Abstract
The proximal domains of mitochondria and the endoplasmic reticulum (ER) are linked by tethering factors on each membrane, allowing the efficient transport of substances, including lipids and calcium, between them. However, little is known about the regulation and function of mitochondria-ER contacts (MERCs) dynamics under mitochondrial damage. In this study, we apply NanoBiT technology to develop the MERBiT system, which enables the measurement of reversible MERCs formation in living cells. Analysis using this system suggests that induction of mitochondrial ROS increases MERCs formation via RMDN3 (also known as PTPIP51)-VAPB tethering driven by RMDN3 phosphorylation. Disruption of this tethering caused lipid radical accumulation in mitochondria, leading to cell death. The lipid radical transfer activity of the TPR domain in RMDN3, as revealed by an in vitro liposome assay, suggests that RMDN3 transfers lipid radicals from mitochondria to the ER. Our findings suggest a potential role for MERCs in cell survival strategy by facilitating the removal of mitochondrial lipid radicals under mitochondrial damage.
Similar content being viewed by others
Introduction
Previous studies suggested that cellular organelles play independent biological roles, but increasing evidence has shown that organelles are not actually independent structures, and that there are physical connections between them1,2. The mitochondria-endoplasmic reticulum (ER) contacts (MERCs) are specific regions where the ER is physically associated with mitochondria3. Several MERC tethering factors have been identified, including MFN1/2-MFN24,5,6, RMDN3/PTPIP51-VAPB7, PDZD8-FKBP88,9, and IP(3)R-VDAC10, and their functional differences and regulatory mechanisms are gradually being elucidated11. MERCs are known to serve as scaffolds for a variety of responses, including lipid metabolism, calcium homeostasis, apoptosis, autophagy, mitochondrial quality control, the unfolded protein response, and ER stress11,12. However, much of the relationship between MERC dynamics and intracellular signaling responses remains unknown.
The most common methods for quantifying MERCs are transmission electron microscopy (TEM), proximity ligation assay (PLA), and the use of split-GFP reconstituted proteins; TEM and PLA require fixation, while split-GFP systems are less reversible, making it difficult to analyze the dynamic behavior of MERC formation in real time12. In recent years, technological developments in protein-protein interaction (PPI) detection have progressed, and methods for the comprehensive analysis of organelle contact factors in living cells using APEX and Turbo-ID have been successively developed13,14. In this study, we focused on NanoBiT (NanoLuc Binary Technology)15 to detect the reversible dynamics of MERCs in living cells. NanoBiT is composed of two subunits, LgBiT (18 kDa) and SmBiT (1.3 kDa), which are split fragments of a luciferase from deep-sea shrimp. LgBiT and SmBiT are optimized for the analysis of reversible PPIs (KD = 190 μM) and are so highly luminescent that they can easily reveal MERCs in living cells. These tools allow us to detect MERCs at high throughput and to analyze the more physiological dynamic behavior of ER-mitochondria contacts.
Mitochondria are the major organelles responsible for the production of reactive oxygen species (ROS)16. A total of 0.2% to 2.0% of oxygen molecules consumed in mitochondria are reduced to superoxide anions and converted to other ROS such as hydrogen peroxide (H2O2) and hydroxyl radicals (OH•)17,18. Hydroxyl radicals (OH•) are short-lived but highly reactive and promote oxidative damage to DNA, protein denaturation, and peroxidation of lipids including phospholipids19,20. Intracellular hydroxyl radicals (OH•) are generated primarily by the Fenton reaction from iron (Fe2+) and copper (Cu2+) with H2O221. Glutathione Peroxidase 4 (GPx4) is the most well-characterized enzyme involved in the direct reduction of lipid hydroperoxides and is thought to function in the cytoplasm and mitochondria, although little is known about the function of mitochondrial GPx422,23. Mitochondria-derived ROS are mainly generated in the electron transfer chain during the oxidative phosphorylation process24. During electron transfer, electrons leak out and interact with oxygen molecules to form superoxide (O2–)25. The major sources of superoxide and hydrogen peroxide in mitochondria are the flavin mononucleotide (FMN) sites in complex I and the Q cycle in complex III25. To respond to the toxicity of ROS, mitochondria have several antioxidant enzyme systems that scavenge ROS as soon as they are generated; for example, because it is highly toxic, superoxide is catalyzed by the superoxide dismutase (SOD) family and then rapidly degraded by specific enzymes such as GSH-Px and by the intervention of glutathione (GSH)26.
A recent study discovered a mitochondria-specific GSH transporter, SLC25A39, which is thought to efficiently scavenge mitochondrial ROS by increasing mito-GSH27. Meanwhile, hydropersulfides including glutathione persulfide, which contain excess sulfur added to the thiol (SH) group of cysteine, have been shown to react efficiently with ROS28. Glutathione persulfide is synthesized from cysteine persulfide by cysteinyl-tRNA synthetases (CARSs); in particular, CARS2 synthesizes a cysteine persulfide in the mitochondrial matrix29. Although accumulating evidence indicates the existence of a mitochondrial ROS scavenging system, the machinery for the removal of mitochondrial lipid peroxides is largely unknown.
ER-mitochondrial contact is thought to modulate ROS-mediated signaling30,31. Redox-related enzymes such as calnexin, Ero1α, and selenon/SEPN1 have been reported to localize and function in MERCs31. In addition, mitochondrial ROS are released into MERCs through the opening of the mitochondrial permeability transition pore (mPTP), leading to an oxidative environment within them30, although the effect of mitochondrial ROS on MERC formation remains unclear.
In this study, based on analysis using the NanoBiT system, we found that mitochondrial ROS generated by the inhibition of mitochondrial respiratory complex I/III or inhibition of glutathione persulfide specifically increased MERC formation via RMDN3-VAPB tethering. Furthermore, this RMDN3-VAPB-mediated ER-mitochondria tethering was shown to be involved in cell survival through the RMDN3-dependent transfer of lipid radicals from mitochondria to the ER. We propose a function for MERCs in attenuating cell death caused by mitochondrial lipid radical toxicity under conditions of mitochondrial damage.
Results
Assay system for reversible MERCs quantification – MERBiT
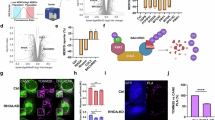
In this study, we applied the NanoBiT system to quantify reversible MERC formation in living cells. A retroviral system was employed to stably express the V5-tagged SmBiT fragment conjugated to the C-terminus of TOMM20 (full length), a mitochondrial outer membrane protein, and the 3×HA-tagged LgBiT fragment conjugated to the N-terminus of Sec61β (full length), an ER-localized protein, in HeLa cells (Fig. 1a). We named this system “MERBiT” (Mitochondria-Endoplasmic Reticulum contact reporter applying NanoBiT) and HeLa cells expressing the MERBiT system “MERBiT cells” (Fig. 1b). Immunostaining analysis revealed the colocalization of TOMM20-V5-SmBiT with the mitochondrial marker HSP60 and LgBiT-Sec61β-3×HA with the ER marker calnexin (Fig. 1c). Western blot analysis confirmed that TOMM20-V5-SmBiT and LgBiT-Sec61β-3×HA were correctly expressed in HeLa cells (Fig. 1d). The MERBiT cells showed a strong luminescence signal compared with both wild-type HeLa cells and HeLa cells stably expressing only TOMM20-V5-SmBiT (Fig. 1e). Our findings also showed that the MERBiT system does not affect the formation of MERCs (Supplementary Fig. 1a, b). When the MERBiT cells were subjected to HBSS starvation, which is known to increase ER-mitochondria contacts32, the luminescence was significantly increased (Fig. 1f). Furthermore, when the cells were replaced with normal medium (10% FBS-containing DMEM) after HBSS stimulation, the luminescence decreased to basal levels after 20 min (Fig. 1f). To investigate the effects of known ER-mitochondria tethering factors on MERBiT luminescence, we knocked down MFN2, PDZD8, RMDN3, and VAPB and observed a decrease in luminescence due to the suppression of each factor (Fig. 1g, h). Simultaneous knockdown of MFN2, PDZD8, RMDN3, and VAPB showed that some double-knockdown combinations had an additive effect in reducing MERC formation (Supplementary Fig. 1c). In contrast to the knockdown experiments, overexpression of MFN2, PDZD8, RMDN3, and VAPB increased MERC formation (Supplementary Fig. 1d). In addition, an increase in luminescence was observed when MERCs were artificially induced by expression of the MERC linker (Fig. 1i, Supplementary Fig. 1e–g). These results indicate that the MERBiT system can monitor the reversible dynamics of MERCs in living cells.
a Schematic of the construct and strategy for detection of MERCs. b Schematic of the MERBiT system. c Representative images of V5-TOMM20-SmBiT and LgBiT−3×HA-Sec61β localization in HeLa cells stably expressing V5-TOMM20-SmBiT and LgBiT−3×HA-Sec61β (MERBiT cells). Cells were stained with V5, HA, HSP60 and calnexin antibodies. HSP60 is used as a mitochondrial marker and calnexin is used as an ER marker. d, Representative immunoblots for each component of MERBiT cells. The lysates of MERBiT cells were analyzed by immunoblotting for V5 (V5-TOMM20-SmBiT), HA (LgBiT-3×HA-Sec61β), TOMM20, HSP60, calnexin, and α-tubulin. Black and white arrowheads indicate tagged and endogenous TOMM20, respectively. e Luminescence of MERBiT cells. Quantification of the luminescence of HeLa cells, MERBiT cells, and stably expressing V5-TOMM20-SmBiT HeLa cells. Data are mean ± s.e.m. (n = 9). f Quantification of MERCs reduction during recovery from starvation in MERBiT cells. Cells were starved in HBSS for 1 h and then recovered in 10% FBS DMEM for the indicated times before luminescence was measured. Data are mean ± s.e.m. (n = 3, triplicate). g, h Effects of knockdown of different MERCs tethering factors on MERBiT luminescence in MERBiT cells. Cells were transfected with the indicated siRNAs and then luminescence was measured or WB was performed with the indicated antibodies to confirm protein expression levels. Data are mean ± s.e.m. (n = 3, triplicate). i MERCs linker increases luminescence. MERBiT cells were transfected with MERCs linker (pCAG-AKAP1(1-30 aa)-mTagBFP-V5-SACM1L (521-587 aa)) and luminescence was detected. Data are mean ± s.e.m. (n = 3, triplicate). Statistical significance was analyzed by one-way analysis of variance (ANOVA) (e, f, g) or Student’s t-test, Two-tailed (i). P values are indicated as; **p < 0.01; ****p < 0.0001.
