Abstract
While it has been suggested that alterations in the composition of gut microbial metabolites may play a causative role in the pathophysiology of autism spectrum disorder (ASD), it is not known how gut microbial metabolites are associated with ASD-specific brain alterations. In this cross-sectional, case-control observational study, (i) fecal metabolomics, (ii) task-based functional magnetic resonance imaging (fMRI), and (iii) behavioral assessments were obtained from 43 ASD and 41 neurotypical (NT) children, aged 8–17. The fMRI tasks used socio-emotional and sensory paradigms that commonly reveal strong evoked brain differences in ASD participants. Our results show that fecal levels of specific tryptophan-related metabolites, including kynurenate, were significantly lower in ASD compared to NT, and were associated with: 1) alterations in insular and cingulate cortical activity previously implicated in ASD; and 2) ASD severity and symptoms (e.g., ADOS scores, disgust propensity, and sensory sensitivities). Moreover, activity in the mid-insula and mid-cingulate significantly mediated relationships between the microbial tryptophan metabolites (indolelactate and tryptophan betaine) and ASD severity and disgust sensitivity. Thus, we identify associations between gut microbial tryptophan metabolites, ASD symptoms, and brain activity in humans, particularly in brain regions associated with interoceptive processing.
Similar content being viewed by others
Introduction
The gut microbial ecosystem generates an estimated 40% of all metabolites in the body’s circulation, including neuroactive and inflammatory molecules1,2. Within the brain-gut-microbiome (BGM) system, neuroactive gut metabolites can modulate brain activity directly, via the systemic circulation, or via vagal and spinal afferents. Ninety percent of vagal fibers are afferents, underscoring the magnitude of information that is transmitted from the gut, its microbiome, and from vagal signals to the brain3,4. Together, these regulatory signals from the gut microbiome can influence socio-emotional and sensory processing, cognition, and behavior5,6,7,8. Pre- and postnatal microbial and neural disruptions may have profound effects on the development of the enteric and central nervous system9. Such disruptions have been linked to the etiology of neurodevelopmental disorders, such as autism spectrum disorder (ASD)10,11,12,13,14.
Indeed, longitudinal studies in ASD indicate that gut microbial dysbiosis can be found in infancy15 and persist through adulthood16,17. Early life gut dysbiosis may be associated with compromised barriers within the BGM system, contributing to the development of neurodevelopmental disorders13,18,19. Further, there is evidence from cross-sectional human studies indicating that gut microbiome and metabolite alterations correlate with ASD symptoms (i.e., difficulties in socio-emotional behavior and sensory perception), which can be ameliorated with probiotics or fecal microbiota transplants from neurotypical (NT) children20,21,22,23. The high prevalence of gastrointestinal (GI) symptoms in ASD (46–84%) further underscores the gut as an important component in ASD pathophysiology3,5,24,25.
Of the gut microbial metabolites that may be relevant to ASD pathophysiology, there has long been an interest in those in the tryptophan/serotonin pathway (Fig. 1). Roughly 30% of individuals with ASD show elevated blood serotonin levels, which have been linked to dysregulation of symptoms such as mood, appetite, and social interactions26. In addition, individuals with ASD with hyper-serotonemia are more likely to experience GI issues27,28. However, reducing dietary tryptophan, which is the precursor for 95% of the body’s serotonin, has been associated with increased ASD symptomatology29,30,31,32. Serotonin, generated by intestinal enterochromaffin cells, is almost completely taken up by platelets. However, as serotonin does not cross the blood–brain barrier, serotonin signals are thought to reach the brain via serotonin-mediated vagal signaling26,33,34. Gut microbes also play an important role in the metabolism of dietary tryptophan into indoles, kynurenate (KA) and kynurenine (KYN), which, in contrast to serotonin, are able to cross the blood–brain-barrier. While KA has neuroprotective effects, KYN has neurotoxic effects especially during critical periods of development, influencing brain microstructure, activity, and development5,35,36. Thus, there is evidence that dysregulation in the tryptophan metabolism and KYN pathway may influence ASD symptomatology both by peripheral and central mechanisms, at least in a subset of individuals.
Arrow thickness represents the strength of the pathway under normal conditions.
Some animal studies have examined how the brain may mediate relationships between gut metabolites and behavior13,37. However, to date, there are no human ASD studies that have looked at interactions between gut metabolites and brain activity, and their impact on ASD symptomatology. In this cross-sectional comparative study, 43 ASD and 41 neurotypical (NT) youth (aged 8–17) underwent functional magnetic resonance imaging studies (fMRI), fecal metabolomics focused on the tryptophan pathway, and comprehensive clinical and behavioral assessments. We focused on interoceptive, emotional, and sensory processing brain regions of interest (ROIs) based on prior studies in children with ASD and their relevance to vagal processing (see “Methods: fMRI Tasks”). The fMRI tasks all involved socio-emotional processing (processing emotions, facial expressions, others’ somatosensory experiences); such socio-emotional processing has been shown to be modulated by tryptophan metabolites, like serotonin31,32,38. Our primary hypotheses were that the relative abundances of tryptophan-related gut metabolites differs between ASD and NT groups, and correlate with ASD symptomatology as well as with atypical brain activity in regions consistently implicated in ASD (i.e., insular subregions, pregenual anterior, and mid-cingulate). While the study design did not allow us to address causality, we additionally explored the hypothesis that task-based brain activity mediates metabolite and behavioral relationships.
Results
Behavioral measures
Independent samples t-tests and Fisher’s exact tests were used to determine significant ASD-NT differences in demographic and behavioral variables. As shown in Supplemental Table 1, compared to the NT group (N = 41; 20F; mean age = 11.6), the ASD group (N = 43; 11F; mean age = 12.06) showed significantly higher values for: body mass index (BMI), prenatal antibiotic use, gastrointestinal (GI) symptoms, sensory sensitivities, attention deficit/hyperactivity disorder (ADHD) symptoms, disgust sensitivity, disgust propensity, and social difficulties. In addition, the ASD group showed significantly lower scores on the full-scale intelligence quotient (FSIQ) and sleep quality (all ps < 0.05). No between-group differences in diet were found (see Supplemental Table 1 and Supplemental fig. 1). Refer to Supplemental Table 2 for the participant’s race and ethnicity by group.
Between-group differences in tryptophan metabolites
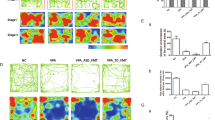
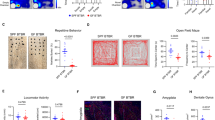
Regarding stool-derived metabolites, as compared to the NT group, the ASD group displayed significantly lower KA levels (β = 0.78, se = 0.23, q = 0.02, Cohen’s d = 0.82; Fig. 2). Other tryptophan pathway metabolites such as xanthurenate (Cohen’s d = 0.49), 5-hydroxypicolinic acid (Cohen’s d = 0.41), and indolepropionate (Cohen’s d = 0.39) had small to moderate effect sizes but did not achieve FDR-corrected statistical significance at q < 0.05 (Supplemental Table 3).
A contrast analysis within the framework of the GLM indicated significant differences between ASD (N = 43) and NT (N = 41) groups in kynurenate (KA; q = 0.02, two-tailed). NT: minimum = −1.15, 1st quartile = −0.23, median = 0.51, 3rd quartile = 1.00, maximum = 1.45. ASD: minimum = −2.68, 1st quartile = −0.98, median = −0.18, 3rd quartile = 0.75, maximum = 1.30. Source data are provided as a Source data file.
Given that in our sample there were greater reports of GI symptoms in ASD compared to NT, in a subset of participants with this data available, we performed a sensitivity analysis including GI symptoms as a covariate. The significant group differences in KA abundance persisted (β = 1.11, se = 0.29, q = 0.009, Cohen’s d = 1.15). Further, indoleacrylate (Cohen’s d = 0.76), indolepropionate (Cohen’s d = 0.62), 5-hydroxypicolinic acid (Cohen’s d = 0.58), and xanthurenate (Cohen’s d = 0.53) had large effect sizes but did not achieve FDR-corrected statistical significance at q < 0.05 (Supplemental Table 4). In addition, including diet as a covariate did not change the results.
Brain-metabolite correlations in the ASD group
We primarily focused on brain regions involved in interoceptive, vagal, and emotion processing, as these may be especially impacted in altered gut-brain interactions. In addition, brain regions of interest (ROIs) were selected from those which showed significant differences in ASD compared to NT based on prior studies and our fMRI tasks (tasks included: processing others’ facial expressions/actions; disgust processing; processing somatosensory experiences; see Methods: fMRI Tasks and Supplemental Table 5). As Fig. 3 shows, ROIs included the: cingulate cortex (right mid-cingulate, pregenual anterior cingulate, anterior mid-cingulate cortex/dorsal medial prefrontal cortex), bilateral insula (anterior, dorsal anterior, mid, ventral, posterior), right fusiform face area (FFA), right inferior gyrus pars opercularis (IFGop), and right primary somatosensory cortex (S1).
