Abstract
The morphogenesis and cellular interactions in developing retina are incompletely characterized. The full understanding needs a precise mapping of the gene expression with a single-cell spatial resolution. Here, we present a spatial transcriptomic (ST) resource for the developing human retina at six developmental stages. Combining the spatial and single-cell transcriptomic data enables characterization of the cell-type-specific expression profiles at distinct anatomical regions at each developmental stage, highlighting the spatiotemporal dynamics of cellular composition during retinal development. All the ST spots are catalogued into consensus spatial domains, which are further associated to their specific expression signatures and biological functions associated with neuron and eye development. We prioritize a set of critical regulatory genes for the transitions of spatial domains during retinal development. Differentially expressed genes from different spatial domains are associated with distinct retinal diseases, indicating the biological relevance and clinical significance of the spatially defined gene expression. Finally, we reconstruct the spatial cellular communication networks, and highlight critical ligand-receptor interactions during retinal development. Overall, our study reports the spatiotemporal dynamics of gene expression and cellular profiles during retinal development, and provides a rich resource for the future studies on retinogenesis.
Similar content being viewed by others
Introduction
The development of the vertebrate retina is a highly coordinated process. During the later stage of gastrulation, the optic cup forms from the surface ectoderm1, followed by differentiation into neural retina in the inside and retinal pigment epithelium in the outside2,3. In the inner layer of the optic cup, retinal progenitor cells (RPCs) give rise to six major classes of neurons, including rod and cone photoreceptor cells (PCs), horizontal cells (HCs), bipolar cells (BCs), amacrine cells (ACs), and retinal ganglion cells (RGCs), as well as one class of glia, Müller glia (MG). Retinal cells form specific layer structures, which exhibit significant changes in their layer organization during embryonic development. After later stages of retinal development, retinal cells are organized into three distinct layers, each with their morphologic and functional characteristics4. Briefly, the cell bodies of PCs are located in the outer nuclear layer (ONL); the cell bodies of interneurons such as HCs, BCs, ACs and the glial cells, MG, are located in the inner nuclear layer (INL); and RGCs form the ganglion cell layer (GCL). The ONL and INL originate from the neuroblast layer (NBL) during the earlier stages of retinal development. Retinal vasculature, composing of pericytes and endothelial cells, is located in the periphery of this orchestrated layered structure. In addition to the precise spatial organization, the differentiation of retinal cells from RPCs is also arranged in temporal order. RGCs and HCs are generated relatively early, while PCs, ACs and BCs are generated relatively late5.
Given that developing retinal cells display distinguishing characteristics and high spatiotemporal dynamics that are associated with their functions, more specific transcriptome profiling of each retinal cell types during development needs to be investigated. Bulk RNA sequencing (RNA-seq) and single-cell RNA sequencing (scRNA-seq) studies on human retina and retinal organoids have offered rich resources6,7,8,9,10,11,12. These studies have delved depth into the post-mitotic transitional cell populations in the human embryonic retina, leading to the identification of three types of intermediate progenitor cells, i.e., neurogenic cells, amacrine-horizontal precursors (AC-HC Precurs), and bipolar-photoreceptor precursors (BC-Photo Precurs). These advancements have enriched our understanding on retinogenesis and the specification of retinal cell fate.
It is, however, not known to what extent the localization of each retinal cell type changes during development in human. Spatial transcriptomic (ST) technologies can provide high-throughput and spatially resolved analysis of gene expression, and have been applied on multiple embryonic tissue sections13,14,15. Combining scRNA-seq and ST methods can help overcome the limitations of individual techniques, facilitate unbiased cell type annotation and allow high-resolution spatiotemporal mapping of tissue development. More recently, scRNA-seq and ST technologies have been applied on retinal samples and organoids to explore the spatial and temporal dynamics of gene expression during retinal development16,17. In addition, a set of genes implicated in diverse retinal diseases have been identified18,19,20. The coverage of the developmental stages explored by existing studies is generally limited, particularly lacking samples before post-conception week (PCW) 14. However, the differentiation and maturation of ACs, cone and rod PCs from RPCs cannot be observed before PCW1417. Moreover, the developmental transitions between retinal cell types, as well as the spatiotemporal specificity of the retina-disease-associated genes during retinal development have not been well characterized. Furthermore, although the gene expression profiles of each retinal cell type during specific developmental stages have been described8,9,10, how these cell types communicate with each other in vivo is still not well studied in human.
In this study, we presented a spatiotemporal resource describing the spatial and cellular heterogeneity of the developing human retina at six developmental stages: PCW9, PCW12 early, PCW12 late, PCW14, PCW16 and PCW17. By integrating spatial and single-cell transcriptomic data, we revealed the spatiotemporal dynamics of retinal cell type composition during retinal development. We further identified diverse spatial domains of ST spots, and associated them with specific anatomical regions and biological functions. By analyzing the developmental changes of spatial domains, we prioritized a set of regulatory genes that were putatively critical for retinogenesis. Furthermore, we examined the localizations of genes associated with retinal diseases within the spatiotemporal maps. Finally, we established spatial cellular communication network for each developmental stage, highlighting the critical ligand-receptor interactions during retinal development. Overall, our study provided a rich resource for investigating spatiotemporal dynamics of gene expression during the early development of human retina.
Results
Spatially resolved transcriptomic profiling of developing human retina
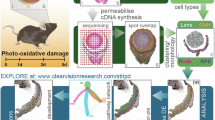
To comprehensively characterize the spatiotemporal gene expression during retinal development, we generated spatial expression profiles of human fetal retinal tissues from PCW9 to PCW17. For each developmental stage, we obtained one fresh frozen sample from the location near the optic nerve, followed by high-throughput sequencing run on the 10x Genomics Visium platform (Fig. 1a). A total of 1541 ST spots in the retinal region were retained with a median of 11,800 unique molecular identifiers (UMIs) and a mean of 4746 genes per spot (Supplementary Data 1). We then annotated the anatomical location of each spot in a radial and tangential system simultaneously to precisely and uniformly describe the spatial coordinates for each spot (Fig. 1b). Briefly, we initially identified NBL and GCL based on hematoxylin and eosin (H&E) staining4,9,21. Considering the actual thickness of the retinal samples (Supplementary Fig. 1a), as well as the relative thickness of the NBL and GCL (Supplementary Fig. 1b), we then divided the NBL and GCL into four (NBL 1/4, NBL 2/4, NBL 3/4, NBL 4/4) and two (GCL 1/2, GCL 2/2) equal-width sublayers, respectively (Supplementary Fig. 2a, b; see Methods). We also tangentially named the retinal regions with an angle deviating from the optic nerve (from −135˚ to 135˚) (Supplementary Fig. 2c). The well-known GCL marker genes (TUBB3 and SNCG) and NBL marker genes (SOX2 and SOX9)8,22 exhibited significantly higher expression in their corresponding sublayers (P < 3.02 × 10−26, one-sided Wilcoxon rank-sum test; Fig. 1c; Supplementary Fig. 3a), confirming the consistency between our annotations and the spatial context in the developing retina.
a The human embryonic retinal samples used in this study. b Schematic showing the data preprocessing framework. The 10x Genomics Visium spatial transcriptomes (i.e., gene expression profiles at spot level) and the corresponding anatomical regions of ST spots annotated based on H&E staining are integrated for downstream analysis. c Spatial visualization of the marker genes of NBL (SOX2 and SOX9) and GCL (TUBB3 and SNCG) at PCW9, PCW14 and PCW17. One section was obtained from each retinal sample. Scale bar, 200 μm. d UMAP plot of gene expression profiles using ST spots from all Visium slices. The spots are colored by radial sublayer annotations, as well as shown with varying sizes by tangential annotations.
We then explored the global similarity among all ST spots of retinal samples through a uniform manifold approximation and projection (UMAP) visualization23 (Fig. 1d; Supplementary Fig. 2d). In the UMAP, the ST spots from the same and/or adjacent sublayers across different developmental stages were clustered together, and the spot coordinates were aligned noticeably with their tangential angles, highlighting the pronounced spatial specificity of the developmental programs in human retina. We identified a set of differentially expressed genes (DEGs) for each of the six sublayers between neighboring developmental stages (FDR < 0.05; Supplementary Data 2). We found that the GCL showed more DEGs than the NBL in earlier stages (Supplementary Fig. 3b), confirming that the gene expression programs in developing human retina exhibited spatiotemporal asynchrony4. In addition, we confirmed that the DEGs of the sublayers from NBL and GCL were enriched in distinct biological functions (Supplementary Fig. 3c; Supplementary Data 3).
