Abstract
Anchorage-independent survival of ovarian tumor cells in ascites is the initial and critical step for peritoneal metastasis. How ovarian tumor cells achieve anchorage independence remains unclear. Here we show that a noncanonical role of spermidine/spermine N1-acetyltransferase 1 (SAT1) dictates anchorage-independent cell survival and potentiates metastatic dissemination in ovarian cancer. SAT1-high cancer cells are prevalent in ascitic tumors, and high SAT1 expression in primary tumors is linked to increased peritoneal metastasis rates in ovarian cancer patients. Mechanistically, SAT1 noncanonically acetylates H3K27 domains in multiple mitosis-regulating genes, increasing their transcriptional levels and protecting disseminating cells from aberrant mitosis and mitotic cell death. Notably, the acetylation of H3K27 by SAT1 depends on the reductive carboxylation of glutamine to supply acetyl-CoA in the nucleus. SAT1 inhibition with the small-molecule inhibitor ginkgolide B attenuates the metastatic tumor burden in mouse models. We conclude that SAT1 inhibition is a promising therapeutic strategy for metastatic ovarian cancer.
Similar content being viewed by others
Introduction
Ovarian cancer is the eighth most common cancer in women and is responsible for more than 200,000 deaths worldwide each recent year1. Epithelial ovarian cancer (EOC), the most common type of ovarian cancer, often spreads into the peritoneum, resulting in extensive peritoneal deposits and malignant ascites at the time of diagnosis2,3. The current standard therapy for EOC is debulking surgery combined with adjuvant platinum-based chemotherapy4,5. A subset of patients with homologous recombination deficiency (HRD) or who respond to first-line platinum-based chemotherapy may benefit from maintenance therapy with poly (ADP‒ribose) polymerase (PARP) inhibitors6,7,8. However, both chemoresistance9,10,11 and resistance to PARP inhibitors12,13,14 are commonly observed in EOC. Effective therapies for patients with EOC represent an urgent unmet clinical need, and insights into pathogenesis are needed to identify potential therapeutic opportunities.
Malignant ascites with disseminating tumor cells is frequently associated with the metastasis and recurrence of ovarian cancer15,16,17,18. When deprived of attachment to the extracellular matrix (ECM), cells normally undergo a form of programmed cell death called anoikis19. However, ovarian cancer cells in ascites survive detachment from the ECM and disseminate extensively in the peritoneal cavity independent of anchorage. Anchorage-independent cell survival may be particularly critical for peritoneal metastasis20. Previous studies have shown that the anchorage independence of cell survival is associated with metabolic reprogramming21,22. Notably, metabolic support for cancer cells in terms of proliferation vs. dissemination could be strikingly different23. However, the upstream signals and downstream targets of metabolic reprogramming that enable anchorage independence and peritoneal metastasis are largely unknown.
In this work, given the above findings, we initially seek to investigate the role of metabolic reprogramming, which plays a key role in anchorage-independent cell survival and peritoneal metastasis in EOC. Strikingly, we uncovered a non-canonical role for SAT1, the rate-limiting enzyme in polyamine catabolism, in regulating H3K27ac epigenetic marks in genes critical for mitotic control and chromosome segregation. This mechanism promotes anchorage-independent survival and drives peritoneal dissemination of the ovarian cancer cells.
Results
SAT1 promoted anchorage-independent cell survival and peritoneal metastasis
To gain insight into the metabolic alterations that contribute to peritoneal metastasis, we used a lentiviral short hairpin RNA (shRNA) library to individually knock down 111 rate-limiting enzymes24 in the human ovarian cancer cell line SKOV3 (Table S1). After stable viral transduction, SKOV3 cells were detached from monolayers and automatically aggregated into anchorage-independent cell clusters. Among these enzymes, SAT1 emerged as a lead hit, and its knockdown strongly reduced the survival of detached tumor cells (Fig. 1a, Table S2). SAT1 is a rate-limiting enzyme in the catabolic pathway of polyamine metabolism that catalyzes the acetylation of spermidine and spermine. Cell apoptosis assays further confirmed that SAT1 knockdown significantly exacerbated the death of detached cells and that rescuing the expression of shRNA-resistant SAT1 fully restored the viability of SAT1-depleted cells, ruling out the possibility that this was an off-target effect of the SAT1 shRNA (Figs. 1b, S1a, b). Conversely, forced overexpression of SAT1 reduced apoptosis and increased the viability of detached cells (Fig. S1c–e). These data indicate that SAT1 supports anchorage-independent cell survival.
a SKOV3 were transduced with individual shRNAs that targeted each of the 111 rate-limiting metabolic enzymes. The transduced cells were detached 24 h before cell viability measurement. Data are presented as a volcano plot of three biologically independent experiments (two-tailed Student’s t test). shRNAs with P < 0.05 and a relative cell viability level (normalized to the control pLKO) of <0.8 or >1.2 were biologically meaningful. The top hit genes are labeled. b Apoptosis of detached SAT1-knockdown cells rescued with the control vector or the SAT1 recombinant vector rSAT1. c Viability of attached or detached cells (detachment 48 h) with or without SAT1 knockdown. d SAT1 expression detected by western blot in attached or detached cells. e Gene set enrichment analysis (GSEA) of RNA-seq data showing enrichment of KEGG hallmark hypoxia in detached cells relative to attached cells; GEO accession: GSE267614. f Representative microscopy images of detached ovarian cancer cells forming clusters and Image-iT green hypoxia reagent staining. Scale bars represent 20 μm. g Binding of HIF-1α to the SAT1 promoter in detached cells determined by a ChIP assay followed by qPCR. h–l Effect of SAT1 knockdown on the ID8 peritoneal dissemination model. h Experimental design (upper panel) and confirmation of SAT1 knockdown by western blot (lower panel). i–j, Average photon flux (i) and representative bioluminescence image (j) of tumors at the indicated timepoints. k, l Representative images (k) and quantification (l) of hemorrhagic ascites and peritoneal tumor nodules in mice at the endpoint. Data shown are the means ± SDs from 3 biological replicates for (b), (c), and (g) and are representative of 3 independent biological experiments for (d) and (f). The error bars represent the SDs of the tumor luminescence signals (i). Statistical analysis was performed via 1-way ANOVA for (b), (c) and (g); 2-way ANOVA for (i); and two-tailed Student’s t test for (l). The samples derive from the same experiment but different gels for SAT1 and ACTB were processed in parallel for (d) and (h). See also supplementary Figs. S1–S3.
Interestingly, the effect of SAT1 depletion on cell viability was observed only in the detached cells but not in the attached monolayer cells (Fig. 1c). This might be because SAT1 expression was almost negligible in the attached cells but was strongly induced upon detachment (Fig. 1d). We compared detached vs. attached cells via RNA-seq and identified hypoxia as one of the most enriched pathways using gene set enrichment analysis (GSEA) (Fig. 1e, Tables S3–S4). Cells normally form clusters upon detachment. Consistent with the results of the RNA-seq analysis, strong hypoxia was detected in the detached cells, especially in the cells inside the cell cluster using hypoxia probe (Fig. 1f). A ChIP assay using an anti-HIF-1α antibody revealed enrichment of HIF-1α at the SAT1 promoter, which contains the typical hypoxia response element (HRE) (Fig. 1g). Knocking down HIF-1α abrogated the expression of SAT1 under detached conditions (Fig. S1f). A luciferase reporter assay further revealed that detachment failed to induce the enrichment of HIF-1α at the SAT1 promoter with the mutated HRE sequence, which lost its binding affinity with HIF-1α (Fig. S1g). These data may explain why SAT1 expression was strongly promoted by cell clustering upon detachment.
Peritoneal metastasis of ovarian cancer requires anchorage independence, which probably requires SAT1 to support the survival of disseminating tumor cells. We tested this hypothesis in an ID8 peritoneal dissemination mouse model. As shown in Fig. 1h, mouse ovarian cancer cell line ID8 was injected intraperitoneally into mice. A parallel experiment was also performed to confirm that SAT1 expression was markedly induced in peritoneal ID8 cells at the early stage after injection (Fig. 1h). Importantly, Sat1 knockdown strongly suppressed tumor cells at the early stage after inoculation and resulted in very low metastatic peritoneal colonization, as shown by bioluminescence tumor imaging (Figs. 1i–j, S2a–c). Consistently, forced SAT1 overexpression supported tumor cells at the early stage after inoculation and promoted overall tumor progression (Fig. S2d–f). Moreover, mice injected intraperitoneally with ID8 cells expressing Sat1 shRNA ultimately developed significantly fewer hemorrhagic ascites and fewer peritoneal tumor nodules than did mice injected with cells expressing a nontargeting shRNA (Fig. 1k–l), indicating that SAT1 knockdown decreased tumor progression. Previous studies have shown that T cells play a dominant antitumor role in ovarian cancer25,26, and it is possible that SAT1 knockdown decreases tumor progression via T cells. However, we observed that SAT1 knockdown similarly suppressed tumor progression in the BALB/c-nu T-cell deficient mouse model, as it did in the immune-component C57BL/6 mouse model (Fig. S3a–c), implying that at least T cells from the immune system are not necessary for the decreased tumor growth potential upon SAT1 knockdown. Moreover, we inoculated control or shSat1-expressing ID8 cells subcutaneously into C57BL/6 mice. Strikingly, the difference in tumor growth potential between the control and SAT1-knockdown groups was almost negligible (Fig. S3d–f), suggesting that cell detachment may account for the decreased tumor growth potential of cancer cells with SAT1 knockdown. These data collectively demonstrate that SAT1 plays a critical role in supporting anchorage-independent cell survival and peritoneal metastasis in ovarian cancer.
SAT1 expression is correlated with peritoneal metastasis in ovarian cancer
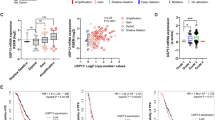
To investigate the potential clinical relevance of SAT1 in the peritoneal metastasis of ovarian cancer, we first examined the protein expression level of SAT1 in a large panel of ovarian cancer cells isolated from the matched primary tumor tissues and malignant ascites of 54 patients with ovarian cancer (Table S5). SAT1 protein levels were analyzed via western blotting followed by ImageJ quantification. The analysis revealed that in 66.7% (36 out of 54) of patients, SAT1 expression was significantly elevated in metastasizing tumor cells isolated from ascites compared with that in primary tumor cells isolated from the matched primary tissue (Fig. 2a). Furthermore. IHC staining of samples from patients with ovarian cancer revealed that SAT1-high cancer cells were highly enriched in the ascitic tumor cluster, whereas both SAT1-high and SAT1-low cancer cells were present in the matched primary tumor tissue (Fig. 2b). These data indicate a close link between SAT1 expression and peritoneal metastasis in the clinic.
a Western blot analyses of SAT1 levels in ovarian tumor cells from primary tumor tissue and matched ascites from 54 patients. ImageJ software was used to compare the relative level of SAT1 in ascites and primary tissues. b Immunohistochemistry analyses of SAT1 expression in primary tumors and matched ascitic tumor clusters from 4 representative patients with ovarian cancer. Scale bars represent 100 μm. c Immunohistochemistry analyses of SAT1 expression in primary tumors from the ovarian cancer tissue array (n = 128). The patient tissues were classified into three groups on the basis of SAT1 expression evaluated by a pathologist (high: homogeneous high SAT1 staining; heterogeneous: high/low SAT1 staining; and low: homogeneous low SAT1 staining). Scale bars represent 50 μm. d–g Primary tumor size (d), peritoneal metastatic status (e), progression-free survival (f), and overall survival (g) in patients in the high/heterogeneous SAT1-expressing group vs the low SAT1-expressing group. The data shown are representative of 2 independent biological experiments for (a) and (b). The statistical analysis was performed via two-tailed Student’s t test (d). The samples derive from the same experiment but different gels for SAT1 and ACTB were processed in parallel for (a). See also supplementary Fig. S4.
