Abstract
The rapid adoption of Artificial Intelligence (AI) in medical imaging raises fairness and privacy concerns across demographic groups, especially in diagnosis and treatment decisions. While federated learning (FL) offers decentralized privacy preservation, current frameworks often prioritize collaboration fairness over group fairness, risking healthcare disparities. Here we present FlexFair, an innovative FL framework designed to address both fairness and privacy challenges. FlexFair incorporates a flexible regularization term to facilitate the integration of multiple fairness criteria, including equal accuracy, demographic parity, and equal opportunity. Evaluated across four clinical applications (polyp segmentation, fundus vascular segmentation, cervical cancer segmentation, and skin disease diagnosis), FlexFair outperforms state-of-the-art methods in both fairness and accuracy. Moreover, we curate a multi-center dataset for cervical cancer segmentation that includes 678 patients from four hospitals. This diverse dataset allows for a more comprehensive analysis of model performance across different population groups, ensuring the findings are applicable to a broader range of patients.
Similar content being viewed by others
Introduction
Recent studies indicate that AI systems can exhibit biases against demographic groups based on attributes such as age, race, gender, and socioeconomic status, particularly in medical imaging tasks like disease detection and treatment recommendations, which raises significant ethical concerns1,2. To address privacy issues related to medical data, FL has emerged as a preferred approach, with several key methods tackling fairness in different ways. FedAvg3 averages model updates from clients but may lead to biased outcomes if certain demographic groups are underrepresented. FedProx4 introduces a proximal term to keep local models aligned with the global model, mitigating the impact of Out-of-Distribution (OoD) data. FedNova5 normalizes local updates based on the number of training steps to balance contributions from clients with varying data amounts, enhancing convergence. SCAFFOLD6 employs control variates to stabilize learning and reduce bias from OoD data distributions. Due to its privacy-preserving capabilities, FL has gained significant attention in the medical field in recent years.
However, most FL research has focused on performance fairness—achieving consistent accuracy across clients—while overlooking group fairness, a gap that risks exacerbating healthcare disparities by underrepresenting diverse demographic groups. Addressing this issue requires models that go beyond privacy preservation to actively ensure fairness. Prioritizing such models is critical to tackling ethical concerns, fostering equitable treatment across demographic groups, and mitigating biases in healthcare applications. Moreover, these advancements would enhance the robustness and generalizability of medical AI systems, paving the way for more inclusive and dependable healthcare solutions.
Several algorithms aim to enhance group fairness in centralized learning7,8,9,10. For example, Fair-Mixup10 generates a distribution path connecting sensitive groups and regularizes the smoothness of the path to improve the generalization of group fairness metrics. Some works11,12 formulate the constrained optimization problem as a two-player game and analyze the solutions and generalization bounds. Additionally, Fair-CDA13 illustrates that group fairness can be promoted by regularizing models along the transitional paths of sensitive attributes among groups. While these strategies show promise for achieving better accuracy and fairness across various benchmarks, their application in medical settings often requires data from multiple centers, which raises privacy concerns. Integrating FL with fair machine learning presents a potential solution by allowing institutions to collaboratively train models without sharing raw patient data14. However, directly addressing group fairness in heterogeneous distributed settings often necessitates feature exchange10,13, which can lead to privacy leaks and contradict the fundamental principles of FL.
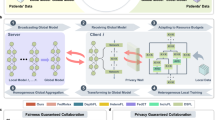
Most importantly, achieving fairness while preserving privacy in medical imaging tasks remains a significant challenge15,16,17. FairFed18 addresses this challenge by using fairness-aware aggregation to enhance fairness in FL. However, FairFed primarily focuses on group fairness criteria and is not specifically designed to ensure performance fairness, which we refer to as equal accuracy (EA) in this study. To address these limitations, we propose FlexFair, a FL framework that integrates three fairness criteria: EA19, demographic parity19 (DP), and equal opportunity8 (EO). FlexFair was evaluated across four distinct medical imaging tasks: polyp segmentation, fundus vascular segmentation, cervical cancer segmentation, and skin disease classification. Specifically, for cervical cancer segmentation, we curated a multi-centre and diverse dataset of 678 patients from four hospitals, reflecting the demographic diversity in clinical settings. By leveraging data from multiple institutions, FlexFair enhances generalizability and ensures its approach can be effectively applied in real-world scenarios. The pipeline of FlexFair is shown in Fig. 1. Our framework adaptively balances trade-offs between fairness and accuracy, and theoretical analysis shows it can accommodate different fairness metrics by modifying a component of the loss function. Results demonstrate that FlexFair achieves high performance and robustness while adhering to fairness criteria like DP, EO, and EA, providing an effective mechanism for ensuring fairness and privacy protection in medical imaging research.
a Overview of the proposed FlexFair and its comparison with both the centralized learning and the vanilla FL method, FedAvg. FlexFair effectively mitigates prediction disparities from task models through a weighted penalty mechanism while prioritizing data privacy by integrating a federated framework. b Detailed design of FlexFair. FlexFair addresses fairness and privacy challenges in federated environments by incorporating multiple sensitive attributes, e.g., age, gender, and site, into its framework. It evaluates fairness using metrics like EA, DP, and EO, and integrates these attributes into a weighted regularized loss to ensure the training process promotes fairness across all groups.
Results
To evaluate the performance of FlexFair, we conducted experiments across various medical imaging tasks, including segmentation and diagnostic challenges, covering real-world scenarios with diverse data distributions and complexities. To ensure a thorough assessment, we employed a comprehensive set of evaluation metrics that simultaneously measured accuracy and fairness. These metrics included dice scores for segmentation and overall accuracy for diagnostic tasks, alongside fairness criteria such as EA, DP and EO. Each experiment was executed across five distinct random seeds to enhance the reliability of our findings, and we reported our results with accompanying statistical analyses to provide a comprehensive understanding of FlexFair’s performance and consistency across different initialization.
