaxis-related genes. Special attention must also be given to Homeos2, which expresses high levels of P2RY12 and CX3CR1, and we annotated as “homeostatic”. Indeed, Homeos2 presents a distinct profile specifically driven by high expression of GRID2, PRDM11 and ADGRB3, which segregates it clearly from the other subpopulations. Interestingly, a similar microglia population, denominated AD2 was identified in an external snRNA-seq study. This AD2 subpopulation was found to be expanded in AD compared to controls and was characterized by upregulation of P2RY12, CX3CR1, GRID2 and ADGRB3. Additionally, this gene expression pattern was also associated with synapse assembly and specifically related to a response to tau-bearing dying neurons35. Moreover, in our clustree analysis, the group of cells corresponding to Homeos2 (or cluster 4), segregated from the main microglia cluster at very low resolutions. This raises questions regarding the true nature of Homeos2, which still requires further studies to elucidate whether it behaves as a homeostatic subpopulation or represents a distinct pathology-related state, and even if it follows the standard microglial ontogeny.
Previous studies had already demonstrated an association between microglia activation and neuropathology. In these, the typical strategy often involves assessing differential gene expression in microglia from patients compared to healthy controls10,11,12,13,14,15,17,19,22,23, with a general description of a pathology-related pro-inflammatory and activated state. In addition, some studies have shown a differential distribution of microglia subpopulations between patients and controls, with the transcriptomic profile of these subpopulations enriched in genes associated with activation. This includes microglia subpopulation resembling the DAM phenotype observed in mice, as well as those expressing genes associated with genetic risk factors for neurological diseases10,11,14,18. The DAM phenotype has indeed been a valuable tool in evaluating the role of microglia in neurodegeneration. It has been shown to be prevalent not only in AD mice, but also in a model of amyotrophic lateral sclerosis8. Furthermore, the aforementioned DAM enrichment in pathology-related microglia extend beyond AD10,11,14, encompassing other conditions such as autism18 and severe COVID-1922, emphasizing the inter-disease and unspecific nature of DAM. Indeed, the role of the DAM phenotype in neuropathology, particularly in human AD, has been a topic of debate, and its replicability has been questioned15. Moreover, from the methodological standpoint, it has been shown that snRNA-seq microglia express lower levels of main DAM genes in comparison to scRNA-seq microglia44. However, in the case of the HuMicA, we used both single-cell and single-nucleus data and were still able to widely demonstrate detectable and varying levels of DAM genes across the microglia populations of our datasets.
It is reasonable to conclude that the DAM phenotype alone may not be sufficient to fully understand microglia’s behaviour in pathology. The original study proposed a sequential two-stage process for DAM activation, with the first stage being Trem2-dependent stage 1 and the second stage being Trem2-independent stage 28. However, a recent ground-breaking study by Silvin and colleagues (2022) provided further insights “deconstructing” the DAM signature. These authors developed an encompassing myeloid cell map of mice brain samples by integrating scRNA-seq data from different neuropathological settings. Within this “M-verse”, they observed two DAM populations with different ontogenies and concluded that one of these populations is composed by infiltrating monocyte-derive “disease-inflammatory macrophages” (DIMs). The authors demonstrated that bona fide DAMs were Trem2-dependent, while DIMs were Trem2-independent33. However, the authors did not elaborate on whether the DIMs could represent what was initially described as the stage 1 DAM based on the dual profile of Trem2-dependency. The experimental design used in our study did not allow to infer on TREM2 dependency, as TREM2 expression was observed consistently across all populations. We describe the prevalence of a significant number of cells carrying the DIM signature across all studied conditions, both patients and controls, which is indicative of the potential relevance of this cell group not only in response to injury and pathology, but also in physiologic settings. Our results regarding the DAM signature move away from the simplified on-and-off or two-way activation route. In this regard, we conclude that the general DAM phenotype, initially described in mice, is detectable in human microglia, but insufficient to represent specific cell states. Here, we identify four populations, in addition to the DIMs, that derive from the original DAM signature but are distinct from each other in their own right. These included cells with clear pro-inflammatory nature (Inflam.DAM) upregulating genes associated with antigen presentation and phagocytosis. Two other distinct clusters shared a similar profile associated with the ribosome complex, a pattern that requires further investigation to understand the role of these cells in the human brain. Although both these populations show striking enrichment for ribosome-related pathways, each present distinct traits and have clustered separately in our pipeline. We could speculate that these represent two biologically distinct populations or are a result of statistical bias from unbalanced tissue region representation. At last, we describe a microglia population expressing genes associated with the lysosome, fatty acid beta-oxidation, lipid metabolism and lipoproteins. All these reflect signatures had been described elsewhere, validating the strength of this study. Furthermore, the transcription factor (TF) analysis outlines MYC as a major driver of microglia activation related to the ribosome and inflammatory profiles. Peroxisome proliferator-activated receptor alpha (PPARA) was the sole activated TF in Lipo.DAM. PPARA is an important modulator of lipid oxidation, and its agonists have been shown to inhibit microglia activation50. In addition, in mice, the microglia-specific knockout of PPARA lead to decreased retinal thickness and overexpression alleviated diabetes-induced retinal degeneration and dysfunction51, raising the question of the potential neuroprotective nature of this population. The DIMs, on