Previous studies have demonstrated that cancer cells require elevated intracellular polyamine levels to sustain their proliferative capacity27. In theory, low SAT1 expression reduces the catabolic flux of polyamines, thereby supporting the proliferation of cancer cells in the primary tumor. Nevertheless, our findings indicate that high SAT1 expression is crucial for cell survival upon detachment. Consequently, the presence of SAT1-high cells in primary tumor tissue likely indicates metastasis. To test this hypothesis, we conducted a study in a second cohort of patients with ovarian cancer. A commercial cancer tissue array containing primary tumor tissue from 128 ovarian cancer patients was analyzed via immunohistochemistry (IHC). A pathologist evaluated the intensity of SAT1 in the tumor tissue array (Table S6). A total of 65.6% (84 out of 128) of the tumors presented high or heterogeneous high SAT1 expression, whereas 34.3% presented low SAT1 expression (44 out of 128) (Fig. 2c). The volume of primary tumors in the low SAT1-expressing group was not significantly different from that in the high/heterogeneous SAT1-expressing group (Fig. 2d). However, patients with high/heterogeneous SAT1-expressing tumors presented a greater incidence of peritoneal metastases, with 75% (63 out of 84) of patients affected, than 52% (23 out of 44) of patients with low SAT1-expressing tumors (Fig. 2e). Metastasis is the main cause of cancer-related death28. We also observed that patients with high/heterogeneous SAT1-expressing tumors had significantly shorter progression-free survival and overall survival times than did those with low SAT1-expressing tumors (Fig. 2f–g). Moreover, the clinical data also suggest that high SAT1 expression might correlate with serous histotypes the most. Epithelial ovarian cancer mainly includes four histotypes, namely, serous, endometrioid, clear cell and mucinous, with serous cases being the most common. There are a total of 128 samples in the commercial cancer tissue array, with the serous subtype being the most common, comprising more than half, followed by the mucinous and endometrioid subtypes, with the remaining types being relatively rare (Fig. S4a). Within the serous subtype, the proportion of samples with high/heterogeneous expression of SAT1 was very high (79%), much greater than that in the mucinous (41%) and endometrioid subtypes (Fig. S4b). Indeed, chi-square analysis revealed that the SAT1 level is not independent of histotype (Fig. S4c). However, when the serous subtype was excluded from the analysis, the SAT1 level was independent of the histotype (Fig. S4d). This analysis may align with clinical observations. Serous ovarian cancer, mostly high-grade serous cancer, is characterized by early onset of peritoneal dissemination via the formation of clusters in the peritoneal cavity. Specifically, within the serous type, we found that in the group with high SAT1 expression, 41 patients had peritoneal metastasis compared to 12 patients without, resulting in a metastasis rate of 77.35%. In contrast, among those with low SAT1 expression, the numbers were 7 patients with and 7 patients without peritoneal metastasis, yielding a metastasis rate of 50%. A chi-square analysis confirmed the significance of the association between SAT1 expression levels and the presence of metastasis in the serous type (Fig. S4e–f). Collectively, the data suggest that high or heterogeneous SAT1 expression is associated with peritoneal metastasis in ovarian cancer, with a notable relevance to the serous subtype of ovarian cancer.
SAT1 protected cells from mitotic catastrophe upon detachment
Next, we investigated how SAT1 expression promotes cell survival upon detachment and potentiates dissemination. To elucidate the underlying mechanism, we initially carried out RNA-seq analysis of detached ovarian cancer cells with or without SAT1 knockdown and identified 1,829 DEGs (P < 0.05) in the detached SAT1-knockdown cells (Table S7). Gene Ontology (GO) term enrichment analysis revealed that the DEGs associated extensively with the mitotic process, most likely reflecting a role of SAT1 in cell cycle control (Fig. 3a). Previous studies have also shown that cancer cells within anchorage-independent cell clusters typically proliferate at a slower rate in a slow cycling state22. We applied gene-set enrichment analysis (GSEA) to the RNA-seq data and found a top enrichment for gene sets that are hallmarks of the G2/M cell cycle checkpoint and mitotic spindle in SAT1-proficient cells compared with SAT1-depleted cells, suggesting tight control of entry into mitosis and chromosome segregation by SAT1 in ovarian cancer cells upon detachment (Fig. 3b). Compared with normally attached proliferating cells, loss of anchorage causes metabolic imbalances, including hypoxia, mitochondrial perturbations, and enormous ROS production29,30, thus possibly resulting in high levels of replication stress and requiring tight control of entry into mitosis and chromosome segregation to maintain genomic integrity and prevent mitotic catastrophe, a form of death during mitosis, in detached cells31,32. Indeed, through monitoring the cell cycle distribution via the click chemistry-based EdU incorporation assay, we observed that SAT1 knockdown caused premature entry into mitosis, leading to a marked increase in the number of cells with incompletely synthesized DNA, a deleterious form of DNA damage (1 C < DNA content <2 C/EdU–) (Fig. 3c, Fig. S4g). Further examination of the cell population with incompletely synthesized DNA revealed that it expressed a substantially greater amount of γH2AX, indicating that the cells had severe DNA damage (Fig. 3d). By contrast, either shRNA-mediated knockdown or forced overexpression of SAT1 in attached ovarian cancer cells did not induce DNA damage (Fig. S5a, b). Imaging of the detached cells via a neutral comet assay revealed that the tail moments of the SAT1-depleted cells increased dramatically, suggesting that extensive DNA damage occurred upon SAT1 knockdown (Fig. 3e). DNA damage persisting in mitosis could jeopardize chromosome segregation and cause mitotic arrest, leading to mitotic catastrophe. As measured by metaphase spreads, chromosomal abnormalities dramatically increased in detached cells with SAT1 knockdown (Fig. 3f). Tracking the SKVO3 cell line stably expressing mCherry-tubulin and GFP-histone H2B via time-lapse microscopy revealed that SAT1 depletion strongly prolonged the duration of mitosis and substantially induced more mitotic cell death (Fig. 3g, h). Taken together, these data demonstrate that SAT1 protects detached cancer cells from mitotic catastrophe by regulating entry into mitosis and chromosome segregation.
a Dot plot pathway enrichment map showing most significantly over-represented pathways from differentially expressed genes (DEG) in detached cells upon SAT1 knockdown via clusterProfiler. Adjusted P values are shown. b Gene set enrichment analysis (GSEA) showing enrichment of the KEGG hallmark G2M checkpoint and mitotic spindle in pLKO cells relative to SAT1-depleted cells upon detachment. GEO accession for (a-b): GSE267614. c Flow cytometric analysis of DNA synthesis (EdU) and DNA content (Hoechst 33342) at the indicated time points. The percentages of cells with 1 C < DNA < 2 C/EdU− were quantified (n = 3, each). d γH2AX levels in cell lysates from the indicated populations of SAT1-knockdown cells detached for 12 h (lower panel). The cell populations were sorted by flow cytometry based on cell cycle distribution (upper panel). e Comet assay performed in detached cells with or without SAT1 knockdown. For each of five biologically independent experiments, approximately 100 individual cells from 3 random fields were scored for the proportion of DNA in the COMET “tail” (n = 5, each). f Metaphase spread assay of chromosomal aberrations in detached cells with or without SAT1 knockdown. g Cells expressing mCherry-tubulin (red) and GFP-H2B (green) were detached and analyzed via time-lapse imaging to monitor mitosis. Representative images of cells at anaphase following nuclear envelope breakdown (NEBD) are shown (right panel). Quantification of the time from NEBD to anaphase (left panel). The cells were maintained in a low-oxygen chamber for SAT1 induction. h Donut plots showing the proportion of cell survival (white) and cell death in mitosis (red) or interphase (pink) following mitotic arrest. Scale bars represent 100 μm for (e), 2 μm for (f), and 10 μm for (g). The data are representative of 2 independent biological experiments for (d) and are presented as the means ± SDs from biological replicates for (c) and (e). Statistical analysis was performed via one-way ANOVA for (c) and via two-tailed Student’s t tests for (e) and (g). The samples derive from the same experiment but different gels for γH2AX and ACTB were processed in parallel for (d).
SAT1 depletion reduced H3K27ac in mitosis-regulating genes
Given the profound alterations in the transcription levels of multiple mitosis-regulating genes in response to SAT1 depletion, we wondered whether SAT1 functions epigenetically in detached ovarian cancer cells. Analysis of various histone modifications in SAT1-depleted ovarian cancer cells upon detachment revealed a modest reduction in the level of H3K27ac (Fig. 4a). H3K27ac is a robust marker of active promoters and distal enhancers that are tightly coupled to transcription factor binding and gene expression33. Consequently, we conducted chromatin immunoprecipitation and sequencing (ChIP-seq) analysis of H3K27ac under detachment conditions and mapped gene regions with significantly altered H3K27ac. The analysis revealed 1870 genes with increased H3K27ac peaks and 1556 genes with decreased H3K27ac peaks in SAT1-deficient ovarian cancer cells compared with control cells. Profiling of the ChIP peaks revealed that H3K27ac is located mainly in the promoter region of genes (Fig. 4b, c, Table S8). The integrated analysis of the RNA-seq and H3K27ac ChIP-seq datasets revealed that 11.05% (93) of the genes with reduced H3K27ac peaks were transcriptionally downregulated (Fig. 4d, Table S9). Notably, these genes were broadly associated with the mitotic cell cycle process, chromosome organization, and chromosome segregation, including CCNB1, BUB1B, FANCD2, CENPA and TOP2A, (Fig. 4e, f). We also validated these findings via ChIP‒qPCR and found that H3K27ac enrichment was significantly reduced in the promoter regions of the representative genes associated with mitosis regulation and chromosome segregation in multiple cell lines (Figs. 4g, S5c). Consistently, the mRNA levels of these genes also decreased with SAT1 knockdown, as detected by RT‒qPCR (Fig. S5d). Moreover, forced overexpression of SAT1 increased H3K27ac enrichment in promoter regions and the transcription levels of these representative genes (Fig. S5e, f). To further demonstrate that SAT1 safeguards cells against mitotic catastrophe through these mitotic-related genes, we selected five representative genes—namely, CCNB1, BUB1B, FANCD2, CENPA, and TOP2A—for co-overexpression and performed a rescue experiment upon SAT1 knockdown in detached cells (Fig. S6a). Overexpressing these genes resulted in a significant rescue effect, markedly decreasing cell apoptosis and the population of cells with incomplete DNA synthesis in SAT1-knockdown cells under detachment (Fig. S6b, c). By performing a comet assay and monitoring the mitotic process, we further confirmed that rescuing the expression of the five representative genes markedly decreased DNA damage and reduced the prolonged duration of mitosis in SAT1-depleted detached cells (Fig. S6d, e).
a Western blot analysis of common histone modifications in cells with or without SAT1 knockdown under detachment for 12 h. Histone H3 or histone H4 was used as a loading control. b, c Average profile of H3K27ac ChIP peaks binding to the TSS region (b) and annotation of the location of H3K27ac peaks in terms of genomic features (c) in detached cells with or without SAT1 knockdown via ChIPseeker tools (n = 3, each). GEO accession number of the ChIP-Seq data is GSE267703. d Venn diagram showing the overlap between genes with reduced expression and genes with reduced H3K27ac deposition in detached SAT1-depleted cells compared with pLKO control cells (n = 3, each). e Analysis of the top GO terms associated with the biological processes associated with the overlapping differentially expressed genes using clusterProfiler tool. Adjusted P values are shown. f Cnetplot visualizing the genes that are involved in the significantly enriched biological process terms using clusterProfiler tools. Adjusted P values are shown. g ChIP‒qPCR analysis of H3K27ac enrichment in several key genes identified in (f) and related to the mitotic cell cycle process, chromosome organization, and chromosome segregation in detached cells with or without SAT1 knockdown. The data are representative of 2 independent biological experiments for (a) and are presented as the means ± SDs from 3 biological replicates for (g). Statistical analysis was performed via two-tailed Student’s t test for (G). The samples derive from the same experiment but different gels for all indicated histone antibodies were processed in parallel for (a). See also supplementary Figs. S5 and S6.