We conducted a comparative analysis of FlexFair against several established FL and fair machine learning methods, such as FedAvg, FedProx, FedNova, SCAFFOLD, FairFed, and FairMixup. This allowed us to assess its ability to balance accuracy and fairness in diverse scenarios. The results demonstrated FlexFair’s superior capacity to manage the inherent variability in medical imaging tasks while reducing performance disparities across clients. By enhancing the overall effectiveness of FL in medical applications, FlexFair also highlights its potential to promote equitable healthcare delivery, particularly in multi-institutional and resource-limited settings.
FlexFair achieves flexible fairness in diverse medical imaging scenarios
FlexFair exhibits consistent improvements in both fairness and performance metrics, as demonstrated by the comparative analysis in Fig. 2. The figure highlights FlexFair’s superior performance across multiple datasets, including polyp, fundus vascular, cervical cancer, and skin disease, where it consistently achieves lower fairness gaps, indicating more equitable outcomes. By running each method across five random seeds and averaging the results, the robustness of FlexFair’s performance is confirmed. The Pareto front plots, which report the top 20 test results under different weights, further underscore FlexFair’s ability to balance high accuracy (measured by dice scores and accuracy) with minimal fairness gaps (EA, DP, EO). This comprehensive evaluation reveals that FlexFair not only excels in predictive accuracy but also maintains fairness across various demographic attributes, making it a promising approach for both segmentation and diagnostic tasks in diverse medical applications.
We compare FlexFair with six baseline methods (FedAvg, FedNova, FedProx, SCAFFOLD, FairFed, and FairMixup) across four datasets: polyp, fundus vascular, cervical cancer, and skin disease. Each method is evaluated on fairness (EA, DP, EO) and accuracy metrics (dice score for segmentation tasks and accuracy for diagnostic tasks). a–c illustrate the Pareto front for segmentation datasets, highlighting trade-offs between fairness and accuracy. FlexFair highlighted with red color consistently achieves superior dice scores and fairness gap. d–f depict maximum gap values for dice scores, where lower values indicate greater fairness. FlexFair outperforms other methods by minimizing the max dice gap across sites. g–j analyze fairness and accuracy in diagnostic tasks on the skin disease dataset, emphasizing FlexFair’s ability to balance demographic parity and equal opportunity across age and gender attributes. k–n confirm that FlexFair achieves the lowest max dice gap values, ensuring equitable performance across all metrics and datasets. Source data are provided as a Source Data file.
In terms of EA, we evaluated three segmentation tasks and chose the site as the sensitive attribute. We measured the maximum dice performance gap across different sites, calculated the overall mean dice performance, and reported the maximum gap between the dice performance of each site and the overall mean dice as the fairness gap, as shown in Equation (1). Figure 2a–c presents the Pareto front, highlighting the trade-off between dice scores and fairness across various methods. FlexFair, depicted in red, stands out for its superior performance, achieving high dice scores while maintaining robust fairness metrics. Among the baseline methods, most advanced FL approaches outperform FedAvg, which yields moderate dice scores but often falls short in terms of fairness. FedProx demonstrates competitive performance but struggles to strike an optimal balance between fairness and accuracy, frequently exhibiting higher max dice gaps that indicate greater unfairness in segmentation outcomes. SCAFFOLD shows inconsistent results, performing well on the fundus vascular dataset but significantly underperforming on the cervical cancer dataset, as shown in Fig. 3. Due to its poor performance on the cervical cancer dataset, the Pareto front for SCAFFOLD is omitted. More detailed results are shown in Table 1. Figure 2d–f presents the maximum gap values of dice performance across different sites, serving as a measure of unfairness-higher maximum gap values indicate greater unfairness. The minimal maximum gap values for each method, exceeding a certain performance threshold, are reported. In Fig. 2, FedAvg exhibits the highest maximum gap, signalling the greatest unfairness, while FlexFair achieves the lowest maximum gap, indicating the most equitable performance. In Fig. 2e for the fundus vascular Dataset, the maximum gap values span from ~0.11 to 0.13. FedAvg again demonstrates the highest maximum gap, suggesting the highest level of unfairness, whereas FlexFair maintains the lowest maximum gap, indicating the most balanced performance. In Fig. 2f for the cervical cancer dataset, SCAFFOLD shows the highest maximum gap, reflecting significant unfairness, while FlexFair again exhibits the lowest maximum gap, reinforcing its effectiveness in ensuring fairness.
We evaluate FlexFair against baseline methods on segmentation tasks across three datasets: cervical cancer, polyp, and fundus vascular. Violin-box plots depict the distribution of the top 20 test results for each method across different weight configurations and random seeds. The boxes represent the interquartile range (IQR), with the median marked by the white line and the mean indicated by the red 'x'. Whiskers extend to data points within 1.5 times the IQR, with black diamonds showing outliers. Dice scores, serving as a metric for segmentation accuracy, highlight FlexFair’s consistently superior performance across all tasks, characterized by a tighter distribution around higher median and mean values compared to the baseline methods. Source data are provided as a Source Data file.
In terms of DP, we ensured that the predictor \(\hat{Y}\) treats different sensitive attribute groups equally by requiring that the prediction probabilities remain the same regardless of the value of the sensitive attribute A. We selected age and gender as sensitive attributes for this metric. To measure the expected predictions under a binary classification task, we applied the softmax function to the output logits of samples and selected the output value at index 1 to represent the probability of positive labels. We then calculated the overall expected predictions and reported the maximum gap between the expected predictions of each sensitive group and the overall expected predictions as the fairness gap, as shown in Equation (2).
In terms of EO, we ensured that the predictor \(\hat{Y}\) maintains EO for correct predictions across different sensitive attribute groups. We applied age and gender as sensitive attributes for this metric as well. To measure the expected predictions under a binary classification task, we applied the same approach as for DP: we applied the softmax function to the output logits of samples and selected the output value at index 1 to represent the probability of positive labels. For samples with positive labels, we calculated the overall expected predictions and reported the maximum gap between the expected predictions of each sensitive group and the overall expected predictions as the fairness gap, as shown in Equation (3). Similar to segmentation tasks, Fig. 2g–n presents the Pareto front and bar charts for the skin disease diagnosis task. The Pareto front uses accuracy as the performance metric, while DP and EO considering different attributes (age and gender) are used as fairness metrics. Notably, FlexFair (represented by the red line) consistently demonstrates superior performance across these metrics. For instance, in the age DP plot (g), FlexFair shows a significant improvement in accuracy as DP value increases. Similarly, the gender DP plot (i) highlights FlexFair’s ability to maintain fairness across genders while improving accuracy. These trends are consistently observed in the age EO (h) and gender EO (j) plots as well, further reinforcing FlexFair’s balanced and equitable performance across different demographic groups. The bottom row bar charts (k-n) provide a comparative analysis of the fairness metrics across different methods. In all four charts—age DP (k), age EO (l), gender DP (m), and gender EO (n)—FlexFair consistently exhibits lower values compared to other methods. The lower maximum gap value across all metrics indicates that FlexFair achieves more equitable performance, effectively reducing bias related to age and gender in diagnostic tasks. Overall, these visualizations underscore FlexFair’s superior performance in maintaining high accuracy while ensuring fairness across diverse patient groups.
