Abstract
Affinity labeling of biomacromolecules is vital for bioimaging and functional studies. However, affinity probes recognizing glycans with high specificity remain scarce. Here we report the development of glycan recombinant affinity binders (GRABs) based on mutant bacterial sialidases, which are enzymatically inactive but preserve stringent specificity for sialoglycan substrates. By mutating a key catalytic residue of Streptococcus pneumoniae neuraminidase A (SpNanA) and Ruminococcus gnavus neuraminidase H (RgNanH), we develop GRAB-Sia and GRAB-Sia3 recognizing total sialoglycans and α2,3-sialosides, respectively. The GRABs exhibit strict substrate and linkage specificity, and tetramerization with streptavidin substantially increases their avidity. The GRABs and tetrameric GRABs (tetra-GRABs) are effective tools for probing sialoglycans in immunoblotting, flow cytometry, immunoprecipitation, and fluorescence imaging. Furthermore, multiplex analysis with tetra-GRABs uncovers spatially distinct sialoglycans in the various mouse organs. This work provides a versatile toolkit for labeling and analyzing sialoglycans with high specificity, sensitivity, and convenience.
Similar content being viewed by others
Introduction
All cells are covered with a dense layer of structurally complex and dynamically regulated glycans, mainly manifesting as glycoproteins, glycolipids, and proteoglycans1. Serving as ligands for molecular recognition, cell-surface glycans are essential for a variety of physiopathological processes2,3,4. To decipher the biological functions of glycans, it is a prerequisite to precisely label and analyze glycans in complex biological systems5. Affinity labeling has been a central tool for visualizing and analyzing biomolecules. For most proteins, specific antibodies are commercially available or can be readily generated. However, glycan-specific antibodies with high specificity and affinity are notoriously difficult to generate6,7. This is to some extent due to the conservation of glycan sequences and structures across vertebrates, rendering glycans low-immunogenic antigens8. Furthermore, the available antibodies for glycans are often generated using glycopeptide antigens and therefore commonly bind specific polypeptide sequences in addition to the glycan epitopes, resulting in bias towards different glycopeptides carrying the same glycans9. In addition, glycans often minimally activate T-cell-dependent antibody responses, generating the immunoglobulin M isotype with lower affinity and specificity10,11. Alternatively, lectins—a large family of glycan-binding proteins—have been exploited as affinity probes for glycan labeling and analysis5,12,13. Lectins are often plant-originated, not produced by the immune system, and possess limited affinity and specificity.
Nature has ingeniously evolved glycoenzymes including glycosyltransferases and glycosidases which synthesize and hydrolyze glycans with stringent specificity, respectively14,15. In the past decades, glycoenzymes have been successfully engineered as glycan labeling tools. For example, the chemoenzymatic glycan labeling strategy has been developed by utilizing glycosyltransferase variants to accept unnatural sugar nucleotides modified with bioorthogonal functional groups. After transferring the unnatural sugars onto specific glycans, bioorthogonal reactions are performed to attach functional probes16,17. In contrast, glycosidases only interact with glycans, which may be exploited for developing glycan binders if the enzymatic activity could be mutated while preserving the binding specificity. In support of this, pioneering works have demonstrated that mutations in specific amino acid residues of a bacterial O-GlcNAcase and a bacterial O-glycoprotease generated variants that were enzymatically inactive while preserving the substrate-binding capabilities for O-GlcNAc and mucin-domain glycoproteins, respectively18,19.
With N-acetyl-neuraminic acid (Neu5Ac) as the most common form, sialic acids are a family of nine-carbon monosaccharides that terminate various cell-surface glycans in mammals20. Sialic acids can be α2,3- or α2,6-linked to galactose (Gal), α2,6-linked to N-acetylgalactosamine (GalNAc), or α2,8-linked to another sialic acid. The resulting sialoglycans are attached to N-linked and O-linked glycoproteins as well as glycolipids. Cell-surface sialoglycans play important roles in regulating molecular recognition, cell-cell communications, and pathogen-host interactions. In particular, recognition of sialoglycans on cancer cells by sialic acid-binding immunoglobulin-like lectins (Siglecs) of immune cells contributes to the regulation of immune cell function, which has recently attracted great interest because of its implication in cancer immunotherapy21,22. Importantly, Siglecs exhibit distinctive specificities towards the sialoside linkages. For example, Siglec-2 (CD22) has evolved strict specificity for α2,6-sialoglycans. Siglec-7 primarily binds α2,8-sialosides, while Siglec-8 exhibits preference for Neu5Acα2-3[6SO4]Galβ1-4GlcNAc23,24. To label and analyze sialoglycans on cancer cells, MAL-II (Maackia amurensis lectin) and SNA (Sambucus nigra lectin), two lectins preferentially bind α2,3- and α2,6-sialoglycans, respectively, have been commonly used5. However, SNA also binds α2,3-sialoglycans although with a lower affinity. Non-specific binding between MAL-II and glycans containing 3-O-sulfated galactose has been demonstrated by the glycan array analysis25. Additionally, there are concerns about potential interactions between lectins and other cellular biomolecules such as proteins, because the specificity validation typically hinges on glycan arrays26,27. On the other hand, the available antibodies recognizing specific sialylated glycomotifs are mostly limited to those for polysialic acid (PSA)28, sialyl-Lewis x (sLeX)29, and sialyl-Tn (STn)30. Although several pan-specific sialoglycan antibodies are commercially available (e.g., CABT-Z313R from the Creative Diagnotics and LS-C664155 from the LSBio), the affinity and specificity of which have not been thoroughly evaluated. Interestingly, the commercially available sialidase-derived sialoglycan binders (i.e., SiaFindTM Lectenz) appear to be generated by computational design and yeast display-based evolution31. However, the detailed design, mechanism, and characterization of these reagents have neither been fully disclosed nor been published via peer review.
Herein, we report the development and comprehensive characterization of glycan recombinant affinity binders (GRABs) based on mutant glycosidases, which are featured with high specificity and affinity for sialoglycans, ease of use, and convenience in production. A key catalytic residue of the pan-specific sialidase Streptococcus pneumoniae neuraminidase A (SpNanA) and α2,3-sialidase Ruminococcus gnavus neuraminidase H (RgNanH) was mutated so that they were enzymatically inactive while retaining the recognition for total sialoglycans and α2,3-sialosides, respectively, with high specificity. The resulting GRAB-Sia and GRAB-Sia3 were further tetramerized with streptavidin to improve the avidity. The GRABs and tetra-GRABs were compatible with a variety of experimental settings such as immunoblotting, flow cytometry, immunoprecipitation, and fluorescence microscopy. Using tetra-GRABs, the spatial distribution of total and α2,3-linked sialoglycans was visualized in several mouse organs including the heart, kidney, lung, and intestine. GRABs based on mutant glycosidases thus represent a versatile toolkit for affinity labeling of glycans, which should find broad applications in investigating the functional roles of glycans.
Results
Pan-specific sialoglycan-binding GRAB
To develop a pan-specific GRAB recognizing sialic acids independent of their glycosidic linkages, we exploited SpNanA, a pan-sialidase that hydrolyses all terminal sialic acids with α2,3, α2,6, and α2,8 linkages across various sialoglycans32. SpNanA comprises a type-40 carbohydrate-binding module (CBM40), a catalytic domain belonging to the glycoside hydrolase family 33 (GH33), and a C-terminal domain (C-domain). There are three key catalytic residues conserved in the GH33 family, which typically include a tyrosine as the catalytic nucleophile, a glutamate as a base catalyst that deprotonates the tyrosine hydroxyl group, and an aspartate as an acid catalyst to protonate the glycosidic oxygen33 (Fig. 1a). Following the intermediate formation, the conserved aspartate serves as a base catalyst, activating the incoming water molecule (for hydrolysis) or glycosyl acceptor (in the case of trans-sialidase). Accordingly, structural analysis has identified D372, E647, and Y752 of SpNanA as the three key residues of catalytic acid/base and nucleophile34 (Fig. 1b). To produce GRAB-Sia from SpNanA, we recombinantly expressed the catalytic domain (residues 318-792) with D372 or E647 mutated and a hexahistidine (His tag) fused at the N-terminus, which were referred to as SpNanAD372N, SpNanAD372A, SpNanAE647A, and SpNanAE647Q, respectively (Fig. 1c and Supplementary Fig. 1a). The CBM40 domain and C-domain were truncated to avoid potential non-specific binding to other glycans, as the substrate specificity of CBMs can be distinct from the catalytic domains to which they are fused35. The wild-type (WT) SpNanA catalytic domain (hereafter referred to as SpNanA) was capable of hydrolyzing sialic acids from the model sialoglycoprotein fetuin, resulting in a decrease of molecular weight of fetuin as shown by the sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis (Fig. 1d). By contrast, none of the mutants altered the molecular weight of fetuin, indicating the abolishment of their sialidase activity. Similar results were also observed by using a complementary sialidase activity assay based on a fluorogenic sialoside substrate (Supplementary Fig. 1b).
a Schematic showing the catalytic mechanism of sialidases in the GH33 family. b Close-up view of the catalytic pocket from the previously determined crystal structure of SpNanA in complex with Neu5Ac (PDB: 3H72)60. Three catalytic residues of SpNanA are labeled and shown as sticks; the Neu5Ac molecule is shown in yellow. c Schematic showing the domain organization of SpNanA and the recombinant SpNanA catalytic domain. d SDS-PAGE analysis of fetuin incubated with WT SpNanA, SpNanA variants, or vehicle. CBB, Coomassie brilliant blue. Representative results are shown from three replicates. e HeLa cells were incubated with vehicle, His-tagged WT SpNanA, or His-tagged SpNanA variants, followed by staining with APC-anti-His tag antibody and flow cytometry analysis. Representative flow cytometry histograms and the bar graph of statistical analysis are shown in the left and right panels, respectively. f WT Hela cells with or without pretreatment of SpNanA and CMAS−/− HeLa cells were incubated with vehicle or His-tagged SpNanAD372N, followed by staining with APC-anti-His tag antibody and flow cytometry analysis. In (e, f), MFI median fluorescence intensity, a.u. arbitrary units. Error bars represent the standard deviation (s.d.) from three biological replicates with cells cultured in different wells/dishes in a single experiment. P values were calculated by one-way ANOVA. Source data are provided as a Source Data file.
