Abstract
Metastasis in cancer is influenced by epigenetic factors. Using an in vivo screen, we demonstrate that several subunits of the polybromo-associated BAF (PBAF) chromatin remodeling complex, particularly Brd7, are required for maintaining breast cancer metastatic dormancy in the lungs of female mice. Brd7 loss induces metastatic reawakening, along with modifications in epigenomic landscapes and upregulated oncogenic signaling. Breast cancer cells harboring Brd7 inactivation also reprogram the surrounding immune microenvironment by downregulating MHC-1 expression and promoting a pro-metastatic cytokine profile. Flow cytometric and single-cell analyses reveal increased levels of pro-tumorigenic inflammatory and transitional neutrophils, CD8+ exhausted T cells, and CD4+ stress response T cells in lungs from female mice harboring Brd7-deficient metastases. Finally, attenuating this immunosuppressive milieu by neutrophil depletion, neutrophil extracellular trap (NET) inhibition, or immune checkpoint therapy abrogates metastatic outgrowth. These findings implicate Brd7 and PBAF in triggering metastatic outgrowth in cancer, pointing to targetable underlying mechanisms involving specific immune cell compartments.
Similar content being viewed by others
Introduction
Metastasis accounts for >90 percent of cancer-related deaths1,2, and efforts to combat this hallmark of systemic malignancy remain the “holy grail” of clinical oncology. Thwarting metastasis is particularly important in the clinical management of breast cancer, the most common malignant tumor type affecting females3, where patient prognosis can be quite favorable in the setting of disease confined to the primary site. Breast cancer predominantly metastasizes to the lung, bone, liver, and brain with the triple-negative variant demonstrating the highest proclivity for lung metastasis. Importantly, more than 60 percent of breast cancer patients eventually succumbing to disease harbor lung metastasis4, and the median survival for patients with breast cancer lung metastasis is only 21 months5. As such, breast cancer lung metastasis is a clinically obstinate problem in urgent need of improved mechanistic understanding. Prior work from multiple groups has shown that metastasis-initiating cells disseminated across secondary organs enter a dormant state while evading different therapeutic and immune insults, only to reawaken at a later time to induce macroscopic metastatic outgrowth6. Within this framework, it is thought that at the time of diagnosis, tumor dissemination to secondary sites has already occurred in >50% of cancer patients7,8, while ~ 80% of lung metastases are thought to arise from early disseminated tumor cells9. Pre-metastatic dormancy would, therefore, seem to represent a tractable setting for appropriately targeted therapeutic intervention to prevent full metastatic relapse. However, the precise molecular mechanism(s) mediating escape from dormancy and subsequent metastatic outgrowth at sites like the lung remain unclear.
Epigenetic abnormalities and their impact on the cellular state have been widely implicated across human cancer10. More specifically, it has been postulated that disseminated cancer cells undergo complex epigenetic rewiring to acclimatize and thrive within distant sites11. Moreover, in the absence of proliferation, these dormant cancer cells may co-opt epigenetic modifications to guard themselves against immune insults. In this functional scheme, effectively manipulating interactions with surrounding microenvironmental immune constituents becomes as important as cell-autonomous considerations12. Much remains to be deciphered, however, with regard to the specific epigenetic events mediating metastatic dormancy and outgrowth, along with relevant immune microenvironmental actors amenable to therapeutic targeting.
To address these clinically relevant knowledge gaps, we employed in vivo shRNA epigenetic screening in previously developed murine breast cancer models of metastatic dormancy and reactivation13,14. This approach identified the polybromo-associated BRG1-associated factor (PBAF) major chromatin remodeling complex component Bromodomain-7 (Brd7) as a critical and potent mediator of breast cancer dormancy. Specifically, Brd7 loss induced shifts in chromatin accessibility and enhancer landscapes genome-wide to promote metastatic outgrowth in the lung through tumor cell-autonomous and immune microenvironmental mechanisms. Examining the microenvironment further, through both flow cytometry and single-cell transcriptomics, we found that Brd7 loss mobilized tumor-promoting inflammatory and transitional neutrophils and modulated T-cell profiles toward stressed and exhausted compartments. Moreover, these microenvironmental disruptions were reversible with appropriate translational pharmacological interventions, pointing to tractable treatment strategies against breast cancer metastatic relapse moving forward. Taken together, this work reveals a role for the Brd7 epigenetic regulator and the PBAF complex in the dormancy, relapse, and metastatic outgrowth of malignant breast cancer, while also characterizing targetable cellular and molecular mechanisms underlying this pathobiology.
Results
In vivo epigenetic screening reveals that PBAF loss promotes breast cancer lung metastasis
To identify epigenetic regulators of breast cancer lung metastatic outgrowth, we performed an in vivo loss-of-function screen, employing the 4T07-TGL murine triple-negative breast cancer cell line. Upon intravenous injection, 4T07-TGL cells characteristically home to the lungs where they persist as solitary cells or micro-metastatic lesions without significant outgrowth. Accordingly, 4T07-TGL cells exhibit properties of a “dormant” lung metastatic line13,15,16. We transduced luciferase-expressing 4T07-TGL cells with a focused shRNA library targeting 242 genes encoding established epigenetic factors and injected experimental and control derivatives into syngeneic Balb/cJ mice. We reasoned that macroscopic lung tumors forming in this context would reflect the silencing of key genes required for suppressing metastatic outgrowth. Barcode sequencing of metastatic lesions derived from the lungs of sacrificed mice revealed that 4 out of 36 gene hits enabling tumor formation (Brd7, Smarcb1, Arid2, and Smarca2) were subunits of the PBAF complex, an established multi-component epigenetic regulator within the SWI/SNF (SWItch/ Sucrose Non-Fermentable) family (Fig. 1A and Supplementary Fig. 1A, B). Moreover, amongst these 4 candidates, Brd7 (bromodomain-7), which encodes a PBAF-specific subunit, emerged as the top-scoring hit (Supplementary Fig. 1B). The SWI/SNF complex has been widely implicated as a tumor suppressor17,18 and frequent inactivating mutations in multiple SWI/SNF subunits (e.g., ARID1A, ARID1B, ARID2, PBRM1, BRD7, and BRG1) have been causally linked to multiple cancers19,20. Nevertheless, the precise role of PBAF complex subunits in triggering metastatic reactivation and outgrowth remains unclear.
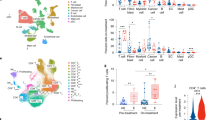
A Design of in vivo loss of function screen. After transduction with the mouse epigenetic shRNA libraries, the 4T07-TGL cells were injected i.v. into syngeneic Balb/cJ mice. Candidate suppressors of metastasis were ascertained by sequencing analysis of lung metastatic lesions and were further validated by additional shRNA studies. B, C 4T07-TGL cells with scrambled sgRNA vector (sgCtr) or Brd7 knockout were injected i.v. into syngeneic mice. (4T07-TGL-sgctr, n = 15 mice; 4T07-TGL-Brd7 KO1, n = 15 mice; 4T07-TGL-Brd7 KO2, n = 15 mice). The extent of lung colonization was measured by bioluminescent imaging. Panels show representative images (B), and quantified normalized photon flux at the indicated time points, error bars represent SD (C). D Kaplan-Meier survival analysis for mice injected with the indicated cell lines. E Representative H&E-stained images of lungs extracted from mice injected with the indicated cell lines at the experimental endpoint. F Quantified lung metastatic lesions in control and Brd7 KO mice (mean ± SEM.; 4T07-TGL-sgctr, n = 15 mice; 4T07-TGL-Brd7 KO1, n = 15 mice; 4T07-TGL-Brd7 KO2, n = 15 mice). Error bars represent SEM. G Representative images of lung sections subjected to IF for the indicated markers. Control (sgCtr) and Brd7 knockout (KO1 and KO2) 4T07-TGL cells were inoculated i.v., and lung sections stained with anti-Ki-67, anti-GFP and DAPI (G). H Bar graph showing the percentage of tumor cells with positive Ki-67 IF at the indicated times. (4T07-TGL-sgctr, n = 5 mice; 4T07-TGL-Brd7 KO1, n = 5 mice; 4T07-TGL-Brd7 KO2, n = 5 mice) *p < 0.05, **p < 0.01. Error bars represent SEM. I, J Kaplan-Meier analysis of distant metastasis-free survival (I) and relapse-free survival (J) in publicly available breast cancer datasets of the indicated subtype (source: KM Plotter for breast cancer). Patients were divided according to BRD7 expression level with the cutoff at 50%. HR, hazard ratio. K PBAF subunit (BRD7, PBRM1, and ARID2) expression in breast cancer primary and metastatic tumors; dataset indicated. *p < 0.05, **p < 0.01. [BRD7: Primary (n = 15 patients)- Minima = 6.966, Maxima = 10.996, Median expression = 9.627, Metastasis (n = 60 patients) – Minima = 8.380, Maxima = 10.759, Median expression = 9.354; PBRM1: Primary (n = 15 patients)- Minima = 10.087, Maxima = 12.162, Median expression = 11.349, Metastasis (n = 60 patients)- Minima = 9.471, Maxima = 12.045, Median expression = 10.115; ARID2: Primary (n = 15 patients)- Minima = 8.755, Maxima = 12.656, Median expression = 11.490, Metastasis (n = 60 patients)- Minima = 8.272, Maxima = 11.718, Median expression = 9.574 (L) PBAF specific subunit expression in normal breast tissue, breast primary tumor and metastases from TCGA invasive breast cancer dataset [Normal (n = 113 patients)- Minima = 775,Maxima = 5805, Median expression = 2018; Primary (n = 1210 patients)- Minima = 110,Maxima = 10285, Median expression = 1563; and Metastasis (n = 7 patients)- Minima = 611,Maxima = 1960, Median expression = 836). M Representative immunohistochemistry (IHC) staining for paired primary tumor and lung metastasis sections in breast cancer tissue microarrays. BRD7 expression was quantified by stain intensity across n = 3 independent tumor regions obtained from n = 1 female breast cancer patient. ****p < 0.0001. Statistical analysis by unpaired two-tailed t test, in all panels, error bars represent SD. 95% CI was used to analyze the results. Source data are provided as a Source Data file.
To validate the results of our screen, we generated Brd7, Arid2, Smarcb1, and Pbrm1 knockdown 4T07-TGL derivatives (Supplementary Fig. 1C) and injected these cell lines into Balb/cJ hosts. As expected, we found that repression of any of these genes induced metastatic outgrowth in the lung, as measured by in vivo bioluminescent imaging (BLI) (Supplementary Fig. 1D–F). As Brd7 represented the top hit in our screen, we chose to further investigate its role in metastatic dormancy and elucidate the mechanism(s) by which its inactivation promotes metastatic outgrowth in the lung. We first generated Brd7 knockout 4T07-TGL lines by CRISPR/CAS9 (Supplementary Fig. 2A, B) and found that complete Brd7 expression loss significantly increased in vivo lung metastasis, as assessed by both in vivo BLI and post-mortem histopathology, while also reducing overall survival (Fig. 1B–F). Brd7 inactivation had no effect on 4T07-TGL cells implanted in the mammary fat pad, with Brd7 KD and control lines exhibiting equivalent growth kinetics (Supplementary Fig. 2C). Moreover, knocking down Brd7 in the D2A1d dormant cell line also led to increased lung metastasis formation (Supplementary Fig. 2D). Brd7-KO 4T07-TGL lung metastases demonstrated a high level of proliferation by Ki-67 immunofluorescence (IF), while lungs from mice injected with 4T07-sgCtr controls were comparatively inert, consistent with the known metastatic dormancy of the parental line (Fig. 1G–H). No significant Brd7-dependent effect was observed on the number of solitary quiescent cells at either 7- or 14-day timepoints (Supplementary Fig. 2E). Finally, Brd7 overexpression in the murine triple-negative 4T1 cell line (Supplementary Fig. 2F), which is highly metastatic at baseline, significantly curbed mammary (primary site) and metastatic lung tumor burden in vivo, while significantly downregulating colony formation, tumor sphere formation, and migration in vitro (Supplementary Fig. 2G–I and Supplementary Fig. 3A–D). Consistent with this latter observation, we found by RT-PCR that Brd7 and Bcl7c were the only PBAF, BAF, SWI/SNF core, or SWI/SNF ATPase subunits demonstrating significantly downregulated expression in 4T1 cells relative to both dormant 4T07-TGL cells and a second murine breast cancer line (168FARN), also characterized by limited metastatic capacity (Supplementary Fig. 4A–D). Moreover, increased metastasis in the Brd7-deficient context was largely restricted to the lung, although the bioluminescence signal was modestly upregulated in the liver; no similar evidence of metastasis was observed in the kidney, heart, and bone (Supplementary Fig. 5A–D). Taken together, these findings reveal a selective role for Brd7 loss in lung metastatic outgrowth. That Brd7 overexpression in 4T1 cells and Brd7 KO in 4T07-TGL cells both impacted metastatic growth, while only the former affected mammary fat pad growth was somewhat surprising but may reflect different absolute protein level set points required to influence in vivo tumor growth in primary and metastatic locations. Consistent with this conjecture, we found by western blot that Brd7 KO in 4T07-TGL cells resulted in protein levels comparable to those seen in parental 4T1 cells, while Brd7 overexpression in 4T1 cells resulted in much higher protein levels than those seen in parental 4T07 cells (Supplementary Fig. 6A).
