Abstract
Hypertensive disorders of pregnancy (HDP), including preeclampsia, affect 1 in 6 pregnancies, are major contributors to maternal morbidity and mortality, yet lack precision medicine strategies. Analyzing transcriptomic data from a prospectively-collected diverse cohort (n = 9102), this study reveals distinct RNA subtypes in maternal blood, reclassifying clinical HDP phenotypes like early/late-onset preeclampsia. The placental gene PAPPA2 strongly predicts the most severe forms of preeclampsia in individuals without pre-existing high risk factors, months before symptoms, and its overexpression correlates with earlier delivery in a dose-dependent manner. Further, molecular subtypes characterized by immune genes are upregulated in less severe forms of HDP. These results reclassify HDP clinical phenotypes into two distinct molecular subtypes, placental-associated or immune-associated. Validation performance for placental-associated HDP yields an AUC of 0.88 in the advanced maternal age population without pre-existing high risk factors. Molecular subtypes create new opportunities to apply precision-based medicine in maternal health.
Similar content being viewed by others
Introduction
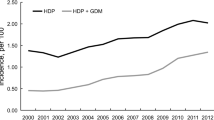
Hypertensive disorders of pregnancy (HDP) are a leading cause of morbidity and mortality among pregnant individuals1, and incidence has nearly doubled in the U.S. from 2007 to 20192 to affect 1 in 6 pregnancies. Though pathogenesis is poorly understood, HDP is presumed to be a multi-etiology disorder with hypothesized origins including viral causes3, immune reactions4, and placental dysfunction5,6. HDP encompasses a wide range of clinical phenotypes, from pregnant individuals who require extreme preterm delivery due to severity of maternal illness, to individuals with mildly elevated blood pressures after delivery requiring no intervention. Despite the spectrum of severity of these phenotypes, clinical recommendations to assess risk are the same7. Given the diverse presentation of disease we hypothesize there are unique molecular subtypes for the distinct clinical phenotypes of HDP. To test this hypothesis a prospective cohort with comprehensive clinical and molecular profiling is required.
Currently risk assessment for HDP is not specific to clinical phenotypes and is traditionally based on clinical and demographic risk factors7,8. Individuals with one high risk factor1,7,8, such as chronic hypertension or a history of preeclampsia, are considered at increased risk. Yet only 5% of pregnant individuals in the U.S meet such criteria9. For 95% of individuals, clinicians rely on a combination of weakly predictive “moderate” risk factors including race and socio-economic status, which can be difficult to medically define and may introduce bias in predictions1,7,8,10. As a consequence of generalized and ambiguous risk factors, the uptake of low-risk interventions such as aspirin has been found to be less than 50% even among high risk patients11. In contrast, for pregnant individuals already hospitalized with symptoms of HDP, prognostic tools such as sFLT1 as a ratio to PGF are available12. This context underscores the urgent need for unambiguous, biologically-driven predictive tools to target preventive treatment strategies, months before symptoms manifest13.
Here, we collected a large, multi-centered, racially, ethnically and geographically diverse prospective cohort and completed transcriptome analyses to explore how molecular subtypes associate with clinical phenotypes. The molecular subtypes were rigorously validated in a large, independent hold-out set (n = 3000).
Results
Prospective cohort with comprehensive clinical and molecular profiling
To understand the underlying biology that drives different pregnancy complications we conducted a multi-center, prospective, observational study: The Miracle of Life Study14. The study’s primary objective is to identify and validate cell-free RNA (cfRNA) classifiers that predict complications during pregnancy, months in advance of symptoms. 10,745 individuals over age 18 with singleton pregnancies were prospectively enrolled and blood samples were obtained in the second trimester, between 17 weeks and 4 days and 22 weeks and 0 days of gestation, timed to coincide with prenatal care for a fetal anatomy scan. To maximize diversity, samples were collected across 11 clinical sites, as well as by mobile phlebotomists covering a total of 808 zip codes. By design, this prospective cohort provides generalizability and molecular analysis unrestricted by predefined clinical phenotypes.
For the HDP analysis presented here, we included all samples that passed accessioning criteria and were collected between June 1st, 2020 and March 31, 2023, from all study sites with at least 200 samples (n = 11 sites and mobile phlebotomy), resulting in 9102 individuals included (Fig. 1, Table 1, Supplementary Table 1). To achieve a held-out validation group of n = 3000, we implemented a temporal split within each site such that the first 62% of samples collected from eligible sites were allocated to training and the last 38% collected were allocated to validation. For sites where source data verification was not possible, all samples were allocated to training (Fig. 1). After accounting for all samples that were dropped due to either incomplete clinical data or lab failures (Fig. 1, Methods), we retained 5399 subjects for training and 2829 subjects for validation of which, respectively, 4402 and 2243 do not have a high risk factor for HDP7.
HDP hypertensive disorders of pregnancy, GHTN gestational hypertension, PE preeclampsia. *For sites where source document verification was possible, otherwise site is only used in training.
HDP was defined using standard clinical and diagnostic criteria as defined by the American College of Obstetricians and Gynecologists (ACOG)8. The ACOG and U.S. Preventive Services Task Force (USPSTF) guidelines7,8 list the high risk factors for preeclampsia as chronic hypertension, preeclampsia in prior pregnancy, diabetes mellitus, chronic kidney disease, systemic lupus erythematosus, and antiphospholipid antibody syndrome. While having a multifetal pregnancy is also a high risk factor, these individuals were not enrolled in this study. Individuals with no high risk factors account for 82% of singleton pregnancies in our study. For 157 of the participants the clinical information was unclear as to case (HDP) or control status (not HDP). These records were adjudicated by an Endpoint Adjudication Committee composed of three board-certified Maternal Fetal Medicine experts that were not members of any of the enrollment sites. The adjudication process involved a review of Case Report Form data and redacted medical records for each subject and required a unanimous agreement by all committee members. Committee members were blinded to RNA sequencing results carried out as part of this study, and decisions were based solely on information from the medical records.
The study was conducted in accordance with Good Clinical Practice Guidelines of the International Conference on Harmonization15 and Institutional Review Board approval was obtained at each participating site. All participants provided written informed consent and those included in the final analysis met protocol-defined eligibility criteria.
We extracted cfRNA from all samples and generated transcriptomes, which were converted to gene counts for analyses and modeling (Methods). In order to adjust for variation in fetal fraction between pregnancies, we derived a method based sum of Y-chromosome counts to adjust counts in placental genes between samples (Methods). Counts were also corrected for fetal sex and gestational age at collection given the known variation16 (Methods).
Molecular subtypes reclassify common clinical phenotypes
Clinical guidelines7,8 define HDP as a disorder including gestational hypertension and preeclampsia with or without severe features, characterized by clinical metrics8. To initiate exploration of how cfRNA transcriptomic profiles align with clinical definitions, we identified differentially expressed genes between preeclampsia (PE) cases and all other samples, using the training data (n = 5399). In addition to measurement of statistical significance of group differences with the Mann–Whitney U statistic, we used Cohen’s d effect size17 as a statistical measure of the magnitude of separation to provide a more meaningful interpretation of differences18. The top two genes by effect size, are PAPPA2, highly expressed in the placenta, (Cohen’s d of 0.42 [95% CI 0.33–0.50], Mann–Whitney padj = 5.3 × 10−13), and CD163, highly expressed in immune cells (Cohen’s d = 0.36 [95% CI 0.28–0.44], Mann–Whitney padj = 5.3 × 10−13). These genes are consistent with prior biomarker findings that demonstrate involvement of placental and/or immune pathways in preeclampsia19, but it is not known if they represent different etiologies of the same disease. With our large prospective cohort we are now able to query whether molecular profiling can identify distinct phenotypes of HDP and provide biological insight as to their differential etiology.