We then evaluated the sialic acid-binding capability of the SpNanA mutants by measuring their binding to cell-surface sialoglycans using a flow cytometry assay (Supplementary Fig. 2). HeLa cells were incubated with His-tagged SpNanA mutants and stained with an allophycocyanin (APC)-conjugated anti-His tag antibody. Strong binding to HeLa cells was observed for SpNanAD372N and SpNanAD372A, while the two E647 mutants completely lost the binding capability (Fig. 1e). Of note, SpNanA resulted in no fluorescence labeling owing to its sialidase activity. Incubation of WT HeLa cells with SpNanA to remove cell-surface sialic acids markedly reduced the binding with SpNanAD372N (Fig. 1f). Moreover, the N-acylneuraminate cytidylyltransferase (CMAS), which encodes the sole synthetase of cytidine 5′-monophosphate-sialic acid (CMP-Sia), was knocked out in HeLa and HEK293T cells by using the CRISPR/Cas9 system (Supplementary Fig. 3a). The high-performance anion exchange chromatography (HPAEC) analysis showed that CMAS−/− cells lost the capability of biosynthesizing CMP-Sia, the nucleotide sugar donor for sialyltransferases (Supplementary Fig. 3b), thus becoming devoid of terminal sialic acids on cell-surface glycans. Binding of SpNanAD372N on CMAS−/− HeLa cells was completely abolished, confirming the strict binding specificity towards sialoglycans (Fig. 1f).
To elucidate the sialoglycan subtypes to which SpNanAD372N bound, we used fetuin as a model sialoglycoprotein and first validated that it carried multiple types of sialoglycan. Lectin blotting using SNA showed that treatment of fetuin with the pan-specific sialidases, SpNanA and neuraminidase from Arthrobacter ureafaciens (AuNan), but not the α2,3-sialidases, RgNanH36 and neuraminidase C from Streptococcus pneumoniae (SpNanC), largely abolished SNA staining, indicating the presence of α2,6-sialoglycans on fetuin (Supplementary Fig. 4a). Staining of Ricinus communis agglutinin I (RCA I), which recognizes exposed Gal upon desialylation, was greatly enhanced by treatment of fetuin with α2,3-sialidases, indicating the presence of α2,3-sialoglycans on fetuin (Supplementary Fig. 4b). A biotin was site-specifically introduced at the N-terminus of SpNanAD372N by using the BirA-AviTag system37, yielding homogeneously biotinylated proteins with high batch-to-batch reproducibility (Supplementary Fig. 5). Binding of the resulting monobiotinylated SpNanAD372N to fetuin was analyzed by blotting. Treatment of fetuin with the pan-sialidases completely abolished the binding, while the α2,3-sialidases resulted in a partial decrease of binding (Supplementary Fig. 6). These results indicate that SpNanAD372N binds to sialoglycoproteins and presumably sialoglycans with different glycosidic linkages including α2,3- and α2,6-sialosides.
In addition to cell-surface staining and blotting, we evaluated whether SpNanAD372N could be used for affinity capture experiments. Fetuin was efficiently captured, while bovine serum albumin (BSA), a non-glycosylated protein, was not enriched by SpNanAD372N (Supplementary Fig. 7a). Moreover, a variety of proteins in the lysate of HEK293T cells but not that of CMAS−/− HEK293T cells or HEK293T cells pretreated with SpNanA were captured with SpNanAD372N (Supplementary Fig. 7b). These results indicate that SpNanAD372N may serve as a GRAB for pan-specific sialoglycan labeling (hereafter referred to as GRAB-SiaTotal or GRAB-Sia).
α2,3-sialoside-binding GRAB
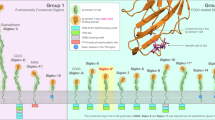
Aiming to develop a GRAB specific for α2,3-sialosides, we engineered the α2,3-sialidase RgNanH, which also belongs to GH33 and possesses the conserved catalytic residues, including D282 and E559 as the acid/base catalysts and Y677 as the nucleophile36 (Fig. 2a). The catalytic domain of RgNanH (residues 243-723) and the D282A and D282N mutants were recombinantly expressed and purified with an N-terminal His tag (Fig. 2b and Supplementary Fig. 8a). As expected, mutation of D282 abolished the sialidase activity of RgNanH (Fig. 2c and Supplementary Fig. 8b). Both RgNanHD282N and RgNanHD282A bound to HeLa cells, with RgNanHD282A showing a higher binding affinity (Fig. 2d). With the α2,3-sialidase activity, RgNanH did not stably bind to the cell surface. To CMAS−/− HeLa cells, no binding of RgNanHD282A was observed, demonstrating the sialic acid-dependence of RgNanHD282A binding (Fig. 2e). Importantly, the binding of monobiotinylated RgNanHD282A to fetuin was sufficiently abrogated by treatment with the α2,3-sialidases to selectively cleave α2,3-linked sialic acids as well as the pan-specific sialidases removing all sialic acids (Fig. 2f and Supplementary Fig. 8c). These results indicate that RgNanHD282A (hereafter referred to as GRAB-Sia3) binds to sialoglycoproteins bearing α2,3-sialosides, probably by recognizing α2,3-sialoglycans. More detailed characterizations of the glycan substrate specificity of both the GRABs were performed in the following sections.
a Close-up view of the catalytic pocket from the previously determined crystal structure of the RgNanH in complex with 2-deoxy 2,3-didehydro Neu5Ac (Neu5Ac2en) (PDB: 4X47)36. Three catalytic residues of RgNanH are labeled and shown as sticks; the Neu5Ac2en molecule is shown in yellow. b Schematic showing the domain organization of RgNanH and the recombinant RgNanH catalytic domain. c SDS-PAGE analysis of fetuin incubated with WT RgNanH, RgNanH variants, or vehicle. Representative results are shown from three replicates. d HeLa cells were incubated with vehicle, His-tagged WT RgNanH, or His-tagged RgNanH variants, followed by staining with APC-anti-His tag antibody and flow cytometry analysis. Representative flow cytometry histograms and the bar graph of statistical analysis are shown in the left and right panels, respectively. e WT Hela cells with or without pretreatment of SpNanA and CMAS−/− HeLa cells were incubated with vehicle or His-tagged RgNanHD282A, followed by staining with APC-anti-His tag antibody and flow cytometry analysis. f Fetuin was incubated with vehicle, RgNanH, SpNanA, SpNanC, or AuNan, followed by blotting with monobiotinylated RgNanHD282A. HRP-conjugated streptavidin was used for visualization. Representative results are shown from three replicates. g WT and CMAS−/− HEK293T cells were incubated with vehicle, biotinylated GRABs (100 μg/mL), SNA (0.2 μg/mL), MAL-II (1 μg/mL), pan-specific SiaFindTM Lectenz (version 1.0 or 2.0, 5 μg/mL), or α2,3-specific SiaFindTM Lectenz (25 μg/mL), followed by staining with streptavidin-AF647 and flow cytometry analysis. Fold changes of MFI are shown above each pair of bars. In (d, e, g), MFI median fluorescence intensity, a.u. arbitrary units. Error bars represent the s.d. from three biological replicates with cells cultured in different wells/dishes in a single experiment. P values were calculated by one-way ANOVA. Source data are provided as a Source Data file.