Induction of mitochondrial ROS increase MERCs formation
It has been reported that HBSS stimulation induces mitochondrial elongation33. To examine whether mitochondrial elongation affects MERC formation, we measured luminescence after the knockdown of Drp1, a mitochondrial fission factor. Consistent with a previous report34 and in contrast to the findings under HBSS starvation, Drp1 knockdown reduced MERC formation (Supplementary Fig. 2a–c). Next, to examine the effect of mitochondrial dysfunction on MERCs, we treated the MERBiT cells with CCCP and valinomycin, mitochondrial oxidative phosphorylation uncouplers; oligomycin, a mitochondrial respiratory chain V inhibitor; rotenone, a mitochondrial respiratory chain I inhibitor; and antimycin A, a mitochondrial respiratory chain III inhibitor. Treatment with CCCP, valinomycin, and oligomycin had no effect on MERCs, whereas treatment with rotenone or antimycin A, which does not affect NanoLuc luminescence (Supplementary Fig. 2e), increased MERCs (Fig. 2a–c, Supplementary Fig. 2d). Stimulation with rotenone or antimycin A is known to produce excessive mitochondrial ROS35,36. Treatment with Mito-TEMPO, a mitochondria-specific ROS scavenger37, markedly suppressed the induction of MERCs by rotenone and antimycin A treatment (Fig. 2a), suggesting that mitochondrial ROS, but not OXPHOS changes, are the trigger inducing MERC formation. To obtain further insight into the relationship between ROS and MERC formation, MERBiT cells were treated with rotenone and antimycin A with or without the oxidized form of N-acetylcysteine (oxNAC) and N-acetylcysteine tetrasulfide (NACS2)38, synthetic sources of glutathione and persulfidated glutathione (Fig. 2d, e). The addition of oxNAC and NACS2 suppressed the formation of mitochondrial ROS-derived MERCs (Fig. 2d, e). Furthermore, MERCs were also increased by treatment with tBHP, an inducer of oxidative stress, and this increase in MERCs was suppressed by the addition of oxNAC and NACS2 (Supplementary Fig. 2f). Based on these results, we hypothesized that glutathione, especially glutathione persulfide, could strongly suppress mitochondrial ROS, which promotes the formation of MERCs. Consistent with this, knockdown of CARS2, an enzyme critical for the production of endogenous Cys-SSH and persulfide, and SLC25A39, a mitochondria-specific glutathione transporter, increased mitochondria and ER colocalization and MERBiT luminescence, which was suppressed by Mito-TEMPO (Fig. 2f, Supplementary Fig. 2g–i and Fig. 3a–c). Furthermore, mitochondrial ROS-derived MERCs were not suppressed by the addition of oxNAC, but NACS2 could only compensate for CARS2 knockdown, not SLC25A39 knockdown (Supplementary Fig. 3d), suggesting that glutathione persulfide, but not glutathione, is critical for scavenging mitochondrial ROS that promote MERC formation. Although Trx2 and Prx3 also localize to mitochondria and inhibit mitochondrial ROS, knockdown of both proteins did not affect MERBiT luminescence (Supplementary Fig. 3e). To demonstrate that mitochondrial ROS specifically induce MERC formation, a mitochondrial matrix-targeted D-amino acid oxidase (DAAO) construct (mitoDAAO) and a non-targeted construct (CytoDAAO) were generated and compared for H2O2-induced MERC formation (Fig. 2g, h)39,40. D-Amino acid oxidase (DAAO) catalyzes the oxidative deamination of D-amino acids (stereoisomers of naturally occurring L-amino acids), producing 2-oxo acids, hydrogen peroxide (H2O2), and ammonia. The results obtained using this tool showed that mitochondrial ROS generation, but not cytoplasmic ROS, promoted the formation of MERCs (Fig. 2h). Finally, using a flow cytometer, we divided MERBiT cells into those with high or low mitochondrial ROS and sorted each population (Fig. 2i). The mitochondrial ROS-high MERBiT cells formed a high number of MERCs (Fig. 2j), which could be abrogated by NAC treatment (Fig. 2k), in contrast to the mitochondrial ROS-low MERBiT cells.
a Mitochondrial ROS produced by rotenone and antimycin A treatment increases luminescence in MERBiT cells. The cells were treated with rotenone (50 nM) and antimycin A (50 nM) with or without Mito-TEMPO for 1 h and then luminescence was measured. b Antimycin A and rotenone stimulation increases MERCs in EM imaging. Representative electron micrographs of HeLa cells treated with or without antimycin A (50 nM) or rotenone (50 nM) for 1 h. Mitochondria and ER were labeled with green and magenta, respectively. ER-mitochondria contact sites are indicated by dashed white lines ( < 25 nm distance between membranes). c Quantification of MERCs length normalized to mitochondrial circumference. Data are mean ± s.e.m. n = 30 cells for control, antimycin A treatment, and rotenone treatment in HeLa cells. d, e oxNAC and NACS2 treatment reduces MERCs formation by removing excessive ROS in MERBiT cells. The cells were treated with rotenone (50 nM) and antimycin A (50 nM) with or without oxNAC (30 µM) and NACS2 (30 µM) for 1 h and then luminescence was measured. oxNAC and NACS2 were treated for 6 h before addition of rotenone and antimycin A. f Mitochondrial ROS production by CARS2 and SLC25A39 knockdown increases MERCs formation in MERBiT cells. The cells were transfected with indicated siRNAs for 3 days and treated with or without Mito-TEMPO for 1 h before luminescence measurement. g, h Mitochondrial, but not cytosolic, H2O2 generation increases MERCs formation in MERBiT cells. MERBiT cells were transfected with the cytoDAAO and mitoDAAO expressing constructs for 1 day and treated with or without D-Met (1 mM) for 1 h before luminescence measurement. i, j The MERBiT cells with higher mitochondrial ROS production form more MERCs than cells with lower mitochondrial ROS production. The cells were stained with MitoSOX 10 µM for 30 min and the MitoSOX high or low MERBiT cells were sorted by a cell sorter. Then the reseeded cells were measured for luminescence. k NAC treatment decreases MERCs formation in MERBiT cells and mitochondrial ROS high MERBiT cells. The cells or mitochondrial ROS high cells were treated with NAC for 2 days and then luminescence was measured. Data are mean ± s.e.m. (n = 3, triplicate), and statistical significance was analyzed by one-way analysis of variance (ANOVA) (a, c, e, f, h, j, k). P values are indicated as **p < 0.01; ***p < 0.001; ****p < 0.0001; n.s., not significant.
a RMDN3 and VAPB are critical tethering factors for MERCs formation induced by antimycin A and rotenone stimulation. MERBiT cells were transfected with the indicated siRNAs for 3 days and treated with or without rotenone (50 nM) and antimycin A (50 nM) for 1 h before luminescence measurements. Data are mean ± s.e.m. (n = 3, triplicate). b Interaction between RMDN3 and VAPB increase in rotenone and antimycin A treatment. HeLa cells were transfected with the indicated vectors and treated with rotenone (50 nM) or antimycin A (50 nM) for 1 h. Cell lysates were subjected to IP assay (left). Ratio of RMDN3-VAPB interaction (right). Data are mean ± s.e.m. (n = 3). c Schematic model of the RMDN3 domain. d The expression levels of RNAi-resistant RMDN3 vectors. The HeLa cells were transfected with RMDN3 siRNA for 2 days and then transfected with the indicated vectors such as RMDN3 RNAi-resistant vectors for 1 day. e FFAT but not TPR domain is important for mitochondrial ROS-induced MERCs formation. The MERBiT cells were transfected with the indicated siRNAs for 2 days. Then transfected with empty vectors or indicated RNAi-resistant vectors for 1 day. Before measuring luminescence, cells were treated with or without rotenone (50 nM) and antimycin A (50 nM) for 1 h. Data are mean ± s.e.m. (n = 3, triplicate). f Threonine 160 mutant of RMDN3 decrease phosphorylation by antimycin A stimulation. HeLa cells were transfected with the indicated vectors and then treated with or without antimycin A (50 nM). Cell lysates were subjected to IP assay and then beads were incubated with or without lambda phosphatase (λPP). Pull-down lysates were subjected to Phos-tag-PAGE or SDS-PAGE. g Phosphorylation of RMDN3 T160 was important for interaction with VAPB by antimycin A treatment. HeLa cells were transfected with the indicated vectors and treated with antimycin A (50 nM) for 1 h. Cell lysates were subjected to IP assay and IB assay (left). Ratio of RMDN3-VAPB interaction (right). Data are mean ± s.e.m. (n = 3). h Phosphorylation of RMDN3 T160 is important for MERCs formation induced by antimycin A stimulation. The MERBiT cells were transfected with the indicated siRNAs for 2 days and then transfected with vectors for 1 day before treatment with or without antimycin A (50 nM) for 1 h, and then the luminescence was measured. Data are mean ± s.e.m. (n = 3, triplicate). Statistical significance was analyzed by one-way analysis of variance (ANOVA) (a, b, e, g, h). P values are indicated as *p < 0.05; **p < 0.01; ****p < 0.0001; n.s., not significant.
Mitochondrial ROS-derived MERCs depend on phosphorylated RMDN3 and VAPB binding
To identify the tethering factor involved in the formation of mitochondrial ROS-derived MERCs, we measured the luminescence upon the knockdown of four known tethering factors. Mitochondrial ROS-derived MERCs induced by rotenone and antimycin A treatment were not reduced by the knockdown of MFN2 or PDZD8, but were significantly reduced by the knockdown of RMDN3 or VAPB (Fig. 3a). Similarly, MERC formation, which was increased by the knockdown of CARS2 or SLC25A39, was also reduced by the knockdown of RMDN3 or VAPB (Supplementary Fig. 4a). Immunoprecipitation assay showed that RMDN3-VAPB binding was increased under conditions of rotenone and antimycin A treatment or CARS2 and SLC25A39 silencing (Fig. 3b, Supplementary Fig. 4b), indicating that the increase in mitochondrial ROS-derived MERC formation is achieved by specific RMDN3-VAPB binding. Correlating with these results, RMDN3 expressed in mitochondrial ROS-high cells (F954 cells) derived from a patient with the mitochondrial disease MELAS syndrome carrying the m.3243 A > G mutation (92% level of heteroplasmy) was found to bind VAPB more strongly than that in mitochondrial ROS-low cells or normal fibroblast cells (NHDF cells) (Supplementary Fig. 4c). RMDN3 contains an FFAT-like motif7, which binds to the major sperm protein (MSP) domain conserved among VAPA, VAPB, and possibly MOSPDs41,42,43, and a tetratricopeptide repeat (TPR) domain with lipid binding and lipid transfer activities44 (Fig. 3c). There was a difference between the decrease in luminescence upon silencing RMND3 and that upon silencing VAPB, as shown in Fig. 3a. For confirmation, we performed an experiment involving the double silencing of VAPA and VAPB and confirmed its additive effect on decreasing MERC formation, suggesting that VAPA is also involved in mitochondrial ROS-derived MERC formation (Supplementary Fig. 4d). To examine whether the FFAT-like motif and TPR domain are involved in the formation of mitochondrial ROS-derived MERCs, we generated RNAi-resistant RMDN3 constructs: resi-wild-type RMDN3, resi-ΔFFAT RMDN3, and resi-ΔTPR RMDN3 (Fig. 3d). Consistent with previous reports, the TPR domain did not affect VAPB interaction and its localization (Supplementary Fig. 4e, f). Although the expression of both resi-wild-type RMDN3 and resi-ΔTPR RMDN3 with si-RMDN3 rescued mitochondrial ROS-derived MERC formation, the expression of resi-ΔFFAT RMDN3 did not (Fig. 3e, Supplementary Fig. 4g). Next, we investigated the molecular signaling response to promote MERC formation. Since previous reports showed that the phosphorylation of RMDN3 FFAT-like motif regulates interaction with the VAPB MSP domain, we hypothesized that mitochondrial ROS promote RMDN3 FFAT phosphorylation and VAPB interaction. As expected, mitochondrial ROS induced by antimycin A treatment increased the phosphorylation of RMDN3, possibly the FFAT-like motif at threonine 160 (T160), a phospho-site previously reported to be essential for interaction with VAPB41 (Fig. 3f, Supplementary Fig. 4h). Expression of the RMDN3 T160A mutant fails to enhance both VAPB interaction and antimycin A-induced MERC formation, indicating that RMDN3 T160 phosphorylation enhances VAPB interaction and is essential for mitochondrial ROS-induced MERC formation. (Fig. 3g, h). Finally, we analyzed the effect of RMDN3-dependent MERCs formation, which is promoted by antimycin A and rotenone stimulation, on calcium transfer to mitochondria using calcium indicator CEPIA245. Consistent with previous report46, suppression of RMDN3 and VAPB reduced histamine-induced calcium uptake into mitochondria (Supplementary Fig. 4i, j). Furthermore, in the suppression of RMDN3, the calcium uptake was reduced during antimycin A and rotenone stimulation compared to unstimulated cells (Supplementary Fig. 4k). These results suggest that mitochondrial ROS trigger RMDN3 T160 phosphorylation and increase VAPB interaction to promote MERC formation.