These findings imply that SAT1 might be important for H3K27ac enrichment in the promoter region and that the transcriptional activation of these genes is required for mitosis regulation and chromosome segregation. Importantly, however, on the basis of current data, we cannot rule out the possibility that SAT1 directly activates sequence-specific transcription factors, thereby increasing the expression of their mitosis-related target genes.
SAT1 directly acetylated H3K27 to regulate H3K27ac
We next explored how SAT1 expression is associated with H3K27ac in detached ovarian cancer cells. Previous studies have reported that alterations in cellular polyamine concentrations affect histone modifications34,35. Therefore, we first monitored the total polyamine concentration in detached ovarian cancer cells. Surprisingly, we found that SAT1 knockdown did not significantly affect the total cellular polyamine concentration, as measured by a fluorometric assay (Figs. 5a, S7a). Similarly, SAT1 overexpression did not significantly alter the cellular polyamine level (Fig. S7a). We further performed polyamine-targeted metabolomics and analyzed individual polyamines, including putrescine (Put), spermidine (Spd) and spermine (Spm), as well as acetylspermidine (Ac-spd) and acetylspermine (Ac-spm), in detached cells via quantitative LC‒MS/MS. We found that while SAT1 knockdown decreased the levels of Ac-spd and Ac-spm in detached cells, the levels of three major polyamines (Put, Spd and Spm) remained largely unchanged with SAT1 knockdown (Fig. S7c). More importantly, the level of the polyamine pool was largely controlled by Put, Spd, and Spm. The total level of these three major polyamines (approximately 0.31 μg/mg) was approximately 6000-fold greater than that of Ac-spd and Ac-spm (approximately 0.05 ng/mg) in the detached cells, as calculated by quantitative MS analysis (Fig. S7c). Therefore, the effect of SAT1 knockdown on cell cycle control may not result from changes in polyamine levels. Since the polyamine level may be affected by a balance between anabolism and catabolism of polyamine and direct uptake from the environment, it is possible that SAT1 alone cannot control the polyamine level. We found that the efficiency of polyamine uptake remained unchanged regardless of SAT1 expression (Fig. 5b). However, the conversion of arginine into polyamine was impaired upon SAT1 knockdown in detached cells. Examination of the RNA-seq data revealed that the expression of enzymes in the polyamine metabolic pathway was altered in detached cells with SAT1 knockdown (Fig. 5c). In the two parallel pathways catalyzing arginine-to-polyamine conversion, agmatinase (AGMAT) was transcriptionally downregulated when SAT1 was depleted, whereas arginase 1 (ARG1) was not detected in detached ovarian cells (Fig. 5c). Moreover, the level of spermine synthase (SMS), the enzyme responsible for the conversion of spermidine to spermine and overall polyamine production flux, also decreased in detached cells with SAT1 knockdown (Fig. 5c). We also observed that the levels of some arginine transporters, including SLC7A3, SLC7A7 and SLC7A6, as well as the level of cellular arginine uptake, were decreased in SAT1-knockdown cells upon detachment (Fig. 5c, d). Taken together, these data suggest that, probably due to the downregulation of arginine uptake and arginine‒polyamine conversion, the intracellular polyamine pool was still relatively stable in SAT1-depleted detached cells. Therefore, H3K27ac alteration and dysregulation of the cell cycle induced by SAT1 knockdown may not be caused by cellular polyamines in detached cells.
a Total polyamine content in the indicated detached cells (n = 5, each). b Relative polyamine uptake into the indicated detached cells determined via 3H-putrescine labeling (n = 5, each). c Schematic representation of polyamine metabolism. Heatmap showing the changes in the mRNA levels of the enzymes in detached SAT1-depleted cells compared with control pLKO cells via RNA-seq analysis (n = 3, each). GEO accession: GSE267614. d Relative 3H-arginine uptake into detached pLKO and SAT1-knockdown cells (n = 5, each). e Cellular location of SAT1 in detached cells analyzed via confocal microscopy. Nuclei were counterstained with DAPI. Scale bars represent 30 μm. f Analysis of interaction between SAT1 and H3 in detached cells. g Molecular docking of H3 with SAT1. SAT1 and H3 proteins are shown as cartoons and colored silver and cyan, respectively. Residues mediating the interaction of H3 and SAT1 are highlighted and shown in stick representation. h Analysis of H3 acetylation in the in vitro acetylation assay using 3H-ac-CoA. Recombinant His-H3 protein was incubated with recombinant flag-SAT1 WT or flag-SAT1 K166A mutant protein in the presence of ac-CoA (n = 10, each). i Analysis of enzyme activity of recombinant SAT1 WT or SAT1 K166A mutant protein using spermine as a substrate (n = 5, each). j Analysis of the interaction between wild-type (WT) or K166A mutant (K166A) SAT1 and His-tagged H3 in detached cells. k, l γH2AX levels (k, timepoint=18 h) and apoptosis (l, timepoint=24 h) in SAT1-depleted cells rescued with the recombinant rSAT1 WT or the rSAT1 K166A mutant upon detachment (n = 5, each). The data are presented as the means ± SDs of biological replicates for (a), (b), (d), (h), (i), (k) and (l) and are representative of two independent biological experiments for (e), (f) and (j). Statistical analysis was performed via two-tailed Student’s t test for (a), (b), and (d) and via 1-way ANOVA for (h, (i), (k) and (l). The samples derive from the same experiment but different gels for all indicated antibodies were processed in parallel for (f) and (j). See also supplementary Figs. S7–S10.
We further demonstrated this by altering the cellular polyamine level. We treated detached cells with a series of concentrations of putrescine and observed an increasing trend in the total polyamine level in the detached cells (Fig. S8a). However, we did not observe a significant change in cell apoptosis, mitotic slippage, or H3K27ac enrichment in the promoters of representative genes related to cell cycle control (Fig. S8b–d). Additionally, while SAT1 knockdown in detached ovarian cancer cells reduced Ac-spd and Ac-spm levels, supplementing these cells with Ac-spd or Ac-spm did not significantly alter total H3K27ac levels, apoptosis, or mitotic slippage relative to untreated controls (Fig. S8e–g). In addition, we also attempted to decrease the cellular concentration of polyamine by knocking down the ODC1 gene, which has been shown to play a key role in controlling the levels of the cellular polyamine pool. ODC1 knockdown significantly decreased the level of the polyamine pool (Fig. S8h). Knocking down ODC1 significantly curtailed the proliferation and induced apoptosis in ovarian cancer cells under attached conditions (Fig. S8h–j). However, no significant changes in cell apoptosis, mitotic slippage, or H3K27ac enrichment in the promoters of representative genes were detected in ODC1-knockdown cells compared with control cells under detached culture conditions (Fig. S8k–n). The findings imply that SAT1’s role in regulating the cell cycle is likely dissociated from polyamine levels in detached cells.
Next, we investigated whether SAT1 could directly affect H3K27ac in detached ovarian cancer cells. KAT3A (CBP) and KAT3B (P300) are the reported acetyltransferases responsible for H3K27ac in normal attached cells36. Interestingly, the SAT1 level increased dramatically to a much greater level than did KAT3A and KAT3B in detached cells (Fig. S9a). Similar to the inhibition or knockdown of CBP/P300, the knockdown of SAT1 decreased H3K27ac levels in detached cells (Fig. 9b, c). Also, we found that the SAT1 protein was located in the nucleus and directly interacted with H3 in detached ovarian cancer cells, as shown by immunofluorescence staining and inverse immunoprecipitation (Figs. 5e–f, S10a). Therefore, we speculate that SAT1, as an acetyltransferase, is likely to directly acetylate H3K27 in detached cells. An in vitro acetylation reaction was performed. We incubated the bead-bound His-tagged histone H3 protein with the SAT1 protein in reaction buffer containing 3H-labeled acetyl-CoA, and then washed away the SAT1 protein and 3H-labeled acetyl-CoA, and measured the acetylation level of histone H3 via a scintillation counter. We found that SAT1 strongly acetylated histone H3, and as a control, nonhistone proteins, including P53 and STAT3, were not acetylated by SAT1(Fig. S10b). Knockdown of SAT1 also did not seem to influence the acetylation status of p53 or STAT3 at typical lysine residues in the detached ovarian cancer cell lines (Fig. S10c). We further validated that H3K27 is acetylated by SAT1 using western blot with a set of specific antibodies targeting histone acetylation. (Fig. S10d). Next, we investigated the structural basis underlying the SAT1-mediated catalysis of H3K27ac. Molecular docking simulations revealed that SAT1 binds directly to the H3 protein through multiple interacting sites. More importantly, the H3K28 site (conventionally referred to as H3K27) is directly present in the active center of SAT1 and is in close proximity to the SAT1 catalytic site Y140. In addition, the positioning of the loop containing the H3K28 site in the active center is likely guided by the H3S29 site, which interacts with both the SAT1 R142 site and the SAT1 K166 site (Fig. 5g). Site mutation of SAT1 K166 to alanine, which may affect the positioning of the H3K28 site in the active center of SAT1 based on the simulation results, significantly suppressed the acetylation of the H3 protein by SAT1 in vitro (Fig. 5h). Moreover, unlike SAT1 WT, rescued expression of the SAT1 K166A mutant failed to restore the level of H3K27ac in detached cells with SAT1 knockdown (Fig. S10e, f). Notably, the SAT1 K166A mutation did not affect the acetylation of polyamine by SAT1, possibly because the SAT1 K166 site is more specific for H3 protein binding and not among the sites mediating polyamine binding (Fig. 5i). Indeed, immunoprecipitation assays revealed that the SAT1 K166A mutation significantly decreased the binding affinity of the SAT1 and H3 proteins (Fig. 5j). Consistently, the level of the polyamine pool was similar in the SAT1-depleted detached cells, with rescued expression of either the SAT1 WT or the K166A mutant (Fig. S10g). In contrast, restored levels of γH2AX (Fig. 5k) and cell apoptosis (Fig. 5l) were observed with rescued expression of WT but not K166A mutant SAT1 in SAT1-depleted detached cells, highlighting the critical role of direct acetylation of H3K27 by SAT1 in regulating the survival of detached cells. Taken together, these data suggest that SAT1 may directly acetylate H3K27 to regulate H3K27ac rather than through polyamine catabolism in detached cancer cells.