To determine how the molecular profile maps to the different clinical phenotypes, we ranked all HDP cases into 8 clinical subgroups by severity using common clinical criteria: gestational age at initial PE diagnosis, gestational age at delivery, and the presence of severe features (Fig. 2a)8; individuals with postpartum PE and gestational hypertension (GHTN) were treated as separate subgroups. For each of the three categories we observe strong dose-dependence to PAPPA2 (Supplementary Fig. 1), validating their use for molecular stratification. We evaluated the effect of PAPPA2 and CD163 expression for each of the 8 subgroups and assessed subgroup similarity based on statistically significant separation of expression levels. This yielded two distinct molecular subtypes: the first comprising subgroups 1–3, which have the most severe clinical outcomes, and the second comprising subgroups 4–8 (Fig. 2b). Compared to non-HDP individuals, subgroups 1–3 each had a large effect size for PAPPA2 (joint Cohen’s d = 1.05 [95% CI 0.91–1.19]), whereas subgroups 4–8 each had small effect sizes for PAPPA2 (joint Cohen’s d = 0.18 [95% CI 0.12–0.25]), a highly significant increase in effect size of 5.7-fold (95% CI 4.0x–9.3x, p < 2e−5) suggesting a strong placental involvement for subgroups 1–3. Differential gene expression analysis between subgroups 1–3 relative to subgroups 4–8 showed that 5 of the 6 differentially expressed genes are upregulated and highly expressed in the placenta (Methods). In contrast the effect size for CD163 was similar across all 8 subgroups. Subgroups 4–8 had larger effect size for CD163 (joint Cohen’s d = 0.26 [95% CI 0.19–0.32]) than PAPPA2, suggesting an immune origin for these subgroups, which is further supported by gene set enrichment analyses and differential gene expression analyses for those groups (Supplementary Fig. 2, and see below).
a Samples are grouped by their clinical phenotype (diagnosis time <37w, delivery time <37w, and/or presentation of severe features), into 8 groups and non HDP controls. 13 samples were missing sufficient information to include in subgroups not shown. b Effect size of PAPPA2 is shown next to each clinical phenotype, blending into molecular subgroups, placental-associated or immune-associated. Statistical comparisons of placental-associated or immune-associated versus non HDP (baseline) calculated using a two-sided Mann–Whitney U test. c Effect size of PAPPA2 for placental-associated and immune-associated HDP split by presence/absence of maternal high risk factors (RF). Statistical comparisons of each subgroup versus the non HDP (baseline) with the corresponding presence/absence of maternal high RF calculated using a two-sided Mann–Whitney U test. d PAPPA2 expression in early-onset PE split into placental-associated and immune-associated HDP among individuals with no high risk factors. e PAPPA2 expression in ante- and intrapartum PE with severe features split into placental-associated and immune-associated HDP among individuals with no high risk factors. Box and whisker plots display a box with lower bound, center line, and upper bound at the 25th, 50th, and 75th percentile, respectively, where whiskers extend to 1.5 times the inter-quartile range with points outside this range depicted individually as circles. Statistical comparisons depicted in box and whisker plots are calculated using a two-sided Mann–Whitney U test. HDP hypertensive disorders of pregnancy. Combined mean effect size for PAPPA2 for PE is represented in white, above-average in reds and below average in blues.
Therefore, we arrived at two distinct molecular subtypes of HDP: 1) Placental-associated HDP comprised of subgroups 1–3, i.e. individuals who were diagnosed before 37 weeks leading to preterm delivery and/or severe features and 2) Immune-associated HDP comprised of subgroups 4–8, including most term PE, postpartum PE and GHTN.
Others have reported differences in gene expression for PE in individuals with CHTN20, so to explore the potential for more biologically homogeneous populations we studied the relationship between HDP molecular subtype and pregnancy status by presence of at least 1 high risk factor (n = 997) or absence thereof (n = 4402). Subgroups 1–3 are strongly stratified by placental molecular subtype: Individuals with a high risk factor had a medium to large effect size for PAPPA2 (joint Cohen’s d = 0.65 [95% CI 0.45–0.86], Fig. 2c); while individuals with no high risk factors had an increased and very large effect size both compared to the non HDP population (joint Cohen’s d = 1.59 [95% CI 1.38–1.81], Fig. 2c) and when compared to all other samples (Cohen’s d = 1.54 [95% CI 1.32–1.75]). In contrast, the immune molecular subtype did not stratify by high risk factor (Fig. 2c).
The molecular stratification of HDP maintains the dose-dependence with gestational age at delivery in the samples with no high risk factors, where higher PAPPA2 expression correlates with earlier delivery (r = −0.42 [95% CI −0.58–−0.22], p = 6.3 × 10−5, Supplementary Fig. 3a). Further, PAPPA2 overexpression is specific to preterm delivery due to preeclampsia; in individuals with spontaneous preterm birth but without preeclampsia (sPTB, n = 246) PAPPA2 is not overexpressed (Cohen’s d = −0.06 [95% CI −0.18–0.07]). This suggests that PAPPA2 is not a signal for delivery date per se, but rather signifies risk of HDP severe enough to require preterm delivery.
The placental-associated molecular subtype is more nuanced as compared to the traditional clinical labels of early-onset or severe features. Early-onset preeclampsia, i.e. those diagnosed less than 34 weeks gestation, stratify into groups aligned with the molecular subtype, with significantly higher PAPPA2 expression among those who also deliver preterm, but not those that deliver at term or have postpartum PE (Fig. 2d, p = 2.8 × 10−4 in the population with no high risk factors). Similarly, preeclampsia with severe features separates into groups aligned with the overall molecular subtype (Fig. 2e, p = 2.7 × 10−9 in the population with no high risk factors). Additionally, differentiation within the early-onset or severe features groups are independent of high risk factors and expands to the full groups as well (Supplementary Fig. 3b, c).
Less severe forms of HDP share immune-associated molecular subtype
To understand the molecular structure within the immune-associated HDP samples, we compared two groups, GHTN (n = 615) and term PE (n = 453), excluding those diagnosed before 37 weeks who later develop severe features (Fig. 2a), and found no genes that pass a Benjamini-Hochberg adjusted p-value threshold at the 0.05 alpha level to qualify for differential expression. The absence of differential genes for these subgroups supports high molecular homogeneity between them. Furthermore, when individually comparing each group to non-HDP controls the tails are enriched for immune genes (Fig. 3a, b), a result also supported by gene ontology analyses (Methods). When comparing the overlaps of all significant differentially expressed genes between the two groups, we see a highly significant overlap (p = 6.7 × 10−36, Fig. 3c). If we compare placental-associated HDP to immune-associated HDP in a standard gene set enrichment analysis, the top 20 gene sets upregulated in immune-associated HDP are exclusively related to inflammatory pathways, whereas several of the top enriched gene sets in placental-associated HDP are related to platelet function (Supplementary Fig. 2). The same conclusion can be reached from gene set enrichment analyses between immune-associated HDP and non-HDP controls and is supported by statistically significant gene-ontology overlaps between the groups (Methods).
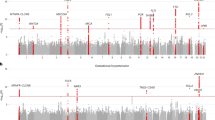
Quantile-quantile (QQ) plot of ranked Mann–Whitney U p-values, colored by absolute Cohen’s d effect size for a immune-associated PE versus non HDP (genomic inflation factor λ = 1.2, 29 significantly differentially expressed genes), b immune-associated GHTN versus non HDP (λ = 1.8, 38 significantly differentially expressed genes). Top 10 genes are named, and gene names are bolded based on overlap with immune and inflammatory GO biological processes. Statistical comparisons in QQ plots are calculated using a two-sided Mann–Whitney U test with multiple test correction by Benjamini–Hochberg. c Overlap in differentially expressed genes. Significance of the overlap was evaluated using a hypergeometric test, where the reported probability corresponds to the likelihood of observing an overlap of a given size or larger by random chance under the null hypothesis that the sets are independent. PE, preeclampsia; GHTN, gestational hypertension; HDP, hypertensive disorders of pregnancy.
Placental-associated molecular subtype predicts preterm PE and adverse neonatal outcomes
Understanding that PE requiring preterm delivery confers the greatest morbidity and mortality to both mother and infant, we then sought to develop a classifier to predict this group based on the molecular subtypes. To build the classifier for placental-associated HDP for individuals with no high risk factors, we first performed feature discovery to identify genes that complement PAPPA2 to improve prediction accuracy. We performed searches across the training data, using either all samples (Fig. 4a) or only samples with no high risk factors (Fig. 4b) to identify the top differentially expressed genes. We optimized and evaluated the performance of our feature selection method using cross-validation with multiple random splits of our training data to avoid the common pitfall of data leakage.
Quantile-quantile (QQ) plot of ranked Mann–Whitney U p-values and colored by absolute Cohen’s d effect size for a placental-associated HDP versus all others (genomic inflation factor λ = 2.2), and b placental-associated HDP versus all others with no high risk factors (λ = 5.4). Statistical comparisons in QQ plots are calculated using a two-sided Mann–Whitney U test with multiple test correction by Benjamini–Hochberg. c Effect size for classifier genes in a population with no high risk factors, split into clinical categories based on diagnosis time <37wks, delivery time <37wks, and/or presentation of severe features. 11 samples were missing sufficient information to include in subgroups and were thus excluded from this plot. d Classifier prediction across placental-associated HDP by gestational age at delivery in individuals with no high risk factors. HDP Hypertensive disorders of pregnancy.