We next evaluated publicly available patient data for associations between BRD7 expression and breast cancer patient outcomes. Low BRD7 levels strongly correlated with significantly decreased distant metastasis-free and relapse-free survival for triple-negative breast cancer in the BRCA_Subtype_PAM50 dataset (Fig. 1I, J)21. Moreover, breast cancer patients with metastasis exhibited significantly lower PBAF subunit (BRD7, PBRM1, and ARID2) expression than metastasis-free counterparts in the Siegel et al. dataset (Fig. 1K)22. Results were similar in the invasive breast cancer TCGA dataset, where we found that expression of a PBAF subunit signature consisting of BRD7, PBRM1, ARID2, and PHF10 was significantly lower in patients with metastasis than in those without (Fig. 1L). Using immunohistochemistry (IHC) in tissue microarrays, we then found that lung metastases demonstrated significantly reduced BRD7 staining intensity relative to case-matched primary breast cancers (Fig. 1M). These tissue-based results further support a central role for BRD7 and PBAF in the suppression of lung metastatic outgrowth in breast cancer.
Brd7 loss modulates cell autonomous and immune microenvironmental functionalities through shifts in chromatin accessibility and enhancer elements
To better ascertain potential molecular mechanisms driving metastatic outgrowth in the context of Brd7 loss, we performed RNA-seq on our isogenic 4T07-TGL lines, subjecting differentially expressed genes to Hallmark and Gene Ontology (GO) correlative analyses. Brd7 KO transcriptional profiles demonstrated positive associations with an array of pathways and functionalities previously implicated in breast cancer, metastasis, and oncogenesis, notably including Hedgehog signaling, epithelial-to-mesenchymal transition, PI3K-AKT-mTOR signaling, neurogenesis, gland development, and chemotaxis (Fig. 2A, B and Supplementary Fig. 6B–D)23,24,25,26,27,28. Brd7 loss also increased levels of pAKT as assessed by western blot (Fig. 2C). By contrast, we found that Brd7 KO transcriptional profiles exhibited negative associations with a variety of immune-related pathways, including interferon response, antigen presentation, MHC class 1 (MHC-1)-related networks, and T cell receptor signaling (Fig. 2D–F). Both antigen presentation and MHC-1 expression on tumor cells dictate the extent to which effective immunosurveillance occurs. Moreover, attenuated interferon response has been previously reported to promote metastasis by downregulating tumor immunogenicity29,30. We found that several MHC-1 genes—namely H2-Q4, H2-Q7, H2-K1, H2-Q10, H2-Q6, and H2-D1—along with established markers of interferon response, were downregulated in Brd7 KO 4T07-TGL cells relative to isogenic controls (Fig. 2E). To further interrogate the impact of Brd7 loss on antigen presentation, we performed FACS analysis for MHC-1 (H-2Kd/H-2Dd) on our 4T07-TGL isogenics, documenting markedly decreased expression in Brd7 KO cells compared to controls (Fig. 2G–H). These findings were recapitulated by IHC for MHC-1 performed in lung sections obtained from mice injected with either 4T07-TGL control or Brd7-KO cells (Fig. 2I, J). Brd7 loss also led to modest but significant increases in colony formation (Fig.2K, L and Supplementary Fig. 6E, F), tumorsphere formation (Fig. 2M, N), cell migration (Fig. 2O, P), and invasion (Supplementary Fig. 6G). These functional assessments broadly correlated with transcriptional profiling results, further implicating both tumor cell-autonomous and immune microenvironmental processes in Brd7-deficient lung metastatic outgrowth.
A Gene Set Enrichment Analysis (GSEA) of transcriptional profiles from Control (sgCtr) and Brd7 KO (KO1 and KO2) 4T07-TGL cells. The top 10 enriched pathways in control or Brd7 KO cells were ranked by normalized enrichment score (NES) with FDR < 0.25, p < 0.05. B Gene Ontology (GO) analysis of upregulated genes in Brd7 knockout 4T07-TGL cells. p < 0.05. C Control (sgCtr) and Brd7 knockout (KO1 and KO2) 4T07-TGL cells were subjected to immunoblotting for AKT phosphorylation at Ser473, along with total AKT, Brd7, and GAPDH (loading control). Quantification of the Western Blot results obtained from N = 3 independent experiments. The samples were derived from the same experiment, but different gels for Brd7, GAPDH, another for pAKT, AKT were processed in parallel. For Brd7 and GAPDH, the membrane was incubated overnight with Rabbit anti-Brd7 and Mouse anti-GAPDH antibody at same time, and then incubated with anti-rabbit IGg and anti-mouse IGg antibody at same time. D Gene Ontology (GO) analysis of downregulated genes in Brd7 KO 4T07-TGL cells. p < 0.05. E Heat map of top differentially expressed genes related to antigen presentation and response to interferon between control and Brd7 KO 4T07-TGL cells. F Hallmark_ALPHA_RESPONSE and HALLMARK_T_CELL_ RECEPTOR_SIGNALING gene sets were significantly enriched in control 4T07-TGL cells compared to Brd7 KO cells. G, H H-2Kd/H-2Dd expression in Control (sgCtr) and Brd7 KO (KO1 and KO2) 4T07-TGL cells as detected by flow cytometry. The mean intensity of H-2Kd/H-2Dd was quantified (H). Quantification of the FACS results was obtained from N = 3 biological replicates (I, J) Representative images of MHC-1 immunohistochemistry staining performed on lung tissue sections obtained from the indicated mice cohorts (I), and quantification of MHC−1 mean intensity is plotted (sgCtr = 3 mice; Brd7-KO = 4 mice)(J). K–P In vitro colony formation (K), tumor sphere (M), and migration (O) assays for Control (sgCtr) and Brd7 KO (KO1 and KO2) 4T07-TGL cells. Bar graphs show quantification assay results obtained from N = 3 independent biological replicates obtained from indicated groups (L, N, P). Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. 95% CI was used to analyze the results. Source data are provided as a Source Data file.
We then sought to elucidate the extent to which loss of BRD7 influences chromatin landscapes, thereby facilitating metastasis-promoting transcriptional profiles. As BRD7 and the PBAF complex had been widely implicated in regulating chromatin accessibility31,32,33,34, we performed ATAC-seq as an early study on our initially generated Brd7-KD 4T07-TGL lines. As expected, Brd7 suppression resulted in widespread shifts in chromatin architecture, both increasing and decreasing accessibility genome-wide (Fig. 3A, B). Impacted loci were most enriched for distal intergenic and intronic regions (Fig. 3C). Gene Ontology analysis performed on the 44 upregulated genes (RNA-seq) showing overlap with regions of increased ATAC-seq accessibility demonstrated significant associations with cell motility and developmental pathways (Fig. 3D). Motif analysis for these loci revealed enrichment for multiple pro-metastatic transcription factors, such as Atf435, ETV136, MAFK37, ERG38, ETV439, SIX140, ETS141, and TEAD442 (Fig. 3E). Gene Ontology analysis performed on the 69 downregulated genes (RNA-seq) showing overlap with regions of decreased ATAC-seq accessibility demonstrated significant associations with immune-related pathways like response to interferon-beta and T cell selection, concurring with transcriptional and functional data (Fig. 3F). Moreover, motif analysis of these same loci demonstrated enrichment for putative JunB, AP-1, Fos, and BATF transcription factors (Fig. 3G). Prior work has documented JunB as both a tumor suppressor in breast cancer and a regulator of innate and adaptive immune response43 that also forms dimeric complexes with AP-1, Fos, and BATF family proteins.
A ATAC-seq was performed in Control (shNTC) and Brd7-KD (Brd7-sh1 and Brd7-sh4) 4T07-TGL cells. Chromatin accessibility heat maps for gained (top panel) and lost (bottom panel) peaks in Brd7-KD 4T07-TGL cells. Peaks were rank ordered by intensity per million mapped reads (RPM). B Overall aggregated enrichment signals within 3 kb of peak centers for differentially accessible regions. C Genomic annotation of gained peaks (left panel) and lost peaks (right panel). D GO analysis of overlap between upregulated genes in Brd7-KD samples (RNAseq) and genes associated with gained ATAC-seq peaks in Brd7-KD samples. E Top 13 gained peak motifs from these overlapping loci and corresponding transcriptional factors. F GO analysis of overlap between downregulated genes in Brd7-KD samples (RNAseq) and genes associated with lost ATAC-seq peaks in Brd7-KD samples showing enrichment for immune regulation pathways. G Top 8 enriched binding motifs from these overlapping loci and predicted transcriptional factors. H Heatmap showing ChIP-Seq signals for H3K27ac peak regions, rank ordered by intensity per million mapped reads (RPM). I Overall H3K27ac peak signal between − 3 kb and 10 kb relative to peak center for gained or lost peaks in Brd7-KO cells. J Genomic annotation of gained (left panel) and lost (right panel) H3K27ac peak distribution. K, L Gene Ontology (GO) analysis of overlap between upregulated genes (RNAseq) and genes associated with gained H3K27ac peaks in Brd7-KO samples (K; N = 224)) and between downregulated genes (RNAseq) and genes associated with lost H3K27ac peaks in Brd7-KO samples (L; N = 55). M, N Top binding motifs in Brd7-KO cells associated with lost (M) and gained (N) H3K27ac and predicted transcriptional factors.
Beyond their impact on chromatin accessibility, PBAF and SWI/SNF complex elements have been shown to maintain epigenomic architecture at enhancer elements throughout the genome44,45. To ascertain Brd7-dependent enhancer profiles in our 4T07-TGL isogenics, we performed ChIP-seq for H3K27ac, a canonical enhancer mark, in our KO and control lines. We found that Brd7 loss altered the extent of H3K27ac labeling at thousands of genomic sites, with the majority showing upregulated signal intensity (Fig. 3H, I). Most loci of increased H3K27ac localized to promoter regions, while loci of reduced H3K27ac mapped primarily to the distal intergenic space (Fig. 3J). Integrating these results with RNA-seq data, we found extensive overlap between upregulated genes and loci of increased H3K27ac, and between downregulated genes and loci of decreased H2K27ac (Fig. 3K, L). By gene ontology analysis, upregulated transcripts showing overlap with altered enhancer elements (N = 224) demonstrated significant links with functionalities related to cellular motility (e.g., actin filament-based process, Rho protein signal transduction), while downregulated counterparts (N = 55) demonstrated immune microenvironmental correlations (e.g., antigen processing and presentation, response to interferon-beta). These transcriptional findings echo in vitro functional studies (Fig. 2). Finally, motif analysis of genomic loci exhibiting reduced H3K27ac with Brd7 loss revealed multiple transcription factors with key roles in adaptive immune cell lineages (NFAC3, NFATC2, SMAD2) (Fig. 3M), while motif analysis for areas of increased H3K27ac demonstrated enrichment for pro-metastatic and pro-tumorigenic transcription factors (ZFX, Meis1, Zac1, ZNF711, TEAD4, TWIST2) (Fig. 3N). Taken together, these transcriptional and epigenomic data point to functionally relevant cell-autonomous and immune microenvironmental pathways potentially mediating lung metastatic outgrowth in the BRD7-deficient context.
Brd7 loss reprograms the tumor immune microenvironment, upregulating inflammatory and transitional neutrophils and effector T cell populations to promote metastatic outgrowth
As indicated above, the impact of Brd7 loss on 4T07-TGL cell migration, sphere formation, and clonogenicity (Fig. 2K–P), while functionally compelling and statistically significant, was relatively modest and did not correlate with the dramatic metastatic outgrowth phenotype in vivo (Fig. 1C, F). Our integrated transcriptional and epigenomic analyses also implicated immune microenvironmental modulation as a potential driver of metastatic outgrowth, findings that were further supported by notably reduced MHC-1 expression in Brd7-KO cells. Moreover, recent studies have clearly demonstrated the extent to which epigenetic mechanisms can enable cancer cells to evade immunosurveillance machinery, supporting escape from dormancy and malignant evolution46,47. Finally, 4T07-TGL cells injected intravenously into either Nu/Nu and NOD SCID immunodeficient murine strains resulted in extensive lung metastases within 3 weeks (Supplementary Fig. 7), underscoring the importance of the immune system for suppressing the metastatic outgrowth in this model system.