the other hand, presented a completely different TF enrichment, associated with standard pro-inflammatory factors like NFkB, STAT1, HIF1A and MITF, which can be considered an additional indication of the different ontogeny between these cells and bona-fide microglia. An important aspect used to interpret distinct microglia subpopulations is the differential expansion or depletion in different pathological conditions. A depletion of homeostatic microglia in pathology would be a reasonable assumption; however, in this study we were unable to observe a consistent decrease of the proportions of the homeostatic subpopulations in the different pathologies, the same way others also failed to describe such trend36. The only exception was the depletion of Homeos1, the main homeostatic population, in MS. Additional studies, with more well-balanced data across pathologies and tissue-regions, will be required to fully access if this MS-specific depletion is of pathological relevance, a result of statistical artifact or related to the tissue region, since the white matter samples used here come exclusively from a study developed in MS (Jakel 2019)16. In regard to the subpopulations we considered under the DAM-verse, these have consistently been shown to be expanded in pathology, particularly in AD8,10,33, whilst there is no clear understanding on the differential distribution of DIMs in human pathology primarily due to the very recent description this cell type33. The most evident pattern of differential distribution in pathology was the expansion of Lipo.DAM in both AD and MS, which was only validated by in situ RNAscope hybridization in postmortem temporal cortex sections for AD. We may conclude that this population may present a particularly relevant role in AD, eventually associated with response to Aβ plaques. However, we cannot completely exclude a potential expansion in other diseases, since AD was the predominant pathology within the HuMicA. Moreover, the lack of validation of the Lipo.DAM expansion in other pathologies may be associated with the mismatch between the evaluated regions in the in situ hybridization and the single-cell object, or the lack of representability of tissue regions with higher degree of association with the pathophysiology of the studied diseases. This expansion had already been described in AD36. Our results suggest that this is not exclusive to AD and may rather represent a general pathology-associated mechanism. Despite only demonstrating it in AD and MS, as abovementioned, a more homogeneous and well-balanced study design would help elucidate if such expansion is observed in more pathologies. Contrarily to what is considered to other CNS cell types, particularly excitatory and inhibitory neuron subtypes, microglia are commonly considered to behave in a more general manner, nonspecifically from the disease, given its role in neuroinflammation, a phenomenon common to all neuropathologies. The HuMicA supports the complexity of microglia in relation to the heterogeneity of existent subpopulations, showing that microglia exist under the form of transcriptionally distinct groups of cells, some of which may be in fact derived from peripheral monocytes, contrarily to what was formerly considered. However, we did not observe clear tendencies characteristic of a single pathology. The expansion of border-associated macrophages only in COVID-19 patients can be explained by the acute nature of the pathology, distinct from the legitimate neurodegenerative diseases. The pseudobulk differential expression analysis further validates the claim of a nonspecific behavior, considering the high degree of overlap of the DEGs between pathologies. Within our data, there is variability in number of cells per sample, and the number of samples per study, being important to address the limitations of pseudobulk analysis in unbalanced datasets and the need for subsequent studies.
An additional microglial population has been described in the literature, one that showed detectable levels of oligodendrocyte markers, such as ST18, PLP1, and MBP. In that study, the authors incubated human primary microglia with myelin particles and observed the localization of oligodendrocyte-derived mRNA within the phagocyting microglia and close to the nuclei17. Similar patterns of phagocytic microglia engulfing oligodendrocyte-derived mRNA were described in mice scRNA-seq data49. In the preliminary stages of the development of the HuMicA, we observed this pattern of high levels of myelin-related genes within a fraction of the microglia cells. Given the significant size of this cluster, it remained prevalent even after quality control filtering and doublet removal and was only excluded in the clustree preprocessing. We opted for a more conservative approach and ultimately excluded these cells from the final integrated object. Despite the existence of myelin-engulfing phagocytic microglia not being fully understood, we believe that it is relevant to shed light on them and incentivize further research on the matter.
Using the in silico approach in our study, we successfully compensated for the common limitation of low sample size in individual human microglia scRNA-seq and snRNA-seq datasets, which have significantly prevented the ability to fully characterize microglial subpopulations. We have described and validated the HuMicA, in which we describe multiple homeostatic microglia states as well as pathology-related phenotypes. We intend that the HuMicA is used as a public resource to study microglia under multiple experimental settings thus contributing for the evolution of the overall knowledge on microglial biology.
Methods
Patients and ethical approval
The human brain tissue data used in this study was obtained from 19 public snRNA-seq and scRNA-seq datasets (Table 1). The original studies were responsible for the ethical approval for sample collection. For the in situ RNAscope validation, samples were obtained from the Pathology Department and the Vall d’Hebron Neurological Tissue Bank (PR(AG)200/2013) at the Vall d’Hebron University Hospital (Barcelona, Spain). Samples and data from patients included in this study were provided by the Vall d’Hebron University Hospital Biobank (PT20/00107), integrated in the Spanish National Biobanks Network, and the Biobanco de Tejidos CIEN, and they were processed following standard operating procedures with the appropriate approval of the Ethical and Scientific Committees.