Next, we benchmarked the binding specificity of GRABs with two widely used lectins recognizing sialoglycans, SNA and MAL-II. In addition, the SiaFindTM Lectenz kits were also compared in parallel. The commercially available biotinylated SNA, MAL-II, and SiaFindTM Lectenz were employed to ensure the comparability with biotinylated GRABs. WT and CMAS−/− HEK293T cells were incubated with the biotinylated lectin, SiaFindTM Lectenz, or GRAB, followed by staining with streptavidin-AF647 and flow cytometry analysis. All lectins, SiaFindTM Lectenz, and GRABs efficiently bound to WT HEK293T cells at a wide range of concentrations (Supplementary Fig. 9). In CMAS−/− HEK293T cells, GRABs exhibited negligible non-specific binding at concentrations up to 100 μg/mL. In contrast, clear binding to CMAS−/− HEK293T cells was observed for SNA and MAL-II at concentrations down to 0.2 μg/mL, which was in agreement with the previously observed non-specific binding of SNA and MAL-II25,27. Similarly, pan-specific SiaFindTM Lectenz (both version 1.0 and 2.0) at 5 μg/mL and α2,3-specific SiaFindTM Lectenz at 25 μg/mL resulted in marked binding to CMAS−/− HEK293T cells (Fig. 2g and Supplementary Fig. 9). At the concentrations that yielded similar fluorescence intensities in WT HEK293T cells, the GRABs exhibited the highest specificity (fold change of MFI between labeled WT and labeled CMAS−/− cells) and lowest non-specific binding (fold change of MFI between labeled and unlabeled CMAS−/− cells) among all the sialic acid binders (Fig. 2g). These results demonstrate the superb specificity of GRABs towards sialoglycans.
Tetramerization of GRABs and substrate specificity
We next quantitatively measure the binding affinity of GRABs with the isothermal titration calorimetry (ITC) analysis. GRAB-Sia displayed comparable affinities for α2,3-sialyllactose (3′SL) and α2,6-sialyllactose (6′SL), with a dissociation constant (Kd) of 17.8 μM and 19.1 μM, respectively (Fig. 3a, b). GRAB-Sia3 bound to 3′SL with a Kd of 7.15 μM, but not to 6′SL (Fig. 3c, d). These results were consistent with the sialoside linkage preference of GRAB-Sia and GRAB-Sia3 observed in the fetuin binding assay.
GRAB-Sia was subjected to ITC analysis with 3′SL (a) and 6′SL (b) as the ligand. GRAB-Sia3 was subjected to ITC analysis with 3′SL (c) and 6′SL (d) as the ligand. In (a–d), the ITC titration and binding curves are shown. e Monobiotinylated GRABs generated using the BirA-AviTag system were mixed with streptavidin conjugates to yield tetra-GRABs. Created in BioRender. Liang, S. (2025) https://BioRender.com/g78q224. Binding of tetra-GRAB-Sia (f) and tetra-GRAB-Sia3 (g) to different glycan structures was evaluated by glycan array analysis. The bar chart showed relative fluorescence units (RFU) of the 166 glycans by their order of glycan ID on the x-axis. Data are presented as mean ± s.d. of four replicates. Glycan structures are grouped based on the terminal monosaccharide. h Representative glycan substrates of tetra-GRAB-Sia and tetra-GRAB-Sia3. Source data are provided as a Source Data file. The source data for (f, g) are provided in a separate Excel file as Supplementary Data 1.
Considering that the binding affinity of GRAB-Sia and GRAB-Sia3 was lower than that of typical antibodies, we sought to enhance the avidity by multimerization of GRABs with streptavidin. The approach has been successfully demonstrated for the MHC tetramer technology, which leverages the assembly of MHC monomers into tetrameric forms to improve binding to T cell receptors38. Besides, multimerization of glycan-binding proteins has also been used to enhance the avidity towards high-density glycans26,39,40. To generate GRAB tetramers, monobiotinylated GRAB-Sia and GRAB-Sia3 were assembled with streptavidin (Fig. 3e). Size exclusion chromatography (SEC) analysis showed that tetra-GRABs were efficiently formed when the molar ratio of monobiotinylated GRAB and streptavidin was at 1:1 (Supplementary Fig. 10). Owing to the superb binding affinity between biotin and the highly stable streptavidin tetramer, the tetra-GRABs could be directly used without the need of further purification.
With tetra-GRABs, we elucidated the substrate specificity of GRABs at the glycan structure level by glycan array analysis, in which the binding of glycan binders towards various immobilized glycans is quantitatively analyzed in a high-throughput manner41. The glycan array we used contained a collection of 166 distinct glycans, covering a wide spectrum of mammalian glycans (Supplementary Fig. 11). As expected, tetra-GRAB-Sia exhibited high affinities for all classes of sialoglycans regardless of their glycosidic linkages, except for sialoglycans with branches at the subterminal galactose residue (Fig. 3f, Supplementary Fig. 12a and Supplementary Data 1). In these instances, the steric hindrance from the branched residues probably impeded the access of sialic acid to the binding pocket of GRAB-Sia. Conversely, tetra-GRAB-Sia3 exhibited a remarkable preference for α2,3-sialoglycans and excluded all α2,6- and α2,8-sialoglycans from the recognition repertoire (Fig. 3g, Supplementary Fig. 12b and Supplementary Data 1). Though GRAB-Sia and GRAB-Sia3 showed similar binding affinities for their respective substrates based on ITC and flow cytometry analysis, lower signal intensities were observed for tetra-GRAB-Sia3 in the glycan array analysis. This was possibly due to the deeper binding pocket of RgNanH that rendered the recognition of α2,3-sialoglycans immobilized on the solid surface of glycan arrays less efficient than those on the cell surface and in the aqueous solution (Supplementary Fig. 13a). Interestingly, sLeX with an α2,3-linked sialic acid was not recognized by tetra-GRAB-Sia3, which was possibly attributed to the steric hindrance posed by the fucose α1,3-linked to N-acetylglucosamine (GlcNAc) (Supplementary Fig. 12b and Supplementary Data 1). Of note, minimal binding towards sulfated glycans was observed for both tetra-GRAB-Sia and tetra-GRAB-Sia3 (Fig. 3f, g, Supplementary Fig. 12, and Supplementary Data 1). Furthermore, tetra-GRAB-Sia and tetra-GRAB-Sia3 recognized sialylated glycomotifs from multiple types of glycoconjugates including glycolipids, N-glycans, and mucin-type O-glycans (Fig. 3h).
To evaluate whether GRABs can recognize other forms of sialic acids in addition to Neu5Ac, we conducted another glycan array analysis that covered sialoglycans with N-glycolylneuraminic acid (Neu5Gc) and deaminated neuraminic acid (Kdn) as well as other modified sialic acids (Supplementary Fig. 14 and Supplementary Data 2). Sialoglycans terminated with Neu5Gc and Kdn were mostly not recognized by tetra-GRAB-Sia or tetra-GRAB-Sia3. Interestingly, tetra-GRAB-Sia3 was also bound to sialoglycans with 8-O-methylated Neu5Ac (Neu5Ac8Me). The fact that both tetra-GRAB-Sia and tetra-GRAB-Sia3 possess binding preference towards Neu5Ac-containing sialoglycans might be correlated to the conserved amino acids around the N-acetyl group of Neu5Ac in their binding pockets (Supplementary Fig. 13b).
Enhanced avidity of tetra-GRABs
We then evaluated whether tetramerization enhanced the avidity of GRABs. Compared to the labeling of HeLa cells with monobiotinylated GRABs followed by staining with streptavidin-AF647, labeling with tetrameric GRAB-Sia-streptavidin-AF647 (tetra-GRAB-Sia-AF647) and GRAB-Sia3-streptavidin-AF647 (tetra-GRAB-Sia3-AF647) resulted in a 93-fold and 225-fold increase in fluorescence intensity, respectively (Fig. 4a, b). Comparable labeling intensities were observed for tetra-GRABs or SNA under the same concentrations (Supplementary Fig. 15a), which was in contrast to monomeric GRABs (Supplementary Fig. 9). The tetra-GRABs preserved the superb specificity towards sialoglycans, as demonstrated by the loss of binding on CMAS−/− HeLa cells and WT HeLa cells pretreated with SpNanA (Fig. 4c, d). Moreover, adding free Neu5Ac or 9-O-acetylated Neu5Ac (Neu5,9Ac2) significantly reduced the labeling signals of both tetra-GRABs to a similar level, suggesting that tetra-GRABs should also bind Neu5,9Ac2-containing sialoglycans (Supplementary Fig. 15b, c). Addition of cytidine 5′-monophosphate-Neu5Ac (CMP-Neu5Ac), serving as a sialoside, also remarkably reduced the labeling signal (Supplementary Fig. 15b, c). The fluorescence intensity of labeled HeLa cells was dependent on the concentration of tetra-GRABs (Supplementary Fig. 15d). At concentrations as low as 1 µg/mL, a signal-to-noise ratio of 1427 and 1186 was achieved with tetra-GRAB-Sia-AF647 and tetra-GRAB-Sia3-AF647, respectively, owing to the high avidity (Fig. 4a, b). Furthermore, the labeling reached equilibrium within 10 min, demonstrating fast binding kinetics between tetra-GRABs and cell-surface sialoglycans (Supplementary Fig. 15e).