Genes for the final classifier were selected by applying this framework on the full training dataset only (n = 5399). The training data were then used to train a logistic regression model with an L1 regularization penalty, and training samples with no high risk factors were used to set a decision boundary for the classification calls. Six genes were included in the classifier, comprising four placental genes (PAPPA2, KISS1, XAGE2, and ADAM12) and two inflammatory and immune related genes (CD163, VSIG4). These six genes have all previously been associated with preeclampsia or early onset preeclampsia21,22,23,24,25,26. Effect sizes for all classifier genes by clinical phenotype in individuals with no high risk factors are shown in Fig. 4c. Although CD163 and VSIG4 are upregulated in placental-associated HDP, their gene networks are down-regulated relative to immune-associated HDP, suggesting these two genes carry a broader signal predictive in both groups (Methods).
We used the Human Protein Atlas tissue-specific abundance data, to evaluate the origin of the model transcripts. For each of the model genes we confirm PAPPA2, KISS1, XAGE2 and ADAM12 are most likely placental transcripts, which is also supported by their low expression in non-pregnant individuals (Supplementary Fig. 4, Methods), and that CD163 and VSIG4 are mostly derived from the spleen and other immune tissues (Supplementary Fig. 4). Extending this technique to annotated cell types, we found that the top three most correlated cell types for each of PAPPA2, KISS1, XAGE2, and ADAM12 are syncytiotrophoblasts, cytotrophoblasts and extravillous trophoblasts (Supplementary Fig. 5, Methods). For CD163 and VSIG4, the top three are Hofbauer cells, Kupffer cells, and macrophages (Supplementary Fig. 5, Methods).
Prior work has shown that some clinical metrics are mildly predictive of the development of HDP7. Thus, we investigated whether any of these individual metrics could provide additional predictive information to the RNA-based predictors. We focused on available collected metrics in the medical record related to high and moderate risk factors for the clinical risk assessment for preeclampsia by the USPSTF, including mean arterial blood pressure most proximal before the time of blood draw, nulliparous status, body mass index (BMI) from the first recorded prenatal visit, race and age. Of those, only mean arterial blood pressure and BMI were used as additional features in the classifier, due to additive performance with the genes in training cross validation. In our training data a genes-only model achieved a mean cross-validation AUC of 0.86 in participants with no high risk factors, and addition of mean arterial pressure and BMI increased the mean performance to 0.88. Importantly, within our heterogeneous training samples, we confirm that race did not enhance predictive information beyond the RNA-based predictors in cross validation in individuals with no high risk factors in agreement with our previous observation in a retrospective study16. The full classifier consists of transcript levels for 6 genes and two clinical metrics; classifier features, coefficients, and decision boundary were all locked prior to classifier validation.
We observed a median lead time of 108 days and 132 days between blood draw and delivery in the placental-associated or immune-associated HDP, respectively. To investigate if the particular signal for the earliest deliveries are driven by proximity of blood draw to delivery or the pathophysiology necessitating earlier delivery, we looked at the impact of both on the model output. In the 6-gene model, earlier delivery has stronger signal than proximity within training cases, supporting that the molecular signal is for severity rather than proximity (Methods). In the placental-associated subtype the test positive cases on average deliver 11.3 days before a test negative case, and importantly no cases below 32 weeks at delivery are missed in the training set (Fig. 4d).
To verify performance on a clinically relevant metric that was not used in training, we looked at the days spent in the neonatal intensive care unit (NICU) for individuals with no high risk factors with respect to a PE diagnosis in the test positive and test negative populations within the training dataset (Methods). Among the individuals who were admitted to the NICU the test positive group spent on average 14 days (n = 81) compared to 5 days (n = 154) in the test negative population (two-sided Kolmogorov–Smirnov test p = 6.1 × 10−4). Additionally, these results replicate in a cross-validation on the training set (Methods). This in combination with the previous observation that test positive individuals deliver earlier (Fig. 4d) supports that individuals who share the placental-associated molecular subtype have more adverse outcomes.
Classifier validation for pregnancies with no high risk factors for PE
Upon completion of training (n = 5399) the classifier was locked with a pre-specified decision boundary. Validation was also pre-specified for individuals with no high risk factors (n = 2243) in the held-out validation set (n = 3000). Validation results reported below are for individuals with either only moderate risk factors or no risk factors.
An important endpoint of the study was to validate the placental-associated molecular subtype i.e. individuals who were diagnosed before 37 weeks leading to preterm delivery and/or severe features, using a classifier based only on RNA features. The results validated that the RNA classifier stratifies placental-associated cases from all other samples (p = 6.3 × 10−7); stratifies placental-associated cases from immune-associated cases (p = 7.4 × 10−4); and also stratifies immune-associated cases from non-HDP controls (p = 9.9 × 10−9).
The full classifier for all individuals with no high risk factors in the validation cohort (n = 2243) yielded an AUC of 0.74 (Fig. 5a, blue line, maternal age: ≥18, predicted outcome: placental-associated HDP). When the case-label is shifted to placental-associated HDP with delivery at or before 35 weeks of gestation (n = 2243), the validated AUC increases to 0.83 (Fig. 5a, green line, maternal age: ≥18, predicted outcome: placental-associated HDP with delivery ≤35 weeks), confirming the observed dose-dependence for gene expression (Supplementary Fig. 1a). The 35 week threshold is chosen to align with where most hospitals require admission to the NICU27. Among the moderate risk factor groups, nulliparous (n = 984), Black race (n = 476), and obesity (BMI > 30, n = 801), the classifier performed similarly to all individuals, while for advanced maternal age (AMA) individuals (n = 403) the validated AUC was 0.88 (Fig. 5a, orange line, maternal age: ≥35, predicted outcome: placental-associated HDP). AMA is associated with earlier preterm deliveries28 where the placental-associated effect size is stronger.
a Receiver operator characteristic (ROC) for all individuals with predicted outcome as placental-associated HDP (blue); all individuals with predicted outcome as placental-associated HDP and ≤35 week delivery (green); individuals with advanced maternal age (≥35 years) with predicted outcome as placental-associated HDP (orange). Performance at the prespecified cutpoint is annotated with a X on each curve. b Comparison of the positive likelihood ratio for each of the groups with USPSTF performance shown in striped lighter tone and the full classifier in solid color matching the ROC. AMA advanced maternal age (≥35 years), HRF High Risk Factor, USPSTF United States Preventive Services Task Force.
As a relevant comparator for a US-based study, we used the USPSTF guidelines7,8. For each group described above the positive likelihood ratios are improved over the guideline performances (Fig. 5b). For AMA individuals, where age itself is a moderate risk factor, the guidelines classify those with one or more additional moderate risk factor(s) as increased risk. For test positive individuals in the AMA population, this corresponds to a fifteen-fold improvement in increased risk to develop placental-associated HDP over the standard guidelines (+17% vs +249%). For test negative individuals in the AMA population, the negative likelihood ratio for the validated classifier was 0.12 compared to 0.38 for USPSTF guidelines. For placental-associated HDP the NPV is 99.7% and the PPV is 8.9%. The PPV for all preeclampsia is 24.6% and for all HDP is 42.9%. Though the validated classifier is optimized to identify the most severe cases, milder cases of PE or GHTN are also enriched in test positive individuals, with a positive likelihood ratio of 3.0 for all preeclampsia and 2.9 for HDP.
Discussion
Through transcriptome analysis in a large (n = 9102), multi-center, ethnically and geographically diverse, prospective cohort, we have defined and validated two distinct molecular subtypes that allow the molecular stratification of hypertensive disorders of pregnancy (HDP). Placental-associated HDP maps clinically to severe/preterm preeclampsia while immune-associated HDP maps clinically to most term preeclampsia and gestational hypertension. Both molecular subtypes can be detected via a standard blood draw at the time of the anatomy scan, between 18 and 22 weeks of gestation, with a median lead time of 108 and 132 days to delivery, respectively, creating new opportunities for preventative interventions29.
Prospective validation of the lead gene, PAPPA2, confirms earlier findings of its relevance at both the cellular30 and circulatory16 levels. The molecular subtypes are specific to HDP: the placental-associated HDP is only present in those with a preterm birth associated with HDP or preterm HDP diagnosis with severe features and it is absent in those individuals having a spontaneous preterm birth. These findings underscore the need to discover specific molecular subtypes that drive other adverse pregnancy outcomes.
