Abstract
Lysine acylation is a ubiquitous post-translational modification (PTM) that plays pivotal roles in various cellular processes, such as transcription, metabolism, protein localization and folding. Thousands of lysine acylation sites have been identified based on advances in antibody enrichment strategies, highly sensitive analysis by mass spectrometry (MS), and bioinformatics. However, only 27 lysine methacrylation (Kmea) sites have been identified exclusively in histone proteins. It is hard to separate, purify and differentiate the Kmea modification from its structural isomer lysine crotonylation (Kcr) using general biochemical approaches. Here, we identify Kmea sites on a non-histone protein, Cyclophillin A (CypA). To investigate the functions of Kmea in CypA, we develop a general genetic code expansion approach to incorporate a non-canonical amino acid (ncAA) ε-N-Methacryllysine (MeaK) into target proteins and identify interacting proteins of methacrylated CypA using affinity-purification MS. We find that Kmea at CypA site 125 regulates cellular redox homeostasis, and HDAC1 is the regulator of Kmea on CypA. Moreover, we discover that genetically encode Kmea can be further methylated to ε-N-methyl-ε-N-methacrylation (Kmemea) in live cells.
Similar content being viewed by others
Introduction
Lysine acylation is a ubiquitous PTM with crucial functions in biological systems, and dysregulation of lysine acylation has been associated with various diseases, such as cancer, cardiovascular diseases, neurological disorders, and metabolic diseases1,2.
In addition to acetylation, new types of lysine acylation have been continuously discovered3,4,5. Kcr, one of lysine acylation was discovered as PTM in 20116, Since then multiple ‘writers’ and ‘erasers’ of Kcr have been identified7,8,9,10,11, thousands of Kcr sites have been found in both histone and non-histone proteins to regulate diverse biological processes12,13,14,15. Recently, Kmea, a structural isomer of Kcr was identified in histone proteins as an enzyme-catalyzed PTM16. Methacrylyl-CoA, an intermediate in valine metabolism, is the donor of Kmea17. Deficiencies of enoyl-CoA hydrolase (ECHS1) and 3-hydroxyisobutyryl-CoA hydrolase (HIBCH) in the valine catabolic pathway result in the accumulation of highly reactive methacrylyl-CoA, which has been observed in patients with Leigh syndrome, a severe neurological disorder in infancy18,19. However, it is unknown whether methacrylyl-CoA can methacrylate non-histone proteins.
To study the functions of lysine acylation (Kac), lysine to glutamine substitution is often used as a mimic of acetylated lysine20. However, this strategy is not suited for studying Kcr or Kmea due to the dramatic difference in chemical structures. Although proteins can be crotonylated by overexpressing acylation transferases in cells or the incubation of proteins with acylation transferases in vitro, this method gives limited site specificity and selectivity. Moreover, it is challenging to separate Kcr and Kmea modified proteins by chromatography or differentiate them with MS, because Kcr and Kmea are structural isomers. To overcome the above limitations, the genetic code expansion technology has been applied to site-specifically incorporate PTMs into proteins in live cells21,22,23,24. Previously, ε-N-Crotonyllysine (CrK) and photo-cross-linkable CrK have been successfully incorporated into histone proteins, but Kmea hasn’t been genetically encoded yet25,26.
Cyclophilin A (CypA) is an intracellular target of the immunosuppressive drug cyclosporin A (CsA), and it is also known to catalyze proline cis-trans isomerization27. CypA has been associated with several human diseases, including viral infection, cancer, and amyotrophic lateral sclerosis (ALS)28,29,30,31. Its interaction with CD147 is a potential target of anticancer therapy32. Lysine acetylation of CypA has been shown to regulate key functions of CypA in immunity and viral infection33,34, and 9 Kcr sites of CypA have been identified in a large-scale proteomics analysis13. However, there is no report about Kmea in CypA.
Here, we identify Kmea in CypA and develop a genetic code expansion approach to incorporate Kmea into proteins to investigate the function of Kmea of CypA. We identify interacting proteins of methacrylated CypA and find that Kmea enhance the interaction of CypA and oxidoreductase. We reveal that Kmea attenuate suppression of CypA on reactive oxygen species (ROS), and HDAC1 is identified as a demethacrylase which can remove Kmea in CypA. We find that genetically encoded Kmea can be further methylated to Kmemea in live cells.
Results
Identification of Kmea in a non-histone protein CypA
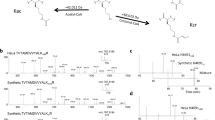
Thioredoxin is a key regulator in redox processes by maintaining a reducing environment in cells to protect cells from the detrimental effects of ROS35. Previously, we incorporated a cross-linkable unnatural amino acid BprY into thioredoxin 1 (TXN1) and applied the OpenUaa software to identify the interacting proteins of TXN136,37. To systematically map the direct interactomes of TXN1, we also developed two cross-linkers BVSB and PDES and performed whole proteome cross-linking in live cells followed by affinity purification and mass spectrometry analysis38. A BVSB mediated cross-linked peptide between TXN1 and Cyclophilin A (CypA) was identified (Fig. 1a), suggesting a direct protein interaction between TXN1 and CypA. CypA is involved in the regulation of ROS by activating peroxidase activity of peroxiredoxins, degrading thioredoxin-interacting proteins, or forming intra-protein disulfide bond39,40,41. To validate this interaction, we performed an affinity-purification mass spectrometry (AP-MS) experiment of CypA and identified TXN1 in the CypA immunoprecipitation (IP) sample (Fig. 1b). Co-IP experiments followed by Western blot analysis further validated this interaction (Supplementary Fig. 1a). CypA has been reported as a substrate of TXN1 transnitrosylation and target of chloroplast thioredoxin42,43.
a Tandem mass spectrum of BVSB mediated cross-linked peptide between TXN1 and CypA. b Previously, protein interaction of TXN1 and CypA was identified in TXN1 His-tag pull down38. To validate protein interaction of TXN1 and CypA, CypA IP experiment was performed and TXN1 was identified in CypA IP samples. c, d HEK293T cells were transfected with CpyA-Strep, Strep-tag purified CypA was digested with trypsin. Digested peptides were analyzed by mass spectrometry. Tandem mass spectrum of peptides contained methacrylation on CypA Lys131 (c) and Lys125 (d). The K in red represents the Kmea modification. e Validation of Kmea on CypA by Western blot analysis. When CypA Lys125 was mutated to Arg, the Kmea level of CypA dramatically decreased. The experiment was repeated three times with similar results. When CypA Lys131 was mutated to Arg, the Kmea level of CypA partially decreased. f Proposed fragmentation pathway of CycIm ion from methacrylated peptide. g CycIm ion was mapped on MS/MS spectrum of methacrylated peptide (Kmea on CypA Lys131). h Confirm CycIm ion with chemically synthesized standard methacrylated peptide.
We performed an open search using pFind 344 to analyze the AP-MS data. Interestingly, we found two tryptic peptides with lysine modifications at CypA K125 and CypA K131 (mass difference of +68.023 Da), which is equal to the mass of Kcr or Kmea (Fig. 1c, d). To differentiate these two modifications, pan-specific anti-Kmea and anti-Kcr antibodies were used. Using synthetic peptides, we demonstrated that the anti-Kmea antibody is highly specific towards Kmea, while some cross-reactivities for Kmea and Kcr were observed for the anti-Kcr antibody (Supplementary Fig. 1b). Western blot analysis using the anti-Kmea antibody showed an intense Kmea band. The band intensity was decreased, when either site 125 or 131 was mutated to Arg (Fig. 1e and Supplementary Fig. 1c). We also treated cells with different concentrations of sodium methacrylate to increase the production of methacrylyl-CoA. As expected, the Kmea band intensity also increased with the increasing doses of sodium methacrylate (Supplementary Fig. 1d). These data indicate that CypA is methacrylated at both site 125 and 131. To the best of our knowledge, methacrylation has not been previously reported on a non-histone protein. Previously, a total of 9 Kcr sites on CypA were identified in a large Kcr dataset from HeLa cells, which contained the sites 125 and 13113. Similar to cyclic immonium (CycIm) ions found in the tandem mass spectra of lysine lactylated peptides45, we also proposed a fragmentation pathway of CycIm ion in methacrylated peptides (Fig. 1f). We observed CycIm ion at m/z 152.107 in the tandem mass spectra of Kmea peptide and synthesized Kmea peptide (Fig. 1g, h and Supplementary Fig. 1e).