Disruption of mitochondrial ROS-derived MERCs reduces cell viability
To understand the physiological significance of mitochondrial ROS-induced MERCs, the effects of knockdown of tethering factors on cell viability were examined. We found that knockdown of RMDN3 and VAPB, but not MFN2 and PDZD8, significantly reduced cell viability upon long-term treatment with rotenone and antimycin A or by silencing CARS2 and SLC25A39 (Fig. 4a, Supplementary Fig. 5a). As in the case of MERC formation (Fig. 2e, Supplementary Fig. 3d), cell death was suppressed by the addition of NACS2, but not oxNAC, following CARS2 and RMDN3 knockdown (Fig. 4b, Supplementary Fig. 5b). This suggests that mitochondrial ROS-induced tethering of RMDN3-VAPB is critical for inhibiting cell death.
a Rotenone or antimycin A treatment of RMDN3 and VAPB knockdown cells reduced cell viability. The HeLa cells were transfected with the indicated siRNAs for 5 days with or without rotenone (50 nM) and antimycin A (50 nM) for 2 days before cell viability was measured. Cell viability was determined and expressed as a fold change of si-NT. Viable cells were detected by cell viability assay using Cell Counting Kit-8. b The HeLa cells were transfected with the indicated siRNAs for 5 days with or without rotenone (50 nM) and antimycin A (50 nM) for 2 days before measuring cell viability. oxNAC (50 µM), NACS2 (50 µM) and mito-TEMPO (100 nM) were treated for 3 days before measuring cell viability. Cell viability was determined and expressed as a fold change of si-NT. Viable cells were detected by cell viability assay using Cell Counting Kit-8. c Lack of TPR domain does not rescue cell viability of RMDN3 knockdown with rotenone or antimycin A treatment. HeLa cells were transfected with RMDN3 siRNA for 5 days and with the indicated vectors for 3 days before measuring cell viability. Rotenone (50 nM) and antimycin A (50 nM) were treated for 2 days before cell viability was measured. Cell viability was determined and expressed as a fold change of si-NT. Viable cells were detected by cell viability assay using Cell Counting Kit-8. FLAG-resi-RMDN3 WT, FLAG-resi-RMDN3ΔFFAT, and FLAG-resi-RMDN3ΔTPR are RMDN3 RNAi-resistant vectors. d HeLa cells were transfected with RMDN3 siRNA for 5 days with or without rotenone (50 nM) and antimycin A (50 nM) for 2 days before measuring cell viability. The indicated inhibitors were treated for 2 days before measuring cell viability. Cell viability was determined and expressed as a fold change of si-NT. Viable cells were detected by cell viability assay using Cell Counting Kit-8. z-VAD-FMK (20 µM), necrostatin-1 (20 µM), ferrostatin-1 (Fer-1) (5 µM), and deferoxamine (DFO) (100 µM). Data are mean ± s.e.m. (n = 3, triplicate), and statistical significance was analyzed by one-way analysis of variance (ANOVA) (a-d). P values are indicated as ****p < 0.0001; n.s., not significant.
To determine whether cell death is inhibited by the lipid transfer activity of RMDN3, cell viability was examined by expressing RNAi-resistant RMDN3 constructs, resi-wild-type RMDN3 and resi-ΔTPR-RMDN3, after the suppression of CARS2 and SLC25A39 or by long-term treatment of RMDN3-knockdown cells with rotenone and antimycin A (Fig. 4c, Supplementary Fig. 5c). Since the expression of resi-ΔTPR-RMDN3 and resi-ΔFFAT-RMDN3 cannot rescue cell viability in RMDN3-knockdown cells, the lipid transfer ability and binding to VAPB of RMDN3 are important for cell survival. To further identify the type of cell death induced by the inhibition of MERC formation using rotenone or antimycin A, we applied the apoptosis inhibitor Z-VAD-FMK, the necroptosis inhibitor necrostatin-1, and the ferroptosis inhibitor Fer-1; however, none of these inhibitors inhibited cell death (Fig. 4d). Meanwhile, the addition of DFO, an iron chelator, inhibited cell death, suggesting that hydroxyl radical generated by the iron-mediated Fenton reaction induces mitochondrial toxicity under MERC disruption. Taken together, these results indicate that RMDN3-VAPB tethering and the lipid transfer activity of RMDN3 inhibit mitochondrial ROS-induced cell death.
Excessive lipid peroxidation caused by MERCs disruption
ROS are known to induce lipid peroxidation and cell death47. Mitochondria are particularly rich in iron, which is thought to promote the production of hydroperoxides through the Fenton reaction48,49. Since cell death was suppressed by the iron chelator DFO (Fig. 4d) and the lipid transfer activity of RMDN3 is important for cell death suppression (Fig. 4c), we considered the possibility that cell death during the inhibition of MERC formation is triggered by lipid peroxidation in mitochondria. We examined mitochondrial peroxidation using MitoPeDPP, a mitochondria-specific lipid peroxidation detection probe50, and found that RMDN3 knockdown with CARS2 and SLC25A39 suppression increased fluorescence intensity compared with that in the control and upon RMDN3, CARS2, or SLC25A39 knockdown alone (Supplementary Fig. 6a, b). Similar results were obtained with VAPB knockdown (Supplementary Fig. 6c). Furthermore, an increase in fluorescence intensity was also observed when RMDN3-knockdown cells were treated with antimycin A and rotenone (Supplementary Fig. 6d). These increases in fluorescence could be suppressed by resi-wild-type RMDN3 expression, but not by resi-ΔTPR-RMDN3 expression (Supplementary Fig. 6e–h), suggesting that mitochondrial ROS-induced MERC formation suppresses lipid peroxidation through the lipid transfer activity of RMDN3.
RMDN3 removes lipid radicals from the mitochondria via the MERCs
Since analysis using a MitoPeDPP probe suggested that the lipid peroxides accumulate in mitochondria, we next attempted to directly observe lipid radicals using LipiRADICAL51. In the RMDN3-knockdown cells treated with rotenone or antimycin A, the LipiRADICAL signal was colocalized with MitoTrackerDeep Red, a mitochondrial marker dye, and there was decreased colocalization of ER-RFP, an ER marker (Fig. 5a–c). To further investigate whether these lipid radicals are regulated by the lipid transfer activity of RMDN3, we generated wild-type-RMDN3-IRES-EBFP and ΔTPR-RMDN3-IRES-mCherry constructs, providing a second readout system to visualize gene expression. In cells expressing wild-type RMDN3 (EBFP-positive cells), LipiRADICAL signals were absent or not colocalized with mitochondria, but partially colocalized with ER (Fig. 5d–g, Supplementary Fig. 7a–f). In contrast, the LipiRADICAL signal remained colocalized with mitochondria in cells that did not express wild-type RMDN3 (EBFP- and mCherry-negative cells) or in cells expressing ΔTPR (mCherry-positive cells) (Fig. 5d, e, Supplementary Fig. 7b, c). Furthermore, since colocalization of LipiRADICAL signal with mitochondria was also observed in cells expressing ΔFFAT (mCherry-positive cells), which is a VAPB interaction motif, MERC formation via RMDN3-VAPB is indispensable for lipid radical scavenging (Fig. 5f, g, Supplementary Fig. 7d, e). These results suggest that lipid radicals, lipid peroxyl radicals, or lipid peroxides are removed from mitochondria by the lipid transfer activity of RMDN3.
a Suppression of RMDN3 and antimycin A treatment leads to accumulation of lipid radicals on mitochondria. Representative LipiRADICAL Green staining images in HeLa cells transfected with RMDN3 siRNA for 3.5 days before staining with LipiRADICAL Green. Cells were treated with antimycin A (50 nM) for 3 h before staining with LipiRADICAL Green and also labeled with MitoTracker as mitochondrial marker. b Quantification of colocalization between LipiRADICAL Green signal and MitoTracker in relation to (a). c Quantification of colocalization between LipiRADICAL Green signal and ER-RFP as an ER marker. Cells were treated with antimycin A (50 nM) for 3 h before staining with LipiRADICAL Green. d Representative images of LipiRADICAL Green staining images in HeLa cells transfected with RMDN3 siRNA for 3.5 days and with the indicated vectors for 2 days before staining with LipiRADICAL Green. Cells were treated with Antimycin A (50 nM) for 1 day before staining with LipiRADICAL Green and MitoTracker. FLAG-resi-RMDN3WT-IRES-EBFP and FLAG-resi-RMDN3ΔTPR-mCherry are RMDN3 RNAi resistant vectors. Blue line indicates cells expressing FLAG-resi-RMDN3WT-IRES-EBFP. Red line indicates cells expressing FLAG-resi-RMDN3ΔTPR-IRES-mCherry. e Quantification of LipiRADICAL Green staining patterns from (d) following patterns in Supplementary Fig. 7a. EBFP (FLAG-resi-RMDN3WT-IRES-EBFP) and mCherry (FLAG-resi-RMDN3ΔTPR-IRES-mCherry) negative cell were counted as non-expressing cells (n = 106 cells). Resi-RMDN3WT cells were counted for EBFP-positive cell staining patterns (n = 92 cells). Resi-RMDN3ΔTPR cells were counted for mCherry-positive cell staining patterns (n = 74 cells). f Representative images of LipiRADICAL Green staining images in HeLa cells transfected with RMDN3-siRNA for 3.5 days and with the indicated vectors for 2 days before staining with LipiRADICAL Green. Cells were treated with antimycin A (50 nM) for 1 day before staining with LipiRADICAL Green and MitoTracker. FLAG-resi-RMDN3WT-IRES-EBFP and FLAG-resi-RMDN3ΔFFAT-mCherry are RMDN3 RNAi-resistant vectors. Blue line indicates cells expressing FLAG-resi-RMDN3WT-IRES-EBFP. Red line indicates cells expressing FLAG-resi-RMDN3ΔFFAT-IRES-mCherry. g Quantification of LipiRADICAL Green staining patterns from (f) following patterns in (Supplementary Fig. 7a). EBFP (FLAG-resi-RMDN3WT-IRES-EBFP) and mCherry (FLAG-resi-RMDN3ΔTPR-IRES-mCherry) negative cell were counted as non-expressing cells (n = 87 cells). Resi-RMDN3WT cells were counted as EBFP-positive cell staining patterns (n = 104 cells). Resi-RMDN3ΔFFAT cells were counted as mCherry-positive cell staining patterns (n = 106 cells).