Spatiotemporal dynamics of cellular distribution during retinal development
Given that there are numerous individual cells within each ST spot, we aimed to examine how variation in the proportions of retinal cell types contributes to variation in spot expression. To this end, we estimated the relative proportions of various retinal cell types for each sample. First, we observed good concordance between our Visium spot-level expression and the single-cell expression during retinal development9 (Fig. 2a). We then deconvolved the spot-level expression matrix by using the single-cell signatures to estimate cell proportions across all samples (see Methods). As a validation, for each cell type, we calculated the Spearman’s Correlation Coefficient (SCC) between the estimated proportions and the expression levels of the corresponding cell type marker genes across all spots (Supplementary Data 4). The cell type marker genes were collected from previous studies on adult and embryonic retina, and retinal organoids8,10,11. As expected, 74.0% of the marker genes showed a significant correlation (SCC > 0, FDR < 0.05). In addition, we performed similar analyses and visualized the expression of cell type marker genes using another recently published retinal ST data17. We compared the results from both datasets, and confirmed the reproducibility of our results (Supplementary Fig. 4a and 4b). These results suggested that the relative proportions of the basic retinal cell types estimated from the single-cell signatures were biologically meaningful and reliable for the downstream analysis.
a Gene expression correlation between the Visium ST data from our study and the single-cell reference data from Lu et al.9. b Spatial distributions of various differentiated cell types during development. c Relative cell count of each retinal cell type during development. Left: differentiated cell types, including AC (amacrine cell), BC (bipolar cell), Cone, HC (horizontal cell), RGC (retinal ganglion cell) and Rod. Right: precursor cell types, including AC-HC Precur (amacrine-horizontal precursor), BC-Photo Precur (bipolar-photoreceptor precursor), neurogenic cell and RPC (retinal progenitor cell).
We then investigated the spatial distribution of the cell types at each developmental stage (Fig. 2b; Supplementary Fig. 5a). Across all developmental stages, RPCs dominated the entire NBL, while intermediate progenitor neurogenic cells dominated the inner (i.e., NBL 3/4 and NBL 4/4) sublayers of NBL. In contrast, RGCs generally dominated in the GCL. Cone and rod PCs preferably co-located with RPCs in the NBL, while ACs generally showed co-occurrence with RGCs in the GCL (Supplementary Fig. 5b). Furthermore, the locations of precursor cell types such as RPCs and neurogenic cells gradually changed from outer layers to inner layers during retinal differentiation, which consistent with previous studies24,25. In addition, we found pronounced differences in the distribution of various cell types along the tangential direction (Supplementary Fig. 5c). In the region near the ciliary body (i.e., >90°), there was a higher abundance of RPCs, while the cell counts of differentiated cell types were notably lower.
We further examined the relative abundances of the precursor cell types versus the differentiated cell types during retinal development. As expected, the differentiated cell types generally showed a steady increase in terms of their relative cell counts, while the precursor cell types showed a decrease (Fig. 2c; see Methods). Notably, we observed more RGCs in PCW9 than other differentiated cell types, suggesting an earlier specification window of RGCs than other differentiated cell types8,9. The estimated relative cell counts of cone and rod PCs, ACs and BCs had a significantly increase at PCW14 (Fig. 2c). Consistently, we observed a dramatic decrease in the relative cell count of AC-HC Precurs, the precursors of ACs and HCs, at PCW14.
In summary, we characterized the spatiotemporal distribution of the retinal cell types at each developmental stage, demonstrated the emergence of differentiated cell types during development, and found that the differentiation of the precursor cell types showed a direction from the outer layers to the inner layers, as well as from the posterior region to the anterior region.
Spatiotemporal distribution and cellular identity of consensus spatial domains
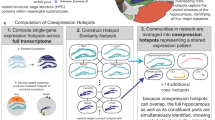
To leverage the spatial information from ST data, we aimed to identify the consensus spatial domains for all the spots. Briefly, we integrated gene expression profiles and spatial adjacency matrices for spatial domain detection (see Methods). We identified 9 spatial domains across all the developmental stages based on the spatially variable genes (SVGs) using the optimal approach (Fig. 3a; Supplementary Fig. 6; Supplementary Fig. 7a; Supplementary Data 5). Each spatial domain was further associated to specific embryonic cellular identity (Fig. 3b). As expected, there was an enrichment of multiple major cell types within the same spatial domain. For example, spatial domain D1 was enriched in RGCs and ACs, while spatial domain D6 was enriched in cone PCs, rod PCs and BC-Photo Precurs.
a UMAP plot of gene expression profiles using ST spots from all Visium slices. The spots are colored by spatial domain annotations (D1–D9). b Heatmap illustrating the enriched cell types associated with each spatial domain. AC amacrine cell, BC bipolar cell, HC horizontal cell, RGC retinal ganglion cell, RPC retinal progenitor cell, AC-HC Precur, amacrine-horizontal precursor, BC-Photo Precur bipolar-photoreceptor precursor. c Spatial distributions of each spatial domain, related to the radial sublayer (top) and absolute tangential angle (bottom). n = 1541 spots were examined. For box plots, boxes represent the interquartile range (25th to 75th percentiles), center lines indicate medians, and whiskers extend to the minima and maxima. d Visualization on the spatial distributions of spatial domains at PCW17. Scale bar, 200 μm. e The four gene modules (M1–M4) identified by WGCNA. For each gene module, the most enriched biological function, and the enrichment in each spatial domain are shown. n = 1541 spots were examined. For box plots, boxes represent the interquartile range (25–75th percentiles), center lines indicate medians, and whiskers extend to the minima and maxima. f Visualization on the spatial distributions of gene modules at PCW17. Scale bar, 200 μm.
We performed DE analysis for all the spatial domains, and identified a set of DEGs for each spatial domain (Supplementary Fig. 7b, c; Supplementary Data 6 and 7). We found that the spatial domains were in line with the cell type-specific marker genes identified through deconvolution. For example, NEFL, NEFM and GAP43 were up-regulated in spatial domains D1 and D2 (FDR < 5.93 × 10−20, log2 fold change >0.603), indicating the presence of RGCs; while RCVRN and NRL were up-regulated in spatial domain D5 (FDR < 4.47 × 10−29, log2 fold change >1.27), indicating the presence of PCs.
The anatomical regions of the spatial domains exhibited significant variations in terms of radial sublayers and tangential angles (Fig. 3c, d; Supplementary Fig. 7d). Within retinal layers, the distribution of various spatial domains was aligned with our prior results on the localizations of specific cell types, with spatial domains D5, D6, and D7 were enriched in photoreceptor cells located in the NBL, while spatial domains D1, D2, and D3 were enriched in RGCs and ACs in the GCL. Along the tangential direction, we identified three spatial domains with uneven distributions. Among these, spatial domain D9 was associated with the optic nerve, significantly enriched in the posterior region of the retina (P = 6.86 × 10−12, one-sided Wilcoxon rank-sum test; Fig. 3c), and was removed in subsequent analysis; spatial domains D4 and D8 were notably enriched in the anterior region of the developing retina (P = 4.88 × 10−109, one-sided Wilcoxon rank-sum test; Fig. 3c). We explored the expression of 24 shared DEGs between spatial domains D4 and D8 using scRNA-seq data from human embryonic retina9 and adult human ciliary body26 (Supplementary Fig. 8a; Supplementary Data 8), and found that it was significantly overexpressed in RPCs and non-pigmented epithelium (NPE) in ciliary body (P < 2.2 × 10−16, one-sided Wilcoxon rank-sum test; Supplementary Fig. 8b–h). These results indicated that the retinal stem cells in the adult ciliary body were associated with these two spatial domains (i.e., D4 and D8) in the anterior region26,27,28.
Spatiotemporal specificity of functional gene modules
We performed weighted gene co-expression network analysis (WGCNA)29 for all the ST spots across different developmental stages, resulting in four major genes modules (Supplementary Fig. 9a–c; Supplementary Data 9). The identified gene modules were enriched in distinct biological functions (Fig. 3e; Supplementary Fig. 9f; Supplementary Data 10): gene module M1 was related to axonogenesis; gene module M2 was related to extracellular matrix organization; gene module M3 was related to eye development and various biological regulations; while gene module M4 was related to photoreceptor cell development. Notably, the gene modules exhibited strong enrichment to specific spatial domains as described before (Fig. 3f; Supplementary Fig. 9d, e).