Genetically encoding Kmea into proteins in E. coli and mammalian cells
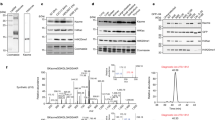
To investigate the functions of Kmea, we attempted to incorporate a ncAA (MeaK, Fig. 2a) into CypA. Based on the published protein structure, the amino acid binding pocket of wild-type Methanosarcina barkeri pyrrolysyl-tRNA synthetase (MbPylRS) was engineered to obtain a synthetase of MeaK (Fig. 2b). We constructed 5 mutants of MbPylRS and tested their incorporation efficiency. The MbPylRS (Y349F)/MbtRNApylCUA pair showed the high incorporation efficiency of MeaK and fidelity (Supplementary Fig. 2). Therefore, this mutant was selected to incorporate MeaK into a model protein MBP-Z. An MBP-Z construct containing a TAG codon at site 24 and a C-terminal His-tag was co-expressed with the MbtRNApylCUA/MbPylRS (Y349F) pair in E. coli DH10B cells to successfully produce the full-length MBP-Z protein in the presence of MeaK (Fig. 2c and Supplementary Fig. 3). The purified MBP-Z (theoretical molecular weight: 50893.21 Da) was confirmed by intact protein mass analysis (Fig. 2d). The incorporation of MeaK into EGFP was also supported by high- resolution tandem MS (Fig. 2e), showing the signature CycIm fragment ion (Fig. 2f). These results demonstrated that Kmea modified proteins can be prepared in E. coli for in vitro biochemical assays.
a Chemical structure of Meak. b Left: X-ray crystal structure of the MmPylRS complex with pyrrolysyl-AMP (PDB ID: 2Q7H), active-site residues for mutation are highlighted in green. Right: residue numbering is based on MbPylRS and MmPylRS. c SDS-PAGE gel of His-tag purified MBP-Z E24MeaK. Protein Z is IgG Fc-binding domain62, MBP-Z is maltose binding protein tagged protein Z. d Intact protein mass analysis of purified protein from E. coli expressed MBP-Z E24MeaK. e Tandem mass spectrum of MeaK incorporated peptide of EGFP D190MeaK. The red U represents MeaK incorporation site. f CycIm ion was mapped on the MS/MS spectrum of the MeaK incorporated peptide. g Genetically encode Meak into EGFP protein in mammalian cells. FACS analysis of HEK293T cells after MeaK incorporation into EGFP-151TAG. h Left: Fluorescence images of EGFP Y151MeaK showing incorporation of MeaK into EGFP in HEK293T cells. Right: Select images from different fields of view under a 10× objective lens, and use ImageJ software to calculate the mean fluorescence intensity (n = 5). Error bar represents ± one standard deviation (SD). i Intact protein mass analysis of HEK293T cells expressed EGFP Y151MeaK. j Tandem mass spectrum of MeaK incorporated peptide of EGFP Y151MeaK.
To assess the potential application of MeaK incorporation in mammalian cells, we engineered 3 aminoacyl-tRNA synthetase mutants (named MeaKRS6-8) of chPylRS46. The chPylRS Y384F/MmtRNApylCUA pair showed the highest incorporation efficiency of MeaK (Supplementary Fig. 4a). Different concentrations of MeaK were tested to culture cells, and the best condition with the final concentration of 1 mM was chosen for MeaK incorporation (Supplementary Fig. 4b). We co-expressed the chPylRS Y384F/MmtRNApylCUA pair with C-terminal Strep-tagged EGFP Y151TAG in HEK293T cells (Supplementary Fig. 3a). The suppression of 151TAG codon by chPylRS Y384F will produce full-length functional EGFP to render cells fluorescent. After adding MeaK to the cell culture, strong EGFP fluorescence was detected in cells by flow cytometry (Fig. 2g). The incorporation of MeaK into EGFP was also validated and quantified by fluorescence microscopy (Fig. 2h). Furthermore, MeaK incorporated EGFP (theoretical molecular weight: 30,003.42 Da) was confirmed by intact protein mass analysis and tandem MS sequencing (Fig. 2i, j), and again the CycIm ion was observed in the tandem mass spectrum (Supplementary Fig. 5). Western blot analysis also validated the MeaK incorporation into EGFP (Supplementary Fig. 3a). To further test the robustness of encoding MeaK in mammalian cells, we incorporated MeaK into CypA K131TAG and histone H3 K18TAG. Full-length CypA and H3 were successfully detected on Western blot (Supplementary Fig. 6). These data demonstrated that MeaK can be site-specifically encoded in mammalian cells to study functions of Kmea.
Comparing interacting proteins of wild-type CypA, CypA mutant, CrK incorporated CypA, and MeaK incorporated CypA
The K125 residue of CypA is highly conserved in mammals (Fig. 3a). K131 is also conserved, but it was mutated to methionine or arginine in Felis Catus and Oryctolagus cuniculus, respectively (Fig. 3a). In several cancer types with available quantitative acetylation proteomic data47, the ratio of acetylated CypA over total CypA is higher in lung adenocarcinoma cells than adjacent normal tissues (Fig. 3b and Supplementary Fig. 7). To gain further insight into the functions of Kmea and how Kmea affects interactions of CypA with other proteins, we constructed wild type (WT) CypA, CypA K125A and CypA K125MeaK, and these constructs all contain the Strep-tag for immunoprecipitation (IP). Each construct was transfected into HEK293T cells. Tandem MS analysis confirmed that MeaK was successfully incorporated at residue 125 of CypA (Fig. 3c). We performed affinity purification and label-free quantitative MS analysis for each construct to investigate how methacrylation affects the interactome of CypA (Fig. 3d and Supplementary Fig. 8a). PCA analysis on identified interacting proteins showed separation for proteins from different IP experiments (Supplementary Fig. 8b).
a Alignment of K125 and K131-flanking sequences of CypA from multiple vertebrate species. CypA K125 and CypA K131 are highlighted in red. b Comparison of CypA K125 acetylation levels between tumor and adjacent tissues from different cancer types. Raw data were obtained from six acylated proteomes of CPTAC (Clinical Proteomic Tumor Analysis Consortium) database. **P < 0.01 (two-tailed t-test). p = 0.0228 and 0.0046, respectively. The center line denotes the median value, the box contains the 25th to 75th percentiles. The whiskers mark the minima and maxima. Sample numbers are provided in the Source Data file. c Tandem mass spectrum of MeaK incorporated peptide from HEK293T cells expressed CypA K125MeaK. The U in red contains the Kmea modification. d SDS-PAGE gel analysis of CypA (K125MeaK, K125A, WT) IP with three biological replicates. Nanobody comes from alpaca-derived heavy chain antibody fragment (VHH) carried by Strep beads. e Valcano plot of semi-quantification result on CypA and MeaK incorporated CypA IP. Red dots, significantly enriched hits (Fold Change ≥2, p-value < 0.05). Gray dots, depleted hits or non-differentially expressed proteins. Data quantified with the Empirical Bayes test two-sided. f Biological process analysis of the enriched hits in (e) and unique proteins identified in CypA K125MeaK in Strep-IP.