While case-control studies have traditionally been used in biomarker discovery studies to explore pregnancy biology, the results often fail to validate, due to their dependence on existing clinical labels that are molecularly heterogeneous and/or their limited ability to sample population variance31. Comprehensive clinical and molecular profiling in a large, diverse, prospective cohort, enables the molecularly driven subtyping of clinical labels, to overcome previous limitations. We discover two molecular subtypes of HDP aligned with disease severity and then validate a molecular classifier for the most severe disease designated as placental-associated HDP. The molecular signal separates samples independent of common clinical phenotypes like presence or absence of severe features and early- or late-onset; classifications others have also found insufficient32. Further, we demonstrate that gestational age at delivery has a strong dose-dependence to PAPPA2. It is known that PAPPA2 plays a role in regulation of the insulin-like growth factor 1 and 2 pathways, which have been implicated in hypertension and preeclampsia33,34,35. Though further mechanistic work is needed to establish causality of PAPPA2 overexpression in HDP, studies in mice indicate that siRNA targeted to PAPPA2 can alleviate the effect in a model system with low extravillous invasion36. Furthermore the dysregulation of PAPPA2 is persistent through the end of pregnancy as evidenced by single-cell sequencing of placenta30
Unexpectedly, placental-associated HDP is most strongly predictive for individuals with no high risk factors, who comprise the vast majority of pregnant individuals9 and for whom there are currently no effective ways to assess risk. A likely explanation is the higher biological homogeneity among individuals with no high risk factor. Other risk prediction models largely identify individuals with pre-existing high risk factors37,38, whereas the test presented here focuses on shared biological signals in the population without pre-existing high risk factors. Specific molecular subtypes are likely to exist for each high risk factor group, such as chronic hypertension or prior PE, but to disentangle those, future studies with a larger sample size of individuals with each high risk factor would be needed. Molecular subtypes for individuals with no risk or 1 or more moderate risk factors, which identify individuals at risk for HDP months prior to any clinical symptoms, present a new opportunity for risk stratification and utilization of preventive strategies.
Having an objective measurement of risk in a population without clinical high risk can help drive adherence39,40 to preventative care29 and is a necessary first step to improve the worsening maternal health status1.
Advancements in medical fields like oncology have been catalyzed by understanding the underlying pathobiology to trigger the development of targeted treatment options for each molecular subtype of disease, and progress towards molecular classification continues41. The advent of early detection of molecular subtypes for HDP may enable new therapeutic targets, such as vascular/endothelial targets for placental-associated HDP and immunomodulatory approaches to immune-associated HDP13,42. Importantly, current clinical trials to treat or prevent preeclampsia43 can now benefit from an improved efficacy-safety profile through targeting patient enrollment by molecular subtype. As examples, statins have been proposed as therapeutic agents in the management of preeclampsia based on the benefits that they provide to patients with angiogenic imbalance, endothelial dysfunction, oxidative injury, and inflammatory injury44 and may benefit from stratification by the placental-associated and immune-associated molecular subtypes. Specifically, higher doses of aspirin may be more effective in individuals with the placental-associated subtype as they have been shown to be more effective in prevention of preterm PE45, which are highly enriched in placental-associated HDP. Importantly, this stratification now provides the opportunity to test and/or develop therapeutics to reverse the rising prevalence of HDP and its significant global contribution to maternal and neonatal morbidity and mortality.
More immediately, a simple, stand-alone blood test to identify pregnant individuals at increased risk of HDP, months in advance of symptoms, is expected to motivate the prescription of and adherence39 to existing, evidence-based interventions to improve outcomes29. The HDP molecular subtypes presented here are objective, biologically driven, and independent of race. They present an opportunity to shift the paradigm away from generalized ‘moderate’ risk factors that are nonspecific, susceptible to bias or difficult to assess clinically, and toward a precision-medicine approach to maternal health.
Methods
Specimen collection
Participants were recruited via either direct recruitment at one of eleven medical centers in the United States or via direct-to-participant recruitment using social media. The clinical recruitment sites were Womans Hospital in Baton Rouge, LA; Magee-Women’s Research Institute at UPMC in Pittsburgh, PA; University of Texas Medical Branch in Galveston, TX; Ochsner Medical Center in New Orleans, LA; Multicare Healthcare System in Tacoma, WA; Washington University Medical Center in St. Louis, MO; University of California San Diego in San Diego, CA; Beth Israel Medical Center in Boston, MA; Brigham and Women’s Medical Center in Boston, MA; The Ohio State University Wexler Medical Center in Columbus, OH; and Thomas Jefferson University Hospital in Philadelphia, PA. Whole blood specimens were collected via standard non-fasting venipuncture using two to three 10 mL RNA Complete BCT for Cell-Free RNA® (cfRNA) tubes (Streck, La Vista, NE) which stabilizes and preserves cfRNA in plasma as well as cellular RNA in white blood cells. Specimens were shipped overnight at ambient temperature via FedEx to the Mirvie laboratory in South San Francisco, California. Only specimens that were received within 48 h were included in the study.
Limited plasma is available from each participant, and requests for plasma cannot be accommodated.
Non-pregnant individuals for baseline RNA measurement were acquired from Discovery Life Sciences (Huntsville, AL) and collected using the same protocol as the clinical study.
Clinical data collection
Shortly after the end of the pregnancy clinical data was abstracted from the medical records encompassing the prenatal record, laboratory results, imaging results, labor and delivery notes, and discharge summaries. All information captured was de-identified and stored in a password-protected and secure web-based electronic data capture system (Medrio, San Francisco, CA). Clinical monitoring was performed by trained clinicians and any queries were resolved with on-site principal investigators and/or by a panel of three independent maternal-fetal medicine physician specialists. All HDP related outcomes were abstracted and adjudicated in accordance with ACOG clinical guidelines8. The clinical data collection was performed by staff separate from the laboratory processing team. Both teams were blinded to the other team’s results during lab processing and clinical data collection. The study was designed and powered to explore cfRNA predictors of adverse pregnancy outcomes and molecular subtypes with a prevalence >1%.
Institutional Review Board approval was obtained at each participating site, reviews were done by Woman’s Hospital Foundation Institutional Review Board (RP-20-027), Beth Israel Deaconess Medical Center Committee on Clinical Investigations (2022P000224), and Advarra Institutional Review Board (Pro00050765). Written informed consent was obtained from all participants.
A total of 444 individuals were excluded from the training cohort and 138 individuals from the validation. Individuals were excluded from the study for the following reasons, did not meet eligibility criteria (training n = 31, validation n = 10), lost to follow-up or incomplete data (training n = 333, validation n = 91), participant or sponsor requested withdrawal (training n = 7, validation n = 2), out-of-window sample collection (training n = 15, validation n = 7), or excluded per adjudication committee (training n = 58, validation n = 28).
Information on the 49 donors for the baseline study were reported by Discovery Life Sciences and the donors were 32 (22–40) years with an average BMI of 32 (19–61) at time of blood draw. The self-reported race for the non-pregnant individuals was 29 Black and 20 White.
Data blinding
Access to clinical data was closely controlled and only made available on a need-to-know basis to protect participant privacy and confidentiality, and to preserve data integrity for analyses. The lab team did not have access to participant information to avoid any bias in sample processing. The data analysis team only had access to data for the training cohort to enable model development, any identifiable information was removed. For the validation cohort only the clinical operations team and the software team had access to data. Prior to running any analyses on the validation data, the prediction model was locked and analyses were pre-specified and documented. The data analysis team provided a script to the software team to output results from the validation data. Access to the validation data is still restricted from the data analysis team to enable future studies.
Sample processing
Similar to methods previously described for the Mirvie RNA Platform, blood specimens received in the laboratory are processed and plasma separated, cfRNA is extracted, libraries are created and sequencing is performed16,46. Total nucleic acid was isolated using QIAamp Circulating Nucleic Acid kit (Qiagen, Germantown, MD) followed by DNAse treatment using Baseline-ZERO DNAse (LGC Biosearch Technologies, Middlesex, UK). An RNA spike-in was added to the samples during extraction. After DNAse treatment, cfRNA was eluted using the Monarch RNA clean-up kit (New England Biolabs, MA). cfRNA libraries were prepared using the SMARTer Stranded Total RNAseq Kit v2 - Pico Input Mammalian (Takara Bio, San Jose, CA) and enriched for the human transcriptome using the SureSelect Target Enrichment kit (Agilent Technologies, Santa Clara, CA).