See Supplemental Table 5 for peak MNI coordinates for each ROI. While our predominant focus was on subregions of the insula and cingulate due to their involvement in interoceptive and emotional processing, we additionally considered other ROIs with significant ASD vs. NT differences in our fMRI tasks and from prior ASD studies. Please see “Methods” for how ROIs were selected. R Right, L Left, insular subregions (dAI left dorsal anterior insula, vAI ventral anterior insula, MI mid-insula, PI posterior insula); cingulate subregions (pACC pregenual anterior cingulate cortex, MCC mid-cingulate cortex, dmPFC/aMCC dorsal medial prefrontal cortex/anterior mid-cingulate cortex); IFGop inferior frontal gyrus, pars opercularis, S1 primary somatosensory cortex.
General linear models (GLMs) were applied to assess associations between brain regions showing differences on the stimulus-evoked tasks with metabolites, controlling for age, sex, FSIQ, and BMI. See Table 1 for metabolite-brain association results for the ASD group (see Supplemental Table 8 for NT results).
During facial expression processing, increased activity in the right IFGop was significantly associated with higher levels of anthranilate, while decreased activity in the right mid-cingulate cortex (MCC) was significantly associated with higher levels of tryptophan betaine. Activity in the right MCC when processing hand actions was significantly associated with increased abundance of N-acetyltryptophan. In addition, higher right MCC activity during processing of non-emotional faces was significantly associated with increased c-glycosyltryptophan.
For the disgust processing task, significant ROI-metabolite correlations primarily involved insular subregions. Notably, lower levels of indolelactate were significantly associated with increased activity in the right mid-insula when viewing disgusting foods. When viewing disgust facial expressions, increased activity in the left dorsal anterior insula was associated with higher levels of KA.
For the somatosensory task, increased levels of indole-3-carboxylate were significantly associated with increased activity in right S1 and dorsal medial prefrontal cortex /anterior mid-cingulate cortex (dmPFC/aMCC) when processing object touch. By contrast, decreased levels of indole-3-carboxylate were associated with activity in the left posterior insula when processing social touch.
Brain-behavioral association results within the ASD group are shown in Table 2 (see Supplemental Table 6 for additional significant associations and Supplemental Table 10 for NT results). Of note, increased right MCC activity related to non-emotional facial processing was positively associated with disgust sensitivity across various stimuli conditions. Further, when looking at disgusting foods, decreased activity in the right mid-insula was significantly associated with autism severity (ADOS-2 RRB and ADOS-2 total score), and activity in the left ventral anterior insula was associated with restricted, repetitive, and stereotyped patterns of behavior (ADI-R RRB).
Behavioral associations with select tryptophan metabolites within the ASD group
The above results indicated that 7 tryptophan metabolites were significantly correlated with alterations in brain activity in ASD. We applied GLMs to investigate how these 7 metabolites may also be related to behavior in the ASD group, while controlling for age, sex, BMI, and FSIQ. As Table 3 shows, we found large effect size associations between metabolites and behavioral variables. Higher levels of tryptophol were significantly correlated with sensory sensitivities (vestibular/proprioception, social, and non-social). In addition, higher levels of anthranilate were significantly correlated with greater quality of sleep. Refer to Supplementary Table 7 for the results of a post hoc analysis of tryptophan metabolite-behavior associations within ASD with all tryptophan metabolites. NT results are presented in Supplemental Table 9.
Mediation analysis
Based on previous theoretical work and animal studies indicating that metabolites affect brain activity which in turn affect behavior13,37,39, we conducted exploratory mediation analyses, with the brain activity in regions that differed between ASD and NT as the mediator between tryptophan metabolite-behavior relationships. As Fig. 4 shows, mediation models (controlling for brain-related variables, age and FSIQ) indicated significant mediating effects for the right mid-insula and the right mid-cingulate. Specifically, activity in the right mid-insula during disgust processing statistically mediated the relationship between indolelactate and ASD severity as measured by the ADOS RRB score (indirect effect: Std. β = 0.341, SE = 0.227, 95% CI [0.016,0.967] and ADOS total score (indirect effect: Std.β = 0.440, SE = 0.585, 95% CI [0.005, 0.984]). In addition, activity in the right mid-cingulate during facial expression processing mediated the relationship between tryptophan betaine and disgust sensitivity (indirect effect: Std. β = −0.328, SE = −0.944, 95% CI [−0.610, −0.075]). We note that while the significant statistical mediation effects observed in these models are consistent with the conceptual model whereby gut metabolites impact brain activity increasing ASD symptomatology, longitudinal and/or interventional experiments (e.g., fecal transplant studies at critical periods of development) are necessary to determine causation. A summary of the main results in ASD is displayed in Table 4. The summary of NT results is available in Supplemental Table 11.
Significant mediation models within the ASD group, with the brain as the mediator between specific metabolites and behavior. For each mediation model, the figure contains the standardized beta and standard error: Std. β (SE) for direct effects. The indirect effect between the metabolite and behavior is listed below the arrow in brackets. * indicates significant indirect effects. A Right mid-insula: disgusting foods vs. rest, indolelactate, and ADOS RRB. B Right mid-insula: disgusting foods vs. rest, indolelactate, and ADOS total score. C right mid-cingulate: non-emotional faces vs. rest, tryptophan betaine, and disgust sensitivity. ADOS Autism Diagnosis Observation Schedule, ADI-R Autism Diagnostic Interview-Revised, RRB Restricted and Repetitive Behaviors, DPSS-R Disgust Propensity and Sensitivity Scale - Revised, r_MI right mid-insula, MCC mid-cingulate cortex. Source data are provided as a Source data file.
Discussion
The current results confirmed our main hypotheses that tryptophan-related fecal metabolites differ between ASD and NT groups and show significant associations with known ASD brain alterations as well as symptomatology, with medium to large effect sizes. Here, we report data consistent with the conceptual model that brain activity mediates the relationship between tryptophan metabolites and ASD symptomatology. Although the current study was not designed to demonstrate causality, these findings represent an important step toward a better mechanistic understanding of ASD.
As Fig. 1 shows, in the gut, more than 90% of dietary tryptophan is metabolized into kynurenine (KYN) and to a lesser degree into serotonin as well as other metabolites. Importantly, any changes in regulation of the KYN pathway, in particular a reduction in plasma levels of kynurenate (KA) and an increase in the KYN/KA ratio, can lead to neurotoxic effects, impact blood serotonin levels, and have profound behavioral and cognitive effects40. Indeed, such dysregulation has been associated with several neuropsychiatric and neurological disorders such as ASD, schizophrenia, depression, and epilepsy41,42,43,44. Here, we found significantly decreased levels of KA in our ASD group compared to NT. This result is consistent with prior research by Bryn et al. (2018) showing lower blood levels of KA in ASD compared to NT children. In that study, the authors suggested that decreased plasma KA levels and an increase in the KYN/KA ratio in ASD may result in less neuroprotection during development41.
There are conflicting data about whether differences in plasma and/or fecal metabolites in ASD may be driven by the high degree of ASD participants with GI issues23,45. However, consistent with Needham et al. (2021), we did not find any significant relationships between fecal KA levels and GI distress in ASD, indicating that our results may not be limited to those with GI symptoms. Whether KA abundance can be used as a biomarker of ASD remains a question for future research23.
In this study, we focused on interactions of tryptophan metabolites with brain activity specifically in brain regions important for interoceptive, vagal, disgust, and socio-emotional processing which have previously been implicated in ASD38,46,47,48,49. Indeed, activity in these ROIs was significantly related to ASD symptomatology and severity (see Table 2). Importantly, we show that in ASD, activity in these ROIs is significantly correlated with fecal abundance of gut-derived metabolites in the tryptophan pathway, including indoles, which have been implicated in previous ASD research50,51. Specifically, we found that different fecal tryptophan metabolites are significantly correlated with task-based brain activity (left pregenual anterior and middle cingulate cortex, insular subregions, right S1) and with ASD severity and symptomatology.
Notably, in line with our hypothesis, we found that in ASD, brain activity in the mid-cingulate associated with social processing modulated the relationship between tryptophan betaine and disgust sensitivity. Tryptophan-related metabolites (i.e., indoles, serotonin) are important peripheral modulators of vagal activity52. Their altered abundance in the gut has been significantly linked to a number of psychological, neurological, and medical symptoms, such as memory loss, long COVID, depression, sleep disturbances, and anxiety52,53. Thus, the altered abundance of fecal tryptophan-related metabolites may modify activity in vagal and enteric afferent terminals in the gut, or after absorption, directly activate brainstem nuclei, influencing upstream MCC brain activity during socio-emotional tasks and impacting emotional processing. The current data support a hypothesized mechanistic link between common ASD differences in brain activity (MCC, insula) and behavior (disgust sensitivity)49,54.
