Abstract
Amyotrophic lateral sclerosis (ALS) is a fatal and rapidly evolving neurodegenerative disease arising from the loss of glutamatergic corticospinal neurons (CSN) and cholinergic motoneurons (MN). Here, we performed comparative cross-species transcriptomics of CSN using published snRNA-seq data from the motor cortex of ALS and control postmortem tissues, and performed longitudinal RNA-seq on CSN purified from male Sod1G86R mice. We report that CSN undergo ER stress and altered mRNA translation, and identify the transcription factor CREB3 and its regulatory network as a resilience marker of ALS, not only amongst vulnerable neuronal populations, but across all neuronal populations as well as other cell types. Using genetic and epidemiologic analyses we further identify the rare variant CREB3R119G (rs11538707) as a positive disease modifier in ALS. Through gain of function, CREB3R119G decreases the risk of developing ALS and the motor progression rate of ALS patients.
Similar content being viewed by others
Introduction
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disease that manifests as a progressive muscular weakness and paralysis, leading to death within only two to three years upon symptom onset1. ALS results from the progressive degeneration of two neuronal populations involved in voluntary motor control: the glutamatergic corticospinal neurons (CSN, or upper motor neurons) in the motor cortex, and the cholinergic motoneurons (MN, or lower motoneurons), in the brainstem and spinal cord1.
Whereas the vast majority of patients are considered sporadic, about 10% have a familial history, and causative genes have been identified in more than half of these cases, the most represented being C9orf72-SMCR8 complex subunit (C9orf72), Superoxide Dismutase 1 (SOD1), TAR DNA Binding Protein (TARDBP) and FUS RNA binding protein (FUS)2. While ALS genetics is complex, causative mutations remarkably converge on recurrent dysregulated pathways including DNA repair, RNA and protein metabolism, intracellular trafficking and mitochondrial dysfunction2. This suggests that at least a subgroup of genetic disease modifiers may modulate, either positively or negatively, the same cellular pathways that accompany the degeneration of vulnerable neurons. The most recent and largest GWAS to date identified 15 ALS-associated risk loci with enriched expression in excitatory glutamatergic neurons3, a cell-type specificity that we4 and others5 recently confirmed. We thus reasoned that unravelling the deregulated cellular pathways that accompany CSN degeneration during the course of ALS may inform on putative genetic disease modifiers and therapeutic targets. Pioneering snRNAseq of post-mortem motor cortex from patients and controls recently provided molecular access to vulnerable cortical excitatory neurons and offered valuable snapshots of these neuronal populations at disease end-stage5,6. Yet, the population of CSN is altogether under-represented in the healthy human brain and further depleted during the course the disease5, and its access limited to post-mortem time point further restricts our ability to understand its disease trajectory and means to cope with disease progression.
To circumvent this limitation, we employed cross-species transcriptomics, combining snRNAseq data from ALS patients, frontotemporal dementia (FTD) patients, and healthy individuals5 to longitudinal RNAseq data of CSN purified from the motor cortex of a mouse model of ALS, the well described Sod1G86R mouse line7. These animals recapitulate early and progressive degeneration of the CSN8, in a manner that is somatotopically related to MN degeneration, similarly to what is reported in sporadic ALS patients9,10,11,12.This allowed us to hypothesize that a conserved signature of CSN degeneration exists across species, and to reason that the comparative integration of mouse and human transcriptomics could improve access to disease-relevant neuronal populations.
We report that comparative transcriptomics identified disease-vulnerable human neuronal populations in ALS patients, that primarily undergo ER stress and altered mRNA translation. We further identified the transcription factor CREB3 and its regulatory network as a resilience marker in human and mice, not only amongst vulnerable neuronal populations, but across all neuronal populations as well as other cell types. Finally, through genetic analyses, we identified a missense rare variant of CREB3 (rs11538707, p.Arg119Gly), which, through gain of function mechanisms, confers a 40% reduction in the risk of developing ALS and is associated with a slower disease progression rate in ALS patients, extending disease duration by 11.6 months.
Results
Cross-species RNAseq integration prioritizes disease-vulnerable human neuronal populations
To unequivocally identify the disease vulnerable CSN amongst human neuronal populations, we integrated the snRNAseq dataset obtained by Pineda and collaborators from 33 ALS patients (16 sporadic and 16 C9orf72), 24 frontotemporal lobar degeneration (FTLD) patients (13 sporadic and 11 C9orf72) and 17 aged-matched controls5, to the cross-species snRNAseq dataset generated by Bakken and collaborators that allowed consensus classification of cell types13 (Fig. 1a). Our analysis yielded a total of 245,143 nuclei integrated across the independent datasets allowing the identification of a total of 116 clusters (Fig. 1b). These were further hierarchically organized based on transcriptomic similarities and associated with their positioning across the cortical layers13, providing a full representation of glutamatergic excitatory neurons, GABAergic inhibitory neurons, and non-neuronal cells (Supplementary Fig. 1). To further maximize power with differential expression analysis, we used aggregated hierarchical clustering to regroup the initially identified 116 cell clusters into 30 cell populations. Noteworthily, the population of cortical layer 5 extra-telencephalic neurons (L5-ET), suspected to represent ALS-vulnerable human CSN, yielded the smallest number of neurons (~ 1000 neurons, corresponding to 0.6% of the total nuclei and 1.7% of all excitatory neurons) (Supplementary Data 1). We observed that the gene-level dispersion was much higher for low to medium expressed genes and negatively correlated with the number of nuclei sequenced per individual (Supplementary Fig. 2). Together, this indicates that rare cell types, such as degenerating CSN, may hinder differential expression analysis and discovery of potential target genes.
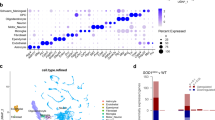
a Experimental design displaying integration of single nuclei RNAseq from ALS patients and healthy controls5, the human motor cortex13 and our database of mouse retrogradelly labelled CSN and control cells (mCSN and mCtl cells) from WT and Sod1G86R male mice. b UMAP plot of the 116 clusters identified in the human M1 motor cortex (left) and expression of the mCSN and mCtl cells enriched genes in each cluster (right). c Experimental design of the cross-species DEG enrichment analysis wherein DEG identified in mCSN were intersected with each of the 30 DEG cell populations in human ALS, FTD and healthy controls. d Cluster dendrogram and heatmaps showing enrichment of mCSN and mCtl cells geneset in each of the 30 DE human cell populations (Cellular identity) as well as enrichment of mCSN DEG in all DE ALS or FTLD cell populations (Cellular vulnerability). We observed a significant enrichment of the mCSN DEG in human excitatory neurons (two-tailed hypergeometric test: Bonferroni’s adjusted *p < 4e-04). Based on mCSN geneset enrichment and cross-species DE analysis, a cumulative score was calculated for each DE group and ranked to prioritize the most affected cell populations in ALS patients (Cellular rank). Created in BioRender. Dieterle, S. (2025) https://BioRender.com/d49e840.
To circumvent the limitation of the small representation of CSN in the human dataset, but also to gain insight into their disease trajectory, we generated a dataset of anatomically-identified, disease-vulnerable mouse corticospinal neurons (mCSN) for comparison with the human dataset. Mouse models of ALS recapitulate CNS degeneration (reviewed in ref. 14). We selected the well described Sod1G86R mouse line which overexpresses the G86R mutation of the murine gene (equivalent to G85R mutation of the human gene), abolishing SOD1 enzymatic activity and allowing to avoid using a wild-type murine Sod1 overexpressing mouse line as a control7. Sod1G86R mice present with muscle denervation and progressive motor impairments by 90 days, followed by ultimate spinal MN degeneration and death of the animals by 105–120 days of age7,8 (Supplementary Fig. 3a). We formerly thoroughly characterized the progressive degeneration of mCSN in male Sod1G86R mice that present with a loss of ≈8% as early as 60 days of age and up to ≈33% at disease end-stage8 (Supplementary Fig. 3a), and demonstrated that mCSN degeneration precedes MN degeneration8. In addition, in Sod1G86R mice, mCSN and MN degenerations were found somatotopically related8 similarly to what is reported in sporadic ALS patients9,10,11,12, and mCSN along with other L5-ET neurons were found detrimental to their downstream spinal targets, highlighting their contribution to disease progression15. In addition to mCSN degeneration, Sod1G86R mice recapitulate cortical hyperexcitability, a typical hallmark of sporadic and familial ALS16. mCSN were retrogradelly labelled from the cervical dorsal funiculus of Sod1G86R male mice and their WT male littermates (Supplementary Fig. 3b), and purified and pooled by FACS at presymptomatic (30 and 60 days), and symptomatic (90 and 105 days) ages (Supplementary Fig. 3b, c, and Fig. 1a). Non-labelled cells from cortical layers 2/3 (L2/3) were also purified from the same animals to serve as a control cellular population (mCtl cells) (Supplementary Fig. 3b, c and Fig. 1a). 64 samples were initially sequenced (n = 4 biological replicated × 2 genotypes × 2 cell types × 4 ages), 7 were excluded because they did not express the expected markers or were altered, and a total of 57 samples were further processed (mCSN: 30 d: n = 4 WT and 3 Sod1G86R; 60 d: n = 4 WT and 4 Sod1G86R; 90 d: n = 4 WT and 4 Sod1G86R; 105 d: n = 4 WT and 2 Sod1G86R; mCtl cells: 30 d: n = 4 WT and 3 Sod1G86R; 60 d: n = 2 WT and 3 Sod1G86R; 90 d: n = 4 WT and 4 Sod1G86R; 105 d: n = 4 WT and 4 Sod1G86R). Unsupervised hierarchical clustering separated mCSN from mCtl cells (Supplementary Fig. 4a and Supplementary Data 2) and indicated that a larger part of the variance observed between WT and Sod1G86R mice is attributable to mCSN (Supplementary Fig. 4b–d) and not to mCtl cells (Supplementary Fig. 4b, e, f). We reasoned that cross-species RNAseq integration could be employed to (i) identify the human neuronal populations that more closely resemble mCSN, independently of the disease condition or genotype (i.e., ‘cellular identity’, Fig. 1b, c), (ii) identify the human neuronal populations whose differential gene expression (DGE) in ALS or FTLD conditions mimics mCSN DGE between Sod1G86R and control mice, (i.e., ‘cellular vulnerability’, Fig. 1c), and (iii) prioritize human neuronal populations based on the combination of ‘cellular identity’ and ‘cellular vulnerability’ (i.e., ‘ranking’, Fig. 1c).
We first selected the top differentially expressed genes (DEG) between mCSN and mCtl cells (Supplementary Data 2) to create a mCSN geneset and its homologous human geneset (Supplementary Data 3)13. Many genes of the mCSN geneset showed a strong expression in the human L5-ET (Supplementary Fig. 5a), including BCL11 Transcription Factor B (BCL11B) and FEZ Family Zinc Finger 2 (FEZF2), known to exert critical functions in the development of cortico-thalamic and subcerebral projection neurons17, or Neurofilament Heavy Chain (NEFH) which has been shown to be enriched in mCSN18. We also observed enrichment of the gene encoding the Sodium voltage-gated channel Beta subunit 4 (SCN4B), primordial for ultrafast spiking of CSN and execution of fine motor skills19.
