Abstract
Bidirectional enhancer RNA (eRNA) transcription is a widespread response to environmental signals and glucocorticoids. We investigated whether single nucleotide polymorphisms (SNPs) within dynamically regulated eRNA-transcribing regions contribute to genetic variation in asthma. Through applying multivariate regression modeling with permutation-based significance thresholding to a large clinical cohort, we identified novel associations between asthma and 35 SNPs located in eRNA-transcribing regions implicated in regulating cellular processes relevant to asthma, including rs258760 (mean allele frequency = 0.34, asthma odds ratio = 0.95; P = 5.04E-03). We show that rs258760 disrupts an active aryl hydrocarbon receptor (AHR) response element linked to transcriptional regulation of the glucocorticoid receptor gene by AHR ligands, which are commonly found in combusted air pollution. The role of rs258760 as a protective variant for asthma was independently validated using UK Biobank data. Our findings establish eRNA signatures as a tool for discovery of functional genetic variants and define a novel association between air pollution, glucocorticoid signaling and asthma.
Similar content being viewed by others
Introduction
Asthma is a highly prevalent disease for which pathogenesis and treatment responsiveness are governed by complex stochastic interactions between genetic background and the environment1. Genome-wide association studies (GWAS) have defined numerous single nucleotide polymorphisms (SNPs) and associated pathways linked to risk of developing asthma2,3,4. For example, a large-scale GWAS of asthma in multiple ethnic populations identified 16 high-confidence SNP associations, annotated to HLA-DQA1 and IL1RL1/IL18R1, which have established functional roles for asthma5. However, the sum of heritability explained by these SNPs is estimated to account for only 2.5% of the genetic risk of asthma6,7,8. Moreover, the majority of asthma-associated SNPs defined through GWAS are in non-protein coding portions of the genome9, and direct regulatory effects of most such SNPs have not been established10. These relative weaknesses of GWAS-based approaches are likely due to several underlying factors, including unaccounted effects of environment on asthma risk, linkage disequilibrium, and the very stringent p-values needed to identify significant genetic associations on a genome-wide basis11. These drawbacks have also limited the discovery of genetic variants relevant to specific asthma phenotypes, such as responsiveness to therapeutics. For example, clinical heterogeneity in responses to glucocorticoids in asthma is well described, however, relatively few genetic variants have been discovered that directly modify transcriptional responses to glucocorticoids or that directly link glucocorticoid targets to asthma risk12.
Specialized regions within the genome that control gene expression, frequently referred to as cis-regulatory elements or enhancers, are enriched for functional genetic variants that are linked to disease heritability13,14. As a method to reduce the p-value burden of traditional GWAS and increase the likelihood of identifying putative functional variants, strategies that incorporate enhancer features into genetic association studies have been developed and applied to various diseases15. However, enhancers, as defined using standard assays for chromatin accessibility or histone modifications, can span hundreds of kilobases16, which impacts resolution and statistical power. Moreover, these approaches often fail to define relationships between enhancers and regulation of specific genes17, especially in the context of disease-associated signaling pathways that reshape both enhancer activity and gene transcription.
Sequencing of non-polyadenylated or nascent transcripts has revealed that RNA is transcribed from active enhancers18,19, and that concomitant with changes in gene expression, significant changes in enhancer RNA (eRNA) transcription20,21 occur in response to a range of stimuli, including steroid hormones, inflammatory signals and environmental pollutants. Although the biological functions of enhancer RNAs remain poorly understood, eRNA transcription, which can be identified based on sites of bidirectional transcription exclusive of annotated gene transcription start sites, has emerged as a very sensitive marker of enhancer activity22. Moreover, discrete sites of RNA polymerase II (RNAPII) loading and transcription initiation within enhancers, which frequently serve as a nidus for the dynamic binding of regulatory transcription factors19,23, including the glucocorticoid receptor, can be bioinformatically inferred based on bidirectional transcription signatures23. We hypothesized that regions localized to sites of RNAPII loading within eRNA-transcript signatures that change dynamically in response to anti-inflammatory glucocorticoids or pro-inflammatory signals in airway epithelial cells harbor functional genetic variants associated with asthma risk. Here, we combine multivariate logistic regression modeling with permutation-based significance testing to filter SNPs within a large asthma cohort based on localization within dynamic eRNA-transcript signatures, and we perform fine mapping, annotation and functional validation of a subset of these novel asthma-associated SNPs.
Results
Localization of SNPs within dynamically regulated enhancers and asthma associations in a large clinical cohort
We previously employed Global Run-on Sequencing (GRO-seq) to analyze nascent RNA transcription on a genome-wide basis in BEAS-2B cells in response to 30-minute treatment with the prototypical anti-inflammatory glucocorticoid, dexamethasone (dex), and/or TNF, a canonical inducer of NFkB signaling, widely used to model therapeutic glucocorticoid receptor (GR) cross-talk with inflammatory signals20. Revised analysis of these data with updated algorithms (see Methods) defined, exclusive of transcription start sites (TSSs), a set of 1714 unique bidirectional enhancer RNAs (eRNAs) that responded significantly to dex (672 upregulated, 207 downregulated) or TNF (752 upregulated, 83 downregulated), and 137 eRNAs that responded to more than one condition (Supplementary Data 1; padj < 0.05 vs vehicle). This compares to over 8000 sites of inducible GR or NFkB (the p65 subunit) occupancy identified in our published ChIP-seq analysis in the same cell type24.
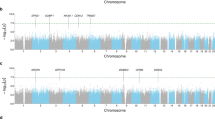
Next, we used Tfit modeling to systematically infer and annotate the sites of RNAPII loading and bidirectional transcription initiation25, frequently referred to as Mu (μ), within the dex- and/or TNF-regulated eRNA set. To investigate potential relationships between these eRNA-transcribing genomic regions and asthma risk in the non-Hispanic white population within the Genetic Epidemiology Research in Adult Health and Aging (GERA) clinical cohort5,26, we leveraged regression modeling, permutation testing and fine mapping approaches27 to create an analysis pipeline (Fig. 1A) for discovery of genetic variants associated with asthma based on proximity to μ. Based on this pipeline, we first screened the GERA dataset to filter SNPs that were annotated within the dynamically regulated enhancer regions, for each treatment separately (dex or TNF). A total of 143,869 SNPs were mapped to dex-responsive enhancer regions and 137,339 were mapped to TNF-responsive enhancer regions. Next, since RNAPII loading sites within eRNA transcribing regions are known to be enriched for functional transcription factor binding motifs23, we filtered these enhancer-SNP sets to include only those that were annotated to regions within 50 bp (+/-) of the Tfit-predicted RNAPII loading site, μ (μ-localized SNPs, or μ-SNPs). Within dex-responsive regions, we identified 866 total annotated μ-SNPs (661 upregulated; 205 downregulated), and within TNF-responsive regions we found 826 μ-SNPs (743 upregulated; 83 downregulated). The lower number of μ-SNPs observed within downregulated regions follows the predicted biological function of enhancers, i.e., dex or TNF perturbation are increasingly recognized as directly affecting gene transcription through upregulating enhancer site function28,29.
A Schematic of pipeline and filtering approaches to identify high confidence μ-SNPs associated with asthma. B–C GRO-seq tracks from BEAS-2B cells treated as indicated for 30 min and visualized in the Integrative Genomics Viewer (IGV) genome browser based on counts per million mapped reads (vertical scales). Blue indicates reads annotated to the sense strand while red indicates reads annotated to the antisense strand. The transcription start site (TSS) and direction of transcription are marked by arrows at the top of each screenshot. Magnified regions show the locations of μ originally calculated by Tfit (green) and following manual refinement (gray) relative to the indicated SNP within dynamically regulated enhancers containing high (B) or low (C) confidence μ-SNPs.
The subsets of μ-SNPs grouped by changes in eRNA transcription with treatment were then tested for association with asthma status using multivariate regression modeling, and the significance of associations was determined using a permutation-based approach implemented in the statistical genetic analysis software, PLINK (v. 1.9)30. Across the four treatment groups, representing 1692 total annotated μ-SNPs, 75 μ-SNPs were initially identified as significantly associated with asthma status (25, dex upregulated; 15, dex downregulated; 32, TNF upregulated; 3, TNF downregulated). SNPs within these regions were further characterized using in silico fine mapping to investigate possible functional impacts and prioritize candidate hits for follow-up study. Ultimately, this analysis pipeline output 39 significantly associated μ-SNPs with compelling evidence for functional roles based on robustness of association with asthma (Table 1) and proximity to μ (Supplementary Table 1). Of these, there were 36 unique μ-SNPs, with 3 μ-SNPs identified within an eRNA region that was regulated by both TNF and dex (rs78553489, rs186724791, and rs150808839). Correlation with our published ChIP-seq analysis, which was performed in BEAS-2B cells24, identified sites of inducible GR or p65 occupancy located within ~ 1 kb of 25 of the 36 unique μ−SNPs (Supplementary Table 1).