We focused on enriched hits in CypA K125MeaK IP and binding proteins identified only in CypA K125MeaK IP because it showed a lower expression level compared to CypA K125A and WT (Fig. 3d and Supplementary Fig. 8a). In comparison to the WT CypA, interactions of 68 proteins with CypA were enriched after MeaK incorporation and 10 binding proteins were only identified in CypA K125MeaK IP samples (Fig. 3e and Supplementary Dataset 1). Similarly, compared to CypA K125A IP, a total of 70 proteins were enriched in CypA K125MeaK IP and 10 binding proteins were identified in CypA K125MeaK IP only (Supplementary Fig. 8c). A total of 52 proteins were enriched in CypA K125MeaK IP compared with WT CypA and CypA K125A IP (Supplementary Fig. 8d). Biological process analysis showed that some of these proteins have protein-disulfide reductase activities or involved in cell redox homeostasis (Fig. 3f).
We also carried out CypA (K131MeaK, K131A and WT) IP, MeaK was successfully incorporated into site 131 of CypA (Supplementary Fig. 9a). Similarly, the expression level of CypA K131MeaK is lower than CypA K131A and WT (Supplementary Fig. 8b, c), PCA analysis showed that identified proteins from different IPs could be separated (Supplementary Fig. 9d). A total of 33 proteins were enriched in CypA K131MeaK IP compared with CypA (K131A, WT) IP (Supplementary Fig. 9e, f and Supplementary Dataset 2).
To compare the effect of Kmea and Kcr on the interactome of CypA. We performed CypA125 CrK IP and CypA125 MeaK IP (Supplementary Fig. 10a). A total of 478 proteins were identified in CypA125 CrK IP and 465 proteins were identified in CypA K125MeaK IP (Supplementary Fig. 10b, Supplementary Dataset 3). In biological processes, the unique proteins of CypA K125MeaK IP are mainly involving in nucleic acid processing and cell differentiation (Supplementary Fig. 10c). Compared to Kcr at the CypA125 site, the presence of Kmea would likely affect cell differentiation and enhancement of hits related to the cGAS/STING and PI3K/AKT signaling pathway (Supplementary Fig. 10d).
Methacrylation at K125 repressed the antioxidant capacity of CypA
Cyclophilin interacts with multiple oxidoreductases and has been identified as a target protein of chloroplast thioredoxin in spinach43,48. It also links redox and light signals to stress response in Arabidopsis49. CypA can be glutathionylated and S-Nitrosylated in mammalian cells42,50, and it binds to peroxiredoxin VI to enhance its antioxidant activity of reducing peroxides to corresponding alcohols39. Recently, CypA has been reported to repress ROS in cancer cells40,41. The overexpression of CypA reduced ROS generation in hypoxic non-small-cell lung cancer (NSCLC) cells, and the knock-down of CypA elevated ROS levels40. Similar effects were also observed in colorectal cancer cells41. CsA, a peptide binder of CypA, has been shown to inhibit the antioxidant capacity of CypA41.
The encoded MeaK is similarly robustly incorporated into the protein in HeLa cells (Supplementary Fig. 11). To test whether methacrylation at K125 can impact the ability of CypA to repress ROS, we incorporated MeaK at residue 125 of CypA in HeLa cells. We generated constructs of CypA (WT or K125MeaK) with mCherry linked via a self-cleavable P2A sequence51. CypA WT-mCherry or CypA K125MeaK-mCherry expressed HeLa cells were treated with or without CsA, and then the cells were incubated with a ROS probe 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) to evaluate the ROS levels. DCFH-DA can be converted to 2′,7′-dichlorodihydrofluorescein (DCFH) by intracellular stearases, and further oxidized to green fluorescent 2′,7′-dichlorofluorescein (DCF) in the presence of ROS52. Cells were first sorted for mCherry fluorescence by flow cytometry, followed by selection based on green fluorescence of DCF (Fig. 4a–c). Compared to WT control cells, cells expressing CypA 125MeaK showed a significantly higher ROS level. Consistent to the previous report41, treating WT cells with CsA also elevated the ROS level (Fig. 4d). These findings suggest that Kmea at the site 125 of CypA may perturb CypA’s ability to suppress ROS (Fig. 4e). Interestingly, no significant change in the ROS level was observed after treating CypA 125MeaK expressing cells with CsA (Fig. 4c). Previously, acetylation of CypA at site 125 has been reported to inhibit binding between CypA and CsA by disrupting water network34. Similarly, using microscale thermophoresis (MST), we confirmed that methacrylation at K125 also reduced the binding affinity of CsA to CypA (Supplementary Fig. 12).
a–c HeLa cells were transfected with CypA-WT-mCherry or CypA-K125MeaK-mCherry, which were either treated with/without 4 µM CsA for 36 h. ROS was measured by flow cytometry. Fluorescence intensity of FITC channels was detected after selection of the same number of CypA+ cells by ECD channel. d Quantitative comparison of ROS levels in different cells. n = 3 independent experiments. ns, not significant; *p < 0.05; **p < 0.01, p = 0.0402 and 0.0048, respectively; (two-tailed t-test/One-way ANOVA analysis). Error bar represents ±one standard deviation. e Model showing CypA 125Kmea modification increases ROS level of cells and it blocks the suppression of CsA on CypA.
CypA Kmea is regulated by HDAC1
To test whether genetically encoded Kmea can be recognized by endogenous deacetylases, we developed a fluorogenic probe EGFP K85MeaK with MeaK incorporated at position 85. This probe is nonfluorescent due to the modification at K85. However, after the removal of the methacrylation modification by demethacrylase, this probe will become fluorescent53. Therefore, if the activity of deacetylases is suppressed, EGFP probe will remain nonfluorescent (Fig. 5a). We generated a construct with the fluorescent probe and mCherry linked by a self-cleavable P2A sequence (Fig. 5b). This construct was co-expressed with the chPylRS Y384F/tRNApylCUA pair with or without MeaK treatment in HEK293T cells. Microscopic imaging showed that red fluorescence was consistent in all samples, but the green fluorescence was dependent on the presence of MeaK. Moreover, green fluorescence was suppressed by pan histone deacetylase (HDAC) inhibitor sodium butyrate (NaBu) and pan-Sirtuin inhibitor nicotinamide (NAM) (Fig. 5b, c and Supplementary Fig. 13). Therefore, genetically encoded Kmea can be correctly recognized by endogenous deacetylase.
a Genetically encoded EGFP K85TAG-based probe for de-conjucation enzymes of Kmea. b Top: Construct of Fluorescent reporter containing EGFP K85TAG, self-cleavable peptide P2A and mCherry. Down: Fluorescence imaging of HDACs demethacrylation activity in HEK293T cells. The cells were co-transfected with two plasmids: one containing chPylRS Y384F/tRNAPylCUA and another containing mCherry-EGFP K85TAG; MeaK (final concentration 1 mM) and NaBu (final concentration 10 mM) were added after transfection 6 h. Imaging was carried out after 36 h. Scale bars, 150 µm. c Normalizing signal of EGFP with mCherry. The error bars represent standard deviation from the mean. n = 3 independent experiments. ns, not significant; ***p < 0.001, p = 0.0004. d Binding of deacetylases to methacrylated CypA, which were detected by AP-MS from HEK293T cells expressed CypA K125MeaK. e Interactions between methacrylated CypA and HDACs with Co-IP experiments. f Western blot analysis showing successful demethacrylation assay by HDAC1 in vivo. g Purificated CypA K125MeaK protein was incubated with or without recombinant HDAC1 for 2 h at 30 °C. Samples were then analyzed by Western blot.