Second, we integrated human and mouse datasets, and identified mCSN geneset in L5-ET (Fig. 1b), cortical layers 5/6 non-projecting (L5/6-NP), cortical layer 5 intra-telencephalic (L5-IT) and cortical layers 2/3 intra-telencephalic (L2/3-IT) neurons (Supplementary Fig. 1). mCtl cell genes instead were enriched in non-neuronal populations: endothelial cells, microglia, and oligodendrocytes (Supplementary Fig. 1). This confirmed that the mCSN geneset is expressed in a large proportion of L5-ET (~ 60%) and a significant proportion (~ 40%) of L2/3-IT-1 neurons (Fig. 1d, Cellular identity heatmap). These neurons express high amounts of NEFH (Supplementary Fig. 6), typical of long-range projection neurons. This also confirmed the identity of mCtl cells as non-neuronal populations (Fig. 1d, Cellular identity heatmap). Together, this shows that several sub-populations of excitatory neurons from the human motor cortex express the gene signature of mCSN, including L5/6-NP, L5-IT and L2/3-IT populations, in addition to the expected L5-ET neurons.
Third, we intersected DEG between WT and Sod1G86R mCSN (Supplementary Data 4) with DEG between controls and ALS patients, or DEG between controls and FTLD patients for each of the 30 cell populations (Fig. 1d, Cellular vulnerability heatmap Supplementary Data 5 and 6). This analysis confirmed a significant enrichment of mCSN DEG in human excitatory neuron populations as opposed to inhibitory and non-neuronal cell populations (Supplementary Data 7, Bonferroni’s adjusted *p < 4e-04). More precisely in ALS, this identified L5/6-NP, L6b, L6-IT, L2/3-IT-1, -2 and -4, and L5-IT-1, -3 and -5 (Fig. 1d, Cellular vulnerability heatmap). The same populations were identified in FTLD, together with L5-ET, L2/3-IT-3, and L5-IT-2 and -4 (Fig. 1d, Cellular vulnerability heatmap). While the absence of enrichment of DEG in L5-ET population in the ALS dataset may seem surprising at first, it could also reflect the degeneration of this vulnerable population in these patients compared to FTLD patients. We indeed observed a non-significant reduction of L5-ET neurons in the motor cortex of ALS (Wilcoxon rank-sum test adjusted for all cell types tested, p = 0.064, Supplementary Fig. 7 and Supplementary Data 8) indicating that some neurodegenerative process is taking place in the L5-ET neurons in ALS but less so in FTLD patients.
Finally, we ranked each human cell population based on the combination of the ‘cellular identity’ and ‘cellular vulnerability’ results, which prioritized L5-ET and L2/3-IT-1 neurons as the human populations that more closely recapitulate the transcriptomic signature of diseased mCSN (Fig. 1d, Cellular rank heatmap). In whole, cross-species RNAseq integration allowed us to prioritize L5-ET and L2/3-IT-1 populations for further analyses.
WGCNA consensus on prioritized cell populations reveals conserved gene regulatory network in ALS
We next aimed at unravelling disease-associated pathways and employed weighted-gene co-expression network analysis (WGCNA) consensus across species and prioritized cell types, i.e., L5-ET, L2/3-IT-1, and mCSN (Fig. 2a). WGCNA analysis identified four mRNA modules significantly correlated with the genotype (mice) or disease condition (humans) and labelled as lightyellow, darkgreen, turquoise (upregulated) and darkgrey (downregulated) modules according to the WGCNA conventions (Benjamini-Hochberg p < 0.05; Fig. 2b and Supplementary Data 9). We then constructed a graphical network of significantly associated modules to highlight hub genes, related weight of the genes within the network and connectivity (Fig. 2c), that illustrates the independent and relative proportions of non-overlapping networks. Assessing the conservation across species further allowed us to prioritize the turquoise and lightyellow modules that respectively showed strong (Zscorepreservation > 10) and moderate preservation (5 < Zscorepreservation < 10, Fig. 2d).
a Experimental design of the WGCNA approach. b Histogram showing conserved signed correlation of WGCNA modules to ALS in prioritized cell types, corrected using the Benjamini-Hochberg method. Dashed lines highlight correlation FDR < 0.05. c UMAP gene network of the selected modules, each dot represents individual gene which size is proportional to its importance (signed kME). d Dot plot showing cross-species conservation of significant modules with strong (Zscore > 10) to moderate (5 < Zscore < 10) preservation. e GO analysis of the turquoise module with genes having signed kME (> 0.6) in the network (the red dashed line indicates an FDR < 0.05). f Violin plot showing turquoise eigengenes expression (two-way ANOVA: Finteraction group*cell = 4.88, **p < 0.01; Tukey-multiple comparisons: *p-adjusted = 0.037 and ***p-adjusted < 0.001). g Heatmap representing the turquoise module expression in healthy controls (HC) and ALS patients (one-tailed Wilcoxon rank sum test: *p-adjusted < 0.05 for significant cell-type). h GO analysis of the lightyellow module with genes having signed kME (> 0.6) in the network (the red dashed line indicates an FDR < 0.05). i Violin plot showing lightyellow eigengene expression (two-way ANOVA: Finteraction group*cell = 5.007, **p < 0.01; Tukey-multiple comparisons: ***p-adjusted < 0.001). j Violin plot showing turquoise eigengene expression in BA9 and cerebellum of ALS patients (two-tailed t-test: t = 6.74, ***p < 0.0001 and t = 1.99, p = 0.058, respectively) and in post-mortem human cortical neurons associated or not with TDP-43 loss-of-function LOF (two-tailed t-test: t = 3.20, p = 0.0079). k Turquoise module expression in Sod1G86R mCSN (two-way ANOVA: Fgroup = 45.25, p < 0.001; Tukey post-hoc: WT vs Sod1G86R at 30 d p = 0.070; 60 d p = 0.27; 90 d ***p < 0.001; 105 d p = 0.089). Boxplots show median and quartile distributions, upper and lower lines represent the 75th and 25th percentiles. l Representative images of puromycin incorporation in mCSN (Arrowheads point at L5 neurons with large CTIP2-positive nuclei) from 90d-old WT and Sod1G86R animals. Scale bar = 20 µm. m Dot plot showing puromycin incorporation quantification in mCSN. Each dot represents one animal (n = 5 WT [163 neurons], 5 Sod1G86R [199 neurons] at 30 d, and n = 8 WT [165 neurons], 8 Sod1G86R [337 neurons] at 90 d). Mixed effect model followed by one-tailed Fisher’s LSD test; p = 0.9802 at 30 d and *p = 0.0271 at 90 d; mean +/− SEM). Created in BioRender. Dieterle, S. (2025) https://BioRender.com/d49e840.
Gene Ontology (GO) analysis of the turquoise module revealed a strong enrichment in RNA-binding genes such as Eukaryotic Translation Initiation Factor 4A2 (EIF4A2), Valosin Containing Protein (VCP), Heterogeneous Nuclear Ribonucleoprotein A0 (HNRNPA0), Splicing Factor 3a Subunit 3 (SF3A3) and Tubulin Alpha 1b (TUBA1B) (Fig. 2e and Supplementary Data 10), and terms associated with mRNA translation, ribosome, and mitochondrial translation (FDR < 0.05). GO analysis on the lightyellow module highlighted terms related to ubiquitin activity such as HECT Domain E3 Ubiquitin Protein Ligase 1 (HECTD1), Ubiquitin Like With PHD And Ring Finger Domains 2 (UHRF2), Ubiquitin Conjugating Enzyme E2 K (UBE2K), and terms associated with translation initiation factor activity such as Eukaryotic Translation Initiation Factor 2B Subunit Delta (EIF2B4), Eukaryotic Translation Initiation Factor 3 Subunit L (EIF3L), Eukaryotic Translation Initiation Factor 2D (EIF2D), and Eukaryotic Translation Initiation Factor 2 Subunit Alpha (EIF2S1) that encodes a master regulator of the endoplasmic reticulum stress response (ER stress) (Fig. 2h and Supplementary Data 10). As expected, both the turquoise and the lightyellow modules were found significantly enriched in two of the three input populations, i.e., L2/3-IT-1 and mCSN (Fig. 2f and i and Supplementary Data 11), and showed a tendency to increase in L5-ET as well. Lack of significance in L5-ET could result from the smaller size of this population due to the neurodegenerative process, as opposed to the two other WGCNA input populations. Alternatively, the turquoise module could be interpreted as a resilience signature found in neurons that have survived until harvesting in mouse and postmortem stage in human. Lack of significance in the L5-ET population could thus reflect its milder resilience ability and greater susceptibility to neurodegeneration. This second hypothesis is favored by the upregulation of the turquoise module across the 28 other cellular populations (Fig. 2g and Supplementary Data 12), suggesting that this module is likely to reflect a resilience signature, as opposed to the neurodegeneration signature that we initially sought. These results suggest that this resilient signature is present in the motor cortex across all major cell types. We then wondered whether the same signature would also be observed in bulk-RNA sequencing from affected tissues. We observed that the turquoise “resilient” signature is significantly upregulated in the frontal cortex (Broadman area 9) of ALS patients compared to healthy controls (Fig. 2j, t-test ***p < 0.001) while no difference was observed in the cerebellum (Fig. 2j, t-test p = 0.058) suggesting that the resilience signature is specifically increase in ALS-affected tissues. Finally, we reveal that pathological loss of nuclear of TDP-43 function (LOF) in postmortem human cortical neurons20 associates with a significant decrease of the turquoise module (Fig. 2j, t-test **p < 0.01, Supplemental Data 13).
To test the dynamics of the turquoise module eigengene regulation over time, we took advantage of our longitudinal mCSN neurons dataset. We observed an increased expression of the turquoise module eigengenes in Sod1G86R compared to WT mCSN (Fig. 2k and Supplementary Datas 14 and 15), that was significant at the symptomatic age of 90 days (Fig. 2k, adjusted-p < 0.001). To validate our findings, we tested whether mCSN displayed altered mRNA translation, as suggested by GO analyses of the two prioritized modules. In vivo puromycin assay21 revealed a significant decrease of puromycin incorporation in motor cortex L5 CTIP2-potitive neurons from 90 day-old Sod1G86R mice compared to WT, indicative of a slower rate of protein synthesis in disease vulnerable neurons (*p = 0.0271, Fig. 2l, m). This phenotype was not present at the presymptomatic age of 30 days (p = 0.9802, Fig. 2m). Thus, our results identified a cross species transcriptomic footprint of resilience associated to ER stress and related mRNA translation.