As a further step in validating the pertinence of this μ-SNP set in relationship to asthma, searches of the NHGRI-GWAS catalog, ClinVar, and PubMed databases were performed. None of the SNPs were reported as previously associated with asthma by GWAS, suggesting these could represent novel asthma associations based on our methods. Since our analysis is not driven by identifying GWAS signals, but rather as a tool to define functional variants and associated target genes, we alternately interrogated for prior associations at the gene level, specifying overlapping or nearest genes to each μ-SNP as the target. Using published data available in the NHGRI-GWAS catalog and a literature search of PubMed, we identified multiple associations with asthma and/or asthma treatment response, including SLC16A1231 (rs78124271), CXCL832,33 (annotated as the closest gene downstream of rs12506479), DUSP434 (annotated as the closest gene downstream of rs186639917 and rs80028982), and CLDN135,36 (annotated as the closest gene downstream of rs60249091). In aggregate, based on proximity, this preliminary analysis revealed that at least 47% (17/36) of our reported loci were previously related to respiratory disease and relevant immune response phenotypes. These data are summarized in Supplementary Data 2.
Refined mapping and analysis of μ-SNP associations
Confidence in predicting RNAPII loading sites based on Tfit-annotated enhancers depends on the complexity and magnitude of the bidirectional eRNA signature. To focus on SNPs that co-localize with high confidence sites of RNAPII loading (μ) and are thus more likely to be directly associated with enhancer function, we visually inspected the GRO-seq tracks within each enhancer region of interest to manually annotate and refine the bidirectional calls (Supplementary Table 2). One non-unique μ-SNP (rs150808839) was excluded here due to lack of visual confirmation of a clear bidirectional signature in its surrounding region. The remaining 35 regions were passed to a custom script that calculates the predicted location of μ based on each manually refined bidirectional annotation. A total of 24 μ-SNPs (69%) were located within 500 bp (+/-) of the refined μ sites; these were classified as high confidence μ-SNPs and prioritized for further analysis (Supplementary Data 3). Examples of high confidence and lower confidence sites of μ-SNP co-localization are depicted in Fig. 1B, C, respectively. Notably, of the 24 high confidence μ-SNPs, 22 (92%) resided in enhancers that, based on similar transcriptional patterns in response to dex and/or TNF (Supplementary Data 4 and 5; padj < 0.05 vs vehicle), could be linked to regulation of at least one proximal gene within a one megabase (Mb) radius (Supplementary Data 3). Gene functions linked to individual SNPs in this manner encompass signaling processes (e.g., BMP137 and PLK238), ubiquitin ligase activity (HERC439) and transcriptional regulation (CEBPB40).
To further assess the regulatory function of the high confidence μ-SNP regions, we cloned ~450-1100 bp spanning 9 of the high confidence μ-SNP regions into a minimal promoter luciferase reporter (Supplementary Data 6). We assayed responses of these reporters to dex and TNF in BEAS-2B cells in comparison to two previously described control glucocorticoid-responsive reporters (pFKBP541, pTNFAIP342) and two reporters derived from low confidence μ-SNP regions (pCOL8A1, pCFLAR). We found that 7 of the 9 high confidence enhancer reporters (78%) exhibited dynamic changes in activity (Fig. 2A) consistent with regulation of the corresponding endogenous eRNAs (Fig. 2B; representative examples). These data indicate that the high confidence μ-SNPs are likely to confer biologically relevant regulatory function.
A Mean (±SD) normalized luciferase activity of indicated enhancer reporter constructs or empty vector (EV) control in BEAS-2B cells treated as indicated for 8 hr (n = 6 technical replicates per treatment). pFKBP5 and pTNFAIP3 were included as canonical positive controls for dex and TNF responses. *Padj < 0.05 vs same reporter+veh; one-way ANOVA corrected for multiple comparisons using Bonferroni’s method (pFKBP5: Pdex < 0.0001, PTNF+dex = 0.0017; pTNFAIP3: Pdex = 0.0008, PTNF < 0.0001, PTNF+dex < 0.0001; pCOL8A1: PTNF < 0.0001, PTNF+dex < 0.0001; pCFLAR: PTNF < 0.0001, PTNF+dex < 0.0001; pRHOB: Pdex < 0.0001, PTNF+dex < 0.0001; pKCMF1: Pdex < 0.0001, PTNF < 0.0001, PTNF+dex < 0.0001; pANKRD1: Pdex = 0.0087; pCEBPB: Pdex < 0.0001, PTNF < 0.0001; pERRFI1: PTNF < 0.0001, PTNF+dex < 0.0001; pIER3: Pdex = 0.0002, PTNF < 0.0001, PTNF+dex < 0.0001; pOSR1: Pdex < 0.0001, PTNF < 0.0001, PTNF+dex < 0.0001; pNR3C1: Pdex < 0.0001, PTNF < 0.0001, PTNF+dex < 0.0001; pGCNT1: Pdex < 0.0001, PTNF = 0.0003, PTNF+dex < 0.0001.) Source data are provided as a Source Data file. B GRO-seq tracks, as described for Fig. 1, for select regions interrogated by enhancer reporter assay.
Functional analysis characterizes novel SNPs relevant to GR signaling
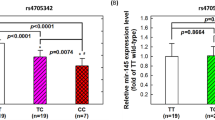
To identify high confidence μ-SNPs that are most likely to exert a functional role in controlling gene expression, we used MatInspector43 to annotate canonical transcription factor binding motifs within 30 bp (+/-) of each SNP site. Through comparing motifs identified for both the major and variant alleles, rs149411423 and rs258760 emerged as strong candidates for functional regulation. rs149411423 is a C > T transversion (minor allele frequency (MAF) in GERA = 2.95E-05; Table 1) located in an enhancer region within PRUNE2 that is ~175 kb from the TSS of GCNT1, which encodes a glycosyltransferase implicated in pulmonary immune responses and allergy44,45. As shown in Fig. 3A, both GCNT1 (log2FC = 2.19, padj = 3.93E-09) and the enhancer harboring rs149411423 (log2FC = 2.57, padj = 0.003) show increased transcription after dex treatment in BEAS-2B cells, whereas PRUNE2 expression was unchanged (log2FC = 0.27, padj = 0.999). GR ChIP-seq analysis in BEAS-2B20,24 (Fig. 3B, top) and primary human airway smooth muscle (HASM) cells46 (Fig. 3B, bottom), another critical effector cell type in asthma, revealed dex-inducible GR occupancy near the rs149411423-μ-SNP region, supporting a direct regulatory role for this enhancer in the induction of GCNT1 transcription and protein expression (see Supplementary Fig 1) by glucocorticoids.
A GRO-seq tracks, as described for Fig.1, for the GCNT1 locus. Scaling applies through panel (D). B Glucocorticoid receptor (GR) ChIP-seq tracks from BEAS-2B (gray; top) and primary human airway smooth muscle (HASM) cells (black; bottom) ±1 hr dex visualized in the IGV browser based on counts per million mapped reads (vertical scales). C Micro-C chromatin contacts in BEAS-2B cells with arcs connecting 1 kb interacting regions across genomic space; darker color indicates higher contact frequency; blue indicates contacts of interest. Arc height is proportional to distance. D Heat map of chromatin contacts in publicly available Micro-C data generated in human embryonic stem cells39; arrows indicate contact between regions of interest. E Magnified view of GRO-seq tracks within rs149411423-μ-SNP region shown in panel (A). Scaling applies through Panel (F). F Aligned ATAC-seq tracks from primary human airway epithelial cells cultured at air-liquid interface (ALI; green), based on counts per million mapped reads (vertical scales), and magnified view of GR ChIP-seq peaks described in panel (B). G Consensus binding logo (MatBase) for GR and sequence of match identified by MatInspector within the GCNT1 enhancer harboring rs149411423. H Mean (±SD) normalized luciferase activity of parent or point mutation GCNT1 reporter constructs in cells treated ±dex for 8 hr (BEAS-2B; n = 6 technical replicates except for pFKBP5 + dex, n = 5) or 24 hr (primary HASM; n = 6 technical replicates except for HASM1: pGCNT1+dex, n = 5 and HASM2: pFKBP5+veh, n = 5). *Padj < 0.05 vs same reporter+veh; unpaired two-sided t-tests corrected for multiple comparisons using Holm-Sidak method (BEAS-2B: all P’s<0.000001; HASM1: P = 0.0028 for pFKBP5, P = 0.00005 for pGCNT1; HASM2: P = 0.0109 for pFKBP5, P = 0.0027 for pGCNT1, P = 0.0047 for pGCNT1 C > T), $Padj < 0.05 vs EV+veh; one-way ANOVA corrected for multiple comparisons using Bonferroni’s test (BEAS-2B: P < 0.0001 for pGCNT1 and pGCNT1 C > T; HASM1: P = 0.0003 for pGCNT1; HASM2: P < 0.0001 for pGCNT1), #Padj < 0.05 vs parent construct+dex; one-way ANOVA corrected for multiple comparisons using Bonferroni’s test (BEAS-2B: P < 0.0001 for pGCNT1 C > T; HASM1: P < 0.0001 for pGCNT1 C > T; HASM2: P < 0.0001 for pGCNT1 C > T). Source data are provided as a Source Data file.