Single nucleus (sn) and single cell (sc) RNA-seq datasets
Detailed technical and clinicodemographic characterization of the used samples from the snRNA-seq and scRNA-seq datasets is presented in Supplementary Data 1. We included six datasets for AD (Mathys, Grubman, Lau, Leng, Morabito, Zhou)10,11,12,13,14,15, two for MS (Jakel, Schirmer)16,17, three for epilepsy (Mancuso, Pappalardo, Thrupp)44,52,53, one with both AD and epilepsy patients (Olah)34, two for COVID-19 (Yang, Fullard)22,23, two for LBD (including Parkinson’s, Parkinson’s with dementia and dementia with Lewy bodies) (Feleke, Smajic)19,54 and one for ASD (Velmeshev)18. The samples consisted mostly of postmortem, but also tissue collected during epileptic resective surgery. The control groups within each study were included in the “No Neuropathology” group, plus data extracted from two additional datasets developed exclusively with individuals with no neuropathological alterations (Tran, Franjic)28,55 After pre-processing, 241 individual samples contributed to the atlas.
Dataset preprocessing
A dataset downloaded as raw fastq files (Jakel) was processed using the CellRanger v.6.1.2 software56. Individual count data was obtained with the count function, aligning to the reference genome GRCh38-3.0.0. The --include-introns option was used to map both unspliced pre-mRNA and mature RNA. Individual count matrices were generated for each library and concatenated into a single count matrix.
Most datasets were obtained already in the count matrix format, either at the raw or preprocessed stages. Data had been already demultiplexed and sequences aligned to the human reference genome (GRCh38), accounting for both intronic and exonic regions in snRNA-seq datasets. For each dataset, count data from each subject was concatenated in one single count matrix. Data processing and subsequent analysis was performed using the Seurat (v4.0.2) R package57. Quality control filtering was uniformized to all datasets. Genes were only included if detected in at least 3 nuclei. For the snRNA-seq datasets, nuclei with unique genes inferior to 200 or superior to 5000, total UMI counts less than 500 or over 20,000, mitochondrial RNA content superior to 20% or ribosomal RNA content superior to 5% were filtered out. The same filtering strategy was used for scRNA-seq data, except for the exclusion based on the ribosomal RNA content. Furthermore, a list of 105 genes described as related to postmortem interval in cerebral cortex was excluded58. The doubletFinder_v3 function59 was used to estimate doublets from each individual subject, which were removed from each dataset’s Seurat object, followed by SCTransform normalization60 with default 3000 variable genes. Dimensionality reduction was performed with Principal Component Analysis (PCA) followed by Uniform Manifold Approximation and Projection (UMAP) accounting for the 30 main principal components (PCs). After preprocessing, the distribution of number of unique genes, UMI counts, percentage of mitochondrial RNA was considered homogeneous between all dataset sources.
Cell-type annotation and isolation of immune cells
Cell clustering was performed with FindNeighbors (30 PCs) and FindClusters (res = 0.05-0.08) functions. The low resolution used allowed for a broad characterization of the 6 main cell types in brain tissue: neurons, oligodendrocytes, OPCs, astrocytes, endothelial cells, and immune cells (Supplementary Fig. 1 and Supplementary Fig. 2). The immune cell clusters encompassed microglia and other infiltrating or resident immune cells, such as T cells and macrophages. Cell type annotation was based on the identification of known canonical gene obtained using FindAllMarkers (min.pct = 0.1 and logfc.threshold = 0.25, test.use = “wilcox”). CD74, DOCK8, APBB1IP, HLA-DRA, PTPRC, P2RY12, C1QB, CX3CR1, C3, CSF1R and AIF1 were considered for identification of the immune cell clusters. The annotated clusters of immune cells did not present any of the known canonical genes for other brain cell types among its markers. Individual subjects with markedly low nuclei/cell number in relation to the remaining dataset were removed before integration. As a side note, we must refer that the “RM102” sample from the Mancuso et al. (2019) study53 was excluded before integration, since preliminary tests indicated that it behaved as an outlier and clustered isolated from all other samples.