CREB3 is a master regulator of excitatory neuron function and a protective factor from ALS
Since the turquoise module was strongly associated with ALS and preserved across species, we sought to identify potential upstream regulators based on three criteria: (i) direct binding of the transcription factors (TFs) to the turquoise module genes, (ii) WGCNA connectivity and (iii) human genetics (Fig. 3a). Intersection of the turquoise module genes with a list of TFs from the Encyclopedia of DNA Elements (ENCODE) project (Fig. 3a and Supplementary Data 16)22, identified 18 TFs (Fig. 3b). Of those, 7 displayed a strong connectivity with the genes of the turquoise module (Fig. 3b and Supplementary Data 17). To assess their putative role in ALS, we tested whether their genes accumulated rare variants in ALS and/or healthy individuals, by performing a rare missense variant burden analysis (RVBA) on a cohort of 1873 ALS patients and 3926 healthy controls which we meta-analyzed to the ALS Project MinE cohort (http://databrowser.projectmine.com/)23. This last step eventually prioritized the CAMP Responsive Element Binding Protein 3 (CREB3, Fig. 3b and Supplementary Data 18). We then assessed CREB3 expression across disease and sporadic versus familial cases, and observed that CREB3 upregulation was selective of ALS as opposed to FTLD, independently of the sporadic or familial group membership (Fig. 3b).
a Experimental design to identify transcription factors (TFs) upstream of the WGCNA turquoise module which genes were intersected with TFs from ENCODE. Each TF was then investigated using an integrative approach combining WGCNA-based connectivity, expression profile of the TF-target genes and burden of rare variants. b Cluster dendrogram and heatmap showing TF-target gene expression in each of the 30 DE cell populations in ALS patients compared to controls. Row clustering of TFs based on their target gene expression identifies two clusters (blue and orange). The orange cluster (dashed line rectangle) was further prioritized based on a higher expression level of its target genes in L5-ET. The right-sided heatmap shows WGCNA-based TF connectivity in prioritized cell populations, association of missense rare variants in each TF with ALS risk, and target genes expression profile in mCSN at a presymptomatic (30–60 d) and symptomatic stages (90–105 d). The bottom heatmap shows the prioritized TF CREB3 expression profile in ALS and FTD patients compared to controls. c Quantile-quantile plot of the meta-analyzed gene burden of rare missense variants in a cohort of 1873 ALS patients and 3926 healthy controls showing CREB3 and ALS known genes. d Locus zoom plot showing the SNP (+/− 500KB) rs11538707 (R119G) association with ALS in the discovery cohort3 and the replication of 1873 ALS cases and 3926 healthy controls. Orange dashed lines shows the genome-wide significant SNP at a p-value < 5e-08 and colored dots represent LD with the lead variant (red diamond). e Forest plot of gene burden association of CREB3 gene together with known ALS genes. Aggregation of rare missense variants in CREB3 confers a reduced risk of ALS (OR = 0.66 95%CI 0.51–0.87; Firth-logistic regression p = 2.9e-03). Genome-wide RVBA association after Bonferroni correction 0.05/14245 = 3.51.e−06. f Forest plot showing association of the missense variant rs11538707 (R119G) on CREB3 with ALS risk, compared to formerly identified associations (see Supplementary Data 25). Errors bars represent 95% confidence interval to the odds ratios. P-values were calculated using a linear-mixed model and represent uncorrected genome-wide association. Genome-wide significance is fixed at p < 5e−08.
Rare variant analysis revealed, as expected, association in all known ALS genes such as SOD1 (Odds ratio (OR) = 6.93 [95%CI: 3.19–15.06]; Pmeta = 5.02e−07), TANK Binding Kinase 1 (TBK1) (OR = 1.83 [95%CI:1.30–2.56]; Pmeta = 2.21e−04) or TARDBP (OR = 2.11 [95%CI:1.35–3.29]; Pmeta = 5.1e−04), conserved across cohorts, and highlighted CREB3, unravelling the protective effect of rare missense variants with a minor allele frequency of 0.0021 in ALS cases, compared to 0.0047 in control individuals, and which replicated across independent cohorts (OR = 0.61 (95%CI:0.46–0.81); Pmeta = 2.79e-04) (Fig. 3c–e and Supplementary Data 18). We then tested whether individual genetic variants could be associated with ALS risk through a sequenced-based genome-wide association study (seqGWAS)24. A null logistic model was fitted on the cohort of 1873 ALS cases and 3926 controls. We observed a moderate inflation of the test statistics (λGC = 1.05), and linkage disequilibrium score regression yielded an intercept of 0.983 (s.e. = 0.0068), indicating that most of the inflation was due to the polygenic signal in ALS (LDSC: ℎ2lhl2 = 0.69, s.e. = 0.0932). We were able to replicate most of the ALS GWAS variants identified by van Rheenen et al., 202223 (Supplementary Data 19) such as C9orf72 (rs2484319, Pmeta = 1.50.10−45), Unc-13 Homolog A (UNC13A) (rs12608932, Pmeta = 7.93.10−26), Kinesin Family Member 5 A (KIF5A) (rs113247976, Pmeta = 6.01.10−13) and Cilia And Flagella Associated Protein 410 (CFAP410) (rs75087725, P meta = 4.62.10−13) (Fig. 3f). In addition, we identified three loci through inverse-variance meta-analysis on chromosome 15 (rs12907456, Pmeta = 2.9.10−08), chromosome 17 (rs12907456, Pmeta = 2.75.10−08) and chromosome 9 (rs11538707, Pmeta = 3.96.10−08) (Fig. 3d–f and Supplementary Data 19). Mapping rs11538707 to the human genome revealed a missense variant p.Arg119Gly (R119G) in CREB3 which confers a 40% reduction in the risk of developing ALS (OR = 0.61, 95%CI 0.51–0.73, Fig. 3f). In our cohort we observed 37 heterozygous carriers in controls (MAF = 0.0045) and 8 heterozygous carriers in cases (MAF = 0.0021), while in the Project MinE Cohort we observed 34 heterozygous carriers (MAF = 0.0070, N = 2406) and 50 carriers in cases (MAF = 0.0038, N = 6525). In general, we observed that the CREB3R119G mutation is twice as much frequent in control than in ALS cases. Overall, our cross-species transcriptomic analysis yielded a specific gene network associated with ALS from which CREB3 appears as a major master regulator, and genetic data integration further identified CREB3 as a protective factor in ALS.
CREB3 regulatory network as a resilience marker in ALS
To determine if CREB3 is actually causing altered gene expression in vulnerable neuronal populations, we performed a series of orthogonal approaches. First, we investigated the expression of CREB3 in RNAseq data from mCSN. Both CREB3 mRNA (Fig. 4a FDR <*p < 0.05, Supplementary Data 4) and CREB3-target genes (Fig. 4b and Supplementary Data 20) were significantly upregulated in Sod1G86R mCSN. Taking advantage of the data obtained from C9orf72 mutation carriers by Pineda et al.5, we further investigated CREB3-target gene regulation across diseases and cortical areas (Fig. 4c and Supplementary Data 21). Increased CREB3 regulon expression was observed in human motor and frontal cortices from ALS patients, but not from FTLD patients (Fig. 4c and Supplementary Data 21), suggesting that CREB3 hyperactivity is disease-specific, and rather reflects neuronal resilience than neuronal vulnerability. To confirm changes of CREB3 expression at the cellular level, we performed fluorescent in situ hybridization on WT and Sod1G86R tissues at symptomatic ages, which confirmed a significant increase of CREB3 mRNA in L5 Fezf2+ neurons (Fig. 4d, p = 0.0255 and Supplementary Fig. 8). In a recent report, Moya and collaborators identified within the cortical L5 neurons of the SOD1G93A mouse model of ALS, a subpopulation of disease vulnerable neurons projecting to the spinal cord and expressing GPRIN Family Member 3 (Gprin3), and a subpopulation of disease-resistant neurons projecting to the pons and expressing Collagen Beta(1-O)Galactosyltransferase 2 (Colgalt2)25. Leveraging these published TRAPseq data, we observed a significant increase of the turquoise module and of CREB3-target genes in disease vulnerable Gprin3-positive neurons compared to Colgalt2-positive neurons and whole motor cortex (Supplementary Fig. 9a–c). To test the extent of our findings in other ALS-vulnerable neurons, we investigated the recently published snRNAseq database of SOD1G93A spinal cord26, and observed a significant upregulation of the turquoise module and of CREB3-target genes selectively in MN, compared to astrocytes, microglia, endothelial cells, and other neuronal populations (Supplementary Fig. 9d–f). We also observed an enrichment of CREB3L2, a gene of the CREB3 family in human and mouse spinal MN27,28 (Supplementary Datas 23 and 24) (Supplementary Fig. 9g–l).
a Violin plot showing increased CREB3 mRNA expression in pre- and symptomatic Sod1G86R mCSN (edgeR-QL F-test, *FDR = 0.028) b Violin plot showing CREB3-target gene expression in pre- and symptomatic Sod1G86R mCSN of (two-tailed permuted-p: ***p = 0.00019). The boxplots show median and quartile distributions, the upper and lower lines representing the 75th and 25th percentiles. c Heatmaps showing increased CREB3-target gene expression in the motor and frontal cortices of ALS and FTLD patients versus controls (*posthoc-Tukey significance in both tissues after Bonferroni correction 0.05/40 = *p < 0.001). d Representative images of Creb3 and Fezf2 mRNA expression in L5 of the motor cortex of 90d-old WT and Sod1G86R mice (left, scale bar = 20 µm), and dot plot showing Creb3 probe integrated intensity quantification (right), each dot representing one neuron (98 neurons from n = 1 females and 4 WT males, and 112 neurons from n = 2 females and 3 Sod1G86R males; data are presented as mean values +/− SEM; two-tailed nested i-test; *p = 0.0255). e Representative images of CREB3 and CTIP2 immunoreactivity in L5 of the motor cortex of end-stage Sod1G86R mouse and WT littermate (left, scale bar = 20 µm), and dot plot showing CREB3 immunoreactivity quantification in CTIP2+ neurons (right), each dot representing one neuron (532 neurons from n = 2 females and 4 WT males, and 474 neurons from n = 2 females and 4 Sod1G86R males; averaged age at perfusion = 102 d for WT, 103 d for Sod1G86R; data are presented as means +/− SEM; two-tailed nested t-test; p = 0.4161). f Correlation between CREB3 protein levels averaged for each Sod1G86R animal and their respective disease duration (n = 2 females and 4 Sod1G86R males; one-tailed Pearson correlation, r2 = 0.5753; *p = 0.0402). g Western blot revealing CREB3 protein in post-mortem motor cortex extracts from sporadic ALS patients and controls (left; Source data are provided as a Source Data file), and CREB3 protein level quantification (right; n = 3 controls and 3 ALS patients; data are presented as means +/− SEM; one-tailed t-test; *p = 0.0369). Created in BioRender. Dieterle, S. (2025) https://BioRender.com/d49e840.