To identify which enzyme is the ‘eraser’ for CypA Kmea, we checked identified interacting proteins in the CypA K125MeaK IP experiment and found that HDAC1-3 interact with CypA K125MeaK (Fig. 5d). We carried out Co-IP experiments to validate the interactions between CypA K125MeaK and HDAC1-3. Western blot analyses demonstrated that CypA K125MeaK indeed interacted with HDAC1-3 (Fig. 5e). In addition, the level of Kmea modification on CypA decreased when Flag-tagged HDAC1 was co-expressed in HEK293T cells (Fig. 5f). The incubation of recombinant Flag-HDAC1 with purified CypA-K125MeaK in vitro also greatly reduced the Kmea level in Western blot signal (Fig. 5g), implying that HDAC1 is a potential regulator of Kmea on CypA. However, no change was observed when HDAC2 or HDAC3 was co-expressed (Supplementary Fig. 14). As previously reported, CypA has 9 Kac sites and 9 Kcr sites13. It is possible that HDAC2 and HDAC3 are involved in deacylation of other acylation sites on CypA. We also tested interactions between methacylated CypA and surtuins. SIRT7 interacted with CypA K125MeaK (Supplementary Fig. 15).
Methylation on genetically encoded Kmea
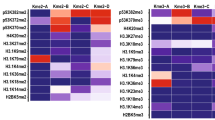
Recently, lysine ε-N-acetyl-ε-N-methylation (Kacme), a PTM containing methylation and acetylation on the same lysine side chain, has been found on histone H4 in multiple species (Fig. 6a)54. To investigate whether genetically encoded Kmea can be further methylated (Fig. 6b), we check the MS data of CypA K125MeaK and CypA K131MeaK IP samples. A mass shift of ~14.016 Da was detected on tryptic peptides containing K125MeaK and K131MeaK (Fig. 6c, d) with a series of continuous fragment ions precisely supported the identification of this modification. This result suggests that genetically encoded Kmea can be further methylated to Kmemea in live cells. We also propose a possible fragmentation pathway of the linear ammonium (LinIm) ion from methyl-methacylated peptide (Fig. 6e). The LinIm ion of Kmemea at m/z 183.149 was observed on tandem mass spectra of methyl-methacylated peptides containing CypA K125Kmemea and CypA K131Kmemea (Fig. 6f, g), providing strong evidence of this modification.
a Chemical structure of Kacme. b Chemical structure of Kmemea. c Tandem mass spectrum of methylation on CypA K125MeaK. d Tandem mass spectrum of methylation on CypA K131MeaK. e Proposed fragmentation pathway of LinIm ion from methyl-methacrylated peptide. f LinIm ion on tandem mass spectrum of peptide containing CypA K125Kmemea. g LinIm ion on tandem mass spectrum of peptide containing CypA K131Kmemea.
Discussion
Methacrylation is an enzyme-catalyzed biological process, and HAT1 and SIRT2 have been identified as methacryltransferase and demethacrylase, respectively. Here, we found that HDAC1 can demethacrylate CypA at site 125. Methacrylyl-CoA is the metharyl-group donor used by methacryltransferase to form Kmea16. As a metabolite in the mitochondrial catabolism of valine, Methacrylyl-CoA is converted by ECHS1 to 3-hydroxyisobutyryl-CoA, which is further converted to 3-hydroxyisobutyrate by HIBCH55. Deficiencies of ECHS1 and HIBCH will cause accumulation of Methacrylyl-CoA and lead to Leigh or Leigh-like syndrome18. However, it is not clear which enzymes were responsible for methacrylation and de-methacylation in mitochondria. It is also not clear whether methacrylation in mitochondria is enzyme catalyzed. These questions need to be addressed in the future. Although the stoichiometry of Kmea has not been characterized, Kac has been reported to be at very low stoichiometry (median 0.02%)56. Therefore, it is difficult to obtain a large amount of site-specifically acylated proteins for biochemical assays by purifying endogenous proteins. Genetically encoded MeaK paves the way for preparing methacrylated proteins and site-specifically introducing Kmea into proteins in live cells to investigate the functions of methacrylated proteins associated with the Leigh syndrome.
From IP experiments using CypA K125MeaK, we have identified well-known binding proteins of CypA, such as CD147, TDP-43, HNRNPA1, HNRNPA2B1, etc. Cytoplasmic aggregation and mislocalization from nucleus to cytoplasm of TDP-43 are pathological hallmarks of ALS57, and the perturbation of the CypA/TDP-43 interaction can cause “TDP-43” pathology, it has been shown that acetylation of K125 on CypA promote CypA/TDP-43 interaction and CypA is necessary for the assembly of TDP-43 in heterogeneous nuclear ribonucleoprotein (hnRNP) complexes and their stability31. We have shown that K125MeaK slightly enhanced CypA/TDP-43 interaction, and the interactions between CypA K125MeaK and hnRNP proteins (HNRNPA1, HNRNPA2B1 and HNRNPAB) were significantly enhanced (Supplementary Fig. 16). These data suggest that CypA K125Kmea possibly keep hnRNP proteins soluble, stabilizes hnRNP complexes and involves in “TDP-43” pathology.
Peroxiredoxins, a ubiquitous family of antioxidant enzymes, catalyze the reduction of peroxides58. Among the six members (PRDX1-6) in mammalian cells, PRDX6 was reported to bind CypA and CypA enhances antioxidant activity of PRDX1-6, activity of PRDX4 was enhanced 3.5 fold by CypA39. In our CypA IP experiments, PRDX4 in CypA K125MeaK IP is 3.5 fold less than intensity of PRDX4 in CypA K125A IP. There is no significant change on the intensity of PRDX4 in CypA K125MeaK IP and CypA WT IP. These data suggest that Kmea of CypA may perturb interaction between CypA and PRDX4 and suppress antioxidant activity of CypA.
In this study, we have identified methacrylation at K125 and K131 of CypA. Previously, mono-methylation has been identified at K49 and K125 of CypA in KYSE-150 cells using antibody enrichment by a pan-specific mono-methyl-lysine antibody followed by mass spectrometry analysis59. Here, we discovered further methylation on genetically encoded Kmea (Kmemea). Although we have not identified endogenous Kmemea on CypA, our data demonstrated that Kmea can be further methylated in live cells. Based on the reaction mechanism of lysine methyltransferases and acetyltransferases, Lu-Culligan et al. proposed that Kacme could be generated by first lysine mono-methylation followed by acetylation. They demonstrated that acetyltransferase P300 is a writer of Kacme. Although it is also possible that Kacme is formed by methylation of Kac54.
We believe that a pan-specific Kmemea antibody is needed to comprehensively identify and investigate endogenous Kmemea. We have performed data mining on various high-throughput proteomics data. So far, no endogenous Kmemea site has been identified. Peptide-level enrichment with pan-specific Kmemea antibody could be the key step for mapping endogenous Kmemea sites. Novel ncAAs will also be important tools. For example, ε-N-Methyl-ε-N-Methacryllysine (MemeaK) with a cross-linkable warhead or the incorporation of MemeaK and cross-linkable ncAA EFSY60 into the same protein will allow mapping of readers, writers and erasers of Kmemea. Diazirine is a photo-activated cross-linking group61, equipping MemeaK with a diazirine group (xMemeaK) and incorporating this ncAA xMemeaK into protein can covalently capture binding proteins of Methyl-methacrylated proteins in live cells and increase sensitivity on identifying interacting proteins. If incorporation of MeaK/MemeaK and EFSY at different sites on the same protein, MeaK/MemeaK can recruit readers, writers, and erasers of Kmemea, and EFSY can covalently fix these protein-protein interactions and enrich to these proteins through its alkyne group.
In summary, we have developed a strategy to site-specifically incorporate lysine methacrylation (Kmea) into proteins in live cells. Interacting proteins of methacrylated CypA were identified, revealing that oxidoreductases were enriched in methacrylated CypA IP. Our finding indicates that the presence of Kmea on CypA reduces the ability to suppress ROS, and HDAC1 can demethacrylate Kmea on site 125 of CypA, suggesting a regulatory mechanism for this PTM. We demonstrated that genetically encoded Kmea can be further methylated in live cells. Identification of new types of PTM and new sites of modification far outpaced functional studies of these sites. Our method to site-specifically incorporate Kmea into proteins in live cells and prepare methacrylated proteins with high yields will accelerate functional annotation of Kmea sites.