RMDN3 TPR domain has lipid radical transfer activity
Since it has been suggested that the RMDN3 TPR domain may have lipid radical transfer activity, we investigated this using a liposomal lipid transfer assay (Fig. 6a). First, we confirmed the RMDN3 TPR domain-mediated transfer activity of TMR phosphatidic acid (PA)-derived lipid radicals generated by AAPH stimulation. The liposome assay showed that the TMR fluorescence signal of acceptor liposomes was dose-dependently increased by the RMDN3 TPR domain and that AAPH stimulation enhanced the TMR fluorescence signal, suggesting that the RMDN3 TPR domain binds to and transfers lipid radicals (Fig. 6b). To directly demonstrate the lipid radical transfer activity of the RMDN3 TPR domain, lipid radicals were labeled with LipiRADICAL Green, a lipid radical-specific probe, and the LipiRADICAL signals in acceptor liposomes were measured. We found that, only in the presence of AAPH stimulation, the LipiRADICAL signal of RMDN3 TPR domains increased in a dose-dependent manner (Fig. 6c). In addition, LipiRADICAL signals were enhanced in the presence of an acceptor and RMDN3 TPR domains compared with those in the presence of only an acceptor (Fig. 6d). These results suggest that the RMDN3 TPR domain has lipid radical transfer activity.
a Schematic of the liposome assay to monitor lipid radical transfer activity. b RMDN3 shows higher lipid transfer activity in the presence of lipid radicals. The y‐axes in the graphs show the percentage of acceptor liposome fluorescence, which was calculated using the following formula: 100×Facceptor /(Facceptor+Fdonor). c Liposome assay to measure lipid radicals transfer by RMDN3 TPR. Donor liposome (containing biotinyl-cap lipids and unsaturated lipids), which bind to streptavidin magnetic beads, were treated with or without AAPH (500 µM), a radical inducer. After washed out AAPH, acceptor liposomes and RMDN3 TPR were mixed with donor liposome-bead lysates. LipiRADICAL Green was then added to the mixture. After removing donor liposome-bead by binding to magnetic racks, the supernatant (including acceptor liposomes and RMDN3 TPR mixture) LipiRADICAL Green fluorescence signals were measured. d The RMDN3 TPR binds and transfers lipid radicals. The donor liposome (containing biotinyl-cap lipids and unsaturated lipids), which bind streptavidin magnetic beads, lysates were treated with or without AAPH (500 µM), a radical inducer. After washed out AAPH, acceptor liposomes and RMDN3 TPR or RMDN3 TPR alone were mixed with donor liposome-bead lysates. Then, LipiRADICAL Green was added to the mixture. After removing donor liposome-bead by binding to magnetic racks, the supernatant (including acceptor liposomes and RMDN3 TPR or RMDN3 TPR alone) LipiRADICAL Green fluorescence signals were measured. e Schematic illustration: induction of mitochondrial ROS promotes MERC formation, and RMDN3 transfers lipid radicals from mitochondria to the ER at MERCs to attenuate mitochondrial oxidative stress. Data are mean ± s.e.m. (n = 3, triplicate), and statistical significance was analyzed by one-way analysis of variance (ANOVA) (b–d). P values are indicated as ****p < 0.0001.
RMDN3 suppresses lipid peroxidation in primary brown adipocytes under stimulation mimicking cold exposure
To observe the suppression of lipid radical toxicity by RMDN3-VAPB tethering in physiological conditions, we focused on brown adipocytes. Brown adipocytes are known to produce mitochondrial ROS in response to cold stimuli to enhance mitochondrial heat production52. First, we evaluated whether the suppression of RMDN3 affected the differentiation of primary brown adipocytes by examining various differentiation markers and lipid droplet formation, and found that it did not affect differentiation (Fig. 7a–g). Since there was no effect on differentiation, we decided to perform an experiment mimicking cold exposure using norepinephrine (NE) stimulation after the induction of differentiation. The results show that NE treatment of primary brown adipocytes increased the signal of MitoPeDPP, which reflects the amount of lipid peroxide in mitochondria (Fig. 7h). Furthermore, RMDN3 and VAPB binding was increased during NE stimulation, and this increase was attenuated by the addition of Mito-TEMPO (Fig. 7i, j). In addition, MitoPeDPP signal was increased when NE was treated under RMDN3 suppression compared with that in control cells. Furthermore, the expression of siRNA-resistant wild-type RMDN3, but not RMDN3 T160A, in RMDN3-suppressed primary brown adipocytes attenuated the signal of MitoPeDPP (Fig. 7k), suggesting that RMDN3 binding to VAPB is increased under a cold stimulus in brown adipose tissue, which suppresses the accumulation of lipid peroxide in mitochondria.
a, b Suppression of RMDN3 does not affect the induction of thermogenic genes. Cells were transfected with the indicated siRNAs and harvested at 0 or 4 days after differentiation. The cell lysates were analyzed by immunoblotting with the indicated antibodies (a). mRNA levels of differentiation markers were measured by qRT-PCR. Data were normalized to s18 mRNA and expressed relative to si-NT on day 4 (b). c Representative images of lipid droplets (LDs) in cells treated with the indicated siRNAs. Cells were fixed on day 4. The LDs and mitochondria were labeled with LipidTOX and anti-TOMM20 antibodies. The LDs and mitochondria were quantified for total LD area in (d), average LD size in (e), total mitochondrial area in (f), and the ratio of total LD area to total mitochondrial area in (g) from the ROI of (c). 2−3 cells from three independent experiments for the control and si-RMDN3#1 cells, respectively. Data are mean ± s.e.m. (n = 3) h Lipid peroxide production increases with NE stimulation. MitoPeDPP (10 µM) was stained 30 min after stimulation with or without NE (1 µM) for 1 h and MitoPeDPP signals were detected in brown adipocytes (day 6). i, j RMDN3 and VAPB binding is increased by mitochondrial ROS generated under NE stimulation. Cell lysates were subjected to IP assay with anti-RMDN3 antibody and IB assay with the indicated antibodies (j). Ratio of RMDN3-VAPB interaction, plotted data for NE (1 µM) with or without Mito-TEMPO (10 µM) treatment versus control. Data are mean ± s.e.m. (n means three independent experiments). k Phosphorylation of RMDN3 by mitochondrial ROS and binding with VAPB is necessary for the suppression of lipid peroxide production. Cells were transfected with the indicated siRNAs and expressed human RMDN3 or human RMDN3 T160A before measuring MitoPeDPP fluorescence. NE was treated for 1 h. Data are mean ± s.e.m. (n means three independent experiments). Statistical significance was analyzed by one-way analysis of variance (ANOVA) (b, j, k) or Student’s t-test, Two-tailed (d–h). P values are indicated as *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; n.s., not significant.
Discussion
Several assay systems have been developed to measure MERC formation, using FRET, split-GFP, BiFC, and ddFP. These approaches have been used to investigate the dynamics of MERC formation in response to various pathological conditions, mitochondrial morphological alterations, and mitochondrial function53. In this study, we developed an assay system using NanoBiT technology to measure MERC dynamics in a more physiological manner, which we named “MERBiT.” This assay system can be applied not only to elucidate the mechanism of MERC formation, but also to screen for drugs that ameliorate mitochondrial dysfunction under various cellular stresses. It is thus expected to provide a versatile tool for future studies. Using MERBiT-expressing cells, we found that induction of mitochondrial ROS promote MERC formation for cell survival through enhanced RMDN3-VAPB binding. This binding ameliorates cytotoxicity caused by lipid radicals accumulated on mitochondria by promoting their retrograde transport from the mitochondria to other organelles, mainly the ER. Other MERC tethering factors, including MFN2 and PDZD8, are not involved in mitochondrial ROS-derived MERC formation and lipid radical scavenging, demonstrating that RMDN3-VAPB tethering specifically and strictly regulates MERC formation under oxidative stress. Thus, our findings reveal that MERCs are functionally targeted by specific tethering factors depending on their purpose.
Next, the molecular mechanism by which mitochondrial ROS promote the binding of RMDN3 to VAPB was investigated. It has been reported that RMDN3 binding to VAPB is enhanced by T160 phosphorylation in the FFAT domain of RMDN341. Consistent with this, we found that T160 phosphorylation is induced by mitochondrial ROS, resulting in enhanced binding of RMDN3 to VAPB. It is likely that an unidentified MERC-localized kinase activated by ROS is involved in the T160 phosphorylation of RMDN3. For example, PERK has been shown to be required at the ER-mitochondrial contact sites to mediate apoptosis following ROS-based ER stress54, suggesting that PERK is one of the candidates responsible for the T160 phosphorylation of RMDN3. In addition to PERK, mitochondrial ROS have been shown to activate several protein kinases, including JNK, p38 MAPK, PKCδ, Akt, and AMPK55,56,57. JNK, p38 MAPK, and PKCδ are thought to play roles in promoting the induction of cell death, so they are unlikely to be involved in the phosphorylation of RMDN3. On the other hand, Akt and AMPK activate cell survival signals, so the possible involvement of these kinases in RMDN3 phosphorylation is certainly worth considering, while other kinases may also feature. In any case, it is likely that the MAM-localized protein kinase is responsible for the phosphorylation of RMDN3. We are currently using a knockdown screening approach to identify the kinase responsible for RMDN3 phosphorylation.
The TPR domain, a lipid transfer domain in RMDN3, has been reported to function in the bidirectional transport of phosphatidic acid between mitochondria and the ER44. The fact that deletion of the TPR domain of RMDN3 failed to remove lipid radicals accumulated in mitochondria suggests that RMDN3 uses its TPR domain to transfer phosphatidic acid-related lipid peroxides in mitochondria to the ER. Indeed, liposome assays suggested that RMDN3 may transport lipid radicals with a higher affinity than phosphatidic acid (Fig. 6). Thus, we speculate that RMDN3 selectively recognizes lipid radicals and oxidized lipids and transfers them to the ER. However, the mechanism by which RMDN3 achieves this selective recognition and how the lipid transfer activity is regulated are currently unknown and require further detailed analysis.