Integration, clustering and filtering of the Human Microglia Atlas
The immune cells from all datasets were integrated using the Seurat integration pipeline61 adapted for datasets normalized with SCTransform. The integration and all analyses posterior to it were performed using Seurat v562. The FindIntegrationAnchors and IntegrateData functions considered the 20 main PCs and the 3000 most variable features across all datasets. The final integrated Seurat object consisted of a total of 102,390 cells/nuclei. Dimensionality reduction was performed using PCA and UMAP (50 PCs). Clustering was performed using FindNeighbors (50 PCs) and FindClusters. An empiric approach was used to infer on the optimal resolution using the clustree package27. Firstly, a clustree was constructed with a resolution range between 0.01 and 0.075 (Supplementary Fig. 3a). At a 0.025 resolution two clusters separate from the main myeloid population and, based on known canonical cell type markers, those correspond to T cells (CD247, TRAC) and a cluster of cells expressing markers from other CNS cell types (SNAP25, SYT1, PLP1, ST18, SLC1A2) which we deemed doublets (Supplementary Fig. 3b and c). To obtain the HuMicA, cluster 0 at 0.025 resolution, corresponding to myeloid cells, was isolated. Furthermore, from this new set of cells, we excluded the cells/nuclei from all individual samples with a cell count inferior to 50, with a final number of individual samples of 241. The final HuMicA object consisted of 90,716 nuclei/cells. Dimensionality reduction and reclustering was performed in the new object with same abovementioned settings, and a clustree was developed with a resolution range of 0.01-1 to assess the optimal clustering resolution. In a clustree a high degree of cell exchange between nodes is indicative of overclustering. We considered res = 0.2 to represent the optimal point in terms of the balance in cluster size and low rate of cell exchange (Supplementary Fig. 3d).
Evaluation of differential cluster distribution
We accessed the proportions of distribution of each HuMicA cluster across the main studied variable (“Group”), which relates to the pathological setting, but also in relation to other variables of interest, namely “Methodology” (scRNA-seq and snRNA-seq) (Supplementary Fig. 4a–e), “Sex” (Supplementary Fig. 4f–j), “Tissue state” (postmortem and surgically resected) (Supplementary Fig. 4k–o), “Age” (Supplementary Fig. 5a–e) and “Tissue region” (Supplementary Fig. 5f–j). Of note, the information on age was converted into a categoric variable representing age intervals. The Fullard dataset does not supply sex nor age information; thus, the age of these individuals was annotated as “NA”, and sex was computed based on the expression of chromosome Y-specific genes (ZFY, UTY and RPS4Y1). Moreover, tissue regions were grouped into more encompassing categories based on anatomic distribution and function, to compensate for high heterogeneity of data distribution (“Frontal”: medial frontal cortex, motor-premotor cortex, prefrontal cortex, superior frontal gyrus; “Temporal”: middle temporal gyrus, temporal neocortex, temporo-parietal cortex; “Limbic”: amygdala, anterior cingulate cortex, dentate gyrus, entorhinal cortex, hippocampus, nucleus accumbens and subiculum; “Midbrain”; “White matter”).
The sccomp package46 was used to infer the significance of the differential distribution of each cluster in relation to each of the variables. The sccomp_glm function calculates the differential composition using a Bayesian method based on sum-constrained independent Beta-binomial distributions. The results are presented by of a credible interval of the slope (95% confidence) and FDR (calculated using an Empirical Bayes approach63). The hypothesis testing calculates the posterior probability of the composition being larger than an established fold-change threshold (default = −0.2-0.2). A credible interval of the slope (95% confidence) higher or lower than zero represents expansion or depletion, respectively, of a cluster for a determined category. A significant differential composition is considered for FDR < 0.05. For the “Group” variable, the “contrasts” option was used to calculate the differential composition of each pathology group in relation to the “No Neuropathology” group. For the remaining variables, the differential composition was calculated in comparison to the largest category (“Methodology” - “Nuclei”; “Sex” - “Male”; “Tissue state” - “Postmortem”; “Age” – “> 80”; “Tissue region” – “Frontal”).
Inference of covariate influence
The integrated Seurat object was converted using the as.SingleCellExperiment function. The percentage of variance explained, considering the genes in the “integrated” assay, was calculated using the plotExplanatoryVariables function from the scater package64 for the variables accounting for individual sample (“Sample_ID”, origin dataset (“Study”), pathological setting “Group”, type of dataset (“Methodology”, scRNA-seq or snRNA-seq), the brain region (“Tissue region (specific)” for the original annotation; “Tissue region (broad)” for the larger grouping), nature of the sample (“Tissue state”, postmortem or surgically ressected), “Sex” and “Age” (categorized by year intervals) variables.
Cluster characterization and calculation of cluster markers
For the calculation of cluster markers, the “RNA” assay from the integrated object was normalized using the NormalizeData function with standard settings. Markers were calculate with the FindAllMarkers function (min.pct = 0.25 and logfc.threshold = 0.25, only.pos = TRUE, test.use = MAST), and were considered significant if FDR < 0.05. For the visualization of the expression of specific genes or sets of genes, we used the normalized counts of the “RNA” assay, as abovementioned. The average expression of sets of multiple genes was obtained with AddModuleScore. Representation of gene expression in the UMAPs was accomplished with the FeaturePlot (Seurat) function. The average expression by cluster (with the AverageExpression function from Seurat) of the normalized “RNA” assay counts was used to plot the heatmap of cluster markers, with the ComplexHeatmap function65; and to plot the cluster dendrogram accounting for the list of unique cluster markers (Supplementary Data 2). For the dendrogram, the “spearman” and “ward.D” methods were considered for distance and clustering, respectively.
