Abstract
The Oncotype DX® Recurrence Score (RS) is an assay for hormone receptor-positive early breast cancer with extensively validated predictive and prognostic value. However, its cost and lag time have limited global adoption, and previous attempts to estimate it using clinicopathologic variables have had limited success. To address this, we assembled 6172 cases across three institutions and developed Orpheus, a multimodal deep learning tool to infer the RS from H&E whole-slide images. Our model identifies TAILORx high-risk cases (RS > 25) with an area under the curve (AUC) of 0.89, compared to a leading clinicopathologic nomogram with 0.73. Furthermore, in patients with RS ≤ 25, Orpheus ascertains risk of metastatic recurrence more accurately than the RS itself (0.75 vs 0.49 mean time-dependent AUC). These findings have the potential to guide adjuvant therapy for high-risk cases and tailor surveillance for patients at elevated metastatic recurrence risk.
Similar content being viewed by others
Introduction
Hormone receptor-positive disease without HER2 overexpression or amplification (HR + /HER2-) is the most common subtype of early breast cancer (EBC), accounting for approximately 70% of diagnoses1. A major challenge in the management of this disease has been identifying the cancers for which adjuvant chemotherapy meaningfully reduces the risk of recurrence. Risk stratification of HR + /HER2- EBC relies upon the integration of traditional clinicopathological features (e.g., tumor size, nodal status, Nottingham grade) with multigene assays to estimate the risk of recurrence and personalize adjuvant therapy. Among the commercially available assays, Oncotype DX (ODX) ® (Exact Sciences, Madison, WI) is the most extensively validated and widely used in clinical practice. By measuring the transcriptional abundance of 16 genes, including ESR1, PGR, HER2, MKI67, and MMP11, against the abundance of five reference genes using reverse transcription quantitative real-time PCR2, ODX calculates a recurrence score (RS) ranging from zero to 100 with both prognostic and predictive value2,3,4,5,6,7,8,9.
Substantial clinical evidence from retrospective and prospective trials has shown that ODX can improve clinical decision-making in breast cancer. Retrospective analyses of the NSABP B142 and TransATAC5 trials demonstrated the prognostic value of ODX in stratifying the risk of recurrence for HR + /HER2- EBC patients. Similarly, analyses of the NSABP B206 and SWOG88147 clinical trials established the predictive value of ODX by uncovering a survival benefit with the addition of adjuvant chemotherapy to endocrine therapy for patients with a high risk of disease relapse. These studies provided the rationale for the prospective evaluation of ODX in the TAILORx8 (>10,000 patients with node-negative disease) and RxPONDER9 (5083 patients with one to three positive lymph nodes) trials and established ODX as the preferred genomic assay for adjuvant treatment-decision-making in HR + /HER2- EBC10,11.
While guidelines have recommended the use of ODX or other assays for more than a decade10,11,12, reimbursement restrictions and global accessibility barriers have limited universal adoption13. Beyond the United States, the cost of around 4000 USD per sample14,15 and turnaround time delaying the start of therapy have created barriers to adoption, despite analyses indicating downstream savings from more tailored adjuvant therapy16. Some efforts have been undertaken to develop nomograms based on clinical and pathologic features annotated during the standard of care, aiming to predict ODX scores17. However, such tools require manual extraction of relevant inputs from the unstructured electronic healthcare record and leave room for improvement in terms of performance, with the assay itself still providing greater cost-effectiveness than these tools16.
We investigated the use of whole-slide images (WSIs) from routinely available formalin-fixed paraffin-embedded (FFPE) tissue slides stained with hematoxylin and eosin (H&E) to predict RS. As previous studies have demonstrated, these slides can be effectively analyzed using deep learning algorithms to predict relapse risk18,19,20,21,22,23,24,25,26. Such algorithms have already been approved for colorectal cancer27,28 in Europe, though their widespread adoption is yet to be realized. One possible reason for this delay could be the limited clinical validation against the standard of care29. However, the field of deep learning is progressing rapidly. Recently, two techniques have markedly enhanced system performance: transformers and self-supervised learning30 (SSL). Furthermore, recent studies have shown that integrating histopathologic imaging with additional modalities, such as genomics, text, and clinical imaging, uncovers intermodal relationships and often improves predictive performance31,32,33,34.
In this study, we develop Orpheus, a multimodal deep learning model to infer the ODX RS from H&E-stained whole-slide images and validate it across three independent patient cohorts for the identification of high-risk patients. Moreover, a head-to-head comparison is made between the Orpheus and ODX RS to identify patients with documented metastatic recurrence. This work advances a multimodal machine learning paradigm in precision oncology, applies it to accurately infer the ODX RS from routine histopathology images, and outperforms the ODX RS in identifying the risk of metastatic recurrence in patients with low ODX RS. This study has the potential to extend access to the established and exploratory applications of the well-validated Recurrence Score and to refine sub-stratification for patients treated using the current standard paradigm.
Results
Data assembly
We assembled three independent cohorts comprising 6172 patients with HR + /HER2- EBC with surgically resected primary tumors (Fig. 1a). Tissue samples were subjected to H&E staining and immunohistochemical (IHC) analysis for hormone receptors and HER2 according to ASCO/CAP guidelines35,36, and samples were submitted for calculation of RS per clinical practice. For a subset, genomic data from clinical MSK-IMPACT targeted sequencing were also available (Fig. 1b). These derivative data were subsequently digitized (Fig. 1c) and used for multimodal modeling (Fig. 1d). We curated a retrospective cohort of 5145 (Fig. 1e) patients with HR + /HER2- EBC (MSK-BRCA; Fig. 1a; Supp. Figure 1) for model training, validation, and testing, whose primary tumors had H&E-stained FFPE tissue specimens available, textual pathology reports, and targeted panel sequencing for a subset (n = 481; Fig. 1b). We allocated these patients a priori into either a withheld test set (20%) or a set used for training and validation (80%; Supp. Tab. 1). Moreover, we assembled two additional independent cohorts of WSIs derived from patients with HR+/HER2- EBC, IEO-BRCA (452 patients) and MDX-BRCA37 (575 patients), for external validation. The patient’s age, sex, and race are reported in Supp. Tab. 2. A patient is considered high-risk with a molecular RS > 25, following TAILORx8.
Early-stage breast tumors are a resected, b profiled histologically, c digitized, and d used for downstream modeling of recurrence risk. e Number of pathologic slides, pathology reports, and patients included in each split. f Histogram depicting number of slides with a given number of tiles. g Tissue detection, tessellation, transformer-based modeling of CTransPath-derived tile embeddings, pathology report scraping, tokenization and transformer-based modeling, nuclear segmentation for interpretation, tensor fusion for multimodal integration. Created in BioRender. Boehm, K. (2025) https://biorender.com/s94u717.
Model training
We developed a transformer model to directly regress the ODX RS from WSIs of EBC. To train this architecture, we employed a two-step process. First, we projected each slide’s tissue-containing tiles (Fig. 1f) into an informative space using a frozen model trained using SSL on over 30,000 slides (Fig. 1g)38. Subsequently, we adapted a transformer architecture39, which was previously validated in a large multicenter study of colorectal cancer40, to map the phenotypic-genotypic correlation between the extracted features and the ODX RS (Fig. 1g). The unimodal and multimodal models were trained to regress RS as a continuous variable (Fig. 1g).