The identification of these gene modules can provide biological insights into the specific spatial regions in the developing retina. For example, gene module M1 was enriched in the spatial domains (i.e., D1, D2, D3 and D4) located in the GCL, indicating that the GCL could be critical in axonogenesis. As another example, gene module M3 was enriched in the spatial domain (i.e., D5) located in the inner sublayers of NBL (NBL 3/4 and NBL 4/4). The spatial domain D5 showed significantly higher proportion of neurogenic cells than in other spatial domains (P = 2.30 ×10−71, one-sided Wilcoxon rank-sum test). Taken together, these results indicated that the inner sublayers of NBL could be the critical region for the functionality of neurogenic cells in regulating retinal development.
We also explored the temporal dynamics of the gene modules during retinal development (Supplementary Fig. 9d). During the development of NBL, the dominant gene module(s) showed a replacement of M3 with M2, then followed by an enrichment of M3 and M4 (Fig. 3f). By contrast, during the development of GCL, the dominant gene module was consistently M1. These results indicated that the NBL exhibited more complex functional changes during retinal development.
Putative regulatory genes of spatial domains during retinal development
Although we have characterized the spatiotemporal localizations of the spatial domains, the putative critical genes regulating the transitions of spatial domains between developmental stages are still unclear. Previous studies have highlighted transcription factors (TFs) and RNA-binding proteins (RBPs) to play critical roles in the regulation of retinal development30,31,32,33,34. To identify putative critical regulatory genes for ST spatial domains, we systematically compared the DEGs of the spatial domains and the repository of regulatory genes (including TFs and RBPs) collected from Lambert et al.35 and Hentze et al.36 respectively. The DEGs of the spatial domains exhibited significant overlap with the TFs and RBPs (odds ratio >1.37, P < 4.12 ×10−5, two-sided Fisher’s exact test; Supplementary Fig. 10a). Notably, the genes HMGA1, JUN, REPIN1, ZNF207, ZNF385A, YBX1, YBX3, CXXC5, CHCHD3 and SAFB were identified in all three gene sets, indicating putative roles in both transcriptional and post-transcriptional regulation during retinal development.
These results inspired us to focus on the regulatory genes that may function at the spatial and temporal resolution. To this end, we identified a set of regulatory genes with elevated expression restricted in the same spatial domain between the neighboring developmental stages (Fig. 4a, b; Supplementary Data 11; see Methods). Interestingly, a great proportion (60.0% of total 215 genes) of these up-regulated regulatory genes during stage transition were RBPs, indicating the critical roles of RBPs and post-transcriptional regulation during retinal development. Although TFs are usually considered as the key regulatory factors for retinal development, the post-transcriptional regulation coordinated by RBPs can be also important and should not be overlooked. Notably, a few up-regulated regulatory genes were found as shared across majority of the spatial domains (7 out of 9 spatial domains), including NRL and HES5 among TFs, and LDHA, ZFP36L2, CRYAB, ENO1, and HNRNPDL among RBPs (Supplementary Fig. 10b, c), suggesting their potential broad and critical roles in retinal development.
a Up-regulated TFs and RBPs for each spatial domain between neighboring developmental stages. b Examples illustrating the expression dynamics of putative critical regulatory genes in each spatial domain during development. c Expression dynamics of growth factors that are related to retinal development in each spatial domain during development.
We also reconstructed the gene regulatory networks (GRNs) between TFs and their target genes (TGs) using expression data across all ST spots (see Methods). We highlighted several TFs with stronger interaction strength, such as ASCL1, ATF4, CREBZF and HBP1 (Supplementary Fig. 11a, b). interestingly, a considerable proportion (22.6%) of the TFs in the GRNs were also found in the set of up-regulated regulatory genes during stage transition (Supplementary Data 12), confirming their important regulatory functions for retinal development.
Next, we identified the enriched biological functions for these up-regulated regulatory genes, and found that the results were consistent with our previous findings from WGCNA (Supplementary Fig. 10d; Supplementary Data 13). For example, the enriched biological functions included eye and visual system development (e.g., NRL and SOX2), and regulation of neuron differentiation37,38 (e.g., NEUROD1 and MEIS1). We also identified a significant enrichment of genes associated with RNA splicing, including RBM3, RBM5 and RBM3939. Interestingly, in the up-regulated regulatory genes from spatial domains D2, D3, D4 and D5 during the transition from PCW16 to PCW17, we observed the genes implicated in glial cell fate commitment (including PAX6 and HES5), indicating the critical spatial regions and temporal windows for the differentiation of glial cell types, such as Müller cells40.
Previous studies have revealed the key roles of growth factors during the process of retinal development41,42,43,44,45. Here, we systematically explored the spatiotemporal expression dynamics of different growth factor families, including vascular endothelial growth factors (VEGFs), insulin-like growth factors (IGFs), fibroblast growth factors (FGFs), transforming growth factors (TGFs), neurotrophins, and epidermal growth factors (EGFs). Some growth factors exhibited dramatic changes of spatial and temporal distributions during retinal development, such as VEGFA, CCN2 (also known as CTGF) (Fig. 4c; Supplementary Fig. 12). We also validated the temporal dynamics of the growth factors using bulk RNA-seq data from retinal organoids46, and found that the expression trends of some critical growth factors (e.g., VEGFA, PGF, CCN2, and VGF) were consistent with the results from our data (Supplementary Fig. 13a, b). Among these, VEGFA and PGF have been implicated in retinal vascularization42,47, CCN2 has been reported to be functional in the formation of the retinal macula9, and VGF can affect neurogenesis48,49.
Biomedical relevance of spatial domains in developing retina
To bridge the gap between the spatiotemporal expression profiles of retinal development and retinal diseases, we then explored the expression patterns of disease-associated genes in spatial domains (Fig. 5a; Supplementary Data 14). Most of the gene sets associated with retinal diseases (9 out of 15 diseases) showed elevated expression in spatial domains D5 and D6 that were enriched in PCs, which have been implicated in the pathogenesis of ocular-retinal developmental disease (ODD), cone or cone-rod dystrophy (COD/CORD), retinitis pigmentosa (RP) and age-related macular degeneration (AMD)50,51,52. NDP, the only gene implicated in retinopathy of prematurity (ROP), showed higher expression in spatial domain D5, which were enriched in neurogenic cells. In contrast, the gene sets of optic atrophy (OA) and syndromic/systemic diseases with retinopathy (SDWR) were highly expressed in spatial domains D4 and D8. We found that the genes associated with retinal diseases were more significantly enriched for DEGs of spatial domains than putative critical regulatory genes (odds ratio = 2.04, P = 2.55 × 10−7, two-sided Fisher’s exact test; Fig. 5b). The genes CRX, NEUROD1, NRL, NR2F1, OTX2 and RAX2 were identified in all three gene sets.
a Clustering of retinal diseases based on the average expression levels of the disease-associated genes across different spatial domains. BBS Bardet-Biedl syndrome, CD chorioretinal atrophy or degeneration, COD/CORD cone or cone-rod dystrophy, CSNB congenital stationary night blindness, DA/DS deafness alone or syndromic, LCA Leber congenital amaurosis, MD macular degeneration, ODD ocular-retinal developmental disease, OA optic atrophy, RP retinitis pigmentosa, SDWR syndromic/systemic diseases with retinopathy, USH Usher syndrome, AMD age-related macular degeneration, ROP retinopathy of prematurity. b Overlap between the disease-associated genes, the putative regulatory genes for spatial domains, and the DEGs of spatial domains. Two-sided Fisher’s exact test. c Comparison of expression levels of genes involved with fatty acid and fat-soluble vitamin metabolism in retinal (n = 1541 spots) versus non-retinal (n = 14,700 spots) regions. ****P < 10−4; ns not significant, one-sided Wilcoxon rank-sum test. FAM fatty acid metabolism, FSVB fat-soluble vitamin biosynthetic, FSVC fat-soluble vitamin catabolic, VAM vitamin A metabolic, VDM vitamin D metabolic, VEM vitamin E metabolic, VKM vitamin K metabolic. For box plots, boxes represent the interquartile range (25th to 75th percentiles), center lines indicate medians, and whiskers extend to the minima and maxima. d Expression dynamics of the genes related to fatty acid and fat-soluble vitamin metabolism during development. e Visualization of spatial distribution for the genes associated with vitamin A metabolism across 6 retinal samples. Scale bar, 1 mm.