To further assess putative regulation of GCNT1 transcription by the rs149411423-μ-SNP enhancer region, we analyzed three-dimensional chromatin contact maps in BEAS-2B cells using Micro-C data we generated previously47. This was visualized as arcs connecting interacting genomic regions across the locus of interest (Fig. 3C). These data identified three-dimensional contacts between the rs149411423-μ-SNP region and the GCNT1 TSS. Moreover, publicly available Micro-C data generated in human embryonic stem cells48 and visualized in the UCSC Genome Browser defined a topologically associated domain (TAD) that contains both the rs149411423-μ-SNP and the GCNT1 TSS (Fig. 3D).
Further inspection the rs149411423-μ-SNP enhancer (Fig. 3E) illustrates clear overlap of the eRNA bidirectional center with a chromatin accessibility peak in differentiated primary human airway epithelial cells within ATAC-seq data we generated previously49 (Fig. 3F, top) and the dex-inducible GR ChIP-seq peak observed in both BEAS-2B and HASM cells (Fig. 3F, bottom), supporting physiological relevance of the region. MatInspector analysis of the sequence surrounding rs149411423 indicated that the transversion resides within a putative glucocorticoid response element (Fig. 3G). Through applying a customized position weight matrix we created to predict relative GR binding affinities50, we found that the predicted affinity of the sequence containing the major C allele for binding GR is ~2-fold greater than that containing the variant T allele, suggesting the SNP may affect enhancer function. To assess whether rs149411423 directly alters GR-regulated transcription, we generated a reporter harboring the variant allele using site-directed mutagenesis and assayed its activity compared to the parent construct with the major C allele in BEAS-2B and primary HASM cells. Relative to the parent construct, the mutant reporter exhibited dramatically attenuated induction by dex in all tested cell types (Fig. 3H).
Based on these data, we used a CRISPR-based method to target the rs149411423-μ-SNP region (see Supplemental Methods) for deletion in Beas-2B cells51. PCR testing of bulk transfected cells indicated successful editing of the region (Supp Fig. 2) at an estimated efficiency range of 30–75% in 3 independent experiments. Analysis of GCNT1 expression in relationship to estimated editing efficiency indicated that dex-mediated induction of GCNT1 expression is largely mediated through the rs149411423-μ-SNP enhancer (Supp Fig. 2). Taken together, our data implicate rs149411423 as reducing GR-mediated induction of an enhancer that regulates transcription of GCNT1, strongly supporting a functional role for this SNP.
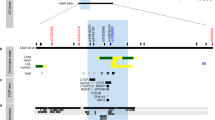
The second strong candidate SNP, rs258760, is a C > T transversion (MAF in GERA = 3.40E-01; Table 1) within an ARHGAP26 intron. This genomic location is approximately 200 kb downstream of the TSS for NR3C1, which encodes the glucocorticoid receptor. To assess whether the enhancer region harboring rs258760 is associated with transcriptional control of ARHGAP26 or NR3C1, we compared eRNA transcription patterns to nascent transcription of both genes after dex or TNF+dex treatment. Figure 4A (GRO-seq tracks) illustrates that eRNA transcription from the rs258760 region is reduced by dex (log2FC = -1.8, padj = 0.035) and TNF+dex treatment (log2FC = -2.01, padj = 0.011). Whereas ARHGAP26 transcription is increased by dex (log2FC = 0.76, padj = 0.035) and TNF+dex (log2FC = 1.41, padj = 1.04E-07), NR3C1 transcription is significantly reduced by treatment with TNF+dex (log2FC = -1.36, padj = 0.0008). As no other gene within a 1 Mb window of the rs258760 region showed significant expression changes with either dex or dex+TNF treatment, these data suggest that rs258760 resides in a regulatory element that controls expression of NR3C1. Further supporting this notion, in our published analysis of nascent transcriptional responses to wood smoke particles (WSP) using Precision Run-on Sequencing (PRO-seq)52, eRNA transcription from the same +200 kb enhancer was significantly increased after 30 minutes of WSP exposure in BEAS-2B cells (log2FC = 1.99, padj = 6.47E-62), with a congruent increase in transcription of NR3C1 (log2FC = 0.90, padj = 9.66E-43), which we validated previously using RT-qPCR52. Expression of ARHGAP26 (log2FC = -0.11, padj = 0.071; Fig. 4A, PRO-seq tracks), however, was unchanged by WSP treatment. Together, these data strongly suggest that the enhancer harboring rs258760 controls transcription of NR3C1, rather than ARHGAP26. This μ-SNP region also aligns with an ATAC-seq peak in differentiated human-derived airway epithelial cells49, providing additional support for the physiologic relevance of this enhancer (Fig. 4B). Furthermore, we previously showed that this enhancer region is occupied by the aryl hydrocarbon receptor (AHR)52, and MatInspector analysis of transcription factor binding motifs in this region in the context of the major C vs variant T alleles revealed that the C allele is a critical nucleotide within a canonical binding motif for AHR. This AHR binding motif is lost in the context of the T allele (Fig. 4C), raising the possibility that AHR ligands, which are found at significant concentrations in wood and wildfire smoke53,54,55, control the activity of this enhancer in an allele-specific manner.
A IGV screenshots of BEAS-2B GRO-seq tracks aligned with PRO-seq tracks (sense strand light blue; antisense light purple) from BEAS-2B cells ±30 min wood smoke particles (WSP) treatment. B ATAC-seq tracks from primary human airway epithelial cells cultured at ALI. C Consensus binding logo (MatBase) for AHR and sequence match identified within the region containing rs258760. D Mean (±SD) normalized luciferase activity of indicated reporters (6 technical replicates) in BEAS-2B cells treated ±TCDD for 8 hr. ^Padj = 0.05 vs EV+veh and *Padj < 0.05 vs same reporter+veh; unpaired two-sided t-tests corrected here and throughout figure for multiple comparisons using Holm-Sidak test (P = 0.0039 for EV; P = 0.0003 for pMT2A, P < 0.0001 for pNR3C1, P = 0.0085 for pNR3C1 C > T), #Padj < 0.05 vs parent construct+TCDD; one-way ANOVA corrected for multiple comparisons using Bonferroni method (P < 0.0001 for pNR3C1 C > T). E qPCR analysis of dex-mediated gene regulation following 4 hr TCDD pre-treatment in BEAS-2B cells (4 technical replicates). Bars depict mean (±SD) CT values on a log2 scale relative to DMSO+veh-treated cells. *Padj < 0.05 vs same gene + DMSO+dex (unpaired two-sided t-tests corrected P = 0.0003 for FKBP5; P = 0.0006 for TNFAIP3; P = 0.0002 for NR3C1). F Magnified view of PRO-seq tracks from (A). G BEAS-2B Micro-C contacts as described for Fig. 3C (blue indicates relevant contacts). H Micro-C interactions39 as described for Fig. 3D; arrows indicate relevant contacts. I Visualization of SNPs by GWAS p-values for associations with forced vital capacity (FVC)57 across the chr5:143,100,000-143,400,000 (NR3C1) topologically-associated domain in the Lung Disease Knowledge Portal; association strength indicated by the color scale. J Mean (±SD) normalized luciferase activity of indicated reporters in BEAS-2B cells treated ±WSP for 8 hr (6 technical replicates). ^Padj < 0.05 vs EV+veh and *Padj < 0.05 vs same reporter+veh (unpaired two-sided t-tests corrected P = 0.0005 for EV, P < 0.0001 for pMT2A, P < 0.0001 for pNR3C1, P = 0.0015 for pNR3C1 C > T, P = 0.0005 for prs258753, P = 0.0081 for prs258753 T > C, P = 0.0005 for prs864354, P = 0.0002 for prs864354 G > T), $Padj < 0.05 vs EV+veh; one-way ANOVA corrected for multiple comparisons using Bonferonni test (all P’s<0.0001), #p < 0.05 vs parent construct+WSP; one-way ANOVA Bonferonni corrected (P < 0.0001 for pNR3C1 C > T). Source data are provided as a Source Data file.
To determine whether AHR and rs258760 regulate activity of this presumptive NR3C1 enhancer, we used site-directed mutagenesis to create a reporter with the minor, asthma-protective T allele, and assayed its activity compared to the parent construct in BEAS-2B cells. As illustrated in Fig. 4D, the parent C allele (pNR3C1) reporter shows robust induction following exposure to the dioxin, TCDD, a potent AHR ligand responsible for toxic effects of Agent Orange56. This induction was markedly reduced in the mutant reporter harboring the minor T allele, with pMT2A serving as a positive control for TCDD responsiveness. Thus, rs258760 disrupts a functional AHR response element linked to regulating expression of NR3C1 in airway epithelial cells in response to AHR ligands. Further, pre-treatment of BEAS-2B cells with TCDD prior to dex exposure potentiates induction of canonical GR transcriptional targets (Fig. 4E), such as FKBP541. These findings implicate the major C allele, which is associated with increased risk of asthma, as integral to an active AHR response element controlling a functional AHR-NR3C1 signaling axis in airway epithelial cells.