To assess the protein expression of CREB3, we performed immunostaining on WT and Sod1G86R tissues at symptomatic ages. This demonstrated an overall increase of CREB3 protein in L5 CTIP2+ neurons of the motor cortex of mutant animals, with a high heterogeneity across samples (Fig.4e, p = 0.4161). To test whether this heterogeneity may be associated to neuroprotection, we assessed CREB3 protein levels in Sod1G86R animals in regard to their disease duration, calculated as the time between disease onset, when the animals stopped gaining weight, and their survival. We observed that animals that expressed the highest levels of CREB3 protein displayed increased disease duration (Fig. 4f, r2 = 0.5753, p = 0.0369). Of note, other ER-stress related proteins such as Activating Transcription Factor 4 (ATF4), Activating Transcription Factor 6 (ATF6) or the phosphorylated form of Eukaryotic Translation Initiation Factor 2A (Phospho-eI2Falpha) were found decreased in L5 CTIP2+ neurons of the motor cortex of symptomatic mutant animals (Supplementary Fig. 10). Finally, we sought for CREB3 protein expression in the motor cortex of ALS patients using Western blot, and revealed increased CREB3 expression in postmortem samples of patients versus controls (Fig.4g, p = 0.0369), in accordance with CREB3 increased expression across various neural cell types (Fig. 2g). Overall, the data confirm that mCSN and human excitatory neurons display increased CREB3 mRNA and CREB3 protein expression, associated with the activation of CREB3 regulatory networks in ALS, and shows that this signature characterizes disease-vulnerable neuronal populations.
CREB3 variant slows disease progression through gain of function
To further test whether increased transcriptional activity of CREB3-regulated genes would confer neuroprotection in ALS patients, we first identified, using CREB3-targeted chromatin immunoprecipitation and co-expression analysis, the CREB3 “regulon” which contains genes that would reflect CREB3 activity in tissues (Fig. 5a). We show that CREB3 regulon activity in the motor cortex (Fig. 5b, c) and in the blood of ALS patients (Fig. 5d, e) positively correlated with survival, and demonstrate an increased survival of ~15 and ~22 months respectively (Fig. 5c, e). The data highlight CREB3 regulatory network as a resilience marker in ALS.
a Schematic of CREB3 regulon identification from RNAseq brain and blood encompassing CREB3-target genes identified through Chip-seq and co-regulated genes identified through weighted gene co-expression analysis. b Scatter plot showing positive correlation between CREB3 regulon activity in the motor cortex of ALS patients and survival (two-tailed Pearson R2 = 0.28; **p = 0.0088). c Survival curves showing disease duration of ALS patients with high (~ 45 months) versus low (~ 30 months) brain CREB3 regulon activity (multivariate cox-ph survival: HR = 2.31 (95%CI:1.28–4.16); **p = 0.0053). d Scatter plot showing positive correlation between CREB3 regulon activity in the blood of ALS patients and survival (two-tailed Pearson R2 = 0.337, t = 3.47; ***p = 0.00076). e Survival curves showing disease duration of ALS patients with high ( ~ 80 months) versus low ( ~ 58 months) blood CREB3 regulon activity (multivariate cox-ph survival: HR = 1.64 (95%CI:1.04–2.56); *p = 0.03). f ALS-FRS slope progression in rs11538707-A and rs11538707-G carriers. g Violin plot showing a slower motor progression rate in rs11538707-G carriers (mean ALS-FRS slope = −0.50 pts/months) compared to rs11538707-A carriers (mean ALS-FRS slope = −0.88 pts/months) (*two-tailed permuted-p = 0.033, Cohen’s d effect = 0.27). h Violin plots showing disease duration of rs11538707-G carriers compared to the rs11538707-A carriers (mean disease duration ~ 55.5 months versus ~ 43.9 month; *two-tailed permuted-p = 0.041, Cohen’s d effect = 0.36). Boxplots show median and quartile distributions, the upper and lower lines representing the 75th and 25th percentiles. i Genome browser Chip-seq track showing CREB3 binding on SEC24A promoter (light orange highlight). j Design of CREB3 and CREB3-R119G expression vectors and SEC24A-GLuc-ON reporter plasmid transfected in HEK293 cells (left), and violin plots (right) showing increased SEC24A promoter activity in CREB3-transfected cells compared to control empty vector (one-way Nested ANOVA: F2–15 = 45, p < 0.0001; Bonferroni’s test: ***p < 0.0001), further enhanced by the R119G missense mutation (Nested ANOVA: F2–15 = 45, p < 0.0001; Bonferroni’s test: **p = 0.0068). Created in BioRender. Dieterle, S. (2025) https://BioRender.com/d49e840.
Finally, we asked whether the R119G variant would also act as a disease modifier. Since we observed a higher allele frequency in control (minor allele frequency = 0.0047) compared to patients (minor allele frequency = 0.0021), phenotyping characterization of rs11538707 carriers in ALS patients is limited. However, leveraging a large cohort of whole-genome sequenced datasets, we show that carriers of the rare minor allele rs11538707-G have a slower progression rate characterized by a slower decline in ALS-FRS score over time (mean ALS-FRS slope = −0.50 pts/month) compared to rs11538707-A allele (mean ALS-FRS slope = −0.88 pts/month) (Fig. 5f g, *permuted-p = 0.033, Cohen’s d = 0.27). To test whether CREB3R119G would increase survival, we took advantage of the largest whole genome sequencing cohort of ALS patients to date (N = 7089). To maximize power and prevent bias due to large unbalanced ratio of carriers (N = 43) and non-carriers (N = 7046), we performed random down sampling of non-carriers and repeated multivariate cox proportional analysis for each permutation which leads to a trend to increase survival in CREB3R119G carriers (cox-ph HR = 0.72 (95%CI; 0.51–1.01), p = 0.066) (Supplementary Fig. 11a, b). Also, we went on to compare mean disease duration in a univariate framework and we observed that the carriers of the rs11538707-G show a significant increase mean survival of ~ 55.5 months compared to the rs11538707-A carriers whose mean survival is 11.6 months shorter (mean disease duration ~ 43.9 month) (Fig. 5h, *permuted-p = 0.041, Cohen’s d = 0.36). Overall, this data confirms that the R119G mutation has a protective role in ALS and demonstrates that ALS patients that do carry the variant have a significantly slower rate of motor progression and increased mean disease duration.
Finally, to test whether the R119G mutation could modulate CREB3 transcriptional activity, we designed a reporter-based assay. We identified SEC24 homolog A, COPII coat complex component (SEC24A) as a canonical CREB3-target gene with multiple CRE-responsive elements in its promoter region (Fig. 5i), and generated a reporter plasmid by placing the luciferase gene downstream of the SEC24A promoter (Fig. 5j). Co-transfection of a wild type CREB3 expression plasmid increased SEC24A promoter activity by a factor of 2, confirming SEC2A4 as a bona fide CREB3-target gene (Fig. 5j and Supplementary Data 22). Co-transfection of a CREB3 R119G expression plasmid further enhanced the SEC24A promoter activity by 48% (Fig. 5j). Thus, the CREB3 R119G variant confers protection against ALS via a higher transcriptional activation of its target genes.
Discussion
In this study, we tested the possibility that at least a subpart of genetic disease modifiers of ALS may directly influence the molecular pathways selectively activated in vulnerable neurons as the disease progresses, and concentrated on CSN for their selective vulnerability to ALS and their excitatory glutamatergic neuron identity3,4,5. We drew inspiration from work carried out in the fields of evolution and development13,29,30,31, and implemented comparative cross-species transcriptomics from post-mortem human snRNAseq and anatomically defined mCSN RNAseq from the Sod1G86R ALS mouse model. We identified, as expected, a strong similarity between mCSN and human L5-ET neurons, which include Betz cells32, but also with human L2/3-IT-1, a population which, noteworthily in non-human primates, is vulnerable to degeneration in late-stage Alzheimer’s disease33. Using weighted gene co-expression analysis of mCSN, L2/3-IT-1 and L5-ET neurons, we identified a conserved dysregulated gene network with a strong enrichment of genes associated with perturbed translational machinery and ER stress, a signature typical of neurodegeneration34, and repeatedly reported in ALS models and patients35,36,37,38,39. Whereas ER stress and unfolded protein response (UPR) were expected in disease-vulnerable neuronal populations, we here demonstrate their presence across populations of excitatory neurons, indicating that in ALS there is no cell-type specific transcriptomic signature selective of disease-vulnerable neurons, but rather a conserved dysregulation of gene networks present in broad populations of cortical neurons. This result could be interpreted as an intrinsic bias of the purification approaches. Indeed, all individual nuclei that were purified from post-mortem brain tissues arose from cells that were present at the moment when the individual passed away, whether they came from so-called disease-vulnerable or disease-resistant populations. Similarly, in the mouse, only cells that were alive at the time of harvesting could be purified and contribute to the pooled cellular population. Therefore, if the data could potentially reflect what will eventually make a neuron prone to die, as we initially planned to unravel, they might as well reflect what had made a neuron able to survive.
By integrating single nuclei transcriptomics with human genetics and epigenetics, we further identified a set of 6 transcription factors whose target gene expression was increased in excitatory neurons of ALS patients. Using gene and variant based analysis, we further prioritized CREB3. CREB3 and members of the CREB3 family (CREB3, CREB3L1, CREB3L2, CREB3L3, CREB3L4) belong to the large bZIP family of transcription factors, which regulates the expression of a large variety of genes involved in lipid metabolism, development, and protein secretion40. CREB3 family of transcription factors are single-pass membrane proteins anchored in the ER membrane, with a long cytoplasmic N-terminal end serving as TF upon cleavage, and a smaller C-terminal end localized within the ER lumen, that contributes to the detection and adaptation to ER stress40. Release of the N-terminal part from the ER membrane and translocation to the nucleus is permitted upon cleavage by Site 2 and Site 1 proteases (S2P and S1P)40,41. Within its family, CREB3 displays the highest expression in the CNS, not only in neurons but also in astrocytes, oligodendrocytes, microglia and endothelial cells42. Upon axonopathy of sensory neurons, CREB3 was demonstrated to be released from axonal ER, retrogradely transported to the soma and involved in axonal regeneration through upregulation of the UPR and cholesterol biosynthesis43,44,45. CREB3 target genes include Homocysteine-induced ER protein (HERP) and ER degradation enhancing alpha mannosidase-like protein 1 (EDEM1) via the unfolded protein response elements46,47, Period homolog 1 (PER1), Dopamine decarboxylase (DCD), FK506 binding protein (FKBP5) via the glucocorticoid responsive element48, as well as Peroxisome Proliferator-Activated Receptor Gamma Coactivator 1-Alpha (PPARGC1A) via the cAMP responsive element (CRE)49. We identified previously described CREB3 target genes such as Cyclin F (CCNF), FUS, Heterogeneous Nuclear Ribonucleoprotein A1 (HNRNPA1) and TBK1, that are particularly relevant to ALS2.