Bassen-Kornzweig syndrome (BKS) is a genetic disease characterized by RP, and is often treated with fat-soluble vitamin E in the clinical practice53,54. To evaluate the effects of fatty acid metabolism (FAM) and fat-soluble vitamins on retinal development, we investigated the spatiotemporal expression of gene sets related to them. Compared to non-retinal regions, the retinal regions showed significantly higher expression of the gene sets associated with FAM, fat-soluble vitamin catabolic process (FSVC), vitamin A metabolic process (VAM) and vitamin K metabolic process (VKM) (P < 1.77 × 10−11, two-sided Wilcoxon rank-sum test; Fig. 5c). In addition, we observed expression patterns with a rising peak from PCW12 late to PCW14 for these aforementioned gene sets (Fig. 5d). The spatial expression profiles of these gene sets revealed that they were enriched for specific regions during retinal development. For example, the genes related to VAM were located in the outer sublayers of NBL (i.e., NBL 1/4 and NBL 2/4) from PCW12 late to PCW16, indicating putative associations with PCs and their precursor cells (i.e., spatial domains D5, D6 and D7) (Fig. 5e; Supplementary Fig. 14).
Spatial cellular communications reveal the key ligand-receptor interactions
Given the significance of cell-cell communications in the functionality of retina55,56, a more detailed characterization of cellular communication networks base on ST data is in an urgent need. To this end, we reconstructed spatial cellular communication networks using COMMOT57 for each ST sample (see Methods). We visualized the spatial signaling directions of all ligand-receptor (L-R) interactions for each developmental stage (Fig. 6a). Interestingly, we discovered that the spatial signals generally propagated between different retinal layers in all developmental stages, moving from the both outer sides (i.e., NBL and GCL) towards the inner layers, particularly enriched in the region where spatial domain D5 was located. We then established domain-domain communication networks for each sample, and found that spatial domains D1, D3, and D5 were implicated in spatial cellular communication processes across all developmental stages (Supplementary Fig. 15a).
a Spatial signaling directions in each developmental stage. b SCSs for each L-R interaction across all samples. The L-R interactions are colored by interaction types including cell-cell contact and secreted signaling. c Visualization of spatial distribution for the ligand gene MDK and the receptor gene LRP1. d The immunohistochemistry experiments showing the co-occurrence of the ligand gene MDK and the receptor gene LRP1 during development. The retinal sections were obtained from three samples at PCW12, PCW14 and PCW16. Scale bar, 100μm.
We then identified the critical L-R interactions from the spatial cellular communications by summarizing the spatial communication scores (SCSs) for each L-R interaction across all samples (Fig. 6b; Supplementary Data 15). Among these key L-R interactions, those associated with secreted signaling accounted for 85.7%, while those related to cell-cell contact comprised only 14.3%. To link these key ligand-receptor interactions in spatial cellular communications with their cell types, we supplemented our analysis with cell-cell communication results from single-cell data9 (Supplementary Fig. 15b; Supplementary Data 16). We found that L-R interactions associated with cell-cell contact (i.e., the Notch signaling pathway58,59) were extensively involved in cell-cell communications between four precursor cell types (i.e., RPC, AC-HC Precur, BC-Photo Precur and neurogenic cell) and other cell types, while L-R interactions related to secreted signaling participated in cell-cell communications between RGCs, ACs and other cell types. For example, we observed widespread expression of the ligand gene CNTN1 and the receptor gene NOTCH1 in the Notch signaling pathway (Supplementary Fig. 15c), as well as the ligand gene PTN and the receptor gene PTPRS in the ECM-receptor interaction pathway60,61 (Supplementary Fig. 15d).
In addition, we highlighted cell-cell communications that have not been fully characterized in the developing human retina. Previous studies have revealed the spatiotemporal expression patterns of ligand gene MDK during retinal development in mouse and chicken62,63. We found that MDK-LRP1 exhibited the highest SCS, indicating its extensive involvement in spatial cellular communication during retinal development (Fig. 6b). We confirmed the co-occurrence of MDK and LRP1 in ST data (Fig. 6c), which can be experimentally validated (Fig. 6d). Furthermore, we discovered that the MDK-LRP1 interaction was involved in the cell-cell communications from four precursor cell types to RGCs (Supplementary Fig. 15b), indicating its potential key roles in regulating RGC maturation.
Discussion
Here, we show a spatial transcriptomic resource for the developing human retina, highlighting the spatiotemporal dynamics of the cellular organization and cell-cell communications during retinal development. Our study commenced with constructing a radial and tangential system for the developing retina, precisely locating ST spot in the spatial navigation system. Subsequently, we analyzed the spatiotemporal dynamics of cellular distribution, revealing the cell differentiation direction during retinal development. We identified a set of spatial domains, and explored the enriched biological functions and critical regulatory genes implicated in the spatial domains. Lastly, our study reconstructed the cell-cell communication networks during retinal development, and uncovered key ligand-receptor interactions.
Our results reveal the spatiotemporal expression patterns in the developing human retina. Given the observation showing that the ACs, BCs, and PCs exhibited elevated cellular proportions, we hypothesized that this period could be critical for retinal cell differentiation. We confirmed that RGCs appeared at PCW9, earlier than other differentiated cell types8,9. Our study revealed that RPCs were located in the NBL, while intermediate progenitor neurogenic cells were restricted in the inner sublayers of NBL. We also observed that cone and rod PCs preferably co-located with RPCs in the NBL, while ACs generally showed co-occurrence with RGCs in the GCL. Based on these results, we hypothesized that there would be a differentiation direction of the precursor cell types from the outer layers to the inner layers, as well as from the posterior region to the anterior region of the developing retina, consistent with previous studies4,10. Although we observed that rod and cone PCs co-localized with RPCs, the rod and cone PCs is more related to neurogenic cells in terms of cellular lineage9,30 (Supplementary Fig. 8c). We observed that the neurogenic cells located in the inner side of RPCs during development (Supplementary Fig. 5a, b). Taken together, these results indicate that the differentiation of some retinal cell types within the NBL undergo a reverse and inside-out process.
Furthermore, we found that the two spatial domains D4 and D8 we identified showed co-localization with the ciliary marginal zone (CMZ), which is always regarded to be associated with retinal stem cells64,65. Notably, the spatiotemporal distributions of D4 and D8 were distinct from other spatial domains. In addition, we defined a set of DEGs shared by D4 and D8, and found that these DEGs were highly expressed in the NPE of the adult ciliary body (Supplementary Fig. 8f–h). Taken together, our results provide evidence on an embryonic origin of potential retinal stem cells within the ciliary body. We found that D4 and D8 exhibited an elevation of FGF2 expression during early embryonic development (Fig. 4c). Interestingly, FGF2 has been used in the reprogramming of ciliary epithelium into other retinal neurons66. As expected, the overall expression of FGF2 showed dramatic decrease during retinal development (Fig. 4c), consistent with the observation from retinal organoids46 (Supplementary Fig. 13b).
We also identified four functional gene modules (M1–M4) of the developing human retina by WCGNA (Supplementary Data 9). Notably, there were significant shifts in the gene modules enriched in the NBL, showing a replacement of M3 with M2, and then followed by an enrichment of M3 and M4. During this process, M2 can be considered as the intermediate state with extracellular matrix (ECM) organization as its most enriched biological function (Fig. 3e). The ECM plays a critical role in regulating retinal cell polarity and tissue architecture67, and its synthesis is known to be modulated by vitamin A68. Our findings revealed that the expression level of the genes related to vitamin A metabolism was higher in the retinal compared to non-retinal regions (Fig. 5c). The genes related to vitamin A metabolism exhibited co-localization with M2 (Fig. 5e; Supplementary Fig. 9d). These results suggest that ECM-involved biological processes are implicated in regulating vitamin A synthesis and metabolism, and provide evidence on the critical time window for supplementation of fat-soluble vitamins during pregnancy.
TFs have been known for their pivotal roles in controlling temporal patterning, neurogenesis, and cell-fate specification during mammalian retinal development30,69. In contrast, the regulatory mechanism of RBPs in retinal development remains unclear. In our study, we identified a set of upregulated regulatory genes, including both TFs and RBPs, between neighboring developmental stages. Interestingly, we found that there were more up-regulated RBPs (129 genes) than TFs (91 genes), indicating their critical roles in retinal development. In particular, most of the up-regulated regulatory genes were RBPs before the stage of PCW12 late, while after PCW12 late, the number of TFs increased (Fig. 4a). Previous studies have revealed the critical roles of RBPs in regulating brain development and pathogenesis70,71,72. The functionality of RBPs in retinal development still needs further investigations.