FlexFair enhances accuracy in both segmentation and diagnostic tasks
Figure 3 presents a comparative analysis of various FL methods alongside FlexFair on segmentation tasks for three medical conditions. The dice scores, which serve as a measure of segmentation accuracy, consistently indicate that FlexFair outperforms the other methods under comparison. This superior performance is reflected in the violin plots, which visualize central tendency measures (mean and median), interquartile ranges, and outliers. The results highlight FlexFair’s ability to deliver both accuracy and robustness across diverse datasets, emphasizing its effectiveness.
To determine whether there are statistically significant differences in performance between FlexFair and the baseline methods, we performed t tests on each method’s accuracy, calculating the p value for each comparison. Table 1 and Table 2 present the mean and standard deviation from five random seeds for both segmentation and diagnostic tasks, which include polyp segmentation, fundus vascular segmentation, cervical cancer segmentation, and skin disease classification. The p values, which highlight the statistical significance of the results, were derived by comparing each method’s accuracy with that of FlexFair.
In these tables, FlexFair consistently outperforms other methods under similar fairness gaps. For example, in the polyp dataset, FlexFair achieves a dice score of 0.885 ± 0.004, and in the fundus vascular dataset, it scores 0.658 ± 0.007. FlexFair also excels in the cervical cancer dataset with a dice score of 0.801 ± 0.003. For skin disease classification, FlexFair records an accuracy of 0.824 ± 0.004 for age DP and 0.824 ± 0.003 for gender DP. The low p values (<0.05) in these tables confirm the statistical significance of FlexFair’s superior performance, indicating that its improvements are not due to random variations.
FlexFair exhibits consistent improvements in fairness and performance while maintaining user privacy
FlexFair prioritizes user privacy through its FL framework, which ensures that sensitive data remains decentralized. By allowing individual users to retain control of their data, FlexFair mitigates the risks associated with data sharing and potential breaches. This decentralized approach not only aligns with privacy regulations but also fosters a collaborative environment for model training without compromising user confidentiality. The distributed training approach enables multiple clients to work together effectively, training a shared model while keeping their raw data secure and local.
In addition to its decentralized architecture, FlexFair incorporates a flexible regularization term that accommodates various fairness criteria, addressing the inherent variability in medical imaging tasks. This adaptability is crucial for reducing disparities in model performance across different demographic groups, ensuring that healthcare delivery is equitable. Through empirical validation across diverse datasets, FlexFair demonstrates its ability to maintain high accuracy and fairness while safeguarding user privacy, making it an ideal solution for sensitive applications in resource-constrained settings.
Discussion
FlexFair stands out by effectively integrating three critical fairness criteria: EA, DP and EO. These criteria are essential for ensuring that machine learning models provide equitable outcomes across diverse demographic groups while safeguarding user privacy, particularly in sensitive applications like healthcare. In terms of EA, FlexFair demonstrates significant superiority over existing methods. FairMixup emphasizes fairness and achieves better Pareto front results than some baseline methods, though it remains less competitive compared to FlexFair. However, FairMixup’s approach comes with a notable drawback: it compromises privacy, a crucial concern in FL scenarios. On three segmentation datasets, FairFed underperforms compared to FlexFair, suggesting that it is not specifically tailored for these scenarios. In terms of DP and EO, FairFed occasionally attains higher accuracy. Despite this, FairFed struggles to achieve a better trade-off between fairness and accuracy, highlighting that while it excels in specific conditions, FlexFair demonstrates more consistent and superior performance across various tasks. Also, FlexFair’s simple implementation allows for easy integration and plug-and-play use, making it accessible for a wide range of applications. By consistently achieving lower fairness gaps and higher performance metrics, FlexFair showcases its ability to address the inherent variability in medical imaging tasks while maintaining robust and accurate outcomes. This dual capability highlights FlexFair’s potential as a valuable tool in achieving both fairness and accuracy in FL settings.
FlexFair distinguishes itself from traditional FL and centralized learning methods by its ability to meet various fairness criteria while ensuring privacy. Traditional FL methods often struggle to guarantee group fairness, as this typically requires centralized access to raw data or features, which is challenging to achieve without compromising data privacy. FlexFair addresses these limitations by incorporating a flexible regularization term within an FL framework. While FairFed aims to enhance group fairness in FL, it struggles to achieve a better balance between fairness and accuracy compared to FlexFair. In conclusion, FlexFair enables decentralized data processing, preserves user privacy, and promotes equity, making it particularly vital for applications in healthcare where both fairness and privacy are critical.
Another key contribution of this work is the collection of a private dataset for the important yet data-scarce disease of cervical cancer. By gathering clinical data from multiple centres and establishing a comprehensive multi-centre data setting, we have validated FlexFair’s effectiveness in a real-world scenario. This dataset not only enhances the demographic diversity required for thorough evaluation but also supports the development of an efficient diagnostic model for cervical cancer. It underscores the practical applicability and potential of FlexFair to support equitable and precise diagnostic outcomes, ultimately contributing to improved healthcare delivery in multi-institutional and resource-constrained settings.