Deep learning infers recurrence risk score from whole-slide images
First, we develop and test the WSI-based model across the three cohorts to measure the generalizability of its performance. In the withheld MSK-BRCA test set, the unimodal WSI-based model achieved a Pearson correlation of 0.60 (95% C.I. 0.55–0.65, p < 10-4; Supp. Fig. 2a) and concordance correlation coefficient (CCC) of 0.57 (95% C.I. 0.52–0.62), along with the area under the precision-recall curve (AUPRC) of 0.55 (95% C.I. 0.47–0.64; Supp. Fig. 2d) and area under the receiver operating characteristic curve (AUROC) of 0.85 (95% C.I. 0.81–0.88; Fig. 2a) for high-risk disease. In the external IEO-BRCA test set, the same model achieved a Pearson correlation of 0.60 (95% C.I. 0.55–0.65; p < 10-4; Supp. Fig. 2b) and CCC of 0.58 (95% C.I. 0.52–0.63; Fig. 4b) along with AUPRC of 0.69 (95% C.I. 0.61–0.76; Supp. Fig. 2e) and AUROC of 0.81 (95% C.I. 0.77–0.85; Fig. 2b). In the external MDX-BRCA test set, which used an inferred, ODX-like RS (see Methods), the same model achieved a Pearson correlation of 0.58 (95% C.I. 0.53–0.63; p < 10-4; Supp. Figure 2c) and CCC of 0.40 (95% C.I. 0.35–0.45) along with AUPRC of 0.71 (95% C.I. 0.65–0.78; Supp. Fig. 2f) and AUROC of 0.80 (95% C.I. 0.76–0.84; Fig. 2c). Full results are detailed in the other panels of Supp. Figs. 2, 3 and in Supp. Data 1. In summary, the WSI-based model robustly infers RS and accurately identifies high-risk diseases across three test cohorts derived from different medical centers and countries.
a–c The whole-slide image (WSI)-based model reliably identifies high-risk disease (RS > 25) as defined by TAILORx across the MSK-BRCA (n = 1029), IEO-BRCA (n = 452), and MDX-BRCA (n = 572) test cohorts. d The multimodal model outperforms the WSI- and text-based unimodal models. Error bars by 1000-fold bootstrapping. e, f The multimodal model outperforms a clinicogenomic nomogram in identifying high-risk diseases. g Calibration plot and h predicted score frequencies. All results are shown for test sets.
Deep learning infers recurrence risk scores from text-based reports
Second, we develop a unimodal text report-based model that achieves a Pearson correlation of 0.59 (95% C.I. 0.53–0.65, p < 10-4) and CCC of 0.53 (95% C.I. 0.47–0.58; Supp. Fig. 4c) along with AUPRC of 0.53 (95% C.I. 0.45 - 0.61; Supp. Fig. 4f) and AUROC of 0.81 (95% C.I. 0.76–0.85; Supp. Fig. 4i) in the MSK-BRCA test set. Full results are detailed in Supp. Fig. 4 and Supp. Data 1. For the subset of male patients in the MSK-BRCA test set, the model demonstrates comparable performance (Supp. Tab. 3).
Multimodal integration of images and text improves recurrence risk score prediction
Finally, we evaluate whether the multimodal model integrating WSIs and text-based reports, Orpheus, improves predictive performance relative to the unimodal models. In the MSK-BRCA test set, the multimodal model achieved a Pearson correlation of 0.70 (Supp. Fig. 5a; 95% C.I. 0.65–0.74, p < 10-4) and CCC of 0.67 (95% C.I. 0.62–0.72). For classification of high-risk (RS > 25) disease, the AUPRC was 0.65 (p < 10-4; 95% C.I. 0.57–0.72), with a macro-averaged F1 score of 0.75 (Supp. Figure 5b). The CCC and Pearson’s correlation based on multimodal scores were higher than those based on unimodal scores (Fig. 2d; Supp. Fig. 5e). The AUROC was 0.88 (Supp. Fig. 5d; 95% C.I. 0.86–0.91, p < 10-4). A confusion matrix for the withheld test set is depicted in Supp. Fig. 5e, showing minimal misclassification between extreme categories, with moderate errors between intermediate and extreme categories (p < 10-4).
Multimodal recurrence risk model outperforms clinicopathologic nomogram
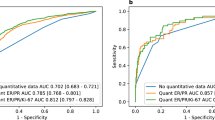
Next, we compare Orpheus to the state-of-the-art nomogram17 for predicting the ODX RS. Specifically, we analyzed the subset of the MSK-BRCA test set with available tumor grades and IHC-derived HR status in the text report as extracted by regular expressions. We compared the ability to discriminate high-risk disease of a nomogram based on clinical and pathologist-annotated features17 to that of the Orpheus, the multimodal (Fig. 2e; Supp. Fig. 6a), text-based (Supp. Fig. 6b, d), and whole-slide image-based (Supp. Fig. 6c, e) model. Orpheus achieved an AUROC of 0.89 and AUPRC of 0.67 (95% C.I. 0.58–0.75), the vision model achieved an AUPRC of 0.55 (95% C.I. 0.46–0.65), and the language model achieved an AUPRC of 0.57 (95% C.I. 0.48–0.66). By comparison, the nomogram17 achieved an AUPRC of 0.40 (95% C.I. 0.32–0.50). For the multimodal model, we suggest an operating point of 31.8 with 93% precision and 23% recall (Fig. 2f). The model was well calibrated for risk stratification based on >25 as a high-risk threshold, with most predicted scores being a low-risk (Fig. 2g, h).
Orpheus outperforms molecular risk score to identify patients with recurrent disease
Using 6814 hand-annotated cases, we tuned a large language model (see “Methods”) to infer recurrences from the electronic medical record, achieving an accuracy of 0.96 (Supp. Fig. 7). By analyzing cases with at least two years of follow-up and ODX RS values ≤ 25, we first used linear models to develop Orpheus+ models to infer risk of distant recurrence from H&E-stained WSIs. In the combined MSK-BRCA test and validation set, Orpheus+ ascertained the risk of distant recurrence with an AUROC of 0.77 (95% C.I. 0.68, 0.85)). This was superior to the Oncotype DX ® RS itself, which was uninformative in the RS ≤ 25 cohort (AUROC = 0.51 (95% C.I. 0.41, 0.61)). Scores differed significantly for cases with or without metastatic recurrence for Orpheus+ (p ≤ 1e-4), but not for ODX RS (p > 0.05; Fig. 3a).
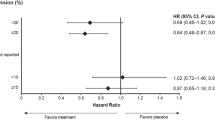
a, b Orpheus+ risk scores versus Oncotype DX ® (ODX, N = 1464) and Multiplex DX ® (MDX, N = 575) recurrence score (RS) values for cases with or without distant recurrence. p-values by Mann–Whitney-Wilcoxon test, two-sided. c Time-dependent areas under the receiver operating characteristic curve for Orpheus+ and ODX RS scores against recurrence in the MSK-BRCA test set for cases with ODX RS ≤ 25, with associated calibration plots. The mean value represents AUC over 2–8 years. d Receiver operating characteristic curves for Orpheus+ and MDX RS scores against recurrence in the MDX-BRCA test/validation set, with associated calibration plots. **** denotes p ≤ 1e−4, *** denotes p ≤ 1e-3, ** denotes p ≤ 1e-2, and * denotes – ≤ 5e-2. In box plots, boxes denote 25th–75th percentiles with lines at the median, whiskers denote the range without outliers, and individual points denote outliers. Exact p-values, top to bottom, for a are 5e-7 and 0.80, and for b, are 0.0013 and 0.0011.
Next, we calculated the dynamic time-dependent AUROC41 for Orpheus+ in a time-frame from two to eight years after surgery, accounting for censoring events to capture time-to-event dynamics encompassing the clinically relevant 5-year prediction window. Orpheus+ achieved a mean time-dependent AUROC of 0.75, whereas ODX RS attained a value of 0.49 (Fig. 3c). In the MDX-BRCA test set, we observed comparable identification of recurrence risk by Orpheus+ (AUROC = 0.64 (95% C.I. 0.56, 0.72)) to that by the Oncotype-like Multiplex DX ® laboratory assay risk score (AUROC = 0.64 (95% C.I. 0.57, 0.70)) itself (Fig. 3b, d). These results show the potential of Orpheus to improve the identification of recurrence risk for patients classified as low risk by TAILORx categories. Considering patients with any ODX RS (rather than only ≤25), the Orpheus+ performance is preserved, with a mean time-dependent AUROC of 0.72, and ODX RS achieves a value of 0.59 (Supp. Fig. 8a). Considering only patients in the MSK-BRCA test set (rather than the combined test and validation set) for cases with ODX RS ≤ 25, the Orpheus+ performance is preserved, with a mean time-dependent AUROC of 0.88, and ODX RS achieves a value of 0.50 (Supp. Fig. 8a). The characteristics of the distant recurrences in the test set are shown in Supp. Data 2. For full results, see Supp. Fig. 8.