A recent study has disclosed the importance of cell-cell communications associated with MDK in retinal development8. We noticed that MDK-LRP1 interaction obtained the highest SCS in ST data (Fig. 6b), and was predicted to present between the source cell types such as neurogenic cells, RPCs, BC-Photo Precurs and AC-HC Precurs and the target cell type RGCs (Supplementary Fig. 15b). Given that the MDK-LRP1 interaction is involved in the biological processes such as cell growth, survival and division73,74, it might play key roles in regulating RGC differentiation and maturation.
Overall, we have generated a spatiotemporal transcriptome resource of the human embryonic retina across six developmental stages. By leveraging this resource, we unveiled the spatiotemporal dynamics of the cellular compositions, identified the consensus spatial domains, prioritized the putative critical regulatory genes, and reconstructed spatial cellular communication networks during human retinal development. Additionally, we report the spatiotemporal distribution of the genes implicated in various retinal diseases, which could be investigated by further studies.
Due to the difficulties of obtaining embryonic retinal samples across multiple developmental stages, our study cannot include biological replicates for the samples in each stage, which may hinder the capability of validating the spatiotemporal dynamics of cellular distributions. Recently, more transcriptomic datasets from human retina have been publicly available, such as scRNA-seq datasets from embryonic retinal samples9, scRNA-seq datasets from adult ciliary body samples26, ST datasets from embryonic retinal samples17, and bulk RNA-seq datasets from retinal organoids46, which could be valuable to be incorporated in our study as external validation. We compared the results from our ST data with the spatiotemporal patterns of the cellular distributions and gene expression from these studies, confirming the reliability of our discoveries. With the acquisition of more embryonic retinal samples and maturity of ST technologies with improved resolution, more high-quality ST data of developing human retina will be included in future studies to minimize the potential bias from the data.
We anticipate that multiple future directions can be explored based on our work. First, although the Visium platform enables high-throughput characterization of spatially resolved gene expression, the resulting data lack real single-cell resolution when applied to the developing human retina. This hinders our ability to precisely localize the expression of cell-type-specific marker genes within retinal tissues and detect the spatial distribution of retinal cell types in high resolution. In this study, the NBL and GCL with only three or four spots in thick were divided into 6 sublayers based on the radial location of the spot centroids. This may lead to ambiguity in relating the gene expression of spots to their anatomical locations. Therefore, future advancements in ST technologies with enhanced sensitivity and resolution will allow us to gain a more detail and accurate view of the spatiotemporal dynamics at different cell types during retinal development. The size of the Visium slices in the study constrains the inclusion of retinal samples from later developmental stages. Even though the majority of the early developmental stages (i.e., from PCW9 to PCW17) have been covered in the study, optimizing the experimental designs, especially utilizing larger slice size, will enable a more comprehensive spatiotemporal map of the developing human retina. Finally, while we characterized the spatiotemporal dynamics of gene expression and cellular composition during retinal development, we lack a clear picture of the spatially resolved epigenetic regulation, which can be achieved by incorporating single-cell ATAC-seq data. Putting together multi-omics data of developing human retina in the context of spatial organization will be valuable for understanding the regulatory mechanisms underlying the complex process of retinal development.
Methods
Human tissue samples
All tissue samples used in this study were obtained from Chinese individuals and were sourced from Obstetrics and Gynecology Hospital of Fudan University. The participants were pregnant women with singleton pregnancies and had no known fetal abnormalities prior to abortion. Gestational age was measured in weeks from the first day of the woman’s last menstrual period and ensured by early pregnancy ultrasound. The collection of human embryos was reviewed and approved by Human Subject Research Ethics Committee of Obstetrics and Gynecology Hospital of Fudan University (2022-92). All participating women provided informed consent for scientific research.
Tissue processing and spatial transcriptome
Tissues were gently washed with cold PBS, then embedded in OCT-filled molds and snap frozen in chilled isopentane. Sagittal microtome sections of the retinal tissues near the center of the lens were cut at a thickness of 10 μm (Leica, cm1950). These sections were placed onto chilled 10x Genomics Visium Tissue Optimization Slides (catalog no. 3000394, 10x Genomics) and Visium Spatial Gene Expression Slides (catalog no. 2000233, 10x Genomics), and firmly adhered by warming the back of the slide. Tissues were permeabilized for 12–18 min, which was determined through tissue optimization time-course experiments.
Tissue sections were fixed in chilled methanol and stained following the guidelines outlined in the Visium Spatial Gene Expression User Guide (catalog no. CG000239 Rev A, 10x Genomics). Brightfield histology images were captured using a 20x objective on Pannoramic MIDI scanner (3D HISTECH). The raw images were then stitched together using Pannoramic MIDI and exported as.tiff files with both low- and high-resolution settings.
Libraries were generated following the Visium Spatial Gene Expression User Guide (catalog no. CG000239, 10x Genomics) and sequenced on a NovaSeq 6000 System (Illumina) using a NovaSeq S4 Reagent Kit (200 cycles, catalog no. 20027466, Illumina), with a sequencing depth of approximately 3.00 ~ 3.37 × 108 read-pairs per sample.
Image registration and anatomical annotation
Image registration was performed manually slice by slice. The boundaries of the retinal regions were determined based on H&E staining images. Briefly, the outer and inner boundaries were identified by black (representing the retinal pigment epithelium layer) and white (representing the vitreous), respectively. Additionally, the boundaries between the NBL and the GCL were identified based on color depth (transitioning from dark blue to light blue) and gene expression.
Subsequently, a quantitative representation of the within-layer locations and tangential angles of each spot was obtained through linear interpolation. We considered the structure and thickness of the retina when dividing the NBL and GCL into sublayers. Briefly, we first estimated the thickness of the developing retina, with a median of 186 μm (Supplementary Fig. 1a), as well as the ratio of NBL thickness to GCL thickness during development (Supplementary Fig. 1b). Considering the change of the ratio over time, we used the relative thickness instead of the absolute thickness when dividing the sublayers. In addition, given that the NBL will differentiate into INL and ONL during the later stages, it would be necessary to divide the NBL into multiple sublayers for augmented resolution. Taken together, we divided the NBL and GCL into four (i.e., NBL 1/4, NBL 2/4, NBL 3/4, NBL 4/4) and two (i.e., GCL 1/2, GCL 2/2) equally-sized sublayers from the outside to the inside, respectively (Supplementary Fig. 2a).
Preprocessing of the 10x Genomics Visium data
The raw FASTQ data and histology images were processed into count matrices using Space Ranger (version 2.1.1), which used STAR75 (version 2.7.2a) for read alignment. The read alignment was based on the human reference genome GRCh38. Sequencing file details were available in Supplementary Data 1.
We excluded poor-quality data in the raw count matrices using Seurat76 (version 4.3.0). The spots with a total read count less than 1000 were considered non-tissue or vitreous regions, and were excluded from the downstream analysis. The spots with expression count from mitochondrial genes exceeding 10% of the total counts were also discarded. The genes with expression in less than 10 spots, as well as the ribosomal and mitochondrial genes were also removed from the downstream analysis.
Processing of scRNA-seq data from human embryonic retina
We obtained publicly available scRNA-seq data of human embryonic retinas from Lu et al.9 To ensure comparability with our samples, only retinal samples from GW9 to GW17 were used in our analysis. The cells with a total read count less than 1000 or a number of expressed gene less than 300 were excluded from the downstream analysis. We also filtered out the cells with expression counts from mitochondrial genes exceeding 10% of the total counts. The ribosomal and mitochondrial genes were discarded from the downstream analysis. All retinal cell type annotations were obtained from Lu et al.9 Müller cells were excluded from the analysis due to there were only 4 Müller cells in the samples used.
Processing of scRNA-seq data from human adult ciliary body
We obtained the scRNA-seq data of human adult ciliary body from Lou et al.26 The cells with a total read count less than 500 or a number of expressed gene less than 300 were excluded from the downstream analysis. We also excluded the cells with expression counts from mitochondrial genes exceeding 40% of the total counts. We then processed the data using the Seurat76 package, including log-normalization with a scaling factor of 10,000, selecting the top 2000 highly variable genes, gene-level scaling and principal component analysis (PCA). Next, we applied the Louvain method77 for cell type clustering with a resolution of 0.5. We then annotated the cell types of human adult ciliary body based on the known marker genes as described in Lou et al.26 (Supplementary Fig. 8f). The full list of marker genes used in our analysis can be found in Supplementary Data 8.