To further probe the relevance of the rs258760-μ-SNP in controlling NR3C1 transcriptional regulation, we visualized our BEAS-2B Micro-C data across this region, which indicated three-dimensional physical contacts between the rs258760-μ-SNP locus and the NR3C1 TSS (Fig. 4F, G). These findings were corroborated by Micro-C data generated in human embryonic stem cells48 (Fig. 4H), indicating that the rs258760-μ-SNP and the NR3C1 TSS reside within a TAD roughly encompassing chr5:143,100,000-143,400,000 (hg38). To determine whether rs258760 and/or other SNPs within the chr5:143,100,000-143,400,000 TAD have been independently associated with measures of lung function relevant to asthma, we used the Lung Disease Knowledge Portal (https://lung.hugeamp.org/) to visualize the TAD in genomic space along the x-axis relative to GWAS p-values for associations with lung forced vital capacity (FVC)57, which has been previously defined as relevant to air pollution and asthma58,59,60,61, on the y-axis (Fig. 4I). While rs258760 did not achieve statistical significance, there were significant associations between a number of other SNPs within the chr5:143,100,000-143,400,000 TAD and FVC, suggesting genetic variation within this TAD influences lung function. Of these, the GWAS-defined lead variant in the region, rs864354 (FVC, GWAS p-value = 3.00E-11), lies within a genomic area that has no features of enhancer activity in our nascent sequencing data (see Fig. 4F). In contrast, rs258753 (FVC, GWAS p-value = 1.74E-9) is located within an intergenic enhancer in which eRNA transcription increases in response to WSP (see Fig. 4F). To further explore putative functional relevance of rs864354 and rs258753, we created luciferase reporters driven by the ~500–800 bp genomic regions spanning each transversion site and used site-directed mutagenesis to introduce the relevant variant allele into each reporter context. We directly compared the activities of these reporters to the rs258760-μ-SNP reporter series in response to WSP treatment (Fig. 4J). The reporter spanning the rs864354 SNP did not exhibit appreciable activity relative to the empty vector control, suggesting the region is not regulatory and the SNP is not likely a functional variant. In contrast, the rs258753 reporter showed basal enhancer activity that increased in response to WSP, though not as robustly as the rs258760-μ-SNP reporter, consistent with an absence of AHR/ARNT binding motifs in the vicinity of rs25875 identified using FIMO62 (Supplementary Data 7). And while the rs258753 variant allele reduced responses to WSP relative to the parent construct, the difference was not statistically significant. In aggregate, these data implicate genetic variation in regulatory elements for NR3C1 that respond to WSP as influencing asthma risk and lung function, and establish a novel connection between asthma, combustion-derived pollutants, and control of NR3C1 expression.
To establish generalizability of the eRNA-SNP filtering method and further extend the relevance of the μ-SNP set, we performed a replication study using the UK Biobank (see Supplementary Methods). Demographic data on the 14,302 subject asthma cohort we defined within the UK Biobank are shown in Supplementary Table 3. Of the 11 μ-SNPs with MAF ≥ 5%, 3 high confidence μ-SNPs, including rs258753, replicated with p values from the UK Biobank of < 0.05 and with joint p-values < 0.05, based on the method of Liptak63 (Table 2). Moreover, 7/8 of the remaining μ-SNPs with MAF ≥ 5% exhibited odds ratios that were directionally (e.g., > 1 or < 1) congruent with the odds ratios defined in GERA (aggregate p-value < 0.006). Although replication of the low allele frequency (0.5% ≤ MAF < 5%) and rare (MAF < 0.5%) μ-SNPs was not anticipated due to obligate power constraints associated with less common variants, one rare variant, rs78124271, also replicated as associated with an asthma diagnosis in the UK Biobank (odds ratio 1.45, UK Biobank p-value = 0.027; UK Biobank/GERA combined p-value = 0.00341). Thus, filtering SNPs based on proximity to μ identifies replicable genetic variants associated with asthma.
Discussion
Nascent transcript sequencing is a powerful method for quantifying enhancer activity and has been applied previously to study eQTLs64,65, yet had not been systematically leveraged to identify novel SNP-disease associations. Our results indicate that constraining genetic association analysis based on SNP proximity to bidirectional eRNA transcription initiation sites can facilitate discovery of genetic variants that influence asthma risk, including the identification of disease-associated low frequency and rare variants, which are difficult to associate with disease using traditional GWAS. SNP filtering and analysis based on nascent eRNA transcription signatures offers other unique advantages including enrichment for functional regions within enhancers, which can span thousands of base pairs when defined using other methods, and in facilitating the identification of presumptive target genes for specific SNP-enhancer regions. Moreover, although we focused here on SNPs within eRNA-centered RNAPII loading regions that respond dynamically to glucocorticoids or TNF, this approach is generalizable to eRNAs regulated by other stimuli relevant to asthma, such as Type II cytokines or particulate pollution. Thus, our data provide a systematic method for discovery of functional variants that influence asthma or other airway disease risk in the context of specific endotypes or environmental exposures.
Our experimental approach was empirically designed to address specific limitations of current methodologies. For example, as gene TSSs are well annotated by many methods66, and SNPs located at TSSs do not pose an assignment problem in which the identification of the target gene for a putative regulatory SNP is uncertain, we specifically excluded sites of bidirectional transcription that overlap with gene TSSs from our filtering pipeline. In addition, since sites of bidirectional eRNA transcription are enriched for transcription factor binding19,23, we used narrow 100 bp regions centered on bidirectional eRNA signatures to filter SNPs prior to permutation analysis. This also served to limit the genomic search space, thereby increasing statistical power. However, when we refined the mapping of bidirectional eRNA transcription in relationship to high priority SNPs identified through our pipeline, we expanded the constraint for μ-SNP proximity to 1 kb regions centered on each refined μ, which better reflects the average size of typical enhancers67. Future investigations are needed to determine the optimal size for initial filtering in relationship to μ and to identify centers of bidirectional transcription more efficiently and using scalable bioinformatics methods, with a goal of obviating the manual refinement step we used in this work. Utilizing additional information on enhancer architecture and function, such as ChIP-seq data, may also help inform sequence size optimization and would help clarify potential advantages or drawbacks of applying eRNA signatures as tools for genetic discovery.
We used congruent transcriptional responses between eRNA-transcribing regions and genes located within a 1 Mb region to empirically assign relevant enhancer-gene interactions, with corroborative support provided by chromatin conformation data. This system expands on methods that have used cap analysis gene expression data from different cell types68,69, which can also define enhancer activity, to facilitate enhancer-promoter assignments, and our 1 Mb size window for assignments is derived from these and other data70. A key strength of our system in relationship to defining functional SNP-gene-disease relationships is the use of dynamic eRNA and gene transcription data in response to disease-relevant stimuli, rather than reliance on static expression comparisons across different cell types71. Additional genomic editing using CRISPR-based approaches to directly assess functional interactions between putative enhancer-reporter pairs, as we performed to definitively link the enhancer harboring rs149411423 with control of GCNT1 expression in response to dex, would further validate our approach, although enhancer redundancy can complicate such efforts72.
We undertook several approaches to establish the general relevance of our findings to asthma. First, we performed in silico analysis, which indicated that almost half of the novel μ-SNPs are in proximity to genes previously linked to asthma or immune function in other studies, establishing biologic plausibility of our findings. Second, we tested genomic regions harboring μ-SNPs in reporter assays, and we found that 7 of 9 tested regions conferred regulatory activity in this context, indicating frequent colocalization of μ-SNPs with genomic regulatory elements. Third, we performed a replication study using the UK Biobank, in which 3/11 μ-SNPs with MAF ≥ 5% replicated with p-values of < 0.05%. Moreover, the odds ratios in the UK Biobank for risk of an asthma diagnosis were directionally congruent with the odds ratios in GERA for 10 of the 11 μ-SNPs with MAF ≥ 5%, providing additional statistical support for our method (p < 0.006). These data exceed replication rates reported by others using these cohorts73. Both GERA and the UK biobank use physician-based diagnoses as a criterion for asthma, and standards for physiologic confirmation may differ between the cohorts, potentially adding variability. Although we focused on the non-Hispanic white subgroup within GERA, the ethnic makeup of this subgroup and the UK Biobank are not identical. This serves to further reduce the a priori power of the replication analysis, as prior studies have found that associations between SNPs and an asthma diagnosis are generally dependent on the underlying ethnicity74,75,76, even accounting for ethnicity-related differences in allele frequencies. Similarly, even within ethnically similar cohorts, modest differences in allele frequency can result in opposite allelic effects on disease risk between cohorts77, as we observed here with replication of the rare variant, rs76717004. Future investigations using and/or combining additional cohorts to increase power may provide further confidence in the novel associations we report in this study that have yet to be replicated.
A further limitation of our study is the reliance on BEAS-2B cells as a system to define relevant eRNA regions that respond to dex or TNF. Accordingly, SNPs residing within eRNAs regulated by these signaling pathways in other cell types may not have been interrogated for associations with asthma in our study. In addition, the regulatory and functional associations we defined between eRNA-SNPs and presumptive target genes need to be further validated in additional cell types directly relevant to asthma, including primary airway epithelial cells, airway smooth muscle, and immune cells.