Further, in the ASD group, we specifically found several correlations between indole metabolites, brain activity (insular subregions, dmPFC/aMCC, IFGop, and S1), and symptomatology (autism severity [ADI-R RSI] and alexithymia). In contrast to serotonin, indoles are exclusively generated by gut microbes from tryptophan and therefore strongly implicate gut microbial alterations in ASD symptomatology55. Interestingly, supporting our last hypothesis, we found that disgust processing activity in the right mid-insula mediates the relationship between indolelactate and two measures of autism severity (ADOS total score and ADOS RRBs). This is particularly noteworthy given that: 1) disgust processing is strongly related to interoceptive processing, making it particularly suited for gut metabolite influence; 2) children with ASD commonly have differences in disgust processing49,54; and 3) the mid-insula is a hub of interoceptive, emotion, and chemosensory processing, and is particularly known to show atypical activity and connectivity in ASD46,54,56. Thus, although these findings do not prove causality between gut microbial metabolites and brain and behavior alterations, they are consistent with the concept that indole metabolites generated by gut microbes (and other tryptophan-related metabolites) directly or indirectly impact brain function, contributing to some ASD-related behavioral symptomatology.
Prior studies have also found that differences in levels of fecal indole metabolites in ASD correlate with autism symptomatology23,57,58. For example, Needham et al. (2021) found significant negative correlations between fecal levels of several indoles, such as indolepropionate, indole, n-formlyanthranalic acid, and indole-3-carboxylate and ASD severity (ADI-R)23. The current results expand upon these previous findings by adding that activity in the right mid-insula may be an important mediator between gut microbial generated indole metabolite and ASD behavior.
Interestingly, we found that a history of prenatal antibiotic exposure was significantly higher in the ASD group. Prenatal maternal antibiotic exposure has been shown to affect the maternal microbiome, which plays an important role in the fetal brain’s exposure to altered maternal microbial metabolites. This prenatal mechanism has been hypothesized to play a role in the etiology of ASD14,59. However, in a post hoc analysis using independent samples t-tests, we found no significant differences in metabolite abundances in the tryptophan pathway between ASD participants with and without prenatal antibiotic exposure; further studies are needed on this topic.
There are several important limitations to the current study. We had strict inclusion criteria for the fMRI component (right handedness, FSIQ > 79, 8–17-year-olds, exclusion of other neuropsychiatric and neurological disorders), which limited the sample heterogeneity and size. To account for this, we utilized a hypothesis-driven approach considering only metabolites within the tryptophan pathway and a priori brain ROIs to limit the number of comparisons, as well as multiple comparisons corrections in all GLM analyses. Another limitation is that we had a larger ratio of males to females in our ASD group compared to the NT group. This was largely due to the exceptional challenge of enrolling in-person participants during the COVID-19 pandemic, which made our initial effort to match groups by sex unviable in a timely manner. However, we accounted for this by including sex as a covariate in all our analyses. Further, the study design was cross-sectional and did not allow the assessment of a causal relationship of the gut microbial metabolites with the brain or behavior. While the mediation models are consistent with our hypotheses, they do not prove causality. Future replication studies are needed as well as studies with more heterogeneous, larger samples with groups matched in sex, well-powered longitudinal designs aimed at studying critical periods of development, interventional designs to demonstrate causality, analysis of broader metabolite pathways, targeted metabolomics, as well as metagenomics to better understand relationships between the brain, gut microbiome, and behavior in ASD. Future studies could also leverage Mendelian randomization (MR) analysis to evaluate potential causal relationships between stool metabolite levels and ASD. However, this approach is currently limited by the lack of adequate data. The only stool-based metabolite genome-wide association study (GWAS) available has a very small sample size and is not publicly accessible60. While serum-based GWAS data exist, they are constrained by relatively low sample sizes, which complicates MR analyses. Specifically, the limited number of genome-wide significant single nucleotide polymorphisms (SNPs) that are independent (i.e., not in linkage disequilibrium) poses challenges for their use as robust instrumental variables61. Despite these current limitations, we believe that this may be an important future technique to better assess causality. Finally, the current study focused on fecal metabolites due to their importance in impacting the brain via vagal signaling. However, given the fact that some tryptophan metabolites can pass the intestinal barrier and the blood–brain barrier (e.g., KYN, indoles), future studies should focus on serum metabolites for a more comprehensive picture of metabolite-brain-behavior associations.
We note that a study by Yap and colleagues (2021) found that microbiome differences between ASD and NT samples were driven by less diverse diets, likely due to significantly restricted ASD food preferences, and thus cautioned against claiming a causative role of the microbiome in ASD pathophysiology62. However, in the current study, a post hoc analysis using independent samples t-tests found no group differences in metabolite concentrations between the observed diet types and no associations with diet between groups or significant correlations between diet and metabolites (Supplemental Fig. 2, Supplemental Table 1). We cannot rule out the possibility of unreliable reporting of dietary patterns by ASD participants or their parents could be responsible for the lack of group differences. Future research should include more reliable and in-depth dietary analyses of participants (and their mothers during the prenatal stage), and use tools such as the foodMAST platform’s foodomic analysis63 to circumvent issues with recording children’s diet (e.g., parent recall bias, calculating accurate portion size)64. Nevertheless, the current study addresses many critiques of prior studies suggested by Yap and colleagues62. We included microbiome-relevant factors as covariates (age, sex, BMI), used multiple comparisons corrections in all GLM analyses, excluded participants on antibiotics, prebiotics, and probiotics, and measured and considered other factors potentially affecting the microbiome (sleep, delivery method, GI issues, medication usage).
In summary, our study reveals that atypical activity in several brain regions previously implicated in ASD pathophysiology is associated with several tryptophan metabolites and symptomatology in youth with ASD. Further, the mediation analysis was consistent with a model that the fecal metabolite abundance is associated with brain activity in regions of interest, contributing to ASD symptomatology. Although our study design is unable to demonstrate causality, these findings represent an important step toward mechanistic integrated models of body-brain-behavior relationships in ASD, with potential implications for future interventions. Future studies incorporating longitudinal designs focusing on critical periods of pre- and postnatal development, as well as interventional designs, are needed to further explore these relationships and their relevance in ASD.
Methods
Participants
Participants were recruited from healthcare clinics in Los Angeles, through advertising in the local community and social media, and by word-of-mouth. Inclusion criteria for all participants included: (a) aged 8–17 years old; (b) IQ of at least 79 on either Full-Scale Intelligence Quotient (FSIQ), or Verbal Comprehension Index (VCI) of the Wechsler Abbreviated Scale of Intelligence 2nd edition (WASI-II)65; and c) right-handed as assessed by a questionnaire adapted from Crovitz and Zener66. Exclusion criteria for all participants included: (a) history of head injury with loss of consciousness greater than 5 min; (b) not sufficiently fluent in English or parent who did not have English proficiency (as not all assessments have been validated in other languages); (c) born before 36 weeks of gestation; (d) contraindications to participating in MRI; (e) on probiotics/prebiotics for the past two weeks; and (f) on antibiotics in the past month.
Additional inclusion criteria for the NT group were: (a) no first-degree relatives diagnosed with ASD; (b) a t-score < 65 on the Conners-3AI parent67 indicating no attention deficit hyperactivity disorder; (c) a t-score < 60 on the Social Responsiveness Scale 2nd edition (SRS-2)68 indicating low likelihood of ASD; and (d) no psychological or neurological disorders. For the ASD group, the Autism Diagnostic Observation Schedule (ADOS-2)69 and the Autism Diagnostic Interview-Revised (ADI-R)70 were administered by a research-certified assessor to confirm ASD diagnosis. Of the ASD participants, 9 were taking antidepressants, 1 was taking anticonvulsants, 11 were on stimulants, and 3 were taking antipsychotics at the time of participation (metabolites were adjusted for medication usage, see Methods subsection “Preprocessing of metabolomic data”). No NT participants were taking medication at time of participation. All participants were instructed to abstain from antibiotic usage for 30 days and probiotics for 14 days prior to participation.
This study was approved by the University of Southern California’s Institutional Review Board (Approval Number: UP-19-00522). Prior to their participation, all participants and parents/legally authorized representatives were emailed a copy of the informed consent and assent forms to give them adequate time to review the forms and ask questions. In person, the informed consent, assent forms, and study procedures were reviewed with the parents/legally authorized representatives and children prior to obtaining written informed consent and assent. Data were collected and stored on REDCap.
Behavioral measures
The study took place over two days. On the first day, behavioral measures and assessments were completed, and fMRI task training and desensitization took place. Within 72 h prior to their second visit, participants collected a stool sample. On the second day, participants brought their stool sample into the lab and then participated in the fMRI sessions.