Deconvolution of spatial transcriptome data using scRNA-seq data
To estimate the cell proportions of each cell type from human embryonic retina, we decomposed the spatial transcriptome data from each developmental stage using the scRNA-seq data from Lu et al.9 We implemented the Cell2location78 package (version 0.1.0) with the settings “N_cells_per_location=20” and “detection_alpha=20”. The deconvolution results were normalized to a summation of 1 for each spot.
We then corrected for differences in cell abundance between spots using H&E staining images to obtain relative cell counts. Specifically, we enhanced the images using the CLAHE algorithm79 and performed stain normalization using the Macenko algorithm80. Next, we employed OpenCV (version 4.9.0) for adaptive thresholding to obtain binary images of cell nucleus. By multiplying the density of nuclear pixels within each spot, we transformed the cell type proportions into relative cell counts.
Dimension reduction and spatial domain detection of ST spots
We first employed the SPARK-X algorithm81 (version 1.1.1) to identify spatially variable genes (SVGs) sample by sample, with the parameter set to “option=mixture”. We applied Bonferroni correction and retained genes with adjusted P less than 0.05 as SVGs.
We then performed processing of the Visium data using the Seurat76 package. The expression counts were log-normalized with a scaling factor of 10,000. SVGs from all stages were used for subsequent analysis. Gene-level scaling and PCA were performed, followed by Harmony82 for data integration. We then applied the uniform manifold approximation and projection (UMAP)23 for visualization.
To identify spatial domains, we took the union of the k-nearest neighbor graph based on Harmony-embedding and the spatial neighbor graph based on spot locations, creating a fused graph. We then identified the spatial domains based on the fused graph using the graph-based Louvain method77, in which we tried a series of the resolutions ranging from 0.5 to 1.5 with a step of 0.1.
In addition, we compared the single-cell data integration results from Harmony with other similar methods including Seurat-CCA76 (version 4.3.0) and scVI83 (version 1.1.5), as well as the methods specified to ST data including BASS84 (version 1.1.0) and GraphST85 (version 1.1.1). Both Seurat-CCA and scVI were used to construct fused graphs in a way similar to Harmony. The performance of Harmony and the other four methods was evaluated in several ways:
-
1.
UMAP visualization (Supplementary Fig. 6a), which was used to assess how the spatial domains were separated.
-
2.
iLISI metric82 (Supplementary Fig. 6b), which can evaluate the data integration quality across different samples.
-
3.
Classification accuracy (Supplementary Fig. 6c), which was used to evaluate the classification robustness of the spatial domains. Briefly, we used the same ten-fold cross-validation to estimate the classification accuracy of the spatial domains using different classifiers including Logistic Regression, Random Forest, and Support Vector Machines.
To determine the optimal resolution, we calculated the minimum number of DEGs for each spatial domain at every resolution, and then identified the point where this number stabilized (at a resolution of 1.1) as the optimal resolution.
Weighted gene co-expression network analysis (WGCNA)
Signed co-expression networks in developing retina were built using the R package WGCNA29 (version 1.71) across different developmental stages. The genes with zero expression in more than 80% of the spots were removed in this analysis. For all genes included, a pairwise correlation matrix was computed, and an adjacency matrix was calculated by raising the correlation matrix to a power. The power was set to 7. We then implemented the blockwiseModules function in the WGCNA29 package with the following settings: “networkType=signed”, “minModuleSize=20”, “corType=bicor”, and “maxPOutliers=0.1”. Modules were defined as branches of a hierarchical cluster tree using the dynamic tree cut method86. Each module was summarized by an eigengene (i.e., the first principal component of the scaled module expression), which explained the maximum amount of variation of the module expression levels.
Differential expression analysis and gene ontology enrichment analysis
In this article, we identified DEGs using the FindAllMarkers function in the Seurat76 package with the parameters “test.use=wilcox”, “min.pct=0.1” and “logfc.threshold=0.25” and then filtered the genes with a FDR threshold of 0.05. We employed clusterProfiler87 (version 4.10.0) to identify enriched gene ontology terms (biological process) for each group of DEGs.
Identification of putative regulatory genes
We compiled a set of 1639 TFs from Lambert et al.35 and 1393 RBPs from Hentze et al.36 and then retained these genes with expression in at least 10% of spots for downstream analysis. These genes were categorized into three groups: “TF” annotated as only transcription factors, “RBP” annotated as only RNA-binding proteins, and “TF & RBP” annotated as both. The spatial domain D9 was not included in the analysis due to its limited sample size. We then performed differential expression analysis for the spots belonging to the same spatial domain between each pair of neighboring stages. We applied Bonferroni correction, and retained the genes with an adjusted P < 0.05 as up-regulated regulatory genes.
Gene regulatory network inference
We utilized SCRIPro88 (version 1.1.30) to calculate the interaction strength between transcription factors (TFs) and the target genes (TGs) based on ST data of all spots. First, we set the parameter “Cell_num=3” when constructing the supercells. We then implemented the SCRIPro_RNA function with parameter “assays = [‘Direct’,‘DNase’,‘H3K27ac’]”, followed by predicting the chromatin landscapes of the supercells from the bulk epigenome reference using LISA89. Next, we assessed the effects and significances of expression response of the marker genes in a specific supercell to the perturbation of transcriptional regulators. We then selected the TFs with P < 0.05 in at least one supercell. The TF-TG interactions were predicted using the get_tf_target function, where for each TF-TG interaction, the maximum value across all supercells was selected to represent the strength of that interaction. Finally, 164 TFs with at least one interacted TG were retained.
Expression analysis of the clinically relevant gene sets
We calculated and visualized the average expression levels of a set of clinically relevant gene sets within each spatial domain during retinal development. The disease-associated gene sets were obtained from RetNet (https://web.sph.uth.edu/RetNet), a website providing 291 genes associated with 15 retinal diseases (Supplementary Data 14). The gene sets related to fatty acid and fat-soluble vitamin metabolism were obtained from the MSigDB database90, in which we selected the gene sets associated with FAM (fatty acid metabolism), FSVB (fat-soluble vitamin biosynthetic), FSVC (fat-soluble vitamin catabolic), VAM (vitamin A metabolic), VDM (vitamin D metabolic), VEM (vitamin E metabolic), and VKM (vitamin K metabolic) using their entry name “KEGG_FATTY_ACID_METABOLISM”, “GOBP_FAT_SOLUBLE_VITAMIN_BIOSYNTHETIC_PROCESS”, “GOBP_FAT_SOLUBLE_VITAMIN_CATABOLIC_PROCESS”, “GOBP_VITAMIN_A_METABOLIC_PROCESS”, “GOBP_VITAMIN_D_METABOLIC_PROCESS”,
“GOBP_VITAMIN_E_METABOLIC_PROCESS”, and “GOBP_VITAMIN_K_METABOLIC_PROCESS”, respectively.
Inference of spatial cellular communication networks
We utilized COMMOT57 (version 0.0.3) to compute the spatial cellular communication networks, and used the CellPhoneDB91 (version 4.0.0) as the ligand-receptor interaction reference database. Ambiguous ligand-receptor interactions within the database were manually removed. Each ST sample was independently inputted to calculate spatial cellular communication, and we reconstructed the domain-domain communication networks with the parameters “n_permutations=1000” and “pathway_name=None”.
To associate spatial cellular communications with cell types, we inferred cell-cell communications based on human embryonic retinal scRNA-seq data9 as well. The communication pair of each source cell type and target cell type can be quantitatively estimated based on the co-expression levels of the ligand gene and the receptor gene from the source and target cell type, respectively.
Immunohistochemistry of retinal tissues
Frozen human fetal retinal tissue samples were cut to 20 μm sections and collected on ColorCoat microscope slides (Citotest Scientific) using a Leica CM1950 cryostat. Before immunofluorescence, the samples on slides were fixed with prechilled anhydrous methanol for about 1 minute, followed by washing 3 times with PBS. The samples were then permeabilized and blocked in blocking buffer (1% bovine serum albumin, 5% goat serum and 0.3% triton-100 in PBS at pH 7.4) for 2 h, and incubated in primary antibodies diluted in blocking buffer overnight at 4 °C. The primary antibodies used were rabbit anti-LRP1 (1:200, ab92544, Abcam), mouse anti-MDK (1:50, sc-46701, Santa Cruz). On the next day, the samples were washed three times in PBS, followed by incubation in secondary antibodies for 2 h at room temperature. The secondary antibodies used were goat anti-rabbit 594 (1:500, ab150080, Abcam) and goat anti-mouse 488 (1:500, ab150113, Abcam). Cell nuclei were stained with DAPI (1:2000, Yeasen), and mounted. Images were acquired on a Nikon AX R Confocal microscope.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The spatial transcriptomic data of the human embryonic retina samples generated in this study has been deposited at the Genome Sequence Archive for Human (GSA-Human) under the accession number HRA006282. The publicly available scRNA-seq data of the human embryonic retina and adult ciliary body can be accessed under the accession number GSE138002 and GSE206026, respectively. Source data are provided with this paper.