Mitochondria are constantly exposed to ROS due to their respiratory activity, but MnSOD, Prx-Trx, and GSH-GPx rapidly scavenge ROS. The recent development of new measurement techniques has made it possible to measure cysteine polysulfides, which consist of multiple sulfur linkages of the thiol group of cysteine, revealing that many proteins in cells contain cysteine polysulfides. Cysteine polysulfides and glutathione polysulfides are highly reactive to ROS58. Indeed, it has been reported that deficiency of CARS2, a cysteine polysulfide synthase present in mitochondria, causes mitochondrial dysfunction29. In the present study, we found that MERC formation was increased when the expression of SLC25A39, a mitochondrial-specific GSH transporter, and CARS2 was suppressed, and that the addition of Mito-TEMPO, mitochondrial-specific ROS scavengers, reversed this increase in MERC formation. Furthermore, MERC induction by CARS2 knockdown was rescued by the addition of NACS2, a synthetic source of cysteine polysulfides, but not by SLC25A39 knockdown. We speculate that NACS2 may not be an active form due to mitochondrial GSH depletion, or that the uptake of glutathione polysulfides such as GSSH into mitochondria was reduced. In addition, the mitochondrial ROS that induced MERCs were not removed by Prx3 or Trx2, suggesting that the ROS that induce MERC formation are removed by cysteine polysulfides or glutathione polysulfides. At present, it is not clear whether cysteine polysulfides or glutathione polysulfides suppress the mitochondrial ROS that induce MERC formation. In conclusion, our results strongly support the hypothesis that cysteine polysulfides and/or glutathione polysulfides primarily react with and scavenge mitochondrial ROS.
Our study shows that some lipid radicals and lipid peroxide produced by mitochondrial ROS were transferred to the ER. It has been reported that a large amount of lipid peroxide accumulates in the ER29. It is thus believed that there is an efficient lipid peroxide scavenging system in the ER. Since the ER occupies nearly 70% of the cell volume59, it is possible that lipid peroxidation does not accumulate in a small area such as the mitochondria, but is distributed throughout the ER to attenuate lipid radical toxicity. This could be a strategy to increase the frequency of catalysis of GPx4, a lipid peroxide scavenging enzyme located in the cytoplasm. Further analysis is needed to elucidate the molecular pathway by which oxidized lipids are scavenged in the ER.
In this study, we used brown adipocytes to show that RMDN3-mediated removal of mitochondrial lipid peroxide toxicity is observed under physiological conditions, such as after stimulation with norepinephrine. Given that RMDN3 is expressed in almost all cells and tissues, this suggests that an RMDN3-dependent lipid radical transport mechanism may be preserved in various cells and tissues.
Methods
Cell culture and transfections
HeLa cells (RIKEN, #RCB0007) were cultured in Dulbecco’s modified Eagles’s medium (DMEM) with high glucose supplemented with 10% fetal bovine serum (FBS). Cells were transfected with plasmids using FuGENE HD Transfection Reagent (Promega) or siRNAs using Lipofectamine RNAiMAX Transfection Reagent (Invitrogen) according to the manufacturer’s instructions. The Brown preadipocytes were cultured according to previously described methods60 with modifications. The stromal vascular fraction (SVF) was obtained from interscapular brown adipose tissue of newborn ICR mice; SVF was minced and incubated in ISO buffer (0.125 M NaCl, 5 mM KCl, 1.3 mM CaCl2, 5 mM glucose, 100 mM HEPES-KOH, pH 7.4, 4% bovine serum albumin) with 1 mg/ml collagenase (Wako, 034-22363) at 37 °C for 30 min. The digested SVF was gently mashed on a 100 μm nylon cell strainer using a syringe plunger with ISO buffer and isolated by centrifugation at 1000 g for 5 min. Cell pellets were washed with ISO buffer and incubated in DMEM (Nacalai Tesque, 08458-16) supplemented with 20% FBS and penicillin/streptomycin at 37 °C in a 5% CO2 incubator. After 3 days culture in DMEM supplemented with 20% FBS and penicillin/streptomycin (day 0), the culture medium was changed to differentiation induction (DI) medium (DMEM supplemented with 20% FBS, 20 nM insulin [Sigma-Aldrich, I1882], 1 nM triiodo-L-thyronine [T3; Nacalai Tesque, Inc. 35006-44], 5 μM dexamethasone [Nacalai Tesque, 11107-64], 0. 125 mM indomethacin [Tokyo Kasei Kogyo, I0655], 0.5 mM IBMX [Nacalai Tesque, 19624-86] and 1 μM rosiglitazone [Tokyo Chemical Industries, R0106)]) for 2 days (day 2). Cells were then cultured in differentiation-enhancing (DE) medium (DMEM supplemented with 20% FBS, 20 nM insulin, and 1 μM T3) for an additional 2 or 4 days (day 4 or day 6). For retroviral experiments, cells were infected with retroviral vectors 3 days before differentiation induction (day -3). siRNA experiments were performed 2 days before differentiation induction (day -2). 30 nM siRNA was introduced into cells using Lipofectamine RNAiMax (Invitrogen). All animals were maintained under university guidelines for the care and use of laboratory animals. The experiments were performed after securing protocol approval from the Gakushuin University and Tokyo University of Pharmacy and Life Sciences Animal Use Committee.
Small Interfering RNA
Knockdown of endogenous human MFN2, PDZD8, RMDN3, VAPB, VAPA, DRP1, Prx3, Trx2 and SLC25A39 or mouse RMDN3 was performed using the siRNAs that were transfected with Lipofectamine RNAiMAX Transfection Reagent. The target sequences were: hMFN261: 5′-AATCCCAGAGGGCAGAACTTTGTCC-3′; hPDZD8: 5′-GTTTAAGCCGTTTTTTCCATT-3′; hVAPB62: 5′-GCTCTTGGCTCTGGTGGTTTT-3′; hPrx3: 5′- AAGCCAAGUCCAGCUGCUUCC -3′; hTrx2: 5′- GGAUCUCCUUGACAACCUU-3′; hSLC25A39 #2: 5′- ACUUAAGGAAUUAACAAGGG -3′′; mRMDN3 #1: 5′- GCUUUAGCUUCAAGGAACA-3′; mRMDN3 #4: 5′- GAACUGCCAGACGUCACUAAU-3′ purchased from Japan Bio Services. The target sequences were: hCARS2 #1: 5′-GCAAAUCAACAGUACGUUUtt-3′; hCARS2 #2: 5′-GCUCAUAUGUUAGAUUUGAtt-3′ purchased from Thermo Fisher Scientific. CARS2: #1 (s35867), #2 (s35868). The target sequences were: hRMDN3 #1: 5′-GAGCGGGACTCTGACAAAGAA-3′; hVAPA: 5′-AUGCGUAUUUGAAAUGCCCAA-3′; hDRP1 #1: 5′-CAGGAGCCAGCTAGATATTAA-3′; hDRP1#2: 5′-CACGGTGGGCGCCGACATCAT-3′; hSLC25A39 #1: 5′-AGCCCTGTACTGGTTCAACTA-3′′were purchased from QIAGEN. Scrambled siRNA (si-NT) was also purchased from QIAGEN (#1027281).
Plasmids and reagents
To generate stably expressing cells, TOMM20-V5-SmBit and LgBiT-3HA-Sec61β were subcloned into the pCX4 retroviral vector63. The expression plasmids for human RMDN3 and VAPB were constructed as previously described64. RMDN3 (ΔTPR) is truncated at the C-terminal transmembrane domain (236-470 aa). RMDN3 (ΔFFAT) is truncated at the FFAT-like domain (157-174 aa). All RMDN3 siRNA-resistant forms were generated by introducing silent mutations in the hRMDN3 siRNA target sequence (5′-GAGCGGGACTCTGACAAAGAA-3′). FLAG-resi-RMDN3 WT, FLAG-resi-RMDN3 ΔFFAT and FLAG-resi-RMDN3 ΔTPR are RMDN3 RNAi-resistant vectors. mCherry-Sec61β (Addgene #49155) were purchased from Addgene. Carbonyl cyanide m-chlorophenylhydrazone (CCCP) was purchased from Wako. Antimycin A, ferrostatin-1, and valinomycin were purchased from Cayman Chemical Company. Rotenone, Mito-TEMPO (hydrate), and tert-butyl hydroperoxide (tBHP) were purchased from TCI Chemicals (Japan, Tokyo). Z-VAD-FMK was purchased from the Peptide Institute. Oligomycin and Necrostatin-1 were purchased from AdipoGen. D-Met was purchased from TCI Chemicals. oxNAC (10 mM) and NACS2 (10 mM) were prepared according to the previously reported method38.
Antibodies
For immunoblotting: Mouse VAPB antibody (66191-1-Ig, 1:1000), rabbit PDZD8 antibody (25512-1-AP, 1:1000), rabbit polyclonal anti-PTPIP51 (RMDN3) antibody (20641-1-AP, 1:1000) and rabbit monoclonal anti-TOMM20 antibody (11802-1-AP, 1:2000) were purchased from Proteintech. Mouse monoclonal anti-MFN2 antibody (sc-100560, 1:5000) and mouse polyclonal HSP60 antibody (sc-136291, 1:2000) were purchased from Santa Cruz Biotechnology. Rabbit anti-RMDN3 antibody (HPA009975, 1:1000) was purchased from Atlas. Mouse monoclonal anti-calnexin antibody (699401, 1:1000) was purchased from Biolegend. Mouse polyclonal HSP60 (sc-136291, 1:1000) was purchased from Santa Cruz Biotechnology. Anti-HA (3F10) peroxidase antibody (2-013-819, 1:5000) was purchased from Roche. Anti-FLAG M2 peroxidase (A8592, 1:1000) was purchased from Sigma-Aldrich. For immunofluorescence: Rabbit monoclonal anti-TOMM20 antibody (11802-1-AP, 1:2000) was purchased from Proteintech. Mouse monoclonal anti-calnexin antibody (699401) was purchased from Biolegend. Mouse polyclonal HSP60 (sc-136291, 1:2000) was purchased from Santa Cruz Biotechnology. Alexa Fluor-conjugated anti-mouse IgG, rabbit IgG, and rat IgG used as secondary antibodies for immunofluorescence were purchased from Invitrogen (1:1000).