Flow cytometry analysis of HeLa cells incubated with tetra-GRAB-Sia-AF647 (a) or tetra-GRAB-Sia3-AF647 (b). Alternatively, HeLa cells were incubated with monobiotinylated GRAB-Sia (a), GRAB-Sia3 (b), or vehicle (a, b), followed by staining with streptavidin-AF647 and flow cytometry analysis. Representative flow cytometry histograms and the bar graph of statistical analysis are shown in the left and right panels, respectively. WT Hela cells with or without pretreatment of SpNanA and CMAS−/− HeLa cells were incubated with vehicle (c, d), tetra-GRAB-Sia-AF647 (c), or tetra-GRAB-Sia3-AF647 (d) and subjected to flow cytometry analysis. e HeLa cells were individually incubated with vehicle, tetra-GRAB-Sia3-AF647, tetra-GRAB-Sia-AF488, or simultaneously incubated with both in a one-pot manner, followed by quantification of the fluorescence intensity via flow cytometry analysis. Error bars represent the mean ± s.d. from three biological replicates with cells cultured in different wells/dishes in a single experiment. P values were calculated by one-way ANOVA. Source data are provided as a Source Data file.
Since 50 µg/mL of tetra-GRABs did not reach saturation for labeling HeLa cells (Supplementary Fig. 15d), 1 µg/mL of tetra-GRABs should label only a small fraction of cell-surface sialoglycans. In line with this, elevating the cell concentrations resulted in decreased labeling intensity (Supplementary Fig. 15f, g). We then exploited the substoichiometric labeling for simultaneous and multiplexed analysis of total and α2,3-linked sialoglycans in the same samples. HeLa cells were simultaneously incubated with tetra-GRAB-Sia-AF488 and tetra-GRAB-Sia3-AF647 in a one-pot manner. The AF488 and AF647 fluorescence intensities were at the same level compared to cells stained with individual tetra-GRABs (Fig. 4e). Importantly, tetra-GRAB staining did not cause observable cytotoxicity in HeLa cells as well as in primary mouse embryonic cells (Supplementary Fig. 16a, b). Furthermore, HeLa cells stained with tetra-GRABs exhibited proliferation rates similar to that of unstained cells (Supplementary Fig. 16c). Together, these results demonstrate the high efficiency, multiplexing ability, and biocompatibility of tetra-GRABs for analyzing sialoglycans in living cells.
Sialoglycan imaging with tetra-GRABs in cells
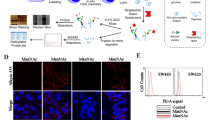
We next applied tetra-GRABs for imaging cell-surface sialoglycans. HEK293T cells were fixed and stained with tetra-GRAB-Sia-AF647, tetra-GRAB-Sia3-AF488, and SNA-Cy3. Strong fluorescence signals representing the total, α2,3-linked, and α2,6-linked sialoglycans were observed on the plasma membranes of WT cells, whereas such signals were absent in SpNanA-treated or CMAS−/− cells (Fig. 5a). Treatment with RgNanH partially decreased the labeling of tetra-GRAB-Sia, completely abolished the labeling of tetra-GRAB-Sia3, and did not affect SNA staining (Fig. 5a). These results again confirmed the linkage specificity of tetra-GRABs. Next, we wondered if tetra-GRABs could be used for imaging intracellular sialoglycans. HeLa cells were fixed, permeabilized, and stained with tetra-GRAB-Sia-AF647. Confocal fluorescence microscopy analysis revealed co-localization of the AF647 signals with the Golgi apparatus, where sialylation occurred42 (Supplementary Fig. 17).
a Confocal fluorescence images of fixed WT and CMAS−/− HEK293T cells stained with tetra-GRAB-Sia-AF647, tetra-GRAB-Sia3-AF488, and SNA-Cy3. WT HEK293T cells pretreated with SpNanA or RgNanH were used for comparison. Scale bars, 10 μm. b Confocal fluorescence images of mouse heart and kidney sections with or without pretreatment of SpNanA incubated with tetra-GRAB-Sia-AF647 and tetra-GRAB-Sia3-AF488. The glomerulus, luminal side of the proximal tubule, and basement membrane are highlighted with white, yellow, and magenta arrows, respectively. Scale bars, 50 μm. In (a, b), the nuclei were stained with Hoechst 33342. Representative images were shown from three independent biological replicates.
Sialoglycan imaging with tetra-GRABs in tissues
Next, tetra-GRABs were utilized for imaging sialoglycans in tissue sections. The heart, kidney, and lung from C57BL/6 mice were fixed, cryosectioned, and stained with tetra-GRAB-Sia-AF647 and tetra-GRAB-Sia3-AF488, followed by confocal fluorescence microscopy analysis. Both tetra-GRABs exhibited robust labeling in the heart and kidney but not in the samples pretreated with SpNanA (Fig. 5b). In the heart, strong fluorescence for total and α2,3-linked sialoglycans was observed on the surfaces of both cardiomyocytes and cardiac fibroblasts. In the kidney, fluorescence staining of sialoglycans was primarily localized in the glomerulus, possibly attributed to the elevated expression levels of the highly sialylated glycoprotein podocalyxin in podocytes43 (Fig. 5b and Supplementary Fig. 18). Notably, the luminal side of the proximal tubule was strongly stained with tetra-GRAB-Sia3 but not tetra-GRAB-Sia, indicating the polarized expression of sialoglycans with different linkages in the tubular epithelial cells (Fig. 5b). In addition, we noted remarkable labeling of tetra-GRAB-Sia and tetra-GRAB-Sia3 on the glomerular and tubular basement membrane, consistent with the fact that these structures are highly enriched with extracellular sialoglycoproteins44 (Fig. 5b and Supplementary Fig. 18). We then compared tetra-GRABs and lectins side-by-side in staining mouse kidney tissues. Residual signal of SNA was observed after SpNanA treatment, indicating possible non-specific binding (Supplementary Fig. 19a). Similarly, non-specific binding of MAL-II was evidenced by residual labeling after SpNanA or RgNanH treatment (Supplementary Fig. 19a, b). Conversely, both tetra-GRABs exhibited superb specificity, as SpNanA and RgNanH completely abolished the fluorescence signals of tetra-GRAB-Sia and tetra-GRAB-Sia3, respectively (Supplementary Fig. 19a, b). In the lung tissue, signals of tetra-GRAB-Sia3 but not tetra-GRAB-Sia were well-colocalized with the epithelial marker EpCAM, indicating the enrichment of α2,3-sialoglycans in alveolar cells (Supplementary Fig. 20). These results demonstrate that tetra-GRABs are versatile probes for imaging sialoglycans with high specificity in tissue samples.
Analyzing sialoglycan distribution in mouse intestine with tetra-GRABs
We next applied tetra-GRABs to analyze the sialoglycan distribution in the mouse intestine. The intestinal epithelium expresses a diverse array of N- and O-linked glycans that serve as a protective barrier against foreign toxins and pathogens45. Consequently, intestinal glycans play a key role in maintaining mucosal homeostasis despite being constantly challenged by diverse foreign stimuli46. Sialoglycans have been recently found to be essential for mucus integrity and protect against excessive bacterial proteolytic degradation47. In addition, as the gut microbiota utilize mucosal sialoglycans as nutrients, the spatial distribution of sialoglycans also contributes to microbiota homeostasis48.
By staining mouse intestine slices with tetra-GRAB-Sia-AF647 and tetra-GRAB-Sia3-AF555, the spatial distribution of sialoglycans along the gastrointestinal tract was visualized by confocal fluorescence microscopy. In the small intestine including the jejunum and ileum, total and α2,3-linked sialoglycans exhibited very similar patterns (Supplementary Fig. 21a). By co-staining EpCAM as the marker of epithelial cells, we found that the majority of sialoglycans were located on EpCAM-negative cells, indicating that epithelial cells in the small intestine had a low level of sialylation. The low level of sialoglycans on epithelia cells was also observed in the large intestine. However, the large intestine displayed a more diverse sialoglycan distribution (Fig. 6a, b). Among different regions of the large intestine, the cecum and rectum exhibited the strongest signal of tetra-GRAB-Sia, while the signal of tetra-GRAB-Sia3 was enriched in the distal colon. In the cecum, signals of total sialoglycans exhibited a punctum-like pattern with an average diameter of 15 μm, likely originating from mucus synthesized by goblet cells (Fig. 6a). In comparison, the distal colon was featured with significantly upregulated α2,3-sialoglycans in similar puncta with higher density and fluorescence intensity (Fig. 6a). Of note, the puncta enriched with total sialoglycans were also observed in the distal colon, but only partially colocalized with the α2,3-sialoglycan-enriched puncta, indicating that specific subtypes of sialoglycans were spatially regulated in the colon. To correlate sialoglycan distribution with transcriptional regulations, we analyzed a published RNA-seq dataset on the mouse intestine49. This revealed a remarkable upregulation of genes involved in sialic acid biosynthesis (i.e., Cmas, Gne, and Nans) and sialyltransferases in the distal colon (Fig. 6c, Supplementary Fig. 21b–d), which was in accordance with the GRAB staining results. Among α2,3-sialyltransferases, St3gal4 and St3gal6 exhibited the highest expression levels in the distal colon (Supplementary Fig. 21b). Moreover, St3gal4 and St3gal6 appeared to be the predominant α2,3-sialyltransferases in the colon, since other ones including St3gal1 and St3gal2 were expressed in a much lower level in the whole colon (Supplementary Fig. 21b). The high α2,3-sialoglycan staining can therefore be possibly attributed to the upregulation of the genes involved in sialic acid biosynthesis and the predominate α2,3-sialyltransferases in the distal colon.
a Mouse large intestine sections were incubated with AF488-labeled anti-EpCAM antibody, tetra-GRAB-Sia-AF647, and tetra-GRAB-Sia3-AF555, followed by fluorescence microscopy analysis. Scale bars, 500 μm and 50 μm for zoomed-out and zoomed-in images, respectively. Representative images were shown from three independent biological replicates. b Schematic showing the spatial distribution of total and α2,3-linked sialoglycans in the mouse intestine revealed by tetra-GRAB staining. Created in BioRender. Liang, S. (2025) https://BioRender.com/g78q224. c Heatmap showing the scaled expression of genes involved in sialoglycan biosynthesis at the proximal and distal colon. The heatmap was generated using a public RNA-seq dataset49. Source data are provided as a Source Data file.