Pseudobulk differentially expressed genes
Differentially expressed genes (DEGs) between pathologies were calculated using a pseudobulk strategy. The raw counts of the “RNA” assay were summed for all cells/nuclei in each individual sample using the AggregateExpression function from Seurat, which was done separately for the complete HuMicA object and for each subset of HuMicA subpopulations (Homeos: clusters 1, 4 and 8; DAM: clusters 1, 3, 5, 6; DIM: cluster 2; MAC: cluster 7). Beforehand, lowly expressed genes were excluded and only genes with more than one count in at least 10 cells were considered. The aggregated count values were adjusted for batch effect of the variable “Methodology” (single-cell or single-nucleus), shown to elicit significant confounding effect, using the Combat_Seq function of the sva package66. The DESeq2 package67 was used for calculation of DEGs using the adjusted count matrixes. No covariate was used in the negative binomial generalized linear model, and the DEGs were calculated for each pathology group in comparison to the healthy population. Significant DEGs were considered for FDR < 0.05 and log2FoldChange > 1, thus considering only upregulated genes. The significance of the overlap between lists of DEGs was performed using the GeneOverlap package68, and the number genes that passed the quality control abovementioned in this section was used as background. These values were 17,943, 14,903, 15,209, 13,113, and 7965 for the DEGs calculated for the full HuMicA object, Homeos (clusters 0,4 8), DAMs (clusters 1,4 8), DIMs (cluster 2) and MAC (cluster 7, macrophages). For the overlaps between HuMicA subsets within each pathological setting, the 17,943 genes from the full object were used (related to Supplementary Fig. 9l). The enrichment is represented by the Jaccard index, which varies between 0 and 1 and represents the similarity between two lists.
Enrichment analysis
Gene ontology (GO) enrichment was obtained using the enrichGO function from the clusterProfiler package69. The simplify function from enrichGO was used to remove redundancy of the most significantly enriched terms. The fold change of GO enrichment was calculated with the equation (gene_pos/gene_total)/(bg_pos/bg_total), which represents the ratio between the positive hits in the input gene list and the total number of genes in that list and the positive hits and full number of genes in the background. The viper70 package was used to assess TF activity, accounting for the TF-target information from the CollecTRI37 repository from the decoupleR package71. The fgsea package72 was used to perform GSEA from differential expression comparisons. For both TF enrichment and GSEA, the results from FindAllMarkers (min.pct = 0.1 and logfc.threshold = 0, test.use = MAST) were used as input, and the avg_log2FC and the p_val_adj settings were used for ranking. This less significant cutoff intends to account for the broad spectrum of differential expression and not only the significant differentially expressed markers.
RNAscope
Human postmortem temporal cortex paraffin sections (5 µm) on Superfrost Plus Slides (Fisherbrand; Cat#12-550-15) were processed following the recommended protocol by ACD Biotechne RNAscope multiplex fluorescent reagent v2 assay (#323285). RNAscope probes were as follows: Hs-P2RY12-C2 (ACD Biotechne 450391-C2) and Hs-GPNMB-C1 (ACD Biotechne 413521-C1). Primary and secondary antibodies for IBA1 were rabbit recombinant monoclonal (Abcam #ab178846, 1:500) and goat anti-rabbit Alexa Fluor 488 (Thermo Scientific # A-11008, 1:1000), respectively. To quench autofluorescence, we applied TrueBlack (Biotium; Cat#23007) for 90 s after the final step (DAPI application) of the RNAscope assay procedure, and gently rinsed with PBS prior to coverslipping and imaging. Maximum projection images (z-stack with a 0.20 µm interval) were taken in a confocal microscope (LSM 980; Zeiss) with AiryScan detection and an 40x oil immersion objective. For probe quantification, 10 images were taken throughout the tissue section by an investigator blind to the experimental group. Fiji (ImageJ 1.54 f) was used to identify IBA1-positive cells and quantify the number of P2RY12 and GPNMB spots in each IBA1-positive cell of each image. Dots were only counted if a cell had both IBA1 staining and a nucleus, stained by DAPI. We included a total of 1784 IBA1 + /DAPI+ cells with a mean of 66 and a range of 13–139 per tissue sample.
Statistics and Reproducibility
The statistical analyses of sc and snRNA-seq data were conducted using R language. No statistical method was used to predetermine sample size. Samples were excluded based on the aforementioned quality control pipeline.