Methods
Reagents and strains
Unless otherwise noted, all commercial reagents were used directly without further purification. The following antibodies were used: anti-6×His-tag (Mouse, Proteintech, cat. no. HRP-66005, 1:10,000), anti-FLAG tag (Mouse, HUABIO, cat. no. M1403-2, 1:10,000), anti-Strep-tag II (Mouse, Wuhan Dian Biotechnology, cat. no. 2098, 1:10,000), Goat anti-Mouse IgG (H+L) HRP conjugate (Jackson ImmunoResearch, Code. 115-035-003, 1:10,000), Goat anti-rabbit IgG (H+L) HRP conjugate (Jackson ImmunoResearch, Code. 115-035-003, 1:10,000), anti-Methacryllysine (Mouse, PTO BIO, cat. no. PTM-1501, 1:2000), anti-Crotonyllysine (Mouse, PTO BIO, cat. no. PTM-502, 1:2000), nti-β-Tubulin (Mouse, Abmart, cat, no. M20005, 1:5000), anti-V5 tag (Mouse, Thermo, cat. no. MA5-15253, 1:2000), anti-Cyclophilin A (Rabbit, HUABIO, at. no. ET1703-33,1:5000). The DH10B strain was laboratory stored. It was used for plasmid amplification and protein expression. HEK293T and HeLa cells were obtained from the American Type Culture Collection (ATCC).
Plasmid construction
To incorporate MeaK in E. coli, the pSupAR-MbPylRS-MbtRNApylCUA plasmid was constructed by introducing Y349F mutation to MbPylRS. The genes encoding target proteins including EGFP and MBP-Z were cloned into the pBAD vector with a His-tag. The Amber stop codon TAG was introduced at the incorporation site by site-directed mutagenesis.
To incorporate MeaK and CrK in mammalian cells, the pNEU-chPylRS-MmtRNApylCUA plasmid was constructed by introducing Y384F mutations into the chimeric pyrrolysyl-tRNA synthetase. The genes encoding target proteins including His-tagged EGFP, TXN1 and H3, CypA with Strep-tag II, were cloned into pRK5M vector. All other eukaryotic expressed proteins carried a Flag-tag at the carboxyl terminus were cloned into the pRK5M vector. The Amber stop codon TAG or other mutations were introduced by site-directed mutagenesis.
Recombinant protein expression
For prokaryotic expression, the plasmid pBAD-MBP-Z-E24TAG was co-transformed with pSupAR-MbPylRS Y349F-MbtRNApylCUA into DH10B competent cells. The transformed cells were plated on LB (Luria-Bertani) agar plate with ampicillin (Sangon) and chloramphenicol (Sangon) and incubated overnight at 37 °C. A single colony was picked and incubated with 1 mL LB medium, which was further diluted into 50 mL LB. When OD600 reached 0.6–0.8, the cell culture was induced with 0.2% arabinose in the absence or presence of 1 mM MeaK. After incubation at 30 °C for 16 h, cell pellets were collected by centrifugation at 4000 × g for 30 min at 4 °C and stored at −80 °C.
For eukaryotic expression, HEK293T cells were seeded in two 10cm-cell culture dishes containing 8 mL of DMEM media with 10% fetal bovine serum (FBS) (Gibco) and 1% penicillin–streptomycin (P/S), and grown at 37 °C in a CO2 incubator. The plasmid pNEU-chPylRS Y384F-MmtRNApylCUA was co-transfected with pRK5M-EGFP Y151TAG, pRK5M-CypA K125TAG or pRK5M-CypA K131TAG into target cells with polyethyleneimine (1 mg/mL) in DMEM media, separately. Six hours post transfection, the cells were treated with or without 1 mM MeaK and cultured for 36 h. After incubation, the cells were washed with PBS and cell pellets were collected by centrifugation at 500 × g for 5 min at 4 °C and stored at -80 °C.
Protein purification
For the purification of His-tagged MBP-Z E24MeaK, cell pellets were resuspended in His-tag lysis buffer (50 mM Tris-HCl pH 8.0, 500 mM NaCl, 20 mM imidazole, 1% v/v Tween 20, and protease inhibitors). The cell suspension was lysed by sonication (Fisher Scientific, φ6, 30% output, 10 min, 5 s off, 3 s on) followed by centrifugation (16,200 × g, 30 min, 4 °C). The supernatant was incubated with 200 μL pre-equilibrated Ni-NTA Beads (Smart-Lifesciences) at 4 °C for 2 h. Then the beads were washed with three volumes of wash buffer 1 (50 mM Tris pH 8.0, 500 mM NaCl, 20 mM imidazole) and wash buffer 2 (50 mM Tris pH 8.0, 500 mM NaCl, 40 mM imidazole). The proteins were eluted with His-tag elution buffer (50 mM Tris pH 8.0, 500 mM NaCl, 250 mM imidazole), and the buffer was exchanged to storage buffer (50 mM HEPES, pH 7.5, 150 mM NaCl).
For the purification of CypA K125MeaK and CypA K131MeaK with Strep-tag II, cell pellets were resuspended in Strep-tag lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 1 mM EDTA, 1% NP-40 and protease inhibitors). The cell suspension was lysed by sonication (φ3, 30% output, 3 min, 5 s off, 3 s on) followed by centrifugation (16,200 × g, 30 min, 4 °C). The supernatant was incubated with 200 μL pre-equilibrated Streptactin Beads 4FF (Smart-Lifesciences) at 4 °C for 2 h. Then the beads were washed with six volumes of wash buffer I (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 1 mM EDTA) and eluted with Strep-tag elution buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 1 mM EDTA, 20 mM D-biotin), followed by buffer exchanging to storage buffer (50 mM HEPES, pH 7.5, 150 mM NaCl).
Genetic incorporation of MeaK
In mammalian cells, HEK293T cells were cultured in DMEM medium (Gibco) supplemented with 10% FBS and 1% P/S. pNEU-chPylRS Y384F-MmtRNApylCUA and pRK5M-EGFP Y151TAG were co-transfected at 70% cell confluency. Six hours post transfection, the media containing transfection complex was replaced with fresh DMEM media with 10% FBS and 1% P/S in the presence or absence of 1 mM MeaK. After incubation at 37 °C for 36 h, transfected cells were imaged. For optimization of MeaK concentration, cells were cultured under different concentrations of MeaK for 36 h for imaging.
In E. coli, pBAD-EGFP D190TAG-His-tag was co-transformed with pSupAR-MbPylRS Y349F-MbtRNApylCUA into DH10B. E. coli chemocompetent cells were seeded on LB agar plates containing ampicillin and chloramphenicol. A single colony was picked and incubated into 1 mL 2× YT medium with ampicillin and chloramphenicol at 37 °C. When OD600 reached 0.6–0.8, 500 μL cell culture was supplemented with 1 mM MeaK and 0.2% arabinose, then incubation at 30 °C for 16 h. As a negative control, 500 μL cell culture was induced with 0.2% arabinose at 30 °C for 16 h. Cell pellets were collected by centrifugation at 4000 × g for 10 min. The cells were then rinsed with 2× SDS-PAGE sample loading buffer. The samples were separated on Tricine-SDS-PAGE and immunoblotted with 1:10000 HRP-conjugated 6×His monoclonal antibody.
Flow cytometric analysis of ncAA incorporation in mammalian cells
HEK293T cells were seeded in a 12-well cell culture dish containing 1 mL of DMEM media with 10% FBS and 1% P/S, and grown at 37 °C in a CO2 incubator. The plasmid pNEU-chPylRS Y384F-MmtRNApylCUA was co-transfected with pRK5M-EGFP (Y151TAG) into target cells with polyethyleneimine (1 mg/mL) in DMEM media. Six hours post transfection, the cells were treated with or without 1 mM MeaK and cultured for 36 h. After incubation, the cells were washed with PBS and resuspended in 500 µL PBS. Flow cytometry was performed with Beckman CytoFlex and data analysis and plotting were performed with FlowJo (10.8.1).