Gene expression analysis by real-time PCR
In HeLa cell analysis, qPCR was performed using the THUNDERBIRD Next SYBR qPCR Mix (TOYOBO). All reactions were performed on a LightCycler96 system (Roche, Basel, Switzerland). Each sample was analyzed in duplicate for each PCR measurement. Experiments were performed in triplicate. Melting curves were checked for specificity. All mRNA expression level results were normalized to the expression level of ACTB. The following primer pairs were used in HeLa cells: Actb: forward (Fwd): 5′-GAGAAGATGACCAGATCAT -3′, reverse (Rev): 5′- ATCTCCATGCCCAGGAAGGAAGG -3′; Slc25a39: Fwd: 5′- TGCCCTTCTCAGCCCTGTA -3′, Rev: 5′- GGTTCACTCTCACAGCCTCC -3′; Cars2: Fwd: 5′- GACAGAAGCAGTACAACATCCA -3′, Rev: 5′- CTTGTCGTGAGCAGGTTCAT -3′. For brown adipocyte analysis, total RNA was extracted from primary mouse brown adipocytes using NucleoSpin® RNA (TAKARA). Reverse transcription reaction was performed using RevaTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO). Real-time quantitative PCR was performed using a KAPA SYBR Fast qPCR Kit (Nippon Genetics, KK4602) and a StepOnePlus Real-Time PCR System (Applied Biosystems). The following primer pairs were used for brown adipose cell analysis: s18: forward (Fwd): 5′-TCCAGCACATTTTGCGAGTA -3′, reverse (Rev): 5′- CAGTGATGGCGAAGGCTATT -3′; Rmdn3: Fwd: 5′- ACCCTACCTACGAGGACAAA -3′, Rev: 5′- AGCCCTCACCCAGGAATA -3′; Ucp1: Fwd: 5′- GTGAACCCGACAACTTCCGAA -3′, Rev: 5′- TGAAACTCCGGCTGAGAAGAT -3′; Cidea: Fwd: 5′- TGACATTCATGGGATTGCAGAC -3′, Rev: 5′- GGCCAGTTGTGATGACTAAGAC -3′; Fabp4: Fwd: 5′- GATGCCTTTGTGGGAACCT -3′, Rev: 5′- CTGTCGTCTGCGGTGATTT -3′; Elovl3: Fwd: 5′- TCCGCGTTCTCATGTAGGTCT -3′, Rev: 5′- GGACCTGATGCAACCCTATGA -3′; Pgc1a: Fwd: 5′- CCCTGCCATTGTTAAGACC -3′, Rev: 5′- TGCTGCTGTTCCTGTTTTC -3′.
Immunoblotting
Proteins in the sample buffer were separated by SDS-PAGE and transferred to Immobilon PVDF membranes (IPVH00010, Millipore). Blots were probed with the indicated antibodies, and protein bands on the blot were visualized using Chemi-Lumi One L (07880-70, Nacalai Tesque) or Immobilon Western Chemiluminescent HRP Substrate (WBKLS0500, Millipore). Band images were captured using a LuminoGraph I imager (ATTO). Relative band intensities were quantified using Fiji/ImageJ software65.
Immunoblotting and Phos-tag PAGE
FLAG-tagged RMDN3 was immunoprecipitated with FLAG M2 antibody, and the lysates were separated by SDS-PAGE and transferred to PVDF membranes (Millipore). The blots were probed with the indicated antibodies, and protein bands on the blot were visualized using enhanced chemiluminescence reagent (Millipore). Phos-tag PAGE was performed using 12.5% SDS-PAGE minigels containing 10 μM Phos-tag (Wako) in the presence of 100 μM MnCl2 according to the manufacturer’s protocol.
Immunostaining
Cells were fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 15 min at 37 °C. Fixed cells were washed three times with PBS, permeabilized with 0.1% Triton X-100 in PBS for 10 min then washed three times with PBS and blocked with 10% FBS in PBS for 10 min, all at room temperature (RT). For immunostaining, cells were incubated with the indicated primary antibodies in PBS containing 5% FBS for 2 h at RT, washed three times with PBS, and then incubated with the appropriate Alexa Fluor-conjugated secondary antibodies in PBS containing 5% FBS for 1 h. The sample was washed three times with PBS, and mounted with Fluorescent Mounting Medium (S3023, Dako), and then analyzed using a FV3000 confocal laser scanning microscope (Olympus). Slice images were acquired every 0.2 μm and reconstructed using Fiji/ImageJ software. For the analysis of LDs, a 400 μm2 of region of interest (ROI) was selected from the single-slice images. The total LDs area and the mean LD area in the ROI were obtained using the “Analyze particles” plugin in Fiji with a minimum area of 0.5 μm2. Similarly, the total mitochondrial area in the ROI was obtained using the “Analyze particles” plugin in Fiji with a minimum area of 0.1 μm2. At least 3 ROIs per condition from two independent experiments were analyzed.
Immunoprecipitation
Cells were lysed with NP-40 lysis buffer (1% Nonidet P40 Substitute, 10 mM Tris–HCl pH 7.4, 150 mM NaCl, 0.5 mM EDTA, 10 mM NaF) containing protease inhibitor cocktail (25955-11, Nacalai Tesque) and centrifuged at 15,000 g for 10 min at 4 °C. The supernatant was subjected to immunoprecipitation using the indicated antibodies and Protein G Sepharose (17–0618-01, GE Healthcare). The beads were washed four times with NP-40 lysis buffer and boiled in 2× Laemmli sample buffer (125 mM Tris–HCl pH 6.8, 10% 2-mercaptoethanol, 4% SDS, 10% sucrose, 0.01% BPB).
Luciferase Assay using MERBiT cell
Luciferase assays using the MERBiT system were performed using the Nano-Glo Live Cell Assay; Nano-Glo Live Cell Substrate (N205B) and NanoBiT Buffer (Nano-Glo LCS Dilution Buffer) (N206B) (Promega, Madison, WI, USA) according to the manufacturer’s instructions. For luminescence quantification, MERBiT cells were seeded at 0.5 or 1.0 × 104 /well on a 96-well white cell culture microplate (Greiner bio-one), followed by transfection of expression vectors and treatment with rotenone or antimycin A. The cell culture medium was replaced with 100 μL of buffered FluoroBrite DMEM (Gibco) or Opti-MEM, containing <1% (v/v) fetal bovine serum (FBS). 25 µL of a 1:20 mixture of Nano-Glo Live Cell Substrate (N205B) and NanoBiT Buffer was added and incubated at 37 °C for 10 min. Luminescence was then measured using a Synergy LX multimode reader (BioTek).
Cell viability assay
Cell viability assay was performed using the Cell Counting Kit-8 (Dojindo) according to the manufacturer’s protocol.
Live cell imaging
For MitoPeDPP (Dojindo, M466) staining: Cells transfected with the indicated siRNAs prior to transfection with the RMDN3 siRNA-resistant construct were plated on 9.5 mm glass-bottom dishes (MATSUNAMI, D141400). If necessary, they were transfected with the indicated vectors and then treated with rotenone (50 nM) or antimycin A (50 nM). For staining, the medium was changed to FluoroBrite-DMEM (Gibco) without FBS containing MitoPeDPP (final 1 µM) and incubated at 37 °C for 30 min with protection from light. The supernatant was discarded and the cells were washed twice with FluoroBrite-DMEM. Add fresh FluoroBrite-DMEM and observe the cells under a fluorescence microscope.
For LipiRADICAL (Funakoshi, FDV-0042) staining: Cells transfected with indicated siRNAs prior to transfection with RMDN3 siRNA-resistant construct were plated on 9.5 mm glass bottom dishes (MATSUNAMI, D141400). If necessary, they were transfected with the indicated vectors and then treated with rotenone (50 nM) or antimycin A (50 nM) for 1 day. For staining, the medium was changed to FluoroBrite-DMEM with 2% FBS containing LipiRADICAL (final 2 µM) and incubated at 37 °C for 1 h. If necessary, cells were stained with the mitochondria-specific dye MitoTracke Deep Red FM (100 nM) (M22426, Invitrogen) for 30 min at 37 °C prior to imaging. Discard the supernatant and wash the cells twice with FluoroBrite-DMEM before viewing. Add fresh FluoroBrite-DMEM with 2% FBS and observe the cells under a fluorescence microscope.
Mitochondrial calcium analysis
Cells stably expressing mitochondria-targeted CEPIA2 (CEPA2mt) were seeded on a 96-Well Black/Clear Bottom Plate (Thermo Fisher Scientific). 48 h after transfections of siRNAs, cells were washed three times in a BSS buffer (120 mM NaCl, 5.4 mM KCl, 0.8 mM MgCl2, 6 mM NaHCO3, 5.6 mM D-glucose, 2 mM CaCl2, and 25 mM HEPES [pH 7.3]) before analysis. Fluorescence values were then collected at indicated times, and cells were stimulated with or without 100 μM histamine in BSS. Fluorescence was recorded by a Synergy LX multimode reader (BioTek) with 485/ 528 nm Filter (green, gain 75).
EM Imaging
HeLa cells and MERBiT cells were fixed in 2.5% glutaraldehyde in DMEM supplemented with 10% FBS at 37 °C in a 5% CO2 incubator for 1 h. After being washed with 0.1 M phosphate buffer (0.08 M disodium hydrogen phosphate, 0.02 M sodium dihydrogen phosphate dihydrate), the cells were scraped and collected with 0.2% BSA/0.1 M phosphate buffer followed by centrifugation at 820 g. The cells were post-fixed with 1% OsO4 (Electron Microscopy Sciences) and 1.5% potassium ferrocyanide (Kanto Chemical, catalog no. 32338-30) in a 0.05 M phosphate buffer for 30 min. After being rinsed 3 times with H2O, the cells were stained with 1% thiocarbohydrazide (Sigma-Aldrich) for 5 min. After being rinsed with H2O 3 times, cells were stained with 1% OsO4 in H2O for 30 min. After being rinsed with H2O 2 times at room temperature and 3 times at 50 °C, cells were treated with Walton’s lead aspartate (0.635% lead nitrate (Sigma-Aldrich, catalog no. 203580), 0.4% aspartic acid (Sigma-Aldrich, catalog no. A9256), pH 5.2) at 50 °C for 20 min. After being rinsed with H2O 3 times, the sections were followed by incubations in an ascending ethanol series (10 min each in 50% on ice and 70% on ice, 10 min each in 90%, and 5 min in 95% ethanol/H2O at room temperature), 5 min in 100% ethanol 4 times and 10 min in butyl 2,3-epoxypropyl ether (Fujifilm Wako pure chemical corporation) twice. This was followed by infiltration of Epon812 resin-butyl 2,3-epoxypropyl ether for 14 h at a 1:1 dilution. After incubating with 100% Epon812 for 2 h, the resin was replaced with fresh 100% Epon812 and incubated for an additional 3 h. Finally, the resin was cured at 65 °C for 5 days. Epon812 (Nisshin EM) was made by mixing 7.5 g of MNA (Nisshin EM, M012), 13.7 g of Epon812 (Nisshin EM T026), and 3.8 g of DDSA (Nisshin EM, D027), and 0.4 g of DMP-30 (Nisshin EM, D032). Resin blocks were trimmed with a TrimTool diamond knife (Trim 45; DiATOME). 60 nm thick ultra-thin sections made with a diamond knife (Ultra 45; DiATOME) were collected on a cleaned silicon wafer strip in a Leica Ultramicrotome (UC7). These ultra-thin sections were collected on cleaned silicon wafer strips and imaged with a scanning electron microscope (JSM-IT800SHL, JEOL). Imaging was done at 1 kV accelerating voltage, 5 kV specimen voltage, 250 pA beam current, 1,024 × 960 frame size, 6 mm working distance, 6.4 × 4.8 µm field of view (pixel size 5 nm), and 5.33 µs dwell time, using the Scintillator Backscattered Electron Detector in Beam Deceleration mode.