In addition to the screening measures, parents completed the Sensory Experiences Questionnaire (SEQ-3)71 to assess sensory processing and the Screen for Child Anxiety Related Emotional Disorders (SCARED-P)72 to measure anxiety symptoms. Participants completed the Disgust Propensity and Sensitivity Scale-Revised (DPSS-R)73 to assess frequency of disgusting experiences and the emotional impact of disgusting stimuli, the Alexithymia Questionnaire for Children (AQC)74 to measure alexithymia, and the Body Perception Questionnaire-Short Form (BQP-SF)75 to measure interoception. We also collected data on variables that could impact the gut microbiome. The following variables were collected from the parent or the child including: birth delivery method, prenatal antibiotic usage, antibiotic and probiotic usage during infancy, gastrointestinal symptoms (Gastrointestinal Symptom Rating Scale)76, stool consistency (Bristol Stool Form Scale)77, sleep (Adolescent Sleep Wake Scale [ASWS]78; and Family Inventory of Sleep Habits [FISH])79, and current medication usage. For diet, parents were asked to report the type of diet that best reflects what their child consumes on a regular basis with the following options: Standard American (high consumption of processed foods, red meat, poultry, fish, eggs, dairy products, vegetables and fruits), Modified American (high consumption of whole grains, some processed foods, poultry, fish, eggs, dairy, vegetables and fruits), Mediterranean, Paleo, Vegetarian, Gluten Free, Dairy Free, Low FODMAP, or Other. No parents reported their child followed a Low FODMAP or Vegetarian diet. Standard American and Modified American were then grouped into the category “American” (high consumption of whole grains, some processed foods such as frozen and packaged foods as well as whole grain pasta and breads, limited quantities of poultry, fish, eggs and dairy, and vegetables and fruits are consumed in moderate to large quantities) and the Mediterranean, Paleo, Gluten Free, Dairy Free, and Other were grouped into the “Other” category. Refer to Supplementary Fig. 2 for the diet breakdown by group.
Assessment descriptions
Autism Diagnostic Observation Schedule, Second Edition (ADOS-2)69
The ADOS-2 is a standardized, observational assessment to assess and diagnose autism spectrum disorders. Research reliable staff administered the ADOS-2 to children in the ASD group to verify ASD diagnosis. Module 3 or Module 4 were utilized depending on the child’s age. As the two modules have two different scoring algorithms, for data analysis we utilized the Module 3 algorithm to calculate comparable scores for participants that were assessed with Module 4.
Autism Diagnostic Interview-Revised (ADI-R)70
The ADI-R is a structured interview classically used for diagnosing ASD, planning treatment, and distinguishing ASD from other developmental disorders. The ADI-R has three domains: language/communication, reciprocal social interactions, and repetitive behaviors/interests. The ADI-R was administered by a research staff member with the parents of children in the ASD group.
Alexithymia Questionnaire for Children (AQC)74
The AQC is a standardized, self-reported questionnaire for children aged 9-15 to assess three domains of alexithymia including difficulty describing feelings, difficulty identifying feelings, and externally oriented thinking. The difficulty describing feelings and difficulty identifying feelings scores were combined for a two-factor total. The externally oriented thinking sub-score was not used in this study due to its low Cronbach’s alpha in children80.
Adolescent Sleep Wake Scale (ASWS)78
Self-reported sleep quality was measured with the ASWS. The ASWS has five dimensions: going to bed, falling asleep, maintaining sleep, reinitiating sleep, and returning to wakefulness. A full-scale sleep quality score is obtained by taking the mean of the subscales. The full-scale sleep quality score was used in this study.
Body Perception Questionnaire-Very Short Form (BQP-VSF)75
The BPQ-VSF is a self-reported measure of interoception. The very short form includes 12 items with the highest factor loadings to generate high fidelity scores from the original measure81.
Bristol Stool Form Scale77
The Bristol Stool Form Scale is a visual scale of stool density to assess gastrointenstial transit time.
Conners 3rd Edition ADHD Index-Parent (Conners 3AI)67
The Conners 3AI-Parent is a parent reported measure of attention deficit hyperactivity disorder (ADHD) symptoms for children 6–18 years old. It was used as a screening measure for the NT group but not the ASD group, as ADHD is highly co-occurring with ASD82.
Disgust Propensity and Sensitivity Scale-Revised (DPSS-R)73
The child version of the DPSS-R is a self-reported measure used to assess the frequency of disgust experiences (disgust propensity) and the negative emotional impact of disgusting stimuli (disgust sensitivity).
Family Inventory of Sleep Habits (FISH)79
The FISH is a parent-completed questionnaire to assess sleep habits in children. The scale includes questions related to daytime routine, pre-bedtime habits, sleep environment, bedtime routine, and parental habits. The total score was used in this study as an overall measure of sleep habits.
Gastrointestinal Symptom Rating Scale (GSRS)76
The GSRS is a self-reported instrument to assess the presence of gastrointestinal symptoms.
Screen for Child Anxiety Related Emotional Disorders-Parent (SCARED P)72
The SCARED-P is a parent reported measure of anxiety symptoms in children aged 8–18 years old. It has five subscales: separation anxiety, generalized anxiety, social anxiety, panic or somatic symptoms, and school avoidance, which can be combined to obtain a total anxiety score.
Sensory Experiences Questionnaire, Version 3.0 (SEQ)71
The SEQ is a parent-reported measure of sensory response patterns (hypersensitivity, hyposensitivity, enhanced perception, and sensory interests, repetitions, and seeking), sensory modalities (tactile, auditory, gustatory, auditory, vestibular/proprioception), and sensory contexts (non-social and social). The hypersensitivity, hyposensitivity, enhanced perception, non-social, social, and vestibular/proprioception were used as variables of interest in this study due to their hypothesized relationships with the fMRI tasks and/or metabolites.
Social Responsiveness Scale, 2nd Edition (SRS-2)68
The SRS-2 is a parent reported questionnaire for children aged 4–18 years that is used as both a screening tool and to aid in clinical diagnosis of ASD. The measure includes questions related to restrictive interests, repetitive behaviors, deficits in social interactions, and social communication.
Wechsler Abbreviated Scale of Intelligence (WASI-II)65
The WASI-II is a standardized measure of intelligence for ages 6–90. There are four subtests: block design, vocabulary, matrix reasoning, and similarities. Block design and matrix reasoning are combined to obtain a perceptual reasoning index (PRI) and vocabulary and similarities are combined for a verbal comprehension index (VCI). The full-scale intelligence quotient (FSIQ) consists of all four subtests. The WASI-II was administered by trained research staff.
Stool sample collection
After completion of their day 1 visit, participants were given a stool collection kit (specimen cup, wooden spatula, plastic bag, Fisherbrand Scientific Commode Specimen Collection System, gloves, ice packs, and an insulated transportation container). They were instructed to collect a stool sample within 72 h prior to their MRI, freeze the sample at home, and transport it in the insulated transportation container with ice packs to the lab. Once in the lab, the sample was placed in a −80 °C degree freezer for storage (first at USC, and then at UCLA where they were aliquoted under liquid nitrogen and stored in a −80 °C degree freezer). Aliquoted samples were shipped on dry ice with a stool collection log to Metabolon Inc. for further processing and analysis on their global metabolomics and bioinformatics platform (Metabolon, 617 Davis Drive, Durham, NC). See the section “Metabolite analysis (All information in this section provided by Metabolon, Inc)” for more details on sample analysis.
Brain imaging
Scanning parameters
MRI data were acquired on a 3 Tesla MAGNETOM Prisma (Siemens, Erlangen, Germany) with a 20-channel head coil. A 5-min structural T1-weighted MPRAGE was acquired for each participant (TR = 1950 ms, TE = 3.09 ms, flip angle = 10°, 256 × 256 matrix, 176 sagittal slices, 1 mm isotropic resolution). Each functional scan consisted of an echo-planar imaging (EPI; 150 whole-brain volumes) acquired with the following parameters: TR = 2 s, TE = 30 ms, flip angle = 90°, 64 × 64 matrix, in-plane resolution 2.5 × 2.5 mm, and 41 transverse slices, each 2.5 mm thick, covering the whole-brain with a multiband factor of three. Spin Echo EPI field mapping data was also acquired in AP and PA directions with identical geometry to the EPI data for EPI off-resonance distortion correction (TR = 1020 ms, TE1 = 10 ms, TE2 = 12.46 ms, flip angle = 90°, FOV = 224 × 224 × 191 mm3, voxel size = 2.5 mm isotropic).
Scanning procedure
All participants completed a practice MRI session in a mock MRI scanner prior to the fMRI tasks to become familiarized with the task and the MRI environment and to increase comfortability and minimize head motion. Functional MRI procedure, task stimuli, fMRI acquisition, and data preprocessing were completed following the protocol previously published in Kilroy et al.38. We utilized a head-motion cut-off of absolute FD > 1.5 mm. Five participants (4 ASD, 1 NT) were excluded for head motion in the watching facial expressions and hand actions task, 3 (2 ASD, 1 NT) for disgust processing, and 2 (1 ASD, 1 NT) from watching others being touched. There were no significant differences in relative head motion between the two groups for the disgust processing (t = 0.981, p = 0.33) and watching others being touched (t = −1.029, p = 0.307) tasks, but significant differences were present for the observation of facial expressions and hand actions task (t = −2.572, p = 0.015). Please see section “fMRI processing” for details regarding motion correction.
fMRI tasks
The task-based fMRI paradigms were selected based on existing literature showing significant ASD vs. NT differences during these tasks, their relevance to key ASD symptomatology (socio-emotional processing and sensory sensitivities)38,46,83,84 and/or their relevance to vagally mediated emotional processes (disgust and emotion processing)49. The fMRI tasks included: watching videos of facial expressions/body actions, physical and social disgust processing tasks, and watching videos of others being touched (Supplemental Fig. 1). Stimuli were presented using the Psychophysics Toolbox85 on MATLAB. During all tasks, participants were instructed to simply watch all videos and remain as still as possible. fMRI tasks are described in A–C below.