The in situ hybridization data generated in this study was analysed using the Fiji software. No statistical method was used to predetermine sample size, and the experiments were not randomized. Confocal microscopy images were blindly taken throughout the tissue section by an independent researcher.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The integrated HuMicA object is available in the interactive Shiny-based web interface (https://in7yqx-ricardo-ferreira.shinyapps.io/shinyapp/), generated using ShinyCell73. Source data are provided with this paper. The raw RNAscope in situ hybridization data generated in this study are provided in the Source Data Fig. 4 file. The accession codes and/or links for the original snRNA-seq and scRNA-seq datasets used to generate the HuMicA are as follows: Mathys, syn1848517 (https://www.synapse.org/#!Synapse:syn18485175); Grubman, GSE138852; Leng, GSE147528; Morabito, GSE174367; Lau, GSE157827; Zhou, syn21125841 (https://www.synapse.org/Synapse:syn21125841/wiki/597278); Olah, syn21438358 (https://www.synapse.org/#!Synapse:syn21438358); Pappalardo, GSE157277; Thrupp, GSE153807; Mancuso, GSE137444; Jakel, GSE118257; Schirmer, https://cells.ucsc.edu/?ds=ms; Velmeshev, https://autism.cells.ucsc.edu/; Feleke, GSE178146; Smajic, GSE157783; Tran, Globus endpoint ‘jhpce#tran2021‘ (https://app.globus.org/file-manager?origin_id=74570c7d-98ff-4e10-ba5b-c735d209cf96&origin_path=%2F); Franjic, GSE186538; Yang, GSE159812; Fullard, GSE164485. Source data are provided with this paper.
Code availability
The code used in this study is available at https://github.com/RicardoMartins-Ferreira/HuMicA (doi.org/10.5281/zenodo.14539369) and https://ricardomartins-ferreira.github.io/HuMicA/, including the scripts for quantification developed by the Microscopy Platform at the High Technology Unit from the Vall d’Hebron Research Institute (UAT-VHIR).
References
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Ginhoux, F., Lim, S., Hoeffel, G., Low, D. & Huber, T. Origin and differentiation of microglia. Front Cell Neurosci. 7, 45 (2013).
Martins-Ferreira, R., Leal, B., Costa, P. P. E. & Ballestar, E. Microglial Innate Memory and Epigenetic Reprogramming in Neurological Disorders. Prog. Neurobiol. 101971. https://doi.org/10.1016/j.pneurobio.2020.101971 (2020).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Woodburn, S. C., Bollinger, J. L. & Wohleb, E. S. The semantics of microglia activation: neuroinflammation, homeostasis, and stress. J. Neuroinflammation 18, 258 (2021).
Subhramanyam, C. S., Wang, C., Hu, Q. & Dheen, S. T. Microglia-mediated neuroinflammation in neurodegenerative diseases. Semin Cell Dev. Biol. 94, 112–120 (2019).
Chiu, I. M. et al. A neurodegeneration-specific gene-expression signature of acutely isolated microglia from an amyotrophic lateral sclerosis mouse model. Cell Rep. 4, 385–401 (2013).
Keren-Shaul, H. et al. A Unique Microglia Type Associated with Restricting Development of Alzheimer’s Disease. Cell 169, 1276–1290.e17 (2017).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Grubman, A. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat. Neurosci. 22, 2087–2097 (2019).
Lau, S.-F., Cao, H., Fu, A. K. Y. & Ip, N. Y. Single-nucleus transcriptome analysis reveals dysregulation of angiogenic endothelial cells and neuroprotective glia in Alzheimer’s disease. Proc. Natl Acad. Sci. USA 117, 25800–25809 (2020).
Leng, K. et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat. Neurosci. 24, 276–287 (2021).
Morabito, S. et al. Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer’s disease. Nat. Genet 53, 1143–1155 (2021).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat. Med 26, 131–142 (2020).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Feleke, R. et al. Cross-platform transcriptional profiling identifies common and distinct molecular pathologies in Lewy body diseases. Acta Neuropathol. 142, 449–474 (2021).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Wright-Jin, E. C. & Gutmann, D. H. Microglia as Dynamic Cellular Mediators of Brain Function. Trends Mol. Med 25, 967–979 (2019).
Yang, A. C. et al. Dysregulation of brain and choroid plexus cell types in severe COVID-19. Nature 595, 565–571 (2021).
Fullard, J. F. et al. Single-nucleus transcriptome analysis of human brain immune response in patients with severe COVID-19. Genome Med 13, 118 (2021).
Lopez-Leon, S. et al. More than 50 long-term effects of COVID-19: a systematic review and meta-analysis. Sci. Rep. 11, 16144 (2021).
Huang, L. et al. 1-year outcomes in hospital survivors with COVID-19: a longitudinal cohort study. Lancet 398, 747–758 (2021).
Douaud, G. et al. SARS-CoV-2 is associated with changes in brain structure in UK Biobank. Nature. https://doi.org/10.1038/s41586-022-04569-5 (2022).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience 7, 1–9 (2018).
Tran, M. N. et al. Single-nucleus transcriptome analysis reveals cell-type-specific molecular signatures across reward circuitry in the human brain. Neuron 109, 3088–3103.e5 (2021).
Mrdjen, D. et al. High-Dimensional Single-Cell Mapping of Central Nervous System Immune Cells Reveals Distinct Myeloid Subsets in Health, Aging, and Disease. Immunity 48, 380–395.e6 (2018).
Bechmann, I. et al. Turnover of rat brain perivascular cells. Exp. Neurol. 168, 242–249 (2001).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Galatro, T. F. et al. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat. Neurosci. 20, 1162–1171 (2017).