Flow cytometry analysis
For analysis using MitoPeDPP in Flow cytometry, cells were stained with 10 µM MitoPeDPP for 30 min, cells were detached with trypsin, trypsin solution was removed, cells were resuspended in 1% FBS/PBS solution, and 10000 cells were analyzed. The gating of the MitoPeDPP signal in Flow Cytometry was set so that the signal of unstimulated cells was less than 5%.
Fluorescence-activated cell sorting (FACS)
MERBiT cells were stained with 10 µM MitoSOX (Invitrogen, M36008) for 30 min and sorted for mitochondrial ROS high and low using Cell Sorter (SONY, SH800). NHDF and F964 cells were stained with 100 nM SiDMA for 45 min and sorted for mitochondrial ROS high and low using Cell Sorter (BD Biosciense, FACS Discover S8). All sorted cells were cultured and then used for experiments.
The purification of the RMDN3 TPR domain
The plasmid and the protocol used for the protein purification process are based on the previous study44. Briefly, the RMDN3 TPR domain (PTPIP51_TPR) was expressed in E. coli Rosetta2 (DE3) (Novagen, USA) cells. When the cell density reached an OD600 of 0.6 at 37 °C, 1 mM isopropyl-D-thiogalactoside was added to induce protein expression. The cells were further incubated at 16 °C for 16 h. The cultured cells were harvested and resuspended in a lysis buffer, of which the composition is as follows: 20 mM Tris–HCl pH 7.5, 150 mM NaCl, 5% glycerol, 5 mM β‐mercaptoethanol, and 1 mM phenylmethylsulfonyl fluoride. The resuspended cells were lysed under the pressure of 10,000 psi using a high-pressure homogenizer (Micronox, South Korea). Its supernatant was separated from the content after centrifugation at 39,000 g for 2 h. The supernatant was mixed with Ni-NTA resin (Cytiva, USA), and the target protein was eluted from the resin using an elution buffer (20 mM Tris–HCl pH 7.5, 150 mM NaCl, 5% glycerol, 5 mM β‐mercaptoethanol, and 300 mM imidazole). The fused His-tag was cleaved by tobacco etch virus (TEV) protease at 4 °C for 16 h. The cleaved PTPIP51_TPR was collected as the flow-through of the second Ni-Affinity Chromatography. The cleaved protein was further purified using a Superdex 200 prep grade column (Cytiva, USA) using SEC buffer (20 mM Tris–HCl pH 7.5, 150 mM NaCl, 5% glycerol, and 2 mM dithiothreitol). The purified RMDN3 TPR domain was concentrated to 4.65 mg/mL using an Amicon® ultra centrifugal filter (10 kDa molecular weight cut-off) for further experiments.
Liposome and lipid transfer assay
The following phospholipids were purchased from Avanti Polar Lipids (USA): 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC, cat. no. 850457 C), 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE, cat. no. 850757 C), 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine- N-(cap biotinyl) (sodium salt) (biotinyl cap-PE, cat. nos. 870273 P and 870277 P), 4a-diaza-s-indacene-2-propionyl-amino-hexanoyl-sn-glycero-3-phosphate (ammonium salt) (TopFluor” TMR-PA, cat. no. 810240 C). Lipid solutions in chloroform were mixed to the desired composition, and the solvent was dried using an N2 gas and vacuum desiccator. The liposome film was dissolved in assay buffer (20 mM Tris–HCl [pH 8.0], 150 mM NaCl, and 2 mM EDTA). Liposomes were extruded 20 times through a 0.1 µm polycarbonate filter using a mini-extruder set (Cat. No. 61000, Avanti Polar Lipids, USA). Bead pulldown-based fluorescent lipid transfer assays were performed according to a previous report44 with modifications. Briefly, after binding of the donor liposomes (2% biotinyl cap-PE; 1% TMR-PA; and 97% POPC) and streptavidin-conjugated beads (Dynabeads Streptavidin T1, Cat. No. 65601), 5 µM donor liposomes and 30 µM acceptor liposomes (100% POPC) were incubated with different concentrations of RMDN3 (TPR) (0, 0.5, 1 and 2 µM) for 30 min at 25 °C with gentle agitation. If necessary, donor liposomes were treated with 500 µM AAPH, incubated at 37 °C for 30 min, and then washed three times with ice-cold assay buffer. Supernatants containing acceptor liposomes and RMDN3 (TPR) were separated from the bead mixtures using magnetic racks. If necessary, 10 µM NBP-Pen (LipiRADICAL) was added to the supernatants and incubated at 37 °C for 30 min. Donor liposomes were solubilized from the beads after treatment with 0.4% (w/v) n-dodecyl-b-D-maltoside in the TMR-PA transfer assay. Fluorescence of acceptor and donor liposomes was also measured in a Synergy LX multimode reader (BioTek). Fluorescence was measured at excitation/emission wavelengths of 530/590 nm (red), 485/ 528 nm (green) for TopFluor-TMA-PA and LipiRADICAL Green, respectively. The lipid transfer of TMR-PA was determined using the formula 100 × F acceptor/ (F acceptor + F donor).
Construction of a hypothetical model of the MERBiT system
To visualize the relationship between LgBiT fused via a linker to the cytoplasmic end of Sec61β and SmBiT fused via a linker to the cytoplasmic end of TOMM20, respectably, when they are in close proximity between the ER and mitochondria and assembled into NanoBiT, the following hypothetical model was constructed. First, we fitted the AlphaFold model of the full-length Homo sapiens Sec61β (AlphaFold ID: P60468) to the C-terminal transmembrane helix of Sec61β in the cryo-electron microscopy (cryo-EM) structure of the human Sec61 complex (PDB ID: 8DNY66). Since the AlphaFold structural prediction does not take into account the constraint information of a protein depending on its characteristic environment such as the organelle membrane in vivo, we remodeled the predicted N-terminal flexible loop of Sec61β as a reference to the cryo-EM potential density map of the Sec61 complex (EMDB ID: EMD-2758466) with the addition of the designed linker between LgBiT and Sec61β to upstream of the N-terminus of Sec61β. Next, we fitted the AlphaFold model of the full-length H. sapiens TOMM20 (AlphaFold ID: Q15388) to the C-terminal domain of TOMM20 (PDB ID: 7VC967) visualized by single-particle cryo-EM analysis of the human TOM complex (PDB ID: 7VDD67 and 7VC9). We then remodeled it to a reasonable region based on the density map of the TOM complex (EMDB ID: EMD-31914 and EMD-3188967) and previous NMR structures (PDB ID: 1OM268). We then manually Supplementary the designed linker between TOMM20 and SmBiT from downstream of the C-terminus of TOMM20. Whether the respective linkers form a secondary structure with Sec61β and TOMM20 was confirmed by using the Phyre2 server69. We then connected the N-terminus of the linker (Sec61β) and the C-terminus of the linker (TOMM20) to the C-terminus of the LgBiT and the N-terminus of the SmBiT of the crystal structure of NanoBiT (PDB ID: 7SNX), respectively. A series of model constructions were performed manually by using the COOT70. The predicted models of the full-length Sec61β and TOMM20 were obtained from the AlphaFold Protein Structure Database71. Figures of the structures were drawn using PyMOL (The PyMOL Molecular Graphics System, user-supported open-source software, version 3.0.0 Schrödinger, LLC).
Statistical analysis
Data are presented as mean ± s.e.m. and expressed as percentages or multiples relative to the values in control cells. Statistical significance was analyzed using Prism 9 software (GraphPad Software). Statistical significance was analyzed by either one-way analysis of variance followed by Bonferroni’s test, as appropriate. P values are indicated as *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; n.s., not significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Phillips, M. J. & Voeltz, G. K. Structure and function of ER membrane contact sites with other organelles. Nat. Rev. Mol. Cell Biol. 17, 69–82 (2016).
Prinz, W. A., Toulmay, A. & Balla, T. The functional universe of membrane contact sites. Nat. Rev. Mol. Cell Biol. 21, 7–24 (2020).
Zung, N. & Schuldiner, M. New horizons in mitochondrial contact site research. Biol. Chem. 401, 793–809 (2020).
de Brito, O. M. & Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456, 605–610 (2008).
Naon, D. et al. Critical reappraisal confirms that Mitofusin 2 is an endoplasmic reticulum-mitochondria tether. Proc. Natl. Acad. Sci. USA. 113, 11249–11254 (2016).
Naon, D. et al. Splice variants of mitofusin 2 shape the endoplasmic reticulum and tether it to mitochondria. Science 380, eadh9351 (2023).
De Vos, K. J. et al. VAPB interacts with the mitochondrial protein PTPIP51 to regulate calcium homeostasis. Hum. Mol.Genet. 21, 1299–1311 (2012).
Hirabayashi, Y. et al. ER-mitochondria tethering by PDZD8 regulates Ca(2+) dynamics in mammalian neurons. Science 358, 623–630 (2017).
Nakamura, K. et al. Mitochondrial protein FKBP8 captures PDZD8 to form mitochondria-ER contacts. bioRxiv, https://www.biorxiv.org/content/10.1101/2023.1108.1122.554218v554212 (2023).
Szabadkai, G. et al. Chaperone-mediated coupling of endoplasmic reticulum and mitochondrial Ca2+ channels. J. Cell Biol. 175, 901–911 (2006).
Eisenberg-Bord, M., Shai, N., Schuldiner, M. & Bohnert, M. A Tether Is a Tether Is a Tether: Tethering at Membrane Contact Sites. Dev. Cell 39, 395–409 (2016).
Scorrano, L. et al. Coming together to define membrane contact sites. Nat. Commun. 10, 1287 (2019).
Kwak, C. et al. Contact-ID, a tool for profiling organelle contact sites, reveals regulatory proteins of mitochondrial-associated membrane formation. Proc. Natl. Acad. Sci. USA 117, 12109–12120 (2020).
Antonicka, H. et al. A High-Density Human Mitochondrial Proximity Interaction Network. Cell Metab. 32, 479–497 e479 (2020).
Dixon, A. S. et al. NanoLuc Complementation Reporter Optimized for Accurate Measurement of Protein Interactions in Cells. ACS Chem. Biol. 11, 400–408 (2016).
Zorov, D. B., Juhaszova, M. & Sollott, S. J. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol. Rev. 94, 909–950 (2014).
Duchen, M. R. Mitochondria in health and disease: perspectives on a new mitochondrial biology. Mol. Aspects Med. 25, 365–451 (2004).
Balaban, R. S., Nemoto, S. & Finkel, T. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005).
Imlay, J. A. & Linn, S. DNA damage and oxygen radical toxicity. Science 240, 1302–1309 (1988).
Ayala, A., Munoz, M. F. & Arguelles, S. Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell. Longev. 2014, 360438 (2014).
Sutton, H. C. & Winterbourn, C. C. On the participation of higher oxidation states of iron and copper in Fenton reactions. Free Radic. Biol. Med. 6, 53–60 (1989).