The extracted cfRNA and libraries were assessed for quality using a reverse transcription-quantitative polymerase chain reaction (RT-qPCT) assay to follow 3 targets of interest as follows: a housekeeping gene (actin beta [ACTB]), an RNA spike-in, and a cfDNA contamination assay. Libraries of multiple samples were then pooled and sequenced to an average depth of 67.03 million reads on the Illumina NovaSeq platform. Individual samples more than 3 standard deviations from the mean with respect to lab quality metrics were removed as outliers. On average, we obtained at least 1 count for 21,478 genes after removing duplicate reads. There was no significant difference in sequencing depth between the training and test groups.
Bioinformatics processing
The bioinformatics pipeline is built as a Nextflow workflow (v23.10.0)47 that is executed with Docker (v20.10.25) containers and encompasses multiple steps as follows. Sequencing reads were converted from bcl files to fastq files using Illumina’s bcl-convert v4.2.748, and fastq files were combined into one R1 and one R2 fastq per sample and validated with biopet-validatefastq v0.1.149 and fq v0.11.050. Adapters were trimmed with fastp v0.23.451, which also performed base correction of obvious errors in overlapping regions (to better estimate duplication rates), and the processed fastq files were again linted with biopet-validatefastq49. Reads were then mapped to GRCh38 using the STAR52 aligner v2.7.10b. The mapped reads were deduplicated using sambamba v0.8.253. Two different gene counts tables were calculated including, a primary count file produced by Rsubread’s featureCounts function v2.8.254 and a secondary counts file produced by HTSEQ v2.0.555 used to generate fetal sex probabilities for data pre-processing. Primary counts tables were converted to log2 counts per million (CPM) with a pseudo count of 1 CPM.
Data pre-processing
Major sample processing library batches were corrected genewise such that each batch yielded the same median over control samples in that training batch for each gene. All genes were residualized over the linear effects of gestational age at sample collection to control for the span of 4 weeks and 3 days in collection and over a fetal sex probability estimated from a gene-based logistic model to control for the variation across fetal sex. Both corrections were warranted as various genes, including CD163, ADAM12, XAGE2, VGLL3, SVEP1, and KRT7, carry variance explained by gestational age even when accounting for fetal sex (p < 0.05, ANCOVA Type III). Genes annotated as “elevated” in placental expression from the Human Protein Atlas56 were additionally residualized over VGLL3, SVEP1, and KRT7 genes to control for fetal fraction variation. These linear correctors were implemented with the scikit-learn 1.4.057 robust HuberRegressor fit on training data.
Gene correction for fetal fraction variation
The fraction of cfRNA in maternal blood that originates from the fetus or placenta rather than the pregnant individual (i.e. the fetal fraction) varies among pregnancies. To control for variation in the overall fetal fraction of circulating RNA in maternal plasma, correction genes were identified by searching for placental genes that are correlated to a known fetal signature. Since certain genes on chromosome Y are specific to males, the expression levels for these genes in pregnancies with male fetuses were used to estimate the fetal fraction in cell-free RNA. Correlations against the total gene expression level across chromosome Y genes in log2 counts downsampled to 3M assigned counts per sample were performed against placental genes significant to pregnancy to identify correctors for the variation in fetal fraction. Placental genes were sorted in descending order by the coefficient of determination calculated with a robust linear regression fit (HuberRegressor). The gene that was most correlated to the sum across chromosome Y genes in individuals pregnant with male fetuses was identified as VGLL3, an autosomal gene. Cell free RNA expression levels summed across the placental genes corrected by VGLL3 showed no difference between pregnancies with male or female fetuses (p > 0.05). Additional placental genes SVEP1 and KRT7 were selected as corrections to enhance the predictive power of the placental-associated classifier. All three placental correction genes are not themselves sex-linked and can thus generalize to all samples. Correcting genes with elevated RNA expression in the placenta (Human Protein Atlas)56 by these fetal-fraction covariates regresses out variation in the placental genes due to variation in fetal fraction.
Gene correction for fetal sex variation
The placental correction genes VGLL3, along with the other placental correction genes SVEP1 and KRT7 and the model features ADAM12 and XAGE2 carry variance explained by fetal sex, even after accounting for collection ga (p < 0.05, ANCOVA Type III). This imparted a fetal sex bias in the model outputs that was detected in cross validation during training as a bias in sensitivity and specificity across male and female fetuses measured across the test folds. This bias was corrected by applying the fetal sex correction by residualizing over a gene-based probability the fetus is male. Data pre-processing applies a gene-based model estimate of the fetal sex of a singleton pregnancy (based on the probability that the fetal sex is male) to control for variation in the circulating RNA derived from the fetus and placenta across male and female fetuses, which have been shown to differ in size at a fixed gestational age58. The fetal sex prediction model is based on eleven genes identified to be differentially expressed with high statistical significance between male and female fetus singleton pregnancies: PCDH11Y, RPS4Y1, EIF1AY, DDX3Y, ZFY, USP9Y, UTY, KDM5D, VGLL3, CAPN6, CSHL1. One gene CAPN6 is on chromosome X, while the remaining eight genes excluding VGLL3 (chr 3) and CSHL1 (chr 17) are located on chromosome Y. The classifier coefficients and decision boundaries were trained on log2 gene counts (applying a single pseudocount) processed by HTSEQ. Samples were qualified for prediction based on a minimum of three million counts uniquely assigned to the reference. The fetal sex prediction model was validated on unseen test data and shown to perform with 98.9% accuracy at 100% sensitivity and 98% specificity on the validation test set when classifying fetal sex.
Differential genes between placental-associated and immune-associated HDP
Placental-associated HDP (subgroups 1–3, n = 198) was compared to immune-associated HDP (subgroups 4–8, n = 1055) by differential gene expression analysis across 2036 pregnancy genes based on a two-sided Mann–Whitney U rank test with multiple test correction by Benjamini–Hochberg. Six genes (PAPPA2, XAGE2, PSG11, KISS1, CD163, and CAPN6) were identified to be differentially expressed between the two subtypes of HDP. Five of these differentially expressed genes are placenta genes (described in Methods): PAPPA2, XAGE2, PSG11, KISS1, and CAPN6. All differential genes are upregulated in placental-associated HDP relative to immune-associated HDP. Cohen’s d effect sizes are shown in Supplementary Table 1 annotated with the tissue RNA expression tissue specificity from the Human Protein Atlas56,59,60.
Gene Ontology overrepresentation analysis
Overrepresentation testing of Gene Ontology (GO)61,62 biological processes complete terms in Homo sapiens (release date 2024-01-17) was performed using the PANTHER Classification System analysis tool (v19.0, https://geneontology.org)63 with the PANTHER Overrepresentation Test (Released 20240226). The top 50 differentially expressed genes identified in the immune-associated HDP vs non HDP differential gene expression analysis across 2036 pregnancy genes based on a two-sided Mann–Whitney U rank test with multiple test correction by Benjamini–Hochberg were analyzed. Fisher’s Exact p-values were calculated for the association of these genes with GO terms and corrected for multiple testing by the false discovery rate. GO terms were filtered to pass the adjusted p-value at alpha equals 0.05, and the GO term with the maximum fold enrichment per shared group was reported for each group.
Among immune-associated HDP cases, a differential gene expression analysis with respect to non HDP controls confirms the gene set enrichment analyses and shows a signature dominated by immune genes. In the top 50 significantly differentially expressed genes 28% overlap with GO categories related to immune processes, including complement activation (GO:0006956, 28-fold enrichment, FDR adjusted p = 0.04), adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains (GO:0002460, 11-fold enrichment, FDR adjusted p = 0.03), and innate immune response (GO:0045087, 5-fold enrichment, FDR adjusted p = 0.04). Importantly, ten of these immune genes are shared between those with PE at term without preterm diagnosis with severe features (Fig. 3a) and twelve are shared between those with GHTN (Fig. 3b). Across immune-associated HDP, term PE without preterm diagnosis with severe features, and GHTN, eight genes are related to immune GO biological processes indicating that these distinct clinical phenotypes of HDP are in fact, molecularly similar.
