Abstract
Genetically encoded calcium ion (Ca2+) indicators (GECIs) are widely-used molecular tools for functional imaging of Ca2+ dynamics and neuronal activities with single-cell resolution. Here we report the design and development of two far-red fluorescent GECIs, FR-GECO1a and FR-GECO1c, based on the monomeric far-red fluorescent proteins mKelly1 and mKelly2. FR-GECOs have excitation and emission maxima at ~596 nm and ~644 nm, respectively, display large responses to Ca2+ in vitro (ΔF/F0 = 6 for FR-GECO1a, 18 for FR-GECO1c), are bright under both one-photon and two-photon illumination, and have high affinities (apparent Kd = 29 nM for FR-GECO1a, 83 nM for FR-GECO1c) for Ca2+. FR-GECOs offer sensitive and fast detection of single action potentials in neurons, and enable in vivo all-optical manipulation and measurement of cellular activities in combination with optogenetic actuators.
Similar content being viewed by others
Introduction
A property of most mammalian tissues is that they are most transparent to wavelengths of light between ~600 nm and ~1300 nm, a range often referred to as the optical window1,2. This wavelength range falls between the absorbance profile of hemoglobin, which predominates at wavelengths below ~600 nm, and the absorbance profile of water, which predominates at wavelengths longer than ~1300 nm. Due to the greater tissue transparency in this wavelength range, fluorescent probes that absorb and emit efficiently within the optical window are desirable for in vivo imaging. In addition, fluorescent probes with longer excitation wavelengths are associated with lower phototoxicity and autofluorescence, reduced crosstalk with green fluorescent indicators, and better spectral compatibility with blue or cyan light-activated optogenetics tools.
In efforts to realize the advantages of fluorophores that excite and emit within the optical window, molecular tool engineers strive to shift the excitation and emission wavelengths of genetically encodable fluorescent proteins (FPs), such as standard red FPs (RFPs) with excitation maxima (λex) at 550 to 580 nm and emission (λem) at 580 to 620 nm, into the far-red region of the spectrum. This longstanding effort has yielded a plethora of far-red FPs (Fig. 1A) with λex > 580 nm and λem > 620 nm3,4,5,6,7,8,9,10,11,12. Efforts to engineer biliverdin (BV)-binding FPs, that fluoresce at even further red-shifted wavelengths, have resulted in near-infrared (NIR) FPs with λex > 640 nm and λem > 670 nm13,14,15. The key difference between these two classes of genetically encodable fluorophores is that the red and far-red FPs autocatalytically form their own chromophore and are homologs of the green FP (GFP), but genetically encoded NIR FPs use the biliverdin cofactor as their chromophore.
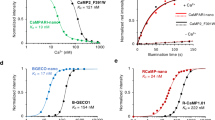
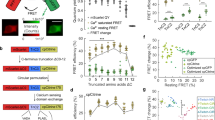
A Selected far-red FPs and FR-GECO genealogy. Far-red FPs (λem > 620 nm) are highlighted in red. B Schematic illustration of FR-GECO1 design and engineering. Linker residues are in grey; asterisk indicates position 35. Magenta and blue-colored amino acids (and correspondingly colored circles in the schematic) represent the positions of ‘gatepost’ residues28 that define the gatepost residues of cpmKelly2. Abbreviations: Calmodulin (CaM); CaM-dependent kinase kinase CaMKK-α/β peptide (ckkap). Absorbance, excitation, and emission spectra of FR-GECO1a (C) and FR-GECO1c (D) with dashed lines representing the fluorescence spectra of the Ca2+-free state and solid lines representing the spectra of the Ca2+-bound state. Two-photon spectra and dynamic range of FR-GECO1a (E) and FR-GECO1c (F). Kd titration (n = 3 technical replicates) with FR-GECO1a (G) and FR-GECO1c (H). pH sensitivity (n = 3 technical replicates) of FR-GECO1a (I) and FR-GECO1c (J). Error bars represent S.E.M. Source data are provided in a Source Data file.
Genetically encoded fluorophores can be engineered into genetically encoded indicators (also known as biosensors). One of the most important examples of these indicators is the family of genetically encoded Ca2+ indicators (GECIs), which can be used for the real-time detection and imaging of cell signaling and neuronal activities. The latest generation of red GECIs for neuronal activity detection includes jRCaMP1a,b, jRGECO1a, K-GECO1, and XCaMP-R, all with single-photon excitation and emission peaks outside of the optical window16,17,18. Among currently available RFP-based GECIs, the most red-shifted variant that uses an FP with an autocatalytic chromophore is CAR-GECO1 (λex ~ 560 nm, λem ~ 609 nm)19. The recently reported near-infrared (NIR) GECI series, NIR-GECO1,2, is based on the BV-binding FP mIFP20,21. NIR-GECO variants have excitation and emission peaks at 678 and 704 nm, respectively, with a 10-fold inverted response in fluorescence intensity upon Ca2+ addition. Although the spectra of NIR-GECO1,2 lie well within the optical window, the molecular brightness of the NIR-GECO1,2 variants is relatively dim compared to RFP-based GECIs20,21. A GECI based on Förster resonance energy transfer (FRET) from a blue-shifted BV-binding FP to a red-shifted BV-binding FP, designated iGECI, provides higher brightness but smaller Ca2+-dependent changes in fluorescence22,23.
In an effort to achieve the ideal combination of red-shifted excitation and emission maxima and fluorescent brightness, we now report the development of a series of intensiometric far-red fluorescent GECI variants. Specifically, we report FR-GECO1a (λex ~ 596 nm, λem ~ 642 nm) and FR-GECO1c (λex ~ 596 nm, λem ~ 646 nm), which are based on the recently engineered far-red FPs mKelly1 (λex ~ 596 nm, λem ~ 656 nm) and mKelly2 (λex ~ 598 nm, λem ~ 649 nm)12. These far-red GECIs open avenues for multicolor Ca2+ imaging in combination with other optogenetic indicators and actuators, as well as deeper tissue in vivo Ca2+ imaging.
Results
Protein engineering
Initial efforts to engineer a far-red Ca2+ indicator followed two parallel strategies. The first strategy was to graft key mutations for far-red fluorescence onto existing red GECIs. We used R-GECO124, CH-GECO125 and K-GECO117 as templates and introduced key mutations from E2-Crimson (λex ~ 611 nm, λem ~ 646 nm, tetrameric)5, RDSmCherry (λex ~ 600 nm, λem ~ 630 nm)10, and mNeptune (λex ~ 600 nm, λem ~ 650 nm)4, respectively (Fig. 1A, Supplementary Table 1). Unfortunately, the resulting prototypes showed complete loss of fluorescence or no substantial spectral red-shifts. The second strategy was de novo engineering of a GECI starting from far-red FP scaffolds. However, our initial attempts to engineer GECI based on mNeptune4 and mCardinal6 did not yield fluorescent prototypes. As both mNeptune and mCardinal retained weak dimerization tendencies4,6,12,26, we suspected that the failure of these prototypes to fluoresce could be due to overlap between the circular permutation site and/or insertion site of calmodulin and its binding peptide with the oligomerization interface, possibly disrupting dimerization that was crucial to the proper folding and/or chromophore maturation of these proteins.
mKelly1 and mKelly2 are far-red FP variants of mCardinal that were engineered to have increased monomericity by a combination of a C-terminal tail deletion, directed evolution, and consensus design12 (Fig. 1A). We reasoned that the enhanced monomericity of mKelly1 and mKelly2 might facilitate circular permutation and further engineering to create GECIs. K-GECO1 was chosen as the starting scaffold because the use of the ckkap peptide, instead of RS20 as the calmodulin binding peptide, contributes to this indicator having high sensitivity, high Ca2+ affinity, good linearity between fluorescence and Ca2+ concentration, and fast response kinetics17. More importantly, it is currently the only GECI engineered using an FP variant derived from eqFP578, from which mKelly2 was also derived (Fig. 1A). We thus replaced the circularly permuted (cp) FusionRed RFP portion of K-GECO1 with cpmKelly2 (Supplementary Fig. 1), the brighter of the two mKelly variants. Thr143 and Trp140 (numbering according to the structure for FusionRed; PDB ID: 6U1A)27 are the gatepost residues of mKelly2, which flank the bulge that has been identified as the privileged site for inserting sensing domains by structural and mechanistic analysis of engineered single FP-based biosensors28. As in K-GECO1, the ckkap peptide was attached to the N-terminal gatepost residue (Thr143 in FusionRed; Thr32Asn in K-GECO1) by a Lys-Tyr linker and CaM was linked to the C-terminal gatepost residue (Trp140 in FusionRed; Trp262 in K-GECO1) by a Glu-Pro-Thr-Asn linker (Supplementary Fig. 1). The initial construct resulted in a dimly fluorescent variant we named FR-GECO0.1 (Fig. 1B). Optimization of linker1 (the amino acids linking the ckkap peptide to cpmKelly2) led to the identification of an improved, but still dim, variant, FR-GECO0.2, with λex = 586 nm, λem = 632 nm and ~7-fold increase of fluorescence upon Ca2+ addition. We then subjected FR-GECO0.2 to eight rounds of iterative directed evolution to improve its brightness and Ca2+ response while ensuring that all the variants chosen to be used as gene templates for the next round maintained a far-red emission peak (Fig. 1B). Ultimately, our efforts led to a bright and sensitive far-red GECI, FR-GECO1a, which incorporated 19 mutations relative to the initial FR-GECO0.1 (Supplementary Table 2). FR-GECO1a has λex = 596 nm, λem = 642 nm, and exhibits a 6-fold increase in fluorescence intensity in the presence of Ca2+ (Fig. 1C).
With the mKelly2-derived FR-GECO1a in hand, we attempted to incorporate mKelly1-specific mutations in an effort to achieve a slightly more red-shifted emission peak and higher photostability12. Notably, mKelly1 was engineered from mCardinal in parallel with mKelly2, and it differed from mKelly2 by six mutations. To test whether these mutations could further improve FR-GECO1a, we screened a combinatorial library of FR-GECO1a containing all possible combinations of the six mKelly1-specific mutations. From this library, we isolated a variant with λex = 596 nm and λem = 646 nm. It also exhibited an 18-fold increase in fluorescence intensity in the Ca2+-bound state, which is an over 2-fold improvement relative to that of FR-GECO1a. This variant, which we named FR-GECO1c (c for contrast) incorporated a single mutation, Tyr35Thr (numbering according to FR-GECO1a sequence, Fig. 1B and Supplementary Fig. 1) which is positioned in close proximity to the first gatepost residue, Leu32 (Fig. 1B)28.