Multimodal interpretability shows concordance with biological processes
Next, we sought to understand the decision-making process of the model’s predictions using Orpheus’ innate interpretability mechanisms. Specifically, using the attention mechanism of the image and text transformers of Orpheus, we visualized the importance of features within the WSIs and pathology reports. Visualizing the attention of the slide (Fig. 4a), the model designates most tiles as background with low-attention scores (Fig. 4b). Higher-attention tiles tend to contain invasive and in situ carcinoma compared to lower-attention tiles, which are more likely to contain fat and stroma (Fig. 4c; Supp. Fig. 9). Analogously to the tiles, the importance of word tokens comprising the synoptic pathology report can be analyzed (Supp. Fig. 10a). For the attention of the reports, analysis shows that words around immunohistochemical analyses for estrogen and progesterone receptors and lymphovascular invasion tend to have highest mean relative attention, alongside punctuation and descriptions of Nottingham grade (Supp. Fig. 10b). Ablating grade or progesterone receptor status from the text report decremented performance in the test set (Supp. Fig. 11). Analyzing the latent space of the learned embeddings of the trained models reveals separation by histologic grade (Supp. Fig. 12a) and progesterone receptor expression (Supp. Fig. 12b) in the MSK-BRCA test set, with the gradients appearing along a learned, lyre-shaped manifold for the multimodal model. The same was observed for the ODX RS itself (Supp. Fig. 12c). We further tested the association of predicted multimodal scores with genomic features. Limiting to cases with MSK-IMPACT, predicted RS was higher for tumors with TP53 mutations, MYC amplifications, PIK3CA amplifications, and BRCA2 mutations (Supp. Fig. 12d–g), and it trended slightly higher for specimens with a greater fraction of genome altered (Supp. Fig. 12h). In the test set alone, the relationships for TP53 and a fraction of genome altered persisted (Supp. Fig. 13).
a Saliency map of contributory foreground tiles, with one tile-attention value pair denoted by the arrows and predicted score. b Histogram of tile-attention values. c The five highest- and lowest-attention tiles at greater magnification. d Association of cellular features with high- and low-risk tissue. Hypothesis testing was performed with the two-sided Mann–Whitney U-test with corrections for multiple testing. e High and f low relative abundance of inflammatory cells. g High and h low standard deviation of neoplastic nuclear area. i–l Quantification of stromal fraction (SF), tumor cell proliferation (Prolif), lymphocyte infiltrating signature score (LISS), and lymphocyte fraction (LF) for predicted low- and high-risk patients (50 each) depicted in blue and orange, respectively, in the MSK-BRCA cohort. p-values are generated using an independent two-sided t-test. In box plots, boxes denote 25th–75th percentiles, whiskers denote the range without outliers, and individual points denote outliers. Scale bars denote 64 µm.
Orpheus provides histologic characterization of high-risk disease
To further explore the model’s capability of correlating histologic features with ODX RS, we analyze the most-attended tiles42 for high- and low-risk disease. The nuclei of these tiles were segmented43, and derivative features of cell type proportions and cellular morphology were tabulated (Fig. 4d). This reveals a relative abundance of inflammatory cells (Fig. 4e, f) and neoplastic cells along with the standard deviation of the neoplastic nuclear area (Fig. 4g, h) as some of the features differ significantly between the groups. Analysis of the tumor microenvironment reveals that high-risk disease exhibited greater stromal fraction (p < 10-4, n = 100) (Fig. 4i), tumor cell proliferation (p < 10-4, n = 100) (Fig. 4j), lymphocyte infiltration signature (p = 2 × 10-3, n = 100) (Fig. 4k), and leukocyte fraction (p < 10-4, n = 100) (Fig. 4l). Extending the tumor microenvironment analysis to external cohorts corroborated these results, especially for tumor cell proliferation which exhibited a significant difference between the predicted high- and low-risk disease patients (p < 10-4, n = 100) in all three cohorts (Supp. Fig. 14). As a further study of differences, we also trained a generative model to synthesize fields of view for informative tiles for high- and low-risk disease (Supp. Fig. 15). Tiles conditioned on the high-risk class depict confluent clusters of tumor cells with moderate to marked nuclear pleomorphism and prominent nucleoli, and tiles conditioned on the low-risk class depicted trabeculae and clusters of tumor cells with moderate nuclear pleomorphism and inconspicuous nucleoli. Tiles conditioned on the background class depicted stroma without epithelial cells.
Orpheus as a conceptual triaging tool for low- and high-risk disease
We test the utility of Orpheus as a pre-screening tool to reduce the load of laboratory testing for breast cancer recurrence risk in clinical workflows. First, we conducted a sensitivity analysis to evaluate the performance of the predicted recurrence risk score in identifying low-risk patients, defined as those with a risk recurrence score <11. The analysis yielded a sensitivity for the test set of MSK-BRCA (n = 1029, 16% predicted low-risk), IEO-BRCA (n = 452, 6% predicted low-risk), and MDX-BRCA (n = 572, 1% predicted low-risk), of 0.90 (Supp. Fig. 16a), 0.96 (Supp. Fig. 16b) and 0.99 (Supp. Fig. 16c), respectively, for the low-risk subgroup. Second, we conducted a specificity analysis to evaluate the performance of the predicted recurrence risk score in identifying high-risk patients, defined as those with a risk recurrence score >25 and who are most likely to benefit from adjuvant chemotherapy8. This resulted in a specificity of 0.94 (Supp. Fig. 16d), 0.68 (Supp. Fig. 16e), and 0.62 (Supp. Fig. 16f) to predict the high-risk subgroup for the test set of MSK-BRCA (n = 1029, 11% predicted high-risk), IEO-BRCA (n = 452, 45% predicted high-risk) and MDX-BRCA (n = 572, 61% predicted high-risk), respectively. Next, we repeated the analyses stratified by age, specifically patients above 50 years of age (Supp. Fig. 17) and patients below or equal to 50 years of age (Supp. Fig. 18), and nodal status, with similar performance metrics in all cohorts regardless of age and nodal status following the TAILORx risk groups8 (Supp. Data 3). Finally, we analyzed the model’s performance on the intermediate-risk (RS 11–25) subgroup using the AUROC, Cohen’s Kappa, F1 score, accuracy, and Matthew’s Correlation Coefficient (Supp. Tab. 4), utilizing additional clinically relevant thresholds of 10, 15 and 25 to binarize the risk predictions8. When observing the model’s performance, specifically on the intermediate-risk group (RS 11–25), we observe a substantial decrease in all metrics compared to all risk groups (RS 0–100).
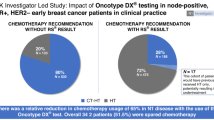
In summary, Orpheus accurately identifies patients with high-risk diseases, as defined by TAILORx, with a high degree of confidence. The model shows potential to guide adjuvant chemotherapy decisions without the need for multigene assay testing (Fig. 5). Specifically, adjuvant chemotherapy could be selectively recommended for a subset of patients classified as high-risk with high confidence. This approach could streamline treatment decisions and reduce the need for additional testing, ultimately improving patient care and resource allocation. Furthermore, for patients who are treated per TAILORx with adjuvant chemoendocrine or endocrine therapy based on ODX RS results, Orpheus identifies distant metastatic recurrences more accurately than ODX RS itself in the test set. With further validation, this prognostic value has the potential to refine patient selection for personalization of adjuvant treatments and follow-up strategies.
The Orpheus multimodal prediction model for recurrence risk prediction is potentially capable of guiding decision-making for adjuvant cytotoxic chemotherapy alongside adjuvant endocrine therapy for predicted low- and high-risk patients. The model is within scope for early-stage hormone receptor-positive (HR+) and HER2- breast cancer patients. Created in BioRender. Marra, A. (2025) https://BioRender.com/l96q019.