After consulting with several clinicians, we gathered valuable feedback on the FlexFair algorithm. These clinicians, with extensive experience in the collection and analysis of multi-centre clinical data, provided insightful input on the algorithm’s real-world applications. They unanimously agreed that FlexFair significantly improved both model fairness and accuracy while ensuring data privacy. The ability of FlexFair to address various fairness criteria, such as DP and EO, was particularly praised, as it is crucial for mitigating diagnostic bias across diverse demographic groups. Additionally, clinicians highlighted its exceptional performance in multi-centre data environments, showcasing its potential for widespread adoption across different medical institutions. They believe that applying this algorithm will substantially enhance the fairness and effectiveness of medical diagnostics, particularly in resource-limited healthcare settings.
In conclusion, FlexFair’s innovative integration of fairness criteria significantly enhances model performance while addressing critical privacy concerns in healthcare applications. By effectively balancing EA, DP, and EO, FlexFair offers a promising approach for promoting equitable outcomes in FL. Despite its advantages, FlexFair is not without limitations. One notable challenge is the potential communication overhead associated with FL frameworks. As model updates are transmitted between local devices and the central server, there may be delays that can impact the speed of convergence during training. Additionally, achieving group fairness often necessitates access to sensitive attributes, which complicates effective performance in scenarios where such labels are not available. Therefore, developing our method in an unsupervised manner is essential. Addressing these challenges will be crucial for maximizing FlexFair’s effectiveness in diverse real-world scenarios, ensuring that it continues to deliver on its promise of fairness and accuracy while upholding user privacy.
Methods
The private dataset for this retrospective study was created under a waiver of informed consent, as the institutional review boards determined that the retrospective design and use of de-identified data posed minimal risk to participants. We conducted the research with a strict commitment to fairness, transparency, and respect, ensuring that all data were meticulously handled and protected. Additionally, we upheld diversity, inclusivity, academic integrity, and ethical guidelines, with no conflicts of interest throughout the study. The ethical considerations underlying this work were rigorously reviewed and approved by the ethics committees of all participating institutions: the Ethics Committee of Sun Yat-sen Memorial Hospital of Sun Yat-sen University (SYSKY-2024-400-01), the Ethics Committee of Guangdong Maternal and Child Health Hospital (202401201), the Ethics Committee of Sun Yat-sen University Cancer Center (SL-G2023-231-01), and the Ethics Committee of Guangdong Province Traditional Medical Hospital (BE2023-146).
Private cervical cancer dataset collection
Cervical cancer
Cervical cancer is the fourth most common malignancy in women, with a 6.5% incidence and 7.7% mortality worldwide. In 2020, there have been more than 340,000 deaths due to cervical cancer were reported worldwide, which remains a significant threat to female health20. External beam radiation therapy (EBRT) and brachytherapy (BT) are the primary radiation modalities for locally advanced cervical cancer21. Accurate segmentation of clinical target volumes and organs at risk is a crucial step for EBRT and BT treatment options, as inaccuracies may result in either over-irradiation of normal tissues or insufficient radiation dose delivery to the tumor22. Magnetic resonance imaging (MRI)-guided treatment planning in EBRT and BT for cervical cancer demonstrates a significant advantage in tumor localization and assessment of tumor infiltration23,24. However, manual MRI image segmentation is a cumbersome process and may be inaccurate due to the inherent bias of the radiation oncologists. This underlines the necessity of rapid and accurate automatic segmentation methods that would improve the workflow efficiency of clinicians and reduce variability in radiotherapy planning.
Cervical cancer dataset collection
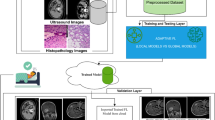
The dataset is composed of multiple pre-treatment pelvic MRI scans in female patients at four institutions, i.e., Sun Yat-sen Memorial Hospital of Sun Yat-sen University (center A), Sun Yat-sen University Cancer Center (center B) Guangdong Province Traditional Medical hospital (center C) and Guangdong Maternal and Child Health Hospital (center D). We define the inclusion criteria as follows: (1) an age ≥18 years; (2) confirmed pathological diagnosis of cervical cancer. We define the exclusion criteria as follows: (1) previous history of chemoradiotherapy therapy at cervical cancer; (2) those presenting tumors with a diameter of <5 mm that were invisible on MRI images; (3) image quality with severe artifact affecting the subsequent analysis. The workflow is illustrated in Fig. 4. The MRI protocols for the four centers are as follows: center A includes an axial T2-weighted sequence (repetition time [TR], 3500 ms; echo time [TE], 129 ms; slice thickness, 5 mm; acquisition matrix 384 × 269); center B includes an axial T2-weighted sequence (TR, 5100 ms; TE, 85 ms; slice thickness, 6 mm; acquisition matrix, 320 × 224); center C includes an axial T2-weighted sequence (TR, 5050 ms; TE, 72 ms; slice thickness, 5mm; acquisition matrix, 256 × 320);center D includes an axial T2-weighted sequence (TR, 3000 ms: TE, 98 ms: slice thickness,6mm; acquisition matrix, 256 × 288).
The cervical cancer dataset is collected across four medical centers with a detailed process outlining patient selection, exclusion criteria, and final cohort composition. From an initial pool of 1144 patients, individuals who meet the inclusion criteria (age ≥18 years and pathology-confirmed cervical cancer) and do not meet exclusion criteria (prior chemoradiotherapy, tumor diameter <5 mm, or severe motion artifacts in MRI) are included in the analysis. After applying these criteria, the final dataset comprises 89, 65, 278, and 246 patients from centers A, B, C, and D, respectively.
Cervical cancer dataset annotation
Radiologists used ITK-SNAP software (www.itksnap.org) to draw regions of interest (ROIs) around the tumor on T2W images to delineate the whole tumor volume. Discrepancies between the readers were resolved through consensus. These labeled ROIs are considered the ground truth data in training.
Dataset statistics
Polyp dataset
The polyp dataset comprises data from two distinct FL clients: CVC-30025 and Kvasir26, with sample sizes of 610 and 1000, respectively, as shown in Table 3. The table also displays the proportion of samples between these datasets, showing Kvasir constituting 62.1% and CVC-300 making up 37.9% of the total dataset. This notable discrepancy in data volume among groups underscores the imperative for models capable of adapting to diverse data sources while preserving accuracy amidst such imbalances.