To ascertain the effects of Brd7 loss on immune-relevant signaling, we performed Luminex analysis for a panel of 40 cytokines on supernatants obtained from 4T07-TGL Brd7-KO and control cells. We found that four pro-metastatic cytokines—CXCL10,CCL5,CCL2, and CCL20—were significantly upregulated in the Brd7-deficient context (Fig. 4A and Supplementary Fig. 8A). Interestingly, all four of these cytokines have been previously implicated in functionally relevant tumorigenic processes, including angiogenesis, invasion, and metastasis, while also promoting recruitment of specific myeloid and lymphoid subtypes48,49,50,51. Having already shown that Brd7 loss upregulated oncogenic PI3K-AKT-mTOR signaling, we evaluated the extent to which Brd7-deficient cytokine release was impacted by LY294002, a broad-spectrum PI3K inhibitor. LY294002 treatment for 5 h significantly downregulated levels of CCL5 and CXCL10 while treatment for 15 h significantly downregulated levels of CCL20 released by Brd7-KO 4T07-TGL cells in conditioned media (Fig. 4B). These effects contrasted with those observed in control 4T07-TGL cells, where cytokine release was either unchanged or increased (for CCL20 alone) by LY294002. Our findings functionally implicate the PI3-AKT-mTOR pathway in Brd7-deficient modulation of the breast cancer tumor microenvironment.
A Luminex analysis on conditioned supernatants showing significantly increased levels of 4 pro-metastatic cytokines—CXCL10, CCL5, CCL2, and CCL20—in Brd7-KO 4T07-TGL cells relative to controls (sgCtr). **p < 0.01, ***p < 0.001. Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. Supernatants were obtained from N = 3 independent biological replicates from each of the indicated groups. B CCL20, CCL5, and CXCL10 expression by Luminex assay in Brd7 KO and sgCtr 4T07-TGL cells treated with either vehicle (DMSO) or the PI3K inhibitor LY294002. Supernatants were obtained from N = 3 independent biological replicates from each of the indicated groups. **p < 0.01, ***p < 0.001. Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. C, D Immunophenotypic flow cytometry on dissociated lungs removed from naïve 8-weeks old Balb/cJ mice or similarly aged littermates 14 days after injection with either control or Brd7 KO 4T07-TGL cells. (**p < 0.01, ***p < 0.001; WT naïve mice lungs, N = 4 mice; 4T07-TGL lungs, N = 4 mice; 4T07-TGL-Brd7 KO1 lungs, N = 4 mice; 4T07-TGL-Brd7 KO2 lungs, N = 4 mice). Statistical analysis by unpaired two-tailed t test. E–G Single-cell analyses of immune cell populations derived from dissociated lungs removed from naïve 8-weeks old Balb/cJ mice or similarly aged littermates 14 days after injection with either control or Brd7-KO 4T07-TGL cells (WT untreated naïve mice lungs, N = 3 mice; 4T07-TGL lungs, N = 3 mice; 4T07-TGL-Brd7 KO1 lungs, N = 3 mice). UMAP plots show clustering and transcriptional assignments for all assessed cells (E). and cells separated by murine cohort (naïve (untreated), control (4T07), and Brd7 KO) (F). Bar graph showing the composition of different immune compartments in the indicated murine cohorts. ***p < 0.001. Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD (G). H–K Represent images of the H&E-stained (H) and immunohistochemistry (IHC)-stained (MPO (I), neutrophil elastase (K), and CD8 (M)) lung sections of mice from the indicated murine cohorts. (J, L, N) Quantification of IHC staining (For MPO and NE staining:4T07-TGL-Brd7 KO, N = 4 mice, 4T07-TGL-sgCtr,N = 3 mice; For CD8 + staining: 4T07-TGL-Brd7 KO, N = 3 mice, 4T07-TGL-sgCtr,N = 3 mice). *p < 0.05, **p < 0.01. Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. O Spearman correlation between BRD7 expression and overall immune infiltration in breast cancer patients (BRCA_MBC patient dataset). 95% CI was used to analyze the results. Source data are provided as a Source Data file.
To interrogate the effects of Brd7 loss on specific immune microenvironmental constituents, we performed immunophenotypic flow cytometry on dissociated lungs removed from naïve 8-week-old mice or similarly aged littermates 14 days after injection with either control or Brd7-KO 4T07-TGL cells (Supplementary Fig. 9A). Intriguingly, we found that Brd7 loss significantly increased neutrophil levels, while also downregulating monocytic cells, macrophages, and dendritic cells in vivo (Fig. 4C, D and Supplementary Fig. 9B). To build on these observations and further delineate the extent to which Brd7 loss alters specific immune cell subpopulations, we subjected dissociated lungs from the same experimental paradigm to scRNAseq (Supplementary Fig. 10A). These studies documented the diverse immune microenvironmental composition of lungs obtained from naïve un-injected mice and littermates injected with either 4T07-TGL control or 4T07-TGL Brd7-KO cells, as depicted by UMAP clustering (Fig. 4E, F). Discrete cellular subsets were then identified by correlative expression signature analysis (Supplementary Fig. 10B, C). Comparing specific immune cell populations, we found that Brd7 loss led to significant increases in neutrophils and effector T cell populations, while downregulating macrophages, megakaryocytes, and plasma cells (Fig. 4G).
Neutrophils have been widely reported to promote cancer cell invasion and migration and have also been recognized as a major driver of breast cancer metastasis through multiple mechanisms52,53,54. To further characterize these neutrophil populations, we performed additional subclustering of our scRNA-seq data, delineating inflammatory and transitional neutrophil variants, both of which demonstrated increased levels with Brd7 KO in dissociated lung specimens 14 days after 4T07-TGL injection (Supplementary Fig. 11A–D). Recent studies have found that the presence of inflammatory and transitional neutrophil subtypes correlates with increased metastasis55,56. In addition, within these neutrophil subclusters, we observed upregulation of Wfdc17, Lrg1, Chd1, and Irf9 (Supplementary Fig. 11E) markers widely implicated in so-called neutrophil extracellular traps (NETs), web-like extracellular DNA structures composed of chromatin and granule proteins that recent reports have implicated in metastatic processes57,58. Further supporting our single-cell transcriptional findings, immunohistochemistry for myeloperoxidase (MPO) and neutrophil elastase (NE), two well-established lineage markers, demonstrated increased neutrophilic infiltrates in Brd7-KO 4T07-TGL lung metastases relative to 4T07-TGL control counterparts (Fig. 4H–L). Effector T cells represent a heterogeneous population of diverse functionality—including but not limited to CD8 + T cells, Tregs, Th17 cells, and γδ T cells—some of which are known to exert pro-metastatic influence on tumors59,60,61,62,63,64. We found that Brd7 KO resulted in a modest but significant increase in CD8 + T cell infiltration by IHC compared to the 4T07-TGL controls (Fig. 4M, N). We also assessed the extent of macrophage infiltration using IHC for F4/80 but found no significant changes associated with Brd7 KO (Supplementary Fig. 12). Taken together, these findings implicate specific immune cell constituents in the Brd7-deficient metastatic outgrowth phenotype.
Finally, we probed existing human data sets to characterize associations between BRD7 and specific immune microenvironmental components. Spearman correlation analysis demonstrated that lower BRD7 expression denoted reduced overall immune infiltration in the BRCA_MBC patient dataset for metastatic breast cancer (Fig. 4O). In addition, in the same dataset, low BRD7 expression correlated significantly with both increased neutrophil abundance and decreased Type 1 regulatory T cells, CD4 + naïve T cells, and nTreg immune cells (Supplementary Fig. 13A).
CD8 + exhausted T cells and stress response CD4 + T cells are enriched in breast cancer lung metastasis upon Brd7 loss
To better ascertain the specific effector T cell populations upregulated in the Brd7-deficient context, we incorporated classifying expression signatures into our scRNAseq analysis, delineating 8 additional sub-clusters: naïve CD4 + T cells, naïve CD8 + T cells, stress response CD4 + T cells, Tregs, effector CD8 + T cells, NK T cells, Ccr7 + memory T cells, and Th17 cells (Fig. 5A–C). We found that Brd7 KO in 4T07-TGL lung metastases was associated with significantly decreased naïve CD4 + T cells and significantly increased stress response CD4+ and effector CD8 + T cells (Fig. 5B). Naïve CD4 + T cells have been shown to stimulate anti-tumor immune defense65 and are essential for recognizing and combating foreign antigens. By contrast, stress response CD4 + T cells, a recently discovered immune sub-compartment, have been associated with aggressive phenotypes in multiple malignancies, including triple-negative breast cancer, and treatment refractivity prior to and after immune checkpoint therapy66. Examining stress response genes and transcriptional signatures in effector T cell populations delineated by our scRNAseq data, we found that levels of two key heat-shock protein transcripts—Hspa1a and Hspa1b—were significantly upregulated not only in stress response CD4 + T cells, but also in Tregs, effector CD8 + T cells, and Ccr7 + memory T cells (Fig. 5D). Moreover, within the stress response CD4 + T cell cluster, Brd7 loss in 4T07-TGL allografts induced strong transcriptional correlations with Hallmark reactive oxygen species and apoptosis pathways, relative to Brd7-intact controls (Fig. 5E). These data demonstrate that lung metastatic outgrowth mediated by Brd7 loss is associated with stress response pathway engagement in multiple relevant T cell compartments.
A UMAP plot showing clustering and transcriptional assignments for CD3 + cell populations derived from dissociated lungs removed from naïve 8-weeks old Balb/cJ mice or similarly aged littermates 14 days after injection with either control or Brd7 KO 4T07-TGL cells. B Bar graph showing the composition of different T cells compartments in the indicated murine cohorts. (WT untreated naïve mice lungs, N = 3 mice; 4T07-TGL lungs, N = 3 mice; 4T07-TGL-Brd7 KO1 lungs, N = 3 mice). Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. C Dot plot showing marker gene expression levels in different T cell compartments. D Hspa1a and Hspa1b expression of different T cell compartments in the indicated murine cohorts. Statistical analysis by unpaired two-tailed Mann-Whitney test; **p < 0.01, ***p < 0.001. E Gene Set Enrichment Analysis (GSEA) of cluster T2 gene expression in response to Brd7 loss in 4T07-TGL cells; HALLMARK REACTIVE OXYGEN PATHWAY and HALLMARK APOPTOSIS gene sets were significantly enriched. F UMAP plots showing the expression of T-cell cytotoxic and exhaustion marker genes (Cd8b1, Gzma, Cd44, Gzmk, Ifng, Ctla4, Lag3, Pdcd1, and Tigit) in cluster T4 subpopulations. G UMAP plots showing CD8 + Teff, Tmem T cells, and Tex T cells in the indicated murine cohorts. H Bar graph showing the composition of CD8 + Teff, Tmem T cells, and Tex T cells in the indicated murine cohorts. I Exhaustion marker gene expression in cluster T4 cells in the indicated murine cohorts. Statistical analysis by unpaired two-tailed Mann-Whitney test; *p < 0.05, ****p < 0.0001, ns: not significant. J, K Representative images (J) and quantification (K) of CTLA4 IHC for lung sections in the indicated murine cohorts. (sgCtr = 3 mice; Brd7-KO = 4 mice) **p < 0.01; statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SD. L, M Gene Set Enrichment Analysis (GSEA) of cluster T4 gene expression in Brd7 KO versus control murine cohorts. The negative regulation of the immune system process gene set was found to be significantly enriched in the Brd7 KO group (M). 95% CI was used to analyze the results. Source data are provided as a Source Data file.
We then performed additional signature-based subclustering of the effector CD8 + T cell pool, designating T effector (Teff), T memory (Tmem), and exhausted T cells from our scRNAseq data (Fig. 5F, G). Comparing our murine cohorts, we found that while levels of Teff and Tmem cells were downregulated with Brd7 loss, CD8 + exhausted T cells were significantly increased in number (Fig. 5H). Tmem cells have been reported to prevent invasion and metastasis, particularly in breast and colorectal cancers67,68. By contrast, exhausted T cells reflect a hyporesponsive state driven by prolonged antigen stimulation and are unable to effectively mitigate tumorigenesis69,70. Further analysis of the effector CD8 + T cell population revealed elevated levels of multiple T cell exhaustion markers—CTLA4, LAG3, TIGIT, PDCD1, TOX, and NFATC1—in response to Brd7 KO allografts (Fig. 5I). Immunohistochemical staining in lung tissue from mice bearing 4T07-TGL metastases confirmed increased CTLA4 expression in lymphocytes within Brd7-KO tumors (Fig. 5J, K). Taken together, these findings implicate targetable mechanisms of immunosuppression induced by Brd7 KO in the process of lung metastatic outgrowth. Further supporting this notion, GSEA analysis of scRNAseq profiles from the T cell cluster demonstrated that Brd7 loss in 4T07-TGL allografts enhanced correlations with tolerance induction, negative regulation of immune system process, negative regulation of immune response, and negative regulation of leukocyte cell adhesion (Fig. 5L, M).