Discussion
The ODX RS is among the most extensively validated predictive and prognostic biomarkers, and given this widespread application, new applications continue to be developed. Our model, Orpheus, provides a way to approximate the continuous RS from routine H&E WSIs, extending the ODX RS and its myriad applications to centers where it is not feasible financially or logistically to order the laboratory assay itself13. In one example application, Orpheus can precisely identify approximately one-quarter of patients with high-risk disease as defined by TAILORx without the need for ODX RS, with superior discrimination of this class compared to a state-of-the-art nomogram, which integrates clinicopathologic features such as IHC-derived progesterone/estrogen receptor positivity, tumor size, lobular versus ductal histology, Nottingham grade, and age17. This would potentially enable physicians to forgo molecular testing in selected cases. Orpheus has the added advantage of not requiring manual curation of these features from the healthcare record. By inferring the continuous RS rather than identifying risk categories, Orpheus further enables emerging applications that tools such as the nomogram would not support, such as identification of risk of local recurrence, clinical trial eligibility44,45, or defining populations that will benefit from the use of neoadjuvant systemic therapies beyond the currently used clinical characteristics46. We further show that this correlation with RS corresponds to meaningful prognostication: for patients who are treated per TAILORx with adjuvant chemoendocrine or endocrine therapy based on ODX RS results, Orpheus identifies distant metastatic recurrences more accurately than ODX RS itself in the test set. With further validation, this prognostic value has implications ranging from tailoring the frequency of surveillance imaging and use in patient decision-making around treatment escalation and compliance with adjuvant endocrine therapy.
Moreover, the findings that multimodal approaches significantly outperform unimodal models further strengthen the broader perspective32,47,48 that integrating multimodal real-world data is a promising direction for AI in oncology. Orpheus is a flexible machine learning framework building on validated unimodal transformer-based architectures and data integration paradigms from the field of sentiment analysis, and the lightweight framework flexibly accepts tokens from the rapidly evolving foundation models for subsequent integration, allowing machine learning practitioners to harness the representative power of deep learning with small cohorts comprising only thousands of patients.
From a biological perspective, our study corroborates via orthogonal histopathologic and inferred transcriptomic analyses that greater immune infiltration portends higher-risk disease. This finding is in accordance with prior studies that tumor-infiltrating lymphocytes are a negative prognostic factor in HR + /HER2- EBC and may be associated with higher RS49,50,51,52. By estimating transcriptomic programs from images using our validated model53, proliferation was also found to be higher in our analysis of patients with predicted high risk, correlating with grade, the MKI67 gene included in the calculation of the RS, the TP53 mutations, and perhaps explaining the empiric association of more heterogeneous areas and perimeters of cancer cells with higher-risk disease. Finally, the greater inferred stromal fraction for higher-risk disease is possibly related to cancer associated fibroblasts and provides support for this line of inquiry54,55 Together, these findings show that deep learning interpretability is greatly improved by orthogonal molecular and cell-level data, which in turn can yield hypothesis driven insights for biological discovery. Future work should include spatial transcriptomic data for direct characterization of clonal heterogeneity and immune programming.
Taken together, this study advances an improved platform to approximate the ODX RS from routine histopathologic WSIs, outperforming a leading existing method in the identification of high-risk disease and—critically—identifying metastatic recurrences for cases with low ODX RS values more accurately than the ODX RS itself. The multimodal model improves performance when pathology text reports are available and is a lightweight and flexible machine learning architecture suitable for application to biomarkers for other histologies. The orthogonal histopathologic and transcriptomic analyses corroborate proliferation and tumor-infiltrating lymphocytes as markers of higher risk. Clinically, Orpheus has the potential to both expand access to precision medicine through ODX RS approximation and enhance its efficacy by identifying patients at risk of distant metastatic recurrence, even among those deemed low-risk by TAILORx-guided treatment.
Methods
Ethics statement
This study complies with all relevant ethical regulations and was approved by the Institutional Review Boards of Memorial Sloan Kettering Cancer Center and IEO Milan, along with the Ethics Commission of the Medical Faculty of the Technical University Dresden. For targeted clinical sequencing, patients provided informed consent for enrollment on a prospective protocol as overseen by the Memorial Sloan Kettering Cancer Center Institutional Review Board. Otherwise, informed consent was waived for this retrospective study, and participants were not compensated.
Statistics and reproducibility
This study was conducted retrospectively. Therefore, no statistical method was used to predetermine sample size, the experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment. The data is split into a training and validation cohort for training of Orpheus, MSK-BRCA, and external validation cohorts, IEO-BRCA and MDX-BRCA. Model training included both male and female cases irrespective of age and nodal status, followed by stratified analyses to evaluate performance in each subgroup. Sex was determined based on sex assigned at birth reported by the institutional database. Before any analysis, the MSK-BRCA cohort was split into a training/validation set and a withheld test set using 80% and 20% of patient IDs, respectively. The model evaluation consisted of two steps: first, comparing Orpheus’s predicted risk scores against OncotypeDX recurrence scores using correlation metrics and risk group classification performance; second, assessing both Orpheus and ODX RS predictions for distant metastatic recurrence through calibration and time-dependent analyses. Unless otherwise specified, we consistently define high-risk as scores >25 and low-risk as scores ≤25, following TAILORx thresholds. This standardized cut-off applies to both Orpheus and OncotypeDX recurrence scores, independent of age or nodal status, throughout the entire manuscript for direct comparability between the predicted Orpheus scores and OncotypeDX recurrence scores. Additional statistical and experimental details, including sample inclusion and exclusion criteria, are provided in the subsequent cohort curation, model training, and model evaluation sections. For Fig. 4c, micrographs were selected by attention scores of one random slide. For Fig. 4e–h, representative fields of view with high or low values of the feature in question were selected. The experiments were not repeated to generate different micrographs.
Cohort curation
For the MSK cohort, cases from 2013-2020 were selected according to Supp. Fig. 1 for this retrospective analysis. All cases were pathologically confirmed HR+/HER2- invasive breast carcinoma without distinction by specific histologic subtype. Pathology reports, and slides were joined by the surgical pathology part number rather than case or block. The synoptic pathology text report for the part used to calculate the RS was included, with fields such as histologic subtype, HR and HER2 IHC percent positivity, histologic grade, anatomic site, and DCIS and LCIS. Oncotype DX results were recorded manually from the healthcare record by a medical oncologist. Regular expressions were used to extract progesterone, estrogen, and HER2 receptor percentage positivity from the pathology text reports. For the external validation cohorts, the cohort from the European Institute of Oncology (IEO-BRCA, Milan, Italy) contained a total of 456 early-stage breast cancer patients who received the official Oncotype DX test. Only histopathology slides and corresponding clinicopathological variables were available for analysis. After filtering down patients based on histology slide availability, 452 patients in the IEO-BRCA cohort were available for external validation. The cohort from MultiplexDX (MDX-BRCA, Bratislava, Slovakia) contained a total of 1013 early-stage breast cancer patients originally obtained for a retrospective study37 from Biobank Graz of the Medical University of Graz, (n = 390, Graz, Austria) and PATH Biobank (n = 592, Munich, Germany), The Biomedical Research Institute of Málaga (n = 27, IBIMA-CIMES-UMA, Malaga, Spain), and a commercial company (n = 4, AMBIO). Histopathology slides, corresponding clinicopathological variables, and research-based Oncotype DX scores derived from RNA-sequencing were available for analysis. After filtering down patients based on histology slide availability and nodal, ER, PR, and HER2 status, which would have been eligible for Oncotype DX, 575 patients in the MDX-BRCA cohort remained for external validation. The research-based scores were calculated using the GeneFu Bioconductor package56, based on the original algorithm to calculate the OncotypeDX score2. Because research-based versions of OncotypeDX use different data inputs (e.g., microarray/RNA-seq) compared to the official OncotypeDX (e.g., RT-qPCR), this may result in scaling effects when comparing research-based scores with official scores, as demonstrated by the OPTIMA trial group57. Consequently, we used their outlined approach57, where they provide a linear equation that models the relationship between research-based versus true OncotypeDX, to rescale the research-based recurrence score into a more realistic range. All included cases were hormone receptor-positive and HER2-negative as defined by the American Society of Clinical Oncology/College of American Pathologists clinical practice guidelines35,36. Experienced breast cancer pathologists from each Institution reviewed the case to confirm the diagnosis of invasive breast cancer and receptor status.