SAT1 acetylation of H3K27 depended on reductive carboxylation of glutamine
Protein acetyltransferases are highly sensitive to the concentrations of their cofactors and the central metabolic intermediate, ac-CoA. Consequently, the activity of protein acetyltransferase is inextricably linked to the metabolic status of the cell36. Therefore, SAT1-mediated H3K27 acetylation could be tightly controlled by metabolic alterations that regulate ac-CoA availability in detached cells. ac-CoA is at the crossroads of cellular metabolism. Glucose and glutamine, the two nutrients most consumed by cancer cells, could be the major sources of the ac-CoA pool in cells. To investigate metabolic changes that might predominantly regulate the ac-CoA pool in detached cells, we first investigated the metabolic state of detached cells by measuring the extracellular acidification rate (ECAR) and oxygen consumption rate (OCR). We found that glycolysis markedly increased in ovarian cancer cells upon detachment (Fig. 6a), which coincided with reduced oxygen consumption rates in detached cells (Fig. 6b), indicating that detached cells are glycolytic, with inefficient respiration and oxidation. On the basis of these observations, we further examined glucose metabolism by tracing 13C6-glucose carbons into metabolites, including pyruvate, lactate, citrate, and ac-CoA. Consistently, we detected strikingly increased labeling of pyruvate and lactate and reduced labeling of citrate and ac-CoA from 13C6-glucose in the detached cells (Fig. 6c, d), indicating that the detached ovarian cancer cells have a more glycolytic phenotype with much less ac-CoA derived from glucose.
a, b Extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) profiles of the indicated cells. Data were normalized to the protein content. c, d Enrichment levels of isotope labeling of pyruvate (Pyr), lactate (Lac), citrate (Cit), and ac-CoA in indicated cells cultured in the presence of 13C6-glucose for 4 h. Peak areas are normalized to cell count; unlabeled: M + 0, labeled: M + 1 to M + n. e Schematic representation showing that 13C5-glutamine is either oxidized the TCA cycle, producing M4 or M2 carbons in the second round (green dots), or undergoes reductive carboxylation to M5 citrate, producing M3 oxaloacetate (Oac), malate (Mal), aspartate (Asp), and fumarate (Fum) (red dots). f–g Enrichment levels of isotope labeling of glutamine metabolic intermediates in indicated cells cultured in the presence of 13C5-glutamine for 4 h. Peak areas are normalized to cell count; unlabeled: M + 0; labeled: M + 1 to M + n. h, i Isotopologues distribution of citrate in indicated cells cultured in the presence of 13C5-glutamine for 4 h. j Cytosolic and nuclear localization of ACLY and SAT1 in indicated cells. LSD1 and ACTB: markers for nucleus and cytosol, respectively. C: cytosol, N: nucleus. k Cytosolic and nuclear levels of ACLY and SAT1 in ACLY-depleted cells rescued with the control vector, rACLY-WT or rACLY-NES under detached conditions. l–n ChIP‒qPCR analysis of H3K27ac marks of indicated mitosis-regulating genes (n = 5, timepoint=12 h), flow cytometric analysis of γH2AX (n = 5, timepoint=18 h) and apoptosis (n = 5, timepoint=24 h) in ACLY-depleted cells rescued with the control vector, rACLY-WT or rACLY-NES under detached conditions with or without SAT1 knockdown. Data are the means ± SDs from 3 technical replicates for (a–g, h, i) and from 5 biological replicates for (l). (a–g, h, i) and (j, k) are representative of 2 independent biological experiments. Data are presented as the means ± SDs of biological replicates for (m, n). Statistical analysis was performed via one-way ANOVA for all the data. The samples derive from the same experiment but different gels for ACLY, SAT1, ACTB, and LSD1 were processed in parallel for (j) and (k). See also supplementary Fig. S11.
Next, we examined the production of acetyl-CoA from glutamine in detached cells. In mammalian cells, acetyl-CoA is produced from glutamine in a two-step process. First, glutamine generates citrate via reductive carboxylation or glutamine oxidation. Then, ATP-citrate lyase (ACLY) converts citrate to acetyl-CoA and oxaloacetate (Fig. 6e). We traced the 13C5-glutamine carbons into ac-CoA and identified a drastic contribution from glutamine to produce ac-CoA, as well as citrate and malate (readily interconverted with oxaloacetate), in the detached cells (Fig. 6f–g), indicating that acetyl-CoA is produced mainly from glutamine metabolism in the detached cells. Notably, we observed significant increases in M + 5 citrate (Fig. 6h–i) and M + 3 malate (Fig. S11a, b) levels in the detached cells, indicating that glutamine routed through reductive carboxylation rather than oxidation to produce citrate, which is converted to acetyl-CoA by ACLY and plays a predominant role in supplying the ac-CoA pool in the detached cells.
Acetylation of H3K27 by SAT1 occurs in the nucleus. Therefore, SAT1 should be more sensitive to nuclear ac-CoA levels. Previous studies have shown that ACLY is located in both the cytosol and nucleus37, which we also confirmed in detached ovarian cancer cells (Fig. 6j). We further tested whether the presence of ACLY in the nucleus is critical for SAT1 acetylation of H3K27. ACLY-depleted ovarian cancer cells were transfected with a GFP empty vector, wild-type (WT) ACLY, or ACLY tagged with a nuclear export signal (NES). As shown by western blotting, NES-tagged ACLY was located mainly in the cytosol, which also did not affect the expression level and nuclear location of SAT1 (Fig. 6k). However, in the ACLY-depleted detached cells, NES-tagged ACLY, in contrast to wild-type ACLY, failed to rescue H3K27ac levels of those representative mitosis-related genes we previously identified, and knocking down SAT1 further abolished the rescue effect of wild-type ACLY (Fig. 6l), indicating that nuclear ACLY is critical for the acetylation of H2K27 catalyzed by SAT1. Consistently, wild-type but not NES-tagged ACLY significantly suppressed the levels of γH2AX (Fig. 6m) and cell apoptosis (Fig. 6n) in the ACLY-depleted detached cells, which were also abrogated when SAT1 was knocked down. Together, these data indicate that the SAT1 acetylation of H3K27 relies on glutamine metabolism, which predominantly provides the co-factor ac-CoA via reductive carboxylation and nuclear ACLY.
Ginkgolide B was identified as an SAT1 inhibitor and reduced peritoneal metastasis as maintenance therapy
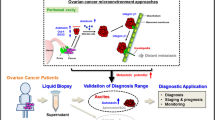
Our findings that SAT1 is markedly induced in detached cells and that attenuation of SAT1 impacts cancer cell peritoneal metastasis implicates SAT1 as a promising antimetastatic target. Regarding treatment options, the antimetastatic effect of SAT1 inhibition suggests that an SAT1 inhibitor could be a suitable candidate for maintenance therapy, especially in the adjuvant setting after debulking surgery, which removes primary tumors. Currently, the only reported SAT1 inhibitors are analogs of MGBG, including berenil and pentamidine, which are antiparasitic amidines38. However, berenil and pentamidine mainly interfere with DNA biosynthesis and indirectly target multiple biological processes, including the host immune response, RNA biosynthesis and ribosome assembly, with unknown mechanisms, raising safety concerns for clinical application in long-term maintenance therapy. In addition, these inhibitors were discovered and verified using polyamines as substrates, which may not efficiently inhibit H3K27ac via SAT1. Therefore, considering both the maintenance treatment option and the use of H3 as a substrate, we designed a series of high-throughput screens to identify H3 substrate-selective SAT1 inhibitors from a “Food as Medicine” library. In the primary screen, an in vitro SAT1 activity assay using recombinant SAT1 and H3 proteins was performed. The top 64 candidates were evaluated on the second screen. We identified a small-molecule compound, ginkgolide B (GB), as an SAT1 inhibitor from 1083 compounds in the library through two rounds of screening (Figs. 7a, S12a, b). In the subsequent DTNB-coupled SAT1 enzyme activity assay using histone H3 and ac-CoA as substrates, GB inhibited the enzyme activity of recombinant, purified active SAT1 proteins with a Ki of approximately 24.18 μM (Fig. 7b). The Lineweaver–Burk plot showed that GB acted as an uncompetitive inhibitor of SAT1 acetylation of H3. Both the Vmax and km values decreased with the addition of GB (Fig. 7c). ChIP‒qPCR and RT‒qPCR further demonstrated that H3K27ac enrichment in the promoter region and the mRNA level of the mitosis-regulating genes were decreased by ginkgolide B treatment in various cell lines under detached conditions (Fig. S13a–h). Consistent with previous observations, total cellular polyamine levels remained unchanged with ginkgolide B and/or SAT1 knockdown in the detached cells (Fig. S13i). Molecular docking simulations also revealed that GB binds to the SAT1:H3 complex at sites including the SAT1 Y163 and H3 G35 sites, supporting the uncompetitive inhibition model (Fig. 7d). More importantly, the tryptophan fluorescence binding assay revealed that mutation of SAT1 Y163 (Y163F) almost completely abolished GB binding to the SAT1:H3 complex, as the Kd value increased from approximately 20.47 μM to 175.4 μM (Fig. 7e), indicating that GB is an effective SAT1 inhibitor.
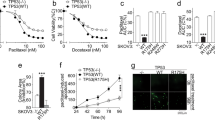
a Schematic illustration of screening to identify SAT1 inhibitor. b, c Activity of purified SAT1 in the presence of increasing concentrations of ac-CoA and ginkgolide B (b, n = 3) and the Lineweaver‒Burk plot (c). d Binding modes of ginkgolide B with the SAT1:H3 complex. SAT1 and H3 are shown as cartoons and colored silver and cyan, respectively. Ligand ginkgolide B (colored in green) and dominant residues of SAT1:H3 complex are shown as sticks. e Tryptophan fluorescence binding assay for determining Kd values of ginkgolide B binding to the indicated SAT1:H3 complex (n = 3). f–h H3K27ac (f, n = 5, timepoint = 12 h), DNA synthesis and DNA content (g), and cell apoptosis (h) in SAT1-depleted cells rescued with SAT1 WT or SAT Y163F mutant under ginkgolide B (20 μM) treatment. (i) Cell viability after treatment with increasing concentrations of ginkgolide B in cancer cells derived from primary tumor tissue (left panel) or from the matched ascites (right panel). Cells were treated under indicated conditions for 24 h. j Schematic illustration of experimental design. k, l Effect of ginkgolide B maintenance therapy on residual model tumor growth. Average photon flux counts (k) and in vivo bioluminescence image (l) at week 10 are shown. (cisplatin 5 mg/kg, Taxol 10 mg/kg, IP injection, four rounds in total), ginkgolide B (5 mg/kg, IP injection, every two days for a total of two weeks), n = 8. m Survival curves of mice (n = 8, each). n Hematology blood tests of mice. Blood was collected retro-orbitally at week 6, one week after ginkgolide B treatment (n = 5, each). o Histological analysis of hematoxylin-eosin-stained tissue sections from representative mice receiving ginkgolide B maintenance therapy. Scale bars represent 500 μm. Data are presented as the means ± SDs of biological replicates for (b, e, g, h, and n). Statistical analysis was performed via 1-way ANOVA for (g, h, and n), 2-way ANOVA for (k), and the log-rank (Mantel‒Cox) test (m). See also supplementary Figs. S12–S14.
We then examined whether the inhibitory effect of GB on H2K27ac marks of mitosis-regulating genes occurred through SAT1. In SAT1 knockdown cells reconstituted with wild-type SAT1 WT or GB binding-deficient mutant SAT1 Y163F, GB treatment attenuated H3K27ac in the representative genes only in cells expressing SAT1 WT but not in those expressing SAT1 Y163F under detachment conditions (Fig. 7f, S14a, b). Consistently, GB treatment also increased the population of incompletely synthesized DNA and increased cell apoptosis in detached cells expressing SAT1 WT but not SAT1 Y163F (Fig. 7g, h). We also examined the effect of GB treatment on the peritoneal metastasis of ovarian cancer in vivo. SAT1-depleted SKOV3 cells reconstituted with SAT1 WT or Y163F mutant expression presented similar levels of tumor progression (Fig. S14c–e). However, with GB drug treatment, significantly different degrees of tumor progression were observed. Compared with SAT1 WT, the SAT1 Y163F mutation significantly rescued tumor growth under GB drug treatment (Fig. S14c–e), indicating that SAT1 is a target of GB in the suppression of peritoneal metastasis, possibly through its interaction with the SAT1 Y163 site.
Using a panel of ovarian cancer cells derived from 54 matched pairs of primary ovarian cancer tissue and malignant ascites (Table S5), we further found that GB treatment markedly reduced the viability of ascites-derived human cancer cells upon detachment. In contrast, the viability of the matched primary tissue-derived cancer cells was not affected by GB treatment under attached conditions, emphasizing the specificity of targeting metastatic cells by GB treatment (Fig. 7i).