In vitro characterization
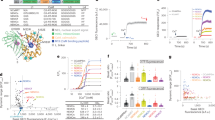
To characterize the photophysical properties of FR-GECO1a and FR-GECO1c, we purified the bacterially-expressed protein. Spectral analysis of the purified proteins revealed that the 596 nm excitation peak of both FR-GECO1a and FR-GECO1c is slightly blue-shifted by 2 nm relative to mKelly2 (598 nm) and that the emission peaks of the FR-GECO1a (642 nm) and FR-GECO1c (644 nm) are also slightly blue-shifted relative to mKelly2 (649 nm) (Fig. 1C, D and Table 1). The one-photon (1P) molecular brightness of FR-GECO1a (9.2) and FR-GECO1c (9.4) in their Ca2+-bound states are approximately 20% brighter than their template FP, mKelly2 (7.7) (Table 1). These results are attributed to a near doubling of the quantum yield (QY or Φ), which offset the effect of the indicators’ smaller effective extinction coefficients (EC or ε) compared to mKelly2. In the Ca2+-free and Ca2+-bound states, FR-GECO1a has ECs of 5710 and 27,000 M−1 cm−1, respectively, and FR-GECO1c has ECs of 4850 and 26,500 M−1 cm−1. The twofold increase in the Ca2+-dependent ΔF/F0 of FR-GECO1c relative to FR-GECO1a (attributable to the Tyr35Thr mutation), was found to result primarily from the two-fold smaller QY in the Ca2+-free state of FR-GECO1c. Under two-photon (2P) conditions, both FR-GECO1a and FR-GECO1c have peak 2P excitation at 1112 nm (Fig. 1E, F), with FR-GECO1a having a ~25% higher 2P molecular brightness (9.2 GM) than FR-GECO1c (7.4 GM) in the Ca2+-bound state (Table 1). Both FR-GECO1a and FR-GECO1c, in Ca2+-saturated states in vitro, are among the brightest red GECIs upon 2P excitation29 (Supplementary Table 3). The optimal wavelength range for 2P excitation, with close to maximum fluorescence change, was between 1050 and 1200 nm (Fig. 1E, F). The overall ΔF/F0 of FR-GECO1a and FR-GECO1c under 1P and 2P conditions are comparable. Taken together, FR-GECO1a and FR-GECO1c exhibit bright far-red fluorescence under both 1P and 2P illumination.
Consistent with other ckkap-based GECIs17,18,30, static Ca2+ titrations revealed that both FR-GECO1a and FR-GECO1c have relatively high affinities for Ca2+ and Hill coefficients (nH) greater than unity (Fig. 1G, H). FR-GECO1a (Kd = 32 nM; nH = 1.49) has a substantially lower Kd and nH than FR-GECO1c (Kd = 87 nM; nH = 1.98). These relatively high Ca2+ binding affinities suggested FR-GECO1a and FR-GECO1c would be well-suited for neuronal activity detection. Kinetic measurements revealed fast fluorescence responses from Ca2+ association and dissociation for both FR-GECO1a and FR-GECO1c (Table 1, Supplementary Fig. 2). Specifically, FR-GECO1a (tauoff = 269 ms) and FR-GECO1c (tauoff = 156 ms) are among the fastest decaying GECIs with high Ca2+ affinity31. The pH dependence (Fig. 1I, J) indicated that both FR-GECO1 variants undergo a shift in pKa upon binding Ca2+ with a pKa of ~9 in the Ca2+-free state and ~8 in the Ca2+-bound state. Both variants exhibited a small peak at pH = 5 in the Ca2+-free state and 5.5 in the bound state, suggesting multiple ionizable groups, as previously observed for the pH profile of the CH-GECO series of red GECIs32. In cultured HeLa cells, both FR-GECO1 variants gave robust responses to histamine-induced Ca2+ oscillations, with an average maximum ΔF/F0 value of 4.1 ± 0.3 for FR-GECO1c and 1.1 ± 0.06 for FR-GECO1a. In response to complete depletion and saturation of Ca2+ (with EGTA/ionomycin and Ca2+/ionomycin, respectively), FR-GECO1c gave an average ΔF/F0 of 10.4 ± 1.0 and FR-GECO1a gave an average ΔF/F0 of 2.8 ± 0.2 (Supplementary Fig. 3). These changes are about 50% of those observed with the purified proteins (Table 1).
To test whether FR-GECOs can be photoactivated by blue light (as observed for the R-GECO family of red GECIs and, to a lesser extent, K-GECO1)16,17,19, purified proteins were imaged using 561 nm (3.70 W/cm2) excitation laser with 488 nm (0.71 W/cm2) blue light laser pulses (50 ms). FR-GECO1a and FR-GECO1c were found to exhibit moderate increases in far-red fluorescence emission with blue light illumination, in both the Ca2+-bound and Ca2+-free states (Supplementary Fig. 4). However, unlike R-GECO1 (t1/2 = 0.56 s for Ca2+-free state)33, the photoactivation rise and decay of FR-GECOs happens within 10 s of milliseconds.
Performance in cultured neurons
The in vitro characterization of FR-GECO1a and FR-GECO1c revealed that, in addition to their red-shifted fluorescence spectra which enable efficient excitation at wavelengths ~596 nm (where jRGECO1a and jRCaMP1a are poorly excitable), these two GECIs had properties, such as large fluorescence changes and high Ca2+ affinities, comparable to state-of-the-art GECIs. Of the two indicators, we expected that FR-GECO1a may be more suitable for labeling fine subcellular neuronal processes, such as axons and dendrites, due to its brighter Ca2+-free state as shown by the characterization in vitro. On the other hand, we expected that FR-GECO1c may provide better performance for the detection of single spikes in neurons in terms of fluorescence change because of its larger Ca2+-dependent ΔF/F0 in vitro relative to FR-GECO1a. To evaluate the performance of FR-GECO1a and FR-GECO1c for imaging neuronal activity and compare them against jRGECO1a and jRCaMP1a, two state-of-the-art red GECIs, we expressed each GECI in dissociated rat hippocampal and cortical neurons. Similar to the red GECIs, we observed that the fluorescence in neurons expressing either FR-GECO1 variant was evenly distributed throughout the cytosol (Fig. 2A). Of the two indicators, FR-GECO1a did, indeed, have brighter basal fluorescence, providing easier identification of transfected cells as well as facilitating visualization of the fine morphology of neural cells (Fig. 2A, H).
A Representative images of neurons expressing jRGECO1a (repeated in 96 wells over 8 plates), jRCaMP1a (repeated in 77 wells over 8 plates), FR-GECO1a (repeated in 58 wells over 5 plates), FR-GECO1c (repeated in 47 wells over 5 plates), at baseline, and stimulation with 3 action potentials (AP), 10 action potentials, and 160 action potentials. All repeats gave similar results. Scale bar is 200 μm in all images. B, C Fluorescence response as a function of time following 3 action potentials (B) and 10 action potentials (C). Fluorescence responses (D), half rise time (E), half decay time (F), SNR (G), and basal fluorescence (H), of jRGECO1a (black; n = 1482 cells), jRCaMP1a (green; n = 1043 cells), FR-GECO1a (red; n = 601 cells), and FR-GECO1c (blue; n = 171 cells). Colors are consistent throughout the figure. All indicators were excited with 565 nm LED and imaged with mCherry filter cubes (excitation 560/40 nm, dichroic mirror 585LP and emission 630/75 nm). For D–G, error bars represent S.E.M. For H, the box indicates the 25th to 75th percentile range, the horizontal line indicates the median, and the whiskers extend from the box to the farthest data point lying within 1.5× the inter-quartile range (IQR). Data points beyond the boundary of the higher whisker were removed.
To evaluate indicator functions, we acquired fluorescence images while delivering electric field stimuli to evoke action potentials (APs) (in trains of 3, 10, 160 APs). FR-GECO1a and FR-GECO1c showed ΔF/F0 of 27% and 28%, respectively, upon 3 APs evoked by the field electrode, similar to jRGECO1a and jRCaMP1a, which showed ΔF/F0 of 34% and 33%, respectively (Fig. 2B–D). When evoked with 160 stimuli, neurons expressing FR-GECO1a exhibited a ΔF/F0 of 82% while FR-GECO1c had a ΔF/F0 of 161% compared to 113% and 107% for jRGECO1a and jRCaMP1a, respectively (Fig. 2D). FR-GECO1a showed a slightly slower rise time than the other three sensors for 3 and 10 APs (half rise time, FR-GECO1a: 0.29 s for 3 AP and 0.32 s for 10 AP), but both FR-GECO1a and FR-GECO1c (half rise time, FR-GECO1a: 0.61 s for 160 AP and FR-GECO1c: 0.59 s) had ~1-fold larger half rise times than jRGECO1a (half rise time: 0.35 s) and jRCaMP1a (half rise time: 0.39 s) with 160 APs (Fig. 2E). Both indicators exhibited relatively fast decay (half decay time, FR-GECO1a: 0.78 s and 0.70 s; FR-GECO1c: 0.80 s and 0.41 s, for 3 and 10 APs, respectively) times comparable to, or faster than, jRGECO1a and jRCaMP1a (half decay time, jRGECO1a: 0.61 s and 0.51 s; jRCaMP1a: 0.52 s and 1.0 s, for 3 and 10 APs, respectively) (Fig. 2F). Of the sensors tested, jRCaMP1a was most notable for its slower decay time (Fig. 2B, C, F). Relative to jRGECO1a and jRCaMP1a, FR-GECO1c exhibited a similar signal-to-noise ratio (SNR) and FR-GECO1a exhibited modestly lower SNR (Fig. 2G). Both FR-GECO1 variants provided robust responses to spontaneous activity in cortical neurons (Supplementary Fig. 5). Overall, these results suggest that both FR-GECO1a and FR-GECO1c enable sensitive detection of neuronal activities. FR-GECO1c, in particular, offers good response times, and large fluorescence changes, comparable to other state-of-art red GECIs16,17,18.