Silvin, A. et al. Dual ontogeny of disease-associated microglia and disease inflammatory macrophages in aging and neurodegeneration. Immunity 55, 1448–1465.e6 (2022).
Olah, M. et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat. Commun. 11, 6129 (2020).
Gerrits, E. et al. Distinct amyloid-β and tau-associated microglia profiles in Alzheimer’s disease. Acta Neuropathol. 141, 681–696 (2021).
Sun, N. et al. Human microglial state dynamics in Alzheimer’s disease progression. Cell 186, 4386–4403.e29 (2023).
Müller-Dott, S. et al. Expanding the coverage of regulons from high-confidence prior knowledge for accurate estimation of transcription factor activities. Nucleic Acids Res 51, 10934–10949 (2023).
Tan, W. et al. Distinct phases of adult microglia proliferation: a Myc-mediated early phase and a Tnfaip3-mediated late phase. Cell Discov. 8, 34 (2022).
Miao, J., Chen, Z., Wu, Y., Hu, Q. & Ji, T. Sp1 Inhibits PGC-1α via HDAC2-Catalyzed Histone Deacetylation in Chronic Constriction Injury-Induced Neuropathic Pain. ACS Chem. Neurosci. 13, 3438–3452 (2022).
Butturini, E., Boriero, D., Carcereri de Prati, A. & Mariotto, S. STAT1 drives M1 microglia activation and neuroinflammation under hypoxia. Arch. Biochem Biophys. 669, 22–30 (2019).
Jayadev, S. et al. Transcription factor p53 influences microglial activation phenotype. Glia 59, 1402–1413 (2011).
Chen, M., Chen, X., Hu, X., Dai, J. & Sun, J. Androgen receptor contributes to microglial/macrophage activation in rats with intracranial hemorrhage by mediating the JMJD3/Botch/Notch1 axis. Neurosci. Lett. 765, 136283 (2021).
Li, Q. et al. Bcl6 modulates innate immunity by controlling macrophage activity and plays critical role in experimental autoimmune encephalomyelitis. Eur. J. Immunol. 50, 525–536 (2020).
Thrupp, N. et al. Single-Nucleus RNA-Seq Is Not Suitable for Detection of Microglial Activation Genes in Humans. Cell Rep. 32, 108189 (2020).
Dolan, M.-J. et al. Exposure of iPSC-derived human microglia to brain substrates enables the generation and manipulation of diverse transcriptional states in vitro. Nat. Immunol. 24, 1382–1390 (2023).
Mangiola, S. et al. sccomp: Robust differential composition and variability analysis for single-cell data. Proc. Natl Acad. Sci. USA 120, e2203828120 (2023).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385.e27 (2023).
Hammond, T. R. et al. Single-Cell RNA Sequencing of Microglia throughout the Mouse Lifespan and in the Injured Brain Reveals Complex Cell-State Changes. Immunity 50, 253–271.e6 (2019).
Li, Q. et al. Developmental Heterogeneity of Microglia and Brain Myeloid Cells Revealed by Deep Single-Cell RNA Sequencing. Neuron 101, 207–223.e10 (2019).
Ramanan, S., Kooshki, M., Zhao, W., Hsu, F.-C. & Robbins, M. E. PPARalpha ligands inhibit radiation-induced microglial inflammatory responses by negatively regulating NF-kappaB and AP-1 pathways. Free Radic. Biol. Med 45, 1695–1704 (2008).
Yuan, T. et al. The Protective Role of Microglial PPARα in Diabetic Retinal Neurodegeneration and Neurovascular Dysfunction. Cells 11, 3869 (2022).
Pappalardo, J. L. et al. Transcriptomic and clonal characterization of T cells in the human central nervous system. Sci. Immunol. 5, eabb8786 (2020).
Mancuso, R. et al. Stem-cell-derived human microglia transplanted in mouse brain to study human disease. Nat. Neurosci. 22, 2111–2116 (2019).
Smajić, S. et al. Single-cell sequencing of human midbrain reveals glial activation and a Parkinson-specific neuronal state. Brain 145, 964–978 (2022).
Franjic, D. et al. Transcriptomic taxonomy and neurogenic trajectories of adult human, macaque, and pig hippocampal and entorhinal cells. Neuron 110, 452–469.e14 (2022).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Zhu, Y., Wang, L., Yin, Y. & Yang, E. Systematic analysis of gene expression patterns associated with postmortem interval in human tissues. Sci. Rep. 7, 5435 (2017).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: Doublet Detection in Single-Cell RNA Sequencing Data Using Artificial Nearest Neighbors. Cell Syst. 8, 329–337.e4 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Stuart, T. et al. Comprehensive Integration of Single-. Cell Data. Cell 177, 1888–1902.e21 (2019).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. https://doi.org/10.1038/s41587-023-01767-y (2023).
Stephens, M. False discovery rates: a new deal. Biostatistics 18, 275–294 (2017).