We further asked whether GB treatment would improve ovarian cancer treatment outcomes in vivo. Since the SAT1 protein sequence is highly conserved, with the Y163 site being the same in humans and mice, we used the ID8 mouse cell line and employed a syngeneic ovarian cancer mouse model to mimic clinical residual disease following debulking surgery and assess the therapeutic benefit of GB in the adjuvant setting as maintenance therapy (Fig. 7j). Adjuvant chemotherapy is the standard first-line treatment following debulking surgery. However, it is frequently ineffective due to the development of chemoresistance in the clinic. In this mouse model, we also observed that tumor growth was significantly attenuated under chemotherapy. However, tumors relapsed in 87.5% (7 out of 8) of the mice approximately two weeks after the full course of chemotherapy was given (Fig. 7k–l), reminiscent of the clinical scenario in which chemotherapy alone is insufficient for the cure or control of residual disease, necessitating the use of maintenance therapy after chemotherapy. Indeed, we found that GB treatment as maintenance therapy markedly suppressed tumor relapse (Fig. 7k–l) and improved the survival of mice (Fig. 7m). In addition, continued exposure to GB was not associated with cumulative toxicity, as no altered hematopoietic properties (Fig. 7n) or significant histopathological changes (Fig. 7o) were observed with the addition of GB in the treatment protocol. Taken together, these results suggest that GB is an SAT1 inhibitor with considerable therapeutic potential as a maintenance therapy.
Discussion
Ovarian cancer (OC) often presents late, with malignant ascites and extensive peritoneal metastasis. Here, we report that spermidine/spermine N1-acetyltransferase 1 (SAT1) noncanonically plays an important role in supporting anchorage-independent cell survival and peritoneal metastasis by directly acetylating H3K27 marks in mitosis-regulating genes and preventing mitotic catastrophe. Upon detachment from the ECM, cancer cells form clusters and undergo strong metabolic perturbation, characterized by hypoxia and reductive carboxylation of glutamine. SAT1 expression was strongly induced by HIF-1α upon detachment, and tumors in the ascites were enriched with SAT1-high cancer cells. Moreover, the acetylation of H3K27 by SAT1 was dependent on nuclear ac-CoA derived from the reductive carboxylation of glutamine. SAT1 serves as an integrated downstream target that links metabolic reprogramming to mitotic control and genomic integrity of ovarian cancer cells in the peritoneal metastasis (Fig. S15).
The peritoneum/omentum is the most frequent site of distant metastasis in individuals with ovarian cancer. Nonetheless, ovarian cancer can metastasize through various pathways, with other organs also commonly affected by distant metastases39,40. Consequently, there is a clear need for future research to explore the function of SAT1 in the metastatic spread of ovarian cancer cells to these alternative sites. The implication of this discovery also suggests that SAT1 expression may play a role in the transformation of benign cells within pre-cancerous lesions into malignant and metastatic cells. In theory, reduced SAT1 expression curtails the breakdown of polyamines, thereby facilitating the proliferation of benign cells within the primary tumor. However, our research reveals that elevated SAT1 expression is essential for cell viability following detachment. Thus, the detection of cells with high SAT1 expression in primary tumor tissue may serve as an indicator of potential metastatic capability.
Metabolic reprogramming is frequently observed in ovarian cancer and could be exploited for therapeutic benefit. Inhibition of the mevalonate pathway synergizes with immune checkpoint blockade (ICB) in ARID1A-inactivated clear cell ovarian carcinoma, and ARID1A mutation also creates a dependency on glutamine metabolism41,42. Recent studies have also shown that SAT1 is closely related to the metabolic status of cells. Acetate derived from cancer-associated fibroblasts promotes cancer development by epigenetically controlling SAT1 expression43, and MUC1-mediated HIF-1α stabilization promotes SAT1 expression in pancreatic cancer44, which supports our findings in detached ovarian cancer cells that SAT1 expression is strongly induced by HIF-1α. However, detachment induces more intensive metabolic perturbation, such as increased production of ROS and depletion of dNTP pools10,21, thereby causing significant DNA replication stress32. The G2/M checkpoint and subsequent mitotic checkpoint are critically important in preventing aberrant mitosis and mitotic catastrophe induced by DNA replication stress in ovarian cancer45,46. Indeed, through a combined analysis of RNA-seq and ChIP-seq data, together with a study on mitosis, we found that SAT1 depletion decreased the global H3K27ac mark within genes required for the G2/M checkpoint and mitosis regulation under detached conditions, leading to unscheduled mitotic entry and increased mitotic cell death.
Interestingly, our data suggest that SAT1 serves as an H3K27 acetyltransferase to directly regulate H3K27ac levels rather than through polyamine metabolism. We showed that the knockdown of SAT1 alone did not change the cellular polyamine level in the detached cells. SAT1 is the rate-limiting polyamine catabolic enzyme that catalyzes the acetylation of spermidine and spermine. The amounts of cellular polyamines are tightly regulated by a complex network that controls their uptake, biosynthesis, and catabolism47,48. However, SAT1 expression is often suppressed in cancer cells. N1,N11-diethylnorspermine (DENSpm, also known as BENSpm), a small molecule agonist that induces SAT1 expression, has been used in preclinical studies for cancer treatment but has very limited efficacy49. Our data suggest that the knockdown of SAT1 in detached cells was accompanied by decreased biosynthesis of polyamines from arginine; thus, SAT1 knockdown did not increase intercellular polyamine levels. While future studies on how the catabolism and biosynthesis of polyamines are coupled are warranted, these data suggest that the global change in H327ac in SAT1-depleted cells may not be due to the role of SAT1 in polyamine catabolism in detached cells.
Our study establishes SAT1 as a specific H3K27 acetyltransferase, though we cannot rule out potential acetylation activity at other unexamined histone residues. Using molecular docking, in vitro acetylation assays, and isogenic cell line pairs, we provide evidence that SAT1 directly catalyzes H3K27 acetylation in the nucleus. Structural modeling suggests that the flexible tail loop of H3 (containing K28) docks into the SAT1 catalytic center, a mechanism analogous to polyamine substrate binding. This similarity may explain SAT1’s substrate specificity for H3K27. However, the precise molecular interface remains unresolved; future structural studies resolving the SAT1/H3 cocrystal will clarify the atomic determinants of this interaction.
Our data revealed that there was only a modest overlap between genes exhibiting decreased H3K27 acetylation and those with downregulated H3K27 acetylation following SAT1 knockdown. Besides, an increase in histone acetylation does not always translate to elevated gene expression36. Notably, while SAT1 could modestly contribute to global H3K27 acetylation, its predominant functional impact lies in enriching H3K27ac marks at promoter regions of genes critical for mitosis regulation and chromosome segregation. However, the precise mechanism by which SAT1 is targeted to the promoters of certain genes to modify H3K27 acetylation remains to be elucidated. We speculated that SAT1 might have a substrate preference, selectively targeting H3K27 acetylation of genes implicated in cell cycle regulation rather than substantially affecting H3K27 global acetylation across the genome. This hypothesis could account for the observation that only a subset of mitotic genes is repressed following SAT1 knockdown, indicating that these genes are direct targets of SAT1. This could also explain why many genes with increased H3K27 levels were still present upon SAT1 knockdown. The genes with elevated H3K27 acetylation levels may be modulated by other histone acetyltransferases, such as p300/CBP. Indeed, the observation that the detection of H3K27ac upon SAT1 depletion strongly suggests that p300/CBP contributes to H3K27ac in detached ovarian cancer cells.
It is also unclear how SAT1 selectively acetylates H3K27 on certain cell cycle-regulating genes and facilitates their transcription. A plausible mechanism involves the specific recruitment of SAT1 to the promoters of these target genes through protein‒protein interactions with certain transcription factors. However, we have not yet identified a consensus sequence for SAT1 that mediates its interaction with transcription factors. The process of SAT1-dependent H3K27 acetylation is likely complex and dynamic and involves additional proteins. Our findings also highlight the importance of ACLY-supplied acetyl-CoA in SAT1-mediated H3K27 acetylation. It is also possible that ACLY could be selectively recruited to the promoters of target genes, thereby influencing SAT1 substrate preference through localized increases in acetyl-CoA levels and, consequently, increasing SAT1 activity. In addition, the present findings do not preclude the possibility that SAT1 may directly activate certain sequence-specific transcription factors, thereby potentiating the expression of mitosis-related target genes. These hypotheses certainly merit further investigation.
Our data indicate that targeting SAT1 via the small-molecule compound ginkgolide B represents a promising maintenance therapy for EOC treatment. From the perspective of the current clinical standard of care, when the initial tumor burden has been reduced to a satisfactory level by surgical debulking, controlling peritoneal metastasis is highly important for preventing deadly relapse during adjuvant or maintenance therapy. Therefore, we performed a drug screen using a “Food as Medicine” library and identified ginkgolide B (BN52021) as a SAT1 inhibitor. Ginkgolide B treatment as maintenance therapy markedly suppressed peritoneal metastasis and tumor relapse in the ID8 syngeneic ovarian cancer residual disease mouse model. Ginkgolide B is an active compound isolated from Ginkgo biloba, a well-known traditional Chinese medicine that has been used for centuries in Asian countries as a cardio- and cerebrovascular therapeutic drug and nutraceutical50. GB treatment relieves endothelial dysfunction and alleviates thrombosis by potently inhibiting platelet activation. Clinical research indicates that individuals with cancer have a 9-fold greater risk of thrombosis than does the general population51, and a significantly increased risk of venous thromboembolism has been associated with cancer surgery52, making thromboembolism a leading cause of death in people with cancer. Therefore, ginkgolide B as a maintenance therapy potentially not only inhibits the peritoneal metastasis of tumor cells but also treats systemic disease caused by cancer, which represents a promising intervention to improve clinical outcomes in patients with EOC. Given the moderate affinity of ginkgolide B for SAT1, with an approximate dissociation constant of 20 μM, further research is needed to explore the use of ginkgolide B as a prototype compound. Optimizing the structure of ginkgolide B to enhance its inhibitory potency against SAT1 could be a crucial step toward its potential clinical application.
Methods
Ethical statement
Animal studies in the study were performed according to protocols reviewed and approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University. The study involving human samples was approved by the Institutional Review Board (IRB) of Sun Yat-sen University, Zhongshan School of Medicine. The study was conducted in accordance with ethical standards and good clinical practice. All clinical samples were collected with informed consent under protocols approved by the IRB of Sun Yat-sen University. Patient clinical information was obtained from the pathology files under the guidelines and with approval from the IRB of Sun Yat-sen University.
Reagents
All key reagents used in this study are listed in Supplemental Table S12. All unique/stable reagents generated in this study are available from the lead contact with a completed Materials Transfer Agreement and are available from the lead contact with a completed Materials Transfer Agreement.
Cell culture
A2780 cells were cultured in complete RPMI-1640 medium supplemented with fetal bovine serum (10%) and penicillin/streptomycin (1%); the OVCA433 cells were cultured in complete DMEM supplemented with fetal bovine serum (10%) and penicillin/streptomycin (1%); the SKOV3 and ES2 cells were cultured in complete McCoy’s 5a medium supplemented with fetal bovine serum (10%) and penicillin/streptomycin (1%); and the 293 T cells were cultured in complete DMEM supplemented with fetal bovine serum (10%) and penicillin/streptomycin (1%). ID8 cells were cultured in complete DMEM with fetal bovine serum (10%) and insulin-transferrin-selenium-ethanolamine (1%). Primary EOC cells were cultured in complete RPMI-1640 medium supplemented with fetal bovine serum (10%), penicillin/streptomycin (1%) and GlutaMax (1%). SKOV3, ES2, and 293 T cells were purchased from the American Type Culture Collection. A2780 and ID8 cells were obtained from Sigma Aldrich. All the cell lines were authenticated via STR profiling and tested for mycoplasma contamination. To culture the cells under detached conditions, the culture plate was coated with 1% autoclaved agarose to inhibit cell attachment to the plate surface, thereby maintaining the cells in a suspended state. In addition to the use of an agarose coating, all other culture conditions, such as the serum concentration and medium type, were identical to those used for cells cultured under standard attachment conditions.