Ca2+ optopatch for all-optical cardiac electrophysiology in vivo in zebrafish embryonic heart
To investigate how FR-GECO1 performs in zebrafish, we compared its performance to state-of-the-art near-infrared GECIs based on BV-dependent FPs. In the spontaneously beating linear heart tubes (LHTs) of 1 day-old embryos, FR-GECO1 showed significantly larger Ca2+-dependent changes in fluorescence (ΔF/F0 = 11% for FR-GECO1a and 30% for FR-GECO1c) than NIR-GECO2 (ΔF/F0 = 3%) and NIR-GECO2G (ΔF/F0 = 3%) (Fig. 3A, B). In our hands, iGECI expressed in the zebrafish heart (Fig. 3C) did not show detectable fluorescence transients in the range of frequencies expected for cardiac calcium dynamics (Fig. 3A, D; Methods). Similar to what we observed during the in vitro characterization, FR-GECO1a had a stronger mean intensity (intensity = 170 arbitrary units (AU)) than FR-GECO1c (intensity = 50 AU) and comparable fluorescence intensities with the other near-infrared GECIs tested (intensity = 200 AU for NIR-GECO2, 120 AU for NIR-GECO2G and 250 AU for iGECI; Fig. 3C). Regardless, both FR-GECO1 variants had significantly larger SNR (FR-GECO1a: 140; FR-GECO1c: 230) than the near-infrared GECIs (SNR, NIR-GECO2: 13; NIR-GECO2G: 15; iGECI: 1.8; Fig. 3D). Exogenous addition of BV did not lead to significant improvements in performance for FR-GECO and the near-infrared GECIs (Supplementary Fig. 6).
Fluorescence responses (A), average intensities (AU arbitrary units) (C), and SNRs (D) of FR-GECO1a, FR-GECO1c, NIR-GECO2, NIR-GECO2G, and iGECI. B Average of 5 spontaneous heartbeats from representative traces for each sensor (left) and representative traces (right) of FR-GECO1a and FR-GECO1c. Colors are consistent with other panels and iGECI was omitted for A and B since no Ca2+-dependent change in fluorescence was observed. A–D n = 23 embryos for FR-GECO1a, 23 embryos for FR-GECO1c, 25 embryos for NIR-GECO2, 25 embryos for NIR-GECO2G, and 12 embryos for iGECI. Error bars are standard deviation. Statistical comparisons were performed using two-sided Welch’s t-test with Benjamini–Hochberg multiple test correction (Shapiro–Wilk test for normality p > 0.5 for all groups); n.s. not significant, *** p < 0.001, **** p < 0.0001. Exact p values are: A FR-GECO1a vs. FR-GECO1c p = 8.526e−4, FR-GECO1a vs. NIR-GECO2 p = 1.321e−5, FR-GECO1a vs. NIR-GECO2G p = 8.468e−6, FR-GECO1c vs. NIR-GECO2 p = 1.7e−5, FR-GECO1c vs. NIR-GECO2G p = 1.594e−5, NIR-GECO2 vs. NIR-GECO2G p = 0.8796; C FR-GECO1a vs. FR-GECO1c p = 2.474e−5, FR-GECO1a vs. NIR-GECO2 p = 0.3249, FR-GECO1a vs. NIR-GECO2G p = 7.260e−2, FR-GECO1c vs. NIR-GECO2 p = 6.933e−10, FR-GECO1c vs. NIR-GECO2G p = 1.71e−4, NIR-GECO2 vs. NIR-GECO2G p = 8.399e−4; D FR-GECO1a vs. FR-GECO1c p = 0.1057, FR-GECO1a vs. NIR-GECO2 p = 2.227e−4, FR-GECO1a vs. NIR-GECO2G p = 2.761e−4, FR-GECO1c vs. NIR-GECO2 p = 5.785e−5, FR-GECO1c vs. NIR-GECO2G p = 6.629e−5, NIR-GECO2 vs. NIR-GECO2G p = 0.6697. Source data are provided in a Source Data file.
Spectrally compatible combinations of channelrhodopsins and GECIs (that is, a Ca2+ optopatch system) can enable the simultaneous all-optical manipulation and measurement of neural activities across large populations of neurons at a cellular, or even subcellular, resolution. However, the spectral overlap between many of the blue light-activated optogenetic actuators and many of the existing red GECIs based on GFP-like FPs has posed a major hurdle for developing such systems. Specifically, blue light illumination can induce transient increases in red GECI fluorescence16 and the red-shifted GECI excitation light can induce channelrhodopsin photocurrents. Currently, maximum spectral separation is achieved with near-infrared GECIs based on BV-binding FPs, which limits their application in physiological contexts with low BV bioavailability. However, the particularly red-shifted fluorescence excitation spectrum of FR-GECO1a,c presents an opportunity to minimize optical crosstalk with blue light-activated channelrhodopsins such as CheRiff, a fast and sensitive blue-shifted channelrhodopsin with a peak λex of ~460 nm34.
Having verified that FR-GECO1 performs well in zebrafish, we then proceeded to examine the spectral orthogonality between FR-GECO and CheRiff. First, we characterized the blue light activation of FR-GECO1a,c in cultured neuron cells. High intensity 470 nm blue light (23 mW/mm2) illumination only induced a moderate transient increase (ΔF/F0 ≤ 0.1) in far-red fluorescence intensity. In accordance with our in vitro characterization with purified proteins, FR-GECO1a,c fluorescence returned to pre-activation baseline level in <10 ms after removal of blue light illumination (Fig. 4A). We then evaluated the photocurrent of CheRiff during illumination with 594 nm light. Typical photocurrents induced by 594 nm light at imaging intensities were between 2–4% of the maximum blue light-induced photocurrent, and are therefore unlikely to trigger spurious action potentials (Fig. 4B).
A Average fluorescence trace of cultured rat hippocampal neurons expressing FR-GECO1a or 1c via transient transfection, under constant 561 nm illumination (13 mW/mm2) and 100 ms flash of 470 nm LED illumination (23 mW/mm2). Time-average of fluorescence increase induced by 470 nm illumination for each neuron was quantified and shown in red (FR-GECO1a) or blue (FR-GECO1c) dots (n = 6 for both FR-GECO1a and 1c). B HEK293T cells transiently transfected with CheRiff were illuminated by alternating pulses of 488 nm and 594 nm light of increasing intensities (top). Photocurrents were measured under a voltage clamp at −65 mV (middle). Photocurrents at 594 nm were then normalized to the maximum photocurrent induced by excitation with 488 nm (bottom). Typical light intensities used for optopatch experiments are highlighted in orange (n = 3 cells). Average fluorescence traces of cultured rat hippocampal neurons expressing FR-GECO1a (or 1c)-P2A-CheRiff-eGFP-Kv2.1 (AAV transduction) under constant 594 nm illumination (10 mW/mm2) and 470 nm optical stimulation for 0.6 s at 0.07 mW/mm2 (C) or 0.35 mW/mm2 (D) (n = 45 neurons for FR-GECO1a; n = 37 neurons for FR-GECO1c). Blue light crosstalk during the stimulation epochs (dotted lines) has been subtracted from the traces. Error bands represent S.E.M. E Isochronal map of calcium wave in an embryonic zebrafish linear heart tube (LHT) at 24 h post-fertilization (hpf). The map was calculated from a spike-triggered average (n = 12 beats). Each isochron is 50 ms, propagating from left to right. Cardiac marker Nkx2.5:ZsYellow marked in green. Scale bar 50 μm. F Mean FR-GECO1c ΔF/F of the LHT in (E). Simultaneous pacing and measurement of Ca2+ dynamics in the LHT using FR-GECO1a combined with optical stimulation at 1 Hz using CheRiff (G) and CoChR (H). FR-GECOs were imaged using constant 594 nm illumination at 24–290 mW/mm2. Channelrhodopsin was excited using 488 nm light at 24 mW/mm2 (25 ms pulse for CoChR, 75 ms pulse for CheRiff). All images were acquired at 33 Hz. Source data are provided in a Source Data file.
Having demonstrated this minimal optical crosstalk, we proceeded to co-express FR-GECO and CheRiff in cultured neurons. Activation of CheRiff with 470 nm blue light produced Ca2+ transients that were reliably reported by both FR-GECO1a and FR-GECO1c under excitation at 594 nm. CheRiff stimulation with 0.6 s of 470 mm light (0.07 mW/mm2) resulted in ΔF/F0 values of 20% and 60% for FR-GECO1a and FR-GECO1c, respectively (Fig. 4C). When activation light intensity was increased to 0.35 mW/mm2, FR-GECO1a and FR-GECO1c exhibited ΔF/F0 values of 100% and 280%, respectively (Fig. 4D). Collectively, these results indicate that, due to their red-shifted spectra, FR-GECO1a and FR-GECO1c are well-suited for multiplexing with the blue-light activated optogenetic actuator CheRiff.
Following the determination of the spectral orthogonality between FR-GECO and CheRiff in cultured cells and the comparison between FR-GECO and available near-infrared GECIs in zebrafish, we further evaluated this Ca2+ optopatch system in vivo in zebrafish. Previous work has shown that activation of channelrhodopsins expressed in sinoatrial cardiomyocytes can trigger heartbeats35. In the heart, Ca2+ influx is involved in action potential generation, and Ca2+-triggered Ca2+ release from the sarcoplasmic reticulum is involved in triggering muscle contraction. Embryonic zebrafish expressing only FR-GECO1c exhibited an average ΔF/F0 of 7% during rhythmic spontaneous Ca2+ oscillations 24 h post-fertilization (hpf) in the zebrafish LHT (Fig. 4E, F Supplementary Movie 1). To simultaneously pace and measure the Ca2+ dynamics in the LHT, both FR-GECO1a and CheRiff were expressed in zebrafish embryos. Optical stimulation of CheRiff using blue light (488 nm light at 24 mW/mm2 and 1 Hz) induced reliable Ca2+ transients that were reported by FR-GECO1a (λex = 594 nm; Fig. 4G). Similar results were observed with the combination of FR-GECO1a and CoChR36, another blue light-activated channelrhodopsin that is capable of generating large photocurrents and that has previously been used in combination with NIR-GECO (Fig. 4H)21. We attribute the apparent lower magnitude of the CoChR-induced transients primarily to slower Ca2+ re-uptake in a more developmentally immature heart37, driving a higher baseline between transients. These results demonstrated that simultaneous expression of FR-GECO1 and blue light-activated channelrhodopsins enables all-optical cardiac pacing and imaging of zebrafish embryonic hearts during development. More broadly, these results show that FR-GECO1a and 1c are compatible with blue light-activated channelrhodopsins for an optopatch approach to manipulating and imaging cells in culture or in vivo.