Twenty-two genes (Fig. 3c) were found to overlap between the 29 differentially expressed genes found in immune-associated PE (Fig. 3a) and the 38 differentially expressed genes found in immune-associated GHTN (Fig. 3b) relative to non-HDP baseline by performing differential gene expression analysis across 2036 pregnancy genes based on a two-sided Mann–Whitney U rank test with multiple test correction by Benjamini–Hochberg. To assess the significance of the observed overlap between two sets, we utilized the hypergeometric distribution. This determines the probability that a given overlap size (or greater) would occur by random chance, assuming the elements are sampled without replacement from a larger population. To calculate the significance of the observed overlap, we computed the cumulative probability of observing an overlap of size X or greater, summing the hypergeometric probabilities for all values from X up to a size of the smallest of two sets. This cumulative probability represents the likelihood of observing an overlap of size X or larger by random chance under the null hypothesis that the sets are independent. We implemented this calculation using the dhyper() function in R64.
Placental-associated classifier
Gene features were selected by differential expression analysis based on a Mann–Whitney U rank test (scipy v1.12.065) for placental-associated cases separately for all samples and for samples with no high risk factors based on counts from featureCounts. Two clinical metrics (mean arterial blood pressure and BMI) were identified to contribute, above and beyond the cfRNA counts, and were included as model features. Gene features and clinical metrics were identified based on performance on unseen test data across many random train/test splits by cross validation. Six genes and two fixed clinical metrics were then scaled and centered using robust statistics as implemented by the RobustScaler in scikit-learn57, and then modeled with a penalized Lasso regression implemented by scikit-learn57 (LogisticRegression). The decision boundary was set to achieve 73% specificity in training samples.
Sample eligibility was defined as those with greater than five million counts uniquely assigned to the reference calculated by featureCounts, more than 96 percent of reads align with strand 2 (the transcript) calculated by Picard66, and total number of detected genes (excluding spike-in RNA) less than 25,000 to protect against samples with DNA contamination. Additionally, samples without library batch information were excluded.
A total of 259 individuals were excluded from the training cohort and 33 individuals from the validation due to the different assay fail criteria. Individuals were excluded from the study for the following reasons: assay fail (training n = 77, validation n = 9), less than 5 million assigned counts (training n = 86, validation n = 20), more than 25,000 genes with counts (training n = 17, validation n = 4), and/or less than or equal to 96% of reads aligns with the strand of the transcript (training n = 118, validation n = 5). Note, a sample can fail multiple metrics, so the sum of individual metrics can have a total greater than the number of excluded samples.
To lower our risk of identifying spurious or false positive features, we reduce our search space from all genes to a subset identified to changes during pregnancy. To establish a pregnancy gene list, 49 nonpregnant females between 22 and 40 years old were recruited. One nonpregnant sample was omitted from subsequent analyses due to missing height and weight records. Gene search space was defined by identifying genes that are differentially expressed between 48 non-pregnant samples and 38 pregnant individuals without annotated pregnancy complications and matched to the non-pregnant individuals on assigned counts and racial demographics and library batch. After matching healthy, uncomplicated pregnant samples were matched on assigned counts, library batch, and race. There were still residual differences in age and BMI, so the subsequent analyses controlled for these. All samples had more than 5.3 million assigned counts and less than 24,000 non-zero count genes. Differential gene expression analysis was performed over 13,342 genes that had at least 10 counts in at least 10% of the combined samples. The analysis evaluated a pregnancy effect in log2cpm with age and BMI as covariates, with a false discovery rate correction for multiple comparisons. No genes were significantly dependent on age and bmi. We include all genes that are differentially expressed with a log2 fold change greater than 0.5 and an adjusted p-value < 0.001. This list resulted in 2036 pregnancy genes. Genes with mean CPM less than or equal to 1 were also excluded from feature searches.
Evaluation of collection proximity to delivery versus HDP severity
PAPPA2, and our cfRNA model more generally, show the strongest signal for the pregnancies that are ultimately delivered earliest. It’s possible that the mothers at most severe risk carry the highest signal, but it’s also possible that event proximity could be driving this pattern, where the signal is noisier when predicting events that are further into the more distant future (i.e. the later deliveries). We can operationalize proximity as the difference between the gestational age at collection and the gestational age at delivery, and because there is some variation in gestational age at collection in our samples (range 17.5–22 weeks), proximity and gestational age at delivery can be distinguished.
To determine whether our model predictions appear to be driven more by severity or proximity, we evaluated a linear regression model gene_model_prediction_logits ~ delivery_weeks + proximity_weeks, fit over all placental-associated HDP training samples (which carry natural variation in gestational age at delivery). Proximity didn’t confer any information that wasn’t already accounted for by delivery GA (bootstrap p = 0.2688) but delivery GA did explain variance in model predictions holding proximity constant (bootstrap p = 0.0042). These data indicate that our model predictions are preferentially sensitive to the biology of more severe risk, rather than mere event proximity.
Given this relationship, we asked whether our 6-gene model could be used to predict variation in gestational age at delivery, holding proximity constant, using the model delivery_weeks ~ gene_model_prediction_logits + proximity_weeks. And indeed, in the same training cases, we saw a significant effect of our model predictions (bootstrap p = 0.0036). Conclusions remained the same even when spontaneous preterm births were removed from this analysis. In other words, even for different samples where the number of days between blood draw and delivery is held constant, our cfRNA signal still stratifies these samples by severity of risk, with the more preterm deliveries (e.g. blood draw at 18 weeks and delivery at 32 weeks) showing the greater signal (compared to e.g. blood draw at 22 weeks and delivery at 36 weeks).
Evaluating days spent in NICU by classifier prediction
Of those newborns who spent time in the NICU, the distribution of days spent in the NICU was compared between individuals predicted to be placental-associated HDP versus those who were not in individuals without high risk factors. In this analysis, individuals without high risk factors included those who have PE and those without PE or spontaneous preterm birth. Individuals were also filtered to exclude individuals who went to the NICU for reasons not related to PE, such as for term birth with chorioamnionitis (n = 21), fetal anomaly (n = 6), maternal diabetes (n = 5), or maternal medication not related to diabetes (n = 4), and spontaneous preterm birth without PE (n = 21). The comparison group is composed of 81 test positive individuals (35 newborns went to the NICU for preterm birth and 46 at term birth) and 154 test negative individuals (8 newborns went to the NICU for preterm birth and 146 at term birth).
Similar results where test positive individuals spent more time on average in the NICU were observed in cross validation on the training data, where the training data was randomly split into a training subset using 65% of the samples with the remainder of samples allocated to a mutually exclusive test subset for evaluation. This random split was repeated 500 times to estimate the distribution of performance. Within each split, features were selected and a model was trained based on the training subset, and the resultant model was evaluated on the corresponding test subset. The 500 model performances across the test sets were combined, and the proportion of times an individual was predicted as placental-associated HDP was calculated for those individuals without high risk factors who spent time in the NICU who either have PE or are without PE or spontaneous preterm birth with exclusions for NICU reasons not related to PE as described above. Those who were predicted to be placental-associated HDP in more than fifty percent of the 500 splits (34 newborns went to the NICU for preterm birth compared to 44 at term birth) were in the NICU on average 9.0 days longer (14.4 days vs 5.4 days, respectively) compared to those who were predicted positive in less than fifty percent of the splits or negative in all splits (9 newborns went to the NICU for preterm birth and 147 at term birth) (two-sided Kolmogorov–Smirnov test p = 1.4×10−3). When considering test negative individuals as those who were never predicted in any of the cross validation splits to be placental-associated HDP, test positive individuals were in the NICU on average 5.96 days longer (11.2 days versus 5.3 days, respectively for test positive (40 newborns went to the NICU for preterm birth compared to 82 at term birth) and test negative (3 newborns went to the NICU for preterm birth and 109 at term birth) individuals (two-sided Kolmogorov–Smirnov test p = 8.0×10−3).
Statistical methods
Unless otherwise specified, results are reported for log2cpm gene counts calculated by featureCounts corrected for batch, gestational age at collection, and inferred fetal sex, as described above. Placental genes were additionally corrected for fetal fraction indicator genes as described above. Cohen’s d effect sizes presented in figures and text are computed using pooled standard deviation, with parametric confidence intervals. Differential gene expression analyses (shown in Figs. 3, 4, Supplementary Figs. and reported throughout the text) were performed via Mann–Whitney U tests over each gene in the search space of pregnancy genes (see above) without continuity correction, and p-values were adjusted by the Benjamini–Hochberg (non-negative) procedure to correct for multiple testing at alpha = 0.05. Confidence intervals for AUC, sensitivity, and specificity presented using holdout validation data in Fig. 5 were computed via bootstrap.