A. Watching videos of facial expressions and hand actions (n = 78; 38 NT [19 female, 19 male], 40 ASD [11 female, 29 male]): One 9-min run with five 15-s blocks of video-stimuli were shown. As Supplemental Fig. 1A shows, blocks consisted of one of three categories of stimuli: emotional facial expressions (e.g., happy expression), non-emotional facial expressions (e.g., tongue to lip), or bimanual hand actions (e.g., playing the xylophone). Each video was presented for 3.75 s followed by a 1.25 s black screen between each stimulus, there were 3 videos per block, and both male and female actors were included in each block. For further details on stimuli, please see Kilroy et al.38.
B. Disgust processing (n = 46; 22 NT [12 female, 10 male], 24 ASD [6 female, 18 male]): There were four categories of stimuli, disgusting foods, disgusted facial expressions, neutral foods, and neutral facial expressions (Supplemental Fig. 1B). The neutral and the disgusted facial expressions were chosen from an online repository (NimStim)86 and from previous research (EmStim)38 then edited and counterbalanced so that each participant saw the same actor displaying a neutral and disgusted facial expression. To ensure that the neutral food images were indeed items the participant truly had no preferential or disgusting feelings for, all participants were administered a questionnaire prior to participating in the study, to assess their preferences for each food stimuli. For each participant, 18 images were used from each stimulus category. One fMRI run was presented to all participants, including six blocks per stimulus category. Within each 15-s block, three different images from the same category were presented with a 250-ms fixation crosshair between each stimulus (e.g., three different disgusting food images). Thus, the fMRI task consisted of 24 blocks (5 per stimulus category), lasting for a single 10-min run.
C. Watching others being touched (n = 37; 19 NT [11 female, 8 male], 18 ASD [5 female, 13 male]): Participants watched four different videos where a person strokes the arm of another person in the MRI scanner with: 1) their hand with glove on (social touch), 2) a dry sponge (object touch), 3) their hand with glove on hovering next to the person’s arm (social touch control), and 4) a dry sponge hovering next to the person’s arm (object touch control) following a similar to the protocol used in Green et al. 2015 (Supplemental Fig. 1C)87. Each video was 15 s long followed by a 15 s rest block. During the rest blocks, participants were shown a black crosshair in the middle of a white screen. Excluding an initial junk block, five blocks of each stimulus condition were alternated with rest in a pseudo-random sequence. Stimuli conditions were counterbalanced across participants. We note that our original intention was to physically touch participants while in the scanner, but as this task was largely conducted during the initial period of the COVID-19 pandemic, we were obliged to remain at a 6-ft distance from our participants, and thus used videos of touch instead, as this has previously been shown to show strong effects88,89.
Analysis
Behavioral group differences
Independent samples t-tests and Fisher’s exact tests were conducted using SPSS Statistics (Version 29.0.2.0) to determine ASD-NT differences in demographic and behavioral variables. Significance was set at p < 0.05.
Metabolite analysis (All information in this section provided by Metabolon, Inc)
Data quality
Instrument and Process Validity: Instrument variability was determined by calculating the median relative standard deviation (RSD) for the internal standards that were added to each sample prior to injection into the mass spectrometers. Overall process variability was determined by calculating the median RSD for all endogenous metabolites (i.e., non-instrument standards) present in 100% of the Client Matrix samples, which are technical replicates of pooled client samples. Values for instrument and process variability met Metabolon’s acceptance criteria.
Sample accessioning
Following receipt, samples were inventoried and immediately stored at −80 °C. Each sample received was accessioned into the Metabolon LIMS system and was assigned by the LIMS a unique identifier that was associated with the original source identifier only. This identifier was used to track all sample handling, tasks, results, etc. The samples (and all derived aliquots) were tracked by the LIMS system. All portions of any sample were automatically assigned their own unique identifiers by the LIMS when a new task was created; the relationship of these samples was also tracked. All samples were maintained at −80 °C until processed.
Sample preparation
Samples were prepared using the automated MicroLab STAR® system from Hamilton Company. Several recovery standards were added prior to the first step in the extraction process for QC purposes. To remove protein, dissociate small molecules bound to protein or trapped in the precipitated protein matrix, and to recover chemically diverse metabolites, proteins were precipitated with methanol under vigorous shaking for 2 min (Glen Mills GenoGrinder 2000) followed by centrifugation. The resulting extract was divided into five fractions: two for analysis by two separate reverse phase (RP)/UPLC-MS/MS methods with positive ion mode electrospray ionization (ESI), one for analysis by RP/UPLC-MS/MS with negative ion mode ESI, one for analysis by HILIC/UPLC-MS/MS with negative ion mode ESI, and one sample was reserved for backup. Samples were placed briefly on a TurboVap® (Zymark) to remove the organic solvent. The sample extracts were stored overnight under nitrogen before preparation for analysis.
QA/QC
Several types of controls were analyzed in concert with the experimental samples: a pooled matrix sample generated by taking a small volume of each experimental sample (or alternatively, use of a pool of well-characterized human plasma) served as a technical replicate throughout the data set; extracted water samples served as process blanks; and a cocktail of QC standards that were carefully chosen not to interfere with the measurement of endogenous compounds were spiked into every analyzed sample, allowed instrument performance monitoring and aided chromatographic alignment. Instrument variability was determined by calculating the median relative standard deviation (RSD) for the standards that were added to each sample prior to injection into the mass spectrometers. Overall process variability was determined by calculating the median RSD for all endogenous metabolites (i.e., non-instrument standards) present in 100% of the pooled matrix samples. Experimental samples were randomized across the platform run with QC samples spaced evenly among the injections.
Ultrahigh performance liquid chromatography-tandem mass spectroscopy (UPLC-MS/MS)
All methods utilized a Waters ACQUITY ultra-performance liquid chromatography (UPLC) and a Thermo Scientific Q-Exactive high resolution/accurate mass spectrometer interfaced with a heated electrospray ionization (HESI-II) source and Orbitrap mass analyzer operated at 35,000 mass resolution. The sample extract was dried then reconstituted in solvents compatible to each of the four methods. Each reconstitution solvent contained a series of standards at fixed concentrations to ensure injection and chromatographic consistency. One aliquot was analyzed using acidic positive ion conditions, chromatographically optimized for more hydrophilic compounds. In this method, the extract was gradient eluted from a C18 column (Waters UPLC BEH C18-2.1 × 100 mm, 1.7 µm) using water and methanol, containing 0.05% perfluoropentanoic acid (PFPA) and 0.1% formic acid (FA). Another aliquot was also analyzed using acidic positive ion conditions; however, it was chromatographically optimized for more hydrophobic compounds. In this method, the extract was gradient eluted from the same aforementioned C18 column using methanol, acetonitrile, water, 0.05% PFPA and 0.01% FA and was operated at an overall higher organic content. Another aliquot was analyzed using basic negative ion optimized conditions using a separate dedicated C18 column. The basic extracts were gradient eluted from the column using methanol and water, however, with 6.5 mM Ammonium Bicarbonate at pH 8. The fourth aliquot was analyzed via negative ionization following elution from a HILIC column (Waters UPLC BEH Amide 2.1 × 150 mm, 1.7 µm) using a gradient consisting of water and acetonitrile with 10 mM Ammonium Formate, pH 10.8. The MS analysis alternated between MS and data-dependent MSn scans using dynamic exclusion. The scan range varied slighted between methods but covered 70–1000 m/z. Raw data files are archived and extracted as described below.
Bioinformatics
The informatics system consisted of four major components, the Laboratory Information Management System (LIMS), the data extraction and peak-identification software, data processing tools for QC and compound identification, and a collection of information interpretation and visualization tools for use by data analysts. The hardware and software foundations for these informatics components were the LAN backbone, and a database server running Oracle 10.2.0.1 Enterprise Edition.
LIMS
The purpose of the Metabolon LIMS system was to enable fully auditable laboratory automation through a secure, easy to use, and highly specialized system. The scope of the Metabolon LIMS system encompasses sample accessioning, sample preparation and instrumental analysis and reporting and advanced data analysis. All the subsequent software systems are grounded in the LIMS data structures. It has been modified to leverage and interface with the in-house information extraction and data visualization systems, as well as third-party instrumentation and data analysis software.