Discussion
Precise glycan labeling is a prerequisite for studying glycan functions. The widely used plant-originated lectins usually suffer from poor specificity and affinity. In this work, we have developed GRABs as a class of versatile sialoglycan binders, with high specificity and affinity. The two GRABs, GRAB-Sia and GRAB-Sia3, were engineered from bacterial sialidase with the catalytic residues mutated so that they became enzymatically inactive while retaining sialoglycan binding. The bacterial origin allows for convenient expression and production of GRABs in prokaryotic systems. These GRABs possess high specificity as compared with SNA, MAL-II, and SiaFindTM Lectenz. More importantly, sialoglycan-deficient cells were employed as a comprehensive and rigorous negative control. Complete lack of GRAB labeling on CMAS−/− cells proved that the GRABs neither bound non-sialylated glycans nor non-specifically interacted with other biomolecules such as proteins and lipids, highlighting the superb specificity that lectins are short of. In the current study, we focused on the use of GRABs primarily in mammalian cells and tissues. Given that bacteria possess sialoside linkages such as α2,9-linked PSA50, the current GRABs, and development of GRABs specific for these linkages will be valuable for studying glycosylation in these species. In addition, the current GRABs recognize only terminal sialic acids considering the exosialidase activity of the parental enzymes. To distinguish more complex sialoglycans including the underlying glycan structures, GRABs may be further developed by direct evolution. Furthermore, GRABs may be used in combination with binders to proteins and lipids for studying specific sialoglycoconjugates such as glycoproteins and glycolipids.
Recognition of glycans by proteins is often multivalent51, which we exploited for improving the avidity of GRABs simply by tetramerizing them with streptavidin. The greatly enhanced labeling efficiency of tetra-GRABs was demonstrated. A variety of streptavidin conjugates such as streptavidin-fluorophore is commercially available, making tetra-GRABs conjugated with functional probes easily accessible. It should be noted that the labeling intensity of tetra-GRABs may depend on both the amount and density of sialoglycans. Although the amount and density are usually correlative, sialylation levels cannot be determined definitively without the use of additional methods such as mass spectrometry. In certain circumstances such as cell membrane microdomains enriched with high-density siaologlycans, high-resolution and super-resolution fluorescence microscopy techniques need to be used to differentiate the two factors. The development of GRAB monomers with enhanced binding affinity will be also valuable for such studies.
As an affinity labeling tool, GRABs provide several advantageous features, thus complementing other methods for sialic acid labeling. For example, sialoglycans can be metabolically incorporated with unnatural sugars containing a bioorthogonal functional group such as the azide, followed by subsequent click-labeling with fluorophores5,52. The metabolic glycan labeling method hinges on the metabolic activity of cells, while GRABs are compatible with the stored clinical samples, many of which have been fixed. Alternatively, cell-surface sialic acids can be selectively oxidized to aldehydes by sodium periodate for subsequent conjugation with fluorophores containing a hydrazide or aminooxy group via imine ligation53. The oxidative conditions are often a concern for sensitive samples. Moreover, GRABs provide specificities for different sialoglycan subtypes. In this regard, two-step chemoenzymatic labeling methods requiring modified nucleotide sugar donors were developed for tagging α2,3-sialoglycans54,55. In addition to being complementary, GRAB labeling possesses the convenience of one-step labeling.
With GRAB-Sia and GRAB-Sia3 now available, the development of GRABs specific for α2,6- and α2,8-sialoglycans should be explored as a future direction. In addition to sialidases, other bacterial glycosidases56 are amenable to engineering into GRABs for binding various glycan types. Of note, the general base and general acid in all glycosidases are typically carboxyl groups57, indicating that the engineering strategy used in this work may have a generic applicability. Finally, the GRABs in this study are generated by rational design and small-scale screening. Integrating computational design and directed evolution to generate GRABs is a promising future direction.
Methods
Ethical statement
This study was conducted in accordance with all relevant ethical regulations. All animal experiments were performed in accordance with guidelines approved by the Institutional Animal Care and Use Committee of Peking University (IACUC protocol number: CCME-ChenX-3) accredited by the Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC) International.
Reagents and antibodies
3′SL (cat. OS03009) and 6′SL (cat. OS03010) were purchased from Glycogene. HRP-conjugated streptavidin (cat. A0303, 1:2,000) was purchased from Beyotime. APC conjugated anti-His tag antibody (cat. 362605, 1:100) and AF488-conjugated anti-mouse EpCAM antibody (cat. 118210, 1:100) were purchased from BioLegend. Streptavidin-AF647 (cat. 2445829) and streptavidin-AF488 (cat. 2506099) were purchased from ThermoFisher. Biotinylated SNA (cat. B-1305-2), SNA-Cy3 (cat. CL-1303-1), biotinylated MAL-II (cat. B-1265-1), and biotinylated RCA (cat. B-1085-1) were purchased from the Vector Lab. Neuraminidase from Arthrobacter ureafaciens (AuNan, cat. P0722S; with a trade name of α2-3,6,8,9 Neuraminidase A) and neuraminidase C from Streptococcus pneumoniae (SpNanC, cat. P0743S; NanC was deduced by the disclosed information of α2,3 specificity, molecular weight, and the species. The trade name is α2-3 Neuraminidase S) were purchased from NEB. Biotinylated pan-specific SiaFindTM Lectenz 1.0 (cat. SK0501B), biotinylated pan-specific SiaFindTM Lectenz 2.0 (SK0502B), and biotinylated α2,3-specific SiaFindTM Lectenz (SK2301B) were purchased from Lectenz Bio. Hoechst 33342 (cat. C0031) was purchased from Biosynthesis Biotechnology. Fetuin (cat. F10555) was purchased from Psaitong. Biotin (cat. B105433) was purchased from Macklin. Neu5Ac (cat. A100555) was purchased from Aladdin. CMP-Neu5Ac sodium salt (cat. R115229) was from Rhawn. Neu5,9Ac2 was chemically synthesized from Neu5Ac as previously reported58.
Plasmids
All constructed plasmids were confirmed by DNA sequencing before use. All primer sequences are listed in Supplementary Table 1. The plasmid encoding SpNanA was generously provided by Prof. Peng Wang at the Southern University of Science and Technology. The DNA sequence encoding RgNanH was obtained by gene synthesis. The DNA sequences encoding the catalytic domain of SpNanA and RgNanH were inserted into the pET15b vector between the NdeI and BamHI sites. Site-directed mutagenesis was performed using the Fast Mutagenesis System (TransGen, cat. FM111-01). To obtain monobiotinylated GRABs, the sequences of GRABs were inserted between the BamHI and SalI sites of the pET-Dual-His-Avi-MCS-BirA vector (Beyotime, D3011). All ligase-free cloning reactions were performed with the ClonExpress® II One Step Cloning Kit (Vazyme, cat. C112).
Protein expression and purification
The plasmids were transformed into E. coli BL21 (DE3) competent cells (TIANGEN, cat. CB105-02), followed by culturing to an OD600 of 0.6 in LB medium and addition of 500 μM isopropyl b-D-1-thiogalactopyranoside (IPTG) overnight at 22 °C. For expressing monobiotinylated GRABs, 50 μM of biotin was added to the culture medium. The cells were harvested by centrifugation at 4500 × g for 20 min and then lysed by ultrasonication in PBS. The supernatants were collected by centrifugation at 13,000 × g for 30 min at 4 °C, followed by purification of His-tagged proteins with nickel nitrilotriacetic acid (Ni-NTA) column (Yeasen). The expressed proteins were eluted with PBS containing 250 mM imidazole. The monobiotinylated GRABs were further characterized using a high-resolution ESI Quadrupole-TOF LC-MS/MS System (Waters Vion) to obtain the deconvoluted mass.