Animal studies
C57BL/6 mice (female, 6 weeks old) and nude mice (BALB/c-nu, female, 6 weeks old) were obtained from Charles River. NOD SCID gamma mice (female, 6 weeks old) were purchased from Jackson Laboratories. The mice were maintained under optimal housing conditions to ensure their well-being and to minimize stress. The housing environment included a controlled light cycle of a 14-h light/10-h dark cycle. Additionally, the temperature in the housing facility was kept within a range of 18–23 °C, and the humidity was maintained between 40–60%. For all the animal studies, the animals were randomly chosen. Concealed allocation and blinding of outcome assessment were used. No statistical method was used to predetermine the sample size. Three types of animal models were used in this study. The approved protocol allows for a maximum tumor size of less than 2000 mm3 and the tumor burden does not impede the free activity of the mice. In the ID8 peritoneal dissemination mouse model and ID8 syngeneic ovarian cancer residual disease mouse model, Mice were observed for tumor growth indicators in the peritoneal cavity, including a 20% rise in abdominal circumference, palpable tumor mass, or blood leakage upon intraperitoneal injection—criteria set as humane endpoints by the Institutional Animal Care and Use Committee of Sun Yat-sen University and adhered to throughout the study.
RNAi screens
Primary screening was performed via a human rate-limiting enzyme lentiviral short hairpin RNA (shRNA) library by transducing SKOV3 ovarian cancer cells with lentivirus pools targeting each gene individually. A detailed list of gene names and target sequences is provided in Table S1. The cells were seeded into 9 replicates and infected with lentivirus in 96-well microplates. Forty-eight hours after infection, the cells were treated with PBS or puromycin (0.5 μg/mL) or transferred into flat bottom ultralow attachment 96-well microplates and cultured under detached conditions in triplicate. After twenty-four hours of the above treatment, cell viability was determined via a CellTiter-Fluor Cell Viability Assay (Promega). Gene candidates that induced more than 20% cell death by shRNA alone, as assessed by the PBS treatment condition, and/or had poor shRNA virus transduction efficacy, as assessed by puromycin selection, were excluded from the final analysis.
Lentivirus production and infection
Briefly, lentiviruses encoding shRNA were produced by transfecting 293 T cells with a mixture of lentiviral vectors (shRNA, psPAX2, pMD2.G). Cells were seeded at 2.2 × 105 per well in 6-well plates and allowed to reach 70% confluence. Transfection involved preparing solution I (500 ng shRNA, 500 ng psPAX2, 50 ng pMD2.G in 30 µL DMEM) and solution II (4 µL TransIT-LT1 in 30 µL DMEM), incubating them for 25 min, and then adding to the cells. After 18 h, the media was refreshed and collected every 24 h, twice. The collected media were filtered (0.45 µM PES) to remove debris, and the virus was harvested, aliquoted, and stored at −80 °C.
Cell apoptosis and cell viability assay
Cell apoptosis was assessed using the FITC Annexin V apoptosis detection kit (BD Biosciences) according to the manufacturer’s instructions. Cell viability was assessed via the CellTiter-Fluor Cell Viability Assay (Promega) according to the manufacturer’s instructions.
Tumor tissue- or ascites-derived cancer cell isolation
Tumor tissue samples and ascites were collected and processed separately. Fresh tumor tissue was harvested, diced into approximately 2 mm sections and crushed with the barrel of a syringe to form a homogenate. The cell homogenate was digested with RPMI 1640 medium containing Type II collagenase (2.5 mg/mL, Thermo Fisher), 1× GlutaMAX (Thermo Fisher), 1% HEPES and 1% penicillin–streptomycin in a 37 °C water bath for 25 min, filtered through a 70 μm filter and centrifuged at 300 × g to pellet the cells. For ascites, heparin was added to the fluid. ACK lysis was performed to remove blood cells if necessary. The samples were subsequently centrifuged at 500 × g for 10 min at 4 °C to harvest the cell pellet. Cancer cells were isolated from cell pellets using the Tumor Cell Isolation Kit human (Miltenyi Biotec, Cat# 130-108-339), following the manufacturer’s protocol. This process involved magnetically labeling non-tumor cells, such as lymphocytes, fibroblasts, and endothelial cells, with a monoclonal antibody cocktail conjugated to microbeads, and then separating them from tumor cells using a magnetic separator. The purified tumor cells were either frozen in RPMI 1640 medium with 20% FBS, 1% penicillin–streptomycin, and 5% DMSO for storage in liquid nitrogen or used directly for subsequent applications.
Immunohistochemistry staining
This study focuses on the peritoneal metastasis of ovarian cancer, thereby exclusively involving female participants. Only data and biological materials from these participants are relevant and analyzed. Tumor tissue and ascites from patients with ovarian cancer were obtained from the First Affiliated Hospital, Sun Yat-sen University, and embedded in paraffin. Paraffin-embedded ovary cancer tissue microarrays (HOvaC151Su02) containing primary tumors with clinical information were obtained from Shanghai Outdo Biotech Co., Ltd. (Shanghai, China). Routine immunohistochemistry (IHC) was performed on paraffin-embedded samples. Following deparaffinization and rehydration, antigen retrieval was performed using an EDTA solution. Slides were then incubated with SAT1 antibody (1:250, 4 °C, overnight) and developed using the ImmPRESS reagent, an avidin-biotin complex system, and 3,3’-diaminobenzidine. A pathologist assessed the staining results.
Western blot
Cellular proteins were extracted using a lysis buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% NP-40, 0.5% Na-deoxycholate) supplemented with a protease inhibitor cocktail (Roche). 30 µg protein were separated on a 10% SDS-PAGE and transferred to a nitrocellulose membrane (Bio-Rad). Membranes were probed with specific primary antibodies overnight at 4 °C, washed with TBST, and then incubated with HRP-conjugated secondary antibodies for one hour at room temperature. Bands were visualized using enhanced chemiluminescence and quantified with ImageJ software.
Immunoprecipitation
To immunoprecipitate His-tagged proteins, HIS-Select Nickel Magnetic Agarose Beads (Sigma, #H9914) were used following the manufacturer’s protocol. As a negative control to assess nonspecific binding, a comparable volume of lysate from cells that did not express the His-tagged protein was used. Each immunoprecipitation was performed using 5 million cells. Cell lysis was carried out in 1% NP-40 buffer [50 mM Tris-HCl (pH 7.5), 150 mM NaCl, and 1% NP-40] supplemented with a protease inhibitor cocktail (Roche, Indianapolis, IN). The beads were equilibrated and then incubated with the cell lysate overnight at 4 °C on an orbital shaker. The bead-bound complexes were subsequently collected via a magnetic separator and washed three times. The immunoprecipitated proteins were finally eluted with 5 gel volumes of elution buffer [50 mM sodium phosphate (pH 8.0), 0.3 M sodium chloride, and 250 mM imidazole] for subsequent analysis.
Immunofluorescence
Detached cells were centrifuged at 500 × g for 5 min at 4 °C to pellet the cells. The cells were fixed with 4% paraformaldehyde, permeabilized in phosphate-buffered saline (PBS) with 0.5% Triton X-100, blocked with normal goat serum, and incubated with a primary antibody against SAT1 overnight at 4 °C. This was followed by incubation with Alexa 488-conjugated anti-rabbit antibody (1:100; A27034, Thermo Fisher Scientific) for one hour at room temperature. The nuclei were then counterstained with DAPI (1 mg/mL). The cell suspension was then loaded onto a slide and mounted with Prolong Gold antifade mountant (P10144, Thermo Fisher Scientific). Images were acquired via a Zeiss confocal microscope.
Hypoxia measurement
Hypoxia levels in the clustered detached cells were measured via an Image-iT Green Kit (Thermo Fisher, Cat# I14834) according to the manufacturer’s instructions.
Comet assay
A neutral comet assay was conducted using the Comet Assay Kit (Trevigen, Cat# 4250-050-K) as per the manufacturer’s guidelines. Cells (5 µL at 1 × 105/ml) were mixed with 50 µL of molten LMAgarose (37 °C) and pipetted onto comet slides, spreading evenly with the pipette tip. Slides were left flat in the dark at 4 °C for 30 min under high humidity. Cell lysis was performed overnight by immersing slides in lysis buffer. Post-lysis, slides were rinsed and immersed in 50 mL of 4 °C neutral electrophoresis buffer for 30 min in the dark. Electrophoresis was run at 1 V/cm for 45 min on wet ice. Slides were then immersed in DNA precipitation solution for 30 min and in 70% ethanol for 30 min at room temperature. After drying at 37 °C for 15 min, slides were stained with SYBR Gold (Thermo Fisher Scientific, S11494) for 30 min in the dark. Comet analysis was performed using a Zeiss microscope and ImageJ software.
Metaphase spreads
Detached cells with or without SAT1 knockdown were treated with 1 µg/ml colcemid (Thermo Fisher Scientific, Somerset, NJ) for 1.5 h to arrest the cells at metaphase. The cells were then suspended in 0.56% KCl for 15 min at 37 °C and fixed with methanol/acetic acid (3:1) (vol/vol) for 20 min at −20 °C. The cells were then dropped onto slides in a 50 °C water bath from a height of approximately 5–7 cm, and metaphase chromosome spreading was checked under a microscope. The slides were air dried at room temperature for two days, the slides were mounted with DAPI, and the chromosomes were detected under a Zeiss confocal microscope.
Live-cell imaging
For analysis of mitotic timing, cells stably expressing GFP-H2B and mCherry-Tubulin were cultured in an uncoated 35 mm glass bottom dish with a 50 µm grid (ibidi Company) under detached conditions. The plates were placed on a stage within an incubator maintained at 37 °C and 5% CO2. Images were acquired via a confocal inverted microscope (Zeiss LSM710) with a 40X objective lens and captured at 5 min intervals for 24 h (using Volocity software). The data were saved as OME-TIFF files and further processed in Imaris 9.5. Only single cells were located and counted. The cells were maintained in a low-oxygen chamber for SAT1 induction at the single-cell level.
RNA-seq
RNA was isolated from SKOV3 cells with or without SAT1 knockdown under attached or detached conditions via an RNeasy Mini Kit (Qiagen). Debris removal solution (Miltenyi Biotec, Cat# 130--109--398) was used for the fast and effective removal of cell debris from viable cells before the extraction of RNA and library construction. Poly-A RNA sequencing was conducted by Novogene according to the manufacturer’s instructions. Fastp (version 0.19.7) was used to perform basic statistics on the quality of the raw reads and obtain clean data. Salmon (version 1.10.2) was used for transcript quantification from the RNA-seq data. The tximport R package (version 1.32.0) was used to import the salmon results into DESeq2. The DESeq2 package (version 1.44.0) was used for the detection and analysis of DEGs. GSEA software and clusterProfiler were used for downstream enriched pathway analysis.
ChIP-seq and ChIP-qPCR
Chromatin immunoprecipitation (ChIP) was performed via an enzymatic chromatin immunoprecipitation (IP) kit (Magnetic Beads) (Cell Signaling Technology). In brief, DNA sequences that are associated with the histone modification H3K27ac coprecipitate as part of the cross-linked chromatin complex. After immunoprecipitation, the H3K27ac-DNA crosslinks were reversed, and the DNA was purified. The immunoenriched DNA samples were used for downstream DNA library construction and sequencing (ChIP-seq) or real-time quantitative PCR of the promoter sequences of the indicated genes (ChIP‒qPCR). ChIP-seq was performed by Novogene according to the manufacturer’s instructions. Fastp (version 0.19.7) software was used to perform basic statistical analysis on the quality of the raw reads and to obtain clean data. Bowtie2 software was used for read alignment. MACS3 software was used for callpeak. SAMtools software was used for index construction and IGV input data processing. ChIPseeker software was used for peak annotation. The clusterProfiler software was used for downstream pathway analysis.