Discussion
GFP- and RFP-based GECIs have been extensively engineered and are widely used for imaging cell signaling and neural activity in mammalian tissue in vivo. However, there remains a need for high-performance GECIs with more red-shifted wavelengths, such as indicators with wavelengths in the far-red or near-infrared range. Indicators with longer excitation and emission wavelengths enable deeper tissue imaging, which may be useful for imaging neural activity in subcortical structures, for example. They are also associated with less phototoxicity and less autofluorescence. Crucially, red-shifted indicators are also generally preferable to blue-shifted indicators as they provide better spectral separation from green indicators and commonly used optogenetic actuation tools such as channelrhodopsins, which allow for the manipulation and control of cells of interest. Here, we have reported the engineering and characterization of FR-GECO1a,c, two far-red fluorescent GECIs that are analogous to the GCaMP series of GECIs. Our initial efforts to engineer a far-red Ca2+ indicator using other RFP scaffolds were unsuccessful, likely due to these scaffolds’ oligomerization tendencies. On the other hand, mKelly2 is reported to be a true monomeric far-red FP, a property that likely helps to explain our success at using it as a scaffold to develop far-red fluorescent GECIs.
Both FR-GECO1 variants are bright relative to other red GECIs under 2P excitation. Their optimal 2P Ca2+-dependent ΔF/F0 occurs between 1050 and 1200 nm, overlapping with the range of common tunable 2P lasers such as titanium-sapphire lasers which typically range from 650 to 1100 nm. Previous study has demonstrated that multiphoton excitation at longer wavelengths around 1200–1400 nm or 1600–1800 nm effectively provides optimal tissue imaging depths when considering both tissue scattering and water absorption38. When excited at 1200 nm, FR-GECO1a and FR-GECO1c, in their Ca2+-saturated states, have very high 2P brightness. At this wavelength, their brightness is comparable to commonly used bright far-red FPs, such as mKate2, tdKatushkha2, and mNeptune39. Both indicators are much brighter upon 2P excitation at 1200 nm than any of the red GECIs characterized so far29 (Supplementary Table 3), suggesting that the far-red mKelly2 scaffold may be more promising than the existing red FP scaffolds for the development of bright fluorescent sensors for 2P excitation.
In addition to greater depth penetration afforded by 2P microscopy, an important driving force for the development of red-shifted indicators is their better compatibility with existing blue-light activatable optogenetic tools. However, one concern is that some red GECIs exhibit transient fluorescence increases (photoactivation) when illuminated with blue light16, complicating their use in optogenetics. Through detailed photophysical characterization of FR-GECO1 photoactivation both in vitro and in cells, we have demonstrated that FR-GECO1 transiently photoactivates and recovers to baseline on the millisecond timescale. This fast rise and decay suggest that the photoactivation behavior of FR-GECOs, while not desirable, could still be easily distinguished from action potential-evoked GECI Ca2+ fluorescence changes, which typically happen on a timescale in the order of seconds due to the response kinetics of GECIs16,18,40. The spectral orthogonality and minimal crosstalk enabled us to combine FR-GECO1 with blue-light activated channelrhodopsins (CheRiff or CoChR) to create Ca2+ optopatch for the all-optical stimulation and recording of excitable cells. Previously reported pairings of CheRiff/jRCaMP1b and eTsChR/jRGECO1a were validated in cultured cells and acute tissue slices, respectively41,42. In this work, we further validated the Ca2+ optopatch construct in vivo in the hearts of embryonic zebrafish.
In addition to the unique spectral profile, the FR-GECO1 series offers other advantages relative to other green, red, and NIR GECIs. FR-GECO1c exhibits a larger Ca2+-dependent ΔF/F0 in vitro than most other red GECIs, such as K-GECO1, XCaMP-R and R-GECO117,18,24. The use of ckkap peptide provides FR-GECO1a and FR-GECO1c with high affinities for Ca2+ (Kd < 100 nM), as well as large fluorescence changes and fast response times for detection of small numbers of action potentials, comparable to other ckkap-based GECIs such as K-GECO1 and the XCaMP series, and RS20-based indicators such as R-GECO1, and the GCaMP and RCaMP1 series17,18,24,33,43. The high sensitivity of FR-GECO1c is likely due to its lower Ca2+-free state fluorescence, larger Ca2+-dependent ΔF/F0, and optimal affinity towards Ca2+. Neurons typically have a basal intracellular [Ca2+] of 50 to 100 nM that can increase to 1 to 10 μM upon stimulation44,45. FR-GECO1c’s affinity (Kd = 83 nM) is positioned in the ideal range to detect small and large changes in Ca2+. Indeed, the affinities of FR-GECO1 are comparable to sensitive variants of the late-generation GCaMP series43,46. Relative to NIR-GECO variants20,21, the FR-GECO1a,c variants are not as red-shifted but are brighter per molecule and have a direct (as opposed to inverse) response to Ca2+. Indicators with a direct response to their ligands are generally favored over indicators with an inverse response due to a better SNR owing to lower background fluorescence. In addition, FR-GECO1c offers better sensitivity and faster kinetics than NIR-GECO2 in neuronal activity detection21.
As first-generation GECIs, FR-GECO1a and FR-GECO1c both have the potential for further improvements. A general trend among FPs is that hypsochromic shifts in peak wavelengths are accompanied by reductions in quantum yield, thus limiting the brightness of available far-red and near-infrared FPs. Future versions of FR-GECO should be optimized for improved brightness to further facilitate imaging in vivo and even faster kinetics. Our pH titrations revealed that both FR-GECO1 variants have a significantly more basic pKa than mKelly2 and so they do not exhibit their maximum brightness under physiological conditions. Accordingly, engineering variants with lower pKa values may lead to improved brightness for imaging in tissue. Additionally, FR-GECO1a,c are the only non-biliverdin-binding GECIs that can be excited by the widely used red helium-neon lasers (~633 nm). Yet, due to the misalignment between the excitation peak (596 nm) and the laser wavelength, the excitation is only ~12.5% of the possible maximum. Further red-shifted FR-GECO1a,c variants with better compatibility with helium-neon lasers could be another important future direction. FR-GECO1a and FR-GECO1c show substantial differences in performance, attributable to a single mutation, Tyr35Thr, at the interface of the FP and the CaM/ckkap domains. To obtain insight into the mechanisms of these biosensors, and the impact of the Tyr35Thr mutation, an experimentally-determined molecular structure of FR-GECO1 will likely be required. Such a structure could further our understanding of the mechanisms underpinning the performance FR-GECO1 and, consequently, serve to guide the development of improved generations of the FR-GECO series.
In summary, we have developed the FR-GECO series of far-red GECIs that expands the spectral palette of available GECIs that utilize archetypical (i.e., non-biliverdin-binding) FPs. Although only the first iteration, FR-GECO1a,c have good brightness, high affinities for Ca2+, and very good sensitivity for neuronal activity detection. We have also successfully shown that FR-GECO1a,c can be used in conjunction with optogenetic actuators in all-optical studies. For these reasons, we are certain that FR-GECO1a,c are valuable additions to the GECI toolkit. Moreover, we anticipate that the cpmKelly2 scaffold of FR-GECO1 can serve as a template for engineering FP-based far-red indicators for ligands other than Ca2+.
Methods
Ethics statement
This research complies with all relevant ethical regulations. Procedures for preparation of primary neurons at Université Laval were approved by the Animal Ethics Committee and were performed according to Canadian Biosafety Guidelines in a Containment Level 2 certified lab. Procedures at Janelia Research Campus were approved by the Institutional Animal Care and Use Committee (IACUC) and the Institutional Biosafety Committee (IBC). All vertebrate experiments at Harvard University were approved by the Institutional Animal Care and Use Committee of Harvard University or the Harvard Medical Area Standing Committee on Animals. Procedures in the United States conform to the National Institutes of Health (NIH) Guide for the Care and Use of Laboratory Animals.
General
Synthetic DNA oligonucleotides and double-stranded fragments (gBlocks) were sourced from Integrated DNA Technologies. CloneAmp HiFi PCR Premix (Takara Bio USA, Inc., 639298) and Q5 polymerase (New England Biolabs, M0491L) were used for high fidelity polymerase chain reactions (PCRs). Random mutagenesis PCRs were performed with Taq polymerase (Thermo Fisher Scientific, EP0406. PCR products were purified by preparative agarose gel electrophoresis and extracted with GeneJET Gel Extraction Kit (Thermo Fisher Scientific, K0692). Restriction enzymes and T4 DNA ligase (EL0011) were purchased from Thermo Fisher Scientific and used according to the manufacturer’s recommended protocols. Digested PCR products were extracted with the GeneJET Gel Extraction Kit before ligation. In-Fusion HD (Takara Bio USA, Inc., 639650) was used for assembly reactions. Plasmid DNA extractions were performed with GeneJET Plasmid Miniprep Kits (Thermo Fisher Scientific, K0503). Sanger sequencing reactions were performed by the Molecular Biology Services Unit at the University of Alberta. Fluorescence measurements were performed on a Safire2 plate reader (Tecan), while absorbance measurements were collected on a DU-800 UV-Visible Spectrophotometer (Beckman).
Protein engineering
Starting template construction
Double-stranded DNA encoding for K-GECO117, with its circularly permuted (cp) FP replaced with cpmKelly212, was commercially synthesized with overlapping sequences with pBAD/HisB (Invitrogen, V43001) in the 5′ and 3′ directions to facilitate insertion between the XhoI and HindIII sites. This fragment was then assembled into pBAD/HisB vector digested with XhoI (Thermo Fisher Scientific, FD0694) and HindIII (Thermo Fisher Scientific, FD0504), and the assembly reaction was used to transform electrocompetent DH10B E. coli (Invitrogen, 18290015 and 12033015) prior to overnight incubation on Lennox Broth (LB; Fisher Scientific, BP1427-500) agar plates supplemented with 400 µg/mL ampicillin (Fisher Scientific, FLBP17605) and 0.02% (w/v) l-arabinose (Alfa Aesar, AAA1192130) at 37 °C. Plasmids were isolated from single colonies grown in 4 mL liquid LB cultures with 100 µg/mL ampicillin, and sequenced by Sanger sequencing.
Plasmids for mammalian expression
Genes encoding FR-GECO1a and FR-GECO1c were amplified by high-fidelity PCR using primers that add vector homologous sequences to the 5′ and 3′ end and assembled into linearized pcDNA3.1/Puro-CAG-ASAP1 to replace the gene encoding for ASAP1 (Addgene plasmid #52519, a gift from Michael Lin)47.
Library construction
To generate random mutagenesis libraries, mutations were introduced into the construct by error-prone PCR amplification with Taq polymerase in the presence of MnCl2 (varies, up to 0.15 mM; Fisher Scientific, M87-100) and a 1:5 ratio of dATP and dGTP to dCTP and dTTP (Thermo Scientific, R0181). The purified PCR product was then assembled with an XhoI and HindIII-digested pBAD/HisB vector. Site-directed mutagenesis libraries were generated by PCR amplification with high fidelity polymerases and primers carrying the codons for the desired mutations followed by DpnI (Thermo Fisher Scientific, FD1703 and FD1704) digestion. The NNK codon was used for complete randomization. For mutations at just one position, amplification was performed with the CloneAmp HiFi PCR Premix, while libraries with mutations at multiple locations were generated by QuikChange Lightning Multi Site-Directed Mutagenesis Kit (Agilent Technologies, 200531).