Code availability
The codes used in this study were provided in GitHub via https://github.com/ZhaoXM-Lab/DevRetinaST92.
References
Cvekl, A. & Zhang, X. Signaling and Gene Regulatory Networks in Mammalian Lens Development. Trends Genet 33, 677–702 (2017).
Gregory-Evans, C. Y., Wallace, V. A. & Gregory-Evans, K. Gene networks: dissecting pathways in retinal development and disease. Prog. Retin Eye Res. 33, 40–66 (2013).
Fuhrmann, S. Eye morphogenesis and patterning of the optic vesicle. Curr. Top. Dev. Biol. 93, 61–84 (2010).
Hoshino, A. et al. Molecular Anatomy of the Developing Human Retina. Dev. Cell 43, 763–779.e764 (2017).
Reese, B. E. Development of the retina and optic pathway. Vis. Res. 51, 613–632 (2011).
Whitmore, S. S. et al. Transcriptomic analysis across nasal, temporal, and macular regions of human neural retina and RPE/choroid by RNA-Seq. Exp. Eye Res. 129, 93–106 (2014).
Lukowski, S. W. et al. A single-cell transcriptome atlas of the adult human retina. EMBO J. 38, e100811 (2019).
Hu, Y. et al. Dissecting the transcriptome landscape of the human fetal neural retina and retinal pigment epithelium by single-cell RNA-seq analysis. PLoS Biol. 17, e3000365 (2019).
Lu, Y. et al. Single-Cell Analysis of Human Retina Identifies Evolutionarily Conserved and Species-Specific Mechanisms Controlling Development. Dev. Cell 53, 473–491.e479 (2020).
Sridhar, A. et al. Single-Cell Transcriptomic Comparison of Human Fetal Retina, hPSC-Derived Retinal Organoids, and Long-Term Retinal Cultures. Cell Rep. 30, 1644–1659.e1644 (2020).
Cowan, C. S. et al. Cell Types of the Human Retina and Its Organoids at Single-Cell Resolution. Cell 182, 1623–1640.e1634 (2020).
Liang, Q. et al. Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling. Nat. Commun. 10, 5743 (2019).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792.e1721 (2022).
Moffitt, J. R., Lundberg, E. & Heyn, H. The emerging landscape of spatial profiling technologies. Nat. Rev. Genet 23, 741–759 (2022).
Chen, T. Y., You, L., Hardillo, J. A. U. & Chien, M. P. Spatial Transcriptomic Technologies. Cells 12, 2042 (2023).
Wahle, P. et al. Multimodal spatiotemporal phenotyping of human retinal organoid development. Nat. Biotechnol. 41, 1765–1775 (2023).
Dorgau, B. et al. Single-cell analyses reveal transient retinal progenitor cells in the ciliary margin of developing human retina. Nat. Commun. 15, 3567 (2024).
Schneider, N. et al. Inherited retinal diseases: Linking genes, disease-causing variants, and relevant therapeutic modalities. Prog. Retin Eye Res. 89, 101029 (2022).
Van Schil, K. et al. Mapping the genomic landscape of inherited retinal disease genes prioritizes genes prone to coding and noncoding copy-number variations. Genet Med. 20, 202–213 (2018).
Menon, M. et al. Single-cell transcriptomic atlas of the human retina identifies cell types associated with age-related macular degeneration. Nat. Commun. 10, 4902 (2019).
Aparicio, J. G. et al. Temporal expression of CD184(CXCR4) and CD171(L1CAM) identifies distinct early developmental stages of human retinal ganglion cells in embryonic stem cell derived retina. Exp. Eye Res. 154, 177–189 (2017).
Quinn, P. M. et al. Human iPSC-Derived Retinas Recapitulate the Fetal CRB1 CRB2 Complex Formation and Demonstrate that Photoreceptors and Müller Glia Are Targets of AAV5. Stem Cell Rep. 12, 906–919 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Quinn, P. M. J. & Wijnholds, J. Retinogenesis of the Human Fetal Retina: An Apical Polarity Perspective. Genes 10, 987 (2019).
Tian, N. Development of Retinal Ganglion Cell Dendritic Structure and Synaptic Connections. In: Webvision: The Organization of the Retina and Visual System (eds Kolb, H., Fernandez, E., Nelson, R.) (University of Utah Health Sciences Center, 1995).
Lou, B. et al. A single-cell transcriptomic atlas of the human ciliary body. Cell Mol. Life Sci. 79, 528 (2022).
Gautam, P. et al. Multi-species single-cell transcriptomic analysis of ocular compartment regulons. Nat. Commun. 12, 5675 (2021).
Frøen, R. et al. Does the adult human ciliary body epithelium contain “true” retinal stem cells? Biomed. Res. Int. 2013, 531579 (2013).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Lyu, P. et al. Gene regulatory networks controlling temporal patterning, neurogenesis, and cell-fate specification in mammalian retina. Cell Rep. 37, 109994 (2021).
Santos-França, P. L., David, L. A., Kassem, F., Meng, X. Q. & Cayouette, M. Time to see: How temporal identity factors specify the developing mammalian retina. Semin Cell Dev. Biol. 142, 36–42 (2023).
Freund, C., Horsford, D. J. & McInnes, R. R. Transcription factor genes and the developing eye: a genetic perspective. Hum. Mol. Genet 5, 1471–1488 (1996).
Dash, S., Siddam, A. D., Barnum, C. E., Janga, S. C. & Lachke, S. A. RNA-binding proteins in eye development and disease: implication of conserved RNA granule components. Wiley Interdiscip. Rev. RNA 7, 527–557 (2016).
Rodriguez, A. R., de Sevilla Müller, L. P. & Brecha, N. C. The RNA binding protein RBPMS is a selective marker of ganglion cells in the mammalian retina. J. Comp. Neurol. 522, 1411–1443 (2014).
Lambert, S. A. et al. The Human Transcription Factors. Cell 172, 650–665 (2018).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Pataskar, A. et al. NeuroD1 reprograms chromatin and transcription factor landscapes to induce the neuronal program. EMBO J. 35, 24–45 (2016).
Islam, M. M., Li, Y., Luo, H., Xiang, M. & Cai, L. Meis1 regulates Foxn4 expression during retinal progenitor cell differentiation. Biol. Open 2, 1125–1136 (2013).
Sutherland, L. C., Rintala-Maki, N. D., White, R. D. & Morin, C. D. RNA binding motif (RBM) proteins: a novel family of apoptosis modulators? J. Cell Biochem. 94, 5–24 (2005).
Roesch, K. et al. The transcriptome of retinal Müller glial cells. J. Comp. Neurol. 509, 225–238 (2008).
Vrensen, WitmerA. N., Van Noorden, G. F. & Schlingemann, C. J. RO. Vascular endothelial growth factors and angiogenesis in eye disease. Prog. Retin Eye Res. 22, 1–29 (2003).
Balser, C., Wolf, A., Herb, M. & Langmann, T. Co-inhibition of PGF and VEGF blocks their expression in mononuclear phagocytes and limits neovascularization and leakage in the murine retina. J. Neuroinflammation 16, 26 (2019).
Grant, M. B. et al. Insulin-like growth factor I acts as an angiogenic agent in rabbit cornea and retina: comparative studies with basic fibroblast growth factor. Diabetologia 36, 282–291 (1993).
Wan, J., Zhao, X. F., Vojtek, A. & Goldman, D. Retinal injury, growth factors, and cytokines converge on β-catenin and pStat3 signaling to stimulate retina regeneration. Cell Rep. 9, 285–297 (2014).
Moon, S. et al. A CTGF-YAP Regulatory Pathway Is Essential for Angiogenesis and Barriergenesis in the Retina. iScience 23, 101184 (2020).
Qu, Z., Batz, Z., Singh, N., Marchal, C. & Swaroop, A. Stage-specific dynamic reorganization of genome topology shapes transcriptional neighborhoods in developing human retinal organoids. Cell Rep. 42, 113543 (2023).
Penn, J. S. et al. Vascular endothelial growth factor in eye disease. Prog. Retin Eye Res. 27, 331–371 (2008).