Total cellular polyamine measurement
Total cellular lysate total polyamine levels in detached cells with or without SAT1 knockdown were measured via a fluorometric total polyamine assay kit (BioVision, Cat# K475) according to the manufacturer’s instructions.
Polyamine uptake assay
3H-putrescine uptake was monitored in detached cells with or without SAT1 knockdown to evaluate the polyamine uptake assay. The cells were cultured under detached conditions for 12 h before the addition of 0.5 μCi putrescine [2,3-3H(N)] dihydrochloride (American Radiolabeled Chemicals, Cat# 0279) for 30 min at 37 °C. Then, the cells were washed twice with cold PBS and lysed with SOLVABLE (PerkinElmer, Cat# 6NE9100). The lysate was transferred to a scintillation vial containing liquid scintillation cocktails and immediately subjected to scintillation counting.
Arginine uptake assay
The detached cells with or without SAT1 knockdown were cultured in DMEM containing 1% arginine for 12 h before the addition of 1 μCi L-[2,3,4-3H]-arginine (American Radiolabeled Chemicals, Cat# 1421) for 60 min at 37 °C. Then, the cells were washed twice with cold PBS and lysed with SOLVABLE (PerkinElmer, Cat# 6NE9100). The lysate was transferred to a scintillation vial containing liquid scintillation cocktails and immediately subjected to scintillation counting.
Molecular docking
For molecular docking between the SAT1 and histone H3 proteins, SAT1 (PDB ID: 2B5G) was downloaded from the PDB database, and H3 (AlphaFoldDB ID: AF-Q6NXT2-F1) was downloaded from the AlphaFold protein structure database. Pymol 2.3.0 was used to remove protein crystal water, original ligands, etc., and the protein structure was imported into AutoDocktools (v1.5.6) for hydrogenation, calculating the charge, assigning the charge, specifying the atomic type, and saving in the “pdbqt” format. The prediction of a series of docking poses characterizing the intermolecular energy landscape in protein interactions is performed on the GRAMM web server. The relevant parameters of the heterotrimeric complex SAT1--H3 were set to center_x = 32.37, center_y = −22.14, and center_z = −18.89; the search space parameters were size_x: 73, size_y: 87, and size_z: 346; and the remaining parameters were set by default. PyMOL 2.3.0 was used to analyze the interaction mode of the docking results.
For molecular docking, the 3D structure of ginkgolide B (PubChem CID: 65243; CAS: 15291-77-7) was obtained from PubChem in SDF format, minimized in ChemBio3D Ultra 14.0 (RMS gradient ≤0.001), and saved as mol2. The optimized molecule was then processed in AutodockTools-1.5.6 for hydrogen addition, charge calculation, and conversion to pdbqt format. The SAT1-H3 complex was prepared using Pymol 2.3.0 to remove crystal waters and original ligands, and further processed in AutoDocktools for hydrogenation, charge calculation, and conversion to pdbqt. POCASA 1.1 identified potential binding sites, and CB-DOCK2 performed the docking. The interaction modes were analyzed using PyMOL 2.3.0.
Detection of in vitro histone H3 acetylation by SAT1 using Western blot
200 ng of histone H3 (His-tagged, Abcam, cat# ab198757) substrate was mixed with 50 ng of recombinant flag-tagged SAT1 purified from 293T-overexpressing cells in HAT buffer (50 mM Tris-HCl, pH 8.0, 10% (v/v) glycerol, 1 mM dithiothreitol, 0.1 mM EDTA, pH 8.0, and 10 μM acetyl-CoA) in a final volume of 30 μL for 45 min at 30 °C, and the mixture was terminated by the addition of SDS‒PAGE sample buffer and heating of the samples to 95 °C for 5 min. Acetylated proteins were visualized via western blotting using the indicated primary antibodies.
Evaluation of in vitro histone H3 acetylation by SAT1 using 3H-acetyl CoA
200 ng of His-tagged H3 and 50 ng of flag-tagged SAT1 recombinant were incubated at 30 °C for 30 min in a 30-μl final reaction mixture consisting of 50 mM Tris-HCl, pH 8.0, 10% (v/v) glycerol, 1 mM dithiothreitol, 0.1 mM EDTA, pH 8.0, and 0.5 μCi of [acetyl-3H]-coenzyme A (Revvity, Cat # NET290050UC). The reaction mixture was then incubated with anti-His tag magnetic beads to capture H3 protein. After washing with TBS buffer 3 times, the His-H3-bound magnetic bead suspension was transferred to a scintillation vial containing liquid scintillation cocktails and immediately subjected to scintillation counting.
DTNB-coupled SAT1 enzyme activity assay
SAT1 activity assays were performed as described elsewhere. SAT1 enzyme activity is coupled with the DTNB (Ellman’s reagent) (5,5-dithio-bis-(2-nitrobenzoic acid) assay. Ellman’s reagent (2 mM DTNB and 1 mM EDTA in 50 mM Tris-HCl, pH 7.5) was freshly prepared prior to the start of the experiment. Fifty nanograms of recombinant flag-tagged SAT1 purified from 293T-overexpressing cells was mixed with substrate buffer (50 mM Tris-HCl (pH 8.0), 1 mM EDTA (pH 8.0), 0.5 mM acetyl-CoA, 1 mM spermidine or 200 ng of histone H3 as indicated) in a final volume of 50 μL for 10 min at 35 °C, and the reaction was stopped by heating the samples at 85 °C for 5 min. After cooling at 14 °C for 2 min, 50 μl of Ellman’s reagent was added to the reaction system and incubated at 25 °C for 30 min. The production of CoA-SH by SAT1 was measured via a 412-nm absorption spectrum to evaluate the SAT1 enzyme activity.
13C Stable Isotope Tracing
13C-labeled glucose (D-Glucose-13C6, Sigma‒Aldrich, Cat#: 389374) and glutamine (L-Glutamine-13C5, Sigma‒Aldrich, Cat#: 605166) were utilized for tracing experiments. For glucose tracing, cells were incubated in glucose-free RPMI-1640 with 11.2 mM 13C6-glucose and 10% dialyzed FBS for 4 h, followed by two washes with cold PBS and metabolite extraction. Similarly, for glutamine tracing, cells were cultured in glutamine-free RPMI-1640 with 2.1 mM 13C5-glutamine and 10% dialyzed FBS under the same conditions. Cell count, determined in parallel experiments, was used for normalization.
Metabolites were extracted using an ice-cold mixture of methanol, chloroform, and H2O (50:50:20). Dried polar metabolites underwent derivatization before GC‒MS analysis using a Thermo 1300 equipped with a 30 m DB-35MS capillary column (Agilent Technologies) and a Thermo ISQ QD MS. Spectra were analyzed with Thermo TraceFinder software, and isotope natural abundance corrections were made using the R package ‘IsoCorrectoR’. Metabolite enrichment was calculated by acquiring peak area values from M + 0 to M + n, normalizing to cell count, and summing 13C-labeled isotopomers (M + 1 to M + n) to determine total 13C enrichment. The M + 0 value represented the unlabeled fraction.
SAT1 inhibitor screen
To screen potential SAT1 inhibitors from the “food as medicine” compound library, two rounds of screening were performed. In the primary screen, an in vitro SAT1 activity assay using 3H-acetyl-CoA was performed in the presence of compounds (10 μM) in the in vitro reaction system. Briefly, 200 ng of His-tagged H3 and 50 ng of flag-tagged SAT1 recombinant were incubated at 30 °C for 30 min in a 30-μl final reaction mixture consisting of 50 mM Tris-HCl, pH 8.0, 10% (v/v) glycerol, 1 mM dithiothreitol, 0.1 mM EDTA, pH 8.0, 0.5 μCi of [acetyl-3H]-coenzyme A and 10 μM candidate compound or DMSO vehicle. The reaction mixture was then incubated with anti-His tag magnetic beads to capture H3 protein, and the SAT1 activity was evaluated by comparing the radioactivity of H3 protein in the compound-treated group with that in the DMSO-treated group via scintillation counting.
In the second round of screening, flag-tagged SAT1-overexpressing ovarian cancer cells were pretreated with 20 μM candidate compound or the DMSO vehicle for 24 h under detached conditions. Subsequently, flag-tagged SAT1 protein in the cells was immunoprecipitated and used in the in vitro SAT1 activity assay as described above to compare cellular SAT1 activity in cells treated with different compounds. Detailed information of the drug library is shown in Table S10.
SAT1 Kinetic Analysis
DTNB-coupled SAT1 enzyme activity assay was used to perform SAT1 kinetic analysis as described above. A SAT1 activity assay with different concentrations of ginkgolide B was performed to determine the Ki value. The data were fitted via nonlinear regression in Prism 8. A series of concentrations of ac-CoA and 200 ng of histone H3 were used as substrates. A linear transform Lineweaver‒Burk plot was used to determine the type of enzyme inhibited by ginkgolide B.
Plasmid construction and mutagenesis
The Gateway cloning system was utilized for plasmid construction. Gene ORFs, with or without His-tag or flag-tag, were PCR-amplified using Q5 high-fidelity DNA polymerase and cloned into the pENTR/D-TOPO vector. Site-directed mutations in the recombinant pENTR/D-TOPO plasmids were introduced using the QuikChange XLII kit (Agilent) and mutation-specific primers. These plasmids were then subcloned into the pLHCX-derived Gateway destination vector for mammalian cell overexpression using LR Clonase II. The ACLY-NES fusion construct was created using primers encoding the PKI NES sequence (LALKLAGLDI). Detailed information on primers, destination vectors, and cloning strategies for each overexpression plasmid is provided in Table S5.
ID8 peritoneal dissemination mouse model
Murine ovarian ID8 cells expressing GFP-luciferase were stably transduced with pLKO control or SAT1 shRNA lentivirus. ID8 cells were selected with puromycin treatment and enriched via GFP sorting. A total of 5 million luciferase-expressing ID8 cells with or without SAT1 knockdown were subsequently injected directly into the abdominal cavity of C57BL/6 mice (female, 6 weeks old; Charles River). In vivo bioluminescence imaging was used to monitor tumor development. Fluorescence analysis was conducted via Living Image software (PerkinElmer, Inc.). In parallel experiments, 2 h or 12 h after luc-ID8 inoculation, disseminating ID8 tumor cells in the abdominal cavity were flushed with PBS and sorted with GFP, after which they were subjected to a western blot analysis to evaluate the SAT1 expression level.
ID8 syngeneic ovarian cancer residual disease mouse model
To mimic the residual disease development, a series of decreasing number of murine ovarian ID8 cells were tested to define the minimum number of cells needed to form metastatic tumor in the ID8 peritoneal dissemination mouse model and confirmed that at least half a million cells is needed to establish the tumor development. In the formal experiment, a total of half a million luciferase-expressing ID8 cells were directly injected into the abdominal cavity of C57BL/6 mice (female, 6 weeks old, Charles River). Mice were treated with vehicle or chemo (cisplatin 5 mg/kg, taxol 10 mg/kg, IP injection) in a total of 4 rounds from week 1 to week 3. For the maintenance therapy group, mice were additionally treated with ginkgolide B for two more weeks from week 3 to week 5. Ginkgolide B was intraperitoneally injected every two days in total two weeks at a dose of 5 mg/kg. In vivo bioluminescence imaging was used to monitor tumor development from week 1 to week 10 when all the mice were alive. Blood was collected retro-orbitally at week 6, one week after ginkgolide B treatment, and used for hematological blood tests. Hematoxylin‒eosin (HE) staining was used to perform histological analysis of tissue sections from representative mice receiving ginkgolide B maintenance therapy.