Training setup of vision, language, and multimodal models
For vision model training, images were preprocessed using STAMP with 1.14 microns per pixel, a tile edge length of 224 pixels, and Macenko normalization58. Tiles were embedded using CTransPath38. After a fully connected layer to project the CTransPath-derived tokens into 512-dimensional space and a ReLU, two PyTorch TransformerEncoderLayers with dimensionality 512 and eight heads were stacked before a final LayerNorm and projection to scalar space. No activation function was used, and no positional encodings were used. For training, a maximum learning rate of 2e-5 with a linear warmup of 1000 steps, learning rate decay by a factor of 0.9999 every step, and L2 decay of 2e-5 were used. A batch size of one slide per GPU across two GPUs with accumulated gradients over four batches was used, with gradients clipped at 0.5. The model was trained for up to 50 epochs with early stopping. This was implemented in PyTorch-Lightning and ran on two NVIDIA Tesla V100 GPUs (CUDA 12.1) on a cluster running Linux. Metrics were tracked using Weights and Biases, and the model with the lowest validation mean squared error was chosen for downstream use. During inference, attention rollout42. was used to attribute attention to each input token, and multiple predictions with dropout enabled were used to estimate confidence intervals. Regression was used instead of classification because classification discards information, and it substantially outperformed classification in our validation set53. For cases with multiple slides corresponding to the same pathologic specimen, all the tiles were bagged prior to any transformer-based analysis.
For language model training, the HuggingFace model tsantos/PathologyBERT was tuned using the HuggingFace BertForSequenceClassification, Trainer, and AutoTokenizer with a batch size of eight-part descriptions per GPU, four Nvidia Tesla V100 GPUs, four gradient accumulations per backprop, a learning rate of 2e-5, L2 decay of 0.01, and ten training epochs. Prior to tokenization, the text corresponding to the part used to measure the Oncotype score was extracted using regular expressions. Addenda, when available, was concatenated to the part description. Names, initials, and the most common logistical comments were removed prior to tokenization. The tsantos/PathologyBERT tokenizer was not modified. The model with the lowest validation loss was chosen for downstream use. During inference, attention rollout was used to attribute attention to each input token, and multiple predictions with dropout enabled were used to estimate confidence intervals.
For multimodal model training, multiple architectures, including simple concatenation of embeddings before dense layers, attention-based integration of unimodal embeddings, or averaging of scores, were explored using the validation set. The final model chosen took the pre-computed unimodal embeddings as input, projected them into 96-dimensional space, performed tensor fusion by prepending unity and taking the Cartesian product59, applied 30% dropout, and passed through a small 96-dimensional regression head to yield a scalar regression score. Finally, linear regression trained on the training set was used to calibrate the weight of the unimodal and multimodal scores to yield a final score. Multiple predictions with dropout enabled were used to estimate confidence intervals. UMAP plots were generated using the Python umap software package with 10 neighbors and min_dist 0.5 for all plots fit on the training set, and only test sets are shown.
Model evaluation
When multiple slides were available for a single measured score, and all contained relevant tissue as per preprocessing, the relevant tissue tiles were bagged prior to inference or training by the vision model. Models were evaluated primarily by Pearson correlation and associated significance and concordance correlation coefficient60. 95% confidence intervals were calculated using bootstrapping (random sampling with replacement) 1000 times. Areas under the precision-recall and receiver operating characteristic curve were calculated using binary thresholding of high- and low-risk disease. Significance was established using 1000-fold permutation tests. We analyzed operating points on the precision-recall curve by varying the threshold from greatest to lowest and tabulating the respective precision and recall for each value. F1 scores were calculated using the weighted average. Confusion matrices were established using the three risk categories (<11, 11–25, >25), and significance was established using McNemar’s test of homogeneity. To further evaluate the performance of our model, we utilized the AUROC, F1 score, accuracy, Cohen’s Kappa, and Matthew’s Correlation Coefficient (MCC). These metrics were assessed across all risk groups (0–100) and specifically within the intermediate-risk group (11–25). To investigate the model’s performance using various clinically-defined thresholds in Supp. Data 3, we applied binary risk stratification (low versus high) at cut-off values of 10, 15, and 25, analyzing both the entire population and subgroups stratified by age and nodal status. This approach allowed us to thoroughly examine the model’s predictive capabilities across populations with different definitions of risk and to evaluate its performance within the clinically challenging intermediate-risk group. To compare Orpheus against the state-of-the-art clinical nomogram, logistic regression for high-risk disease was performed using the formula established by Orucevic et al.17, including age, histological classification, tumor grade, PR regression status, and tumor size, with corresponding coefficients and an intercept term. The features were extracted using regular expressions from the pathology report. Reports with failed extraction (e.g., due to the absence of hormone receptor annotation in the pathology report or unusual formatting precluding the extraction of tumor size) were excluded from the comparison analysis. Precision-recall analyses were performed using the predicted score [0, 100], in the case of our transformer regression, and the logistic regression score [0, 1] in the case of the nomogram’s logistic regression formula. Summarizing, we evaluated Orpheus through a two-pronged approach. First, we assessed its ability to predict ODX RS values by comparing sample-level continuous risk predictions using Pearson correlation and concordance correlation coefficient. We then evaluated its performance in classifying patients into TAILORx risk groups, measuring agreement with the gold-standard RS categorization through multiple metrics: AUROC, AUPRC, F1 score, accuracy, Cohen’s Kappa, and Matthew’s Correlation Coefficient. In the second step, we compared both Orpheus and RS against actual clinical outcomes, specifically distant metastatic recurrence. This comparison utilized calibration plots and time-dependent AUROCs, which account for censoring events to capture time-to-event dynamics beyond the clinically relevant 5-year prediction window.
Patient outcome modeling
We developed a method to identify primary breast cancer patients with metastatic recurrence from a combination of medical oncology notes, radiology notes, tumor marker lab values and internal referral data using natural language processing (NLP) and machine learning methods. Briefly, a pretrained Clinical longformer model pretrained on MIMIC III data (https://huggingface.co/yikuan8/Clinical-Longformer) was selected to predict metastatic recurrence from medical oncology notes. Our model was finetuned on a note dataset consisting of the note closest to the follow-up date of the patient and a random sample of three notes following the date of surgery for N = 6814 patients with disease status labels curated from the Breast Disease Management Team at Memorial Sloan Kettering. Notes prior to the date of recurrence were labeled ‘early’ while notes following the date of local and metastatic recurrence were labeled ‘local’ and ‘metastatic’ respectively. On a held-out dataset of N = 1363 primary breast cancer patients, the finetuned Clinical Longformer model was tested. Ensembling was performed: specifically, the local and metastatic probabilities of the NLP model were further included as features of a Random Forest model that also included outputs from a metastatic sites model (Clinical BERT), progression probability (RoBERTa), positive tumor markers (CEA, CA 15-3, CA 125), and indicators of internal referral. The MDX-BRCA cases were manually annotated as distant, non-recurrent, or uncertain recurrences, where solely cases with ≥24 months follow up time were considered for these analyses. Distant metastatic recurrences in the MSK-BRCA test set were manually validated and tabulated. For the subsequent outcome modeling, bagged WSI-based embeddings were generated for each case. Using the same train/validation/test split of the MSK-BRCA dataset, a logistic regression model implemented in scikit-learn with C = 1e-5 was trained with class weights calculated by the scikit-learn compute_class_weight utility given the class imbalance. Calibration was performed using the sigmoid method with five-fold cross-validation in the training set. The model was applied to the combined MSK-BRCA test and MSK-BRCA validation sets and the MDX-BRCA test set. Orpheus+ models were trained separately for any recurrence, locoregional recurrence, and distant metastatic recurrence. Calibration curves were plotted using the corresponding scikit-learn functions, comparing predicted probabilities from Orpheus with observed outcomes, accounting for censoring events. Moreover, the time-dependent area under the receiver operating characteristic (AUROC) curves were plotted using scikit-survival with three-month bins from 24 to 96 months, accounting for censoring events to capture time-to-event dynamics beyond the clinically relevant 5-year prediction window. Mann–Whitney U-tests were used to compare inferred risk scores for cases with, and without, the type of recurrence under analysis.
Model interpretability
To interpret model predictions, we employed multiple analytical approaches: visualizing attention patterns in text reports for the language component of the model, examining nuclear features, generating tissue representations, quantifying tumor microenvironment components, and conducting gene association analyses across risk groups for the vision component of the model.