Despite these limitations, our detailed analysis of two variants, rs149411423 and rs258760, directly illustrates the effectiveness of our experimental design for discovery of functional SNPs and extends our understanding of GR signaling in the context of asthma risk. rs149411423, a high risk, rare allele, disrupts a functional GRE that regulates GCNT1. Although rs149411423 did not replicate with UK Biobank data, the very low frequency of rs149411423 significantly reduces the statistical power of such replication studies. However, the functional effects of rs149411423 and the CRISPR-confirmed regulatory association with GCNT1, a glycosyltransferase linked to IgE levels, immune signaling and the pulmonary immune response to infection44,78,79, provide strong biologic support for the relevance of rs149411423 to asthma. Furthermore, GCNT1 is expressed in airway epithelial cells, where it is believed to play a role in mucin glycosylation80, and our data show that the rs149411423 enhancer resides in open chromatin in differentiated primary human airway epithelial cells (Fig. 3). Thus, although it remains to be determined whether rs149411423 directly influences the therapeutic effects of GCs in asthma, we elucidate a novel and physiologically plausible GR-GCNT1 axis that is relevant to asthma risk and highlights GCNT1 as a potential therapeutic target.
Interestingly, rs149411423 was present at very low minor allele frequencies in all populations genotyped in GERA, and imputation of genotypes in GERA was robust26. However, rs149411423 was reported at a notably higher frequency in the African-American population genotyped in the 1000 Genomes Project (MAF in 1000 Genomes = 0.0106). Studies that utilize multi-ethnic cohorts with sufficient sample sizes to provide the statistical power to detect robust associations with asthma are needed to further clarify the roles of this and other low frequency and rare variants we identified in this study. While we limited our analysis to SNPs in GERA with imputation quality scores > 0.9, it is possible that the imputation of the rare SNPs may be subject to errors. For rs149411423, our extensive mechanistic analysis links this SNP directly to altered induction of an asthma-relevant target gene, GCNT1, by glucocorticoids, providing additional support for a link between rs149411423 and asthma. Mechanistic studies of the other rare SNPs may similarly strengthen the genetic findings we report here.
We also discovered a genetic variant, rs258760, a low risk, high frequency allele that regulates expression of the glucocorticoid receptor itself in relationship to asthma risk. In this case the variant allele is the most common allele, and minor allele frequencies were again found to vary by ethnicity in other multi-ethnic cohorts such as 1000 Genomes: (rs258760 MAF in 1000 Genomes = 0.34 for European, 0.117 for African-American and 0.008 for East Asian populations). rs258760 was also one of the 3 common alleles (MAF > 5%) that replicated using UK Biobank data. The more common C allele, associated with increased risk of an asthma diagnosis, is a required nucleotide within a functional binding site for AHR. Our data implicate this AHR response element in controlling increased transcription of airway epithelial GR in response to AHR ligands, which are present at physiologically relevant concentrations in particulate matter air pollution81,82. Our results are supported by evidence that rs258760 and NR3C1 reside within a TAD and physically associate with each other in three dimensions. Moreover, within this TAD, a second SNP, rs258753, which achieved genome-wide significance in a GWAS examining lung forced vital capacity (FVC), is located within a distinct wood smoke-induced enhancer. However, this SNP was not predicted to alter binding site preferences for AHR, and accordingly, our data show that only rs258760 significantly influenced enhancer responses to wood smoke. Thus, we establish an unexpected genetic link between combustion-related air pollution, such as wood smoke, regulation of GR expression, and lung function.
Intriguingly, as the loss of a functional AHR element caused by the rs258760 variant allele was associated with a decreased risk of an asthma diagnosis in both the GERA and UK Biobank cohorts, our data support a model in which local inflammatory repression via GR in the setting of air pollution may have deleterious effects. This raises the possibility that empiric use of inhaled steroids in the context of air pollution, a problem of increasing importance given the malign effects of climate change on air quality83,84,85, may entail unappreciated risks dependent on genetic context. This notion aligns with several studies in which the clinical effectiveness of inhaled corticosteroid use in the setting of air pollution was brought into question86,87,88,89, and supports further investigation of genetic associations between the NR3C1 locus and asthma, and also consideration of a powered clinical trial to directly determine whether inhaled corticosteroids are harmful or beneficial in the context of air pollution. While further exploration and validation are underway, particularly in appropriately powered multi-ethnic cohorts that account for environmental factors, in aggregate, we have defined novel genetic links between GR signaling and asthma risk, and we have established a powerful new application of eRNA transcription signatures for discovery of SNP-disease associations.
Methods
Ethical Standards
This research complies with all relevant ethical standards, and the data used from GERA is derived from studies originally approved by Institutional Review Boards of Kaiser Permanente.
Nascent transcription and cistromic datasets
We previously published the BEAS-2B GRO-seq nascent transcript dataset20 used in the current study with linked Gene Expression Omnibus (GEO) accession number GSE124916 (30 min datasets). Duplicate samples for four treatment conditions [vehicle (ethanol), TNF (20 ng/ml), dexamethasone (100 nM) or TNF+dex] at two time points (10 or 30 minutes) were sequenced in the original study. Raw sequencing data in fastq format as well as non-normalized, strand-differentiated coverage files of hg38-mapped reads in bedGraph format are available for download under this accession series. For the current study, computational analysis to identify regions with bidirectional transcriptional activity (eRNAs) was performed exactly as described20 with the following modification: a newly available version (https://github.com/Dowell-Lab/FStitch) of the Fast Read Stitcher (FStitch) program (v. 1.1.1), with improved efficiency and accuracy impacts on downstream processing, was used for file pre-processing prior to application of Tfit (v. 1.0) to annotate the bidirectional eRNA signatures. Revised eRNA annotations were then subjected to differential expression analysis using DESeq2 to identify differentially transcribed eRNAs (Supplementary Data 1) and their inferred sites of RNAPII loading (μ). The GEO accession number for the primary human airway epithelial cell ATAC-seq data (Norm-1 and Norm-2 datasets), described previously49, is GSE157606. BEAS-2B ChIP-seq data were published previously24, with GEO accession number GSE79803. HASM ChIP-seq data were also described previously46, with GEO accession number GSE95632. Here, to create the screenshots depicted in Fig. 3, we remapped the BEAS-2B and HASM ChIP-seq datasets to hg38 using our standardized ChIP-seq pipeline. Briefly, reads are trimmed for adapters, minimum length, and minimum quality using the bbduk tool from the BBMap Suite (v. 38.05). Trimmed reads are then mapped to hg38 using hisat2 (v. 2.1.0). Resulting SAM files are converted to sorted BAM files using samtools (v. 1.9) and to bedGraph coverage format using genomeCoverageBed from the BEDTools suite (v. 2.29.2). Read coverage is normalized to reads per million mapped using a custom python script and files converted to TDF format using igvtools (v. 2.5.3) for visualization in IGV. The BEAS-2B 30-minute wood smoke particle exposure PRO-seq data depicted in Fig. 4 were published previously52, with GEO accession number GSE167371.
Genomic data
We utilized genotype (SNP) and phenotype information (asthma diagnosis, age, sex, body mass index (BMI), length of follow up, smoking status (current, former, never), and the first six ancestral principal components) from the Genetic Epidemiology Research on Adult Health and Aging (GERA) cohort, which includes over 110,000 subjects (68,623 total asthma cases across four ethnic groups) with extensive electronic medical records (EMR) information and genome-wide SNP genotypes (>8 million typed and imputed markers). The QC methods for GERA genomic data were previously described26. Briefly, extensive QC was performed on genotype data prior to imputation. The Affymetrix Axiom chip was used for genotyping and the Axiom Analysis Suite and Array Power Tools include a Rare Heterozygous Adjusted Algorithm90. Genotypes were pre-phased with Shape-IT v. 2.5 and genetic markers were imputed to 1000 Genomes Project (phase 1 integrated release) reference CEU panels using IMPUTE2. For the eRNA-SNP analysis, we applied a threshold for imputed SNPs to exclude SNPs with imputation quality scores (info r2) < 0.9, significantly higher than the commonly applied lower threshold of 0.3-0.591. As GWAS-based associations of genetic risk factors with disease susceptibility can vary significantly by ethnic group within large multi-ethnic cohorts such as GERA5, we limited the current study to a subset of the cohort with the highest percentage of asthma cases (16,274 non-Hispanic white asthmatic cases and 38,269 controls).
μ-SNP co-localization and statistical association with asthma risk
Unless otherwise indicated, R statistical computing software and PLINK30 (v. 1.9; http://pngu.mgh.harvard.edu/purcell/plink) were used for all analyses.