Targeting the immune-suppressive microenvironment abrogates Brd7-deficient breast cancer metastatic outgrowth in lung
Having implicated multiple immune cell subtypes in Brd7-deficient lung metastatic outgrowth, we sought to determine the extent to which targeting these specific compartments would be of potential therapeutic benefit. As indicated above, Brd7-KO 4T07-TGL allografts promoted neutrophil infiltration, which prior work has already implicated in breast-to-lung metastasis71,72. To specifically interrogate the role of neutrophils in our in vivo model system, we subjected mice to treatment anti-Ly6G (300 µg/mice) or isotype control antibody every other day starting 5 days prior to injection with either 4T07-TGL control or 4T07-TGL Brd7-KO cells (Supplementary Fig. 14A). Anti-Ly6G achieved robust depletion of the neutrophil compartment, as assessed by flow cytometric analysis (Supplementary Fig. 14B, C). Imaging allografted mice weekly, we found that anti-Ly6G-mediated neutrophil depletion dramatically reduced lung metastases in the Brd7-deficient context, reverting normalized photon flux to that seen for 4T07-TGL control allografts (Fig. 6A, B and Supplementary Fig. 14D). Control IgG treatment had no effect. As indicated above, recent reports have suggested that neutrophils promote metastasis through NETs57,58. Thus, we sought to quantify NET formation in a neutrophil/cancer cell (100:1) co-culture system ex vivo with Sytox Orange, a non-cell permeable DNA specific dye73,74,75,76 (Fig. 6C). Neutrophil/4T07-TGL Brd7- KO cell co-culture resulted in significantly increased NET formation when compared to co-cultures with 4T07-TGL control cells (Fig. 6D). To disrupt NET formation in our 4T07-TGL Brd7 KO allografts, we utilized the neutrophil elastase inhibitor sivelestat in vivo, which has previously been shown to prevent NETosis54. We found that sivelestat treatment every other day, starting 5 days prior to 4T07-TGL Brd7 KO cell injection (Supplementary Fig.14E), severely abrogated Brd7-KO lung metastases in vivo, as measured by BLI and normalized photon flux (Fig. 6E, F and Supplementary Fig. 14F). These data confirm the importance of neutrophils and NETosis in Brd7-deficient lung metastatic outgrowth.
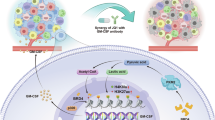
A, B Mice were treated with anti-Ly6G (300 µg/mice) or isotype control antibody every other day starting 5 days prior to injection with either 4T07 control or 4T07 Brd7 KO cells (Brd7 KO-Anti-Ly6G, N = 9 mice;Brd7 KO-IgG, N = 8 mice; 4T07-TGL-Anti-Ly6G,N = 8 mice; 4T07-TGL-IgG,N = 8 mice). Representative bioluminescence images for the indicated murine cohorts at 30 min, 2 weeks, and 4 weeks post tumor cell injection (A). Line graph showing normalized photon flux for the indicated murine cohorts over time (B). ****p < 0.0001. Statistical analysis by unpaired two-tailed Mann-Whitney test; error bars represent SEM. C Experiment design of neutrophil/cancer cell co-culture and NET quantification. D NET extracellular DNA quantification by Sytox Orange fluorescence for the indicated co-cultures. (**p < 0.01; N = 6) Statistical analysis by unpaired two-tailed t test; in all panels, error bars represent SEM. E, F Mice were treated with the NET inhibitor Sivelestat or vehicle every other day starting 5 days prior to injection with either 4T07 control or 4T07 Brd7 KO cells (Brd7 KO-Vehicle treatment, N = 10;Brd7 KO-Sivelestat treatment, N = 10). Represent bioluminescence images for the indicated murine cohorts at 30 min, 1 week, and 4 weeks post tumor cell injection (E). Line graph showing the normalized photon flux for the indicated murine cohorts over time (F). ****p < 0.0001. Statistical analysis by unpaired two-tailed Mann-Whitney test; error bars represent SEM. G, H Mice were treated with isotype control, Anti-LAG3, Anti-CTLA4, or both Anti-LAG3 and Anti-CTLA4 every other day starting 1 day after cancer cell injection until Week 5 (Brd7 KO-Anti-LAG3, N = 10 mice;Brd7 KO-Anti-CTLA4, N = 10 mice; 4T07-TGL-Anti-LAG3 + CTLA4, N = 10 mice; 4T07-TGL-Brd7 KO-IgG, N = 10 mice). Representative bioluminescence images for the indicated murine cohorts at 30 min, 2 weeks, and 5 weeks post tumor cell injection (G). Line graph showing the normalized photon flux for the indicated murine cohorts over time (H). ****p < 0.0001. Statistical analysis by unpaired two-tailed Mann-Whitney test; error bars represent SEM. I Kaplan-Meier survival analysis for the indicated murine cohorts. Statistical analysis by log-rank test. ***p < 0.001, ns, not significant. J Schematic of the mechanism(s) governing BRD7-deficient breast cancer lung metastatic outgrowth. 95% CI was used to analyze the results unless mentioned otherwise. Source data are provided as a Source Data file.
Our scRNAseq analysis also implicated lymphocyte stress response and cytotoxic T-cell exhaustion in breast-to-lung metastatic outgrowth induced by Brd7 loss. As indicated above, the cytotoxic CD8 + T cell compartment demonstrated upregulated expression of several established exhaustion markers in the setting of Brd7 deficiency, including CTLA4, PD1 (PCDC1), LAG3, and TOX. Interestingly, while neutrophils have been shown to generally impair T cell activation and proliferation, NETs have specifically been demonstrated to drive T cell exhaustion77,78. To evaluate this specific effect of NETosis in the Brd7-deficient context, we treated the mice injected with Brd7-KO cells with either Sivelestat or DMSO starting at the 2-week timepoint, which marks the onset of lung metastatic outgrowth (Supplementary Fig. 15A). FACS analysis of dissociated lungs isolated at the 4-week timepoint demonstrated a significant decrease in CTLA4 + CD8 + and PD1 + CD8 + T cells upon Sivelestat treatment (Supplementary Fig. 15B, C). These findings indicate that disrupting NET formation significantly attenuates T cell exhaustion induced upon Brd7 loss.
We also evaluated the extent to which targeting CTLA4 and LAG3, the two immune checkpoint receptors most highly upregulated in the Brd7-deficient context, could impair lung metastatic outgrowth. Both LAG3 and CTLA4 feature prominently in current immunotherapy management. We treated mice subjected to 4T07-TGL injection with the LAG3 inhibitor relatlimab, the CTLA4 inhibitor ipilimumab, or both, monitoring lung tumor burden with BLI imaging (Supplementary Fig. 15D). We found that ipilimumab profoundly reduced lung metastatic burden in mice injected with 4T07-TGL Brd7 KO cells (Fig. 6G, H and Supplementary Fig. 15E). By contrast, targeting LAG3 had minimal impact on lung metastasis alone, and did not appreciably augment the effects of ipilimumab (Fig. 6G, H). Concordant with these response data, ipilimumab, either alone or in combination with relatlimab, significantly prolonged survival in 4T07-TGL Brd7 KO-allografted mice, while relatlimab alone did not (Fig. 6I). H&E staining confirmed the complete abrogation of Brd7-deficient lung metastasis in both the anti-CTLA4 and the anti-CTLA4/anti-LAG3-treated cohorts (Supplementary Fig. 15F). Taken together, these results underscore the key role played by T cell exhaustion in Brd7-deficient lung metastatic outgrowth and raise the possibility of implementing immune checkpoint inhibition in analogous clinical contexts.
Discussion
Breast cancer metastasis, a primary driver of patient mortality, often arises several years after the initial diagnosis of the primary tumor. This likely reflects tumor cells extravasating to distant secondary sites early in the disease course79, followed by an extended period of dormancy characterized by dynamic equilibrium with the surrounding immune microenvironment. Metastatic dormancy allows for tumor cell survival in the face of microenvironmental insults in the premetastatic niche. The signals and molecular factors that promote metastatic dormancy, along with subsequent escape from dormancy and macroscopic expansion remain largely unknown. Through our in vivo loss-of-function epigenetic screen in a physiologically relevant dormancy model, we identified several components of the SWI/SNF family of chromatin remodelers, namely Brd7, Arid2, and Smarcb1, as mediators of breast cancer dormancy whose loss induces lung metastatic outgrowth. Amongst these hits, Brd7 emerged as the most robust dormancy regulator.
Epigenetic alterations in cancer cells have previously been shown to influence a range of cancer-relevant cellular processes, including proliferation, invasion, and migration, through their influence on oncogenic signaling. We found that loss of Brd7 triggered upregulation of PI3K-AKT-mTOR, Hedgehog, and EMT networks. PI3K-AKT-mTOR signaling has been widely implicated in breast cancer metastasis, cell motility, and invasion, and its activation has also been shown to orchestrate escape from dormancy7,80,81. In light of these findings, multiple PI3K inhibitors are currently being evaluated in clinical trials for HER2 + and triple-negative breast cancers82. Moreover, our studies demonstrated that PI3K-AKT-mTOR inhibition impairs the release of pro-tumorigenic cytokines induced by Brd7 loss (see below), providing functional links between this canonical oncogenic signaling pathway and modulation of key immune cell compartments within the lung metastatic niche. Hedgehog signaling has also been implicated in breast cancer metastasis. Specifically, Sonic Hedgehog (SHH) overexpression in triple-negative breast cancer upregulates migration, invasion, and cell proliferation capacity in vitro83 and enhances spontaneous lung metastasis in vivo84. Moreover, canonical hedgehog signaling has been shown to enhance tumor angiogenesis85 and overexpression of hedgehog-associated genes is correlated with poor outcomes in breast cancer patients86. Additional pro-metastasis pathways like RAP1, MAPK, and Ras signaling were also enriched upon Brd7 loss. Finally, we found that Brd7 loss upregulated molecular networks mapping to neurogenesis and mammary gland development, both of which have been shown to facilitate metastasis in recent studies26,27,87,88.
The extent to which dormant cancer cells acclimatize to “foreign soil” is in large part dictated by their ability to evade immune recognition and culling. In this context, the consequences of tumor cell epigenetic reprogramming on the surrounding immune microenvironment represent an emerging field of study89,90. In this report, we demonstrate that loss of Brd7 influences multiple immune cell compartments to facilitate lung metastasis. We found that Brd7 loss leads to downregulation of MHC class 1-related genes and suppression of interferon response, both of which have previously been implicated as elements of tumor-induced immunosuppression91,92,93 while IFNγ has been shown to drive breast cancer dormancy94. Moreover, Brd7 loss promoted the release of pro-metastatic cytokines like CCL2, CCL5, CXCL10, and CCL20. Interestingly, these four cytokines appear to facilitate metastasis by crafting an immune suppressive microenvironment. CXCL10 and CCL20 have been shown to drive invasion and metastasis51,95 and both CCL5 and CCL2 promote angiogenesis, migration, invasion, and metastasis in a variety of tumor types96,97,98 particularly breast cancer99,100 With regards to effects on specific immune constituents, CCL5 has been demonstrated to regulate neutrophil trafficking to lung100, and CCL2 has been found to promote breast cancer lung metastasis by recruiting tumor-associated macrophages (TAMs)101. Similarly, CCL20 has been shown to activate tumor-associated neutrophils102. Interestingly, both CCL2 and CCL5 have been demonstrated to induce NETosis103,104.