Data extraction and compound identification
Raw data was extracted, peak-identified and QC processed using Metabolon’s hardware and software. These systems are built on a web-service platform utilizing Microsoft’s.NET technologies, which run on high-performance application servers and fiber-channel storage arrays in clusters to provide active failover and load-balancing. Compounds were identified by comparison to library entries of purified standards or recurrent unknown entities. Metabolon maintains a library based on authenticated standards that contains the retention time/index (RI), mass to charge ratio (m/z), and chromatographic data (including MS/MS spectral data) on all molecules present in the library. Furthermore, biochemical identifications are based on three criteria: retention index within a narrow RI window of the proposed identification, accurate mass match to the library +/−10 ppm, and the MS/MS forward and reverse scores between the experimental data and authentic standards. The MS/MS scores are based on a comparison of the ions present in the experimental spectrum to the ions present in the library spectrum. While there may be similarities between these molecules based on one of these factors, the use of all three data points can be utilized to distinguish and differentiate biochemicals. More than 3300 commercially available purified standard compounds have been acquired and registered into LIMS for analysis on all platforms for determination of their analytical characteristics. Additional mass spectral entries have been created for structurally unnamed biochemicals, which have been identified by virtue of their recurrent nature (both chromatographic and mass spectral). These compounds have the potential to be identified by future acquisition of a matching purified standard or by classical structural analysis.
Curation
A variety of curation procedures were carried out to ensure that a high-quality data set was made available for statistical analysis and data interpretation. The QC and curation processes were designed to ensure accurate and consistent identification of true chemical entities, and to remove those representing system artifacts, mis-assignments, and background noise. Metabolon data analysts use proprietary visualization and interpretation software to confirm the consistency of peak identification among the various samples. Library matches for each compound were checked for each sample and corrected if necessary.
Metabolite Quantification and Data Normalization
Peaks were quantified using area-under-the-curve.
Preprocessing of metabolomic data
Peak area values were log transformed and KNN imputation was applied for missing data90. Next, data was Z score normalized and adjusted for use of antidepressants, vitamins, supplements, laxatives, antihistamines, stimulants, cognition enhancers, and antipsychotics. Specifically we regressed out significant medication/supplement effects identified using a backward selection approach (function “MASS::stepAIC” in R using the BIC, i.e., log(n) degrees of freedom). The adjusted features were then used in downstream analyses. A priori metabolite targets of interest included 26 named metabolites in the tryptophan pathway (see Fig. 1).
Between-group differences in metabolites
Contrast analysis was applied within the framework of the general linear model (GLM) to determine ASD-NT differences in 26 tryptophan metabolites. The model included group as a factor, and sex, age, and BMI as covariates. As children with ASD in our sample had significantly higher levels of gastrointestinal symptoms compared to the NT children (p < 0.001; Supplemental Table 1) we also ran GLMs controlling for gastrointestinal symptoms (GSRS) and diet to assess potential confounding effects of GI symptoms on the levels of tryptophan-related metabolites. Significance was considered at a false discovery rate of q < 0.05. Cohen’s d was calculated to provide an effect size for differences. R Version 4.2.1 was used for all GLM analysis.
fMRI
fMRI preprocessing
All analyses followed best practices in fMRI analysis, as detailed in our prior studies38,49. The data analytic approach used to address each of our research questions utilized FMRIB’s Software Library 6.0 (FSL)91,92,93,94,95. Standard preprocessing pipeline was performed involving: (a) structural T1 brain extraction and non-brain tissue removal; (b) smoothing with 5 mm FWHM Gaussian kernel; (c) B0 unwarping along y-axis; (d) high pass filter with 100 s cutoff; (e) realignment using MCFLIRT to obtain motion estimates; and (f) Independent component analysis (ICA). Preprocessed data was fed into the ICA AROMA algorithm96, which filtered out noise and motion components from the whole brain signal. Registration to the MNI-152 standard atlas using 12 degrees-of-freedom affine transformation and FNIRT nonlinear registration were performed92,93.
Within-group analyses
Individual participants’ statistical images were subjected to higher-level mixed-effects analyses using FSL’s FLAME Stage 1 algorithm, modeling the stimulus conditions for each participant as separate regressors. For watching facial expressions and hand actions, regressors included: emotional faces, non-emotional faces, and bimanual hand actions. For disgust processing, regressors included: disgusting foods, neutral foods, disgusted facial expressions, neutral facial expressions. Subject-specific head motion parameters were used as nuisance regressors. For observation of others being touched, regressors included social touch and object touch.
Between-group analysis
Between-group comparisons between the NT and ASD groups were performed using higher level mixed-effects analyses with FSL’s FLAME 1 algorithm. Age, Sex, and IQ were used as covariates in all group-level analyses. For watching facial expressions and hand actions, groups were contrasted on: all stimuli>rest; emotional facial expressions>rest; non-emotional facial expressions>rest; all facial expressions>rest; hand actions>rest. For disgust processing, groups were contrasted on: disgusting foods>rest, disgusted faces>rest. For observation of others being touched, groups were contrasted on: social touch>rest and object touch>rest. For the facial expressions/hand action task and the disgust tasks, the resulting group-level images for all models were thresholded at voxel Z > 3.1, with a cluster size probability correction threshold of p < 0.05. For observation of others being touched, a more lenient threshold (Z > 2.3 cluster size probability threshold of p < 0.05) was used to have more sensitivity to detect effects given the more subtle observation (rather than physical touch) task used, due to COVID-19 restrictions (see Methods: fMRI Tasks, Watching others being touched). In addition, for disgust and facial expression/hand action tasks, for hypothesized regions of interest (ROIs), a small volume correction (SVC) analysis with a significance threshold of p < 0.01 using predefined masks for disgust and observation tasks. For the facial expression/hand action task, we used structurally defined anterior insula parcellations from extant literature and the Harvard-Oxford atlas parcellations for the pACC and amygdala. For the IFGop, we used a hand-drawn anatomically derived ROI38 and previously published insula parcellations97. For the disgust task, ROIs for SVC analysis were defined utilizing the Neurosynth database (which performs automated large-scale meta-analyses of fMRI data), using the search terms: “disgust”, “emotional faces”, and “food”, and we also included insula parcellations from extant literature97. Functional ROIs were then masked with structural ROIs from the Harvard-Oxford atlas (thresholded at 30% probability) to ensure they captured non-overlapping regions.
General linear models (GLMs)
GLMs were applied within the ASD and NT groups to test brain-behavior, brain-metabolite, and metabolite-behavior relationships. The GLMs included group as a factor, and sex, age, IQ, and BMI were included as covariates. As a measure of effect size, we report the standardized beta (Std β). Std β between 0.10 and 0.29 is considered small, 0.30–0.49 medium, and greater than 0.50 large98. Brain ROIs were chosen based on group differences in fMRI tasks as well as prior studies supporting atypicalities in brain activity in the chosen ROIs38,46,49,83,84. The Benjamini-Hochberg method to correct for multiple comparisons was used; the false discovery reporting threshold set at 10% (FDR)99. We used FDR correction for the number of dependent variables in each analyses. Specifically for brain-metabolite and brain-behavior analyses, FDR correction was for the number of ROIs compared. For metabolite-behavior analyses, FDR correction was for the number of metabolites compared. To limit the number of comparisons, only metabolites that significantly correlated with brain activity were included in the metabolite-behavior analyses.
As a post hoc analysis, we ran GLMs with all primary variables and all tryptophan metabolites (see Supplemental Table 7). The GLM model included group as a factor, and sex, age, IQ, and BMI were included as covariates.
Mediation models
Exploratory mediation analyses were conducted to determine if the brain regions that differed between ASD and NT statistically mediated the relationships between metabolites and behavior in the ASD group. The variables included in the mediation models were selected based on ROIs that had both significant ROI-metabolite associations and significant ROI-behavior associations. Mediation modeling was performed using lavaan in R. We estimated the bootstrapped 95% percentile confidence intervals for the indirect effects100. Confidence intervals that do not contain zero are considered significant. Because age, sex, and BMI are collinear, we ran analyses using only age and FSIQ as covariates. In addition, in terms of regressors for brain and behavior (as opposed to metabolites), it is less common and relevant to control for BMI, especially those in normal ranges, as was our ASD sample mean (see Supplemental Table 1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Deidentified data are available via the NIMH Data Archive: The Relationship Between Brain Functioning, Behavior, and Microbiota in Autism Spectrum Disorder #4991. Metabolomic data is available through Zenodo (https://zenodo.org/records/1467599). Source data are provided with this paper.
Code availability
MRI codes are available on GitHub: https://doi.org/10.5281/zenodo.14834325 and https://doi.org/10.5281/zenodo.14834337.
References
Rojo, D. et al. Exploring the human microbiome from multiple perspectives: factors altering its composition and function. FEMS Microbiol. Rev. 41, 453–478 (2017).
Miri, S., Yeo, J., Abubaker, S. & Hammami, R. Neuromicrobiology, an emerging neurometabolic facet of the gut microbiome? Front. Microbiol. 14, 1098412–1098412 (2023).
Alharthi, A., Alhazmi, S., Alburae, N. & Bahieldin, A. The human gut microbiome as a potential factor in autism spectrum disorder. Int. J. Mol. Sci. 23, 1363 (2022).