Lewis, J. E., Brameld, J. M. & Jethwa, P. H. Neuroendocrine Role for VGF. Front. Endocrinol. 6, 3 (2015).
Wang, Y., Qin, X., Han, Y. & Li, B. VGF: A prospective biomarker and therapeutic target for neuroendocrine and nervous system disorders. Biomed. Pharmacother. 151, 113099 (2022).
De Silva, S. R. et al. The X-linked retinopathies: Physiological insights, pathogenic mechanisms, phenotypic features and novel therapies. Prog. Retin Eye Res. 82, 100898 (2021).
Madigan, S. E. J., Natoli, M. C., Penfold, R. & Provis, P. L. JM. Cone degeneration in aging and age-related macular degeneration. Arch. Ophthalmol. 127, 483–492 (2009).
Campochiaro, P. A. & Mir, T. A. The mechanism of cone cell death in Retinitis Pigmentosa. Prog. Retin Eye Res. 62, 24–37 (2018).
Lazaro, R. P., Dentinger, M. P., Rodichok, L. D., Barron, K. D. & Satya-Murti, S. Muscle pathology in Bassen-Kornzweig syndrome and vitamin E deficiency. Am. J. Clin. Pathol. 86, 378–387 (1986).
Natarajan, S. Retinitis pigmentosa: a brief overview. Indian J. Ophthalmol. 59, 343–346 (2011).
D’Souza, S. & Lang, R. A. Retinal ganglion cell interactions shape the developing mammalian visual system. Development 147, dev196535 (2020).
White, T. W. & Bruzzone, R. Intercellular communication in the eye: clarifying the need for connexin diversity. Brain Res. Brain Res. Rev. 32, 130–137 (2000).
Cang, Z. et al. Screening cell-cell communication in spatial transcriptomics via collective optimal transport. Nat. Methods 20, 218–228 (2023).
Zhou, B. et al. Notch signaling pathway: architecture, disease, and therapeutics. Sig. Transduct. Target Ther. 7, 95 (2022).
Mills, E. A. & Goldman, D. The Regulation of Notch Signaling in Retinal Development and Regeneration. Curr. Pathobiol. Rep. 5, 323–331 (2017).
Santiago-Medina, MyersJ. P. & Gomez, M. TM. Regulation of axonal outgrowth and pathfinding by integrin-ECM interactions. Dev. Neurobiol. 71, 901–923 (2011).
Perris, R. & Perissinotto, D. Role of the extracellular matrix during neural crest cell migration. Mech. Dev. 95, 3–21 (2000).
Gramage, E., Li, J. & Hitchcock, P. The expression and function of midkine in the vertebrate retina. Br. J. Pharm. 171, 913–923 (2014).
Campbell, W. A. et al. Midkine is neuroprotective and influences glial reactivity and the formation of Müller glia-derived progenitor cells in chick and mouse retinas. Glia 69, 1515–1539 (2021).
Kuwahara, A. et al. Generation of a ciliary margin-like stem cell niche from self-organizing human retinal tissue. Nat. Commun. 6, 6286 (2015).
Bélanger, M. C., Robert, B. & Cayouette, M. Msx1-Positive Progenitors in the Retinal Ciliary Margin Give Rise to Both Neural and Non-neural Progenies in Mammals. Dev. Cell 40, 137–150 (2017).
Del Debbio, C. B., Peng, X., Xiong, H. & Ahmad, I. Adult ciliary epithelial stem cells generate functional neurons and differentiate into both early and late born retinal neurons under non-cell autonomous influences. BMC Neurosci. 14, 130 (2013).
Varshney, S., Hunter, D. D. & Brunken, W. J. Extracellular Matrix Components Regulate Cellular Polarity and Tissue Structure in the Developing and Mature Retina. J. Ophthalmic Vis. Res. 10, 329–339 (2015).
Barber, T., Esteban-Pretel, G., Marín, M. P. & Timoneda, J. Vitamin a deficiency and alterations in the extracellular matrix. Nutrients 6, 4984–5017 (2014).
Qian, J. et al. Identification of regulatory targets of tissue-specific transcription factors: application to retina-specific gene regulation. Nucleic Acids Res. 33, 3479–3491 (2005).
Schieweck, R., Ninkovic, J. & Kiebler, M. A. RNA-binding proteins balance brain function in health and disease. Physiol. Rev. 101, 1309–1370 (2021).
Prashad, S. & Gopal, P. P. RNA-binding proteins in neurological development and disease. RNA Biol. 18, 972–987 (2021).
Ince-Dunn, G. et al. Neuronal Elav-like (Hu) proteins regulate RNA splicing and abundance to control glutamate levels and neuronal excitability. Neuron 75, 1067–1080 (2012).
Filippou, P. S., Karagiannis, G. S. & Constantinidou, A. Midkine (MDK) growth factor: a key player in cancer progression and a promising therapeutic target. Oncogene 39, 2040–2054 (2020).
Gao, S. et al. Identification of HSC/MPP expansion units in fetal liver by single-cell spatiotemporal transcriptomics. Cell Res. 32, 38–53 (2022).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Blondel, V. D., Guillaume, J. L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory E 2008, P10008 (2008).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. 40, 661–671 (2022).
Yadav, G., Maheshwari, S. & Agarwal, A. Contrast limited adaptive histogram equalization based enhancement for real time video system. In: 2014 international conference on advances in computing, communications and informatics (ICACCI)) (IEEE, 2014).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. In: 2009 IEEE international symposium on biomedical imaging: from nano to macro) (IEEE, 2009).
Zhu, J., Sun, S. & Zhou, X. SPARK-X: non-parametric modeling enables scalable and robust detection of spatial expression patterns for large spatial transcriptomic studies. Genome Biol. 22, 184 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Li, Z. & Zhou, X. BASS: multi-scale and multi-sample analysis enables accurate cell type clustering and spatial domain detection in spatial transcriptomic studies. Genome Biol. 23, 168 (2022).
Long, Y. et al. Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with GraphST. Nat. Commun. 14, 1155 (2023).
Langfelder, P., Zhang, B. & Horvath, S. Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics 24, 719–720 (2008).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Chang, Z., Xu, Y., Dong, X., Gao, Y. & Wang, C. Single-cell and spatial multiomic inference of gene regulatory networks using SCRIPro. Bioinformatics 40, btae466 (2024).
Qin, Q. et al. Lisa: inferring transcriptional regulators through integrative modeling of public chromatin accessibility and ChIP-seq data. Genome Biol. 21, 32 (2020).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Zhang, J. et al. Spatiotemporally resolved transcriptomics reveals the cellular dynamics of human retinal development. Zenodo https://doi.org/10.5281/zenodo.14862032 (2025).
Acknowledgements
This work was partly supported by National Natural Science Foundation of China (T2225015, 61932008, 62433008, 82371698), Shanghai Science and Technology Commission Program (23JS1410100, 24JS2810100, 24JS2810400), Key Science and Technology Project of Hainan Province (ZDYF2024SHFZ058), National Key R&D Program of China (2020YFA0113000, 2021YFC2701600, 2021YFC2701601, 2021YFC2701603, 2024YFC2707500, 2023YFF1204800, 2020YFA0712403), Shenzhen Medical Research Fund (B2404004), Shenzhen Key Laboratory of Maternal and Child Health and Diseases (ZDSYS20230626091559006), Shenzhen Medical Research Special Fund Project (C2401035), Sanming Project of Medicine in Shenzhen (SZSM202211032, SZSM202311005, SZSM202311010), Shenzhen Clinical Research Center for Obstetrics and Reproductive Diseases (LCYSSQ20220823091401002) and Clinical Research Plan of SHDC (SHDC2024CRI077).
Author information
Authors and Affiliations
Contributions
X.M.Z., X.L. and Y.T.Y. conceived the project; J.W., Q.Z., Y.W. and C.W. conducted sample collection; J. Zhang performed raw data quality control and preprocessing; J. Zhang and X.Z. analyzed the data; J. Zhang and Y.T.Y. interpreted the results; Z.C. and J. Zhuang performed the immunohistochemistry experiments; J. Zhang and Y.T.Y. and X.M.Z. prepared the manuscript; Z.G. and R.S.M. contributed in manuscript writing. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Manu Setty, Anand Swaroop and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, J., Wang, J., Zhou, Q. et al. Spatiotemporally resolved transcriptomics reveals the cellular dynamics of human retinal development. Nat Commun 16, 2307 (2025). https://doi.org/10.1038/s41467-025-57625-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57625-9