Library screening
DH10B E. coli were transformed with DNA encoding the library and cultured on LB agar plates supplemented with ampicillin and l-arabinose, as described earlier. The E. coli colonies expressing the library were then screened on the plate based on fluorescence intensity using a custom imaging system described previously48. Briefly, this imaging system illuminates a Petri dish with light from a Lambda LS 175 W xenon arc lamp (Sutter Instruments) that has passed through an excitation filter (Chroma, ET560/40×) in a filter wheel (Lambda 10-3 from Sutter Instruments). Fluorescence is viewed either by eye through tinted goggles (Forensics Source), or using a CCD camera (QImaging) connected to a personal computer. Colonies displaying the highest fluorescence intensity were then cultured in 4 mL liquid LB with ampicillin and arabinose overnight at 37 °C. Proteins were then extracted from cells using B-PER (Thermo Fisher Scientific, 78248) and subjected to a secondary screen by measuring their fluorescence intensities in Ca2+-free (30 mM 3-(N-morpholino)propanesulfonic acid (MOPS; Thermo Fisher Scientific, A17214-22), 100 mM KCl (Sigma Aldrich, 529552-250GM), 10 mM ethylene glycol-bis(β-aminoethyl ether)-N,N,N’,N’-tetraacetic acid (EGTA, pH 7.2; MP Biomedicals, ICN1948238), and Ca2+ buffers (30 mM MOPS, 100 mM KCl, 10 mM Ca-EGTA, pH 7.2) to determine their Ca2+-dependent ΔF/F0. Plasmids for variants showing the largest Ca2+-dependent ΔF/F0 and highest fluorescence intensities were extracted, sequenced, and used as the template(s) for the next round of evolution.
In vitro characterization
Protein expression and purification
Electrocompetent DH10B E. coli were transformed with the pBAD/HisB plasmid carrying the gene encoding the protein of interest and grown on solid media. Single colonies from the transformed bacteria were then used to inoculate 4 mL of a starter culture supplemented with ampicillin and l-arabinose that was then shaken at 225 rpm in 37 °C overnight. The starter culture was added to 500 mL of LB with 100 µg/mL ampicillin and the culture was shaken at 37 °C. After 4 h, 0.02% l-arabinose is added to induce expression, and the culture is shaken for another 4 h at 37 °C before harvesting the bacteria by centrifugation. Bacteria were then resuspended in 1× TBS (50 mM Tris-HCl, 150 mM NaCl, pH 7.5; Tris base from Fisher Scientific, BP152-5; NaCl from Fisher Scientific, BP3581), and lysed by sonication. The lysate was clarified by centrifugation and the cleared lysate was incubated with Ni-NTA resin (G Biosciences, 786-408) for at least 1 h. Resin bound to protein was washed with TBS wash buffer (1× TBS with 20 mM imidazole, pH 8.0; imidazole from Thermo Scientific, 396741000), followed by elution with TBS elution buffer (1× TBS with 250 mM imidazole, pH 7.8). The purified protein was then left at room temperature overnight to facilitate protein folding and chromophore maturation before concentrating and buffer exchanging into 1× TBS using 10 kDa centrifugal filter units (Millipore Sigma, UFC801024 and UFC 901024). Unless noted otherwise, all steps were carried out at 4 °C or on ice.
Brightness, affinity, and pH-sensitivity measurement
Effective ECs of FR-GECO variants were measured using the alkali denaturation method26. Briefly, purified protein for each FR-GECO variant was diluted in NaOH, Ca2+-free (30 mM MOPS, 100 mM KCl, 10 mM EGTA, pH 7.2), and Ca2+ buffer (30 mM MOPS, 100 mM KCl, 10 mM Ca-EGTA, pH 7.2), and the absorption spectrum for each sample was collected. Because the same amount of protein was used for each set of measurements for each protein, the concentration of protein present in each sample can be calculated according to Beer’s Law and the previously reported EC of 44,000 for the absorbance peak around ~450 nm, which corresponds to the denatured tyrosine-based chromophore26,49. Further application of Beer’s Law with the calculated concentration for each variant yields the EC in the Ca2+-bound and Ca2+-free states. Measurements were performed in triplicate and the results were averaged. Determination of QY was performed according to previously established protocols using mKelly2 as the standard12,24,48. For each protein, a series of diluted samples were prepared in Ca2+-free and Ca2+ buffer from the samples used for extinction coefficient measurements such that the peak absorbance was equal to, or less than, 0.05. For each dilution series, the emission spectra were collected from 590 to 750 nm with an excitation wavelength of 570 nm. The total fluorescence intensities for each dilution were integrated, plotted against the absorbance, and the slope of each line (m) was calculated. Quantum yields were then calculated using the published quantum yield for mKelly2 (0.18)12 and according to the equation:
Buffers for the Kd determination, with free [Ca2+] ranging from zero to 39 µM, were prepared by mixing appropriate volumes of Ca2+-free and Ca2+ buffers50. Purified FR-GECO variants were diluted in these buffers and the fluorescence intensities of the protein in each solution were measured in triplicate. The readings were plotted against the free Ca2+ concentration on a logarithmic scale, and the data fitted to the Hill equation for the Kd and apparent Hill coefficient. Aliquots of a buffer containing 30 mM trisodium citrate and 30 mM borax (sodium borate) were adjusted using HCl or NaOH so that their pH ranged from 3 to 11. Concentrated protein, as well as 1 μL of either 200 mM EGTA or CaCl2 for the Ca2+-free and Ca2+ conditions, respectively, were then diluted in these buffers for a total volume of 100 μL. The fluorescence intensities were measured in triplicate, and the readings were plotted against pH. Due to the ionization events at pH ~5 to 6, only the data for pH 6.5 to 11 were fitted to the Hill equation for the pKa.
In vitro kinetics measurement
Rapid kinetic measurements of FR-GECO1a and FR-GECO1c in purified protein were made using an Applied Photophysics SX-20 Stopped-Flow Reaction Analyzer using fluorescence detection. The dead time of the instrument was 1.1 ms. The excitation wavelength used was 625 nm with 2 nm bandwidth and emitted light was collected at 690 nm through a 10 mm path at room temperature. For koff measurements, 2 μM of each FR-GECO1a and FR-GECO1c proteins in 1 mM Ca2+ (30 mM MOPS, 100 mM KCl, pH 7.2) were rapidly mixed at 1:1 ratio with 100 mM of EGTA (30 mM MOPS, 100 mM KCl, pH 7.2) at room temperature. Measurements were taken for five replicates. koff values were determined by fitting a single exponential dissociation curve to the signal decay with units of s−1, using Prism GraphPad. For kon, both variants, buffered in 30 mM MOPS and 100 mM KCl (pH = 7.2), were rapidly mixed at 1:1 ratio with varying concentrations of Ca2+ produced by reciprocal dilutions of 10 mM EGTA and 10 mM CaEGTA by using the Calcium Calibration Buffer Kit (Thermo Fisher Scientific, C3008MP). The slope of kobs was used to determine the kon rate in the units of s−1 M−1. The tau values for both on and off rates were calculated as the reciprocal of the kon and koff rates, respectively. Two replicates were used for each Ca2+ concentration.
Photoactivation measurement
Microdroplets of FR-GECO1a and FR-GECO1c in purified protein in either Ca2+ or Ca2+-free buffer (pH 7.2) were prepared using octanol (Sigma-Aldrich). The prepared sample was sandwiched between pre-silanized coverglass and coverslip (Sigma-Aldrich). Using a widefield microscope each droplet was continuously illuminated with a 561 nm laser for 120 s. 488 nm laser was used for photoactivation while the 561 nm laser was on. The 488 nm laser was on for 50 ms with the shutters switching on at various frequencies (0.2, 1, or 2 Hz).
2P characterization
Experimental setup for 2P spectral measurements includes a tunable femtosecond laser InSight DeepSee Dual (Spectra Physics) coupled with a photon-counting spectrofluorometer PC1 (ISS)29,51. The 2P fluorescence excitation (2PE) spectra were measured by automatically stepping laser wavelength and recording total fluorescence intensity at each step. A combination of filters, including FF01-770/SP, FF01-680/SP (both Semrock), and FGL630 long-pass (Thorlabs) was used in the left emission channel of the PC1 spectrofluorometer to eliminate scattered laser light. To scale the 2P excitation spectra, we use a parameter called 2P molecular brightness F2(λ). This parameter depends on fractional contributions to 2P brightness of the anionic and neutral forms at excitation wavelength λ and can be presented as follows:
where φ is the fluorescence quantum yield of form A or N, and σ2 is the 2P absorption cross-section. Since we collect fluorescence spectrum through an FGL630 long-pass filter, technically φN only comprises a part of fluorescence originating from anionic deprotonated chromophore that appears after the excited-state proton transfer from initially excited neutral form. To obtain the 2 P excitation spectrum in units of molecular brightness, we independently measured ρA, φA, and σ2,A(λ) of the anionic form for both FR-GECO1a,c in Ca2+-free and Ca2+-saturated states and normalized the unscaled 2P spectrum to the product ρAφAσ2,A(λ). The cross-section σ2,A(λ) was measured at the wavelength where the contribution of the neutral form is negligible (λ = 1064 nm). This measurement was performed using Rhodamine B (Exciton, 06101) in alkaline ethanol as a reference standard. Its cross-section was measured relative to Rhodamine 6 G (Sigma-Aldrich, 252441) in ethanol (with the average <σ2,RhB/σ2,Rh6G> = 1.072 ± 0.017). Using this number together with the Rhodamine 6G cross-section in ethanol (σ2,Rh6G) obtained after averaging of seven independent measurements51: <σ2,Rh6G> = 12.4 ± 2.6 GM, the cross-section of Rhodamine B in ethanol at 1064 nm was calculated σ2,RhB = 13.3 ± 2.8 GM and used as a reference. In a modified version of previously reported method51, a total (without monochromator) 2P excited (at 1064 nm) fluorescence signals I as a function of laser power P were collected for both the sample and reference solutions (solutions were held in 3 × 3 mm cuvettes (Starna) with a maximum optical density less than 0.1). The fluorescence was collected at 90° to excitation laser beam through FF01-770/SP, FGL630 Longpass, and long wave pass 561 (Edge BasicTM, Semrock) filters, using the left emission channel of a PC1 spectrofluorometer working in a photon counting mode. The power dependences of fluorescence were fit to a quadratic function I = aP2, from which the coefficients aS and aR were obtained for the sample (index S) and reference (index R) solutions, respectively. Second, the 1 P excited fluorescence signals were measured for the same samples and in the same registration conditions. In this case, strongly attenuated radiation of a 561 nm line of a Sapphire 561-50 CW CDRH (Coherent) laser was used for excitation. The fluorescence power dependences for the sample and reference were measured and fit to a linear function: I = bP, from which the coefficients bS and bR were obtained. The 2 P absorption cross-section was then calculated as follows:
Here, λ1 is the wavelength used for 1P excitation (561 nm), λ2 is the wavelength used for 2 P excitation (1064 nm), εR,S(λ1) are the corresponding extinction coefficients, measured at λ1. (In contrast to the one-photon characterization section, we define extinction coefficient (εS) here to be of a particular, anionic, form. It relates to the previously defined extinction coefficient, e.g. shown in Table 1, as follows: εS = ε/ρA) This approach allows us to automatically correct for the laser beam properties (pulse duration and spatial intensity distribution), fluorescence collection efficiencies for 1P and 2P modes, PMT spectral sensitivity, differences in QYs and concentrations between S and R solutions. The molecular brightness of the anionic form was then calculated as a product F2 = ρAφAσ2,A(λm) with the 2P cross-section taken at spectral maximum, λm, for both states of the indicator. The QYs, ECs, and fractional concentrations were all measured independently. Finally, the 2P excitation spectra were scaled to the calculated F2(1064 nm) values.