To examine the extent to which Brd7 loss modulates immune microenvironmental cellular compartments, we performed extensive flow cytometric and single-cell transcriptomic studies in our dormancy-reactivation models. We found increased recruitment of pro-metastatic neutrophils, stress response CD4 + T cells, and exhausted CD8 + T cells upon Brd7 loss. Multiple studies have shown that neutrophils can facilitate breast cancer lung metastasis by promoting a favorable pre-metastatic niche53,54,71. Neutrophils have also been demonstrated to suppress T cell proliferation and impair TCR signaling78. Our analysis revealed recently identified metastasis-promoting and immunosuppressive inflammatory and transitional neutrophil populations to be enriched in the Brd7-KO context. Moreover, neutrophil recruitment in the setting of Brd7 deficiency was accompanied by the heightened formation of NETs, which have already been shown to promote metastasis, reawaken dormant cancer cells in mice, and drive T cell exhaustion77,105,106. Our scRNAseq profiling implicated the stress-response CD4 + T cell compartment in breast cancer lung metastatic reawakening and outgrowth. This T cell population is distinctly characterized by high levels of heat shock protein and is postulated to provoke immunotherapy resistance. Our findings also pointed to a role for CD8 + exhausted T cells in Brd7-deficient lung metastatic outgrowth and indicated that this cell type was at least partially induced by NETs. While we considered experimentally depleting the CD8 + T cell population in the context of our models to further probe functional relevance, we ultimately felt that such an approach would have complex effects, hampering their interpretation, as this lymphocytic compartment consists of both tumor-promoting (e.g., exhausted T cells) and -antagonistic subsets (e.g., Teff and Tmem cells). Accordingly, we cannot formally exclude that some subset of CD8 + T cells contributes to BRD7-deficient breast cancer lung metastasis, similarly to neutrophils. Nevertheless, targeting NETs and/or reinvigorating cytotoxic T cells through immune checkpoint inhibition with ipilimumab effectively abrogated lung metastases in our Brd7-deficient models, pointing to viable avenues for therapeutic advancement107,108. That anti-LAG3 treatment largely failed to impair lung metastasis to a similar degree as anti-CTLA4 in our experimental systems correlates with its underwhelming impact observed thus far as a single agent in human clinical trials.
Crucial elements of the mechanistic findings obtained from our in vitro and in vivo metastatic dormancy models were recapitulated in existing large-scale human datasets. Specifically, we found that reduced BRD7 and/or PBAF component expression correlated with a variety of unfavorable clinical metrics in multiple breast cancer cohorts, including metastasis-free and relapse-free survival. Associations with key immune cell populations, including neutrophils, cytotoxic T cells, and Tregs, were also validated.
In summary, we demonstrate that BRD7 loss and impaired PBAF chromatin remodeling complex functionality coordinates escape from metastatic dormancy to promote breast cancer lung metastatic outgrowth by way of immune microenvironmental modulation (Fig. 6J). Our study augments the understanding of epigenetic and microenvironmental mechanisms promoting relapse and metastatic outgrowth at distant organ sites, while also pointing to tractable strategies for effective therapeutic targeting. We identify BRD7 as a useful prognostic biomarker designating an increased risk of lung metastatic relapse in breast cancer patients. Moreover, we demonstrate that utilizing Sivelestat, neutrophil depletion, or immune checkpoint inhibition could therapeutically curb breast cancer lung metastasis. Accordingly, our findings speak to multiple basic and translational elements of cancer research and inform the advancement of clinical cancer management.
Methods
Animal studies
Mouse studies were conducted in accordance with protocols approved by the Institutional Animal Care and Use Committee of MD Anderson Cancer Center (MDACC). Six- to eight-week-old female mice were used at the commencement of all experiments. Female BALB/c, nude, and NOD/Scid mice were purchased from The Jackson Laboratory. Animals were housed at 20–22 °C on a 12 h light/12 h dark at 50–60% humidity. To examine lung colonization, the indicated cancer cells were resuspended in 100 µl of HBSS and injected into the lateral tail vein. All experiments were performed in syngeneic host mice with an intact immune system unless specifically indicated. Lungs were perfused and collected for FACS, IHC, or IF analysis. For orthotopic injection, 4T1 (5 × 105) cells were resuspended in Matrigel at a 1:1 ratio and injected into the fourth mammary gland. To examine spontaneous metastasis, primary tumors were surgically removed, and lung metastases were analyzed at different time points using the IVIS in vivo imaging system. Tumor growth was measured either by Vernier caliper or BLI with IVIS. Tumor volumes were analyzed using the formula V = (L x W x W)/2, where V: volume, L: length, W: width. BLI was used to examine metastatic outgrowth. For BRD7-KO/KD the lung metastatic phenotype was well-developed at 4 weeks, allowing this to serve as an endpoint for all downstream studies. Five weeks was used for the ICI studies to further document therapeutic effect, until the drug supply was depleted, and survival was further monitored until 50 days. A seven-week endpoint was used for the Smarcb1-KD and Pbrm1-KO study because the lung metastatic phenotype in these mice took longer to develop. For primary tumor growth studies, 1700 mm^3 was the pre-determined endpoint. The maximal IACUC-approved tumor volume was not exceeded. Metastatic lesions were confirmed by macroscopic and microscopic analyses. When indicated, 4T07-TGL cells were injected i.v. into nude and NOD scid mice. Lung colonization was monitored and assessed as described above.
Cell culture, plasmid construction, and lentivirus infection
4T1-CRL-2539 cells were obtained from ATCC and the HEK-293F (R79007) was obtained from Thermo-Fischer. 67NR, 168FARN, and 4T07 cell lines were generously provided by Dr. Fred R. Miller (Wayne State University, Detroit, MI), D2A1-m cell line was obtained from Dr. R.Weinberg, MIT (Derived from D2A1 which was obtained from Dr. Fred R. Miller) and the D2A1-d cell line was obtained from Dr. R.Weinberg, MIT (Derived from D2A1 which was obtained from Dr. Fred R. Miller). The mouse breast cancer cell lines and HEK293F cells were cultured in Dulbecco’s Modified Eagle Medium (Gibco) with 10% fetal bovine serum (FBS, Gibco) at 37 °C in 5% CO2. All cell lines used in this study were confirmed Mycoplasma-negative using the Universal Mycoplasma Detection Kit (ATCC 30-1012 K). Plasmids pCDH-CMV-MCS-EF1-Puro, lentiCRISPR v2, PSPAX2, and PMD2.G were purchased from Addgene. The mouse Brd7 coding sequence was cloned into the pCDH-CMV-MCS-EF1-Puro vector with the EcoRI/BamHI sites. A custom library consisting of 2410 shRNAs focused on chromatin remodeling enzymes were kindly provided by Giulio Draetta109. The Short hairpin RNA (shRNA) specific for mouse Brd7 and Smarcb1 were cloned into the miR-E backbone vector110. The target site sequences for each shRNA were as follows:
shBrd7-1: 5′- CCACATTATGACTCTACATTT-3′
shBrd7-4: 5′- GCAAAGATGATTCTGATTTAA-3′
shSmarcb1-5: 5′- GACAGCATCCTAGAGGATCAA-3′
shSmarcb1-6: 5′- TATGTTCCGAGGTTCTCTGTA-3′
The sgRNAs specific for indicated mouse genes were cloned into the lentiCRISPR v2 vector with standard procedure111. The target sequences are as follows:
sgBrd7-1: 5′- GGCGCAATAAAATCCGTCAC-3′
sgBrd7-2: 5′- CCTCAAAGTCGGGGGGAGCG-3′
sgPbrm1-1: 5′- TCAACCAGACTATTATGAAG-3′
sgPbrm1-2: 5′- GTTGTAGCCACAAATCCATC-3′
sgSmarcb1-1: 5′- AACTACCTGCGTATGTTCCG-3′
sgSmarcb1-2: 5′- CTAGTCGCCGCCAGAGTGAG-3′
For lentivirus production, HEK293F cells at 80–90% confluence were transfected with Pcdh-puro, lentiCRISPR v2-puro, or miE-shRNA-puro plasmid, along with packaging plasmids PSPAX2 and PMD2.G by Lipofectamine 3000 (Invitrogen). After 48 h the supernatant containing virus was collected and filtered through 0.45 μm filters to remove cell debris and stored at 4 °C. For lentivirus transduction, indicated cells at 30–40% confluence were incubated with the indicated viral supernatants with Polybrene at 8 μg/ml for 24 h. The successfully transduced cells were selected by 4 μg/mL puromycin 48 h after transduction.
In vivo epigenetic-focused shRNA library screen
The in vivo epigenetic focused loss-of-function screen was performed as previous described with modification13. Briefly, the dormant mouse breast cancer cell line 4T07-TGL was transduced with the 2.4K- Epigenetic-Focused shRNA library at 25% efficiency. After 48 hr puromycin selection, 300,000 cells were injected into 6 weeks old female syngeneic Balb/cJ mice via tail vein to allow most of the cells to seed in the lung. A total of 20 mice were injected in order to cover all the shRNAs. Lung metastases were monitored by bioluminescence imaging weekly. Individual lung metastasis nodules was isolated, and genomic DNA of the tumors was extracted using the DNeasy Blood & Tissue Kit (Qiagen). PCR was used to amplify the shRNA cassettes, and Sanger sequencing was performed to determine the shRNA barcode. Identified shRNAs were then individually transduced into 4T07-TGL cells to evaluate their effect on lung metastasis. The candidate shRNAs were ranked according to the ability to induce lung metastatic colonization. All mouse studies were conducted in accordance with protocols approved by the Institutional Animal Care and Use Committee of MD Anderson Cancer Center (MDACC).
Bioluminescent imaging
Mice were anesthetized and retro-orbitally injected with 1.5 mg of D-luciferin at the indicated times. Animals were imaged in an IVIS 100 chamber within 5 min of D-luciferin injection, and data were recorded using Living Image software (IVIS Spectrum). To measure lung metastasis, photon flux was calculated by drawing a region of interest encompassing the thorax of the mouse using the Living Image (by PerkinElmer) or Aura imaging (by Spectral Instruments imaging) software. After subtracting a background value obtained from a control mouse injected only with D-luciferin, photon flux was normalized to the value obtained 30 min after tumor cells injection.
Animal drug treatment
4T07-TGL-sgCtr and the 4T07-TGL-Brd7 KO cells were injected via the tail vein on day 0 through tail vein injection in the syngeneic Balb/cJ mice. For neutrophil depletion, the anti-Ly6G (BioXcell, clone 1A8) antibody or rat IgG control (I4131, Sigma) was administered to the mice (300 µg/mice) intraperitoneally on days -5, -3, -1, 1 and on every alternate day thereafter. To confirm neutrophil depletion, FACS analysis for neutrophils was performed on blood collected through retro-orbital puncture 14 days post cellular injection. For immune checkpoint therapy, the mice injected with the 4T07-TGL-Brd7 KO cells (Day 0), were treated with either the anti-CTLA4 ab alone (BioXcell, clone 9D9), anti-LAG3 alone (BioXcell, clone C9B7W) or the anti-CTLA4 plus anti-LAG3 combination intraperitoneally (300 µg/mice) on days 1,3,5 and on every alternate day thereafter until Week 5. The mice in the control group were administered (300 µg/mice) with mouse IgG2b isotype control (BioXcell, clone MPC-11) and the rat IgG1 isotype control (BioXcell, clone HRPN). For Sivelestat (MedChemExpress, HY17443) treatment, the mice injected with the 4T07-TGL-Brd7 KO cells (Day 0) were i.p. injected with Sivelestat (50 mg/kg) or DMSO control on days -5,-3,-1,1 and on every alternate day thereafter until Week 4. Bioluminescent imaging was performed every week starting from 30 mins after tumor cell injection. To evaluate the effect of Sivelestat on T cell exhaustion, mice were treated with Sivelestat (50 mg/kg) starting from week 2 post tumor cell injection, and on every other day thereafter until week 4.
Histology and immunostaining
Paraffin-embedded sections obtained from the mouse lung or human lung were subjected to Hematoxylin and Eosin (H&E) staining or Immunofluorescence (IF) and Immunochemistry staining according to standard paraffin-embedded tissue staining protocols. In brief, after lung tissue sections were dewaxed in xylene and rehydrated, antigen retrieval was carried out in 10 mM sodium citrate (pH 6.0) at 100 °C for 5 min using a microwave. The tissue sections were then blocked with 5% normal goat serum for 1 h at room temperature and further incubated with primary antibodies diluted in blocking buffer overnight at 4 °C. The following antibodies were used for staining: Brd7 (Brd7 (1:200; 15125, Cell Signaling Technology), Ki67 (1:800; 9129, Cell Signaling Technology), MPO (1:100; Human and Mouse MPO, Recombinant Anti-Myeloperoxidase antibody [EPR20257] (ab208670), NE (1:100; Human and Mouse neutrophil elastase, Neutrophil elastase Recombinant Rabbit Monoclonal Antibody (8X4E6), Catalog # MA5-42901, F4/80 [1:500; (D2S9R) XP ® Rabbit mAb: CST 70076 T] and Ctla4 (Human and Mouse CTLA4, Rabbit monoclonal, Recombinant Anti-mouse CTLA4 antibody[CAL49] (1:500; ab237712). Tissue sections were subsequently washed with PBS and incubated with appropriate secondary antibodies for 1 h at room temperature. DAPI (1:1000; 4083, Cell Signaling Technology) was used to mark the nuclei, and fluorescence images were captured by a Zeiss LSM 700 confocal microscope (Zeiss). All image quantification was performed using ImageJ software (version 1.8.0).