Fluorescence response of EGFP K85TAG
HEK293T cells were seeded in a 12-well cell culture dish containing 1 mL of DMEM media with 10% FBS and 1% P/S and grown at 37 °C in a CO2 incubator. The plasmid pNEU-chPylRS Y384F-MmtRNApylCUA was co-transfected with pRK5M-mCherry-P2A-EGFP (K85TAG) into target cells with 3 µL polyethyleneimine in 1 mL DMEM media. mCherry proteins were used as an internal control, and the self-cleavable peptide allows mCherry and EGFP to separate and fold independently after translation. Six hours post transfection, the media were replaced with complete DMEM media with or without 1 mM MeaK, 10 mM NAM and 10 mM NaBu. After incubation for 36 h, the cells were imaged with a microscope. The total intensity of EGFP and mCherry were measured separately using ImageJ software and the signal of EGFP was normalized to the mCherry signal for each group. The normalized data was further analyzed by 2-tailed student’s t-test and plotted with Graph Prism 9.0.
Co-immunoprecipitation (Co-IP)
HEK293T cells were harvested after transfection for 48 h, the cells were washed 2 times with PBS, resuspended with 1 mL PBS and centrifuged (500 × g, 4 °C, 3 min) to remove the upper liquid. The samples were resuspended with 500 µL Strep-tag lysis buffer, and sonicated with Sonic Dismembrator (Fisher Scientific, φ3, 30% output, 1 min, 5 s off, 3 s on) in an ice-water bath, followed by centrifugation (21,000 × g, 30 min, 4 °C). The soluble fractions were collected and incubated with pre-equilibrated Streptactin Beads 4FF at 4 °C for 1 h with constant mechanical rotation. Then centrifuging and removing the upper liquid, and washing the samples 3 times by wash buffer I and 3 times by wash buffer II (100 mM Tris-HCl pH 8.0, 37.5 mM NaCl, 1 mM EDTA). 4× protein loading was added for Western blot analysis. The samples were separated on 10% Tricine-SDS-PAGE and immunoblotted with specific primary antibody solution overnight at 4 °C. After incubation with the matching secondary antibody, chemiluminescent detection was performed using enhanced chemiluminescence (ECL) luminous solution (Vazyme).
Mass analysis of intact proteins
Purified proteins were analyzed on mass spectra of intact proteins obtained using a Triple TOF 6600 (SCIEX) MS System equipped with an electrospray ionization (ESI) source in conjunction with SCIEX Analyst TF software. Protein samples were separated and desalted on a Aeris 3.6 µm WIDEPORE C4 LC Column (200 Å, 2.1 × 50 mm, 3.6 μm, Phenomenex). Solvent A was 0.1% formic acid in water and solvent B was acetonitrile. A constant flow rate of 0.3 mL/min was used. Mass spectral deconvolution was performed using SCIEX OS-Q software (version 2.0, SCIEX Corporation).
Sample preparation for in-solution digestion
Purified proteins were precipitated with 6 times the volume of 100% acetone 2 h at −20 °C. The protein pellets were washed twice with ice-cold acetone and air-dried. The protein pellets were resuspended in 50 mM ammonium bicarbonate and digested with trypsin (1:50) overnight at 37 °C. Then, peptides were reduced with 5 mM tris-carboxyethylphosphine (TCEP) and alkylated with 10 mM iodoacetamide (IAA) in the dark for 1 h at room temperature. Then, stopped by acidification with formic acid (FA) at a final concentration of 5% (v:v). Peptides were desalted with StageTip and dried.
Affinity purification and mass spectrometry (AP-MS)
As described previously in Co-IP assay, after washing with buffer I and II, beads were resuspended in 40 μL 8 M urea in Tris buffer. Protein was reduced with 0.4 μL of 500 mM TCEP for 15 min at room temperature before adding 0.8 μL of 550 mM IAA for 20 min at room temperature in the dark and followed by dilution to 2 M urea with 100 mM pH 8.5 Tris buffer. Proteins were digested on the beads with 1.5 μL 1 μg/μL Trypsin (YaxinBIO) at 37 °C in the dark overnight. The digestion stopped by adding FA (Formic acid) to a final concentration of 5%. The supernatant without beads was transferred to a new tube and was collected in three repetitions with 5% FA. Peptides were desalted using StageTip and dried.
HPLC-MS/MS analysis
Peptides were redissolved in 0.1% FA and separated on C18 (75 μm × 15 cm, 1.9 µm C18, 1 µm tip) with Easy-nLC 1200 system (Thermo Fisher Scientific). Using the following mobile phases: 2% ACN (Acetonitrile) incorporating 0.1% FA (Solvent A) and 80% ACN incorporating 0.1% FA (Solvent B). Samples were analyzed with a 60 min gradient at a flow rate of 450 nL/min. Peptides were ionized by electrospray ionization at +2.1 kV. Tandem mass spectrometry analysis was carried out on a Q‐Exactive HF-X mass spectrometer (Thermo Fisher Scientific) under the control of Xcalibur software (version 4.5.445.18). The mass spectrometer operated in data-dependent acquisition mode, and survey scans were obtained in a full mass scan range of 400–1400 m/z with lock mass activated. In each scan cycle, fragmentation spectra of the 20 most intense peptide precursors in the survey scan were acquired in the higher-energy collisional dissociation (HCD) mode. Full MS scan at resolution of 60,000, followed by MS/MS scans at resolution of 15,000 with an isolation width of 1.6 m/z. The AGC targets for MS1 and MS2 scans were 3 × 106 and 1 × 105, respectively, and the maximum injection time for MS1 and MS2 were 45 and 22 ms, respectively. Precursors of +1, +6, +8 or above, or unassigned charge states were rejected; exclusion of isotopes was disabled; dynamic exclusion was set to 30 s. Raw files of AP-MS were collected as three biological replicates. Other LC-MS/MS experiments were performed once.
MS data analysis
Modified peptides were identified using pFind software. pFind (version 3.2.0) search parameters: precursor mass tolerance 20 parts per million (ppm); fragment mass tolerance 20 ppm; peptide length minimum 6 amino acids and maximum 100 amino acids per chain; peptide mass minimum 600 and maximum 10,000 Da per chain. Carbamidomethyl[C] (+57.0215 Da) was the fixed modification; Acetyl[AnyN-term] (+42.0106 Da) and oxidation[M] (+15.9949 Da) were the variable modification; Trypsin as the digestive enzyme; three missed cleavage sites per chain. maximum false discovery rate (FDR) for peptides and proteins of 1%. The human protein sequences were downloaded from Uniprot. The “Open Search” mode was activated when unknown modifications were present on the peptide.
For AP-MS raw data was processed in the MaxQuant software (version 2.0.3.0), The minimal required peptide length was set to seven amino acids and both protein and peptide identifications were accepted at a FDR of 1%. Database searches were performed according to standard settings with trypsin being used, allowing 2 missed cleavages. Oxidation [M], Acetyl [Protein N-term] and Kmea[K] (+68.023 Da) were allowed as variable modifications with a maximum number of 5. Carbamidomethyl [C] was allowed as a fixed modification. The match between runs feature and the label-free quantitation function were activated. All other parameters were left at default.
Microscale thermophoresis (MST) binding assay
Purification of recombinant EGFP fusions of CypA and CypA K125MeaK. Analysis was performed in buffer (50 mM HEPES, 150 mM NaCl and 5% DMSO). 5 μL of target protein was mixed with 5 μL of 2-fold serially diluted CsA at room temperature. Mixture samples were obtained through Monolith Capillaries (Catalog No.MO-K022). Signal was measured by Monolith NT.115 (NanoTemper) with Nano BLUE detector. Curve fitting was performed by using MO.Affinity Analysis v3.0.6. Data are presented as mean ± SD. The given Kd values were calculated with a 95% confidence level from three experiments.