Forsythe, P., Bienenstock, J. & Kunze, W. A. Vagal pathways for microbiome-brain-gut axis communication. Adv. Exp. Med. Biol. 817, 115–133 (2014).
Chernikova, M. A. et al. The brain-gut-microbiome system: pathways and implications for autism spectrum disorder. Nutrients 13, 4497 (2021).
Chen, Y.-C. et al. Altered gut microbiota correlates with behavioral problems but not gastrointestinal symptoms in individuals with autism. Brain, Behav., Immun. 106, 161–178 (2022).
Zhu, X. et al. Microbiota-gut-brain axis and the central nervous system. Oncotarget 8, 53829–53838 (2017).
Sharon, G. et al. Human gut microbiota from autism spectrum disorder promote behavioral symptoms in mice. Cell 177, 1600–1618.e17 (2019).
Srikantha, P. & Mohajeri, Hasan M. The possible role of the microbiota-gut-brain-axis in autism spectrum disorder. Int. J. Mol. Sci. 20, 2115 (2019).
Borre, Y. E. et al. Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol. Med. 20, 509–518 (2014).
Cryan, J. F., O’Riordan, K. J., Sandhu, K., Peterson, V. & Dinan, T. G. The gut microbiome in neurological disorders. Lancet Neurol. 19, 179–194 (2020).
Goldstein, A. M., Hofstra, R. M. W. & Burns, A. J. Building a brain in the gut: development of the enteric nervous system. Clin. Genet. 83, 307–316 (2013).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Jašarević, E. & Bale, T. L. Prenatal and postnatal contributions of the maternal microbiome on offspring programming. Front. Neuroendocrinol. 55, 100797–100797 (2019).
Zuffa, S. et al. Early-life differences in the gut microbiota composition and functionality of infants at elevated likelihood of developing autism spectrum disorder. Transl. Psychiatry 13, 257–257 (2023).
Zhang, Q. et al. Comparison of gut microbiota between adults with autism spectrum disorder and obese adults. PeerJ 9, e10946 (2021).
Pang, X. et al. Characteristics of the gut microbiota in young adults with autism spectrum disorder. J. Integr. Neurosci. 22, 141–141 (2023).
Aburto, M. R. & Cryan, J. F. Gastrointestinal and brain barriers: unlocking gates of communication across the microbiota–gut–brain axis. Nat. Rev. Gastroenterol. Hepatol. https://doi.org/10.1038/s41575-023-00890-0 (2024).
De Angelis, M., Francavilla, R., Piccolo, M., De Giacomo, A. & Gobbetti, M. Autism spectrum disorders and intestinal microbiota. Gut Microbes 6, 207–213 (2015).
Liu, P., Peng, G., Zhang, N., Wang, B. & Luo, B. Crosstalk between the gut microbiota and the brain: an update on neuroimaging findings. Front. Neurol. 10, 883–883 (2019).
Kang, D. W. et al. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome 5, 10–10 (2017).
Arnold, L. E. et al. Probiotics for gastrointestinal symptoms and quality of life in autism: a placebo-controlled pilot trial. J. Child Adolesc. Psychopharmacol. 29, 659–669 (2019).
Needham, B. D. et al. Plasma and fecal metabolite profiles in autism spectrum disorder. Biol. Psychiatry 89, 451–462 (2021).
Al-Beltagi, M. Autism medical comorbidities. World J. Clin. Pediatr. 10, 15–28 (2021).
Gorrindo, P. et al. Gastrointestinal dysfunction in autism: parental report, clinical evaluation, and associated factors. Autism Res. 5, 101–108 (2012).
Gabriele, S., Sacco, R. & Persico, A. M. Blood serotonin levels in autism spectrum disorder: a systematic review and meta-analysis. Eur. Neuropsychopharmacol. 24, 919–929 (2014).
Mawe, G. M. & Hoffman, J. M. Serotonin signalling in the gut-functions, dysfunctions and therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 10, 473–486 (2013).
Iglesias–Vázquez, L., Riba, G. V. G., Arija, V. & Canals, J. Composition of gut microbiota in children with autism spectrum disorder: a systematic review and meta-analysis. Nutrients 12, 792 (2020).
Daly, E. et al. Response inhibition and serotonin in autism: A functional MRI study using acute tryptophan depletion. Brain 137, 2600–2610 (2014).
Daly, E. M. et al. Serotonin and the neural processing of facial emotions in adults with autism: an fmri study using acute tryptophan depletion. Arch. Gen. Psychiatry 69, 1003–1013 (2012).
MacFabe, D. Autism: metabolism, mitochondria, and the microbiome. Glob. Adv. Health Med. 2, 52–66 (2013).
Yano, JessicaM. et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 163, 258–258 (2015).
Muller, C. L., Anacker, A. M. J. & Veenstra-VanderWeele, J. The serotonin system in autism spectrum disorder: from biomarker to animal models. Neuroscience 321, 24–41 (2016).
Schain, R. J. & Freedman, D. X. Studies on 5-hydroxyindole metabolism in autistic and other mentally retarded children. J. Pediatr. 58, 315–320 (1961).
Bohár, Z., Toldi, J., Fülöp, F. & Vécsei, L. Changing the face of kynurenines and neurotoxicity: therapeutic considerations. Int. J. Mol. Sci. 16, 9772–9793 (2015).
Pappolla, M. A. et al. Indoles as essential mediators in the gut-brain axis. Their role in Alzheimer’s disease. Neurobiol. Dis. 156, 105403–105403 (2021).
Morais, L. H., Schreiber, H. L. & Mazmanian, S. K. The gut microbiota–brain axis in behaviour and brain disorders. Nat. Rev. Microbiol. 19, 241–255 (2021).
Kilroy, E. et al. Unique deficit in embodied simulation in autism: an fMRI study comparing autism and developmental coordination disorder. Hum. Brain Mapp. 42, 1532–1546 (2021).
Needham, B. D. et al. A gut-derived metabolite alters brain activity and anxiety behaviour in mice. Nature 602, 647–653 (2022).
Kennedy, P. J., Cryan, J. F., Dinan, T. G. & Clarke, G. Kynurenine pathway metabolism and the microbiota-gut-brain axis. Neuropharmacology 112, 399–412 (2017).
Bryn, V., Verkerk, R., Skjeldal, OlaH., Saugstad, OlaD. & Ormstad, H. Kynurenine pathway in autism spectrum disorders in children. Neuropsychobiology 76, 82–88 (2018).
Lapin, I. Kynurenines as probable participants of depression. Pharmacopsychiatry 6, 273–279 (1973).
Schwarcz, R., Bruno, J. P., Muchowski, P. J. & Wu, H. Q. Kynurenines in the mammalian brain: when physiology meets pathology. Nat. Rev. Neurosci. 13, 465–477 (2012).
Lee, B. H., Smith, T. & Paciorkowski, A. R. Autism spectrum disorder and epilepsy: disorders with a shared biology. Epilepsy Behav. 47, 191–201 (2015).
Laghi, L. et al. Are fecal metabolome and microbiota profiles correlated with autism severity? A cross-sectional study on asd preschoolers. Metabolites 11, 654 (2021).
Uddin, L. Q. & Menon, V. The anterior insula in autism: under-connected and under-examined. Neurosci. Biobehav. Rev. 33, 1198–1203 (2009).
Pfeifer, J. H. et al. Neural and behavioral responses during self-evaluative processes differ in youth with and without autism. J. Autism Dev. Disord. 43, 272–285 (2013).
Loureiro, F., Ringold, S. M. & Aziz-Zadeh, L. Interoception in autism: a narrative review of behavioral and neurobiological data. Psychol. Res. Behav. Manag. 17, 1841–1853 (2024).
Jayashankar, A. et al. Disgust processing differences and their neural correlates in autistic youth. OSF Preprints. https://doi.org/10.31234/osf.io/dt678 (2025).
Dan, Z. et al. Altered gut microbial profile is associated with abnormal metabolism activity of Autism Spectrum Disorder. Gut Microbes 11, 1246–1267 (2020).
Marler, S. et al. Brief Report: Whole blood serotonin levels and gastrointestinal symptoms in autism spectrum disorder. J. Autism Dev. Disord. 46, 1124–1130 (2016).
Wong, A. C. et al. Serotonin reduction in post-acute sequelae of viral infection. Cell 186, 4851–4867.e20 (2023).
Góralczyk-Bińkowska, A., Szmajda-Krygier, D. & Kozłowska, E. The microbiota–gut–brain axis in psychiatric disorders. Int. J. Mol. Sci. 23, 11245 (2022).
Jayashankar, A. & Aziz-Zadeh, L. Disgust processing and potential relationships with behaviors in autism. Curr. Psychiatry Rep. 25, 465–478 (2023).
Tennoune, N., Andriamihaja, M. & Blachier, F. Production of indole and indole‐related compounds by the intestinal microbiota and consequences for the host: the good, the bad, and the ugly. Microorganisms 10, 930 (2022).
Craig, A. D. How do you feel? Interoception: the sense of the physiological condition of the body. Nat. Rev. Neurosci. 3, 655–666 (2002).