Sialidase activity
In the gel shift assay, 1 μL of purified sialidase variant (1 mg/mL in PBS) was added to 100 μL of fetuin (1 mg/mL in PBS), which resulted in a molar ratio of ~1:100. The resulting sample was incubated at 37 °C for 1 h. The reaction mixture was then mixed with 25 μL of 5X denaturing protein loading buffer (Yeasen, No. 20315ES05) and heated at 95 °C for 10 min before being subjected to SDS-PAGE analysis and CBB staining. Replicates were performed on different days, using samples independently treated and analyzed. The fluorogenic assay for relative sialidase activity was conducted following the manufacturer’s protocol (Beyotime, P0306), using 2 ng of SpNanA or its mutants, or 10 ng of RgNanH or its mutants, as the input. Replicates were performed using the same batch of proteins in different wells of 96-well plate.
Cell lines
HeLa (ATCC®CCL-2) and HEK293T (ATCC®CRL-11268) cells were obtained from American Type Culture Collection. Cells were cultured in DMEM (Gibco) supplemented with 10% (vol/vol) heat-inactivated fetal bovine serum (Gibco), 100 U/mL penicillin, and 100 μg/mL streptomycin (Gibco) at 37 °C under 5% CO2.
Mice embryonic cells and tissue sections
All mice were purchased from Charles River Laboratories. Mouse embryos at days 18–21 were collected from pregnant C57BL/6N mice, euthanized, cut into small pieces, and dissociated with 0.1 mg/mL collagenase I (Sangon, cat. A004194) at 37 °C for 1 h. The cells were then filtered with a 40-μm cell strainer for subsequent analysis. Mouse tissue sections were collected from 6–8 weeks female C57BL/6N mice. Three mice were sacrificed and the heart, kidney, lung, and intestine were collected. After washing with PBS, the tissue was fixed with 4% paraformaldehyde (PFA) at 4 °C overnight followed by the dehydration with 15% sucrose and 30% sucrose for at least 48 h. The tissues were then embedded with O.C.T. (Sakura) and transferred into the freezing microtome. Ten-micrometer sections were used for GRAB staining.
Generation of CMAS-knockout cells
The gRNA sequence (5′-GAAGAACATTAAGCACCTGG-3′) targeting the human CMAS gene was inserted into the px458 vector59. HeLa or HEK293T cells were transfected with the resulting px458-gRNA plasmid using Lipofectamine 2000 according to the manufacturer’s protocol. After 48 h, GFP-positive cells were sorted into 96-well using a BeckMan MoFlo XDP cell sorter. Clonally expanded cells were subjected to genotyping using primers targeting the genomic region of CMAS. Successful knockout was validated by Sanger sequencing with a pair of amplifying and sequencing primers: 5′-TCGGGCGGCGCCGAGCTGAG-3′ and 5′-AGCCCCTCCAGCAGCCCGAC-3′.
Quantification of CMP-Neu5Ac using HPAEC
The cells were dissociated with trypsin, washed twice in cold PBS, resuspended in 1 mL of cold PBS, counted, and centrifuged. Ten million cells per sample were resuspended in 0.5 mL of 75% ethanol in deionized water and lysed by sonication, followed by centrifuging at 15,000 × g for 15 min at 4 °C to remove insoluble debris. The resulting supernatant was evaporated in a vacuum centrifuge. MP-C9-Azido-Sialic-Acid was used as an internal standard. 300 µL solution containing 20 μM MP-C9-Azido-Sialic-Acid (Glycogene) in 10 mM ammonium bicarbonate was added to the sample and stored at 4 °C before HPAEC analysis. HPAEC was performed on a Dionex CarboPacTM PA1 column (4 × 250 mm, analytical) equipped with a DIONEX ICS-5000 HPAEC system with a UV detector set at 254 nm. The eluent A was 1 mM NaOH in deionized water and C was 1 mM NaOH with 1 M NaOAc in deionized water. The gradient used was: 0 min (A 95%; C 5%); 40 min (A 70%; C 30%); 45 min (A 70%; C 30%); 60 min (A 45%; C 55%); 65 min (A 45%; C 55%); 75 min (A 0%; C 100%); and 80 min (A 0%; C 100%). Intracellular concentrations of CMP-Neu5Ac were calculated by estimating the diameter of a HEK293T cell as 13 μm. For replicates, cells were cultured in different wells/dishes in the same incubator using the same batch of culture media, and then independently dissociated and transferred to different tubes for further process and HPAEC quantitation on the same day.
Screening of sialidase variants for cell binding
HeLa cells were dissociated with trypsin and resuspended in PBS. Afterward, cells were incubated with 10 μg/mL His-tagged sialidase variants at room temperature for 1 h with rotation, followed by two rounds of washing with PBS. Then, the cells were stained with APC-labeled anti-His tag antibody. After another two rounds of washing, cells were subjected to flow cytometry analysis with a BD Accuri C6 flow cytometer. Replicates were performed in a single experiment with cells cultured from different wells/dishes. Specifically, the cells for replicates were cultured in different wells/dishes in the same incubator using the same batch of culture media, and then independently dissociated, stained, and washed in separate tubes before loading to a flow cytometer on the same day.
Lectin and GRAB blotting
One microliter of recombinant SpNanA (1 mg/mL in PBS), RgNanH (1 mg/mL in PBS), AuNan (20,000 units/mL), or SpNanC (8000 units/mL) was added to 100 μL of fetuin (1 mg/mL in PBS). The resulting sample was incubated at 37 °C for 1 h. The reaction mixture was then mixed with 25 μL of 5X denaturing protein loading buffer (Yeasen, No. 20315ES05) and heated at 95 °C for 10 min. After loading and running the protein samples by SDS-PAGE and transferring the proteins from the gel to the PVDF membrane, the membrane was blocked for 1 h at room temperature using blocking buffer (5% BSA in TBST). Then, the membrane was incubated overnight with monobiotinylated GRAB (10 μg/mL), biotinylated SNA (0.5 μg/mL), or biotinylated RCA (0.5 μg/mL) in the Primary Antibody Dilution Buffer (Beyotime, P0023A) at 4 °C. After three times of washing with TBST, the membrane was incubated with HRP-conjugated streptavidin (1:5000 for GRAB and 1:10,000 for SNA or RCA) in TBST at room temperature for 1 h, followed by another three washing steps with TBST. A Tanon-5200Multi (Tanon) scanner was used for scanning the membrane. For all the SDS-PAGE and blotting analyses, representative results are shown from three replicates performed on different days.
Enrichment of sialoglycoproteins with GRAB-Sia
Cells were lysed with RIPA buffer [1% Nonidet P-40 (vol/vol), 0.5 % sodium deoxycholate (wt/vol), 150 mM NaCl, 0.1% SDS (wt/vol), 50 mM Tris-HCl, EDTA-free protease inhibitor mixture (Pierce), pH 7.4]. To modify resins with GRAB-Sia, Ni-NTA resins were incubated with His-tagged GRAB-Sia (1:5.5, mL:mg) at 4 °C for 3 h, followed by washing with PBS for three times. For the enrichment of sialoglycoproteins, purified proteins in PBS or cell lysates in RIPA were incubated with GRAB-Sia-modified resins (50:1, mg of lysates:mL of resins) overnight at 4 °C. Afterwards, the resins were washed with PBS for purified proteins or with RIPA for cell lysates for five times, followed by elution with 10 mM Neu5Ac for three times. The combined elutes were concentrated by using 10 kDa ultrafilters for further analysis.
Sialidase treatment on cells and tissue sections
Cells (2 × 106 cells/mL) or tissue sections were incubated with 10 μg/mL of recombinant SpNanA or RgNanH at room temperature for 30 min (cells) or 1 h (tissue sections).
Glycan array analysis
The glycan array chips (KERUIXIN Co., Ltd.) with one covering 166 distinct glycan structures and another covering 53 glycans containing modified sialic acids were used. Each glycan structure was repeatedly immobilized on four (Fig. 3f, g) or six different spots (Fig. S14b) for replication. The simplified nomenclature for glycan structures is shown in Supplementary Figs. 11 and 14. Briefly, the glycan array chip was firstly blocked and washed twice before use. Meanwhile, the monobiotinylated GRAB (15 μg/mL) was pre-incubated with streptavidin-Cy3 (2.5 μg/mL) to generate the tetra-GRABs. Then, the chip was incubated with tetra-GRAB-Cy3 for 1 h at room temperature, followed by another two rounds of washing steps before recording the absorbance using an excitation light at 532 nm with photomultiplier tube set at 500.
Isothermal titration calorimetry
ITC experiments were performed using the PEAQ-ITC system (Malvern, UK). Before titration, the protein samples were further purified by SEC. The 3′SL and 6′SL were dissolved in PBS. The protein concentration in the detection cell was set as 50 μM and the ligand concentration in the syringe was set as 1 mM in all experiments. The Malvern software was used to analyze the data.
SEC analysis of tetra-GRABs
SEC was performed on an AKTA system (GE Healthcare). The proteins were filtered and loaded into a Superdex 200 Increase GL column (GE Healthcare) pre-equilibrated by the same buffer. Data were processed with the ASTRA6 software (Wyatt).