Statistical methods
To identify genetic predictors for asthma that were also present in dynamically regulated enhancer regions, we first screened the GERA subset (54,543 samples) to filter SNPs that overlapped the Tfit-annotated enhancer regions significantly regulated by dex or TNF (Supplementary Data 1), for each treatment separately. Next, we filtered these SNP sets to include only those that were annotated to regions within 50 bp (+/-) of the Tfit-predicted RNAPII loading site (μ), referred to as μ-SNPs. After sorting by minor allele frequency (MAF), μ-SNPs were categorized as either common (MAF ≥ 0.05) or uncommon (MAF < 0.05) variants and tested independently. Tests for association of additive genotype (categorical variable) with asthma status (dichotomized outcome variable) were conducted using multivariate logistic regression models, adjusted for covariates (listed in the “Genomic data” section above). The significance of associations was determined using a permutation-based approach. One method to improve feature prediction is to combine model-based approaches with permutation. Since each set of dynamically regulated enhancer regions contained relatively few (< 1000) SNPs genotyped in GERA, yielding a small number of tests, far fewer SNPs were interrogated overall as compared to a GWAS, which tests millions of SNPs. In this instance, multiple correction methods traditionally applied in GWAS (e.g., Bonferroni) are overly conservative, or over-correct for Type I error. In contrast to the Chi-square test, permutation-based approaches are appropriate for analysis of rare alleles or smaller numbers of samples and preserve the correlational structure between SNPs, requiring a less stringent multiple test correction threshold. Max(T) permutation was applied with 5000 permutations to generate two sets of empirical (pointwise and familywise) P values. The max(T) permutation approach is advantageous for candidate SNP-based approaches, as it is typically sufficient to perform a much smaller number of tests. Following permutation and multiple testing correction using a false-discovery rate (FDR) threshold of 0.05, associations were prioritized based on their level of significance. PLINK v. 1.9 was used to conduct permutation-based analyses while R was employed for SNP filtering, analysis of summary statistics, and prioritization.
Fine mapping and candidate SNP selection
An in silico fine mapping approach was applied using a web-based annotation tool, SNPnexus92 (v. 4; https://www.snp-nexus.org/v4/), to determine the SNP genomic localization, predicted functions (coding, non-coding, etc.), predicted consequences on protein/gene expression or function, relationship to regulatory elements (e.g., transcription factor motifs, CpG windows), and impact on structural variation (e.g., CNVs). Prior reports of associations with asthma, lung function measures, glucocorticoid response or related phenotypes were also determined through literature search using the NCBI PubMed database. After fine mapping, SNPs were grouped according to whether their respective enhancer regions were up- or down-regulated and compared across groups. The resulting set of SNPs with the strongest predictive evidence for causality and closest proximity to μ were prioritized for functional characterization in vitro.
Manual refinement of μ and classification of high confidence μ -SNPs and associated target genes
For each of the 36 unique candidate μ-SNPs prioritized for follow-up, we visually examined the GRO-seq tracks within the surrounding genomic region in order to manually refine the bidirectional calls originally made by Tfit. We passed these manually refined regions of bidirectional transcription (Supplementary Table 2) to a custom script that calculates the midpoint between the peak signal intensity in each half of the region, resulting in a new estimate of μ for each bidirectional relying both on manual refinement and computational analysis. We then averaged the μ values for any sets of replicates in which the specified bidirectional had robust signal and compared the refined μ estimate to the SNP location. SNPs located within 500 bp (+/-) of the refined μ locations were classified as high confidence μ-SNPs and examined further (Supplementary Data 3). A custom python script, in conjunction with bedtools (v. 2.28.0), was used to identify genes within 1 Mb (+/-) of each SNP and append results from differential gene expression analysis (Supplementary Data 4) for easy comparison with the relevant μ-SNP bidirectional signature. To account for the short 30-minute GRO-seq treatment timepoint potentially missing true signal changes in longer gene transcripts, a second, “truncated” differential gene expression analysis was performed. Assuming a conservative polymerase processivity rate of 3 kb/min93, gene transfer format (GTF) annotation files were generated corresponding to the first 90 kb downstream of all UCSC RefGene-annotated gene TSSs (downloaded August 2019); genes smaller than 90 kb were retained in full. Reads were then counted across these GTF files using the featureCounts algorithm within the R (v. 3.6.1) package Rsubread (v. 3.10) and analyzed with DESeq2 (v. 1.26.0) for differential expression (Supplementary Data 5). Genes for which significant expression changes were primarily evident in the truncated DESeq2 analysis, including NR3C1, which spans ~150 kb, are asterisked in Supplementary Data 3; data presented in the Results section for NR3C1 are from this truncated analysis.
Cell culture and reagents
BEAS-2B transformed normal human airway epithelial cells (ATCC) were grown in DMEM with L-glutamine and 4.5 g/L glucose (Corning) containing 10% FBS (VWR) and 1% penicillin/streptomycin (pen/strep; Corning). Primary human airway smooth muscle (HASM) cells (provided by Dr. Reynold Panettieri, Rutgers Biomedical Health Sciences) were cultured in Ham’s F12 Medium with L-glutamine (Corning) containing 10% FBS and 1% pen/strep. All cells were maintained in 5% CO2 at 37 °C.
Dexamethasone (dex; Sigma-Aldrich) was dissolved in sterile 100% ethanol (vehicle) and used at a final concentration of 100 nM. Recombinant Human TNF-alpha Protein (TNF) purchased from R&D Systems was diluted in sterile 1X Dulbecco’s phosphate buffered saline (DPBS) containing 0.1% bovine serum albumin (BSA) and used at a final concentration of 20 ng/ml. 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) obtained from Cambridge Isotope Laboratories was diluted in sterile DMSO (vehicle) and used at a final concentration of 10 nM. Wood smoke particles were obtained and prepared as described previously52 and used at a final concentration of 1 mg/ml.
Micro-C computational analysis
Paired-end Micro-C data were analyzed using the HiC-Pro94(v. 2.11.4) pipeline, which independently aligns R1 vs R2 reads to the hg38 reference genome using bowtie2 (v. 2.2.9) in end-to-end mode. Prior to inputting raw fastq data into HiC-Pro, we merged R1 and R2 reads from each replicate, respectively. HiC-Pro then filters for valid interactions within specified bins of the genome using samtools (v. 1.1), python (v. 2.7), and the python packages bx-python (v. 0.8.9), scipy (v. 1.2.3), pysam (v. 0.16.0.1) and pandas (v. 0.24.2) to generate raw contact counts between corresponding pairs of loci, output as a single coverage file. We configured HiC-Pro to remove all singleton, multimapped, and duplicate reads, and binned reads at 1 kb intervals, in accordance with the highest suggested resolution for Micro-C. To account for systematic biases inherent in chromatin conformation assays, contact counts were normalized using the iced normalization package (v. 0.5.7). Contacts were then filtered to remove those with a normalized count < 1.0 and were visualized as contact maps using the plotgardener library (v. 1.4.2) in R (v. 4.2.3). Micro-C data is deposited under GSE241294.
Cloning, transfection and luciferase assays
Enhancer reporter constructs were amplified from BEAS-2B genomic DNA by PCR, cloned into pCR2.1-TOPO (Life Technologies), and subsequently ligated into the pGL3-Promoter vector backbone (Life Technologies) using SacI/XhoI (pCOL8A1) or KpnI/XhoI (all others). pFKBP5, pTNFAIP3, and pMT2A positive control constructs have been described41,42,50. The QuikChange II Site-directed Mutagenesis (SDM) Kit from Agilent Technologies was used as instructed by the manufacturer to generate reporter constructs harboring variant alleles (pGCNT1 C > T, pNR3C1 C > T, prs258753 T > C, prs864354 G > T). PCR primer sequences used for cloning and SDM and detailed information pertaining to each reporter construct and related μ-SNP are available in Supplementary Data 6.
For transfection experiments, BEAS-2B cells were seeded on 96-well tissue culture plates at 20 K cells/well in complete medium. The next day, cells were transfected with a total of 200 ng DNA/well (10:1 ratio of firefly reporter construct to Renilla luciferase internal control (pSV40-RL; Promega)) using Lipofectamine 2000 (0.5 ul/well) from Life Technologies. Transfection complexes were removed the following day and cells were treated in fresh complete medium for 8 hr prior to luciferase assay. HASM1 cells were seeded on 96-well tissue culture plates at 50 K cells/well in complete medium. Medium was replaced two days later with fresh complete and the following day, cells were transfected with a total of 125 ng DNA/well (10:1:1 ratio of firefly luciferase to renilla luciferase to mCherry plasmid, respectively) using Lipofectamine 3000 (0.3 ul/well) and P3000 Reagent (0.8 ul/well) from Life Technologies. Transfection complexes were removed the following day and cells were treated in fresh complete medium for 24 hr prior to luciferase assay. HASM2 cells were seeded on 96-well plates at 40 K cells/well in antibiotic-free medium. The next day, cells were transfected with 200 ng total DNA/well (10:1:1 ratio as described for HASM1 cells) using FuGENE6 (1 ul/well) from Promega. Transfection complexes were replaced the following day with fresh complete medium and cells allowed to recover an additional 24 hr prior to 24 hr treatment and luciferase assay.
Luciferase assays were performed using the Dual-Luciferase Reporter Assay System (Promega) and Infinite M1000 Plate Reader (Tecan) as previously described95, with the following modifications for cells grown in 96-well plates: each well was lysed in 25 ul 1X lysis reagent and luminescence detected from 5 ul lysate combined with 40 ul of each assay reagent. Each experiment was performed in technical sextuplet using independent wells (except for those employing TCDD, which were run in technical quadruplicate to reduce waste generation) and repeated at least once with qualitatively similar results.