Imaging in cultured cells
Imaging in HeLa cells
The genes coding FR-GECO1a and FR-GECO1c were inserted into the pcDNA3.1 vector (Thermo Fisher Scientific, V79020) between the XhoI and HindIII restriction sites. The cells were cultured in Dulbecco’s Modified Eagle Medium (Nakalai Tesque, 08456-65) supplemented with 10% fetal bovine serum (Sigma-Aldrich, F7524-500ML) and 1% penicillin–streptomycin (Nakalai Tesque, 09367-34). HeLa cells (~104, ATCC, CCL-2) were seeded into 35-mm glass-bottom (collagen coated) cell-culture dishes (Iwaki, 3911-035) with 200 μL opti-MEM (Thermo Fisher Scientific, 31985-070). Cells were transfected with the pcDNA-FR-GECO variants plasmid (2 μg) using 4 μL polyethyleneimine (Polysciences, 24765-1, 1 mg polyethyleneimine diluted in 1 mL ultrapure water). Transfected cells were imaged three days after transfection using a IX83 wide-field fluorescence microscopy (Olympus) equipped with a pE-300 LED light source (CoolLED), a 40× objective lens (numerical aperture = 1.3; oil), an ImagEM X2 EM-CCD camera (Hamamatsu), Cellsens software (Olympus), and a STR stage incubator (Tokai Hit). The filter sets for imaging were 545/20 nm excitation, 645-nm dclp dichroic mirror, and 678/65 nm emission. For imaging of Ca2+-dependent fluorescence, the cells were washed twice with Hank’s balanced salt solution with Ca2+ (HBSS(+); Nakalai Tesque, 09735-75), and the buffer was exchanged to 1 mL HBSS(+) supplemented with 10 mM HEPES (Nakalai Tesque, 17557-94) and incubated at 37 °C. Images were acquired every 5 s with an exposure time of 100 ms. After 1 min of imaging, histamine (Wako, 087-03533, 50 μM final concentration) was added. Imaging continued for 11 min, and then cells were washed twice with HBSS without Ca2+ (HBSS(-)) (Nakalai Tesque, 17461-05), and 1 mL of HBSS(−) was added. After 1 min, EGTA/ionomycin (2.5 mM/1.5 μM final concentration) were added to fully deplete Ca2+. Once a steady fluorescence intensity was reached, the dish was washed twice more and supplemented with HBSS(−). Ca2+/ionomycin (5 mM/1.5 μM final concentration) was then immediately added to saturate the indicator.
Imaging of spontaneous activity in cultured neurons
Primary rat cortical neurons were prepared and maintained by the CERVO Brain Research Centre’s Neuronal Culture Platform as previously described52,53. Briefly, neonatal rats were sacrificed and their cortices were dissected and dissociated with papain (Worthington, LS003126). Dissociated neurons were plated on 12 mm PDL-laminin coated glass coverslips and cultured in neurobasal media (Gibco, 21103049) supplemented with B-27 (Gibco, 17504044), GlutaMax (Gibco, 35050-061), FBS (Hyclone, SH30071.03), and penicillin/streptomycin (Gibco, 15140031), with Ara-C (Sigma Aldrich, C−1768) added 5 days after plating. Neurons were then fed twice a week by replacement of half the growth medium with serum- and Ara-C-free growth medium. At 5 days in vitro (DIV), neurons were transduced with adeno-associated viruses (AAVs) encoding FR-GECO1-WPRE (2.4 × 1010 GC/well) under the control of either the CAG or human synapsin promoter and imaging was performed on DIV 15. Prior to imaging, neurons were washed thrice in HBSS supplemented with 10 mM HEPES (pH 7.4). Images were acquired on a Zeiss Observer Z.1 equipped with a 20× objective (NA = 0.4, LD Plan-Neofluar 20×, Zeiss), a CCD camera (Axiocam 503 m), HXP 120 V light source, and filter set 00 (BP 530-585 nm excitation, FT 600 nm dichroic mirror, and LP 615 nm emission; Zeiss). Images were acquired for 20 min with 5 s intervals, 1.5 s exposure time and 2 × 2 binning. Images were first aligned to correct for drift as needed, and baseline corrected prior to analysis in ImageJ and plotting on Prism GraphPad.
Imaging of cultured neurons with electrical stimulation
Primary rat hippocampal and cortical neurons were prepared and maintained as described previously46. Briefly, neonatal rat pups (Charles River Laboratory) were euthanized, and hippocampal and cortical tissues were isolated and dissociated using papain (Worthington) in 10 mM HEPES (pH 7.4) in Hanks’ Balanced Salt Solution, incubating at 37 °C for 30 min. The cell suspension was triturated, filtered (40 µm), and electroporated with 400 ng of plasmid DNA in nucleofection solution (Lonza). Neurons were plated at 1 × 105 cells per well in poly-D-lysine-coated, 96-well glass-bottom plates (MatTek) with a 4:1 ratio of NbActiv4 (BrainBits) to plating medium (28 mM glucose, 2.4 mM NaHCO3, 100 µg/mL transferrin, 25 µg/mL insulin, 2 mM L-glutamine, 100 U/mL penicillin, 10 µg/mL streptomycin, and 10% FBS in MEM). On the following day, 190 µL of NbActiv4 medium was added to each well and plates were incubated at 37 °C with 5% CO2 for 12–15 days before imaging.
Cultured neurons were transfected with an expression vector with hSyn1-NES-FR-GECO1-WPRE, where FR-GECO1 included the leader sequence of pBAD, using electroporation (Lonza Nucleofector) according to the manufacturer’s instructions. Imaging and the corresponding analysis were performed as described previously46. Specifically, prior to the imaging session, transfected neurons were first washed thrice with imaging buffer (140 mM NaCl, 0.2 mM KCl, 30 mM glucose, 10 mM HEPES, pH 7.4), and treated with synaptic blockers (10 μM CNQX, 10 μM CPP, 10 μM gabazine, and 1 mM MCPG (Tocris Bioscience)). Neurons were imaged using an inverted Olympus IX81 microscope equipped with a 10× objective (NA = 0.4, UPlanSApo, Olympus), an EMCCD camera (Ixon Ultra DU897, Andor), a 565 nm LED for excitation (Cairn Research) and mCherry filter cubes (#49008, Chroma). Neurons were stimulated at room temperature by field stimulation with a custom-built electrode. Trains of 3, 10 and 160 APs were stimulated and fluorescence images were acquired at 206 Hz throughout the stimulation session. Reference images were taken after stimulation to perform cell segmentation during analysis.
For image analysis, cell segmentation was performed using the Suite2P package in Python. Background fluorescence was subtracted from the images before calculating ΔF/F0. The ΔF/F0 value for each segmented cell was determined by dividing the fluorescence intensity after stimulation by the baseline fluorescence (F0) and subtracting 1. Half-rise time and half-decay time were computed by fitting a one-phase exponential rise or decay curve on the fluorescence traces.
Ca2+ Optopatch
Cell culture
HEK293T cells (ATCC, CRL-3216) were grown at 37 °C, 5% CO2, in DMEM with 10% fetal bovine serum (Life Technologies, 10082−147) and 50 U/mL penicillin-streptomycin (Life Technologies, 15060-063). Cell confluency was maintained at <80% by splitting every 2-3 days using trypsin-EDTA (Life Technologies, 25300054). Cell passage numbers were maintained <15. Hippocampal neurons from postnatal day 0 (P0) rat pups were dissected and cultured at a density of 30,000–40,000 cells per cm2 on a laminin (Fisher Scientific, 23-017-015) and poly-D-lysine (PDL, Sigma-Aldrich, P1024) coated glass-bottom dish (Cellvis, D35−14−1.5-N), using Brainphys Neuronal Medium (STEMCELL Technologies, 05792, 05794), following the manufacturer’s protocols.
DNA constructs
A Kv2.1 soma-localization sequence was appended to the C terminus of CheRiff-eGFP54 (Addgene plasmid #51693). To combine optogenetics with Ca2+ imaging, soma-localized CheRiff-eGFP-Kv2.1 and FR-GECO1a (or 1c) were linked with a ribosomal skip sequence P2A and co-expressed from a single vector55. An AAV backbone for Cre-dependent expression under hSyn promoter (Addgene plasmid #126512) was linearized via restriction digestion with NheI (New England Biolabs, R3131S) and AscI (New England Biolabs, R0558S). The FR-GECO, P2A, and CheRiff-eGFP-Kv2.1 were cloned into the AAV backbone via Gibson assembly (New England Biolabs, E2611S) to obtain hSyn-DiO-FRGECO1a(or 1c)-P2A-CheRiff-eGFP-Kv2.1.