Generation of fluorescent tetra-GRABs
One hundred micrograms per milliliter of monobiotinylated GRAB was mixed with 25 μg/mL fluorophore-conjugated streptavidin in PBS, followed by incubation at 4 °C for 30 min before use. The concentration of monobiotinylated GRAB in the solution of tetra-GRABs was used to represent the concentration of tetra-GRABs unless otherwise noted.
Labeling of sialoglycans with GRABs, SiaFind Lectenz, and lectins for flow cytometry analysis
For tetra-GRABs, trypsin-dissociated cells were resuspended in PBS and incubated with 1 μg/mL of fluorescent tetra-GRABs at room temperature for 30 min with rotation. After washing twice with PBS, the cell suspension was subjected to flow cytometry analysis. To compare the avidity of GRABs and tetra-GRABs, 1 μg/mL of monobiotinylated GRAB and 0.5 μg/mL of streptavidin-AF647 were used for the two-step labeling. To compare GRABs with SiaFindTM Lectenz and lectins, cells were incubated with monobiotinylated GRAB, biotinylated lectin, or biotinylated SiaFindTM Lectenz at varied concentrations in PBS at room temperature for 30 min. Subsequently, the cells were washed with PBS twice and incubated with 0.5 μg/mL streptavidin-AF647 at room temperature for 30 min. Finally, the cells were washed with PBS for another two times and subjected to flow cytometry analysis. All flow cytometry data was analyzed using FlowJo v10.9. The median fluorescence intensity (MFI) values are presented in arbitrary units, calculated by dividing the original intensity by a unified factor (e.g., 10,000). Biological replicates were performed in a single experiment with cells cultured from different wells/dishes. Specifically, the cells for replicates were cultured in different wells/dishes in the same incubator using the same batch of culture media, and then independently dissociated, stained, and washed in separate tubes before loading to a flow cytometer on the same day.
Competition of tetra-GRAB binding by sialic acids
Trypsin-dissociated cells were resuspended in PBS and incubated with 1 μg/mL of fluorescent tetra-GRABs at room temperature for 30 min with rotation. After washing twice with PBS, the cell suspension was then incubated with vehicle, 10 mM of Neu5Ac, 10 mM of Neu5,9Ac2, or 10 mM of CMP-Neu5Ac in PBS for 1 h, followed by washing with PBS for another round before flow cytometry analysis.
Cytotoxicity assay
Cells of various types were incubated with 1 μg/mL of tetra-GRAB for 30 min. Then, the cell viability was assayed by using trypan blue staining and analyzed by a TC20 cell counter (Bio-rad).
Cell proliferation assay
HeLa cells were firstly dissociated with trypsin and resuspended in cell medium. After two times of washing with PBS, cells were incubated with or without 1 μg/mL of tetra-GRAB for 30 min. Then, cells were resuspended in cell medium and seeded to a 96-well plate with 2500 cells per well. Cell proliferation was measured by the Cell Counting Kit-8 assay (Yeason) at 24, 48, and 72 h.
Confocal fluorescence microscopy
For cell imaging, cells were fixed with 4% PFA for 15 min at room temperature and washed with PBS for two times. Subsequently, cells were incubated with 1 μg/mL of fluorescent tetra-GRABs for 1 h at room temperature. Then, the cells were washed twice with PBS and stained with Hoechst 33342 (5 μg/mL) for 15 min, followed by microscopy analysis. For imaging intracellular sialoglycans, the cells were permeabilized with 0.1% Triton-X100 at room temperature for 20 min before staining with tetra-GRAB-Sia-AF647. Biological replicates were performed on cells cultured on different days.
For tissue imaging, frozen sections were blocked with 5% BSA in PBS and incubated with 2 μg/mL of fluorescent tetra-GRABs at 4 °C overnight, where the AF488-conjugated anti-EpCAM antibody was optionally added at the same time. For the comparison between lectins and tetra-GRABs, the kidney sections were incubated with 4 μg/mL of fluorescent tetra-GRAB, 10 μg/mL of biotinylated SNA, or 10 μg/mL of biotinylated MAL-II at room temperature for 2 h. For SNA and MAL-II staining, the sections were then washed with PBS for two times and incubated with 1 μg/mL of streptavidin-AF647 at room temperature for 2 h. After washing with PBS for two times, the sections were stained with Hoechst 33342 (5 μg/mL) for additional 15 min before fluorescence microscopy analysis. Biological replicates were performed using multiple mouse tissues from three mice independently collected and stained, before imaging analysis on different days.
Microscopy was performed on a Zeiss LSM 700 laser scanning confocal microscope or an Olympus SLIDEVIEW VS200 slideviewer.
Replicates
Detailed descriptions and rationale of replicates performed for each type of experiment are summarized in Supplementary Table 2. During the development and characterization of GRABs, we independently repeated part of the experiments in Figs. 1–4 on different days, leading to the same conclusions on the molecular properties of GRABs. Examples of repeated flow cytometry experiments are shown in Supplementary Fig. 22.
Statistical analysis
Statistical analyses were performed using GraphPad Prism software (version 7.0) or R v4.3.1. Comparisons of two groups were analyzed using a two-tailed t-test. Comparisons of multiple groups were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons test. In all figures with error bars, data are presented as mean ± s.d.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Correspondence and requests for materials should be addressed to P.D. and X.C. Crystal structures used in the study were previously reported: 3H72 and 4X47. Source data are provided with this paper.
References
Reily, C., Stewart, T. J., Renfrow, M. B. & Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 15, 346–366 (2019).
Haltiwanger, R. S. & Lowe, J. B. Role of glycosylation in development. Annu. Rev. Biochem. 73, 491–537 (2004).
Marth, J. D. & Grewal, P. K. Mammalian glycosylation in immunity. Nat. Rev. Immunol. 8, 874–887 (2008).
Pinho, S. S. & Reis, C. A. Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer 15, 540–555 (2015).
Cheng, B., Tang, Q., Zhang, C. & Chen, X. Glycan labeling and analysis in cells and in vivo. Annu. Rev. Anal. Chem. 14, 363–387 (2021).
Polonskaya, Z., Savage, P. B., Finn, M. G. & Teyton, L. High-affinity anti-glycan antibodies: challenges and strategies. Curr. Opin. Immunol. 59, 65–71 (2019).
Sterner, E., Flanagan, N. & Gildersleeve, J. C. Perspectives on anti-glycan antibodies gleaned from development of a community resource database. ACS Chem. Biol. 11, 1773–1783 (2016).
West, C. M., Malzl, D., Hykollari, A. & Wilson, I. B. H. Glycomics, glycoproteomics, and glycogenomics: an inter-taxa evolutionary perspective. Mol. Cell. Proteom. 20, 100024 (2021).
Tarp, M. A. et al. Identification of a novel cancer-specific immunodominant glycopeptide epitope in the MUC1 tandem repeat. Glycobiology 17, 197–209 (2007).
Weintraub, A. Immunology of bacterial polysaccharide antigens. Carbohydr. Res. 338, 2539–2547 (2003).
Avci, F. Y., Li, X., Tsuji, M. & Kasper, D. L. A mechanism for glycoconjugate vaccine activation of the adaptive immune system and its implications for vaccine design. Nat. Med. 17, 1602–1609 (2011).
Sharon, N. & Lis, H. History of lectins: from hemagglutinins to biological recognition molecules. Glycobiology 14, 53R–62R (2004).
Van Damme, E. J. M. 35 years in plant lectin research: a journey from basic science to applications in agriculture and medicine. Glycoconj. J. 39, 83–97 (2022).
Rini, J. M., Moremen, K. W., Davis, B. G. & Esko, J. D. Glycosyltransferases and glycan-processing enzymes. in Essentials of Glycobiology (Cold Spring Harbor Laboratory Press, 2022).
Cui, T. et al. Glycoenzyme tool development: principles, screening methods, and recent advances †. Chin. J. Chem. 40, 746–758 (2022).
Lopez Aguilar, A. et al. Tools for studying glycans: recent advances in chemoenzymatic glycan labeling. ACS Chem. Biol. 12, 611–621 (2017).
Griffin, M. E. & Hsieh-Wilson, L. C. Tools for mammalian glycoscience research. Cell 185, 2657–2677 (2022).
Selvan, N. et al. A mutant O-GlcNAcase enriches Drosophila developmental regulators. Nat. Chem. Biol. 13, 882–887 (2017).
Shon, D. J. et al. An enzymatic toolkit for selective proteolysis, detection, and visualization of mucin-domain glycoproteins. Proc. Natl. Acad. Sci. USA 117, 21299–21307 (2020).
Chen, X. & Varki, A. Advances in the biology and chemistry of sialic acids. ACS Chem. Biol. 5, 163–176 (2010).
Pillai, S., Netravali, I. A., Cariappa, A. & Mattoo, H. Siglecs and immune regulation. Annu. Rev. Immunol. 30, 357–392 (2012).
van de Wall, S., Santegoets, K. C. M., van Houtum, E. J. H., Büll, C. & Adema, G. J. Sialoglycans and siglecs can shape the tumor immune microenvironment. Trends Immunol. 41, 274–285 (2020).