McCarthy, D. J., Campbell, K. R., Lun, A. T. L. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Leek, J. T. et al. sva: Surrogate Variable Analysis. R package version 3.44.0. (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Shen, L. & ISoMaM, S. GeneOverlap: Test and visualize gene overlaps. R package version 1.34.0 http://shenlab-sinai.github.io/shenlab-sinai/ (2022).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov. (Camb. (Mass.)) 2, 100141 (2021).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet 48, 838–847 (2016).
Badia-I-Mompel, P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinforma. Adv. 2, vbac016 (2022).
Korotkevich, G. et al. Fast gene set enrichment analysis. bioRxiv 60012. https://doi.org/10.1101/060012 (2021).
Ouyang, J. F., Kamaraj, U. S., Cao, E. Y. & Rackham, O. J. L. ShinyCell: simple and sharable visualization of single-cell gene expression data. Bioinformatics 37, 3374–3376 (2021).
Acknowledgements
We would like to acknowledge all the authors that contributed for the development of the studies from which the data used in our HuMicA was collected. The importance of public data access must always be reinforced as a paramount tool for the progression of scientific knowledge, allowing for alternative interpretations of the original data and additional discoveries. We thank the CERCA Programme/Generalitat de Catalunya, the Josep Carreras Foundation and ICBAS-UP for institutional support. E.B. is funded by the Spanish Ministry of Science and Innovation (MICINN) [PID2020117212RB-I00; AEI/10.13039/501100011033] and the Catalan Agency for Management of University and Research Grants (AGAUR, 2021 SGR 01213). R.M.-F. was funded by an FCT (Fundação para a Ciência e Tecnologia) fellowship (SFRH/BD/137900/2018). UMIB is funded by FCT Portugal (UIDB/00215/2020 and UIDP/00215/2020), and ITR (LA/P/006/2020). A.L. is funded by Fondo de Investigación Sanitaria-Instituto de Salud Carlos III (Spain)-FEDER (PI21/01603) and MICINN (Ramón y Cajal [RYC2021-032947-I] financed by MCIN/AEI/10.13039/501100011033 and the European Union-NextGenerationEU/PRTR). EM.‘s work in this publication was supported by the Ramón y Cajal fellowship RYC2021-032359-I, funded by the Spanish Ministry of Science, and by the Catalan Agency for Management of University and Research Grants (AGAUR, 2021 SGR 01586). Our acknowledgments also extent to all members of the Epigenetics and Immune Disease Group at the Josep Carreras Leukaemia Research institute, to Prof. Berta Martins da Silva and the Immunogenetics Laboratory of ICBAS-UP, and to Prof. António Martins da Silva, M.D. and Dr. João Chaves from Hospital de Santo António – Centro Hospitalar e Universitário do Porto (HSA-CHUP) for the continuous collaborations and guidance. We also thank for advice Dr. Marta Martinez-Vicente from the Autophagy and Lysosomal Dysfunction Lab, Neurodegenerative Diseases Research Group, Vall d’Hebron Research Institute-CIBERNED, 08035 Barcelona. We are indebted to brain donors and families for the generous donation of brain tissue for diagnostic and research purposes to the Vall d’Hebron Neurological Tissue Bank (PR(AG)200/2013). We want to particularly acknowledge the patients and the Biobank Banco de Tejidos CIEN for their collaboration. We are grateful to Marta Valeri and Cristina de Dios from the Microscopy Core Facility at the High Technology Unit (UAT) from VHIR for support in confocal imaging; Teresa Moline from the Pathology Department at Vall d’Hebron University Hospital for assistance in histological processing.
Author information
Authors and Affiliations
Contributions
R.M.-F. and E.B. conceived the study. The bioinformatic analysis was performed by R.M-F. and J.C.-S. R.M.-F., J.C.-S., B.L., J.R.-U., E.M., P.P.C., A.L., and E.B contributed to the verification and interpretation of the analysis. A.L. and E.M.S. provided postmortem samples, performed, and analyzed validation experiments. R.M.-F. and E.B. were responsible for the writing of the article. All the authors reviewed, edited, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Marta Olah and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Martins-Ferreira, R., Calafell-Segura, J., Leal, B. et al. The Human Microglia Atlas (HuMicA) unravels changes in disease-associated microglia subsets across neurodegenerative conditions. Nat Commun 16, 739 (2025). https://doi.org/10.1038/s41467-025-56124-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56124-1
This article is cited by
-
Selective neuronal restoration of progranulin does not prevent the frontotemporal dementia like-phenotype of progranulin knockout mice
Journal of Neuroinflammation (2026)
-
Single-nucleus RNA sequencing dataset of diverse tissues from wild-type monkey and Tau-P301L transgenic monkey
Scientific Data (2026)
-
Genome-wide consensus transcriptional signatures identify synaptic pruning linking Alzheimer’s disease and epilepsy
Molecular Psychiatry (2026)
-
TSC-associated microglial hyperactivity: enhanced calcium signaling, metabolism, and phagocytosis
Acta Neuropathologica (2026)
-
Transcriptional and neuroprotective effects of hexokinase-2 inhibitors administered after stroke
Journal of Neuroinflammation (2025)