Imaging and electrophysiology
Cell culture media were replaced by the imaging buffer, consisting of 125 mM NaCl, 2.5 mM KCl, 2.5 mM HEPES, 30 mM glucose, 1 mM MgCl2, 3 mM CaCl2, with a pH of 7.3 and osmolarity of 310 mOsm. For photocurrent measurement, HEK293T cells were transfected with FCK-CheRiff-eGFP (Addgene plasmid #51693) using TransIT-293 (Mirus, MIR2700). After two days, cells were plated on PDL-coated glass-bottom dishes at a density of ~ 100,000 cells/cm2. Cells were allowed to settle and adhere for 6 h prior to measurement. Borosilicate glass capillaries (WPI, 1B150F-4) were pulled to tip resistances of 5-10 MΩ, and filled with internal solution of 125 mM potassium gluconate, 8 mM NaCl, 0.6 mM MgCl2, 0.1 mM CaCl2, 1 mM EGTA, 10 mM HEPES, 4 mM Mg-ATP, 0.4 mM Na-GTP, adjusted to pH 7.3 and 295 mOsm with sucrose. For photocurrent measurement, HEK293T cells were voltage-clamped to -65 mV, then illuminated with alternating 1 s pulses of 488 nm (Coherent, Obis 1226419) and 594 nm laser light (Huber Photonics, 0594-04-01-0100-300) at intensities 1-17 and 0.5–75 mW/mm2, respectively. Current recordings were collected using MultiClamp 700B amplifier (Molecular Devices), filtered at 5 kHz with the amplifier’s internal Bessel filter, and digitized at 10 kHz.
For photoswitching characterization, neurons were transfected with CAG-FR-GECO1a (or 1c) plasmid using Lipofectamine 2000 (Fisher Scientific, 11-668-027) on DIV 7-10, and imaged on DIV 14-18 using a custom microscope described previously42. For photoswitching measurements, neurons were continuously illuminated with 561 nm light (MPB Communications, F-04306-02), at 13 mW/mm2 for 10 s, while 100 ms flashes of 470 nm LED (Thorlabs, M470L3) light at 1 Hz (23 mW/mm2) were used to induce photoswitching. Fluorescence emission was collected using a 10× objective (Olympus, XLPan N 10× SVMP, NA 0.6), filtered through a band-pass filter (Chroma, ET600/50 m), and imaged onto a scientific CMOS camera (Hamamatsu, Orca Flash 4.0) at 100 Hz.
For all-optical neurophysiology measurement, cultured rat hippocampal neurons at DIV 3-4 were transduced using AAVs encoding hSyn-Cre and hSyn-DIO-FR-GECO1a(or 1c)-P2A-CheRiff-eGFP-Kv2.1. Cells were imaged on DIV 14–19. Neurons were illuminated with continuous 594 nm light (Hubner Photonics, Cobolt MamboTM 100 mW) at 10 mW/mm2, while a 600 ms pulse of 470 nm light at 0.07 or 0.35 mW/mm2 was used to stimulate CheRiff. Fluorescence emission was collected using a wide field 2× objective (Olympus, MVPLAPO 2XC, NA 0.5), filtered through a bandpass filter (Semrock, FF02-641-75), and imaged at 30 Hz.
Data analysis
To determine the crosstalk of 594 nm light into CheRiff activation, the 594 nm photocurrents were normalized by the maximum (saturating) 488 nm photocurrent. The 488 nm photocurrents were calculated by averaging the photocurrent during the last 100 ms of a 1 s long stimulus. The 594 nm photocurrents were calculated by averaging the photocurrent during the last 500 ms of a 1 s long stimulus.
For optical crosstalk and all-optical neurophysiology analysis, fluorescence traces were extracted by averaging the movie pixels in manually defined regions of interest containing single neurons. To correct for the fluorescence background of the imaging buffer, pixels surrounding the neuron were averaged then subtracted from the fluorescence traces. The mean optical crosstalk response was calculated by averaging five trials of blue light stimulation in cells expressing FR-GECO1a or 1c and not expressing a channelrhodopsin. To calculate the optical crosstalk artifact (ΔF/F0), we defined F0 as the mean fluorescence during the initial 100 ms of orange-only illumination, and ΔF as the mean increase in fluorescence during the 100 ms blue light stimulus. To calculate fluorescence traces during the all-optical neurophysiology experiments, optical crosstalk was first corrected by subtracting the fluorescence increase at the onset of the blue light stimuli. This procedure relied on the fact that optical crosstalk was effectively instantaneous, while optogenetically induced Ca2+ increases took several tens of milliseconds to develop. F0 was calculated by the 2 s average of the baseline fluorescence and ΔF was calculated as the increase in fluorescence above the baseline.
In vivo imaging in embryonic zebrafish
GECI comparisons in embryonic zebrafish
TgBAC(-36nkx2.5:ZsYellow) embryos were injected with 20 fmol of GECI mRNA, in addition to 25 pg of α-bungarotoxin mRNA (synthesized using pmtb-t7-alpha-bungarotoxin (Addgene plasmid #69542) and the mMESSAGE mMACHINE SP6 Transcription Kit (Thermo Fisher Scientific, AM1340)) and 4 ng of tnnt2a morpholino (Gene Tools). 30× Danieau buffer was prepared according to the following recipe: 1740 mM NaCl, 21 mM KCl, 12 mM MgSO4•7H2O, 18 mM Ca(NO3)2, 150 mM HEPES buffer. 25 µM biliverdin (Sigma, 30891) or vehicle (1% DMSO in 0.3× Danieau buffer, pH 7.4) was added to embryos at 14 hpf. Embryos were raised to 24-30 hpf and dechorionated using 1 mg/mL Pronase (Sigma, 53702). All sensors were imaged for 30 s at 40 Hz frame-rate with 23 mW/mm2 illumination intensity, using an Olympus XLPLN25XWMP2 objective (25×, water immersion). FR-GECO1 variants were excited using a Cobolt Mambo 594 nm laser and emission fluorescence was collected through a 640/57 nm filter. Near-infrared sensors were excited using an Opto-Engine MLL-FN-639 laser and emission fluorescence was collected through a 665 nm long-pass filter (Chroma, ET665lp). Cardiac tissue was identified by the Otsu thresholding technique applied to an image of ZsYellow fluorescence. The point with the highest GECI intensity within the cardiac tissue was selected as the centroid of a circle with radius 3 µm for extraction of signal. Mean intensity and ΔF/F for each signal trace were calculated after subtracting mean intensity of control embryos without GECI injection and performing correction for photobleaching by subtracting the local minimum value in a trace over a moving window of two seconds as described previously34. SNR was calculated by taking the ratio of the mean power in the fluorescence dynamics between 0.1 Hz and 1.5 Hz over the mean power outside of this frequency band. Embryos with SNR < 5 were discarded. Statistical comparisons were performed using Welch’s t-test with Benjamini-Hochberg multiple test correction (Shapiro-Wilk test for normality p > 0.5 for all groups).
All-optical electrophysiology of embryonic zebrafish
Zebrafish Danio rerio were raised at 28 °C. The transgenic line TgBAC(-36nkx2.5:ZsYellow) (AB background) was used to mark the embryonic heart56. Open reading frames of FR-GECOs and channelrhodopsins, CheRiff and CoChR, were cloned into the pMTB plasmid backbone57 and capped mRNA was synthesized using the mMESSAGE mMACHINE™ SP6 Transcription Kit (Thermo Fisher Scientific). 100 pg of FR-GECO mRNA was injected into fertilized eggs at the 1-cell stage, in addition to 25 pg of α-bungarotoxin mRNA58 to arrest tail twitches and 4 ng of tnnt2a morpholino oligonucleotide (CATGTTTGCTCTGATCTGACACGCA, Gene Tools) to arrest heart motion59. 25 pg of CoChR-GFP and 50 pg of CheRiff-GFP RNA were included in the injection mix in the respective experiments. Embryos were raised to 20 hpf and dechorionated using 1 mg/mL Pronase (Sigma). FR-GECOs were imaged using a Cobolt Mambo 594 nm laser (100 mW) and channelrhodopsins were stimulated using a Coherent Obis 488 nm laser (100 mW). The blue light was targeted to the heart using a digital micromirror device to avoid spurious stimulation of ChR in other tissues. Images were recorded using an Olympus XLPLN25XWMP2 objective (25×, water immersion) and a Hamamatsu Orca Flash sCMOS camera. Segmentation was performed by thresholding an nkx2.5:ZsYellow image and manually adjusting it. ΔF/F traces were obtained by subtracting background, performing a linear photobleach correction, and taking the 10th percentile of fluorescence intensity as the baseline. Spike-triggered averages were obtained by performing peak detection on the ΔF/F trace and averaging across samples within a window flanking the peak. Spatial isochrons were calculated by identifying the time to half maximum of the spike-triggered average at every image pixel and spatially smoothing with a mean filter.
Statistics and reproducibility
Details of the exact statistical tests used, and sample sizes used, are provided in the figure legends. Replication information for all data reported in this study is stated in figure legends. All repetitions were deemed successful. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. Plasmid constructs encoding FR-GECO1a and FR-GECO1c with a CAG promoter are available through Addgene (Plasmid IDs #163681 and #163682, respectively). The full DNA and protein sequences of FR-GECO1a and FR-GECO1c are provided in Supplementary Table 4. The previously reported crystal structure for FusionRed which was used as a reference for numbering mutations, is available under PDB ID 6U1A. Source data are provided with this paper.
References
Tromberg, B. J. et al. Non-invasive in vivo characterization of breast tumors using photon migration spectroscopy. Neoplasia 2, 26–40 (2000).
Lyons, S. K., Patrick, P. S. & Brindle, K. M. Imaging mouse cancer models in vivo using reporter transgenes. Cold Spring Harb. Protoc. 2013, 685–699 (2013).
Abbyad, P., Childs, W., Shi, X. & Boxer, S. G. Dynamic Stokes shift in green fluorescent protein variants. Proc. Natl Acad. Sci. USA 104, 20189–20194 (2007).
Lin, M. Z. et al. Autofluorescent proteins with excitation in the optical window for intravital imaging in mammals. Chem. Biol. 16, 1169–1179 (2009).
Strack, R. L. et al. A rapidly maturing far-red derivative of DsRed-Express2 for whole-cell labeling. Biochemistry 48, 8279–8281 (2009).
Chu, J. et al. Non-invasive intravital imaging of cellular differentiation with a bright red-excitable fluorescent protein. Nat. Methods 11, 572–578 (2014).
Shen, Y., Lai, T. & Campbell, R. E. Red fluorescent proteins (RFPs) and RFP-based biosensors for neuronal imaging applications. Neurophotonics 2, 031203 (2015).
Hense, A. et al. Monomeric Garnet, a far-red fluorescent protein for live-cell STED imaging. Sci. Rep. 5, 18006 (2015).
Bajar, B. T. et al. Fluorescent indicators for simultaneous reporting of all four cell cycle phases. Nat. Methods 13, 993–996 (2016).