Gene Set Enrichment Analysis
Gene Set Enrichment Analysis (GSEA) was carried out using the R package fgsea v.1.30.067 using default settings. Difference between group-level mean log2cpm values, which is equivalent to log2 fold change, was used as a gene-level measure in the analysis. Gene lists used in the analysis were from the Molecular Signature Database68, the Human Protein Atlas56,59, and the STRING database69.
Out of 16,008 gene sets annotated by gene ontology terms, 373 were significantly enriched (FDR < 0.05) in placental-associated samples relative to immune-associated samples and 140 were significantly enriched in immune-associated relative to placental-associated. Twenty most enriched sets along with their respective relative leading edge sizes (Fraction of gene set), K-S test adjusted p-value and the normalized enrichment score (NES) are shown in Supplementary Fig. 2.
To analyze a possible tissue or cell-type origin of cfRNA’s we used consensus transcript levels from 55 human tissues compiled from the Human Protein Atlas, HPA56, and The Genotype-Tissue Expression60 data as well as transcript expression levels summarized per gene in 81 cell types from HPA59. The most likely cell type and tissue origin of the model genes’ cfRNA transcription profiles were identified using the following procedure. For each model gene, we established the null correlation distribution: if a model gene is not expressed or lowly expressed in a cell type or tissue, then the distribution of linear correlations between the gene’s cfRNA transcription profile and the cfRNA profiles of genes specific to that cell type represents a quasi-null distribution. These null distributions were aggregated across all cell types or tissues where the model transcript was expressed at a level of less than 10 transcripts per million. The aggregated distributions were used to establish gene-specific thresholds, above which correlations were considered significant.
The fraction of a specific set (elevated in cell types or tissue-specific at tau >0.970) that correlated with the model gene above the threshold corresponded to a conditional likelihood that the model gene’s profile is associated with the tissue-specific transcriptional profile. The reported fractions were calculated at above the 99th percentile of each of the gene-specific null distributions. To reduce the skewness of the null distribution, Pearson correlation coefficients were Fisher transformed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings described in this manuscript are available in the article, in the Supplementary Information or as Source data available from the corresponding author upon request and under controlled access. Given privacy restrictions on identifiable data, non-commercial use of these data is possible upon request, subject to such restrictions and execution of a Data Transfer and Use Agreement with Mirvie Inc. where data are housed. Inquiries for data usage can be directed to research@mirvie.com, and all requests will be reviewed within 30 days.
Code availability
Code is available in the following link: https://doi.org/10.5281/zenodo.14681170.
References
Fink, D. A. et al. Trends in Maternal Mortality and Severe Maternal Morbidity During Delivery-Related Hospitalizations in the United States, 2008 to 2021. JAMA Netw. Open 6, e2317641 (2023).
Cameron, N. A. et al. Trends in the Incidence of New-Onset Hypertensive Disorders of Pregnancy Among Rural and Urban Areas in the United States, 2007 to 2019. J. Am. Heart Assoc. 11, e023791 (2022).
Motomura, K., Morita, H., Naruse, K., Saito, H. & Matsumoto, K. Implication of viruses in the etiology of preeclampsia. Am. J. Reprod. Immunol. 91, e13844 (2024).
Collier, A.-R. Y., Smith, L. A. & Karumanchi, S. A. Review of the immune mechanisms of preeclampsia and the potential of immune modulating therapy. Hum. Immunol. 82, 362–370 (2021).
Roberts, J. M. & Escudero, C. The placenta in preeclampsia. Pregnancy Hypertens. 2, 72–83 (2012).
Roberts, J. M. & Bell, M. J. If we know so much about preeclampsia, why haven’t we cured the disease? J. Reprod. Immunol. 99, 1–9 (2013).
Davidson, K. W. et al. Aspirin use to prevent preeclampsia and related morbidity and mortality: US preventive services task force recommendation statement. JAMA 326, 1186–1191 (2021).
ACOG. Gestational hypertension and preeclampsia: ACOG practice bulletin, number 222. Obstet. Gynecol. 135, e237–e260 (2020).
Wheeler, S. M., Myers, S. O., Swamy, G. K. & Myers, E. R. Estimated prevalence of risk factors for preeclampsia among individuals giving birth in the US in 2019. JAMA Netw. Open 5, e2142343 (2022).
Sacks, D. A. & Incerpi, M. H. Of aspirin, preeclampsia, and racism. N. Engl. J. Med. 390, 968–969 (2024).
Krishnamurti, T. et al. Use of a smartphone app to explore potential underuse of prophylactic aspirin for preeclampsia. JAMA Netw. Open 4, e2130804 (2021).
Zeisler, H. et al. Predictive Value of the sFlt-1:PlGF Ratio in Women with Suspected Preeclampsia. N. Engl. J. Med. 374, 13–22 (2016).
Myatt, L. The prediction of preeclampsia: the way forward. Am. J. Obstet. Gynecol. 226, S1102–S1107.e8 (2022).
ClinicalTrials.gov. https://clinicaltrials.gov/study/NCT06074601 (2024).
E6(R2) Good Clinical Practice: Integrated Addendum to ICH E6(R1)|FDA. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/e6r2-good-clinical-practice-integrated-addendum-ich-e6r1 (2018).
Rasmussen, M. et al. RNA profiles reveal signatures of future health and disease in pregnancy. Nature 601, 422–427 (2022).
Cohen, J. Statistical power analysis for the behavioral sciences, 79–81 (Lawrence Erlbaum Associates, 1988). https://doi.org/10.4324/9780203771587.
Sawilowsky, S. S. New effect size rules of thumb. J. Mod. Appl. Stat. Meth. 8, 597–599 (2009).
Than, N. G. et al. Integrated systems biology approach identifies novel maternal and placental pathways of preeclampsia. Front. Immunol. 9, 1661 (2018).
Hesson, A. M., Langen, E. S., Plazyo, O., Gudjonsson, J. E. & Ganesh, S. K. Placental transcriptome analysis of hypertensive pregnancies identifies distinct gene expression profiles of preeclampsia superimposed on chronic hypertension. BMC Med. Genomics 16, 91 (2023).
El-Sherbiny, W. & Nasr, A. & Soliman, A. Metalloprotease (ADAM12-S) as a predictor of preeclampsia: correlation with severity, maternal complications, fetal outcome, and Doppler parameters. Hypertens. Pregnancy 31, 442–450 (2012).
Macintire, K. et al. PAPPA2 is increased in severe early onset pre-eclampsia and upregulated with hypoxia. Reprod. Fertil. Dev. 26, 351–357 (2014).
Tsoutsouki, J., Patel, B., Comninos, A. N., Dhillo, W. S. & Abbara, A. Kisspeptin in the prediction of pregnancy complications. Front Endocrinol. 13, 942664 (2022).
Textoris, J. et al. Evaluation of current and new biomarkers in severe preeclampsia: a microarray approach reveals the VSIG4 gene as a potential blood biomarker. PLoS ONE 8, e82638 (2013).
Tsuchida, N. et al. Transcriptomic features of trophoblast lineage cells derived from human induced pluripotent stem cells treated with BMP 4. Placenta 89, 20–32 (2020).
Faas, M. M., Spaans, F. & De Vos, P. Monocytes and macrophages in pregnancy and pre-eclampsia. Front. Immunol. 5, 298 (2014).
Joshi, N. S. et al. Admission and care practices in united states well newborn nurseries. Hosp. Pediatr. 13, 208–216 (2023).
Lamminpää, R., Vehviläinen-Julkunen, K., Gissler, M. & Heinonen, S. Preeclampsia complicated by advanced maternal age: a registry-based study on primiparous women in Finland 1997-2008. BMC Pregnancy Childbirth 12, 47 (2012).
Roberts, J. M. et al. Care plan for individuals at risk for preeclampsia: shared approach to education, strategies for prevention, surveillance, and follow-up. Am. J. Obstet. Gynecol. 229, 193–213 (2023).
Admati, I. et al. Two distinct molecular faces of preeclampsia revealed by single-cell transcriptomics. MED 4, 687–709.e7 (2023).
Ghaemi, M. S. et al. Proteomic signatures predict preeclampsia in individual cohorts but not across cohorts - implications for clinical biomarker studies. J. Matern. Fetal Neonatal Med. 35, 5621–5628 (2022).
Yagel, S. et al. Expert review: preeclampsia Type I and Type II. Am. J. Obstet. Gynecol. MFM 5, 101203 (2023).
Cirrik, S. & Schmid-Schönbein, G. W. IGF-1 receptor cleavage in hypertension. Hypertens. Res. 41, 406–413 (2018).