Flow cytometric analysis of ROS level in HeLa cells
HeLa cells were seeded in a 12-well cell culture dish containing 1 mL of DMEM media with 10% FBS and 1% P/S, and grown at 37 °C in a CO2 incubator overnight. The plasmid pRK5M-mCherry, pRK5M-CypA-WT-mCherry, pRK5M-CypA-K125TAG-mCherry and pNEU-chPylRS Y384F-MmtRNApylCUA were transfected into target cells with 3 µL PEI in 1 mL DMEM media. 6 h post transfection, the media were replaced with complete DMEM media with or without 1 mM MeaK and 4 µM CsA. After incubation for 36 h, the cells were washed 2 times with PBS. Then the cells were incubated with intracellular ROS probe DCFH-DA (OwowLife, 10 µM) for 20 min at 37 °C, and washed 2 times with PBS. The cells were then rinsed with 500 µL PBS and analyzed by Beckman CytoFlex. Positive cells (n = 5000) with the mCherry fluorescence signal were collected and then the mean FTIC-A intensity of positive cells was measured by CytoExpert software, which indicates the level of intracellular ROS, and was plotted with FlowJo 10.8.1.
In vitro and in vivo demethacrylation assay
In vitro, 400 ng purificated recombinant HDAC1 protein and 1 µg CypA K125MeaK protein was incubated in reaction buffer for 2 h at 30 °C. The reaction buffer for the deacylation was 25 mM Tris-HCL, with 150 mM NaCl, 2 mM MgCl2, 1 µM ZnCl2, 1 mM DTT. Protein loading was added for the next Western blot analysis.
In vivo, HEK293T cells were seeded in a 12-well cell culture dish. The plasmids pNEU-chPylRS Y384F-MmtRNApylCUA and pRK5M-CypA-K125TAG-Strep were co-transfected with or without plasmid pRK5M-HDAC1-Flag into target cells with 3 µL polyethyleneimine in 1 mL DMEM media. Six hours post transfection, the media were replaced with complete DMEM media with 1 mM MeaK.Then HEK293T cells were harvested after transfection for 48 h. The samples were resuspended with 500 µL Strep-tag lysis buffer, and sonicated in an ice-water bath, followed by centrifugation (21,000 × g, 30 min, 4 °C). Soluble fractions were collected and incubated with pre-equilibrated Streptactin Beads 4FF at 4 °C for 1 h with constant mechanical rotation. Then centrifuging and removing the upper liquid, and washing the samples 3 times by wash buffer I and 3 times by wash buffer II. Protein loading was added for the next Western blot analysis.
Bioinformatics analysis
Unless noted, normalization of raw read counts was done using the preprocessCore package (v.4.6.0) in R. Raw data results of maxquant output were normalized by the quantile normalization function embedded in preprocessCore, finally, the protein expression values were presented in the log2. Principal component analysis (PCA) of the data matrix using the prcomp function in R. Proteins were generated using the normalized raw data excluding individuals with protein expression deficiencies. The proteins were ranked from largest to smallest among individuals using the coefficient of variation, and the top 1000 proteins were taken as input to apply principal component analysis.
GO analysis of genes with differentially expressed proteins was performed by the R package ClusterProfiler package (v.4.0.5), a tool that analyzes functional profiles of gene and gene clusters. The protein IDs included in UniProt were converted to the corresponding gene Entrez IDs. GO terms with p-value < 0.05 were considered significantly enriched for visualization.
Statistics and reproducibility
The experiments in the Figs. 2c; 5e–g were repeated twice with similar results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The primers, protein sequences and chemical compound information data generated in this study are provided in the Supplementary Information. The mass spectrometry data generated in this study have been deposited in the ProteomeXchange Consortium under accession code PXD051167. [https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID=PXD051167]. Source data are provided with this paper.
References
Shang, S., Liu, J. & Hua, F. Protein acylation: mechanisms, biological functions and therapeutic targets. Signal Transduct. Target. Ther. 7, 396 (2022).
West, A. C. & Johnstone, R. W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Investig. 124, 30–39 (2014).
Xie, Z. et al. Metabolic regulation of gene expression by histone lysine beta-hydroxybutyrylation. Mol. cell 62, 194–206 (2016).
Tian, H. et al. Genetically encoded benzoyllysines serve as versatile probes for interrogating histone benzoylation and interactions in living cells. ACS Chem. Biol. 16, 2560–2569 (2021).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Tan, M. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146, 1016–1028 (2011).
Feldman, J. L., Baeza, J. & Denu, J. M. Activation of the protein deacetylase SIRT6 by long-chain fatty acids and widespread deacylation by mammalian sirtuins. J. Biol. Chem. 288, 31350–31356 (2013).
Bao, X. et al. Identification of ‘erasers’ for lysine crotonylated histone marks using a chemical proteomics approach. eLife 3, e02999 (2014).
Sabari, B. R. et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol. Cell 58, 203–215 (2015).
Liu, X. et al. MOF as an evolutionarily conserved histone crotonyltransferase and transcriptional activation by histone acetyltransferase-deficient and crotonyltransferase-competent CBP/p300. Cell Discov. 3, 17016 (2017).
Wei, W. et al. Class I histone deacetylases are major histone decrotonylases: evidence for critical and broad function of histone crotonylation in transcription. Cell Res. 27, 898–915 (2017).
Xu, W. et al. Global profiling of crotonylation on non-histone proteins. Cell Res. 27, 946–949 (2017).
Yu, H. et al. Global crotonylome reveals CDYL-regulated RPA1 crotonylation in homologous recombination-mediated DNA repair. Sci. Adv. 6, eaay4697 (2020).
Zhang, N. et al. Global crotonylatome and GWAS revealed a TaSRT1-TaPGK model regulating wheat cold tolerance through mediating pyruvate. Sci. Adv. 9, eadg1012 (2023).
Jiang, G., Li, C., Lu, M., Lu, K. & Li, H. Protein lysine crotonylation: past, present, perspective. Cell Death Dis. 12, 703 (2021).
Delaney, K. et al. Histone lysine methacrylation is a dynamic post-translational modification regulated by HAT1 and SIRT2. Cell Discov. 7, 122 (2021).
Bachhawat, B. K., Coon, M. J., Kupiecki, F. P., Nagle, R. & Robinson, W. G. Coenzyme A thiol esters of isobutyric, methacrylic, and beta-hydroxyisobutyric acids as intermediates in the enzymatic degradation of valine. J. Biol. Chem. 224, 1–11 (1957).
Peters, H. et al. ECHS1 mutations in Leigh disease: a new inborn error of metabolism affecting valine metabolism. Brain 137, 2903–2908 (2014).
Fitzsimons, P. E. et al. Clinical, biochemical, and genetic features of four patients with short-chain enoyl-CoA hydratase (ECHS1) deficiency. Am. J. Med. Genet. A 176, 1115–1127 (2018).
Wang, X. & Hayes, J. J. Acetylation mimics within individual core histone tail domains indicate distinct roles in regulating the stability of higher-order chromatin structure. Mol. Cell. Biol. 28, 227–236 (2008).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding N(epsilon)-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Hoppmann, C. et al. Site-specific incorporation of phosphotyrosine using an expanded genetic code. Nat. Chem. Biol. 13, 842–844 (2017).
Ding, W. et al. Computational design and genetic incorporation of lipidation mimics in living cells. Nat. Chem. Biol. 20, 42–51 (2023).
Weyh M., Jokisch M. L., Nguyen T. A., Fottner M. & Lang K. Deciphering functional roles of protein succinylation and glutarylation using genetic code expansion. Nat. Chem. 16, 913–921 (2024).
Kim, C. H., Kang, M., Kim, H. J., Chatterjee, A. & Schultz, P. G. Site-specific incorporation of epsilon-N-crotonyllysine into histones. Angew. Chem. Int. Ed. Engl. 51, 7246–7249 (2012).