For report visualization, Django was used to colorize each token in the description by relative attention. A similar color scale in HSV space was applied to tint each tile by its relative attention, with absolute attention plotted on corresponding histograms with logarithmic counts displayed. The five tiles with the highest and lowest absolute attention are displayed for quality control by users.
For nuclear analysis, the three most informative tiles from each slide with the highest 100 and lowest 100 predicted scores were identified. HoVerNet43 with PanNuke-derived weights was used for instance segmentation, and quantitative features such as solidity, area, and perimeter were calculated for the outline of each nucleus. Within each tile, summative statistics were used to aggregate these features. The feature with the highest variance inflation factor was iteratively removed until the highest variance inflation factor was 10 or less. Comparisons for the values of these remaining features between the high- and low-risk groups were made with the two-sided Mann–Whitney U-test with corrections for multiple testing. For the volcano plot, log fold change was calculated as the base-two logarithm of the mean value for the high-risk group over the same for the low-risk group. Tiles with fewer than 50 total nuclei were excluded. For comparisons involving standard deviations, tiles with fewer than ten of the relevant cell types were excluded.
For the generation of tissue representations across the risk groups, we identified the highest-attention tiles from cases with measured recurrence scores below 11 or above 25 as chosen by an Attn-MIL model61 trained on the training set. Subsequently, using these tiles and a random sampling of low-attention tiles across the same slides, we used Studio GAN62 to train a ReACGAN architecture with big_resnet backbone, batch size of 36 per GPU across four NVIDIA Tesla V100 GPUs, and default loss parameters. The conditional architecture encoded three classes: high score, low score, and background. Spectrum plots and canvases were generated as per the default StudioGAN code.
For the quantification of the tumor microenvironment, we used a pretrained deep learning model for the quantification of the tumor microenvironment for the top 50 predicted high- and low-risk patients by the recurrence risk vision prediction model, specifically for the stromal fraction (SF) and leukocyte fraction (LF) as assessed via DNA methylation analysis, lymphocyte infiltrating signature score (LISS) and proliferation (Prolif) as measured by RNA expression63. The deep learning regression model was trained on whole-slide images from a breast cancer cohort from The Cancer Genome Atlas (TCGA) in a weakly-supervised setting53 using the open-source biomarker data from Thorsson et al.63 Statistical significance is measured by an independent t-test, indicating a difference in sample means between predicted high- and low-risk patients (p < 0.05). The scores for the tumor microenvironment quantification are inferred based on the same tile embeddings (1×768) which were used in training the vision model for the recurrence risk prediction.
To analyze the association of genetic patterns with predicted risk groups, the specimens were sequenced by MSK-IMPACT64, annotated by OncoKB65, and accessed via cBioPortal66. Only specimens attributed to a patient without other specimens were considered to avoid part-level mismatch, and specimens across the MSK-BRCA training, validation, and test set were included. Variants in the most commonly altered genes in the MSKCC Clinical Sequencing Cohort (TP53, PTEN, BRCA2, FGF4, KMT2C, PAK1, FGF19, CDH1, MYC, ARID1A, FGF3, GATA3, CBFB, CCND1, RUNX1, PIK3CA, FGFR1, NSD3, MAP3K1) were considered, provided that the variant in question (e.g., TP53 SNV) occurred in at least two of the considered samples. Passenger mutations were ignored. Using Bonferroni correction, genes associated with high- or low-risk status based on the measured, ground truth recurrence score with a significance of q = 0.05 were identified. The fraction of genome altered, tumor mutational burden, and mutation count were analyzed using the Pearson correlation with the same significance and correction.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
In accordance with the institutional policies of IEO and MSK, all requests for data collected or curated in-house will be evaluated on a case-by-case basis to determine whether the data requested and the use case comply with intellectual property and patient privacy obligations. For the MDX-BRCA cohort, patient-level research-based OncotypeDX scores, digitized whole-slide images of H&E, and clinicopathological data cannot be publicly shared due to privacy concerns, existing material and data transfer agreements between MultiplexDX and participating biobanks, and proprietary ownership by MultiplexDX. Qualified researchers may apply for access to these data through the MultiplexDX Data Access Committee (DAC) by sending an initial request to the following email address: info@multiplexdx.com. Then, the qualified researcher would submit a brief research proposal and a standard form describing the project, data/materials requested, applicable ethics, and purpose. Requests will be reviewed and discussed by the DAC based on scientific merit. The time-frame of response to an initial request is about 1-2 months. After approval, the parties will agree on the conditions of a data access/sharing agreement and restrictions of use, which may increase the total time-frame to around 6 months. Source data are provided in this paper.
Code availability
The source code is available under an open-source license on GitHub. The multimodal modeling package, Orpheus, is available at https://github.com/kmboehm/orpheus. The preprocessing pipeline for whole-slide images is found at https://github.com/KatherLab/STAMP, and our code for regressing transcriptomic programs from images is found at https://github.com/KatherLab/marugoto/releases/tag/v1.0.0-regression. The GAN was trained using https://github.com/POSTECH-CVLab/PyTorch-StudioGAN, with our weights and configuration parameters at https://www.synapse.org/breastGAN. The code to calculate nuclear features based on HoverNet inference is at https://gist.github.com/kmboehm/aea77f24a9cdbb1f246dacaae812053d.
References
Giaquinto, A. N. et al. Breast Cancer Statistics, 2022. CA Cancer J. Clin. 72, 524–541 (2022).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 351, 2817–2826 (2004).
Morganti, S. et al. Refining risk stratification in HR-positive/HER2-negative early breast cancer: how to select patients for treatment escalation? Breast Cancer Res. Treat. 192, 465–484 (2022).
Nitz, U. et al. Reducing chemotherapy use in clinically high-risk, genomically low-risk pN0 and pN1 early breast cancer patients: five-year data from the prospective, randomised phase 3 West German Study Group (WSG) PlanB trial. Breast Cancer Res. Treat. 165, 573–583 (2017).
Dowsett, M. et al. Prediction of risk of distant recurrence using the 21-gene recurrence score in node-negative and node-positive postmenopausal patients with breast cancer treated with anastrozole or tamoxifen: a TransATAC study. J. Clin. Oncol. 28, 1829–1834 (2010).
Paik, S. et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24, 3726–3734 (2006).
Albain, K. S. et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, oestrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomised trial. Lancet Oncol. 11, 55–65 (2010).
Sparano, J. A. et al. Adjuvant chemotherapy guided by a 21-gene expression assay in breast cancer. N. Engl. J. Med. 379, 111–121 (2018).
Kalinsky, K. et al. 21-Gene assay to inform chemotherapy benefit in node-positive breast cancer. N. Engl. J. Med. 385, 2336–2347 (2021).
Andre, F. et al. Biomarkers for adjuvant endocrine and chemotherapy in early-stage breast cancer: ASCO guideline update. J. Clin. Oncol. 40, 1816–1837 (2022).
Loibl, S. et al. Early breast cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up†. Ann. Oncol. https://doi.org/10.1016/j.annonc.2023.11.016 (2023).
Gradishar, W. J. et al. NCCN Guidelines® Insights: Breast Cancer, Version 4.2023: Featured Updates to the NCCN Guidelines. J. Natl. Compr. Canc. Netw. 21, 594–608 (2023).
Paulden, M. et al. Cost-effectiveness of the 21-gene assay for guiding adjuvant chemotherapy decisions in early breast cancer. Value Health 16, 729–739 (2013).
Özmen, V. et al. Cost effectiveness of gene expression profiling in patients with early-stage breast cancer in a middle-income country, turkey: results of a prospective multicenter study. Eur. J. Breast Health 15, 183–190 (2019).
de Jongh, F. E., Efe, R., Herrmann, K. H. & Spoorendonk, J. A. Cost and clinical benefits associated with Oncotype DX® test in patients with early-stage HR+/HER2- node-negative breast cancer in the netherlands. Int. J. Breast Cancer 2022, 5909724 (2022).
Berdunov, V. et al. Cost-effectiveness analysis of the Oncotype DX Breast Recurrence Score test in node-positive early breast cancer. J. Med. Econ. 25, 591–604 (2022).
Orucevic, A., Bell, J. L., King, M., McNabb, A. P. & Heidel, R. E. Nomogram update based on TAILORx clinical trial results—Oncotype DX breast cancer recurrence score can be predicted using clinicopathologic data. Breast 46, 116–125 (2019).