Duan, S. & Paulson, J. C. Siglecs as immune cell checkpoints in disease. Annu. Rev. Immunol. 38, 365–395 (2020).
Crocker, P. R., Paulson, J. C. & Varki, A. Siglecs and their roles in the immune system. Nat. Rev. Immunol. 7, 255–266 (2007).
Bai, X., Brown, J. R., Varki, A. & Esko, J. D. Enhanced 3-O-sulfation of galactose in Asn-linked glycans and Maackia amurenesis lectin binding in a new Chinese hamster ovary cell line. Glycobiology 11, 621–632 (2001).
Rillahan, C. D. & Paulson, J. C. Glycan microarrays for decoding the glycome. Annu. Rev. Biochem. 80, 797–823 (2011).
Bojar, D. et al. A useful guide to lectin binding: machine-learning directed annotation of 57 unique lectin specificities. ACS Chem. Biol. 17, 2993–3012 (2022).
Nagae, M. et al. Crystal structure of anti-polysialic acid antibody single chain Fv fragment complexed with octasialic acid: Insight into the binding preference for Polysialic acid. J. Biol. Chem. 288, 33784–33796 (2013).
Matsumura, R. et al. Novel antibodies reactive with sialyl Lewis X in both humans and mice define its critical role in leukocyte trafficking and contact hypersensitivity responses. J. Biol. Chem. 290, 15313–15326 (2015).
Prendergast, J. M. et al. Novel anti-Sialyl-Tn monoclonal antibodies and antibody-drug conjugates demonstrate tumor specificity and anti-tumor activity. mAbs 9, 615–627 (2017).
Yang, L. et al. Sialic acid binding polypeptide. US patent 11434479 (2022).
Xu, G. et al. Three Streptococcus pneumoniae Sialidases: three different products. J. Am. Chem. Soc. 133, 1718–1721 (2011).
Coines, J., Raich, L. & Rovira, C. Modeling catalytic reaction mechanisms in glycoside hydrolases. Curr. Opin. Chem. Biol. 53, 183–191 (2019).
Gut, H., Xu, G., Taylor, G. L. & Walsh, M. A. Structural Basis for Streptococcus pneumoniae NanA Inhibition by Influenza Antivirals Zanamivir and Oseltamivir Carboxylate. J. Mol. Biol. 409, 496–503 (2011).
Owen, C. D. et al. Streptococcus pneumoniae NanC: structural insights into the specificity and mechanism of a sialidase that produces a sialidase inhibitor. J. Biol. Chem. 290, 27736–27748 (2015).
Tailford, L. E. et al. Discovery of intramolecular trans-sialidases in human gut microbiota suggests novel mechanisms of mucosal adaptation. Nat. Commun. 6, 1–12 (2015).
Beckett, D., Kovaleva, E. & Schatz, P. J. A minimal peptide substrate in biotin holoenzyme synthetase‐catalyzed biotinylation. Protein Sci. 8, 921–929 (1999).
Christophersen, A. Peptide‐MHC class I and class II tetramers: from flow to mass cytometry. HLA 95, 169–178 (2020).
Rodrigues, E. et al. A versatile soluble siglec scaffold for sensitive and quantitative detection of glycan ligands. Nat. Commun. 11, 5091 (2020).
Achilli, S. et al. TETRALEC, artificial tetrameric lectins: a tool to screen ligand and pathogen interactions. Int. J. Mol. Sci. 21, 5290 (2020).
Oyelaran, O. & Gildersleeve, J. C. Glycan arrays: recent advances and future challenges. Curr. Opin. Chem. Biol. 13, 406–413 (2009).
Schjoldager, K. T., Narimatsu, Y., Joshi, H. J. & Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 21, 729–749 (2020).
Stotter, B. R. et al. Cosmc-dependent mucin-type O-linked glycosylation is essential for podocyte function. Am. J. Physiol. Physiol. 318, F518–F530 (2020).
Hennet, T. Collagen glycosylation. Curr. Opin. Struct. Biol. 56, 131–138 (2019).
Luis, A. S. & Hansson, G. C. Intestinal mucus and their glycans: a habitat for thriving microbiota. Cell Host Microbe 31, 1087–1100 (2023).
Brazil, J. C. & Parkos, C. A. Finding the sweet spot: glycosylation mediated regulation of intestinal inflammation. Mucosal Immunol. 15, 211–222 (2022).
Yao, Y. et al. Mucus sialylation determines intestinal host-commensal homeostasis. Cell 185, 1172–1188.e28 (2022).
Kudelka, M. R., Stowell, S. R., Cummings, R. D. & Neish, A. S. Intestinal epithelial glycosylation in homeostasis and gut microbiota interactions in IBD. Nat. Rev. Gastroenterol. Hepatol. 17, 597–617 (2020).
Bergstrom, K. et al. Proximal colon-derived O-glycosylated mucus encapsulates and modulates the microbiota. Science 370, 467–472 (2020).
Liao, G., Zhou, Z., Suryawanshi, S., Mondal, M. A. & Guo, Z. Fully synthetic self-adjuvanting a-2,9-oligosialic acid based conjugate vaccines against group C meningitis. ACS Cent. Sci. 2, 210–218 (2016).
Martínez-Bailén, M., Rojo, J. & Ramos-Soriano, J. Multivalent glycosystems for human lectins. Chem. Soc. Rev. 52, 536–572 (2022).
Cheng, B., Xie, R., Dong, L. & Chen, X. Metabolic remodeling of cell-surface sialic acids: principles, applications, and recent advances. ChemBioChem 17, 11–27 (2016).
Zeng, Y., Ramya, T. N. C., Dirksen, A., Dawson, P. E. & Paulson, J. C. High-efficiency labeling of sialylated glycoproteins on living cells. Nat. Methods 6, 207–209 (2009).
Wen, L. et al. Two-step chemoenzymatic detection of N-acetylneuraminic acid−α(2-3)-galactose glycans. J. Am. Chem. Soc. 138, 11473–11476 (2016).
Babulic, J. L. et al. One-step selective labeling of native cell surface sialoglycans by exogenous α2,8-sialylation. ACS Chem. Biol. 18, 2418–2429 (2023).
Armstrong, Z., Rahfeld, P. & Withers, S. G. Discovery of new glycosidases from metagenomic libraries. in Methods in Enzymology Vol. 597 (Elsevier Inc., 2017).
Zechel, D. L. & Withers, S. G. Dissection of nucleophilic and acid-base catalysis in glycosidases. Curr. Opin. Chem. Biol. 5, 643–649 (2001).
Rauvolfova, J., Venot, A. & Boons, G. J. Chemo-enzymatic synthesis of C-9 acetylated sialosides. Carbohydr. Res. 343, 1605–1611 (2008).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Hsiao, Y. S., Parker, D., Ratner, A. J., Prince, A. & Tong, L. Crystal structures of respiratory pathogen neuraminidases. Biochem. Biophys. Res. Commun. 380, 467–471 (2009).
Acknowledgements
This project is supported by the National Natural Science Foundation of China (No. 22377002 to P.D., No. 22250010 and No. 92153301 to X.C.) and the Beijing Natural Science Foundation (No. 5244034 to Q.T.). P.D. is a recipient of Li Ge-Zhao Ning Life Science Research Foundation for Young Scholars, Young Elite Scientists Sponsorship Program by CAST (China Association for Science and Technology), Clinical Medicine Plus X—Young Scholars Project from Peking University (PKU2023LCXQ002) and National High Level Hospital Clinical Research Funding, Interdisciplinary Research Project of Peking University First Hospital (2023IR14). Q.T. is supported by the National Postdoctoral Program for Innovative Talent. X.C. is a recipient of the Xplorer Prize. We thank the Flow Cytometry Core at the National Center for Protein Sciences at Peking University, particularly Dr. Liying Du for technical help.
Author information
Authors and Affiliations
Contributions
P.D. and X.C. conceived the project; S.L. and Q.T. performed most of the experiments. X.G. helped with the tissue imaging. Z.L. helped with the ITC analysis. Y.G. and J.S. helped with the flow cytometry analysis. J.C. helped with the HPAEC analysis. B.C. provided Neu5,9Ac2. Q.S. and J.S. helped with the protein purification. S.L., Q.T., P.D., and X.C. analyzed the data and wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
A Chinese patent application (application no. 202311031342.5) covering the sequence and use of GRAB-Sia and GRAB-Sia3 has been filed in which Peking University is the applicant with X.C., P.D., S.L., and Q.T. being the inventors. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, S., Tang, Q., Guo, X. et al. Mutant glycosidases for labeling sialoglycans with high specificity and affinity. Nat Commun 16, 1427 (2025). https://doi.org/10.1038/s41467-025-56629-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56629-9
This article is cited by
-
Engineering glycosyltransferases into glycan binding proteins using a mammalian surface display platform
Nature Communications (2025)