CREB3 is composed of a transcriptional activation domain (AA 1–92) that includes two LXXLL motifs and a HCFC1-binding motif, a bZIP domain (AA 150–213), and a transmembrane domain (AA 231–247). In silico analyses reveal that the rs11538707-G variant is not predicted to be deleterious, and that the amino acid R119 has a high per-residue confidence score50. Thus, if the SNP we identified falls into the N-terminal part of the protein that displays the function of transcription factor, it does not fall onto an identified domain, and one could simply speculate that the R119G mutation might affect the conformation of the protein, enhancing its transcriptional activity as we demonstrated with the SEC24A promoter activity assay. A recent study in iPSCs-derived MN showed an increase of nuclear accumulation and phosphorylation of CREB1 (pCREB1S133) in ALS-patients-derived cultures in comparison with controls, followed by time-dependent depletion of pCREB1S133 which paralleled the synaptic impairment and preceded neuronal loss, indicative of an initial activation of a neuroprotective signaling that could not be sustained over time51. Thus, one could speculate that, in stressed and disease-vulnerable long-range projection neurons, increased CREB3 activity may represent a protective mechanism allowing neurons to cope with disease-induced cellular changes, and contribute to slow-down the neurodegenerative process52. The correlation between CREB3 expression and survival of the Sod1G86R mice (i.e., still present at symptomatic ages), and the correlation between CREB3 regulon expression in the brain of ALS patients and disease duration, support this hypothesis which will have to be demonstrated in the future. In addition, we show that ALS patients carrying the protective allele rs11538707-G have a significantly longer disease duration compared to rs11538707-A carriers (~ 55.5 vs ~43.9 months), suggesting a neuroprotective effect of the p.Arg119Gly mutation through a CREB3 gain of function. Previous studies have shown the critical role of regulated intramembrane proteolysis of ER-anchored transcription factor such as CREB3 in viral replication53, and carriers of rs11538707-G protective allele were shown to have a significantly increased risk of genital herpes infection54 highlighting the role of CREB3 in antiviral response and immunity.
The most recent and largest GWAS in ALS to date identified numerous genes associated with ALS risk, but very few disease modifiers3, such as the G4C2 expansion in C9orf72 gene and TBK1, both associated with shorter survival3. Yet, it is noteworthy that, similarly to CREB3, both genes have important functions in the immune response to pathogens55 and are highly expressed in immune brain cells56. More recently, using rare variant association analyses applied to whole genome sequencing data, Eitan et al. identified rare variants in the Interleukin 18 Receptor Accessory Protein (IL18RAP) 3′UTR in non-ALS genomes, associated with a fivefold reduced risk of developing ALS57. The authors further demonstrated that the variant IL18RAP 3′UTR attenuated NF-κB activity in microglia57. It is therefore tempting to speculate that the immune-nervous system interaction could be the key driver of disease progression in ALS. Together with IL18RAP 3′UTR57, CREB3 adds to the very short list of protective variants against ALS.
Our work has several limitations, first we undertook a cross-species comparison of vulnerable CSN in ALS while leaving aside MN that are also known to degenerate. While we do observe that CREB3 target genes are increased in MN of Sod1G93A, we were unable to truly identify single-cell specific alterations at the pseudobulk level in human ALS patients-derived MN. However, such analysis is difficult to perform as human MN are known to be rare and difficult to sample28. Second, we have shown that CREB3-targets genes are part of a resilience signature which is stronger in ALS-affected tissues. However, we have yet to understand why some CSN/MN tend to resist to degeneration while other do not. We hypothesize that differential CREB3 activity could be part of this differential vulnerability. Third, we have shown that increased CREB3 activity in central and peripheral tissue is positively correlated with ALS survival. Besides having only correlative conclusions, it is here unlikely that increased CREB3 activity is driven by the rare genetic variation CREB3R119G that we identified in our study because of the low sample size of the cohort. Therefore, increased CREB3 activity could be here associated with non-genetic factors that remained to be elucidated. Finally, we have shown that CREB3R119G is associated with an increased mean disease duration. We did not observe an increased survival for CREB3R119G when fitted using multivariable cox regression. Cox-regression tends to show little power when testing rare variants due to the unbalanced carriers/non-carriers’ ratio. To overcome this limitation and maximize power, we randomly down-sampled non-carriers and repeated the cox proportional analysis. However, we observed a strong heterogeneity of overall survival in non-carriers that is dependent on the country of origin of ALS patients (Supplementary Fig. 11c and Supplementary Data 25). Thus, we believe that random down sampling drawn from a highly heterogenous population can strongly impede survival association which is especially true for rare variants that tend to show population-specific effects58.
Overall, through integrative cell-type specific comparative analysis of mouse and human transcriptomics, combined to large scale genetic and epigenetic data, we identified CREB3 as a protective factor in ALS, that confers neuroprotection through a gain of function mechanism. Future studies will be needed to understand the role of CREB3 in neurons. As of now, our data suggest that the differential cellular vulnerability of CSN is associated with their intrinsic properties of long-range projection neurons, and that boosting CREB3 activity could represent an interesting therapeutic strategy to prevent disease onset or slow-down disease progression.
Methods
mCSN RNAseq
Animals
All animal experiments were performed under the supervision of authorized investigators and approved by the local ethical committee of Strasbourg University (CREMEAS, agreements #00766, #1534, and #28521). BAC transgenic mice with the G86R murine Sod1 missense mutation7 were obtained from the animal facility of the Faculty of Medicine, University of Strasbourg. Mice received water and regular rodent chow ad libitum. Sod1G86R animals were followed daily, and disease progression was rated according to a clinical scale going from score 4 to 0, as we previously described59.
Retrograde labelling of mCSN and FACS purification of mCSN and mCtl cells
Twenty five-day-old WT and Sod1G86R male mice were deeply anesthetized with an i.p. injection of Ketamine (Imalgène 1000®, Merial; 120 mg/kg body weight) and Xylazine (Rompun 2%®, Bayer; 16 mg/kg body weight) and placed on a heating pad. A laminectomy was performed on C3-C4 cervical vertebrae and the animals were positioned below an injector (Nanoject II, Drummond Scientific, PA) mounted on a micromanipulator. A pulled glass capillary loaded with Green IX Retrobeads (Lumafluor) was used to puncture the dura and lowered to the dorsal funiculus. Five pressure microinjections of 23 nl were performed on each side of the dorsal funiculus. 30, 60, 90, 105-day-old injected mice were deeply anesthetized with by an i.p. injection of Ketamine/Xylazine (120 mg/kg; 16 mg/kg) before decapitation. The brains were sectioned in a stainless-steel coronal brain matrix (Harvard Apparatus, MA), and 1 mm-thick sections were transferred under a fluorescence SMZ18 microscope (Nikon). Cortical layer 5 and layers 2–3 from same animals were microdissected from 4 coronal sections of 1 mm each, spanning the entirety of the CSN population, and collected in separate tubes filled with iced HABG (Hibernate A (BrainBits UK), B27, Glutamax (Gibco) and 0.1 N NaOH). The microdissected tissues were enzymatically digested with 34 U/ml papain at 37 °C for 30 min. Cells were mechanically dissociated by gentle trituration in iced HABG, filtered through 70 μm cell strainer (BD Falcon) and subjected to density centrifugation through a three-density step gradient of Percoll (Sigma, MO) as previously described60. Upon centrifugation, the cell pellets were resuspended in cold 0.01 PBS and fixed with 70% EtOH for 30 min at 4 °C. Fixed cells were centrifugated to eliminate EtOH and resuspended in 0.01 M PBS complemented with RNAse inhibitors (Promega). Microsphere-labelled mCSN were purified using the FACS Aria II (BD Biosciences), based on their fluorescence, size and granularity. Unlabelled control cells were purified by FACS, using the same size and granularity as for the mCSN in an attempt to collect cortical upper layer callosal-projection neurons. However, RNAseq analyses revealed that unlabeled control cells had a glial and endothelial identity. Approximately 2000 CSN and exactly 2000 control cells were collected from each adult mouse brain and used as individual biological replicates for RNA sequencing.
RNA sequencing of purified mice CSN and Ctls cells
Full length cDNA was obtained using the SMART-Seq v4 Ultra Low Input RNA kit for Sequencing (Clontech, CA) according to the manufacturer’s instructions. 11 cycles of cDNA amplification were performed using the Seq-Amp polymerase. 600 ng of pre-amplified cDNA were then used as input for Tn5 transposon tagmentation by the Nextera XT kit (Illumina) followed by 12 cycles of library amplification. Following purification using Agencourt AMPure XP beads (Beckman Coulter), the size and concentration of the libraries were assessed on an Agilent 2100 Bioanalyzer. The libraries were then loaded in the flow cell at a concentration of 3 nM, clusters were generated by using the Cbot and sequenced on the Illumina HiSeq 4000 system as paired-end 2 × 50-base reads, following Illumina’s instructions. Raw reads were mapped to the mouse reference genome GRCm38 with STAR version 2.7.061 and default parameters using Ensembl gene annotations (version 87). Gene-level abundance estimates were estimated using the option–quantMode geneCount in STAR. Raw counts were imported into R.4.3.1 and batch effects were removed on raw counts using the Combat-Seq method62. Combat-seq uses a negative binomial regression method that retains the integer nature or raw counts. Batch-corrected counts were then processed following the the ARMOR workflow63. The ARMOR workflow uses the edgeR-QL framework which provides more accurate type I error rate control, as it accounts for the uncertainty of the dispersion estimates. After visual inspection of sample clustering using PCA and definition of DEG contrast, differential expression was performed between presymptomatic and symptomatic Sod1G86R mice compared to controls for both cell types. Significance was fixed at FDR < 0.05.
Human single nucleus RNAseq (snRNAseq) data analysis and cluster annotation
SMART-seq v4
Raw read (fastq) files from the Allen Brain Atlas13 were aligned to the GRCh38 human genome sequence (Genome Reference Consortium, 2011) with the RefSeq transcriptome version GRCh38.p2 (RefSeq, RRID SCR_003496, current as of 13 April 2015) and updated by removing duplicate Entrez gene entries from the gtf reference file for STAR processing as described previously13.
10× Chromium RNA sequencing (Cv3)
Raw FASTQ files were downloaded from Pineda et al.,5 and the Allen Brain Atlas13 and aligned to the pre-mRNA annotated human reference genome GRCh38 using Cell Ranger v4.0 (10× Genomics, Pleasanton CA) except for substituting of the curated genome annotation used for SMART-seq v4 quantification. Introns were annotated as ‘mRNA’, and intronic reads were included to quantify expression. Quality control criteria were used as previously described13, so that for Cv3, criteria were: more than 500 (non-neuronal nuclei) or more than 1000 (neuronal nuclei) genes were detected and doublet score was less than 0.3.
Clustering of snRNA-seq data
Nuclei were grouped into transcriptomic cell types using an iterative clustering procedure64. Read counts were summed, and log2-transformed expression was centered and scaled across nuclei. Clusters were identified with Louvain community and pairs of clusters were merged if either cluster lacked marker genes. Clustering was applied iteratively to each subcluster until clusters could not be further split and robustness was assessed by repeating iterative clustering 100 times for random subsets of 80% of nuclei. Consensus clusters were defined by iteratively splitting the co-clustering matrix. The clustering pipeline is implemented in the R package scrattch.hicat v0.0.22 with marker genes defined using the limma package; the clustering method is provided by the ‘run_consensus_clust’ function (https://github.com/AllenInstitute/scrattch.hicat). Clusters were curated based on quality-control criteria or the expression of markers genes13.
To establish a set of human consensus cell types across all three datasets, we performed a separate integration of snRNA-seq technologies on the major cell classes (glutamatergic, GABAergic, and non-neuronal) as described previously13.