Skin disease dataset
The skin disease dataset leverages the HAM-1000027 and BCN-2000028 datasets, known for their comprehensive annotations encompassing sensitive attributes such as age, gender, and skin type. After filtering samples without gender and age information, the sample sizes are 8819 for HAM-10000 and 7705 for BCN-20000. The gender distribution shows a slight male majority with 52.7% male and 47.3% female in HAM-10,000, and a slight female majority with 48.7% male and 51.3% female in BCN-20000. The age distribution indicates a higher number of samples (80.3% in HAM-10000 and 72.6% in BCN-20000) in the age group under 60 for both datasets. This dataset provides a robust foundation for evaluating the fairness and efficacy of skin disease detection algorithms across varied demographics.
Fundus vascular dataset
The fundus vascular dataset includes three primary datasets: CHASE-DB129, DRIVE30, and STARE31, with sample sizes of 28, 40, and 20, respectively. These datasets are limited in size, as illustrated in Table 3, presenting significant challenges in training robust machine learning models due to the constrained dataset size and the risk of overfitting.
Cervical cancer dataset
As previously mentioned, the cervical cancer dataset is collected from four centers: center A, center B, center C, and center D, with sample sizes of 1383, 1332, 323, and 328 respectively. Table 3 illustrates the distribution of samples among these centers, with center A having the largest share at 41.1%.
Network architecture
We selected SANet32 as our foundational architecture due to its effectiveness in medical image segmentation. SANet addresses challenges across three key dimensions: image color, background noise, and foreground-background distribution. It uses data augmentation methods like random color swapping during training to focus on shape and structural information, rather than lesion color. Attention mechanisms are employed to suppress background noise, and a post-processing strategy balances the distribution of predicted results during inference. For segmentation tasks, SANet serves as the backbone model, while ResNet-5033 is used for classification tasks.
Experimental settings
In our comparative study of FL methods, we assess and improve upon four FL methods: FedAvg3, FedProx4, SCAFFOLD6, FedNova34, and two state-of-the-art group fairness methods: FairMixup10, and FairFed18. For each dataset and task, we ensure that all methods use the same learning rate, batch size, and number of epochs.
To achieve the fairness-accuracy trade-off, we adjust specific hyperparameters within each method. For FlexFair, we adapt the weight of the penalty, while FedAvg remains unchanged as it is not designed for fairness. For FedProx, we adjust the penalty constant μ. Similarly, SCAFFOLD, FedNova, FairMixup, and FairFed have specific hyperparameter adjustments tailored to their frameworks, such as local client stepsize ηt, client momentum factor ρ, penalty weight λ, and perturbation ratio β. These adjustments allow us to comprehensively evaluate and enhance the performance of each method, highlighting the trade-offs between fairness and accuracy.
Statistics & reproducibility
To comprehensively evaluate both the performance and fairness of each algorithm, all experiments were conducted under five different random seeds, and each method was further explored with multiple hyperparameter settings (e.g., μ in FedProx, fairness penalty λ in FlexFair and FairMixup, and local client stepsize ηt in SCAFFOLD). This strategy ensures a thorough search of the fairness-accuracy trade-off space and provides robust conclusions about each method’s sensitivity to initialization and parameter choices.
In the bar charts and tables (Tables 1 and 2), each bar or entry corresponds to the averaged result and associated standard deviation over five seeds at a particular hyperparameter setting. To establish the statistical significance of any observed performance differences, we conducted paired t tests with the null hypothesis that no performance difference exists compared to our proposed method; p values below 0.05 indicate statistically significant improvements.
By systematically varying hyperparameters and repeating experiments under multiple random seeds, we ensure that our comparisons and conclusions about fairness and accuracy are robust to minor changes in initialization and parameter tuning. Sample size selection was based on literature references and practical data considerations. No statistical method was used to predetermine the sample size. No data were excluded from the analyses. Furthermore, the experimental process did not involve randomization, and no blinding was performed.
Evaluation metrics
We utilize three fairness metrics: EA, DP, and EO to assess the fairness of our methods. Specifically, we use EA to measure performance fairness in segmentation tasks, and DP and EO to measure group fairness in diagnostic tasks. Suppose that we have a set of data points {(xi, yi)} drawn from an unknown joint distribution over cx × cy, where cx is a subset of \({{\mathbb{R}}}^{d}\). The attribute A represents a sensitive characteristic that should not influence decision-making differently. \({{{\mathcal{A}}}}_{k}\) represents the condition set satisfying \(\cup {{{\mathcal{A}}}}_{k}={{\mathcal{A}}}\).
EA is a fairness metric used to address disparities in prediction accuracy across different groups. It quantifies unfairness by measuring the maximum difference in prediction accuracy between these groups. We specifically apply it to assess the gap in dice performance for segmentation tasks. The formula for EA is shown in Equation (1). In this context, \({{{\mathcal{A}}}}_{k}\) represents distinct hospitals. A predictor \(\hat{Y}\) satisfies EA if it minimizes the maximum difference in prediction accuracy across these hospital groups.
DP ensures that a predictor \(\hat{Y}\) treats different sensitive attribute groups equally. Specifically, it requires that the prediction probabilities remain the same regardless of the value of the sensitive attribute A: \(P(\hat{Y}| A=0)=P(\hat{Y}| A=1)\). In other words, the model’s predictions should not be influenced by variations in the sensitive attribute. DP emphasizes group fairness, aiming to ensure that individuals within different groups based on sensitive features receive positive decisions at equal rates. To evaluate the fairness of a trained model f under the DP definition, we use a relaxed metric called FDP. This metric quantifies the difference between the expected predictions for different sensitive attribute groups, as shown in Equation (2). The goal is for FDP to approach zero, indicating that the model achieves DP. However, meeting strict DP requirements can lead to reduced prediction accuracy, especially for certain predictions (such as hobbies or expertize) where genuine differences exist between groups. As an alternative, we consider other fairness criteria to address these limitations.
EO ensures that a predictor \(\hat{Y}\) maintains EO of correct predictions across different sensitive attribute groups. Specifically, the EO violation metric is given by Equation (3). Unlike DP, EO considers the correlation between Y and A, allowing for variations in base rates across groups. In real-world applications, EO serves as a fairness criterion when there are strict requirements for accurate predictions, and we prioritize the qualifications of candidates when making decisions. The goal is to minimize FEO while ensuring fairness across different sensitive attribute groups.