Colony formation assays
For 3D colony formation assays, 1 ml of 0.5% agar in DMEM was plated into 6-well culture plates and allowed to solidify at 37 °C for 30 mins. 1000 cells were then suspended in 1 ml 0.3% agar with complete growth medium and then supplied to the upper layer of the solidified 0.5% agar. Colony numbers were counted 14 days after seeding under a microscope, and clusters of >50 cells were defined as a colony112. Photos were taken and quantified by ImageJ.
Transwell migration assay
Transwell migration assays were performed using 24-well plates with 8-μm pore inserts (Corning). Cells were harvested, resuspended at 1 × 105 cells/mL in serum-free media, and 100 µL was added to the upper chamber. The lower chamber contained 600 µL of complete media with 10% FBS as a chemoattractant. After incubating for 12 h at 37 °C with 5% CO2, non-migrated cells on the upper surface were removed with a cotton swab. Migrated cells on the lower surface were fixed with 4% paraformaldehyde, stained with 0.1% crystal violet, and counted under a microscope.
Transwell invasion assay
Transwell invasion assays were conducted using 24-well plates with 8-μm inserts coated with Matrigel (50 µL per insert). After allowing the Matrigel to solidify, 1 × 105 cells were seeded in serum-free media in the upper chamber, with the lower chamber filled with complete media containing 10% FBS as a chemoattractant. After 24 h of incubation at 37 °C with 5% CO2, non-invaded cells were removed. Invaded cells on the lower surface were fixed, stained with crystal violet, and counted under a microscope.
Tumor sphere assays
Tumor sphere assays was performed as previously described14. Briefly, single-cell suspensions of parental or Brd7-KO 4T07-TGL cells were prepared, in 5000-cell aliquots, and seeded on ultra-low attachment 6-well plates (Corning Costar), supplemented with mammary epithelial growth medium (MEBM, BioWhittaker), 1:50 B27 (Invitrogen), 20 µg/ml EGF(PeproTech), 10 µg/ml bFGF (BD Biosciences), 4 μg/ml heparin, 4 μg/ml insulin, and 0.4% BSA (Sigma). After 7–9 days of culturing, photos were taken, and spheroids with a diameter of > 50 μm were quantified using ImageJ.
Western blots
Cells were harvested and lysed in ice-cold lysis buffer containing nonidet P-40, sodium deoxycholate, and Triton X-100, along with a cocktail of protease and phosphatase inhibitors. The protein concentration of the lysate was assessed using a BCA assay (Pierce). Equal amounts of protein were separated by SDS-PAGE and subsequently transferred to PVDF membranes. To block nonspecific interactions, the membrane was incubated with a blocking buffer containing nonfat milk. Primary antibodies targeting specific proteins of interest were applied, such as Brd7 (1:1000; 15125, Cell Signaling Technology), Beta-Actin (1:1000; 3700, Cell Signaling Technology) and RhoGDI (1:1000; 2564, Cell Signaling Technology), followed by corresponding HRP or fluorescence conjugated Goat anti-rabbit IgG and goat anti-mouse IgG secondary antibodies. Proteins were detected using an ECL substrate and visualized with a ChemiDoc imaging system (Bio-Rad).
Bulk RNA sequencing and data analysis
RNA-seq on 4T07-TGL cells with or without Brd7 KO was performed in the MD Anderson Advanced Technology Genomics Core (ATGC). STAR 2.4.2a was used to align the RNA-seq reads to the reference mouse genome (mm10) and to count the number of reads mapping to each gene in the Ensembl GRCm38.80 gene model. Differential expression analysis was performed using the DESeq2 package(v.1.10.0)in R (v.4.3.1;http://www.R-project.org/). Gene set enrichment analysis was performed using GSEA (v.3.0) incorporating gene sets from MSigDB (v.5.0). Normalized enrichment scores (NES) and FDR/q values were used to explore enriched pathways in each phenotype.
Single-cell RNA-seq and data analysis
Single-cell RNA-seq was performed using 10x genomics platform. Mouse lung tissues were harvested and dissociated to single cells. Live CD45-positive immune cells were sorted and resuspended at a concentration of 1 × 106 per ml in PBS with 0.4% BSA for 10x genomics processing. Approximately 10,000 cells were then loaded onto a 10x Genomics Chromium Chip to generate single-cell gel beads in emulsion (GEM). scRNA-seq libraries were prepared according to the Single Cell 3′ Reagent Kits v2 User Guide. The library was then sequenced on an Illumina platform (HiSeq 4000) to yield 2 × 150 bp paired-end reads, achieving sequencing depth at approximately >90% sequencing saturation. Raw gene expression matrices were generated for each sample by the Cell Ranger (v.3.0.2). Quality control was then performed by calculating the number of genes, UMIs, and the proportion of mitochondrial genes using Seurat R package V3 (https://satijalab.org/seurat/), and cells with a low number of covered genes (gene count <500) and high mitochondrial counts (mt-genes >0.1) were excluded. Gene expression matrices were normalized by the NormalizeData function, and 2000 features with high cell-to-cell variation were calculated using the FindVariableFeatures function. For dataset normalization and dimensionality reduction, the RunPCA function was conducted with default parameters on linear-transformation scaled data generated by the ScaleData function. Next, the ElbowPlot, DimHeatmap, and JackStrawPlot functions were used to identify the true dimensionality of each dataset, as recommended by the Seurat developers. Finally, we clustered cells using the FindNeighbors and FindClusters functions and performed nonlinear dimensional reduction with the RunUMAP function using default settings. Marker genes were determined by previous published database and ImmGen database (https://www.immgen.org/). Gene set enrichment analysis (GSEABase_1.68.0) was performed using gene sets from MSigDB (v.5.0) to identify gene sets and specific biological processes that are significantly enriched in different groups. Normalized enrichment scores (NES) and FDR/q values were used to explore enriched pathways for each phenotype.
ATAC-seq and data analysis
For ATAC-seq, freshly harvested cells were directly sent to the MD Anderson Advanced Technology Genomics Core for library construction and sequencing. Raw ATAC-seq reads were processed through a standard bioinformatics pipeline. Initial quality control was performed using FastQC, followed by adapter trimming with Trimmomatic. Reads were aligned to the mouse genome (mm10) using the Bowtie2 (v2.2.4) and duplicate reads were removed using Picard tools (v.1.126) (http://broadinstitute.github.io/picard/). Peak calling was conducted with MACS2, utilizing a q-value cutoff of 0.01. Read counts per peak were quantified with BEDTools. Differential accessibility analysis was performed using DESeq2 or EdgeR to identify significant differences between conditions. ChIPseeker (v1.8.0) R package was used for peak annotations, and motif discovery was performed using HOMER (v4.10). ChIPseeker was used for TSS site visualizations and quality controls. KEGG pathway analysis and Gene Ontology (GO) analysis was performed using the clusterProfiler R package (v3.0.0). Data visualization and generating plots were performed in the R environment (v4.3.1) (https://www.r-project.org/).
ChIP-seq and data analysis
ChIP-seq was conducted on 4T07-TGL cells, parental or Brd7-KO, by utilizing the SimpleChIP® Enzymatic Chromatin IP Kit (Magnetic Beads) (9003, Cell Signaling Technologies) according to established protocols. The H3K27Ac Rabbit monoclonal antibody (1:100; 8173, Cell Signaling Technologies) was applied for chromatin immunoprecipitation. Purified ChIP DNA was sent to the MD Anderson Advanced Technology Genomics Core for library preparation and sequencing. Quality control of raw sequencing data was performed by FastQC v0.11.9. Cleaned reads are aligned to the reference genome by Bowtie 2 (version 2.2.4). Duplicate reads were removed with Picard v1.126. Low-quality mapped reads (mapping quality <20) are filtered out to reduce noise. Peaks were identified using MACS (version 1.4.2). Normalized BigWig files were generated by BEDTools (version 2.17.0) as well as the bedGraphToBigWig tool (version 4). Differential peak analysis was carried out using DESeq2. ChIPseeker (version 1.8.0) in R was used to annotate identified peaks and visualize transcription start sites, and motif discovery was performed by HOMER (version 4.10). Gene Ontology analyses were performed with the clusterProfiler R package (version 3.0.0), and plots generated by ggplot2 in R.
NET quantification using Sytox Orange
Neutrophils were isolated from blood obtained through intracardiac puncture from 8-weeks old female Balb/cJ mice using an immunomagnetic negative selection kit- EasySepTM Mouse Neutrophil enrichment kit (19762, Stemcell Technologies) as per the manufacturer’s protocol. The 4T07-Brd7 KO and the 4T07-TGL cells were seeded at a density of 2500 cells/well in a 96-well Greiner flat-bottom black microplate in phenol-red free DMEM supplemented with 10% FBS and 1% penicillin/streptomycin. The isolated neutrophils were co-cultured with the Brd7 KO and the 4T07-TGL cells at a ratio of 100:1(neutrophils: cancer cells) with 250 K neutrophils/well. The neutrophil- cancer cells co-cultures were incubated for 16 h at 37 °C in 5% CO2, following which the NET quantification with Sytox Orange staining was performed as previously described73. The fluorescence intensity was measured with the BMG Labtech FLUOstar Omega microplate reader at excitation and emission wavelengths of 544 nm and 590 nm, respectively.
Human samples and ethics statement
Human tumor samples of a 57-year-old Asian female patient with breast cancer (primary and metastasis tumor) were obtained from Bioaitech China, with written informed consent obtained from the patient. The study was approved by the local ethics committee of the Guangzhou National Laboratory.
Analysis of public databases
TCGA data were sorted based on BRD7 expression and stratified into quarters. To estimate the immune cell abundance, we utilized the ImmuCellAI tool, correlating with the top and bottom BRD7-expressing quartiles (http://bioinfo.life.hust.edu.cn/ImmuCellAI#!/)113,114. Distant metastasis free survival (DMFS), and Relapse free survival (RFS), stratified by BRD7 mRNA expression, were presented as Kaplan–Meier plots and tested for significance using log-rank tests115.
FACS Analysis and antibodies
To profile lung immunocytes, mice were perfused with 10 ml cold 1X PBS by cardiac puncture, and lungs collected in ice-cold D-PBS. Lungs were then digested in collagenase D (Roche Life Science) and single-cells suspensions prepared as described previously116. Cells were spun down at 400 g for 5 min and resuspended in 1 ml fluorescence-activated cell sorting (FACS) buffer (D-PBS without calcium and magnesium containing 0.5% FBS, 2 mM EDTA, and 0.09% sodium azide). For tissue flow cytometry analysis, cells were blocked with rat monoclonal anti-CD16/CD32 (Fc block antibody, BD Biosciences) in PBS for 30 min at 4 °C. Cells were then stained for 1 h with the FACS antibody cocktail in the dark at 4 °C. The cells were then washed 3 times by centrifugation at 400 × g for 5 mins each at 4 °C, after which the pellet was resuspended in 500 µl of FACS buffer. Cell suspension samples were passed through a 30 μm cell strainer and subjected to flow cytometry on LSRFortessa X-20. Data were analyzed using FlowJo software. Antibodies, including anti-CD45-APC/Cy7 (1:50; clone 30-F11), CD3-Brilliant Violet 510 (1:50; clone 17A2), CD11b-Brilliant Violet 421 (1:100; clone M1/70), CD4-PerCP/Cyanine5.5 (1:100; clone GK1.5), CD8-Brilliant Violet 650 (1:200; clone 53-6.7), Ly6G-PE/Cy7 (1:100; clone 1A8), Ly6C-APC (1:100; clone HK1.4), F4/80-PE (1:50; clone BM8), CD49b-PE/Dazzle (1:100; clone DX5), CD19 (1:100; Brilliant Violet 711, clone 6D5), CD11c-Brilliant Violet 605 (1:100; clone N418), and Fixable Viability Dye Zombie UV™ were purchased from Biolegend. For T cell exhaustion FACS analysis, the antibodies used were PE anti-mouse CD152 (1:100; clone UC10-4B9)-BioLegend Cat# 106305; and APC anti-mouse CD279 (PD-1), (1:100; clone 29 F.1A12)-BioLegend Cat#135209.