Qin, F. et al. Linking chromatin acylation mark-defined proteome and genome in living cells. Cell 186, 1066–1085.e1036 (2023).
Fischer, G., Wittmann-Liebold, B., Lang, K., Kiefhaber, T. & Schmid, F. X. Cyclophilin and peptidyl-prolyl cis-trans isomerase are probably identical proteins. Nature 337, 476–478 (1989).
Liu, W. et al. Cyclophilin A-regulated ubiquitination is critical for RIG-I-mediated antiviral immune responses. eLife 6, e24425 (2017).
Schulze, C. J. et al. Chemical remodeling of a cellular chaperone to target the active state of mutant KRAS. Science 381, 794–799 (2023).
Zhao, Y. et al. TRIM5alpha restricts poxviruses and is antagonized by CypA and the viral protein C6. Nature 620, 873–880 (2023).
Lauranzano, E. et al. Peptidylprolyl isomerase A governs TARDBP function and assembly in heterogeneous nuclear ribonucleoprotein complexes. Brain 138, 974–991 (2015).
Zhu, D. et al. The Cyclophilin A-CD147 complex promotes the proliferation and homing of multiple myeloma cells. Nat. Med. 21, 572–580 (2015).
Kim, S. C. et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell 23, 607–618 (2006).
Lammers, M., Neumann, H., Chin, J. W. & James, L. C. Acetylation regulates cyclophilin A catalysis, immunosuppression and HIV isomerization. Nat. Chem. Biol. 6, 331–337 (2010).
Lee, S., Kim, S. M. & Lee, R. T. Thioredoxin and thioredoxin target proteins: from molecular mechanisms to functional significance. Antioxid. Redox Signal. 18, 1165–1207 (2013).
Yang, B. et al. Proximity-enhanced SuFEx chemical cross-linker for specific and multitargeting cross-linking mass spectrometry. Proc. Natl Acad. Sci. USA 115, 11162–11167 (2018).
Liu, C. et al. Identification of protein direct interactome with genetic code expansion and search engine OpenUaa. Adv. Biol. 5, 2000308 (2021).
Wu, T. et al. Mapping protein direct interactome of oxidoreductases with small molecular chemical cross-linkers in live cells. Redox Biol. 61, 102642 (2023).
Lee, S. P. et al. Cyclophilin a binds to peroxiredoxins and activates its peroxidase activity. J. Biol. Chem. 276, 29826–29832 (2001).
Li, Y. & Yang, L. Cyclophilin a represses reactive oxygen species generation and death of hypoxic non-small-cell lung cancer cells by degrading thioredoxin-interacting protein. Cell Cycle 21, 1996–2007 (2022).
Peng, L. et al. Redox-sensitive cyclophilin A elicits chemoresistance through realigning cellular oxidative status in colorectal cancer. Cell Rep. 37, 110069 (2021).
Wu, C. et al. Distinction of thioredoxin transnitrosylation and denitrosylation target proteins by the ICAT quantitative approach. J. Proteom. 74, 2498–2509 (2011).
Motohashi, K., Kondoh, A., Stumpp, M. T. & Hisabori, T. Comprehensive survey of proteins targeted by chloroplast thioredoxin. Proc. Natl Acad. Sci. USA 98, 11224–11229 (2001).
Chi, H. et al. Comprehensive identification of peptides in tandem mass spectra using an efficient open search engine. Nat. Biotechnol. 36, 1059–1061 (2018).
Wan, N. et al. Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome. Nat. Methods 19, 854–864 (2022).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253 (2017).
Thangudu, R. R. et al. Proteomic data commons: a resource for proteogenomic analysis. Cancer Res. 80, LB-242 (2020).
Motohashi, K., Koyama, F., Nakanishi, Y., Ueoka-Nakanishi, H. & Hisabori, T. Chloroplast cyclophilin is a target protein of thioredoxin. Thiol modulation of the peptidyl-prolyl cis-trans isomerase activity. J. Biol. Chem. 278, 31848–31852 (2003).
Dominguez-Solis, J. R. et al. A cyclophilin links redox and light signals to cysteine biosynthesis and stress responses in chloroplasts. Proc. Natl Acad. Sci. USA 105, 16386–16391 (2008).
Ghezzi, P. et al. Redox regulation of cyclophilin A by glutathionylation. Proteomics 6, 817–825 (2006).
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE 6, e18556 (2011).
Murphy, M. P. et al. Guidelines for measuring reactive oxygen species and oxidative damage in cells and in vivo. Nat. Metab. 4, 651–662 (2022).
Xuan, W., Yao, A. & Schultz, P. G. Genetically encoded fluorescent probe for detecting sirtuins in living cells. J. Am. Chem. Soc. 139, 12350–12353 (2017).
Lu-Culligan, W. J. et al. Acetyl-methyllysine marks chromatin at active transcription start sites. Nature 622, 173–179 (2023).
Peters, H. et al. Metabolite studies in HIBCH and ECHS1 defects: Implications for screening. Mol. Genet. Metab. 115, 168–173 (2015).
Hansen, B. K. et al. Analysis of human acetylation stoichiometry defines mechanistic constraints on protein regulation. Nat. Commun. 10, 1055 (2019).
Suk, T. R. & Rousseaux, M. W. C. The role of TDP-43 mislocalization in amyotrophic lateral sclerosis. Mol. Neurodegener. 15, 45 (2020).
Perkins, A., Nelson, K. J., Parsonage, D., Poole, L. B. & Karplus, P. A. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem Sci. 40, 435–445 (2015).
Olsen, J. B. et al. Quantitative profiling of the activity of protein lysine methyltransferase SMYD2 using SILAC-based proteomics. Mol. Cell. Proteom. 15, 892–905 (2016).
Liu, D. D. et al. Characterize direct protein interactions with enrichable, cleavable and latent bioreactive unnatural amino acids. Nat. Commun. 15, 5221 (2024).
Yang, T., Liu, Z. & Li, X. D. Developing diazirine-based chemical probes to identify histone modification ‘readers’ and ‘erasers. Chem. Sci. 6, 1011–1017 (2015).
Hogbom, M., Eklund, M., Nygren, P. A. & Nordlund, P. Structural basis for recognition by an in vitro evolved affibody. Proc. Natl Acad. Sci. USA 100, 3191–3196 (2003).
Acknowledgements
We thank technical assistance from LSI core facility, Life Sciences Institute (LSI), Zhejiang University. This work was supported by the Chinese National Natural Science Funds (22374128, 22074132 and 91953103 to B.Y.), the National Key R&D Program of China (2022YFF0608402 to B.Y.), the outstanding youth fund of Zhejiang Province (LR20B050001 to B.Y.), the special COVID-19 program of the Sino-German Center for Research Promotion (C-0023 to B.Y.), Open Project Program of the State Key Laboratory of Proteomics (SKLPO201806 to B.Y.).
Author information
Authors and Affiliations
Contributions
T.Y.Z. and S.Y.C. contributed equally to this work. T.Y.Z. and S.Y.C. conducted the majority of the experiments and data analysis. M.Z., H.L., X.Z., and H.S. performed biological validation experiments. T.W. purified methacrylated proteins. E.A., J.W., and H.W. chemically synthesized MeaK compound. B.K.J., H.H., X.W., K.S., and P.L. performed bioinformatics analysis and protein structure modeling. D.D.L. screen tRNA synthetase. Y.F. performed MS data analysis. Y.Y., A.A.S., and L.Z. provided advice for the biological experiments. M.W. performed experiments on biomedical samples. H.W., C.T., and Y.B. directed the project. H.W. and Y.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, TY., Chen, SY., Zhang, M. et al. Genetically encoding ε-N-methacryllysine into proteins in live cells. Nat Commun 16, 2623 (2025). https://doi.org/10.1038/s41467-025-57969-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57969-2
This article is cited by
-
Enrichable cross-linkers for mapping direct protein interactions
Genome Biology (2025)