Each expression matrix was log2(CPM + 1) transformed then placed into a Seurat object and variable genes were determined by down sampling each expression matrix to a maximum of 300 nuclei per scrattch.hicat-defined with n set to 20, to generate a list of up to 20 marker genes per cluster. The union of the Cv3, Cv3-ALS and SSv4 gene lists were then used as input for anchor finding, dimensionality reduction, and Louvain clustering of the full expression matrices. Louvain clustering was performed to over cluster the dataset to identify more integrated clusters than the number of scrattch.hicat-defined clusters. For instance, glutamatergic neurons had 30, 47, and 48 scrattch.hicat-defined clusters, 110 overclustered integrated clusters, and 38 final human consensus clusters after merging for Cv3 and SSv4 datasets, respectively. To merge the over clustered integrated clusters, up to 20 marker genes were found for each cluster to establish the neighborhoods of the integrated dataset.
Cell-type specific differential expression
Differentially expressed genes (DEGs) for a given species were identified by using Seurat’s FindAllMarkers function with a Wilcox test and comparing each cluster with every other cluster under the same subclass, with logfc.threshold set to 0.7 and min.pct set to 0.5. The union of up to 100 genes per cluster with the highest avg_logFC was used. The average log2 expression of the DEGs was then used as input for the build_dend function from scrattch.hicat to create the dendrograms.
Cell type-specific pseudo-bulk DGE groups were built based on hierarchical clustering and Euclidean distance between each cluster leading to 30 DGE cell groups (see Fig. 2). For each of the DGE cell group, differential expression analysis was performed as described in the ARMOR workflow63 for sufficiently abundant cell types using age, sex, and disease group as design covariates and gene-wise single-cell-level variance as weights for the linear model.
CSN gene signature in human snRNAseq
A geneset of GFP-positive cells was built based on DGE analysis of CSN vs and control cells in mouse. Our final set of ~50 genes was used as an input to AUcell65 to evaluate the distribution of AUC scores across all the cells and explore the relative expression of the signature. The function AUCell_exploreThresholds() was used to determine the minimum AUC values where cells are considered to express the CSN geneset. For each cluster, the proportion of cells expressing the CSN geneset was calculated.
To identify overlap between mouse and human snRNAseq gene signatures, cell-type specific DGE genes were intersected with mCSN DEG genes, and significant overlap was calculated using an hypergeometric test with the phyper() function in R.
Cross-species consensus weighted-gene coexpression network analysis (WGCNA)
To facilitate comparison across species, mouse gene identifiers were re-annotated with human Ensembl gene orthologs using biomaRt, an R interface with the Biomart database (www.biomart.org)66. Only identifiers that were common to both human and mouse meta-sets were retained. A consensus network represents a single network arising from multiple sources of data constructed from the weighted average of correlation matrices from both the human and mouse in this study. By definition, consensus modules are the branches of a clustering tree developed from a consensus gene dissimilarity, comparable to the single-network approach; consensus modules contain genes that are closely related in both networks, i.e., the modules are present in both networks. After scaling the network (consensus scaling quantile = 0.2), a threshold power of 14 was chosen (as the smallest threshold resulting in a scale-free R2 fit of 0.9) and the consensus topological matrix was created as follows: consTOM <- consensusTOM(multiExpr,checkMissingData = TRUE,maxBlockSize = Inf, randomSeed = 12345, corType = “pearson”, maxPOutliers = 0.05, quickCor = 0, pearsonFallback = “individual”, power = 14, networkType = “signed”, TOMType = “signed”, networkCalibration = “full quantile”, calibrationQuantile = 0.95,sampleForCalibration = TRUE, sampleForCalibrationFactor = 5000). The consensus tree was then built and the modules identified with the function cutTreeDynamic() with the default parameters and the minModuleSize (= 30). This approach identified 36 modules for which correlation to phenotype (ALS) and adjusted-pvalue (FDR) were calculated and intersected between mouse and human to identify conserved directionality and association. This approach led to the identification of a final set of 20 modules. Module eigengenes (MEs) were used for module–trait association analysis, differential eigengene network analysis, and for DGE analysis. Difference in expression across trait groups was tested using a Kruskall–Wallis one-way analysis of variance. A gene’s module membership (k ME) is defined as the Pearson correlation between each gene and each ME; genes with high k ME values were considered “hub” genes and were highly co-expressed within a subnetwork. Module preservation statistical tests were used to assess how well network properties of a module in one reference data set were preserved in a comparator data set (modulePreservation function in WGCNA). Preservation statistics are influenced by a number of variables (module size, network size, etc). A composite preservation Z-score (Z summary) was used to define preservation relative to a module of randomly assigned genes where values 5 < Z < 10 represent moderate preservation, while Z > 10 indicated high preservation. Genes in each network module were characterized using EnrichR (version 1.2.5)67, and we considered a term to be significant for FDR < 0.05. Genes network modules were constructed using metacells transformed data aggregated by cell class. For glutamatergic metacells matrices, we calculated signed kMEs for each gene and each module identified in the WGCNA analysis. The topological matrix was filtered to contain selected hub genes (2 gene/module) and was used an input to UMAP dimensionality reduction. Finally, each dot size (genes) was scaled to the signed kMEs for the corresponding module.
Violin plot of eigengenes expression
For each module and corresponding cell type, module eigengenes was calculated in control and ALS conditions using the moduleEigengenes() function. Comparison across conditions was calculated using a one-way analysis of variance and considered significant when FDR < 0.05.
ENCODE Chip-seq transcription factors
A list of 281 transcription factors and their bed files were downloaded from the ENCODE consortium (https://www.encodeproject.org/). Bed files were intersected with TSS annotations from the GRCh38 version and peaks were annotated to the closest TSS genes.
A final list of 278 TFs was intersected with genes identified in the WGCNA turquoise module, which lead to a final set of 18 TFs candidates. For each of TFs candidate, target gene Z-scores were average for each cell type and cluster using the hclust() function in R. CREB3 target gene Z-scores were averaged across cell types and compared between FTD and ALS patients.
Genetic data processing and association
Association testing and meta-analysis
Whole genome sequencing from 1945 ALS patients and 3950 controls were processed as described above. Duplicate individuals were removed (king-cutoff = 0.084). Population structure was assessed by projecting 1000 G principal components (PCs) and outliers from the European ancestry’ population were removed (> 4 SD on PC1-4). Finally, samples in common between the individual genotype data and van Rheenen’s study3 were identified using the checksum program id_geno_checksum and were removed from our analyses. In total, 1873 ALS cases and 3926 controls pass quality check analysis and were used for rare variant burden and genome-wide association analyses (GWAS). After quality control, a null logistic mixed model was fitted using SAIGE with principal component (PC)1–PC10 as covariates. The model was fit on a set of high-quality (INFO > 0.9) SNPs pruned with PLINK 2.0 (‘–indep-pairwise 50 25 0.2’) in a leave-one-chromosome-out scheme. Subsequently, a SNP-wise logistic mixed model including the saddle point approximation test was performed using genotype dosages with SAIGE. To assess any residual confounding due to population stratification and artificial structure in the data, we calculated the LDSC intercept using SNP LD scores calculated in the HapMap3 CEU population.
Multivariate cox-proportional hazard analysis of the CREB3 regulon
For survival analysis, we calculated survival time from onset to death, or the censoring date using cox proportional hazard estimates, with comparisons through log-rank tests. Age at onset, sex and site of onset were included as covariates in the brain and blood-derived RNAseq cohorts. We tested for the proportional hazards assumption of the Cox regression model using the cox.ph() function. Survival analyses used the “cox.ph()” function from the “survival” package (version 3.5–5).
Sampled balanced multivariate cox-proportional hazard analysis of the CREB3R119G
After meta-analysis, 7161 whole genome sequencing were available for survival analysis. Outliers were identified based on the 0.5th and 99.5th percentile of the survival distribution across the entire cohort which removed 72 individuals (~ 1% of the total cohort) who had extreme survival values. High unbalanced ratio of carriers/non-carriers can limit power in cox-proportional hazard ratio estimates which prompt us to address the problem by randomly down sampling control individual to maximize power. Power estimates were evaluated using the function “powerCT.default0” with project hazard ratio, carriers/non-carriers ratio and number of events. Our power calculation estimated 99% power for N = 1500 non-carriers and N = 43 carriers. We then randomly down-sampled non-carriers individuals and repeated the cox.ph analysis N = 10,000 times. Survival analyses used the “cox.ph()” function from the “survival” package (version 3.5–5). Age at onset, sex and origin of the patients were used as covariates. Cox-proportional estimates were averaged across all permutations.
Univariate mean survival comparison
A univariate comparison of the mean survival was performed between carriers and non-carriers. Survival data from 7046 non-carriers and 43 carriers were randomly sampled 100,000 times and the average difference of mean survival between carriers and non-carriers was calculated for each permutation (Supplementary Fig. 11d). Significance was achieved if permuted-p < 0.05. Cohen’s d effect size was calculated using difference in mean survival between carriers and non-carriers divided by the standard deviation of the overall survival in the cohort.
Gene burden association analyses
To aggregate rare variants in a genic burden test framework we used the method described in the ALS 2021 GWAS3. For the discovery cohort, 6538 ALS cases and 2415 controls were available (http://databrowser.projectmine.com/) and 1873 ALS patients and 3926 controls for the replication. In short, a variety of variant filters was applied to allow for different genetic architectures of ALS associated variants per gene as was used previously68,69. In summary, variants were annotated according to allele frequency threshold (MAF < 0.01 or MAF < 0.005) and predicted variant impact (“missense”, “damaging”, “disruptive”). “Disruptive” variants were those variants classified as frameshift, splice-site, exon loss, stop gained, start loss and transcription ablation. “Damaging” variants were missense variants predicted to be damaging by seven prediction algorithms (SIFT, Polyphen-2, LRT, MutationTaster2, Mutations Assessor, and PROVEAN). “Missense” variants are those missense variants that did not meet the “damaging” criteria. All combinations of allele frequency threshold and variant annotations were used to test the genic burden on a transcript level in a firth logistic regression framework where burden was defined as the number of variants per individual. Sex and the first 10 principal components were included as covariates. All ENSEMBL protein coding transcripts for which at least five individuals had a non-zero burden were included in the analysis. Meta-analysis was performed using an inverse variance weighted method70.
Validations on mouse tissues
Puromycin administration
Six 90-day-old female Sod1G86R mice and five WT female littermates were anaesthetized with 2% isoflurane / 98% air, and two intracerebroventricular injections (one per hemisphere) were performed in the lateral ventricles to deliver 2.07 µl of puromycin (Sigma, P7255) diluted in saline at 25 µg/µl as previously described21. Animals were allowed to recover from surgery and sacrificed and perfused one hour later. Their brains were collected and processed for immunofluorescence.