Imai, H., Matsuoka, M., Kumagai, T., Sakamoto, T. & Koumura, T. Lipid Peroxidation-Dependent Cell Death Regulated by GPx4 and Ferroptosis. Curr. Top. Microbiol. Immunol. 403, 143–170 (2017).
Maiorino, M., Conrad, M. & Ursini, F. GPx4, Lipid Peroxidation, and Cell Death: Discoveries, Rediscoveries, and Open Issues. Antioxid. Redox Signal. 29, 61–74 (2018).
Brand, M. D. The sites and topology of mitochondrial superoxide production. Exp. Gerontol. 45, 466–472 (2010).
Zhao, R. Z., Jiang, S., Zhang, L. & Yu, Z. B. Mitochondrial electron transport chain, ROS generation and uncoupling (Review). Int. J. Mol. Med. 44, 3–15 (2019).
Mari, M. et al. Mitochondrial Glutathione: Recent Insights and Role in Disease. Antioxidants (Basel) 9 (2020).
Wang, Y. et al. SLC25A39 is necessary for mitochondrial glutathione import in mammalian cells. Nature 599, 136–140 (2021).
Barayeu, U. et al. Supersulfide biology and translational medicine for disease control. Br. J. Pharmacol. https://doi.org/10.1111/bph.16271 (2023).
Akaike, T. et al. Cysteinyl-tRNA synthetase governs cysteine polysulfidation and mitochondrial bioenergetics. Nat. Commun. 8, 1177 (2017).
Booth, D. M., Varnai, P., Joseph, S. K. & Hajnoczky, G. Oxidative bursts of single mitochondria mediate retrograde signaling toward the ER. Mol. Cell 81, 3866–3876 e3862 (2021).
Yoboue, E. D., Sitia, R. & Simmen, T. Redox crosstalk at endoplasmic reticulum (ER) membrane contact sites (MCS) uses toxic waste to deliver messages. Cell Death Dis. 9, 331 (2018).
Cieri, D. et al. SPLICS: a split green fluorescent protein-based contact site sensor for narrow and wide heterotypic organelle juxtaposition. Cell Death Differ. 25, 1131–1145 (2018).
Rambold, A. S., Kostelecky, B., Elia, N. & Lippincott-Schwartz, J. Tubular network formation protects mitochondria from autophagosomal degradation during nutrient starvation. Proc. Natl. Acad. Sci. USA. 108, 10190–10195 (2011).
Yang, Z., Zhao, X., Xu, J., Shang, W. & Tong, C. A novel fluorescent reporter detects plastic remodeling of mitochondria-ER contact sites. J. Cell Sci. 131, jcs208686 (2018).
Li, N. et al. Mitochondrial complex I inhibitor rotenone induces apoptosis through enhancing mitochondrial reactive oxygen species production. J. Biol. Chem. 278, 8516–8525 (2003).
Chen, Q., Vazquez, E. J., Moghaddas, S., Hoppel, C. L. & Lesnefsky, E. J. Production of reactive oxygen species by mitochondria: central role of complex III. J. Biol. Chem. 278, 36027–36031 (2003).
Kagan, V. E. et al. Mitochondrial targeting of electron scavenging antioxidants: Regulation of selective oxidation vs random chain reactions. Adv. Drug Deliv. Rev. 61, 1375–1385 (2009).
Zhang, T. et al. Enhanced Cellular Polysulfides Negatively Regulate TLR4 Signaling and Mitigate Lethal Endotoxin Shock. Cell Chem. Biol. 26, 686–698 e684 (2019).
Halvey, P. J. et al. Selective oxidative stress in cell nuclei by nuclear-targeted D-amino acid oxidase. Antioxid. Redox Signal. 9, 807–816 (2007).
Hoehne, M. N. et al. Spatial and temporal control of mitochondrial H(2) O(2) release in intact human cells. EMBO J. 41, e109169 (2022).
Di Mattia, T. et al. FFAT motif phosphorylation controls formation and lipid transfer function of inter-organelle contacts. EMBO J. 39, e104369 (2020).
Cabukusta, B. et al. Human VAPome Analysis Reveals MOSPD1 and MOSPD3 as Membrane Contact Site Proteins Interacting with FFAT-Related FFNT Motifs. Cell Rep. 33, 108475 (2020).
Subra, M. et al. VAP-A intrinsically disordered regions enable versatile tethering at membrane contact sites. Dev. Cell 58, 121–138.e129 (2023).
Yeo, H. K. et al. Phospholipid transfer function of PTPIP51 at mitochondria-associated ER membranes. EMBO Rep. 22, e51323 (2021).
Suzuki, J. et al. Imaging intraorganellar Ca2+ at subcellular resolution using CEPIA. Nat. Commun. 5, 4153 (2014).
De Vos, K. J. et al. VAPB interacts with the mitochondrial protein PTPIP51 to regulate calcium homeostasis. Hum. Mol. Genet. 21, 1299–1311 (2012).
Gaschler, M. M. & Stockwell, B. R. Lipid peroxidation in cell death. Biochem. Biophys. Res. Commun. 482, 419–425 (2017).
Dixon, S. J. & Stockwell, B. R. The role of iron and reactive oxygen species in cell death. Nat. Chem. Biol. 10, 9–17 (2014).
Tadokoro, T. et al. Mitochondria-dependent ferroptosis plays a pivotal role in doxorubicin cardiotoxicity. JCI Insight 8, e169756 (2023).
Shioji, K., Oyama, Y., Okuma, K. & Nakagawa, H. Synthesis and properties of fluorescence probe for detection of peroxides in mitochondria. Bioorg. Med. Chem. Lett. 20, 3911–3915 (2010).
Yamada, K. et al. Fluorescence probes to detect lipid-derived radicals. Nat. Chem. Biol. 12, 608–613 (2016).
Chouchani, E. T. et al. Mitochondrial ROS regulate thermogenic energy expenditure and sulfenylation of UCP1. Nature 532, 112–116 (2016).
Huang, X., Jiang, C., Yu, L. & Yang, A. Current and Emerging Approaches for Studying Inter-Organelle Membrane Contact Sites. Front. Cell Dev. Biol. 8, 195 (2020).
Verfaillie, T. et al. PERK is required at the ER-mitochondrial contact sites to convey apoptosis after ROS-based ER stress. Cell Death Differ. 19, 1880–1891 (2012).
Zhao, Y. et al. ROS signaling under metabolic stress: cross-talk between AMPK and AKT pathway. Mol. Cancer 16, 79 (2017).
Qi, X. & Mochly-Rosen, D. The PKCdelta -Abl complex communicates ER stress to the mitochondria - an essential step in subsequent apoptosis. J. Cell Sci. 121, 804–813 (2008).
Kulisz, A., Chen, N., Chandel, N. S., Shao, Z. & Schumacker, P. T. Mitochondrial ROS initiate phosphorylation of p38 MAP kinase during hypoxia in cardiomyocytes. Am. J. Physiol. Lung Cell. Mol. Physiol. 282, L1324–L1329 (2002).
Ida, T. et al. Reactive cysteine persulfides and S-polythiolation regulate oxidative stress and redox signaling. Proc. Natl. Acad. Sci. USA. 111, 7606–7611 (2014).
Heinrich, L. et al. Whole-cell organelle segmentation in volume electron microscopy. Nature 599, 141–146 (2021).
Kato, H. et al. ER-resident sensor PERK is essential for mitochondrial thermogenesis in brown adipose tissue. Life Sci. Alliance 3, e201900576 (2020).
Ban-Ishihara, R., Ishihara, T., Sasaki, N., Mihara, K. & Ishihara, N. Dynamics of nucleoid structure regulated by mitochondrial fission contributes to cristae reformation and release of cytochrome c. Proc. Natl. Acad. Sci. USA. 110, 11863–11868 (2013).
Stoica, R. et al. ER-mitochondria associations are regulated by the VAPB-PTPIP51 interaction and are disrupted by ALS/FTD-associated TDP-43. Nat. Commun. 5, 3996 (2014).
Akagi, T., Sasai, K. & Hanafusa, H. Refractory nature of normal human diploid fibroblasts with respect to oncogene-mediated transformation. Proc. Natl. Acad. Sci. USA. 100, 13567–13572 (2003).
Ito, N. et al. MITOL regulates phosphatidic acid-binding activity of RMDN3/PTPIP51. J. Biochem. 171, 529–541 (2022).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Itskanov, S. et al. A common mechanism of Sec61 translocon inhibition by small molecules. Nat. Chem. Biol. 19, 1063–1071 (2023).
Su, J. et al. Structural basis of Tom20 and Tom22 cytosolic domains as the human TOM complex receptors. Proc. Natl. Acad. Sci. USA. 119, e2200158119 (2022).
Abe, Y. et al. Structural basis of presequence recognition by the mitochondrial protein import receptor Tom20. Cell 100, 551–560 (2000).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Casañal, A., Lohkamp, B. & Emsley, P. Current developments in Coot for macromolecular model building of Electron Cryo-microscopy and Crystallographic Data. Protein Sci. 29, 1069–1078 (2020).
Varadi, M. et al. AlphaFold Protein Structure Database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–d375 (2024).
Acknowledgements
We thank Momoka Sakamaki for technical assistance with qRT-PCR and members of the Laboratory of Cell Biology, Biomedical Research Core Facilities, Juntendo University Graduate School of Medicine for technical assistance. This study was partially supported by the MEXT/JSPS KAKENHI [Grant Nos. 22K15399, 22H05574 and 24H01327 (for I.S.), 23K14185 and 22K20637 (for N.I.), 21K06844 (for R.I.), 21H0207, 21H05267, and 23K17979 (for T.S.), and J22H05532 (Y.H.), and 23H02691, 20H04911, and 20H03454 (for SY)], the Uehara Memorial Foundation (for I.S. and S.Y.), the Kowa Life Science Foundation (to I.S.)], and AMED under grant numbers 23gm1610011h0001 (to T.S.), and JP19dm0207082 (to Y.H.), JP17gm5010002, JP18gm5010002, JP19gm5010002, and JP20gm5010002 (to S.Y.).
Author information
Authors and Affiliations
Contributions
I.S. and S.Y. designed the experiments; I.S., N.I., H.O., R.K., Y.I., F.Y., T.N., H.S., K.O., and E.T. performed the experiments; I.S. and S.Y. analyzed and interpreted the data; T.T., S.N., A.S., Y.O., Y.T., B.L., Y.H., Y.F., K.U., T.S., H.K., H.N., and R.I. assisted in the analysis; I.S. and S.Y. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shiiba, I., Ito, N., Oshio, H. et al. ER-mitochondria contacts mediate lipid radical transfer via RMDN3/PTPIP51 phosphorylation to reduce mitochondrial oxidative stress. Nat Commun 16, 1508 (2025). https://doi.org/10.1038/s41467-025-56666-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56666-4
This article is cited by
-
Convergent activation of the integrated stress response and ER–mitochondria uncoupling in VAPB-associated ALS
EMBO Molecular Medicine (2025)
-
GLP-1R associates with VAPB and SPHKAP at ERMCSs to regulate β-cell mitochondrial remodelling and function
Nature Communications (2025)