Statistics & reproducibility
No data were excluded from the analyses. The Investigators were not blinded to allocation during experiments and outcome assessment. GraphPad Prism 8 software was used for statistical analysis and graphical presentation. The statistical analysis of significance was based on a two-tailed Student’s t test or one-way or two-way ANOVA with Bonferroni post hoc multiple-comparisons, as indicated in the figure legend. It should be noted that statistical analyses are based on a set of assumptions, such as homogeneity of variances and a normal distribution. The variance between the groups being statistically compared was found to be similar. P values of 0.05 or less were considered statistically significant. No statistical method was used to predetermine the sample size for any in vivo experiments. Instead, the minimum sample size was employed, as a significant difference was detected in preliminary studies.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data generated in this study are provided in the Supplementary Information/Source Data file. RNA-seq data derived from cells in detached vs attached conditions and SAT1 knockdown cells vs pLKO cells are available at Gene Expression Omnibus under accession code GSE267614. ChIP-seq data derived from detached SAT1 knockdown cells vs pLKO cells in detached conditions are available at Gene Expression Omnibus under accession code GSE267703. Relevant approvals from China’s Ministry of Science and Technology related to the export of genetic information and materials relevant to this work was obtained if applicable. All sequencing data are publicly available. Source data are provided with this paper.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Marks, Z. R. C. et al. Interferon-epsilon is a tumour suppressor and restricts ovarian cancer. Nature 620, 1063–1070 (2023).
Zheng, X. et al. Single-cell analyses implicate ascites in remodeling the ecosystems of primary and metastatic tumors in ovarian cancer. Nat. Cancer 4, 1138–1156 (2023).
Tavares, V. et al. paradigm shift: a comprehensive review of ovarian cancer management in an era of advancements. Int. J. Mol. Sci. 25, 1845 (2024).
Pan, C. et al. Cisplatin-mediated activation of glucocorticoid receptor induces platinum resistance via MAST1. Nat. Commun. 12, 4960 (2021).
Tao, L. et al. Repression of LSD1 potentiates homologous recombination-proficient ovarian cancer to PARP inhibitors through down-regulation of BRCA1/2 and RAD51. Nat. Commun. 14, 7430 (2023).
Kim, H. et al. Combining PARP with ATR inhibition overcomes PARP inhibitor and platinum resistance in ovarian cancer models. Nat. Commun. 11, 3726 (2020).
Gonzalez-Martin, A. et al. Niraparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med 381, 2391–2402 (2019).
Li, J. et al. ROS-regulated phosphorylation of ITPKB by CAMK2G drives cisplatin resistance in ovarian cancer. Oncogene 41, 1114–1128 (2022).
Li, J. et al. Tyrosine catabolism enhances genotoxic chemotherapy by suppressing translesion DNA synthesis in epithelial ovarian cancer. Cell Metab. 35, 2044–2059 e2048 (2023).
Chowdhury, S. et al. Proteogenomic analysis of chemo-refractory high-grade serous ovarian cancer. Cell 186, 3476–3498 e3435 (2023).
Gupta, N. et al. BLM overexpression as a predictive biomarker for CHK1 inhibitor response in PARP inhibitor-resistant BRCA-mutant ovarian cancer. Sci. Transl. Med 15, eadd7872 (2023).
Karakashev, S. et al. EZH2 Inhibition Sensitizes CARM1-High, Homologous Recombination Proficient Ovarian Cancers to PARP Inhibition. Cancer Cell 37, 157–167 e156 (2020).
Gogola, E. et al. Selective Loss of PARG Restores PARylation and Counteracts PARP Inhibitor-Mediated Synthetic Lethality. Cancer Cell 33, 1078–1093 e1012 (2018).
Lahtinen, A. et al. Evolutionary states and trajectories characterized by distinct pathways stratify patients with ovarian high grade serous carcinoma. Cancer Cell 41, 1103–1117 e1112 (2023).
Song, M. et al. IRE1alpha-XBP1 controls T cell function in ovarian cancer by regulating mitochondrial activity. Nature 562, 423–428 (2018).
Chae, C. S. et al. Tumor-Derived Lysophosphatidic Acid Blunts Protective Type I Interferon Responses in Ovarian Cancer. Cancer Discov. 12, 1904–1921 (2022).
Quiralte, M. et al. Proteomic profiles of peritoneal fluid-derived small extracellular vesicles correlate with patient outcome in ovarian cancer. J. Clin. Invest. 134, e176161 (2024).
Weems, A. D. et al. Blebs promote cell survival by assembling oncogenic signalling hubs. Nature 615, 517–525 (2023).
Ford, C. E., Werner, B., Hacker, N. F. & Warton, K. The untapped potential of ascites in ovarian cancer research and treatment. Br. J. Cancer 123, 9–16 (2020).
Jin, L. et al. The PLAG1-GDH1 Axis Promotes Anoikis Resistance and Tumor Metastasis through CamKK2-AMPK Signaling in LKB1-Deficient Lung Cancer. Mol. Cell 69, 87–99 e87 (2018).
Jiang, L. et al. Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature 532, 255–258 (2016).
Rossi, M. et al. PHGDH heterogeneity potentiates cancer cell dissemination and metastasis. Nature 605, 747–753 (2022).
Wang, X. et al. UDP-glucose accelerates SNAI1 mRNA decay and impairs lung cancer metastasis. Nature 571, 127–131 (2019).
Chap, B. S., Rayroux, N., Grimm, A. J., Ghisoni, E. & Dangaj Laniti, D. Crosstalk of T cells within the ovarian cancer microenvironment. Trends Cancer 10, 1116–1130 (2024).
Anadon, C. M. et al. Ovarian cancer immunogenicity is governed by a narrow subset of progenitor tissue-resident memory T cells. Cancer Cell 40, 545–557 e513 (2022).
Holbert, C. E., Cullen, M. T., Casero, R. A. Jr. & Stewart, T. M. Polyamines in cancer: integrating organismal metabolism and antitumour immunity. Nat. Rev. Cancer 22, 467–480 (2022).
Gerstberger, S., Jiang, Q. & Ganesh, K. Metastasis. Cell 186, 1564–1579 (2023).
Labuschagne, C. F., Cheung, E. C., Blagih, J., Domart, M. C. & Vousden, K. H. Cell Clustering Promotes a Metabolic Switch that Supports Metastatic Colonization. Cell Metab. 30, 720–734 e725 (2019).
Boese, A. C. et al. Succinyl-CoA ligase ADP-forming subunit beta promotes stress granule assembly to regulate redox and drive cancer metastasis. Proc. Natl Acad. Sci. USA 120, e2217332120 (2023).
Matthews, H. K., Bertoli, C. & de Bruin, R. A. M. Cell cycle control in cancer. Nat. Rev. Mol. Cell Biol. 23, 74–88 (2022).
Saxena, S. & Zou, L. Hallmarks of DNA replication stress. Mol. Cell 82, 2298–2314 (2022).
Sungalee, S. et al. Histone acetylation dynamics modulates chromatin conformation and allele-specific interactions at oncogenic loci. Nat. Genet 53, 650–662 (2021).
Aye, I. et al. Placental sex-dependent spermine synthesis regulates trophoblast gene expression through acetyl-coA metabolism and histone acetylation. Commun. Biol. 5, 586 (2022).
Puleston, D. J. et al. Polyamine metabolism is a central determinant of helper T cell lineage fidelity. Cell 184, 4186–4202 e4120 (2021).
Shvedunova, M. & Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 23, 329–349 (2022).
Sivanand, S. et al. Nuclear Acetyl-CoA Production by ACLY Promotes Homologous Recombination. Mol. Cell 67, 252–265 e256 (2017).
Libby, P. R. & Porter, C. W. Inhibition of enzymes of polyamine back-conversion by pentamidine and berenil. Biochem Pharm. 44, 830–832 (1992).
Tan, D. S., Agarwal, R. & Kaye, S. B. Mechanisms of transcoelomic metastasis in ovarian cancer. Lancet Oncol. 7, 925–934 (2006).
Bayraktar, E., Chen, S., Corvigno, S., Liu, J. & Sood, A. K. Ovarian cancer metastasis: Looking beyond the surface. Cancer Cell 42, 1631–1636 (2024).
Wu, S. et al. Targeting glutamine dependence through GLS1 inhibition suppresses ARID1A-inactivated clear cell ovarian carcinoma. Nat. Cancer 2, 189–200 (2021).
Zhou, W. et al. Targeting the mevalonate pathway suppresses ARID1A-inactivated cancers by promoting pyroptosis. Cancer Cell 41, 740–756 e710 (2023).
Murthy, D. et al. Cancer-associated fibroblast-derived acetate promotes pancreatic cancer development by altering polyamine metabolism via the ACSS2-SP1-SAT1 axis. Nat. Cell Biol. 26, 613–627 (2024).
Murthy, D. et al. The MUC1-HIF-1alpha signaling axis regulates pancreatic cancer pathogenesis through polyamine metabolism remodeling. Proc. Natl Acad. Sci. USA 121, e2315509121 (2024).
Li, J. et al. ODF2L acts as a synthetic lethal partner with WEE1 inhibition in epithelial ovarian cancer models. J Clin. Invest. 133, e161544 (2023).
Gallo, D. et al. CCNE1 amplification is synthetic lethal with PKMYT1 kinase inhibition. Nature 604, 749–756 (2022).
Mossmann, D. et al. Arginine reprograms metabolism in liver cancer via RBM39. Cell 186, 5068–5083 e5023 (2023).
Lee, M. S. et al. Ornithine aminotransferase supports polyamine synthesis in pancreatic cancer. Nature 616, 339–347 (2023).
Han, X. et al. Activation of polyamine catabolism promotes glutamine metabolism and creates a targetable vulnerability in lung cancer. Proc. Natl Acad. Sci. USA 121, e2319429121 (2024).
Ye, W. et al. Anti-atherosclerotic effects and molecular targets of ginkgolide B from Ginkgo biloba. Acta Pharm. Sin. B 14, 1–19 (2024).
Swanton, C. et al. Embracing cancer complexity: Hallmarks of systemic disease. Cell 187, 1589–1616 (2024).
Bjorklund, J. et al. Risk of Venous Thromboembolic Events After Surgery for Cancer. JAMA Netw. Open 7, e2354352 (2024).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (82422060 to CP; 82202841 to J Li) and the National Key R&D Program of China (2021YFA0911600 to CP).
Author information
Authors and Affiliations
Contributions
G.N., H.T., Q.M., M.W., Y.G., J.W., M.X., J. Liu and S.Y. provided clinical information. X.H. and S.C. performed animal experiments. C. Zheng, J. Lu, T.Y., C. Zhang, and W.P. performed all the other experiments. C.P. and J. Li designed and supervised the study. C.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Joshua Bauer, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, C., Niu, G., Tan, H. et al. A noncanonical role of SAT1 enables anchorage independence and peritoneal metastasis in ovarian cancer. Nat Commun 16, 3174 (2025). https://doi.org/10.1038/s41467-025-58525-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58525-8
This article is cited by
-
ZDHHC9 and spermine metabolism: a palmitoylation-driven pathway to prostate carcinogenesis
Journal of Translational Medicine (2025)
-
Glutamine-driven metabolic reprogramming promotes CAR-T cell function through mTOR-SREBP2 mediated HMGCS1 upregulation in ovarian cancer
Journal of Translational Medicine (2025)
-
Regulation of polyamine interconversion enzymes affects α-Synuclein levels and toxicity in a Drosophila model of Parkinson’s Disease
npj Parkinson's Disease (2025)
-
Metabolic support protects mucosa from ferroptosis in radiation-induced mucositis
Nature Communications (2025)