Su, Z. et al. BCR-Net: a deep learning framework to predict breast cancer recurrence from histopathology images. PLoS ONE 18, e0283562 (2023).
Baltres, A. et al. Prediction of Oncotype DX recurrence score using deep multi-layer perceptrons in estrogen receptor-positive, HER2-negative breast cancer. Breast Cancer 27, 1007–1016 (2020).
Li, H. et al. Deep learning-based pathology image analysis enhances Magee feature correlation with oncotype DX breast recurrence score. Front. Med. 9, 886763 (2022).
Klein, M. E. et al. Prediction of the oncotype DX recurrence score: use of pathology-generated equations derived by linear regression analysis. Mod. Pathol. 26, 658–664 (2013).
Romo-Bucheli, D., Janowczyk, A., Gilmore, H., Romero, E. & Madabhushi, A. A deep learning based strategy for identifying and associating mitotic activity with gene expression derived risk categories in estrogen receptor positive breast cancers. Cytometry A 91, 566–573 (2017).
Romo-Bucheli, D., Janowczyk, A., Gilmore, H., Romero, E. & Madabhushi, A. Automated tubule nuclei quantification and correlation with oncotype DX risk categories in ER+ breast cancer whole slide images. Sci. Rep. 6, 32706 (2016).
Cho, S. Y. et al. Author Correction: Deep learning from HE slides predicts the clinical benefit from adjuvant chemotherapy in hormone receptor-positive breast cancer patients. Sci. Rep. 11, 21043 (2021).
El Agouri, H. et al. Assessment of deep learning algorithms to predict histopathological diagnosis of breast cancer: first Moroccan prospective study on a private dataset. BMC Res. Notes 15, 66 (2022).
Chen, Y. et al. Computational pathology improves risk stratification of a multi-gene assay for early stage ER+ breast cancer. NPJ Breast Cancer 9, 40 (2023).
Skrede, O.-J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360 (2020).
Kleppe, A. et al. A clinical decision support system optimising adjuvant chemotherapy for colorectal cancers by integrating deep learning and pathological staging markers: a development and validation study. Lancet Oncol. 23, 1221–1232 (2022).
Reis-Filho, J. S. & Kather, J. N. Overcoming the challenges to implementation of artificial intelligence in pathology. J. Natl. Cancer Inst. 115, 608–612 (2023).
Xu, H. et al. Vision transformers for computational histopathology. IEEE Rev. Biomed. Eng. 17, 63–79 (2024).
Boehm, K. M. et al. Multimodal data integration using machine learning improves risk stratification of high-grade serous ovarian cancer. Nat Cancer 3, 723–733 (2022).
Boehm, K. M., Khosravi, P., Vanguri, R., Gao, J. & Shah, S. P. Harnessing multimodal data integration to advance precision oncology. Nat. Rev. Cancer https://doi.org/10.1038/s41568-021-00408-3 (2021).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer Cell 40, 865–878.e6 (2022).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual-language foundation model for pathology image analysis using medical Twitter. Nat. Med. 29, 2307–2316 (2023).
Wolff, A. C. et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J. Clin. Oncol. 36, 2105–2122 (2018).
Allison, K. H. et al. Estrogen and progesterone receptor testing in breast cancer: ASCO/CAP Guideline Update. J. Clin. Oncol. 38, 1346–1366 (2020).
Paul, E. D. et al. The spatially informed mFISHseq assay resolves biomarker discordance and predicts treatment response in breast cancer. Nat. Commun. 16, 226 (2025).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Dosovitskiy, A. et al. An image is worth 16x16 words: transformers for image recognition at scale. International Conference on Learning Representations (2021).
Wagner, S. J. et al. Fully transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. https://arxiv.org/abs/2301.09617 (2023).
Kamarudin, A. N., Cox, T. & Kolamunnage-Dona, R. Time-dependent ROC curve analysis in medical research: current methods and applications. BMC Med. Res. Methodol. 17, 53 (2017).
Abnar, S. & Zuidema, W. Quantifying attention flow in transformers. https://arxiv.org/abs/2005.00928 (2020).
Graham, S. et al. Hover-Net: simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 58, 101563 (2019).
White, J. R. et al. NRG-BR007: A phase III trial evaluating de-escalation of breast radiation (DEBRA) following breast-conserving surgery (BCS) of stage 1, hormone receptor+, HER2-, RS ≤18 breast cancer. J. Clin. Oncol. 40, https://doi.org/10.1200/JCO.2022.40.16_suppl.TPS613 (2022).
Braunstein, L. Z. & Taghian, A. G. Molecular phenotype, multigene assays, and the locoregional management of breast cancer. Semin. Radiat. Oncol. 26, 9–16 (2016).
Smith, I. et al. Long-term outcome and prognostic value of Ki67 after perioperative endocrine therapy in postmenopausal women with hormone-sensitive early breast cancer (POETIC): an open-label, multicentre, parallel-group, randomised, phase 3 trial. Lancet Oncol 21, 1443–1454 (2020).
Lipkova, J. et al. Artificial intelligence for multimodal data integration in oncology. Cancer Cell 40, 1095–1110 (2022).
Sammut, S.-J. et al. Multi-omic machine learning predictor of breast cancer therapy response. Nature 601, 623–629 (2022).
Caparica, R. et al. Tumour-infiltrating lymphocytes in non-invasive breast cancer: a systematic review and meta-analysis. Breast 59, 183–192 (2021).
Miglietta, F. et al. Association of tumor-infiltrating lymphocytes with recurrence score in hormone receptor-positive/HER2-negative breast cancer: Analysis of four prospective studies. Eur. J. Cancer 195, 113399 (2023).
Ahn, S. G. et al. Comparisons of tumor-infiltrating lymphocyte levels and the 21-gene recurrence score in ER-positive/HER2-negative breast cancer. BMC Cancer 18, 320 (2018).
Kolberg-Liedtke, C. et al. Association of TILs with clinical parameters, Recurrence Score® results, and prognosis in patients with early HER2-negative breast cancer (BC)-a translational analysis of the prospective WSG PlanB trial. Breast Cancer Res. 22, 47 (2020).
El Nahhas, O. S. M. et al. Regression-based deep-learning predicts molecular biomarkers from pathology slides. Nat. Commun. 15, 1–13 (2024).
Guo, Z. et al. Cancer-associated fibroblasts induce growth and radioresistance of breast cancer cells through paracrine IL-6. Cell Death Discov. 9, 6 (2023).
Wang, B. et al. Cancer-associated fibroblasts promote radioresistance of breast cancer cells via the HGF/c-met signaling pathway. Int. J. Radiat. Oncol. Biol. Phys. 116, 640–654 (2023).
Gendoo, D. M. A. et al. Genefu: an R/Bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics 32, 1097–1099 (2016).
Bartlett, J. M. S. et al. Computational approaches to support comparative analysis of multiparametric tests: Modelling versus Training. PLoS ONE 15, e0238593 (2020).
El Nahhas, O. S. M. et al. From whole-slide image to biomarker prediction: end-to-end weakly supervised deep learning in computational pathology. Nat. Protoc. 20, 293–316 (2024).
Zadeh, A., Chen, M., Poria, S., Cambria, E. & Morency, L.-P. Tensor fusion network for multimodal sentiment analysis. In: Proceedings of the 2017 Conference on Empirical Methods in Natural Language Processing (Copenhagen, Denmark, 2017).
Lin, L. I. A concordance correlation coefficient to evaluate reproducibility. Biometrics 45, 255–268 (1989).
Ilse, M., Tomczak, J. M. & Welling, M. Attention-based deep multiple instance learning. https://arxiv.org/abs/1802.04712 (2018).
Kang, M., Shin, J. & Park, J. StudioGAN: a taxonomy and benchmark of GANs for image synthesis. IEEE Trans. Pattern Anal. Mach. Intell. 45, 15725–15742 (2023).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830.e14 (2018).
Cheng, D. T. et al. Memorial Sloan kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 2017, PO.17.00011 (2017).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, l1 (2013).