Algorithm
In the context of collaborative training in FL, it is possible to create a high-performing global model that may inadvertently incorporate latent discriminatory biases against specific demographic groups in the dataset. To address this issue, we develop a weighted-variance-regularization approach that aims to enhance the model’s fairness while preserving prediction accuracy.
For group fairness assessment, there have been various metrics, which quantify the disparity between the model’s performance on specific demographic groups and the average performance across all groups8,35,36,37,38,39,40,41,42. In this study, we consider the maximum performance gap among different groups as an evaluation measure to gauge the fairness of the learned model. The maximum performance gap is defined as follows:
where w represents a weight vector, ℓ denotes an utility function, \({{{\mathcal{A}}}}_{k}\) represent the k-th group and \({{\mathcal{A}}}={{{\mathcal{A}}}}_{1}\bigcup {{{\mathcal{A}}}}_{2}\bigcup \cdots \bigcup {{{\mathcal{A}}}}_{K}.\) Here \({R}_{k}({{\boldsymbol{\theta }}};{{{\mathcal{A}}}}_{k})\) and \(\bar{R}({{\boldsymbol{\theta }}},{{\bf{w}}};{{\mathcal{A}}})\) are empirical estimates of \({\mathbb{E}}[\ell ({{{\rm{f}}}}_{{{\boldsymbol{\theta }}}}({{\bf{x}}}),y)| {{{\mathcal{A}}}}_{k}]\) and \({\mathbb{E}}[\ell ({{{\rm{f}}}}_{{{\boldsymbol{\theta }}}}({{\bf{x}}}),y)| {{\mathcal{A}}}]\) respectively:
and
Next, we show that both DP and EO can be written in the form of Equation (4).
Example 1 (DP)
The DP violation metric is given by
Therefore, \({F}_{{{\rm{DP}}}}=F({{\mathcal{A}}},\ell,{{\bf{w}}})\) if \(\ell ({{{\rm{f}}}}_{{{\boldsymbol{\theta }}}},z)={\mathbb{I}}\{{{{\rm{f}}}}_{{{\boldsymbol{\theta }}}}({{\bf{x}}})=1\}\) and \({w}_{k}=\frac{{\sum }_{({{\bf{x}}},y)}{\mathbb{I}}\{({{\bf{x}}},y)\in {{{\mathcal{A}}}}_{k}\}}{{\sum }_{k}{\sum }_{({{\bf{x}}},y)}{\mathbb{I}}\{({{\bf{x}}},y)\in {{{\mathcal{A}}}}_{k}\}}.\)
Example 2 (EO)
The EO violation metric is given by
Then, \({F}_{{{\rm{EO}}}}=F({{\mathcal{A}}},\ell,{{\bf{w}}})\) if we change the grouping strategy \(\left\{{{{\mathcal{A}}}}_{k},k=1,\ldots,K\right\}\) to \(\left\{{{{\mathcal{A}}}}_{k}\cap \{ \, y=1\},k=1,\ldots,K\right\}\).
For smooth training, we propose a variance-regularization approach that directly constrains the max performance gap and allows for easy extension to incorporate other types of fairness considerations. The regularization is designed as:
Note that \(F({{\mathcal{A}}},\ell,{{\bf{w}}})\le \sqrt{{{\rm{Penalty}}}({{\boldsymbol{\theta }}},{{\bf{w}}};{{\mathcal{A}}})}\le \sqrt{K}F({{\mathcal{A}}},\ell,{{\bf{w}}}),\) which implies that constraining the variance-regularization is equivalent to constraining the max performance gap. To facilitate optimization and implementation, we use dice loss as \({R}_{k}({{\boldsymbol{\theta }}};{{{\mathcal{A}}}}_{k})\) in the segmentation task and cross entropy loss for the classification task. This approach bears a similarity to the VREx41 method. However, our method is not limited to task loss; \({R}_{k}({{\boldsymbol{\theta }}};{{{\mathcal{A}}}}_{k})\) can be adapted based on specific metrics. This flexibility allows our approach to be tailored to different fairness metrics and objectives.
Our method is specifically designed for the FL context, addressing the unique challenges of FL environments. By incorporating various fairness criteria while ensuring privacy, our weighted-variance-regularization approach provides a robust framework for achieving equitable outcomes. It allows decentralized data processing, preserves user privacy, and promotes equity, making it particularly crucial for applications in healthcare and other sensitive fields.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
In this study, we utilize the following datasets: Polyp dataset. The training and testing datasets, including Kvasir and CVC-300 are obtained from PraNet https://github.com/DengPingFan/PraNet. The training dataset is available at https://drive.google.com/file/d/1lODorfB33jbd-im-qrtUgWnZXxB94F55/view?usp=sharing, and the testing dataset is available at https://drive.google.com/file/d/1o8OfBvYE6K-EpDyvzsmMPndnUMwb540R/view?usp=sharing. Note that since our training only examined the goal of training with the specified dataset (site) and measuring fairness, test datasets other than these two datasets are not considered by us for participation in the training since they do not have a corresponding training set. Fundus dataset. The datasets include CHASE_DB1, DRIVE, and STARE, accessible at https://www.kaggle.com/datasets/khoongweihao/chasedb1, https://www.kaggle.com/datasets/andrewmvd/drive-digital-retinal-images-for-vessel-extraction, and https://paperswithcode.com/dataset/stare, respectively. Skin disease dataset. We use the HAM10000 and BCN20000 datasets, which have been filtered to exclude data lacking the required sensitive attributes. Metadata containing data information is provided in our code repository. The original image data can be obtained from https://www.kaggle.com/datasets/kmader/skin-cancer-mnist-ham10000and https://paperswithcode.com/dataset/bcn-20000. Private dataset. The dataset generated and analyzed in this study is currently stored on Google Drive and can be accessed via our code repository at https://github.com/MaksimXing/FlexFair. In line with Nature’s open-data principles and to ensure permanent linkage, we have deposited the dataset on Figshare (https://doi.org/10.6084/m9.figshare.28639598). The dataset will remain available for the foreseeable future. There are no restrictions on who may request access, and the data may be used for research purposes. Source data are provided with the paper as a single Excel file, in which we have consolidated all raw experimental results. Each sheet in the file corresponds to the raw data for a specific figure or table from the main manuscript and Supplementary Information. This file has been uploaded to our GitHub repository (release version available). Source data are provided with this paper.