Luminex analyses
The BioTechne Mouse Luminex discovery assay was employed for Luminex profiling of 40 cytokines. The 4T07-TGL-sgCtr, 4T07-TGL-Brd7 KO clone 1 and 4T07-TGL-Brd7 KO clone 2 cells were seeded on day 0 in DMEM + 10% FBS on 6-well plates at a density of 0.2 × 106 in triplicate. On day 1, the cell culture media was replaced with DMEM media containing 0.5% FBS. After 24 h, supernatants were collected and centrifuged at 15000 × g for 10 mins at 4 oC before analysis on the Luminex discovery assay using the manufacturer’s protocol. For studies involving inhibitor treatment, cells were treated with varying drug concentrations ranging from 0.1 µm, 1 µm, and 10 µm of LY294002 (MedChemExpress, pan-PI3K inhibitor) for 5 h (CCL2 and CXCL10) and for 15 h (CCL20). Media containing drugs were then removed and replaced with DMEM media containing 0.5% FBS. Supernatants were then collected and processed as described above.
Quantitative RT-PCR
Total RNA was extracted from cells using the RNeasy Mini Kit (Qiagen), and cDNA was generated with the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Quantitative PCR was performed using SYBR Green PCR Master Mix (Applied Biosystems), with transcript levels normalized to Gapdh. All samples were run in triplicate. The primer sequences were as follows:
mGapdh-F: 5′-AGGTCGGTGTGAACGGATTTG-3′
mGapdh-R: 5′-TGTAGACCATGTAGTTGAGGTCA-3′
mArid2-F: 5′- TGGATCTTCACGGGCTCTACA-3′
mArid2-R: 5′- CGTTGGAACAACTTCTGGGAAA-3′
mBrd7-F: 5′- AAGCGACCATGACAAACACAA-3′
mBrd7-R: 5′- ATCCTCCTTGACTCTTCTCCG-3′
mPbrm1-F:5′- CTGTAGATCCTATTGCTGTGTGC-3′
mPbrm1-R:5′- GCTTTGGAGCCCTAATGAACA-3′
mSmarcb1-F: 5′- TCCGAGGTGGGAAACTACCTG-3′
mSmarcb1-R: 5′- CAGAGTGAGGGGTATCTCTTGT-3′
mSmarcc2-F: 5′- AACCGCCAACCAACAAGTCT-3′
mSmarcc2-R:5′- AGGAAACATTTGATCGGCAGT-3′
mSmarcd2-F:5′- CCAGCGCCGAGGGTTAAAG-3′
mSmarcd2-R:5′- GCTTCCTCTCGAAAGCTAAAAGA-3′
mSmarce1-F:5′- AAAAGACCATCTTATGCCCCAC-3′
mSmarce1-R:5′- CCTGTAGTTGTTGTAGGCGAG-3′
mArid1a-F:5′- CTTCCCCAACCACCAGTACAA-3′
mArid1a-R:5′- CTGTGCGAAGGACGAAGAC-3′
mArid1b-F:5′- CGTGCGGAGCTTGTCTTTC-3′
mArid1b-R:5′- CCTCCTTCTCATAGGTCTGTGG-3′
mDpf2-F:5′- AGCTCCTTGGCGAGCAATAC-3′
mDpf2-R:5′- CTCTGGGCTACTCCAGTCTGT-3′
mActl6a-F:5′- GTGTACGGCGGAGATGAAGTT-3′
mActl6a-R:5′- GGGAAATCAACCTTAGGGCAGT-3′
mBcl7c-F:5′- GCGGTACTGATGAACCCCC-3′
mBcl7c-R:5′- GACAGATGGCACTGGCTCAA-3′
mSmarca2-F:5′- CTCCTGGACCAATTCTGGGG-3′
mSmarca2-R:5′- CATCGTTGACAGAGGATGTGAG-3′
mSmarca4-F:5′- CAAAGACAAGCATATCCTAGCCA-3′
mSmarca4-R:5′- CACGTAGTGTGTGTTAAGGACC-3′
mSs18-F:5′- AGGGGCACAGCATTACCAAG-3′
mSs18-R:5′- TCTGTAGGGAGGCATCTGTCG-3′
Statistics and reproducibility
All statistical analyses not previously described were carried out using GraphPad Prism 8 and R. No statistical method was used to predetermine sample size. For animal experiments, each mouse was counted as a biologically independent sample. The mice were randomized prior to commencing an in vivo experiment. The investigators were not blinded to allocation during the experimental procedures and during outcome assessments. Data were analyzed by unpaired Student t test, unless mentioned otherwise. Survival was measured according to the Kaplan–Meier method and analyzed by log-rank (Mantel–Cox) test. P < 0.05 was considered significant. Error bars represent standard error of the mean (SEM) or standard deviation (SD), unless mentioned otherwise.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Bulk RNA-seq, ATAC-seq, ChIP-seq, and scRNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession codes GSE267197, GSE262165, GSE260497 and GSE261385. All data are available in the main text or the Supplementary Information. Source data are provided in this paper.
References
Seyfried, T. N. & Huysentruyt, L. C. On the origin of cancer metastasis. Crit. Rev. Oncog. 18, 43–73 (2013).
Fares, J. et al. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct. Target. Ther. 5, 28 (2020).
Łukasiewicz, S. et al. Breast Cancer-Epidemiology, Risk Factors, Classification, Prognostic Markers, and Current Treatment Strategies-An Updated Review. Cancers 13, 4287 (2021).
Jin, L. et al. Breast cancer lung metastasis: Molecular biology and therapeutic implications. Cancer Biol. Ther. 19, 858–868 (2018).
Xiao, W. et al. Risk factors and survival outcomes in patients with breast cancer and lung metastasis: a population-based study. Cancer Med. 7, 922–930 (2018).
Giancotti, F. G. Mechanisms governing metastatic dormancy and reactivation. Cell 155, 750–764 (2013).
Sosa, M. S., Bragado, P. & Aguirre-Ghiso, J. A. Mechanisms of disseminated cancer cell dormancy: an awakening field. Nat. Rev. Cancer. 14, 611–622 (2014).
Goddard, E. T. et al. Dormant tumour cells, their niches and the influence of immunity. Nat. Cell Biol. 20, 1240–1249 (2018).
Hosseini, H. et al. Early dissemination seeds metastasis in breast cancer. Nature 540, 552–558 (2016).
Baylin, S. B. & Jones, P. A. Epigenetic determinants of cancer. Cold Spring Harb. Perspect. Biol. 8, a019505 (2016).
Robinson, N. J., Parker, K. A. & Schiemann, W. P. Epigenetic plasticity in metastatic dormancy: mechanisms and therapeutic implications. Ann. Transl. Med. 8, 903 (2020).
Massagué, J. & Ganesh, K. Metastasis-initiating cells and ecosystems. Cancer Discov. 11, 971–994 (2021).
Gao, H. et al. Multi-organ site metastatic reactivation mediated by non-canonical discoidin domain receptor 1 signaling. Cell. 166, 47–62 (2016).
Gao, H. et al. The BMP inhibitor Coco reactivates breast cancer cells at lung metastatic sites. Cell. 150, 764–779 (2012).
Aslakson, C. J. & Miller, F. R. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 52, 1399–1405 (1992).
Kumar, D., et al. LncRNA Malat1 suppresses pyroptosis and T cell-mediated killing of incipient metastatic cells. Nat. Cancer 5, 262–282 (2024).
Reisman, D., Glaros, S. & Thompson, E. The SWI/SNF complex and cancer. Oncogene 28, 1653–1668 (2009).
Lu, P. & Roberts, C. W. The SWI/SNF tumor suppressor complex: Regulation of promoter nucleosomes and beyond. Nucleus 4, 374–378 (2013).
Wilson, B. G. & Roberts, C. W. SWI/SNF nucleosome remodellers and cancer. Nat. Rev. Cancer 11, 481–492 (2011).
Mashtalir, N. et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell 175, 1272–1288 (2018).
Nielsen, T. O. et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin. Cancer Res. 16, 5222–5232 (2010).
Siegel, M. B. et al. Integrated RNA and DNA sequencing reveals early drivers of metastatic breast cancer. J. Clin. Invest. 128, 1371–1383 (2018).
Riobo-Del Galdo, N. A., Lara Montero, Á. & Wertheimer, E. V. Role of hedgehog signaling in breast cancer: Pathogenesis and therapeutics. Cells. 8, 375 (2019).
Ye, X. et al. Upholding a role for EMT in breast cancer metastasis. Nature 547, E1–E3 (2017).
Paplomata, E. & O’Regan, R. The PI3K/AKT/mTOR pathway in breast cancer: targets, trials and biomarkers. Ther. Adv. Med. Oncol. 6, 154–166 (2014).
Zhao, Q. et al. The clinicopathological significance of neurogenesis in breast cancer. BMC Cancer 14, 484 (2014).
Wu, D., Thompson, L. U. & Comelli, E. M. MicroRNAs: A link between mammary gland development and breast cancer. Int. J. Mol. Sci. 23, 15978 (2022).
Roussos, E. T., Condeelis, J. S. & Patsialou, A. Chemotaxis in cancer. Nat. Rev. Cancer 11, 573–587 (2011).
Shankaran, V. et al. IFNgamma and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 410, 1107–1111 (2001).
Dunn, G. et al. Cancer immunoediting: from immunosurveillance to tumor escape. Nat. Immunol. 3, 991–998 (2002).
Peng, C. et al. The transcriptional regulation role of BRD7 by binding to acetylated histone through bromodomain. J. Cell Biochem. 97, 882–892 (2006).
Sun, H. et al. Solution structure of BRD7 bromodomain and its interaction with acetylated peptides from histone H3 and H4. Biochem. Biophys. Res. Commun. 358, 435–441 (2007).
Kharel, A. et al. Loss of PBAF promotes expansion and effector differentiation of CD8+ T cells during chronic viral infection and cancer. Cell Rep. 42, 112649 (2023).
Baxter, A. E. et al. The SWI/SNF chromatin remodeling complexes BAF and PBAF differentially regulate epigenetic transitions in exhausted CD8+ T cells. Immunity. 56, 1320–1340 (2023).
Wei, L. et al. Cancer-associated fibroblasts-mediated ATF4 expression promotes malignancy and gemcitabine resistance in pancreatic cancer via the TGF-β1/SMAD2/3 pathway and ABCC1 transactivation. Cell Death Dis. 12, 334 (2021).
Heeg, S. et al. AK. ETS-Transcription factor ETV1 regulates stromal expansion and metastasis in pancreatic cancer. Gastroenterology 151, 540–553 (2016).
Okita, Y. et al. The transcription factor MAFK induces EMT and malignant progression of triple-negative breast cancer cells through its target GPNMB. Sci. Signal. 10, eaak9397 (2017).
Adamo, P. & Ladomery, M. The oncogene ERG: a key factor in prostate cancer. Oncogene 35, 403–414 (2016).
Aytes, A. et al. ETV4 promotes metastasis in response to activation of PI3-kinase and Ras signaling in a mouse model of advanced prostate cancer. Proc. Natl. Acad. Sci. USA 110, E3506–E3515 (2013).
Xu, H., Zhang, Y., Peña, M. M., Pirisi, L. & Creek, K. E. Six1 promotes colorectal cancer growth and metastasis by stimulating angiogenesis and recruiting tumor-associated macrophages. Carcinogenesis 38, 281–292 (2017).
Tomar, S. et al. ETS1 induction by the microenvironment promotes ovarian cancer metastasis through focal adhesion kinase. Cancer Lett. 414, 190–204 (2018).
Liu, M., Hu, W., Meng, X. & Wang, B. TEAD4: A key regulator of tumor metastasis and chemoresistance - Mechanisms and therapeutic implications. Biochim. Biophys. Acta Rev. Cancer 1879, 189050 (2024).
Wutschka, J. et al. JUNB suppresses distant metastasis by influencing the initial metastatic stage. Clin. Exp. Metastasis 38, 411–423 (2021).
Kelso, T. W. R. et al. Chromatin accessibility underlies synthetic lethality of SWI/SNF subunits in ARID1A-mutant cancers. Elife. 6, e30506 (2017).
Alver, B. H. et al. The SWI/SNF chromatin remodelling complex is required for maintenance of lineage specific enhancers. Nat. Commun. 8, 14648 (2017).
Costoya, J. A. & Arce, V. M. Cancer cells escape the immune system by increasing stemness through epigenetic reprogramming. Cell Mol. Immunol. 20, 6–7 (2023).
Cao, J. & Yan, Q. Cancer Epigenetics, Tumor Immunity, and Immunotherapy. Trends Cancer 6, 580–592 (2020).
Kim, M. et al. Role of CXCL10 in the progression of in situ to invasive carcinoma of the breast. Sci. Rep. 11, 18007 (2021).
Khalid, A. et al. Recent advances in discovering the role of CCL5 in metastatic breast cancer. Mini. Rev. Med. Chem. 15, 1063–1072 (2015).
Kitamura, T. et al. CCL2-induced chemokine cascade promotes breast cancer metastasis by enhancing retention of metastasis-associated macrophages. J. Exp. Med. 212, 1043–1059 (2015).