Kelly, R. S. et al. Metabolomics and communication skills development in children; evidence from the ages and stages questionnaire. Metabolites 9, 42 (2019).
De Angelis, M. et al. Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS ONE 8, e76993 (2013).
Sun, Z. et al. Revealing the importance of prenatal gut microbiome in offspring neurodevelopment in humans. EBioMedicine 90, 104491 (2023).
Zierer, J. et al. The fecal metabolome as a functional readout of the gut microbiome. Nat. Genet. 50, 790–795 (2018).
Chen, Y. et al. Genomic atlas of the plasma metabolome prioritizes metabolites implicated in human diseases. Nat. Genet. 55, 44–53 (2023).
Yap, C. X. et al. Autism-related dietary preferences mediate autism-gut microbiome associations. Cell 184, 5916–5931.e17 (2021).
West, K. A., Schmid, R., Gauglitz, J. M., Wang, M. & Dorrestein, P. C. foodMASST a mass spectrometry search tool for foods and beverages. NPJ Sci. Food 6, 22–22 (2022).
Shim, J.-S., Oh, K. & Kim, H. C. Dietary assessment methods in epidemiologic studies. Epidemiol. Health 36, e2014009 (2014).
Wechsler, D. Data from: Wechsler Abbreviated Scale of Intelligence - Second Edition (2011).
Crovitz, H. F. & Zener, K. A group-test for assessing hand-and eye-dominance. Am. J. Psychol. 75, 271–276 (1962).
Conners, C. K. Conners third edition (Conners 3) (Western Psychological Services, 2008).
Constantino, J. N. & Gruber C. P. Social Responsiveness Scale: SRS-2 (Western Psychological Services Torrance, 2012).
Lord, C., DiLavore, P., Risi, S., Gotham, K. & Bishop, S. Autism Diagnostic Observation Schedule (ADOS-2) (2012).
Lord, C., Rutter, M. & Le Couteur, A. Autism Diagnostic Interview-Revised: a revised version of a diagnostic interview for caregivers of individuals with possible pervasive developmental disorders. J. Autism Dev. Disord. 24, 659–685 (1994).
Baranek, G. T. Sensory Experiences Questionnaire (Version 3.0, Unpublished Manuscript) (The University of North Carolina at Chapel Hill, 2009).
Birmaher, B. et al. The Screen for Child Anxiety Related Emotional Disorders (SCARED): scale construction and psychometric characteristics. J. Am. Acad. Child Adolesc. Psychiatry 36, 545–553 (1997).
Fergus, T. A. & Valentiner, D. P. The Disgust Propensity and Sensitivity Scale–Revised: an examination of a reduced-item version. J. anxiety Disord. 23, 703–710 (2009).
Rieffe, C., Oosterveld, P. & Terwogt, M. M. An alexithymia questionnaire for children: factorial and concurrent validation results. Personal. Individ. Differences 40, 123–133 (2006).
Porges, S. Body perception questionnaire. Laboratory of Developmental Assessment, University of Maryland (1993).
Svedlund, J., Sjödin, I. & Dotevall, G. GSRS–a clinical rating scale for gastrointestinal symptoms in patients with irritable bowel syndrome and peptic ulcer disease. Dig. Dis. Sci. 33, 129–134 (1988).
Lewis, S. J. & Heaton, K. W. Stool form scale as a useful guide to intestinal transit time. Scand. J. Gastroenterol. 32, 920–924 (1997).
LeBourgeois, M. K., Giannotti, F., Cortesi, F., Wolfson, A. R. & Harsh, J. The relationship between reported sleep quality and sleep hygiene in Italian and American adolescents. Pediatrics 115, 257–265 (2005).
Malow, B. A. et al. A sleep habits questionnaire for children with autism spectrum disorders. J. Child Neurol. 24, 19–24 (2009).
Loas, G., Braun, S., Delhaye, M. & Linkowski, P. The measurement of alexithymia in children and adolescents: psychometric properties of the Alexithymia Questionnaire for Children and the twenty-item Toronto Alexithymia Scale in different non-clinical and clinical samples of children and adolescents. PLOS ONE 12, e0177982 (2017).
Cabrera, A. et al. Assessing body awareness and autonomic reactivity: factor structure and psychometric properties of the Body Perception Questionnaire-Short Form (BPQ-SF). Int J. Methods Psychiatr. Res 27, e1596 (2018).
Chen, M.-H. et al. Autistic spectrum disorder, attention deficit hyperactivity disorder, and psychiatric comorbidities: a nationwide study. Res. autism Spectr. Disord. 10, 1–6 (2015).
Dapretto, M. et al. Understanding emotions in others: mirror neuron dysfunction in children with autism spectrum disorders. Nat. Neurosci. 9, 28–30 (2006).
Green, S. A. et al. Overreactive brain responses to sensory stimuli in youth with autism spectrum disorders. J. Am. Acad. Child Adolesc. Psychiatry 52, 1158–1172 (2013).
Brainard, D. H. The psychophysics toolbox. Spat. Vis. 10, 433–436 (1997).
Tottenham, N. et al. The NimStim set of facial expressions: judgments from untrained research participants. Psychiatry Res. 168, 242–249 (2009).
Green, S. A. et al. Neurobiology of sensory overresponsivity in youth with autism spectrum disorders. JAMA Psychiatry 72, 778–786 (2015).
Keysers, C. et al. A touching sight: SII/PV activation during the observation and experience of touch. Neuron 42, 335–346 (2004).
Masson, H. L. & Isik, L. Rapid processing of observed touch through social perceptual brain regions: an EEG-fMRI fusion study. J. Neurosci. 43, 7700–7711 (2023).
Do, K. T. et al. Characterization of missing values in untargeted MS-based metabolomics data and evaluation of missing data handling strategies. Metabolomics 14, 128–18 (2018).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. NeuroImage 62, 782–790 (2012).
Bannister, JenkinsonM., Brady, P. & Smith, M. S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. NeuroImage 17, 825–841 (2002).
Jenkinson, M. & Smith, S. A global optimisation method for robust affine registration of brain images. Med. Image Anal. 5, 143–156 (2001).
Smith, J. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. NeuroImage 23, S208–S219 (2004).
Woolrich, M. W., Ripley, B. D., Brady, M. & Smith, S. M. Temporal autocorrelation in univariate linear modeling of FMRI data. NeuroImage 14, 1370–1386 (2001).
Pruim, R. H. R. et al. ICA-AROMA: a robust ICA-based strategy for removing motion artifacts from fMRI data. NeuroImage 112, 267–277 (2015).
Deen, B., Pitskel, N. B. & Pelphrey, K. A. Three systems of insular functional connectivity identified with cluster analysis. Cereb. Cortex 21, 1498–1506 (2011).
Nieminen, P. Application of standardized regression coefficient in meta-analysis. BioMedInformatics 2, 434–458 (2022).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B, Methodol. 57, 289–300 (1995).
MacKinnon, D. P., Lockwood, C. M. & Williams, J. Confidence limits for the indirect effect: distribution of the product and resampling methods. Multivar. Behav. Res. 39, 99–128 (2004).
Acknowledgements
We thank all our participants, research assistants, and the Integrative Biostatistics and Bioinformatics Core at the Goodman Luskin Microbiome Center for their contributions to this study. We also thank Ruty Mehrian-Shai, Antonio Damasio, Sarkis Mazmanian, and Jonas Kaplan for helpful discussions on this study. This work was funded by Eunice Kennedy Shriver National Institute of Child Health and Human Development (R01HD079432; PI: L.A.Z., M.D.) and the Department of Defense through the Idea Development Award (AR170062; PI: L.A.Z., E.A.M.). Opinions, interpretations, conclusions, and recommendations are those of the authors and are not necessarily endorsed by the NIH or the Department of Defense. Additional support was provided by the Nedra Gillette Endowed Research Fellowship E.K.
Author information
Authors and Affiliations
Contributions
Idea conceptualization: L.A.Z., E.A.M., J.S.L., and M.D.; Design: L.A.Z., E.A.M., J.S.L., E.K., C.B., A.J., and M.D.; Data acquisition: E.K., C.B., A.J., and S.M.R.; Specimen handling/preprocessing: J.P.J.; Data analysis: S.M.R., J.S.L., S.T., A.J., C.B., L.A.Z., E.K., R.R.B., and S.M.J.; First draft of manuscript: L.A.Z., S.M.R., J.S.L., and E.A.M.; Manuscript editing/revision: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aziz-Zadeh, L., Ringold, S.M., Jayashankar, A. et al. Relationships between brain activity, tryptophan-related gut metabolites, and autism symptomatology. Nat Commun 16, 3465 (2025). https://doi.org/10.1038/s41467-025-58459-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58459-1
This article is cited by
-
Gut Microbiota and Mitochondrial Dysfunction in Autism: Clinical Correlations and Future Directions
Molecular Neurobiology (2026)
-
Gut microbiota and urine metabolomics signature in autism spectrum disorder children from Southern China
BMC Pediatrics (2025)