Shen, Y., Chen, Y., Wu, J., Shaner, N. C. & Campbell, R. E. Engineering of mCherry variants with long Stokes shift, red-shifted fluorescence, and low cytotoxicity. PLoS ONE 12, e0171257 (2017).
Matela, G. et al. A far-red emitting fluorescent marker protein, mGarnet2, for microscopy and STED nanoscopy. Chem. Commun. 53, 979–982 (2017).
Wannier, T. M. et al. Monomerization of far-red fluorescent proteins. Proc. Natl Acad. Sci. USA 115, E11294–E11301 (2018).
Yu, D. et al. A naturally monomeric infrared fluorescent protein for protein labeling in vivo. Nat. Methods 12, 763–765 (2015).
Rodriguez, E. A. et al. A far-red fluorescent protein evolved from a cyanobacterial phycobiliprotein. Nat. Methods 13, 763–769 (2016).
Shemetov, A. A., Oliinyk, O. S. & Verkhusha, V. V. How to increase brightness of near-infrared fluorescent proteins in mammalian cells. Cell Chem. Biol. 24, 758–766.e3 (2017).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. Elife 5, e12727 (2016).
Shen, Y. et al. A genetically encoded Ca2+ indicator based on circularly permutated sea anemone red fluorescent protein eqFP578. BMC Biol. 16, 9 (2018).
Inoue, M. et al. Rational engineering of XCaMPs, a multicolor GECI suite for in vivo imaging of complex brain circuit dynamics. Cell 177, 1346–1360. e24 (2019).
Wu, J. et al. Improved orange and red Ca2+ indicators and photophysical considerations for optogenetic applications. ACS Chem. Neurosci. 4, 963–972 (2013).
Qian, Y. et al. A genetically encoded near-infrared fluorescent calcium ion indicator. Nat. Methods 16, 171–174 (2019).
Qian, Y. et al. Improved genetically encoded near-infrared fluorescent calcium ion indicators for in vivo imaging. PLoS Biol. 18, e3000965 (2020).
Shemetov, A. A. et al. A near-infrared genetically encoded calcium indicator for in vivo imaging. Nat. Biotechnol. 39, 368–377 (2021).
Shcherbakova, D. M. Near-infrared and far-red genetically encoded indicators of neuronal activity. J. Neurosci. Methods 362, 109314 (2021).
Zhao, Y. et al. An expanded palette of genetically encoded Ca2+ indicators. Science 333, 1888–1891 (2011).
Carlson, H. J. & Campbell, R. E. Mutational analysis of a red fluorescent protein-based calcium ion indicator. Sensors 13, 11507–11521 (2013).
Cranfill, P. J. et al. Quantitative assessment of fluorescent proteins. Nat. Methods 13, 557–562 (2016).
Muslinkina, L. et al. Two independent routes of post-translational chemistry in fluorescent protein FusionRed. Int. J. Biol. Macromol. 155, 551–559 (2020).
Nasu, Y., Shen, Y., Kramer, L. & Campbell, R. E. Structure- and mechanism-guided design of single fluorescent protein-based biosensors. Nat. Chem. Biol. 17, 509–518 (2021).
Molina, R. S. et al. Understanding the fluorescence change in red genetically encoded calcium ion indicators. Biophys. J. 116, 1873–1886 (2019).
Inoue, M. et al. Rational design of a high-affinity, fast, red calcium indicator R-CaMP2. Nat. Methods 12, 64–70 (2015).
Kerruth, S., Coates, C., Dürst, C. D., Oertner, T. G. & Török, K. The kinetic mechanisms of fast-decay red-fluorescent genetically encoded calcium indicators. J. Biol. Chem. 294, 3934–3946 (2019).
Carlson, H. J. & Campbell, R. E. Circular permutated red fluorescent proteins and calcium ion indicators based on mCherry. Protein Eng. Des. Sel. 26, 763–772 (2013).
Akerboom, J. et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front. Mol. Neurosci. 6, 2 (2013).
Hochbaum, D. R. et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat. Methods 11, 825–833 (2014).
Arrenberg, A. B., Stainier, D. Y. R., Baier, H. & Huisken, J. Optogenetic control of cardiac function. Science 330, 971–974 (2010).
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nat. Methods 11, 338–346 (2014).
Jia, B. Z., Qi, Y., Wong-Campos, J. D., Megason, S. G. & Cohen, A. E. A bioelectrical phase transition patterns the first vertebrate heartbeats. Nature 622, 149–155 (2023).
Horton, N. G. et al. In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nat. Photonics 7, 205–209 (2013).
Drobizhev, M., Makarov, N. S., Tillo, S. E., Hughes, T. E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat. Methods 8, 393–399 (2011).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Afshar Saber, W., Gasparoli, F. M., Dirks, M. G., Gunn-Moore, F. J. & Antkowiak, M. All-optical assay to study biological neural networks. Front. Neurosci. 12, 451 (2018).
Farhi, S. L. et al. Wide-area all-optical neurophysiology in acute brain slices. J. Neurosci. 39, 4889–4908 (2019).
Dana, H. et al. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat. Methods 16, 649–657 (2019).
Berridge, M. J., Lipp, P. & Bootman, M. D. The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 1, 11–21 (2000).
Grienberger, C. & Konnerth, A. Imaging calcium in neurons. Neuron 73, 862–885 (2012).
Zhang, Y. et al. Fast and sensitive GCaMP calcium indicators for imaging neural populations. Nature 615, 884–891 (2023).
St-Pierre, F. et al. High-fidelity optical reporting of neuronal electrical activity with an ultrafast fluorescent voltage sensor. Nat. Neurosci. 17, 884–889 (2014).
Ai, H.-W., Baird, M. A., Shen, Y., Davidson, M. W. & Campbell, R. E. Engineering and characterizing monomeric fluorescent proteins for live-cell imaging applications. Nat. Protoc. 9, 910–928 (2014).
Gross, L. A., Baird, G. S., Hoffman, R. C., Baldridge, K. K. & Tsien, R. Y. The structure of the chromophore within DsRed, a red fluorescent protein from coral. Proc. Natl Acad. Sci. USA 97, 11990–11995 (2000).
Tsien, R. & Pozzan, T. Measurement of cytosolic free Ca2+ with quin2. Methods Enzymol. 172, 230–262 (1989).
Drobizhev, M., Molina, R. & Hughes, T. Characterizing the Two-photon Absorption Properties of Fluorescent Molecules in the 680-1300 nm Spectral Range. BIO-PROTOCOL 10, e3498 (2020).
Nault, F. & De Koninck, P. Dissociated Hippocampal Cultures. in Springer Protocols Handbooks 137–159 (Humana Press, Totowa, NJ, 2009).
Wiesner, T. et al. Activity-dependent remodeling of synaptic protein organization revealed by high throughput analysis of STED nanoscopy images. Front. Neural Circuits 14, 57 (2020).
Lim, S. T., Antonucci, D. E., Scannevin, R. H. & Trimmer, J. S. A novel targeting signal for proximal clustering of the Kv2.1 K+ channel in hippocampal neurons. Neuron 25, 385–397 (2000).
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE 6, e18556 (2011).
Zhou, Y. et al. Latent TGF-β binding protein 3 identifies a second heart field in zebrafish. Nature 474, 645–648 (2011).
Xiong, F. et al. Specified Neural Progenitors Sort to Form Sharp Domains after Noisy Shh Signaling. Cell 153, 550–561 (2013).
Swinburne, I. A., Mosaliganti, K. R., Green, A. A. & Megason, S. G. Improved long-term imaging of embryos with genetically encoded α-bungarotoxin. PLoS ONE 10, e0134005 (2015).
Sehnert, A. J. et al. Cardiac troponin T is essential in sarcomere assembly and cardiac contractility. Nat. Genet. 31, 106–110 (2002).
Acknowledgements
We thank Saaya Hario, Dr. Fern Sha, Rosario Valenti, the Janelia Research Campus cell culture facility, and the University of Alberta Molecular Biology Services Unit, for technical assistance. Ronak Patel is a member of the Janelia Research Campus Tool Translation Team (T3); we thank the T3 for providing equipment access and technical support. We thank Dr. Timothy Wannier for sharing the mKelly sequences and the laboratory of Caroline and Geoffrey Burns (Boston Children’s Hospital) for generously providing the TgBAC(-36nkx2.5:ZsYellow) transgenic zebrafish line. This work was supported by grants from the Natural Sciences and Engineering Research Council of Canada (NSERC; RGPIN-2018-04364 to R.E.C.), the Canadian Institutes of Health Research (CIHR; FS-154310 to R.E.C.), the Japan Society for the Promotion of Science (KIBAN(S) 19H05633 and KIBAN(A) 24H00489 to R.E.C.; KIBAN(B) 23K26794 to T.T.), the National Institutes of Health (NIH) BRAIN Initiative (1R01NS126102-01 to R.E.C.), and the Janelia Visiting Scientist Program (Janelia Research Campus, HHMI). We acknowledge NIH/NINDS BRAIN Initiative grant U24NS109107 (M.D.; Resource for Multiphoton Characterization of Genetically-Encoded Probes), NIH grants 1R01NS133755 (A.E.C.) and 1R01NS126043 (A.E.C.), and Chan Zuckerberg Dynamic Imaging grant 2023-321177 (A.E.C.).
Author information
Authors and Affiliations
Consortia
Contributions
Y.S. and R.E.C. conceived the project. Y.S., R.D., and J.W. performed protein engineering. R.D., A.A., Y.Z., Y.S., M.D., R.S.M., A.S.A., and R.P. performed in vitro protein characterization. R.D., K.S., A.A., J.Z., D.R., J.P.H., and The GENIE Project Team performed cultured cell imaging and analysis. Y.Q. performed the Ca2+ optopatch. B.Z.J. performed all-optical electrophysiology of embryonic zebrafish. Y.S., K.P., A.G.T., E.R.S., T.E.H., S.G.M., T.T., M.E.P., A.E.C., and R.E.C. supervised the research. Y.S., R.D., M.D., and R.E.C. wrote the paper. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Samuel Wang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dalangin, R., Jia, B.Z., Qi, Y. et al. Far-red fluorescent genetically encoded calcium ion indicators. Nat Commun 16, 3318 (2025). https://doi.org/10.1038/s41467-025-58485-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58485-z
This article is cited by
-
Active pixel power control for crosstalk-free All-optical neural interrogation
Nature Communications (2026)
-
A red fluorescent genetically encoded biosensor for in vivo imaging of extracellular l-lactate dynamics
Nature Communications (2025)