Acknowledgements
J.N.K. is supported by the German Federal Ministry of Health (DEEP LIVER, ZMVI1-2520DAT111), the German Cancer Aid (DECADE, 70115166), the German Federal Ministry of Education and Research (PEARL, 01KD2104C; CAMINO, 01EO2101; SWAG, 01KD2215A; TRANSFORM LIVER, 031L0312A; TANGERINE, 01KT2302 through ERA-NET Transcan), the German Academic Exchange Service (SECAI, 57616814), the German Federal Joint Committee (TransplantKI, 01VSF21048) the European Union’s Horizon Europe and innovation program (ODELIA, 101057091; GENIAL, 101096312), the European Research Council (ERC; NADIR, 101114631) and the National Institute for Health and Care Research (NIHR, NIHR213331) Leeds Biomedical Research Center. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR, or the Department of Health and Social Care. This work was funded by the European Union. Views and opinions expressed are, however, those of the author(s) only and do not necessarily reflect those of the European Union. Neither the European Union nor the granting authority can be held responsible for them. F.P. is funded in part by an NIH/NCI P50 CA247749 01 grant and by a Starr Cancer Consortium grant. A.M. is supported by the ESMO José Baselga Fellowship for Clinician Scientists founded by AstraZeneca (2023-2025). S.P.S. received funds from the Warren Alpert Center for Computational Pathology and the Ovarian Cancer Research Alliance and is a Susan G. Komen Scholar. This work was also supported in part by a National Cancer Institute Cancer Center Support Grant (P30 CA008748; MSKCC), the Molecular Diagnostics Service in the Department of Pathology (MSKCC), and the Marie-Josee and Henry R. Kravis Center for Molecular Oncology (MSKCC). J.S.R.-F. was funded in part by the Breast Cancer Research Foundation, by a Susan G Komen Leadership grant, and by the NIH/NCI P50 CA247749 01 grant. B.W. is funded in part by the Breast Cancer Research Foundation and Cycle for Survival grants. S.C. is funded in part by the Breast Cancer Research Foundation. Generation of the MDX-BRCA dataset was supported by research funding from MultiplexDX and the European Union’s Horizon 2020 research and innovation program under an EIC Accelerator grant (agreement No 946693) awarded to MultiplexDX s.r.o. (P.C. as PI). E.D.P. and P.C. are partly funded by the European Union’s NextGenerationEU through the Recovery and Resilience Plan for Slovakia under project No. 09I03-03-V03-00101 (Project PI: EDP). P.R. discloses funding from the NIH, Komen, Breast Cancer Alliance, BCRF, Grail, Illumina, Novartis, AstraZeneca, Epic Sciences, Invitae/ArcherDx, Biothernostics, Tempus, Inivata, Biovica, Guardant, Personalis, Myriad.
Author information
Authors and Affiliations
Contributions
K.M.B. and O.S.M.E.N. developed the software and analyzed the data. A.M., F.P., H.W., B.W., F.P., L.B., P.R., P.S., S.C., R.E., K.M.B., and J.N.K. interpreted the results through lenses of clinical oncology and pathology. A.M. collected the MSK-BRCA cohort. N.S., M.W., and J.J. developed the machine learning model for the identification of MSK-BRCA patients with a metastatic recurrence. A.M., E.G., N.F., C.F., Eltjona M, Elisabetta M, S.D., P.Z., G.C. collected the IEO-BRCA cohort. E.D.P. and P.C. collected and conducted analysis for MDX-BRCA cohort. R.E. and C.M.L.L. assisted with data analysis. S.C., J.R.F., S.P.S., and J.N.K. designed the study. S.P.S. and J.N.K. supervised the study. All authors contributed to writing and reviewing the report approved the final version for submission, and have agreed to be personally accountable for the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the report.
Corresponding authors
Ethics declarations
Competing interests
J.S.R.-F. reports receiving personal/consultancy fees from Goldman Sachs, Bain Capital, REPARE Therapeutics, Saga Diagnostics, and Paige.AI, membership of the scientific advisory boards of VolitionRx, REPARE Therapeutics and Paige.AI, membership of the Board of Directors of Grupo Oncoclinicas, and ad hoc membership of the scientific advisory boards of AstraZeneca, Merck, Daiichi Sankyo, Roche Tissue Diagnostics and Personalis, outside the scope of this study. He is also currently employed at AstraZeneca. OSMEN holds shares in StratifAI GmbH. JNK declares consulting services for Owkin, France, DoMore Diagnostics, Norway, Panakeia, UK, Scailyte, Switzerland, Cancilico, Germany, Mindpeak, Germany, MultiplexDx, Slovakia, and Histofy, UK; furthermore, he holds shares in StratifAI GmbH, Germany, has received a research grant by GSK, and has received honoraria by AstraZeneca, Bayer, Eisai, Janssen, MSD, BMS, Roche, Pfizer and Fresenius. The mentioned competing interests are related to cancer and the computational analysis of histopathology slides, which is the main topic of this research. SPS conducts collaborative research with AstraZeneca Inc., outside the scope of this study. FP reports receiving consultancy fees from AstraZeneca and reports membership of advisory boards of MultiplexDx and AstraZeneca. EDP and PC are employed at MultiplexDX. A.M. has received honoraria as a consultant, advisor, or speaker from Roche and Menarini/Stemline and has received travel and accommodation support from AstraZeneca. E.M. received honoraria for Consulting or Advisory Role: Exact Sciences, MSD Oncology, Daiichi Sankyo/AstraZeneca, Pfizer, Seagen, Ipsen. Travel, Accommodations, Expenses: Roche, Pfizer, Lilly, Novartis, Gilead Sciences, AstraZeneca, Pierre Fabre. R.E. has received honoraria from Roche, Eisai, Pfizer, BioNTech, Veracyte (PROCURE), Diaceutics, Mindpeak, AstraZeneca, MEDAC, and Novartis. The institution of R.E. conducts research for AstraZeneca, Roche, Janssen-Cilag, NanoString Technologies, Biocartis, ZytoVision, Novartis, Cepheid, Mindpeak, MSD, Gilead, Palleos Healthcare, Owkin and BioNTech. E.D.P. and P.C. are current employees of and receive research funding from MultiplexDX. P.C. is an equity holder in MultiplexDX. BW reports research funding from Repare Therapeutics. SC discloses compensation for professional services to Boxer Capital, LLC, Eli Lilly and Company, Encore Medical Education, Genesis Therapeutics, Gerson Lehrman Group, Novartis, Nuvalent, Inc., Prelude Therapeutics, SAGA Diagnostics, and eFFECTOR Therapeutics. He also discloses equity and IP rights in Odyssey Biosciences and equity in Totus Medicines, Inc. H.W. is an advisor for AstraZeneca and Daiichi Sankyo. P.R. reports receiving funding from Grail, Illumina, Novartis, AstraZeneca, Epic Sciences, Invitae/ArcherDx, Biothernostics, Tempus, Inivata, Biovica, Guardant, Personalis, and Myriad. P.R. also is an advisor to Novartis, AstraZeneca, Pfizer, Lilly/Loxo, Prelude Therapeutics, Epic Sciences, Foundation Medicine, Inivata, Natera, Tempus, SAGA Diagnostics, Paige.ai, Guardant, and Myriad. P.R. is also a co-founder and board member of Odyssey Biosciences. LZB reports providing professional services to the Cancer Prevention & Research Institute of Texas. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Boehm, K.M., El Nahhas, O.S.M., Marra, A. et al. Multimodal histopathologic models stratify hormone receptor-positive early breast cancer. Nat Commun 16, 2106 (2025). https://doi.org/10.1038/s41467-025-57283-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57283-x
This article is cited by
-
Towards interpretable prediction of recurrence risk in breast cancer using pathology foundation models
npj Digital Medicine (2026)
-
Artificial intelligence in digital pathology diagnosis and analysis: technologies, challenges, and future prospects
Military Medical Research (2026)
-
Computational pathology in breast cancer: optimizing molecular prediction through task-oriented AI models
npj Breast Cancer (2025)
-
Artificial intelligence-driven pathomics in hepatocellular carcinoma: current developments, challenges and perspectives
Discover Oncology (2025)
-
Deep learning assessment of metastatic relapse risk from digitized breast cancer histological slides
Nature Communications (2025)