RNA purification and quantitative RT-PCR (qRT-PCR)
BEAS-2B cells were grown to confluence in 6-well tissue culture plates and treated with vehicle (DMSO) or TCDD (10 nM) for 4 hr. Wells were rinsed 3X in DPBS and then fresh medium containing 100 nM dex was added to all wells for an additional 4 hr. Cells were harvested in TRIzol and RNA purified using the PureLink RNA Mini Kit from Life Technologies prior to qRT-PCR, performed with normalization to RPL19 as previously described95. Primers used for qRT-PCR are described in Supplementary Table 4.
Statistical notes
Statistical tests for comparing reporter assays and RT-qPCR data are described in the Figure Legends. In all cases, tests were performed under the assumption of normality of the underlying distributions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All genomics data used in this manuscript are publicly available, with accession numbers identified above in the relevant methods sections. All computer tools used are available publicly and are referenced. Information for access to GERA data can be found at: https://divisionofresearch.kaiserpermanente.org/research/research-program-on-genes-environment-and-health/. Source data are provided with this paper.
References
Bønnelykke, K. & Ober, C. Leveraging gene-environment interactions and endotypes for asthma gene discovery. J. Allergy Clin. Immunol. 137, 667–679 (2016).
Kim, K. W. & Ober, C. Lessons learned from GWAS of asthma. Allergy Asthma Immunol. Res. 11, 170–187 (2019).
Sleiman, P. M. et al. Variants of DENND1B associated with asthma in children. N. Engl. J. Med. 362, 36–44 (2010).
Bønnelykke, K. et al. A genome-wide association study identifies CDHR3 as a susceptibility locus for early childhood asthma with severe exacerbations. Nat. Genet 46, 51–55 (2014).
Dahlin, A. et al. Large-scale, multiethnic genome-wide association study identifies novel loci contributing to asthma susceptibility in adults. J. Allergy Clin. Immunol. 143, 1633–1635 (2019).
Long, A., Bunning, B., Sampath, V., DeKruyff, R. H. & Nadeau, K. C. Epigenetics and the Environment in Airway Disease: Asthma and Allergic Rhinitis. Adv. Exp. Med Biol. 1253, 153–181 (2020).
Augustine, T., Al-Aghbar, M. A., Al-Kowari, M., Espino-Guarch, M. & van Panhuys, N. Asthma and the Missing Heritability Problem: Necessity for Multiomics Approaches in Determining Accurate Risk Profiles. Front Immunol. 13, 822324 (2022).
Vicente, C. T., Revez, J. A. & Ferreira, M. A. R. Lessons from ten years of genome-wide association studies of asthma. Clin. Transl. Immunol. 6, e165 (2017).
Nasser, J. et al. Genome-wide enhancer maps link risk variants to disease genes. Nature 593, 238–243 (2021).
Alsheikh, A. J. et al. The landscape of GWAS validation; systematic review identifying 309 validated non-coding variants across 130 human diseases. BMC Med Genomics 15, 74 (2022).
Chen, Z., Boehnke, M., Wen, X. & Mukherjee, B. Revisiting the genome-wide significance threshold for common variant GWAS. G3 (Bethesda) 11, jkaa056 (2021).
Ortega, V. E., Meyers, D. A. & Bleecker, E. R. Asthma pharmacogenetics and the development of genetic profiles for personalized medicine. Pharmgenomics Pers. Med. 8, 9–22 (2015).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Demenais, F. et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet 50, 42–53 (2018).
Cano-Gamez, E. & Trynka, G. From GWAS to function: using functional genomics to identify the mechanisms underlying complex diseases. Front Genet 11, 424 (2020).
Grosveld, F., van Staalduinen, J. & Stadhouders, R. Transcriptional regulation by (super)enhancers: from discovery to mechanisms. Annu Rev. Genomics Hum. Genet 22, 127–146 (2021).
Currin, K. W. et al. Genetic effects on liver chromatin accessibility identify disease regulatory variants. Am. J. Hum. Genet 108, 1169–1189 (2021).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Dukler, N. et al. Nascent RNA sequencing reveals a dynamic global transcriptional response at genes and enhancers to the natural medicinal compound celastrol. Genome Res. 27, 1816–1829 (2017).
Sasse, S. K. et al. Nascent transcript analysis of glucocorticoid crosstalk with TNF defines primary and cooperative inflammatory repression. Genome Res. 29, 1753–1765 (2019).
Franco, H. L., Nagari, A. & Kraus, W. L. TNFalpha signaling exposes latent estrogen receptor binding sites to alter the breast cancer cell transcriptome. Mol. Cell 58, 21–34 (2015).
Mikhaylichenko, O. et al. The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription. Genes Dev. 32, 42–57 (2018).
Azofeifa, J. G. et al. Enhancer RNA profiling predicts transcription factor activity. Genome Res. 28, 334–344 (2018).
Kadiyala, V. et al. Cistrome-based cooperation between airway epithelial glucocorticoid receptor and NF-kappaB orchestrates anti-inflammatory effects. J. Biol. Chem. 291, 12673–12687 (2016).
Azofeifa, J. G. & Dowell, R. D. A generative model for the behavior of RNA polymerase. Bioinformatics 33, 227–234 (2017).
Kvale, M. N. et al. Genotyping Informatics and quality control for 100,000 subjects in the genetic epidemiology research on adult health and aging (GERA) cohort. Genetics 200, 1051–1060 (2015).
Yoosefzadeh-Najafabadi, M., Eskandari, M., Belzile, F. & Torkamaneh, D. Genome-wide association study statistical models: a review. Methods Mol. Biol. 2481, 43–62 (2022).
Gerber, A. N., Newton, R. & Sasse, S. K. Repression of transcription by the glucocorticoid receptor: a parsimonious model for the genomics era. J. Biol. Chem. 296, 100687 (2021).
Schmidt, S. F., Larsen, B. D., Loft, A. & Mandrup, S. Cofactor squelching: artifact or fact? Bioessays 38, 618–626 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet 81, 559–575 (2007).
Granell, R. et al. A meta-analysis of genome-wide association studies of childhood wheezing phenotypes identifies ANXA1 as a susceptibility locus for persistent wheezing. Elife 12, e84315 (2023).
Gras, D. et al. Human bronchial epithelium orchestrates dendritic cell activation in severe asthma. Eur. Respir. J. 49, 1602399 (2017).
Qiu, Y., Zhu, J., Bandi, V., Guntupalli, K. K. & Jeffery, P. K. Bronchial mucosal inflammation and upregulation of CXC chemoattractants and receptors in severe exacerbations of asthma. Thorax 62, 475–482 (2007).
Kobayashi, Y. et al. Impaired dual-specificity protein phosphatase DUSP4 reduces corticosteroid sensitivity. Mol. Pharm. 91, 475–481 (2017).
Xia, Y., Cao, H., Zheng, J. & Chen, L. Claudin-1 mediated tight junction dysfunction as a contributor to atopic march. Front Immunol. 13, 927465 (2022).
Fujita, H. et al. Claudin-1 expression in airway smooth muscle exacerbates airway remodeling in asthmatic subjects. J. Allergy Clin. Immunol. 127, 1612–21.e8 (2011).
Hopkins, D. R., Keles, S. & Greenspan, D. S. The bone morphogenetic protein 1/Tolloid-like metalloproteinases. Matrix Biol. 26, 508–523 (2007).
Cozza, G. & Salvi, M. The acidophilic kinases PLK2 and PLK3: structure, substrate targeting and inhibition. Curr. Protein Pept. Sci. 19, 728–745 (2018).
Hochrainer, K. et al. The human HERC family of ubiquitin ligases: novel members, genomic organization, expression profiling, and evolutionary aspects. Genomics 85, 153–164 (2005).
Poghosyan, A., Patel, J. K., Clifford, R. L. & Knox, A. J. Epigenetic dysregulation of interleukin 8 (CXCL8) hypersecretion in cystic fibrosis airway epithelial cells. Biochem Biophys. Res Commun. 476, 431–437 (2016).
Hubler, T. R. & Scammell, J. G. Intronic hormone response elements mediate regulation of FKBP5 by progestins and glucocorticoids. Cell Stress Chaperones 9, 243–252 (2004).
Altonsy, M. O., Sasse, S. K., Phang, T. L. & Gerber, A. N. Context-dependent cooperation between nuclear factor kappaB (NF-kappaB) and the glucocorticoid receptor at a TNFAIP3 intronic enhancer: a mechanism to maintain negative feedback control of inflammation. J. Biol. Chem. 289, 8231–8239 (2014).
Cartharius, K. et al. MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinformatics 21, 2933–2942 (2005).
Fonseca, K. L. et al. Deficiency in the glycosyltransferase Gcnt1 increases susceptibility to tuberculosis through a mechanism involving neutrophils. Mucosal Immunol. 13, 836–848 (2020).
Recto, K. A. et al. Transcriptome-wide association study of circulating IgE levels identifies novel targets for asthma and allergic diseases. Front Immunol. 14, 1080071 (2023).
Sasse, S. K. et al. Glucocorticoid receptor ChIP-seq Identifies PLCD1 as a KLF15 target that represses airway smooth muscle hypertrophy. Am. J. Respir. Cell Mol. Biol. 57, 226–237(2017).