Code availability
The code used in this study is available at the FlexFair GitHub repository43 and has been made citable with the https://doi.org/10.5281/zenodo.14926975. The plotted function, corresponding raw data, and dataset resources used in this study (including skin disease and private cervical cancer datasets) are also accessible via the repository. Notably, the code for training the polyp, cervical cancer, and fundus models is based on a modified version of SANet: https://github.com/weijun-arc/SANet. In addition, we provide pre-trained models to start training for these three datasets. For the dermatology dataset, we used the standard ResNet-50 for training, and running the code automatically downloads the official pre-trained model released by PyTorch: https://download.pytorch.org/models/resnet50-19c8e357.pth.
References
Yang, Y., Zhang, H., Gichoya, J. W., Katabi, D. & Ghassemi, M. The limits of fair medical imaging ai in real-world generalization. Nat. Med. 30, 2838–2848 (2024).
Jin, R. et al. Fairmedfm: fairness benchmarking for medical imaging foundation models. arXiv https://arxiv.org/abs/2407.00983 (2024).
McMahan, B., Moore, E., Ramage, D., Hampson, S. and y Arcas, B.A. Communication-efficient learning of deep networks from decentralized data. In: Artificial intelligence and statistics, 1273–1282 (PMLR, 2017).
Li, T. et al. Federated optimization in heterogeneous networks. Proc. Mach. Learn. Syst. 2, 429–450 (2020).
Li, Q., Diao, Y., Chen, Q. and He. B. Federated learning on non-iid data silos: an experimental study. In: 2022 IEEE 38th international conference on data engineering (ICDE), 965–978 (IEEE, 2022).
Karimireddy, S. P. et al. Scaffold: Stochastic controlled averaging for federated learning. In: International conference on machine learning, pages 5132–5143 (PMLR, 2020).
Zemel, R., Wu, Y., Swersky, K., Pitassi, T., Dwork, C. Learning fair representations. In: International conference on machine learning, 325–333 (PMLR, 2013).
Hardt, M., Price, E. & Srebro, N. Equality of opportunity in supervised learning. Adv. Neural Inf. Process. Syst. 29, 3315–3323 (2016).
Zafar, M. B., Valera, I., Rodriguez, M. G., Gummadi, K. P. and Weller, A. Fairness constraints: mechanisms for fair classification. In Artificial Intelligence and Statistics, 962–970 (PMLR, 2017).
Chuang, C.-Y., Liang, J., Torralba, A. and Jegelka, S. Fair mixup: better generalization for group fairness metrics. In: International Conference on Learning Representations (2021).
Agarwal, A., Beygelzimer, A., Dudik, M., Langford, J. and Wallach, H. M. Reductions for fair classification: a game-theoretic perspective. In: International Conference on Machine Learning, 60–69 (PMLR, 2018).
Cotter, A. et al. Training well-generalizing classifiers for fairness metrics and other data-dependent constraints. In International Conference on Machine Learning, 1394–1403 (PMLR, 2019).
Sun, R. et al. Fair-cda: continuous and directional augmentation for group fairness. In: Proceedings of the AAAI Conference on Artificial Intelligence, 37, 9918–9926 (2023).
Zeng, Y., Chen, H. and Lee, K. Improving fairness via federated learning. arXiv https://arxiv.org/abs/2110.15545 (2021).
Tran, C., Fioretto, F., Van Hentenryck, P. and Yao, Z. Decision making with differential privacy under a fairness lens. In IJCAI, 560–566 (2021).
Tayebi Arasteh, S. et al. Preserving fairness and diagnostic accuracy in private large-scale AI models for medical imaging. Commun. Med. 4, 46 (2024).
Cummings, R., Gupta, V., Kimpara, D. and Morgenstern, J. On the compatibility of privacy and fairness. In: Adjunct publication of the 27th conference on user modeling, adaptation and personalization, 309–315 (2019).
Ezzeldin, Y. H., Yan, S., He, C., Ferrara, E. and Avestimehr, A. S. Fairfed: enabling group fairness in federated learning. In Proceedings of the AAAI conference on artificial intelligence, 37, 7494–7502 (2023).
Castelnovo, A. et al. A clarification of the nuances in the fairness metrics landscape. Sci. Rep. 12, 4209 (2022).
Sung, H. et al. Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Pötter, R. et al. Mri-guided adaptive brachytherapy in locally advanced cervical cancer (embrace-i): a multicentre prospective cohort study. Lancet Oncol. 22, 538–547 (2021).
Eminowicz, G., Rompokos, V., Stacey, C. & McCormack, M. The dosimetric impact of target volume delineation variation for cervical cancer radiotherapy. Radiother. Oncol. 120, 493–499 (2016).
Petrič, P. et al. Uncertainties of target volume delineation in mri guided adaptive brachytherapy of cervix cancer: a multi-institutional study. Radiother. Oncol. 107, 6–12 (2013).
Hellebust, T. P. et al. Dosimetric impact of interobserver variability in mri-based delineation for cervical cancer brachytherapy. Radiother. Oncol. 107, 13–19 (2013).
Vázquez, D. et al. A benchmark for endoluminal scene segmentation of colonoscopy images. J. Healthc. Eng. 2017, 4037190 (2017).
Jha, D. et al. Kvasir-seg: A segmented polyp dataset. In: MultiMedia Modeling: 26th International Conference, MMM 2020, Daejeon, South Korea, January 5–8, 2020, Proceedings, Part II 26, 451–462. Springer (2020).
Tschandl, P., Rosendahl, C. & Kittler, H. The ham10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 1–9 (2018).
Combalia, M. et al. Bcn20000: dermoscopic lesions in the wild. arXiv https://arxiv.org/abs/1908.02288 (2019).
Fraz, M. M. et al. An ensemble classification-based approach applied to retinal blood vessel segmentation. IEEE Trans. Biomed. Eng. 59, 2538–2548 (2012).