Immunofluorescence
Mice were euthanized with an overdose of pentobarbital sodium and phenytoin sodium (120 mg/kg) and transcardially perfused with cold 0.01 M PBS, followed by cold 4% PFA in 0.01 M PBS. Brains were dissected and post-fixed overnight in 4% PFA. Brains were cut coronally in 40 μm-thick sections on a vibratome (Leica Biosystems). To reveal CREB3 expression, brain sections were first heated at 80 °C in 10 mM citrate buffer for 30 min. This step was not used to reveal puromycin. Brain sections were incubated 1 h in blocking solution (8% goat serum, 0.3% BSA, 0.3% Triton in PBS), and 48 to 72 h at 4 °C with primary antibodies. Sections were rinsed, incubated 2 h with the secondary antibodies, rinsed and mounted in Prolong Diamond mounting medium (Invitrogen, # P36970). The primary antibodies used were: mouse@puromycin (DSHB, #PMY-2H4; 1/100), rat@CTIP2 (Abcam, #Ab18465; 1/100), rabbit@CREB3 (Aviva Systems Biology #OAAN03577, 1/50), rabbit@ATF4 (Abcam, #ab31390, 1/100), rabbit@ATF6 (Santa Cruz, #sc-22799, 1/100) and rabbit@Phospho-eI2Falpha (Cell Signaling, #9721, 1/200). Secondary antibodies were from the Alexa series (Life Technologies; 1/1000).
RNAScope
Upon fixation as described above, brains were cryoptotected and cut coronally in 14 μm-thick sections on a cryostat (Leica). We selected sections located 1.1 mm rostral from Bregma where mCSN with a cervical projection are located8. RNAscope® Multiplex Fluorescent Reagent Kit v2 (Advanced Cell Diagnostics, #323100) was employed according to the manufacturer’s instructions, using probes targeting Creb3 and Fezf2 (Advanced Cell Diagnostics #1263611-C3 and #313301-C2). Briefly, sections were incubated with 1× PBS for 5 min to wash out OCT, baked for 30 min at 60 °C, post-fixed with 4% fresh PFA for 90 min, sequentially dehydrated with increasing concentrations of ethanol, baked again for 30 min at 60 °C, and incubated for 10 min in RNAscope™ Hydrogen Peroxide Reagent at RT. Target retrieval was performed for 5 min in a steamer, and RNAscope™ Protease III Reagent was applied for 15 min at 40°. Slides were then hybridized with target probes for 2 h at 40 °C in a hybridization oven (Boekel). Slides were stored with 5× Saline Sodium Citrate overnight at RT and signals were amplified using amplifiers and horse radish peroxidases from the reagent kit and TSA fluorophores (TOCRIS, # 7526 and # 7527). Samples were mounted in Superfrost® Plus slides (VWR, #631-0108) using ProLong™ Diamond Antifade Mountant medium (Invitrogen, #P36965).
Image acquisition and quantification
Images were captured at 63× using an AxioImager.M2 microscope equipped with a structured illumination system (Zeiss) and a high-resolution B/W camera (Hamamatsu), and run by the ZEN 2 software (Zeiss). Image analyses were performed with ImageJ (NIH). Modal Grey Value of puromycin and CREB3 signals was measured in CTIP2+ neurons in layer 5 of the mouse motor cortex, and Creb3 integrated density divided by cell size was measured in Fezf2+ cells.
Western blot on post-mortem human samples
Autopsy samples from motor cortex were obtained from the NeuroCEB biobank (https://www.neuroceb.org/fr/) for three sporadic ALS patients (one 55-year-old woman, one 61-year-old woman, and one 59-year-old man) of and three healthy controls (three men aged of 100, 95, and 85 years). Patients and / or families had provided written informed consent for the tissue collection and analyses. ALS diagnosis was based on the revised El Escorial criteria and was confirmed after autopsy. During autopsy, tissue samples were frozen in liquid nitrogen. Use of the tissue for research was declared at the French ministry for research and higher education. Tissues were homogenized in 1 ml of 0.1 mM ascorbic acid with a stainless-steel bead Tissue Lyser (Qiagen) during 3 min at 30 Hz, sonicated 3 × 10 s at 100 W with a Digital Sonifier (Banson) and centrifuged (20,000 × g, 30 min, 4 °C). Supernatants were collected and protein concentrations assessed with the Protein Assay kit (Bio-Rad). For fractionation, 10% (w/v) brain pellets (30–70 mg) were prepared in PBS and combined 1:1 with 2% sarkosyl (N-lauroylsarcosine) in PBS. Benzonase (50 U/ml; Sigma) was added to digest DNA, and the homogenates incubated with constant agitation at 37 °C for 30 min, followed by ultracentrifugation at 100,000 g for 30 min at 4 °C. The supernatant was removed and saved as the soluble fraction. Pellets were washed in 1% sarkosyl in D-PBS and spun at 100,000 g for 30 min. The supernatant was discarded and the pellet solubilized in 4% SDS, followed by addition of Laemmli sample buffer and heating at 100 °C for 10 min. A total amount of 8 µg of proteins /well were deposited and allowed to migrate on a gradient SDS gel (stain free 4–20%; Bio Rad), transferred onto nitrocellulose and probed with anti-CREB3 primary antibody (Proteintech, #11275-1-AP). Detection was performed with an HRP-conjugated secondary antibody combined with the Immobilon Forte Western HRP substrate (Millipore, #WBLUF0500). Quantification was performed using the Chemidoc software (BioRad); bands were normalized to total proteins.
Dual luciferase assays
HEK-293 cells were cultured in Dulbecco’s modified Eagle’s Medium (DMEM), 10% foetal bovine serum and 1% penicillin-streptomycin at 37 °C in 5% CO2. Cells were plated in a 6-well plate and transfected 24 h after plating in DMEM + 0.1% foetal bovine serum and 1% penicillin-streptomycin using Lipofectamine 2000 (Invitrogen). 800 ng of dual luciferase reporter containing CREB3 response element was co-transfected with 2400 ng of pCMV6, pCMV6-CREB3 or pCMV6-CREB3-R119G plasmid. Dual luciferase assays were performed 24 h after transfection as described by the manufacturer (Promega Secrete-Pair Dual Luminescence Assay Kit # LF033).
Statistics & Reproducibility
The aims of this study were (i) to run comparative transcriptomics between vulnerable neuronal populations of ALS mouse models and ALS patients, (ii) identified upstream regulators of identified common molecular pathways and (iii) run genetic and epidemiologic analyses on identified regulators to unravel putative disease modifiers. Vulnerable mCSN were purified from the motor cortex of the Sod1G86R mouse model of ALS to run RNAseq, and the data were integrated to published snRNAseq data from ALS patients5, using consensus classification of cell types13. Experiments on mouse samples were conducted by investigators blinded to the genotype. No statistical method was used to predetermine the sample size, which was estimated on the basis of former studies in rodents8,15,16,71. No data were excluded. Power calculate was performed in survival analysis to estimate the ratio of carriers and non-carriers in down-sampling method.
Immunofluorescence and RNAScope data are represented as nested scatter dot plots, and were analyzed using two-tailed nested t tests performed on Prism 6 (GraphPad). Results were considered significant when p < 0.05. For the luciferase assay, transfected cells were run as duplicate for a total of 6 independent experiment. A linear mixed-model was fitted to integrate the technical replicate as random effect followed by a Bonferroni’s test for multiple comparison. Results were considered significant when adjusted-p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Mouse CSN RNAseq data generated in this study have been deposited in the ArrayExpress database at EMBL-EBI under accession number https://www.ebi.ac.uk/biostudies/arrayexpress/studies/E-MTAB-7876 and the Gene Expression Omnibus (GEO) repository under the accession code: GSE291695. Human single-nuclei RNA-seq were downloaded through the GEO repository under the accession code: GSE174332 and GSE219281. Human and mouse snRNAseq of the spinal cord were downloaded through the GEO repository under the accession code: GSE161621 and GSE228778. Blood RNA-seq dataset is publicly available and was downloaded through NCBI GEO under the accession code: GSE234297. Post-mortem brain RNAseq from ALS patients are available through the target ALS consortium (https://www.targetals.org/research/funded-consortia/) with restricted access to authorized researcher. Human ALS whole genome sequencing is available to authorized researcher through dbGaP under the accession: phs001963.v2.p1. Whole genome sequencing data from Project Mine Consortium is available to authorized researcher (http://databrowser.projectmine.com/). RNA sequencing data from the New York Genome Consortium is available through the GEO repository under the accession code: GSE153960. Source data are provided with this paper.
Code availability
Relevant codes are posted at: https://github.com/SalimMegat/CREB3_ALS_resilience.
References
Van Es, M. A. et al. Amyotrophic lateral sclerosis. Lancet 390, 2084–2098 (2017).
Goutman, S. A. et al. Emerging insights into the complex genetics and pathophysiology of amyotrophic lateral sclerosis. Lancet Neurol. 21, 465–479 (2022).
van Rheenen, W. et al. Common and rare variant association analyses in amyotrophic lateral sclerosis identify 15 risk loci with distinct genetic architectures and neuron-specific biology. Nat. Genet. 53, 1636–1648 (2021).
Megat, S. et al. Integrative genetic analysis illuminates ALS heritability and identifies risk genes. Nat. Commun. 14, 342 (2023).
Pineda, S. S. et al. Single-cell dissection of the human motor and prefrontal cortices in ALS and FTLD. Cell 187, 1971–1989.e16 (2024).
Li, J. et al. Divergent single cell transcriptome and epigenome alterations in ALS and FTD patients with C9orf72 mutation. Nat. Commun. 14, 5714 (2023).
Ripps, M. E., Huntley, G. W., Hof, P. R., Morrison, J. H. & Gordon, J. W. Transgenic mice expressing an altered murine superoxide dismutase gene provide an animal model of amyotrophic lateral sclerosis. Proc. Natl. Acad. Sci. USA. 92, 689–693 (1995).
Marques, C., Burg, T., Scekic-Zahirovic, J., Fischer, M. & Rouaux, C. Upper and lower motor neuron degenerations are somatotopically related and temporally ordered in the Sod1 mouse model of amyotrophic lateral sclerosis. Brain Sci. 11, 1–18 (2021).
Bede, P. et al. Grey matter correlates of clinical variables in amyotrophic lateral sclerosis (ALS): a neuroimaging study of ALS motor phenotype heterogeneity and cortical focality. J. Neurol. Neurosurg. Psychiatry 84, 7166–7773 (2013).
Jin, J. et al. Dominant heterogeneity of upper and lower motor neuron degeneration to motor manifestation of involved region in amyotrophic lateral sclerosis. Sci. Rep. 9, 20059 (2019).
Ravits, J., Paul, P. & Jorg, C. Focality of upper and lower motor neuron degeneration at the clinical onset of ALS. Neurology 68, 1571–1575 (2007).
Zhang, Q. et al. Side of limb-onset predicts laterality of gray matter loss in amyotrophic lateral sclerosis. BioMed. Res. Int. 2014, 1–11 (2014).
Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 1–50 (2021).
Brunet, A., Stuart-Lopez, G., Burg, T., Scekic-Zahirovic, J. & Rouaux, C. Cortical circuit dysfunction as a potential driver of amyotrophic lateral sclerosis. Front. Neurosci. 14, 363 (2020).
Burg, T. et al. Absence of subcerebral projection neurons is beneficial in a mouse model of amyotrophic lateral sclerosis. Ann. Neurol. 88, 1–30 (2020).