El Kharbili, M. et al. Noncoding SNPs decrease expression of FABP5 during COPD exacerbations. J. Clin. Invest 134, e175626 (2023).
Krietenstein, N. et al. Ultrastructural details of mammalian chromosome architecture. Mol. Cell 78, 554–565.e7 (2020).
Gally, F. et al. The MUC5B-associated variant rs35705950 resides within an enhancer subject to lineage- and disease-dependent epigenetic remodeling. JCI Insight 6, e144294 (2021).
Sasse, S. K. et al. Response Element Composition Governs Correlations between Binding Site Affinity and Transcription in Glucocorticoid Receptor Feed-forward Loops. J. Biol. Chem. 290, 19756–19769 (2015).
Jacobi, A. M. et al. Simplified CRISPR tools for efficient genome editing and streamlined protocols for their delivery into mammalian cells and mouse zygotes. Methods 121-122, 16–28 (2017).
Gupta, A. et al. Deconvolution of multiplexed transcriptional responses to wood smoke particles defines rapid aryl hydrocarbon receptor signaling dynamics. J. Biol. Chem. 297, 101147 (2021).
Young, T. M. et al. Identifying toxicologically significant compounds in urban wildfire ash using in vitro bioassays and high-resolution mass spectrometry. Environ. Sci. Technol. 55, 3657–3667 (2021).
Ghetu, C. C. et al. Wildfire impact on indoor and outdoor PAH Air quality. Environ. Sci. Technol. 56, 10042–10052 (2022).
Wentworth, G. R., Aklilu, Y. A., Landis, M. S. & Hsu, Y. M. Impacts of a large boreal wildfire on ground level atmospheric concentrations of PAHs, VOCs and ozone. Atmos. Environ. 178, 19–30 (2018). (1994).
Sorg, O. AhR signalling and dioxin toxicity. Toxicol. Lett. 230, 225–233 (2014).
Shrine, N. et al. New genetic signals for lung function highlight pathways and chronic obstructive pulmonary disease associations across multiple ancestries. Nat. Genet. 51, 481–493 (2019).
Castelluccio, J. F. et al. Study of correlation between forced vital capacity and demand for healthcare services in severe asthmatics. Multidiscip. Respiratory Med. 10, 22 (2015).
Ackermann-Liebrich, U. et al. Lung function and long term exposure to air pollutants in Switzerland. Study on Air Pollution and Lung Diseases in Adults (SAPALDIA) Team. Am. J. Respir. Crit. Care Med 155, 122–129 (1997).
Miller, A. & Palecki, A. Restrictive impairment in patients with asthma. Respir. Med. 101, 272–276 (2007).
Abbey, D. E. et al. Long-term particulate and other air pollutants and lung function in nonsmokers. Am. J. Respir. Crit. Care Med. 158, 289–298 (1998).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Lipták, T. On the combination of independent tests. Magy. Tud. Akad. Mat. Kut. Int Kozl 3, 171–197 (1958).
Hauberg, M. E. et al. Differential activity of transcribed enhancers in the prefrontal cortex of 537 cases with schizophrenia and controls. Mol. Psychiatry 24, 1685–1695 (2019).
Danko, C. G. et al. Identification of active transcriptional regulatory elements from GRO-seq data. Nat. Methods 12, 433–438 (2015).
Forrest, A. R. R. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Panigrahi, A. & O’Malley, B. W. Mechanisms of enhancer action: the known and the unknown. Genome Biol. 22, 108 (2021).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Furlong, E. E. M. & Levine, M. Developmental enhancers and chromosome topology. Science 361, 1341–1345 (2018).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Cannavò, E. et al. Shadow enhancers are pervasive features of developmental regulatory networks. Curr. Biol. 26, 38–51 (2016).
Edris, A. et al. Asthma exacerbations and eosinophilia in the UK Biobank: a genome-wide association study. ERJ Open Res. 10, 00566–02023 (2024).
Kantor, D. B. et al. Replication and fine mapping of asthma-associated loci in individuals of African ancestry. Hum. Genet 132, 1039–1047 (2013).
Baye, T. M. et al. Differences in candidate gene association between european ancestry and african american asthmatic children. PLOS ONE 6, e16522 (2011).
Chang, X. et al. Genetic architecture of asthma in African American patients. J. Allergy Clin. Immunol. 151, 1132–1136 (2023).
Greene, C. S., Penrod, N. M., Williams, S. M. & Moore, J. H. Failure to replicate a genetic association may provide important clues about genetic architecture. PLoS One 4, e5639 (2009).
Perkey, E. et al. GCNT1-mediated O-glycosylation of the sialomucin CD43 Is a sensitive indicator of notch signaling in activated T Cells. J. Immunol. 204, 1674–1688 (2020).
Kathryn, R. et al. Transcriptome-wide association study of circulating IgE levels identifies novel targets for asthma and allergic diseases. medRxiv 08, 20176479 (2020). 2020.
Chen, G. et al. SPDEF is required for mouse pulmonary goblet cell differentiation and regulates a network of genes associated with mucus production. J. Clin. Invest 119, 2914–2924 (2009).
Weng, C. M. et al. Aryl hydrocarbon receptor activation by diesel exhaust particles mediates epithelium-derived cytokines expression in severe allergic asthma. Allergy 73, 2192–2204 (2018).
Mason, G. G. Dioxin-receptor ligands in urban air and vehicle exhaust. Environ. Health Perspect. 102, 111–116 (1994).
Perera, F. & Nadeau, K. Climate change, fossil-fuel pollution, and children’s health. N. Engl. J. Med. 386, 2303–2314 (2022).
Xu, R. et al. Wildfires, global climate change, and human health. N. Engl. J. Med. 383, 2173–2181 (2020).
Guarnieri, M. & Balmes, J. R. Outdoor air pollution and asthma. Lancet 383, 1581–1592 (2014).
Lewis, T. C. et al. Air pollution and respiratory symptoms among children with asthma: vulnerability by corticosteroid use and residence area. Sci. Total Environ. 448, 48–55 (2013).
Ierodiakonou, D. et al. Ambient air pollution, lung function, and airway responsiveness in asthmatic children. J. Allergy Clin. Immunol. 137, 390–399 (2016).
Zora, J. E. et al. Associations between urban air pollution and pediatric asthma control in El Paso, Texas. Sci. Total Environ. 448, 56–65 (2013).
Gomez, L. et al. Modification of asthma clinical trial treatment efficacy by social and environmental exposures. ISEE Conference Abstracts 2021. https://doi.org/10.1289/isee.2021.O-TO-084 (2021).
Mizrahi-Man, O. et al. Novel genotyping algorithms for rare variants significantly improve the accuracy of Applied Biosystems™ Axiom™ array genotyping calls: retrospective evaluation of UK biobank array data. PLoS One 17, e0277680 (2022).
Roshyara, N. R., Kirsten, H., Horn, K., Ahnert, P. & Scholz, M. Impact of pre-imputation SNP-filtering on genotype imputation results. BMC Genet. 15, 88 (2014).
Oscanoa, J. et al. SNPnexus: a web server for functional annotation of human genome sequence variation (2020 update). Nucleic Acids Res. 48, W185–w192 (2020).
Jonkers, I., Kwak, H. & Lis, J. T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Sasse, S. K. et al. The glucocorticoid receptor and KLF15 regulate gene expression dynamics and integrate signals through feed-forward circuitry. Mol. Cell Biol. 33, 2104–2115 (2013).
Acknowledgements
Funding was provided as follows: NHLBI R01 HL109557 (to A.N.G.); NHLBI R01 HL152244 (to A.D.); NIH R01 GM125871 (to R.D.D.).
Author information
Authors and Affiliations
Contributions
S.K.S. performed, designed, and analyzed experiments, prepared figures, and cowrote the manuscript. A.D. performed, designed, and analyzed experiments, prepared tables, and co-wrote the manuscript. L.S. and M.A.G. analyzed data and reviewed the manuscript. F.G. and A.G. performed, designed, and analyzed experiments and edited the manuscript. A.C.W. and C.I. reviewed the manuscript and curated data. R.D.D. and S.T.W. edited the manuscript, interpreted data, and designed experimental methods. A.N.G. designed and coordinated the study, interpreted data, and cowrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.N.G. and S.K.S. consult for and own shares in Psammiad Therapeutics; R.D.D. is a cofounder of Arpeggio Biosciences; S.T.W. receives royalties from UpToDate and is on the Scientific Board of Histolix. These external interests are not directly relevant to the research presented herein, and did not influence the presented experiments or their interpretation. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Klaus Bønnelykke and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sasse, S.K., Dahlin, A., Sanford, L. et al. Enhancer RNA transcription pinpoints functional genetic variants linked to asthma. Nat Commun 16, 2750 (2025). https://doi.org/10.1038/s41467-025-57693-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57693-x
This article is cited by
-
Implicates of PIP5K1α in asthma-related biological processes: insights into mechanisms and therapeutic potential
Journal of Translational Medicine (2025)
-
Atlas of nascent RNA transcripts reveals tissue-specific enhancer to gene linkages
BMC Genomics (2025)