Lai, W. & Yu, L. Insulin-like growth factor 1 ameliorates pre-eclampsia by inhibiting zinc finger E-box binding homeobox 1 by up-regulation of microRNA-183. J. Cell. Mol. Med. 27, 1179–1191 (2023).
Crosley, E. J., Dunk, C. E., Beristain, A. G. & Christians, J. K. IGFBP-4 and -5 are expressed in first-trimester villi and differentially regulate the migration of HTR-8/SVneo cells. Reprod. Biol. Endocrinol. 12, 123 (2014).
Winship, A. L. et al. Interleukin-11 alters placentation and causes preeclampsia features in mice. Proc. Natl Acad. Sci. USA 112, 15928–15933 (2015).
Ronzoni, S. et al. Preterm preeclampsia screening and prevention: a comprehensive approach to implementation in a real-world setting. BMC Pregnancy Childbirth 25, 32 (2025).
Adil, M. et al. Preeclampsia risk prediction from prenatal cell-free DNA screening. Nat. Med. https://doi.org/10.1038/s41591-025-03509-w (2025).
Cowan, A., Haverty, C., MacDonald, R. & Khodursky, A. Impact of early preeclampsia prediction on medication adherence and behavior change: a survey of pregnant and recently-delivered individuals. BMC Pregnancy Childbirth 24, 196 (2024).
Mamudu, H. M., Paul, T. K., Veeranki, S. P. & Budoff, M. The effects of coronary artery calcium screening on behavioral modification, risk perception, and medication adherence among asymptomatic adults: a systematic review. Atherosclerosis 236, 338–350 (2014).
André, F., Rassy, E., Marabelle, A., Michiels, S. & Besse, B. Forget lung, breast or prostate cancer: why tumour naming needs to change. Nature 626, 26–29 (2024).
Roberts, J. M. et al. Subtypes of preeclampsia: recognition and determining clinical usefulness. Hypertension 77, 1430–1441 (2021).
Deepak, V., El-Balawi, L. & Harris, L. K. Placental Drug Delivery to Treat Pre-Eclampsia and Fetal Growth Restriction. Small e2311165 https://doi.org/10.1002/smll.202311165 (2024).
Tan, L. et al. Statins prevent the deleterious consequences of placental chemerin upregulation in preeclampsia. Hypertension 81, 861–875 (2024).
Rolnik, D. L. et al. Aspirin versus Placebo in Pregnancies at High Risk for Preterm Preeclampsia. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa1704559 (2017).
Camunas-Soler, J. et al. Predictive RNA profiles for early and very early spontaneous preterm birth. Am. J. Obstet. Gynecol. 227, 72.e1–72.e16 (2022).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
BCL Convert Support. https://support.illumina.com/sequencing/sequencing_software/bcl-convert.html (2024).
ValidateFastq. https://biopet.github.io/validatefastq/0.1.1/index.html (2018).
GitHub - stjude-rust-labs/fq: Command line utility for manipulating Illumina-generated FASTQ files. https://github.com/stjude-rust-labs/fq (2024).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Sambamba: process your BAM data faster! https://lomereiter.github.io/sambamba/ (2022).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Putri, G. H., Anders, S., Pyl, P. T., Pimanda, J. E. & Zanini, F. Analysing high-throughput sequencing data in Python with HTSeq 2.0. Bioinformatics https://doi.org/10.1093/bioinformatics/btac166 (2022).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Miltoft, C. B. et al. Contingent first-trimester screening for aneuploidies with cell-free DNA in a Danish clinical setting. Ultrasound Obstet. Gynecol. 51, 470–479 (2018).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 7, eabh2169 (2021).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Ashburner, M. et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Gene Ontology Consortium et al. The Gene Ontology knowledgebase in 2023. Genetics 224, iyad031 (2023).
Thomas, P. D. et al. PANTHER: Making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna, Austria, 2023).
Oliphant, T. E. Python for Scientific Computing. Comput. Sci. Eng. 9, 10–20 (2007).
Picard toolkit. https://broadinstitute.github.io/picard/ (2019).
Korotkevich, G. et al. Fast gene set enrichment analysis. BioRxiv https://doi.org/10.1101/060012 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
Acknowledgements
We thank all blood donors whose contributions made this study possible; Dr. Alison Cowan for critical discussions on the molecular subtypes, input on clinical definitions and organization; Drs. Andrea Edlow, Michelle Owens and Laura Goetzl for serving on the adjudication committee; Michelle Li, Valerie Yu, Christopher Lee, Lorraine Li, and Andrew Yip for assistance with laboratory work; Aram Saravani for input on analyses; Reece McDonald, Frank Troxell and Curebase (Storm Stillman) for help with direct-to-participant recruitment; Nan Zhang for discussions on biostatistics; Joel Adamich for assistance with figures; Hong Frank, Karandeep Kaur, Honiko Morrison, Monica Jenema, Jaquelyn Bartlett, Amanda Wigger, Logan Brown, Erica Strain, Tracy Cruess, Sierra Loomis, Leanne Welshan, Feyishara Oreagba, Deborah Tweedy, Ashley Hassman, Erin Kuroiwa, Althea Agnostopoulos, and Turquoise Mann for help with clinical data abstraction; Research coordinators across all clinical sites for their help securing samples; Precia Group (Donna Russell, Amber Sexton, Samantha Udondem, Amy Rush, Rohit Nariya) for help with identifying clinical sites.
Author information
Authors and Affiliations
Contributions
M.A.E., C.H., M.L., E.N., M.J., and M.Rasmussen conceptualized and designed the study with input from K.R., W.A.G., J.R.B., C.G.-B., and A.J. and the remaining authors. R.B., D.G.K., P.M.S., G.R.S., A.S., E.B.C., T.F.M. V.B., A.Y.C., A.I.F., E.P.-H., L.D.P., E.F.S., K.R., W.A.G., J.R.B., C.G.-B., and A.J. provided samples and data to the study, A.B.M, I.A., and M.L. curated the data collection and obtained approvals for use in this study where required, with input from sample providers. E.N. and M.Reddy designed laboratory protocols; laboratory experiments were carried out by M. Reddy, J.L., K.M., H.C. E.P.S.G, N.D.-B., A.K., F.S., R.N., S.R.Q., M.A.D., and M. Rasmussen conceptualized computational analyses; E.P.S.G, N.D.-B., A.K., F.S., and R.N. implemented and reviewed code. M.A.E., E.P.S.G, N.D.-B., M.J., and M.Rasmussen drafted the manuscript with critical input from all authors.
Corresponding author
Ethics declarations
Competing interests
M.A.E., E.P.S.G, N.D.-B., A.B.M., M. Reddy, A.K., J.L., I.A., K.M., H.C., F.S., R.N., T.F.M., S.R.Q., M.A.D., C.H., M.L., E.N., M.J. and M. Rasmussen have an equity interest in Mirvie. Mirvie Inc. provided funding for the study. The following patents and patent applications cover various aspects described in the manuscript, including detecting presence or elevated risk of pregnancy complications such as preeclampsia in a pregnant subject based on differential expression of RNA markers: US Patent No 11,208,693 (Mirvie Inc, M.J., E.N., M.Rasmussen, F.S., M.Reddy), US Application No. 18/167,322 (Mirvie Inc, M.J., E.N., M.Rasmussen, F.S., M.Reddy, E.P.S.G, A.K., R.N., M.L.), and International Application No PCT/US2024/027,444 (Mirvie Inc, M.J., E.N., M.Rasmussen, F.S., M.Reddy, E.P.S.G., A.K., R.N., M.L., N.D.-B.). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Elovitz, M.A., Gee, E.P.S., Delaney-Busch, N. et al. Molecular subtyping of hypertensive disorders of pregnancy. Nat Commun 16, 2948 (2025). https://doi.org/10.1038/s41467-025-58157-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58157-y
This article is cited by
-
Protein networks are influenced by maternal BMI and differentiate preterm birth types
Communications Medicine (2026)
-
Early gestational prediction of spontaneous preterm birth using a validated three-protein serum biomarker panel
BMC Medicine (2026)
-
Raised Leptin and Pappalysin2 cell-free RNAs are the hallmarks of pregnancies complicated by preeclampsia with fetal growth restriction
Nature Communications (2025)
-
Cell type inference in cell-free nucleic acid liquid biopsy
Nature Biotechnology (2025)
-
Multi-omic insights of preeclampsia and cardiovascular health outcomes
Communications Medicine (2025)