Staal, J., Abramoff, M., Niemeijer, M., Viergever, M. & van Ginneken, B. Ridge-based vessel segmentation in color images of the retina. IEEE Trans. Med. Imaging 23, 501–509 (2004).
Hoover, A., Kouznetsova, V. & Goldbaum, M. Locating blood vessels in retinal images by piecewise threshold probing of a matched filter response. IEEE Trans. Med. imaging 19, 203–210 (2000).
Wei, J. et al. Shallow attention network for polyp segmentation. In MICCAI, 699–708 (Springer, 2021).
He, K., Zhang, X., Ren, S. and Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, 770–778 (2016).
Wang, J., Liu, Q., Liang, H., Joshi, G. & Poor, H. V. Tackling the objective inconsistency problem in heterogeneous federated optimization. Adv. Neural Inf. Process. Syst. 33, 7611–7623 (2020).
Donini, M. Oneto, L., Ben-David, S., Shawe-Taylor, J. S. & Pontil, M. Empirical risk minimization under fairness constraints. Advances in neural information processing systems, 31 (2018).
Kearns, M., Neel, S., Roth, A. and Wu, Z. S. Preventing fairness gerrymandering: auditing and learning for subgroup fairness. In International conference on machine learning, 2564–2572 (PMLR, 2018).
Hébert-Johnson, U., Kim, M., Reingold, O. & Rothblum, G. Multicalibration: calibration for the (computationally-identifiable) masses. In International Conference on Machine Learning, 1939–1948 (PMLR, 2018).
Williamson, R. & Menon, A. Fairness risk measures. In: International conference on machine learning, 6786–6797, PMLR (2019).
Monteiro Paes, L., Long, C., Ustun, B. & Calmon, F. On the epistemic limits of personalized prediction. Adv. Neural Inf. Process. Syst. 35, 1979–1991 (2022).
Hashimoto, T., Srivastava, M., Namkoong, H. & Liang, P. Fairness without demographics in repeated loss minimization. In: International Conference on Machine Learning, 1929–1938 (PMLR, 2018).
Krueger, D. et al. Out-of-distribution generalization via risk extrapolation (rex). In: Proceedings of the 38th International Conference on Machine Learning, 5815–5826 (PMLR, 2021).
Paes, L. M. Suresh, A. T., Beutel, A., Calmon, F. P. & Beirami, A. Multi-group fairness evaluation via conditional value-at-risk testing. arXiv https://arxiv.org/abs/2312.03867 (2023).
Xing, H. et al. Achieving flexible fairness metrics in federated medical imaging. Code repository for FlexFair. https://doi.org/10.5281/zenodo.14926975, (2025).
Acknowledgements
This work was supported by Shenzhen-Hong Kong Joint Funding No. SGDX20211123112401002 (awarded to Z.L.), the Basic Research Project No. HZQB-KCZYZ-2021067 of Hetao Shenzhen HK S&T Cooperation Zone (awarded to S.C.), NSFC with Grant No. 62293482 (awarded to S.C.), Shenzhen General Program No. JCYJ20220530143600001 (awarded to Z.L.), the Shenzhen Outstanding Talents Training Fund 202002 (awarded to S.C.), the Guangdong Research Project No.2017ZT07X152 and No. 2019CX01X104 (awarded to S.C.), the Guangdong Provincial Key Laboratory of Future Networks of Intelligence (Grant No. 2022B1212010001) (awarded to J.R., S.C.), the Guangdong Provincial Key Laboratory of BigData Computing CHUK-Shenzhen (awarded to Z.L.), the NSFC 61931024&12326610 (awarded to Z.L.), the Key Area R&D Program of Guangdong Province with grant No. 2018B030338001 (awarded to Z.L., S.C.), the Shenzhen Key Laboratory of Big Data and Artificial intelligence (Grant No. ZDSYS201707251409055) (awarded to Z.L., S.C.), National Nature Science Foundation of China (NSFC Grant No. 12201048) (awarded to C.X.), by China Association for Science and Technology Youth Care Program (awarded to Z.L.), and by Tencent & Huawei Open Fund (awarded to Z.L.).
Author information
Authors and Affiliations
Contributions
In this work, H.X. leads cervical cancer dataset processing, conducts extensive experiments, develops the federated learning framework, analyzes data, and assists with manuscript editing. R.S. contributes to algorithm design, guides algorithm optimization across varied settings establishes fairness evaluations and leads manuscript writing. J.R. helps with the design of the federated learning framework and co-writes parts of the manuscript. J.W. guides the algorithm implementation of segmentation scenarios and co-writes some parts of the manuscript. C.F. helps with private dataset processing and assists with manuscript preparation and figure design. X.D. (Xuan Ding) participates in private dataset processing and performs statistical experiments. Z.G. helps with some parts of the result analysis. Y.W., Y.H., W.W., X.B., Y.T. and X.L. manage multi-centre data collection and annotation to ensure dataset quality, with W.W. and X.B. providing clinical evaluations. C.X. offers theoretical insights and analysis of algorithms. S.C. contributes to the main research proposal and execution, designs rigorous experimental protocols, and guides the paper writing process to ensure high scholarly standards. X.H.D. (Xiaohui Duan) leads the collection of multi-center data and annotation and guides segmentation experiments. Finally, Z.L. guides the research direction, such as the main research and method design and rigorous experimental protocols design, and is in charge of the entire revision procedure.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Soroosh Tayebi Arasteh and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xing, H., Sun, R., Ren, J. et al. Achieving flexible fairness metrics in federated medical imaging. Nat Commun 16, 3342 (2025). https://doi.org/10.1038/s41467-025-58549-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58549-0
This article is cited by
-
Differential privacy for medical deep learning: methods, tradeoffs, and deployment implications
npj Digital Medicine (2026)
-
Medical support platform for melanoma analysis and detection based on federated learning
Scientific Reports (2026)
-
From pretraining to privacy: federated ultrasound foundation model with self-supervised learning
npj Digital Medicine (2025)
-
Connecting algorithmic fairness and fair outcomes in a sociotechnical simulation case study of AI-assisted healthcare
Nature Communications (2025)