Kadomoto, S., Izumi, K. & Mizokami, A. The CCL20-CCR6 Axis in Cancer Progression. Int. J. Mol. Sci. 21, 5186 (2020).
Wu, L. et al. Breast cancer cell-neutrophil interactions enhance neutrophil survival and pro-tumorigenic activities. Cancers 12, 2884 (2020).
Gong, Zheng et al. Immunosuppressive reprogramming of neutrophils by lung mesenchymal cells promotes breast cancer metastasis. Sci. Immunol. 8, eadd5204 (2023).
Xiao, Y. et al. Cathepsin C promotes breast cancer lung metastasis by modulating neutrophil infiltration and neutrophil extracellular trap formation. Cancer Cell. 39, 423–437 (2021).
McGinnis, C. S. et al. The temporal progression of lung immune remodeling during breast cancer metastasis. Cancer Cell 42, 1018–1031 (2024).
Jablonska, J., Lang, S., Sionov, Vogt & Granot, R. Z. The regulation of pre-metastatic niche formation by neutrophils. Oncotarget. 8, 112132–112144 (2017).
Kaltenmeier, C., Simmons, R. L., Tohme, S. & Yazdani, H. O. Neutrophil extracellular traps (NETs) in cancer metastasis. Cancers 13, 6131 (2021).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147 (2018).
Wang, J. J. et al. Infiltration of T cells promotes the metastasis of ovarian cancer cells via the modulation of metastasis-related genes and PD-L1 expression. Cancer Immunol. Immunother. 69, 2275–2289 (2020).
Haj-Shomaly, J. et al. T Cells promote metastasis by regulating extracellular matrix remodeling following chemotherapy. Cancer Res. 82, 278–291 (2022).
Kos, K. et al. Tumor-educated Tregs drive organ-specific metastasis in breast cancer by impairing NK cells in the lymph node niche. Cell Rep. 38, 110447 (2022).
Huppert, L. A. et al. Tissue-specific Tregs in cancer metastasis: opportunities for precision immunotherapy. Cell Mol. Immunol. 19, 33–45 (2022).
Karpisheh, V. et al. The role of Th17 cells in the pathogenesis and treatment of breast cancer. Cancer Cell Int. 22, 108 (2022).
Coffelt, S. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Kravtsov, D. S., Erbe, A. K., Sondel, P. M. & Rakhmilevich, A. L. Roles of CD4+ T cells as mediators of antitumor immunity. Front. Immunol. 13, 972021 (2022).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Pagès, F. et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N. Engl. J. Med. 353, 2654–2666 (2005).
Christian, L. S. et al. Resident memory T cells in tumor-distant tissues fortify against metastasis formation. Cell Rep. 35, 109118 (2021).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Jiang, Y., Li, Y. & Zhu, B. T-cell exhaustion in the tumor microenvironment. Cell Death Dis. 6, e1792 (2015).
Wculek, S. K. & Malanchi, I. Neutrophils support lung colonization of metastasis-initiating breast cancer cells. Nature. 528, 413–417 (2015).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Jiang, D., Saffarzadeh, M. & Scharffetter-Kochanek, K. In vitro demonstration and quantification of neutrophil extracellular trap formation. Bio. Protoc. 7, e2386 (2017).
Jiang, D. et al. Suppression of neutrophil-mediated tissue damage-a novel skill of mesenchymal stem cells. Stem Cells 34, 2393–2406 (2016).
Yost, C. C. et al. Impaired neutrophil extracellular trap (NET) formation: a novel innate immune deficiency of human neonates. Blood 113, 6419–6427 (2009).
Bruns, S. et al. Production of extracellular traps against Aspergillus fumigatus in vitro and in infected lung tissue is dependent on invading neutrophils and influenced by hydrophobin RodA. PLoS Pathog. 6, e1000873 (2010).
Kaltenmeier, C. et al. Neutrophil extracellular traps promote T cell exhaustion in the tumor microenvironment. Front. Immunol. 12, 785222 (2021).
Bert, S., Nadkarni, S. & Perretti, M. Neutrophil-T cell crosstalk and the control of the host inflammatory response. Immunol. Rev. 314, 36–49 (2023).
Hüsemann, Y. et al. Systemic spread is an early step in breast cancer. Cancer Cell 13, 58–68 (2008).
Miricescu, D. et al. PI3K/AKT/mTOR Signaling pathway in breast cancer: From molecular landscape to clinical aspects. Int. J. Mol. Sci. 22, 173 (2020).
Correa, R. J., Peart, T., Valdes, Y. R., DiMattia, G. E. & Shepherd, T. G. Modulation of AKT activity is associated with reversible dormancy in ascites-derived epithelial ovarian cancer spheroids. Carcinogenesis 33, 49–58 (2012).
Ma, F. & Xu, B. Expert consensus on the clinical application of PI3K/AKT/mTOR inhibitors in the treatment of advanced breast cancer. Cancer Innovation. 1, 25–54 (2022).
Habib, J. G. & O’Shaughnessy, J. A. The hedgehog pathway in triple-negative breast cancer. Cancer Med. 5, 2989–3006 (2016).
Harris, L. G., Pannell, L. K., Singh, S., Samant, R. S. & Shevde, L. A. Increased vascularity and spontaneous metastasis of breast cancer by hedgehog signaling mediated upregulation of cyr61. Oncogene. 31, 3370–3380 (2012).
Chen, W. et al. Canonical hedgehog signaling augments tumor angiogenesis by induction of VEGF-A in stromal perivascular cells. Proc. Natl. Acad. Sci. USA 108, 9589–9594 (2011).
Han, B. et al. FOXC1 activates smoothened-independent Hedgehog signaling in basal-like breast cancer. Cell Rep. 13, 1046–1058 (2015).
Cervantes-Villagrana, R. D., Albores-García, D., Cervantes-Villagrana, A. R. & García-Acevez, S. J. Tumor-induced neurogenesis and immune evasion as targets of innovative anti-cancer therapies. Signal Transduct. Target. Ther. 5, 99 (2020).
Wiseman, B. S. & Werb, Z. Stromal effects on mammary gland development and breast cancer. Science. 296, 1046–1049 (2002).
Garcia-Fabiani, Maria B. et al. “Genetic alterations in gliomas remodel the tumor immune microenvironment and impact immune-mediated therapies.” Front. Oncol. 11, https://doi.org/10.3389/fonc.2021.631037 (2021).
Tomasi, T. B., Magner, W. J. & Khan, A. N. H. Epigenetic regulation of immune escape genes in cancer. Cancer Immunol. Immunother. 55, 1159–1184 (2006).
Taylor, B. C. & Balko, J. M. Mechanisms of MHC-I downregulation and role in immunotherapy response. Front. Immunol. 13, 844866 (2022).
Dhatchinamoorthy, K., Colbert, J. D. & Rock, K. L. Cancer immune evasion through loss of MHC class I antigen presentation. Front. Immunol. 12, 636568 (2021).
von Locquenghien, M., Rozalén, C. & Celià-Terrassa, T. Interferons in cancer immunoediting: sculpting metastasis and immunotherapy response. J. Clin. Invest. 131, e143296 (2021).
Correia, A. L. et al. Hepatic stellate cells suppress NK cell-sustained breast cancer dormancy. Nature 594, 566–571 (2021).
Liu, M., Guo, S. & Stiles, J. K. The emerging role of CXCL10 in cancer (Review). Oncol. Lett. 2, 583–589 (2011).
Aldinucci, D., Borghese, C. & Casagrande, N. The CCL5/CCR5 axis in cancer progression. Cancers 12, 1765 (2020).
Liu, W. et al. CC Chemokine 2 promotes ovarian cancer progression through the MEK/ERK/MAP3K19 signaling pathway. Int. J. Mol. Sci. 24, 10652 (2023).
Qiu, J. et al. CCL5 mediates breast cancer metastasis and prognosis through CCR5/Treg cells. Front. Oncol. 12, 972383 (2022).
Bonapace, L. et al. Cessation of CCL2 inhibition accelerates breast cancer metastasis by promoting angiogenesis. Nature 515, 130–133 (2014).
Hwaiz, R., Rahman, M., Syk, I., Zhang, E. & Thorlacius, H. Rac1-dependent secretion of platelet-derived CCL5 regulates neutrophil recruitment via activation of alveolar macrophages in septic lung injury. J. Leukoc. Biol. 97, 975–984 (2015).
Yoshimura, T., Li, C., Wang, Y. & Matsukawa, A. The chemokine monocyte chemoattractant protein-1/CCL2 is a promoter of breast cancer metastasis. Cell Mol. Immunol. 20, 714–738 (2023).
Kwantwi, L. B. et al. Tumor-associated neutrophils activated by tumor-derived CCL20 (C-C motif chemokine ligand 20) promote T cell immunosuppression via programmed death-ligand 1 (PD-L1) in breast cancer. Bioengineered. 12, 6996–7006 (2021).
Hofbauer, T. M. et al. Neutrophil extracellular traps induce MCP-1 at the culprit site in ST-segment elevation myocardial infarction. Front. Cell Dev. Biol. 8, 564169 (2020).
Rossaint, JanM. et al. Synchronized integrin engagement and chemokine activation is crucial in neutrophil extracellular trap–mediated sterile inflammation. Blood 123, 2573–2584 (2014).
Park, J. et al. Cancer cells induce metastasis-supporting neutrophil extracellular DNA traps. Sci. Transl. Med. 8, 361ra138 (2016).
Albrengues, J. et al. Neutrophil extracellular traps produced during inflammation awaken dormant cancer cells in mice. Science 361, eaao4227 (2018).
Jia, W., Mao, Y., Luo, Q., Wu, J. & Guan, Q. Targeting neutrophil elastase is a promising direction for future cancer treatment. Discov. Oncol. 15, 167 (2024).
Zhang, H. et al. Recent research and clinical progress of CTLA-4-based immunotherapy for breast cancer. Front. Oncol. 13, 1256360 (2023).
Carugo, A. et al. In vivo functional platform targeting patient-derived Xenografts identifies WDR5-Myc association as a critical determinant of pancreatic cancer. Cell Rep. 16, 133–147 (2016).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Hines, D. L. Lipid accumulation and production of colony-stimulating activity by the 266AD cell line derived from mouse bone marrow. Blood 61, 397–402 (1983).
Miao, Y. R. et al. ImmuCellAI-mouse: a tool for comprehensive prediction of mouse immune cell abundance and immune microenvironment depiction. Bioinformatics 38, 785–791 (2022).
Miao, Y. R. et al. ImmuCellAI: A unique method for comprehensive T-cell subsets abundance prediction and its application in cancer immunotherapy. Adv. Sci. 7, 1902880 (2020).
Gyorffy, B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput. Struct. Biotechnol. J. 19, 4101–4109 (2021).
Sheng, J., Ruedl, C. & Karjalainen, K. Most tissue-resident macrophages except microglia are derived from fetal hematopoietic stem cells. Immunity. 43, 382–393 (2015).
Acknowledgements
This study is dedicated to the memory of our mentor, Dr. Filippo Giancotti, who was a pioneer in the field of cancer metastasis and who shall forever be remembered for both his scientific brilliance and compassion for patients. This work was supported by NIH Outstanding Investigator Award R35 CA197566 and partially by CPRIT Recruitment of Established Investigators Award RR160031 (F.G.G.), and the Young Scientists Program of Guangzhou Laboratory, Grant No. QNPG23-17 (J.Z.). J.M. is supported by the Cancer Prevention and Research Institute of Texas (CPRIT) Graduate Scholar Research Training Grant (RP210028) and the Schissler Foundation Fellowship. The use of Core facilities - The Flow Cytometry and Cellular Imaging Core Facility, the Orion Core, the Research Histology Core Laboratory, and the Small Animal Imaging Facility -were supported by P30 CA016672 (MDACC). The Advanced Technology Genomics core was also supported by NIH 1S10OD024977-01 (MDACC).
Author information
Authors and Affiliations
Contributions
F.G.G. and J.T.H. designed the entire study and directed the project. J.M. and J.Z. designed, performed, and analyzed most of the experiments. F.Q., S.L., and D.K. helped in some of the in vivo experiments and IHC staining. J.M. and J.T.H. wrote the initial manuscript and J.M., J.Z., and J.T.H. revised and finalized the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All the authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ana Correia, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mondal, J., Zhang, J., Qing, F. et al. Brd7 loss reawakens dormant metastasis initiating cells in lung by forging an immunosuppressive niche. Nat Commun 16, 1378 (2025). https://doi.org/10.1038/s41467-025-56347-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56347-2
This article is cited by
-
Remodeling the tumor dormancy ecosystem to prevent recurrence and metastasis
Signal Transduction and Targeted Therapy (2026)