Scekic-Zahirovic, J. et al. Cortical hyperexcitability in mouse models and patients with amyotrophic lateral sclerosis is linked to noradrenaline deficiency. Sci. Transl. Med. 16, eadg3665 (2024).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Berg, J. et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature 598, 151–158 (2021).
Zemel, B. M. et al. Resurgent Na+ currents promote ultrafast spiking in projection neurons that drive fine motor control. Nat. Commun. 12, 6762 (2021).
Liu, E. Y. et al. Loss of nuclear TDP-43 is associated with decondensation of LINE retrotransposons. Cell Rep. 27, 1409–1421.e6 (2019).
Shrestha, P. et al. Cell-type-specific drug-inducible protein synthesis inhibition demonstrates that memory consolidation requires rapid neuronal translation. Nat. Neurosci. 23, 281–292 (2020).
Consortium, T. E. P. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 488, 57–74 (2012).
Project MinE ALS Sequencing Consortium Project MinE: study design and pilot analyses of a large-scale whole-genome sequencing study in amyotrophic lateral sclerosis. Eur. J. Hum. Genet. 26, 1537–1546 (2018).
McMahon, A. et al. Sequencing-based genome-wide association studies reporting standards. Cell Genom. 1, 100005 (2021).
Moya, M. V. et al. Unique molecular features and cellular responses differentiate two populations of motor cortical layer 5b neurons in a preclinical model of ALS. Cell Rep. 38, 110556 (2022).
MacLean, M., López-Díez, R., Vasquez, C., Gugger, P. F. & Schmidt, A. M. Neuronal-glial communication perturbations in murine SOD1G93A spinal cord. Commun. Biol. 5, 177 (2022).
Blum, J. A. et al. Single-cell transcriptomic analysis of the adult mouse spinal cord reveals molecular diversity of autonomic and skeletal motor neurons. Nat. Neurosci. 24, 572–583 (2021).
Gautier, O. et al. Challenges of profiling motor neuron transcriptomes from human spinal cord. Neuron 111, 3739–3741 (2023).
Polioudakis, D. et al. A single-cell transcriptomic atlas of human neocortical development during Mid-gestation. Neuron 103, 785–801.e8 (2019).
Trevino, A. E. et al. Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 184, 5053–5069.e23 (2021).
Velmeshev, D. et al. Single-cell analysis of prenatal and postnatal human cortical development. Science 382, eadf0834 (2023).
Nolan, M. et al. Quantitative patterns of motor cortex proteinopathy across ALS genotypes. Acta Neuropathol Commun. 8, 98 (2020).
Bussière, T. et al. Progressive degeneration of nonphosphorylated neurofilament protein-enriched pyramidal neurons predicts cognitive impairment in Alzheimer’s disease: stereologic analysis of prefrontal cortex area 9. J. Comp. Neurol. 463, 281–302 (2003).
Lindholm, D., Wootz, H. & Korhonen, L. ER stress and neurodegenerative diseases. Cell Death Differ. 13, 385–392 (2006).
Gami-Patel, P. et al. Unfolded protein response activation in C9orf72 frontotemporal dementia is associated with dipeptide pathology and granulovacuolar degeneration in granule cells. Brain Pathol. 31, 163–173 (2021).
Ilieva, E. V. et al. Oxidative and endoplasmic reticulum stress interplay in sporadic amyotrophic lateral sclerosis. Brain: J. Neurol. 130, 3111–3123 (2007).
Prudencio, M. et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat. Neurosci. 18, 1175–1182 (2015).
Sasaki, S. Endoplasmic reticulum stress in motor neurons of the spinal cord in sporadic amyotrophic lateral sclerosis. J. Neuropathol. Exp. Neurol. 69, 346–355 (2010).
Saxena, S., Cabuy, E. & Caroni, P. A role for motoneuron subtype-selective ER stress in disease manifestations of FALS mice. Nat. Neurosci. 12, 627–636 (2009).
Sampieri, L., Di Giusto, P. & Alvarez, C. CREB3 transcription factors: ER-Golgi stress transducers as hubs for cellular homeostasis. Front. Cell Dev. Biol. 7, 123 (2019).
Raggo, C. et al. Luman, the cellular counterpart of herpes simplex virus VP16, is processed by regulated intramembrane proteolysis. Mol. Cell Biol. 22, 5639–5649 (2002).
Ying, Z., Zhang, R., Verge, V. M. K. & Misra, V. Cloning and characterization of rat Luman/CREB3, a transcription factor highly expressed in nervous system tissue. J. Mol. Neurosci. 55, 347–354 (2015).
Ying, Z., Misra, V. & Verge, V. M. K. Sensing nerve injury at the axonal ER: activated Luman/CREB3 serves as a novel axonally synthesized retrograde regeneration signal. Proc. Natl. Acad. Sci. USA 111, 16142–16147 (2014).
Ying, Z. et al. The unfolded protein response and cholesterol biosynthesis link luman/CREB3 to regenerative axon growth in sensory neurons. J. Neurosci. 35, 14557–14570 (2015).
Hasmatali, J. C. D. et al. Axotomy induces phasic alterations in luman/CREB3 expression and nuclear localization in injured and contralateral uninjured sensory neurons: correlation with intrinsic axon growth capacity. J. Neuropathol. Exp. Neurol. 78, 348–364 (2019).
DenBoer, L. M. et al. Luman is capable of binding and activating transcription from the unfolded protein response element. Biochem. Biophys. Res. Commun. 331, 113–119 (2005).
Liang, G. et al. Luman/CREB3 induces transcription of the endoplasmic reticulum (ER) stress response protein Herp through an ER stress response element. Mol. Cell Biol. 26, 7999–8010 (2006).
Penney, J., Taylor, T., MacLusky, N. & Lu, R. LUMAN/CREB3 plays a dual role in stress responses as a cofactor of the glucocorticoid receptor and a regulator of secretion. Front. Mol. Neurosci. 11, 352 (2018).
Locke, B., Campbell, E. & Lu, R. CREB3 mediates the transcriptional regulation of PGC-1α, a master regulator of energy homeostasis and mitochondrial biogenesis. FEBS Lett. 598, 1730–1739 (2024).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Catanese, A. et al. Synaptic disruption and CREB-regulated transcription are restored by K+ channel blockers in ALS. EMBO Mol. Med. 13, e13131 (2021).
Szewczyk, B. et al. FUS ALS neurons activate major stress pathways and reduce translation as an early protective mechanism against neurodegeneration. Cell Rep. 42, 112025 (2023).
Yadavalli, T. et al. Standalone or combinatorial phenylbutyrate therapy shows excellent antiviral activity and mimics CREB3 silencing. Sci Rep. 6, eabd9443 (2020).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Weidberg, H. & Elazar, Z. TBK1 mediates crosstalk between the innate immune response and autophagy. Sci. Signal 4, pe39 (2011).
Lenz, K. M. & Nelson, L. H. Microglia and beyond: innate immune cells as regulators of brain development and behavioral function. Front. Immunol. 9, 698 (2018).
Eitan, C. et al. Whole-genome sequencing reveals that variants in the Interleukin 18 Receptor Accessory Protein 3’UTR protect against ALS. Nat. Neurosci. 25, 433–445 (2022).
Gaastra, B. et al. Rare genetic variation in UNC13A may modify survival in amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Frontotemporal Degener. 17, 593–599 (2016).
Rouaux, C. et al. Sodium valproate exerts neuroprotective effects in vivo through CREB-binding protein-dependent mechanisms but does not improve survival in an amyotrophic lateral sclerosis mouse model. J. Neurosci. 27, 5535–5545 (2007).
Guez-Barber, D. et al. FACS purification of immunolabeled cell types from adult rat brain. J. Neurosci. Methods 203, 10–18 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Zhang, Y., Parmigiani, G. & Johnson, W. E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR Genom. Bioinform 2, lqaa078 (2020).
Orjuela, S., Huang, R., Hembach, K. M., Robinson, M. D. & Soneson, C. ARMOR: an automated reproducible modular workflow for preprocessing and differential analysis of RNA-seq data. G3 (Bethesda) 9, 2089–2096 (2019).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Durinck, S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Genovese, G. et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat. Neurosci. 19, 1433–1441 (2016).
Van der Spek, R. A. A. et al. The project MinE databrowser: bringing large-scale whole-genome sequencing in ALS to researchers and the public. Amyotroph. Lateral Scler. Frontotemporal Degener. 20, 432–440 (2019).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Scekic-Zahirovic, J. et al. Evidence that corticofugal propagation of ALS pathology is not mediated by prion-like mechanism. Prog. Neurobiol. 200, 101972 (2021).
Acknowledgements
The work has been supported by a European Research Council (ERC) starting grant #639737, a Marie Skłodowska-Curie career integration grant #618764, an Association Française contre les Myopathies (AFM)-Telethon trampoline grant #16923 and a Neurex grant to C.R. C.M. was supported by a PhD fellowship from the Institut National de la Santé Et de la Recherche Médicale (Inserm) and Région Alsace. Sequencing was performed by the GenomEast platform, a member of the “France Génomique” consortium (ANR-10-INBS-0009). This work was supported in part by the Intramural Research Program of the National Institutes of Health, the National Institute on Aging (1ZIAAG000933). Project MinE has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement n° 772376 – EScORIAL). The authors are thankful to the ENCODE and Project MinE Consortia which generated part of the datasets used in this study, as well as NeuroCEB biobank which provided the human samples. They thank Claudia De Tapia, Annie Picchinenna and Marie-José Ruivo for technical support.
Author information
Authors and Affiliations
Contributions
C.R. and S.M. conceptualized the study. S.M. performed transcriptomic and S.M. and R.A.J.Z. performed genetic analysis. C.M. and M.F. generated mouse RNAseq data. S.M., C.K., and M.A.M.P. analyzed mouse RNAseq data, and SM analyzed human snRNAseq data and genetic data. M.H.G., G.S.L., S.D.G., C.G., and A.B. performed immunofluorescence and RNAscope validations, with image analyses plug-ins designed by P.K. C.S. performed the luciferase assays designed by L.D. L.D. and S.M. established the needed collaborations with geneticists. B.T., L.F., A.C., A.L., L.F., J.H.V., S.W.S., and R.A.J.Z. provided the whole genome sequencing data of ALS patients and controls as well as the clinical data. The manuscript was drafted by C.R. and S.M. and reviewed and accepted by all authors.
Corresponding authors
Ethics declarations
Competing interests
JHV reports to have sponsored research agreements with Biogen, Eli Lilly and Astra Zeneca. The remaining authors declare no competing interests.
Consent for publication
All authors read and approved the publication of this manuscript.
Peer review
Peer review information
Nature Communications thanks Jack Humphrey and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Megat, S., Marques, C., Hernán-Godoy, M. et al. CREB3 gain of function variants protect against ALS. Nat Commun 16, 2942 (2025). https://doi.org/10.1038/s41467-025-58098-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58098-6
This article is cited by
-
Smart assistive technologies for neurodisorders: A review on AI, IoT, and wearable systems for enhanced patient care
Neurological Sciences (2026)